DOI:10.32604/biocell.2021.013198

| BIOCELL DOI:10.32604/biocell.2021.013198 |  |
| Article |
Isolation, purification and structure elucidation of three new bioactive secondary metabolites from Streptomyces lividans AM
1Botany and Microbiology Department, Faculty of Science, Damanhour University, Damanhour, 22511, Egypt
2Chemistry Department, Faculty of Science, Mansoura University, El-Mansoura, 35516, Egypt
3Bioprocess Development Department, Genetic Engineering and Biotechnology Research Institute, City of Scientific Research and Technological Applications (SRTA-City), New Borg El-Arab City, Alexandria, 21934, Egypt
4Biology Department, College of Science, King Khalid University, Abha, 9004, Saudi Arabia
5Chemistry of Natural Compounds Department, Division of Pharmaceutical Industries, National Research Centre, Dokki-Cairo, 12622, Egypt
6Organic and Bioorganic Chemistry, Department of Chemistry, Bielefeld University, Bielefeld, 33615, Germany
*Address correspondence to: Mohammad El-Metwally, mmmyco@sci.dmu.edu.eg
Received: 29 July 2020; Accepted: 03 November 2020
Abstract: Microorganisms are a huge mine of bioactive metabolites, and actinomycetes are one of the very active groups in this area. In this article, we are concerned about the full taxonomical characterization of Streptomyces lividans AM, isolated from Egyptian soil. This isolate produced three new bioactive metabolites, namely: 1-Nona-decanoyl,4-oleyl disuccinate (1), filoboletic acid; (9Z,11E)-8,13-dihydroxy octadeca-9,11-dienoic acid (2), and sitosteryl-3β-D-glucoside (3). Extensive 1D and 2D NMR and HR-mass spectrometry were used to elucidate the structures of the three compounds. Moreover, ten known compounds were also identified. The antimicrobial activity of the producing organism and newly reported compounds (1–3) was investigated against a selected group of pathogenic microorganisms. A full taxonomical characterization of the strain was described as well.
Keywords: Bioactive metabolites; Streptomyces sp.; Taxonomy, Biological activity
Microbial worlds are unlimited resources of bioactive secondary metabolites. Secondary metabolites are crucial players in microbial development and interactions with other organisms. This mine of highly effective metabolites is still deep and needs more and more research. One of these good resources is Streptomyces sp., which is confirmed as one of the most productive sources for innovation of lead drugs (Blunt et al., 2003; Laatsch, 2010). Most Streptomyces species have the potential to synthesize several specific biologically active chemicals (Goodfellow and Haynes, 1984), which present a diversity of biological properties acting as antibiotics (Maiti et al., 2020; Ibrahim et al., 2019; Butler, 2004), antimycotics (Singh and Rai, 2013), antivirals (Raveh et al., 2013; Wei et al., 2014), anticancer (Davies-Bolorunduro et al., 2019), herbicides, pesticides, anti-parasitic, enzyme inhibitors (Bo et al., 2019; Olano et al., 2009), and pharmacologically active agents (Ahmad et al., 2017). However, many habitats of these Gram-positive bacteria are still unexplored as potential sources of new bioactive natural products (Baltz, 2007; Butler et al., 2013). In this article, three new compounds were isolated from full-identified Streptomyces lividans AM additional to ten known compounds. This article represents another good step to discover some elements of microbial, secondary compounds.
General experimental procedure
NMR spectra: 1D (1H NMR, 13C NMR, DEPT) and 2D (COSY, HMQC, and HMBC) NMR spectra were quantified on Bruker Avance DRX 500 and DRX 600 MHz spectrometers employing standard pulse sequences with reference to residual solvent signals. HR-EI-MS had been calculated using the GCT Premier spectrometer. The ultraviolet-visible (UV–Vis) spectrum was measured on Spectro UV–Vis Double Beam PC8 scanning auto Cell UVD-3200, LABOMED, INC. Column chromatography has been performed on the silica gel 60 (0.040–0.063 mm, Merck) and Sephadex LH-20 as the stationary phases. Preparative TLC (0.5 mm thick) and analytical TLC with pre-coated Merck silica gel 60 PF254+366. Rf values and analysis of chromatograms were performed under UV light (254 and 366 nm) and further by heating after spraying with anisaldehyde sulfuric acid reagent.
Isolation of Streptomyces sp. Isolate AM
The Streptomyces isolate AM has been isolated from a soil sample obtained from the governorate of Dakahlia, Egypt using starch nitrate medium (g/L) (Starch, 20; KNO3, 2; K2HPO4, 1; MgSO4.7H2O, 0.5; NaCl, 0.5; FeSO4.7H2O, 0.01; CaCO3, 3; agar, 20 in 1 L of distilled water (Waksman, 1959).
Characterization of Streptomyces sp. Isolate AM
Morphology and cultural properties
Substrate mycelium color, aerial mycelium colors, and diffusible pigment production were observed on ISP media as previously described (Shirling and Gottlieb, 1972). Spore chain morphology and the spore surface of the strain AM were examined after 14 days using Scanning Electron Microscope Jeol JSM-6360 LA.
The ability of the isolate AM to utilize different carbon sources was determined according to Shirling and Gottlieb (1966). Lecithinase activity was determined on egg–yolk plate medium (Nitsch and Kutzner, 1969). Hydrolysis of starch (Goodfellow and Orchard, 1974) and Liquefaction of gelatin (Waksman, 1961) were also investigated. The studied isolate AM was inoculated into skimmed milk medium (Iwasaki et al., 1981) and incubated at 30°C, then the degree of coagulation or liquefaction was recorded after 14 days of incubation. Melanoid production was determined after 4 days of incubation by using different media such as peptone-yeast extract-iron agar, tyrosine agar, and tryptone-yeast broth medium (Shirling and Gottlieb, 1966). The tested isolate was grown on modified Bennett's agar medium that supplemented separately with adenine (1%), hypothanthine (0.4%), allantoin (1%), and uric acid (1%). The degradation was determined after 7, 14, and 21 days of incubation (Jones, 1949). The clear zone formed around the tested isolate growth was recorded as a positive result. The ability of the isolate AM to reduce nitrates to nitrites was checked after 7 and 14 days as reported by Williams et al. (1983). Sodium chloride tolerance of the selected isolate AM was investigated according to Tresner et al. (1968). The capacity of the isolate AM to produce hydrogen sulfide was detected by inserting lead acetate paper strips into the neck of the culture tube containing peptone iron agar media. The formation of hydrogen sulfide was indicated by the blackening of lead acetate strips after 7 days of incubation (Küster and Williams, 1964). The ability of the tested isolate to decompose carboxymethyl cellulose (CMC) was quantified according to Prasad et al. (2013) using a sterilized Hutchinson liquid medium (Crawford and McCoy, 1972) supplemented with CMC 1% (w/v).
Sequence alignment and phylogenetic analysis
The partial 16S rRNA gene sequence of strain AM was aligned with the corresponding 16S rRNA sequences of the genus Streptomyces retrieved from the GenBank databases by using BLAST (www.ncbi.nlm.nih.gov/blast) (Altschul et al., 1997) and the software package MEGA 5 (Tamura et al., 2007) was used for multiple alignment and phylogenetic analysis. The phylogenetic tree was created via the Neighbor-joining method (Saitou and Nei, 1987).
Fermentation, working up and isolation
The spore suspension of S. lividans AM has been inoculated into 100 mL of ISP2 medium and cultivated at 28°C for 3 days. Five mL of seed culture was inoculated into 1L Erlenmeyer bearing modified rice-solid medium: Commercial rice 100 mg; 150 distilled water containing yeast extract 0.4% and malt extract 1% each flask under aseptic conditions. The culture was incubated for 14 days at 37°C. The obtained culture was collected and soaked in methanol, followed by filtration. After filtration, the water/methanol fraction was concentrated in vacuo. The remaining water residue was mixed with ethyl acetate. The ethyl acetate phase containing the microbial organic extract was finally separated and concentrated in vacuo till dryness delivering the desired reddish-brown crude extract.
The crude extract (3.2 g) was fractionated by silica gel column chromatography (column: 60 cm × 3 cm) eluting with a cyclohexane-DCM-MeOH gradient to afford four fractions on the basis of the analysis of TLC. Fraction I (0.91 g) was purified by silica gel CC, eluting with a cyclohexane-DCM gradient to give colorless oil of glycerol linoleate (25 mg), colorless semi-solid of linoleic acid (500 mg), and a colorless oil of 1-nona-decanoyl,4-oleyl disuccinate (1, 22 mg). Fraction II (0.51 g) was divided into sub-fractions IIa (0.22 g) and IIb (0.18 g) by silica gel CC, eluting with CH2Cl2-CH3OH. Purification of IIa by Sephadex LH-20 CC (CH2Cl2/40% MeOH) afforded filoboletic acid (2, 3 mg) as a colorless oil, and colorless solids from ferulic acid (4, 11 mg), indole-3-acetic acid methyl ester (2 mg), 4-hydroxy-phenyl acetic acid (6 mg). Purification of sub-fraction IIb by Sephadex LH-20 CC (DCM/MeOH [60:40]) resulted in 4-hydroxy-phenyl acetic acid (6 mg) as colorless solid. Fraction FIII was subjected to purification on silica gel eluted with DCM-MeOH followed by Sephadex LH-20 to afford two colorless solids of 3-(hydroxy-acetyl)-indole (3 mg), indol-3-carboxylic (2 mg). An application of the last fraction IV to Sephadex LH-20 (MeOH) led to colorless solids of p-hydroxybenzoic acid (3 mg), uracil (12 mg), and sitosteryl-3β-D-glucoside (3, 22 mg).
1-nona-decanoyl,4-oleyl disuccinate (1) C41H74O6 (662): Colorless oil, showing similar chromatographic properties to linoleic acid; Rf = 0.55 (DCM/MeOH [95:5]). For 1H NMR (300 MHz, CDCl3) and 13C NMR (125 MHz, CDCl3) data, see Tab. 3. (+) ESI MS m/z (%): 685 ([M+Na]+). (+) HR-ESI MS m/z: 685.53875 (calcd. 685.53774 for C41H74O6Na).
Filoboletic acid; (9 Z,11 E)-8,13-dihydroxy octadeca-9,11-dienoic acid (2) C18H32O4 (312): Colorless semi solid, UV faint absorbing, stained as dark violet and later as grey. Rf = 0.35 (CH2Cl2/7% CH3OH). The 1H (CDCl3, 300 MHz): δ = 6.18 (m, 1H), 6.05 (m, 1H), 5.61 (m, 2H), 4.10 (m, 2H), 2.39 (brm, 2H), 2.08 (br. m, 2H), 1.65 (brm, 2H), 1.30 (br. s, 12H), 0.91 (m, 3H). (+) ESI MS m/z (%): 335 [M+Na]). (-) ESI MS m/z (%): 311 [M-H]-). (+) HR-ESI MS m/z: 335.21942 (calcd. 335.21928 for C18H32O4Na). (-) HR-ESI MS m/z: 311.22178 (calc. 311.22168 for C18H31O4).
Sitosteryl-3 β-D-glucoside (3): C35H60O6 (576): Colorless solid, UV non absorbing or fluorescence, detected by anisaldehyde/sulfuric acid as dark brown, turned later to black. Rf = 0.25 (CH2Cl2/12% CH3OH). 1HNMR (DMSO-d6, 300 MHz) and 13C NMR (DMSO-d6, 125 MHz) data are listed in Tab. 4. (+) ESI MS: m/z (%): 599 [M+Na]+; (+) HR ESI MS: m/z (%): 599.4267 (calcd. 599.4282 for C35H60O6Na).
Ferulic acid; 3-(4-Hydroxy-3-methoxy-phenyl)-acrylic acid (4) C10H10O4 (194): Yellow solid, showing UV absorbance, and stained pink-violet on spraying with anisaldehyde/sulfuric, Rf = 0.26 (DCM/MeOH, 90:10). 1HNMR (300 MHz, CDCl3): δ = 7.19 (d, 2.0, 1H, H-5), 7.08 (dd, 8.2, 2.0, 1H, H-9), 6.83 (d, 8.1, 1H, H-8), 6.61 (d, 15.9 Hz, 1H, H-3), 6.33 (d, 15.9 Hz, 1H, H-2), 3.90 (s, 3H, OCH3-7). 13C NMR (125 MHz, CDCl3): δ = 169.5 (Cq-1), 149.9 (Cq-7), 147.9 (Cq-6), 145.5 (CH-3), 126.3 (Cq-4), 122.5 (CH-9), 115.0 (CH-8), 114.4 (CH-2), 110.2 (CH-5), 56.2 (7-OCH3). (+) ESI MS m/z (%): 195 ([M+H]+), 217 ([M+Na]+); (-) ESI MS m/z (%): 195 ([M-H]-).
Antimicrobial assay using agar diffusion test
Antimicrobial activity testing of the crude extract of Streptomyces lividans AM and the desired new compounds (1-3) were carried against a collection of microorganisms using the agar diffusion method. Paper-disk diffusion assay (Bayer et al., 1966) with some modifications has been followed to measure the antimicrobial activity. Twenty milliliters of medium seeded with test organism were poured into 9 cm sterile Petri dishes. After solidification, the paper disks were positioned on inoculated agar plates and allowed to distribute the loaded substances at 4°C for 2 h. The plates were incubated for 24 h at 35°C. Both bacteria and yeasts were grown on nutrient agar medium (g/l): Beef extract, 3; peptone, 10; and agar, 20. The pH was adjusted to 7.2. Fungal strain was grown on potato dextrose agar medium (g/l): Potato extract, 4; Dextrose, 20; Agar No. 1, 15 (pH 6). After incubation, the zones of inhibition were measured against the test microorganisms comprising: Gram-positive bacteria; (Bacillus cereus ATCC6633 and Staphylococcus aureus ATCC6538-P), Gram-negative bacteria (Pseudomonas areuginosa ATCC 27853), and yeast (Candida albicans ATCC 10231).
Characterization of the Streptomyces isolate
Morphology and culture properties of Streptomyces lividans AM
Morphological characteristics of the isolate AM were illustrated in Tab. 1. The isolate growth on starch nitrate media, ISP 2 and ISP 4 media was excellent, good on ISP 5 and ISP 7, and weak at ISP 3 and ISP 6. No diffusible pigment was observed in all tested media. Scanning electron microscope images of the isolate AM grown on starch nitrate agar medium showed that the spore chain is spiral where the spore surface ornamentation is smooth (Fig. 1).
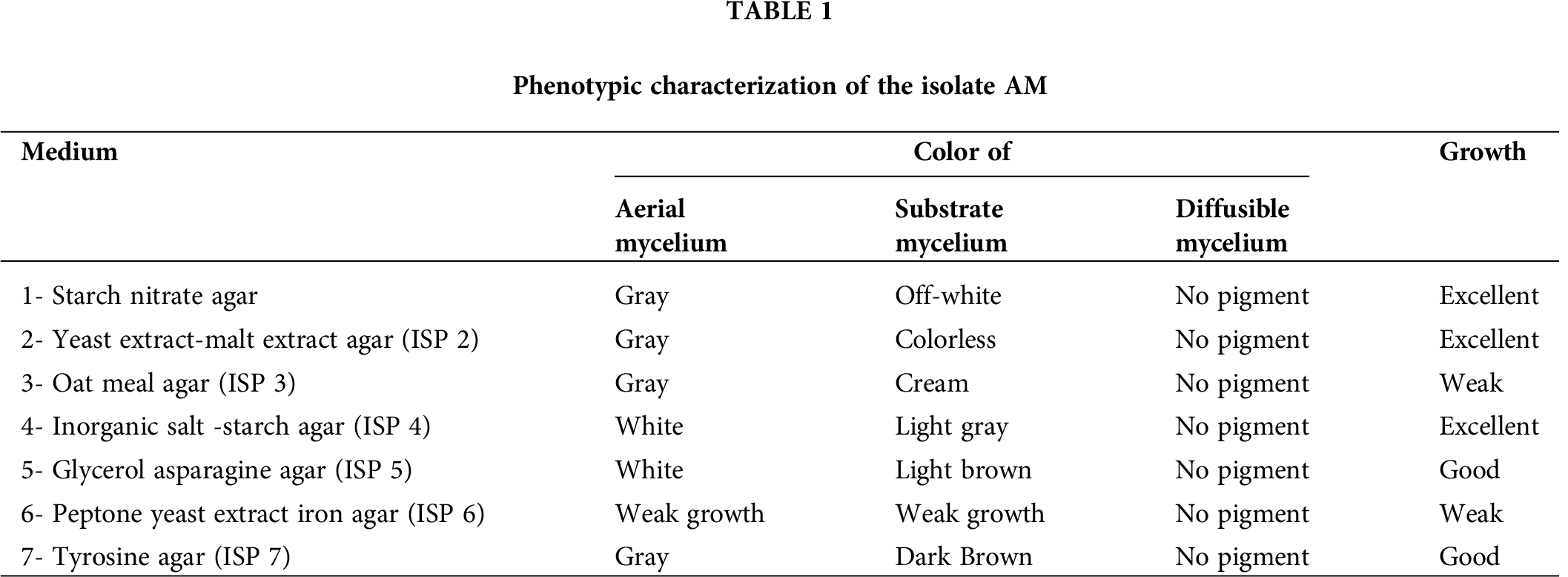

Figure 1: Scanning electron micrograph of the isolate AM grown on starch nitrate agar; (A) spore chain and (B) spore surface ornamentation.
Physiological and chemotaxonomic characteristics of Streptomyces lividans AM
The physiological and chemotypic characteristics of the isolate AM were illustrated in Tab. 2, where no melanoid pigment was formed in peptone-yeast extract iron agar or tyrosine agar. Isolate AM utilizes D-Glucose, L-arabinose, D-xylose, i-inositol, D-mannitol, D-fructose, rhamnose, sucrose, and raffinose for growth, but growth on sucrose or raffinose may be less than on the other carbon sources. It degrades adenine, uric acid, allantoin, hypoxanthine, and cellulose. Coagulation and peptonization of milk were positive, whereas lecithinase activity was negative (Fig. 2). Nitrate hydrolysis and H2S production were positive.


Figure 2: Negative lecithinase activity of Streptomyces lividans AM.
Molecular identification using 16S rRNA
The obtained 479 base pair sequence was analyzed using nucleotides BLAST, where it was compared with other rRNA genes that have been sequenced so far. A phylogenetic tree (Fig. 3) based on 16S rRNA gene sequences of members of the genus Streptomyces was constructed according to the neighbor-joining method using MEGA 5. The sequence analysis showed a close relationship to Streptomyces lividans strain TNAU1 (HQ897160.1) with a maximum identity of 97%. Moreover, the nucleotide sequence (479 base pairs) was deposited in the GenBank sequence database as Streptomyces lividans strain AM, and the accession number MF623053 was obtained.

Figure 3: Neighbor-joining tree based on 16S rRNA gene sequences, showing the phylogenetic relationship between strain AM and related species of the genus Streptomyces. GenBank sequence accession numbers are indicated in parentheses after the strain name.
Fermentation and structure elucidation
The Streptomyces lividans AM was fermented on rice-solid medium and the strain organic extract displayed potent activity versus Pseudomonas aeruginosa (30 mm), Candida albicans (25 mm), Bacillus cereus (22 mm), and moderate activity against Staphylococcus aureus (12 mm). In the chemical screening through TLC analysis, several bands of wide polarity were detected: (a) a group of non-UV-absorbing compounds, detected as intensive violet-blue bands by heating after spraying with anisaldehyde/sulfuric acid reagent; (b) UV absorbing bands, some of them showed pink, brown, violet, and/or orange colorization with anisaldehyde/sulfuric acid; (c) while the remaining UV-absorbing zones showed no color staining with the same reagent. Separation using a different series of chromatographic skills, namely silica gel and Sephadex LH-20 CC, afforded 1-nona-decanoyl, 4-oleyl disuccinate (1), filoboletic acid; (9Z,11E)-8,13-dihydroxy octadeca-9,11-dienoic acid (2), and sitosteryl-3β-D-glucoside (3). Further ten known compounds were afforded: ferulic acid (4) (Al-Refa, 2008; Hamed et al., 2017), glycerol monolinoleate (Naureen et al., 2015), linoleic acid (Mahmoud, 2005), indol-3-acetic acid methyl ester, 4-hydroxy-phenyl acetic acid (Mahmoud, 2005), 2-hydroxy-phenyl acetic acid (Mahmoud, 2005), 3-(hydroxy-acetyl)-indole (Mahmoud, 2005), indol-3-carboxylic (Mahmoud, 2005; Shaaban et al., 2002), p-hydroxybenzoic acid (Mahmoud, 2005), and uracil (Mahmoud, 2005).
1-Nona-decanoyl-,4-oleyl disuccinate
Compound 1 was obtained as a colorless oil with middle polarity, exhibiting a violet staining on spraying with anisaldehyde/sulfuric acid. The molecular weight of 1 was established by ESIMS as 662 Daltons with a corresponding molecular formula of C41H74O6, concluding the existence of five double bond equivalents (DBE). The 1H NMR spectrum exhibited a multiplet signal of 2H at δ 5.71 being for olefinic proton signals, in addition to two triplet signals being for two methylene groups at δ 3.62 and 3.19, which could be attached to sp2 systems, e.g., carbonyl of acid, ester or amide groups. Additional two triplet signals for further methylene groups were observed at δ 2.79 and 2.48 having a similar resonating pattern to those of the pervious triplet methylene groups. However, the lower chemical shifts of the last methylene groups establish their direct adjacent to sp2 systems, having a direct attachment (i.e., ethanediyl group) owing to the shown H-H COSY correlation between H2-2’(δ 2.79) and H2-3’(δ 2.48). Further, multiple signals were observed at δ 1.64 and 1.53 being for additional methylene groups. Finally, the spectrum exhibited a broad signal in the region of 1.39–1.34, being for multi methylene groups, terminated by triplet methyl groups (δ 0.90).
According to the 13C and HMQC spectral data (Tab. 3), compound 3 displayed forty-one carbon signals, classified into: four carbonyls appeared in the region of δ 173.6–173.2, two olefinic CH carbons at δ 129.9 and 127.8. The remaining carbons were visible in the region of δ 47.2–13.0, representing 35 sp3 carbon signals. Based on the HMBC and H-H COSY correlations (Fig. 4, Tab. 3), the two methylene groups at δ = 2.79 and 2.48 were confirmed to be flanked by two carbonyls of carboxylic acid esters visible at δ 173.2 and 173.6, deducing the existence of a succinate system. The lengths of two fatty acid chains were deduced, representing nonadecanoeate and oleate chains, one of them is bearing an olefinic double bond, attached to both sides of the succinate ester as the low chemical shifts of their esterified carbonyls (~173).
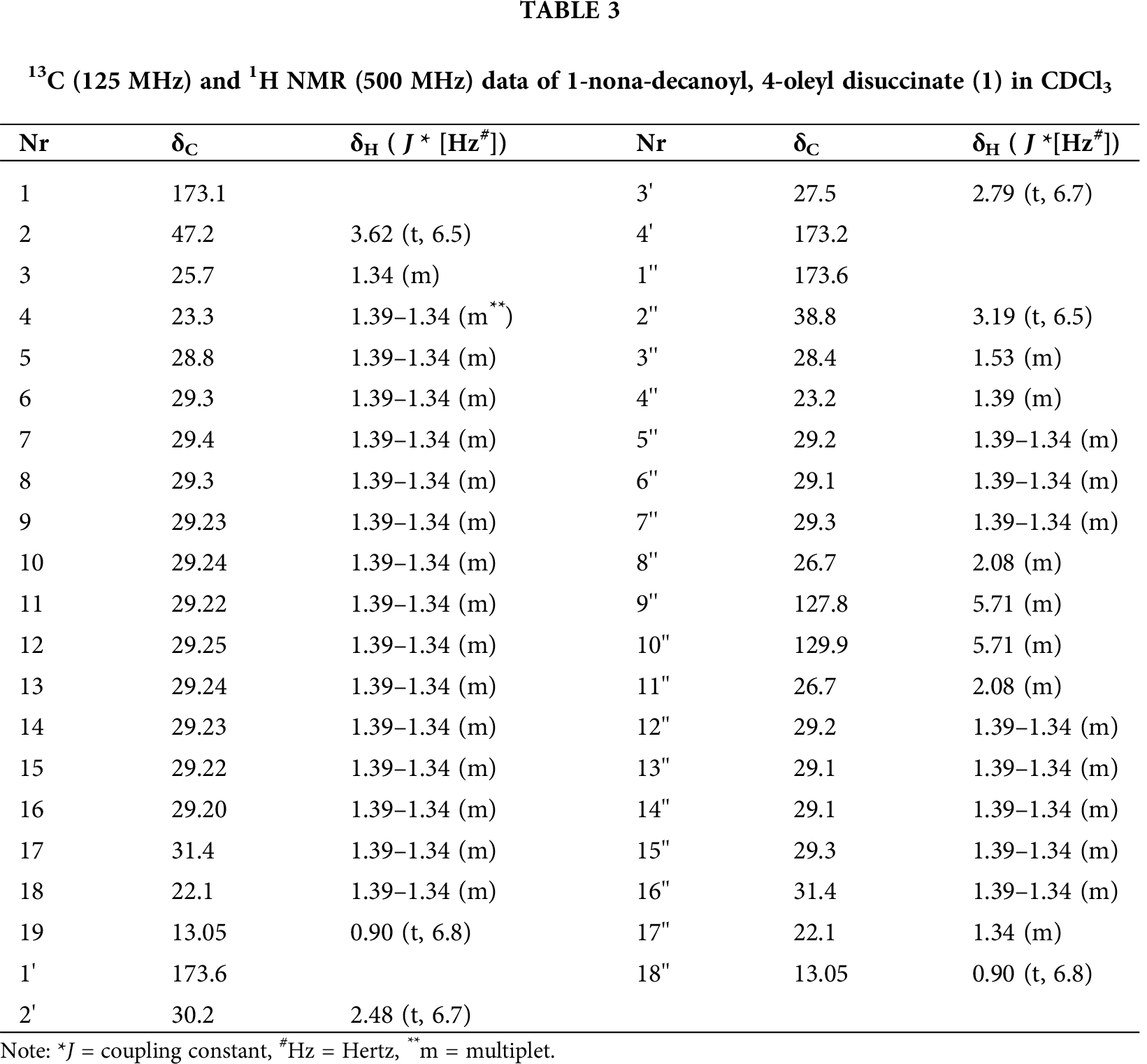

Figure 4: H-H COSY and HMBC connectivities of 1-nona-decanoyl-, 4-oleyl disuccinate (1).
Based on the presented molecular weight, corresponding molecular formula, and intensive study of the NMR data, this directed to structure 1 as 1-nona-decanoyl,4-oleyl disuccinate. It was not possible to fix the position of the olefinic double bond position as the high overlapping of the NMR signals in the structure. However, based on the biosynthetic pathways and the produced closely linoleic acid and glycerol monolinoleate by the same strain, the location of the olefinic bond at 9-position is the most plausible for structure 1. A search in the different databases (AntiBase, DNP and Scifinder) confirmed the natural novelty of 1. Only six structurally related compounds to 1 were reported in the literature: 1,4-didodecanoyl succinate;1,4-di-9-octadecanoyl succinate; 1,2,3-tricosanoyl-2-hydroxy-succinate; 1,4-di-hexadecanoly-2,3-bis(3-carbony-1-oxopropoxy) succinate (Scifinder). Esters of fatty acids, namely (R) and (S)-Glycerol-monolinoleate, were reported to exhibit inhibitory activities with IC50 values of 45.0 and 52.0 µM, respectively, against lipoprotein-associated phospholipase A2 [Lp-PLA2]; the latter is a specific marker of vascular inflammation associated with atherosclerosis (Lee et al., 2005).
As further middle polar colorless oil, compound 3 was isolated, exhibiting faint UV absorbance during TLC, which was detected as dark violet on spraying with anisaldehyde/sulfuric acid. The molecular weight of 3 was established to be 312 Dalton based on the exhibited molecular ion peaks shown at m/z 335 ([M+Na]+) and 311 ([M-H]−) in the ESI positive and negative modes, respectively. The corresponding molecular formula of 2 was deduced as C18H32O4 according to the HRSI MS as well.
The 1H NMR spectrum exhibited three multiple signals with integration of 4H located in the olefinic region at δ 6.18 (1H), 6.05 (1H), and 5.61 (2H). At δ 4.10, multiple signals with integration of 2H being for two hydroxy methines were visible. Two strong broad multiple signals were observed at δ 2.39, 2.08 corresponding to two 2H methylene protons attached mostly to sp2 carbons, being of the carboxylic acid and olefinic carbons, respectively. Three additional broad signals were observed at δ 1.65, 1.30, and 0.91, characteristics most likely of a long chain of methylene carbons ended by a terminal methyl group at δ 0.88. Based on these data and according to the search in AntiBase, filoboletic acid; (9Z,11E)-8,13-dihydroxy octadeca-9,11-dienoic acid was confirmed as the sole matching compound, which we report herein for the first time as a new bacterial metabolite. Filoboletic acid was previously reported as an antiviral compound produced by a fungus belonging to the genus Filoboletus (Simon et al., 1994). However, it is reported herein to first time from bacterial strains, and particularly from Streptomyces sp.
As polar colorless solid, compound 3 was obtained from fraction IV after purification with silica gel column followed by Sephadex LH-20. It showed no UV activity during TLC. However, it was detected as a dark brown when spraying with anisaldehyde/sulfuric acid. The molecular weight of 3 was established as 576 Dalton according to ESI MS, and the corresponding molecular formula was determined as C35H60O6 according to HRESI MS, containing 4 DBE. We report herein the full assignment of the structure (Fig. 5, Tab. 4) as it has not completely been assigned before failing 2D NMR assigning in several articles (Scifinder).


Figure 5: H-H COSY (–) and selected HMBC (H→C) correlations of sitosteryl-3β-D-glucoside (3).
It is worthy to mention that this is the first time to report β-sitosteryl D-glucoside from microorganisms. In contrast β-sitosteryl D-glucoside has been reported commonly from many diverse plant sources, e.g., the juice of Florida Valencia oranges (Citrus sinensis) (Ma and Schaffer, 1953), walnut (Juglans regia) (Jurd, 1956), Rhus trichocarpa (Yasue and Kato, 1957), H. longituba (Takemoto and Kusano, 1966), Zizyphus spina-christi (Aynehchi and Kiumehr, 1974; Aynehchi and Mahoodian, 1973), Euphorbia tinctoria (Aynehchi and Kiumehr, 1974). Oryza sativa (rice) hulls (Chung et al., 2005), represent one of the important sources of β-sitosteryl D-glucoside, and pre-germinated brown rice bran (Usuki et al., 2008). Therefore, it might be that β-sitosteryl D-glucoside has been isolated from the rice medium during the extraction process. However, an application of the same cultivating medium for the growth of fungal strains, followed by the same extracting procedure did not afford the same metabolite. This experiment was done several times confirming the capability of Streptomyces sp. rather than fungal strains to release the desired metabolite owing to their containing of rather different enzymes and genomes than those present in fungi, which have the efficiency to release/bio-transform such metabolite from the main producing media (rice media). In accordance, Streptomyces sp. have either high capability to release rather different enzymes with high efficiency to excrete compound 3, rather than those present in fungal strains or this compound is originally isolated from the rice medium, although this compound was not reported before from the rice bran before. Biologically, the use of sitosteryl β-D-glucoside as an emulsifier or preservative in food and feed was reported to exhibit no risk (Weber, 1988). Biologically, β-Sitosterol-3-O-β-d-glucopyranoside is selectively inhibited the activity of mammalian DNA polymerase λ (pol λ) in vitro (Mizushina et al., 2006).
Biologically, the organic extract of the bacterial strain exhibited potent activity against the Gram-negative bacteria Pseudomonas aeruginosa ATCC 27853 (30 mm), the yeast Candida albicans ATCC 10231 (25 mm), and the Gram-positive bacteria Bacillus cereus ATCC6633 (22 mm), while showed moderate activity against Staphylococcus aureus ATCC6538-P (12 mm). However, the newly reported compounds (1–3) displayed no antimicrobial activity against the referred set of pathogenic microorganisms.
Streptomyces lividans AM, which was isolated from an Egyptian soil sample, could produce three newly bioactive metabolites, namely 1-nona-decanoyl,4-oleyl disuccinate, filoboletic acid, (9Z,11E)-8,13-dihydroxy octadeca-9,11-dienoic acid, and sitosteryl-3β-D-glucoside. Further work needs to be done to evaluate the antiviral, antitumor, antifungal, antiparasitic, and many else of these secondary metabolites.
Acknowledgement: The authors thank Prof. N. Sewald in NMR and MS Departments, Bielefeld University for his support and Lab facilities during the carry out of the research work.
Availability of Data and Materials: All data generated or analyzed during this study are included in the manuscript.
Authors’ Contributions: The authors confirm contribution to the paper as follows: Study conception and design: Mohammad El-metwally and Mohamed Shaaban; data collection: Manal Elfedawy, Gaad Sohsah and Ahmed Rezk; analysis and interpretation of results: Mohammad El-metwally, Mamdouh Abdel-Mogib and Mohamed Shaaban; draft manuscript preparation: Mahmoud Moustafa, Mohammad El-metwally and Mohamed Shaaban. All authors reviewed the results and approved the final version of the manuscript.
Funding Statement: This work was supported by grant from the Deanship of Scientific Research at King Khalid University for Funding under Grant No. (R.G.P 2/90/41); German Academic Exchange Service (DAAD) Project-ID-57166072.
Conflicts of Interest: The authors declare that they have no conflicts of interest to report regarding the present study.
Ahmad SJ, Abdul Rahim MBH, Baharum SN, Baba MS, Zin NM (2017). Discovery of antimalarial drugs from streptomycetes metabolites using a metabolomic approach. Journal of Tropical Medicine 2017: 1–7. DOI 10.1155/2017/2189814. [Google Scholar] [CrossRef]
Al-Refa MHI (2008). New and bioactive secondary metabolites from ma-rine and terrestrial bacteria: Ramthacin A, B, C, and polyene macrolides from genetically modified bacteria. Göttingen: Niedersächsische Staats-und Universitätsbibliothek Göttingen. [Google Scholar]
Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997). Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Research 25: 3389–3402. DOI 10.1093/nar/25.17.3389. [Google Scholar] [CrossRef]
Aynehchi Y, Kiumehr N (1974). Chemical examination of Euphorbia tinctoria Boiss. Acta Pharmaceutica Suecica 11: 185. [Google Scholar]
Aynehchi Y, Mahoodian M (1973). Chemical examination of Zizyphus spina-christi (L.) Willd. Acta Pharmaceutica Suecica 10: 515. [Google Scholar]
Baltz RH (2007). Antimicrobials from actinomycetes: Back to the future. Microbe-American Society For Microbiology 2: 125. [Google Scholar]
Bayer A, Kirby W, Sherris J, Turck M (1966). Antibiotic susceptibility testing by a standardized single disc method. American Journal of Clinical Pathology 45: 493–496. DOI 10.1093/ajcp/45.4_ts.493. [Google Scholar] [CrossRef]
Blunt JW, Copp BR, Munro MH, Northcote PT, Prinsep MR (2003). Marine natural products. Natural Product Reports 20: 1–48. DOI 10.1039/b207130b. [Google Scholar] [CrossRef]
Bo AB, Kim JD, Kim YS, Sin HT, Kim HJ, Khaitov B, Ko YK, Park KW, Choi JS (2019). Isolation, identification and characterization of Streptomyces metabolites as a potential bioherbicide. PLoS One 14: e0222933. DOI 10.1371/journal.pone.0222933. [Google Scholar] [CrossRef]
Butler MS (2004). The role of natural product chemistry in drug discovery. Journal of Natural Products 67: 2141–2153. DOI 10.1021/np040106y. [Google Scholar] [CrossRef]
Butler MS, Blaskovich MA, Cooper MA (2013). Antibiotics in the clinical pipeline in 2013. Journal of Antibiotics 66: 571–591. DOI 10.1038/ja.2013.86. [Google Scholar] [CrossRef]
Chung IM, Hahn SJ, Ahmad A (2005). Confirmation of potential herbicidal agents in hulls of rice, Oryza sativa. Journal of Chemical Ecology 31: 1339–1352. DOI 10.1007/s10886-005-5290-5. [Google Scholar] [CrossRef]
Crawford DL, McCoy E (1972). Cellulases of Thermomonospora fusca and Streptomyces thermodiastaticus. Applied Microbiology 24: 150–152. DOI 10.1128/AM.24.1.150-152.1972. [Google Scholar] [CrossRef]
Davies-Bolorunduro OF, Adeleye IA, Akinleye MO, Wang PG (2019). Anticancer potential of metabolic compounds from marine actinomycetes isolated from Lagos Lagoon sediment. Journal of Pharmaceutical Analysis 9: 201–208. DOI 10.1016/j.jpha.2019.03.004. [Google Scholar] [CrossRef]
Goodfellow M, Haynes J (1984). Actinomycetes in marine sediments. In: Ortiz-Ortiz L, Bojalil LF, Yakoleff V (eds.Biological, biochemical and biomedical aspects of actinomycetes, pp. 453–472. London: Academic Press. [Google Scholar]
Goodfellow M, Orchard VA (1974). Antibiotic sensitivity of some nocardioform bacteria and its value as a criterion for taxonomy. Microbiology 83: 375–387. [Google Scholar]
Hamed A, Abdel-Razek AS, Frese M, Wibberg D, El-Haddad AF, Ibrahim TM, Kalinowski J, Sewald N, Shaaban M (2017). New oxaphenalene derivative from marine-derived Streptomyces griseorubens sp. ASMR4. Zeitschrift für Naturforschung B 72: 53–62. DOI 10.1515/znb-2016-0145. [Google Scholar] [CrossRef]
Ibrahim AA, El-Housseiny GS, Aboshanab KM, Yassien MA, Hassouna NA (2019). Paromomycin production from Streptomyces rimosus NRRL 2455: Statistical optimization and new synergistic antibiotic combinations against multidrug resistant pathogens. BMC Microbiology 19: 18. DOI 10.1186/s12866-019-1390-1. [Google Scholar] [CrossRef]
Iwasaki A, Itoh H, Mori T (1981). Streptomyces sannanensis sp. nov. International Journal of Systematic and Evolutionary Microbiology 31: 280–284. [Google Scholar]
Jones KL (1949). Fresh isolates of actinomycetes in which the presence of sporogenous aerial mycelia is a fluctuating characteristic. Journal of Bacteriology 57: 141–145. DOI 10.1128/JB.57.2.141-145.1949. [Google Scholar] [CrossRef]
Jurd L (1956). The sterol and carbohydrate constituents of the walnut (Juglans regia). Journal of Organic Chemistry 21: 759–760. DOI 10.1021/jo01113a010. [Google Scholar] [CrossRef]
Küster E, Williams S (1964). Production of hydrogen sulfide by streptomycetes and methods for its detection. Applied Microbiology 12: 46–52. DOI 10.1128/AM.12.1.46-52.1964. [Google Scholar] [CrossRef]
Laatsch H (2010). A data base for rapid structural determination of microbial natural products, and annual updates. http://www.user.gwdg.de/~ucoc/laatsch/AntiBase.htm. [Google Scholar]
Lee WS, Kim MJ, Beck YI, Park YD, Jeong TS (2005). Lp-PLA2 inhibitory activities of fatty acid glycerols isolated from Saururus chinensis roots. Bioorganic & Medicinal Chemistry Letters 15: 3573–3575. DOI 10.1016/j.bmcl.2005.05.056. [Google Scholar] [CrossRef]
Ma RM, Schaffer P (1953). β-Sitosteryl D-glucoside and β-sitosterol from commercially dried grapefruit pulp. Archives of Biochemistry and Biophysics 47: 419–423. DOI 10.1016/0003-9861(53)90/478-0. [Google Scholar] [CrossRef]
Mahmoud MAS (2005). Bioactive secondary metabolites from marine and terrestrial bacteria: Isoquinolinequinones, bacterial compounds with a novel pharmacophor (Ph.D. Thesis). Georg-August Universität, Göttingen, Germany. [Google Scholar]
Maiti PK, Das S, Sahoo P, Mandal S (2020). Streptomyces sp SM01 isolated from Indian soil produces a novel antibiotic picolinamycin effective against multi drug resistant bacterial strains. Scientific Reports 10: 1–12. [Google Scholar]
Mizushina Y, Nakanishi R, Kuriyama I, Kamiya K, Satake T, Shimazaki N, Koiwai O, Uchiyama Y, Yonezawa Y, Takemura M (2006). β-Sitosterol-3-O-β-D-glucopyranoside: A eukaryotic DNA polymerase λ inhibitor. Journal of Steroid Biochemistry and Molecular Biology 99: 100–107. DOI 10.1016/j.jsbmb.2005.12.007. [Google Scholar] [CrossRef]
Naureen H, Asker M, Shaaban M (2015). Structural elucidation and bioactivity studies of secondary metabolites from endophytic Aspergillus niger. Indian Journal of Applied Research 5: 76–83. [Google Scholar]
Nitsch B, Kutzner H (1969). Egg-yolk agar as a diagnostic medium for streptomycetes. Experientia 25: 220–221. DOI 10.1007/BF01899136. [Google Scholar] [CrossRef]
Olano C, Méndez C, Salas JA (2009). Antitumor compounds from marine actinomycetes. Marine Drugs 7: 210–248. DOI 10.3390/md7020210. [Google Scholar] [CrossRef]
Prasad P, Singh T, Bedi S (2013). Characterization of the cellulolytic enzyme produced by Streptomyces griseorubens (Accession No. AB184139) isolated from Indian soil. Journal of King Saud University–Science 25: 245–250. DOI 10.1016/j.jksus.2013.03.003. [Google Scholar] [CrossRef]
Raveh A, Delekta PC, Dobry CJ, Peng W, Schultz PJ, Blakely PK, Tai AW, Matainaho T, Irani DN, Sherman DH (2013). Discovery of potent broad spectrum antivirals derived from marine actinobacteria. PLoS One 8: e82318. DOI 10.1371/journal.pone.0082318. [Google Scholar] [CrossRef]
Saitou N, Nei M (1987). The neighbor-joining method: A new method for reconstructing phylogenetic trees. Molecular Biology and Evolution 4: 406–425. [Google Scholar]
Shaaban M, Maskey RP, Wagner-Döbler I, Laatsch H (2002). Pharacine, a natural p-cyclophane and other indole derivatives from Cytophaga sp. strain AM13. 1. Journal of Natural Products 65: 1660–1663. DOI 10.1021/np020019a. [Google Scholar] [CrossRef]
Shirling EB, Gottlieb D (1972). Cooperative description of type strains of Streptomyces: V. Additional Descriptions1. International Journal of Systematic and Evolutionary Microbiology 22: 265–394. [Google Scholar]
Shirling ET, Gottlieb D (1966). Methods for characterization of Streptomyces species. International Journal of Systematic Bacteriology 16: 313–340. DOI 10.1099/00207713-16-3-313. [Google Scholar] [CrossRef]
Simon B, Anke T, Sterner O (1994). Hydroxylated unsaturated fatty acid from cultures of a Filoboletus species. Phytochemistry 36: 815–816. DOI 10.1016/S0031-9422(00)89826-X. [Google Scholar] [CrossRef]
Singh N, Rai V (2013). In vitro antimycotic activity of a new isolate Streptomyces fradiae MTCC 11051 against the multi-drug resistant pathogenic fungi. Journal of Pharmacy Research 7: 331–336. DOI 10.1016/j.jopr.2013.04.024. [Google Scholar] [CrossRef]
Takemoto T, Kusano G (1966). Studies on the constituents of Hemerocallis. I. Constituents of Hemerocallis longituba. Yakugaku zasshi: Journal of the Pharmaceutical Society of Japan 86: 1116. DOI 10.1248/yakushi1947.86.11_1116. [Google Scholar] [CrossRef]
Tamura K, Dudley J, Nei M, Kumar S (2007). MEGA4: Molecular evolutionary genetics analysis (MEGA) software version 4.0. Molecular Biology and Evolution 24: 1596–1599. DOI 10.1093/molbev/msm092. [Google Scholar] [CrossRef]
Tresner H, Hayes JA, Backus E (1968). Differential tolerance of streptomycetes to sodium chloride as a taxonomic aid. Applied Microbiology 16: 1134–1136. DOI 10.1128/AM.16.8.1134-1136.1968. [Google Scholar] [CrossRef]
Usuki S, Ariga T, Dasgupta S, Kasama T, Morikawa K, Nonaka S, Okuhara Y, Kise M, Robert KY (2008). Structural analysis of novel bioactive acylated steryl glucosides in pre-germinated brown rice bran. Journal of Lipid Research 49: 2188–2196. DOI 10.1194/jlr.M800257-JLR200. [Google Scholar] [CrossRef]
Waksman SA (1961). The Actinomycetes. Vol. II. Classification, identification and descriptions of genera and species. The Actinomycetes, vol. 2. DOI 10.5694/j.1326-5377.1962.tb67171.x. [Google Scholar] [CrossRef]
Waksman SA (1959). Strain specificity and production of antibiotic substances. X. characterization and classification of species within the Streptomyces griseus group. Proceedings of the National Academy of Sciences of the United States of America 45: 1043. [Google Scholar]
Weber N (1988). Metabolism of sitosteryl β-D-glucoside and its nutritional effects in rats. Lipids 23: 42–47. DOI 10.1007/BF02535303. [Google Scholar] [CrossRef]
Wei Y, Fang W, Wan Z, Wang K, Yang Q, Cai X, Shi L, Yang Z (2014). Antiviral effects against EV71 of pimprinine and its derivatives isolated from Streptomyces sp. Virology Journal 11: 195. DOI 10.1186/s12985-014-0195-y. [Google Scholar] [CrossRef]
Williams S, Goodfellow M, Alderson G, Wellington E, Sneath P, Sackin M (1983). Numerical classification of Streptomyces and related genera. Microbiology 129: 1743–1813. DOI 10.1099/00221287-129-6-1743. [Google Scholar] [CrossRef]
Yasue M, Kato Y (1957). Components of wood of Rhus trichocarpa. Yakugaku Zasshi (Journal of the Pharmaceutical Society of Japan) 77: 1045–1047. DOI 10.1248/yakushi1947.77.9_1045. [Google Scholar] [CrossRef]
 | This work is licensed under a Creative Commons Attribution 4.0 International License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. |