DOI:10.32604/cmc.2020.012976

| Computers, Materials & Continua DOI:10.32604/cmc.2020.012976 |  |
| Article |
Potential Inhibitory Effect of Vitamins Against COVID-19
1Department of Clinical Laboratory Sciences, College of Applied Medical Sciences, Jouf University, Sakaka, Al Jouf, Saudi Arabia
2College of Pharmacy, Jouf University, Sakaka, Al Jouf, Saudi Arabia
3Department of Microbiology and Molecular Genetics, The Women’s University, Multan, Pakistan
4Department of Computer Sciences, Kinnaird College for Women, Lahore, Pakistan
5Department of Microbiology and Molecular Genetics, University of the Punjab, Lahore, Pakistan
*Corresponding Author: Kashaf Junaid. Email: kjunaid@ju.edu.sa; kashaf_junaid@hotmail.com
Received: 20 July 2020; Accepted: 11 September 2020
Abstract: Coronavirus disease 2019 (COVID-19) is a current pandemic that has affected more than 195 countries worldwide. In this severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) pandemic, when treatment strategies are not yet clear and vaccines are not available, vitamins are an excellent choice to protect against this viral infection. The rationale behind this study was to examine the inhibitory effect of vitamins B, C, and D against the main protease of SARS-CoV-2 and angiotensin-converting enzyme 2 (ACE2), which have critical rolesin the immune system. Molecular docking, performed by using MOE-Dock of the Chemical Computing Group, was used to understand the mechanism. The vitamins all docked within the active sites of the Mpro (PDB ID:6LU7) and ACE2 receptor proteins (PDB ID:6VW1). Vitamins B and C delivered maximum energy scores against both targets, while vitamin D displayed a binding energy score of −7.9532 kcal/mol for Mpro and −7.9297 for ACE2. The efficiency of all three vitamins is higher than the binding energy score of chloroquine (−6.889 kcal/mol), which is now under clinical trials. The use of vitamins is beneficial, being immune system restorative, and they also act as anti-COVID agents. Although the potential beneficial effects of vitamin B and C are revealed through docking studies, further clinical trials are required for the validation of these results.
Keywords: SARS-CoV-2; vitamins; docking; antivirals; chemical computing
Evolution results from persistent and dynamic interactions between a host and pathogens, such thatboth engage in a continual efforts to suppress one another. Genetic transformations in pathogens and natural selection make pathogens more vulnerable, as well as enable them to escape host immune responses. The current COVID-19 pandemic is example of this process [1]. Following the report of the first case of novel coronavirus (COVID-19) was reported in December 2019 in Wuhan, China, it was declared an international public health emergency in January 2020, causing more than 211 K deaths worldwide. According to the World Health Organization (WHO), the mortality rate was 3.4% from December 2019 to April 2020 [2].
Coronaviruses (CoVs) are enveloped viruses with non-segmented, single-stranded, and positive-sense RNA genomes. Of the seven strains demonstrated known to infect humans, SARS-CoV, MERS-CoV, and SARS-CoV-2 are zoonotic viruses that cause severe lower respiratory tract dysfunction. SARS-CoV-2 shows 97% similarity with bat coronaviruses [1]. Despite the spread of COVID-19 across all continents, there is no definite mortality rate yet (1–6%); however, this is less than MERS (~40%) and SARS (~10%). The mortality rate of COVID-19 is due to a lack of approved drugs and its high infectivity rate (Ro 1.4–4.0). The only available defense against COVID-19 is strong immunity. Therefore, developing a potential drug is imperative [3,4].
SARS-CoV-2 encodes a variety of structural, non-structural, and accessory proteins, such as RNA-dependent RNA polymerase (RdRp), 3-chymotrypsin-like (3CL) protease, papain-like protease, helicase, and the spike glycoprotein [5]. Several compounds have been reported that target these proteins and inhibit viral pathogenesis. The cellular receptor of SARS-CoV-2 is angiotensin-converting enzyme 2 (ACE2), an enzyme that is primarily involved in angiotensin (Ang) maturation, which plays a role in vasoconstriction and blood pressure. ACE2 has a peptidase domain at the N-terminal and a collectrin-like domain (CLD) at the C-terminal. The receptor-binding domain (RBD) is recognized by polar residues of the extracellular peptidase domain of ACE2 [6]. The spike glycoproteins on enveloped CoVs aid viral entry into the host cells by binding to host cell receptors via the S1 subunit of the spike protein and later by fusion of the viral and host cell membranes by the S2 subunit. After binding of S1 to the host receptor (ACE2), a new cleavage site on S2 is exposed, which the host protease then cleaves. This step is crucial in the development of infection. The RBD is on S1, which binds with the peptidase domain of human ACE2 receptors. Since the RBD is directly involved in the binding of the virus to host cells, targeting the RBD is a suitable choice for the development of therapeutic agents [2,6].
In the race to develop a vaccine for coronavirus, efforts have focused on diagnosis to identify infected persons and enable their isolation and treatment [7]. Currently, screening of a large number of compounds as potential treatment agents for coronavirus is under way, among which remdesivir is undergoing clinical trials in the USA and China. This drug has previously been used as an inhibitor against several other RNA viruses, including HIV [5]. Likewise, chloroquine (CQ) and hydroxychloroquine (HCQ) sulfate are also under consideration. These immunomodulators have already been in use for the treatment of malaria and other autoimmune disorders. Moreover, these drugs have also shown some promising results against the Ebola virus [3]. Virtual screening through molecular docking increases the success rate of drug discovery [5]. In addition to molecular protein modeling, various bioinformatic tools can provide useful insights allowing the visualization of possible variations in protein structures and complexes.
The study described in this paper aimed to explore the binding of vitamins to viral receptors. The SARS-CoV-2 RBD is directly involved in the binding of the virus to host cells, and hence targeting the RBD may be a soundchoice for the development of therapeutic agents against COVID-19 [2,6]. Therefore, the RBD and ACE-2 were targeted via molecular docking to screen the chemical structures of vitamins B, C, and D as potential inhibitors of this newly emerged coronavirus infection.
The molecular sequences of the main protease of SARS-CoV-2 (PDB ID:6LU7) and the ACE2 receptor (PDB ID:6VWI) were downloaded from the protein database (PDB database).
The 3D structures of the selected ligands, Vitamins B, C, and D, were drawn using the Builder tool incorporated in MOE-Dock from the Chemical Computing Group. The energy level was minimized for each ligand.
For structure-based virtual screening, vitamins B, C, and D were docked with the SARS-CoV-2 protease (PDB ID:6LU7) and PD-ACE2 (PDB ID:6VW1) using MOE-Dock. The binding of different compounds at the receptor site generated binding energies D-G (Gibbs energy), expressed as docking scores in potential energy maps.
During docking, the ligands were regarded asflexible, while the protein was regarded as rigid. The site finder tool incorporated in MOE was used to find the active center of the target proteins. The dummy atoms resulting from alpha spheres with a backbone were generated, and residues were kept fixed. Later, the energy level was minimized. This application also determines the amino acids present at the active sites of proteins that interact with ligands. Results of less than 1.0 Å in positional root mean square deviation (RMSD) were considered ideal and were clustered to determine the most favorable binding conformation. The highest binding energy (most negative) was considered as the ligand with maximum binding affinity.
All the selected compounds docked in the target protein’s active site, and ten different conformations were selected for each ligand. The resulting ligand-protein complex was selected based on energy scores. These ligand-protein complexes were then used to calculate energy parameters through MMFF94x force field energy calculations and predict ligand-protein interactions at the active site.
The ongoing COVID-19 pandemic is disturbing, since this new virus is highly contagious. Since it first appeared in Wuhan, this virus has infected thousands of people in China and has subsequently spread to almost every country world-wide. The highest mortality rates for COVID-19 have been reported in the USA and Italy [8]. The results of the present study could fill the existing gaps in the treatment of this ongoing viral infection. The discovery of a potent drug against COVID-19 is desperately needed, but this will be a time-consuming process. In the current situation, there is a need to look for other alternative treatments with potent antiviral effects, but minimal side effects.
The bioinformatics modeling system known as molecular docking plays an important role in screening novel compounds that can potentially help fight this dangerous virus. Docking is based on the interaction of small molecules, known as ligands or inhibitors, with the target protein’s active sites. This non-covalent binding process predicts the binding affinity between the two molecules. With a low docking score, an active compound is considered an excellent compound to inhibit the target protein.
In this study, two important sites on SARS-CoV-2 were targeted. The first was the main protease (Mpro) of SARS-COV-2, which contains 312 amino acids and has a molecular weight of 34 kDa. The second was the spike protein ACE2 receptor, with a molecular weight of 193 kDa. Based on the binding energy scores, it was found that vitamin C is the most potent inhibitor of the three tested vitamins, with a binding energy score of −11.3518 (Tab. 1). Details of the binding and interaction are given in Tab. 2.
Table 1: Molecular docking analysis of vitamin C, B, and D against the major protease and ACE2 receptor of COVID-19
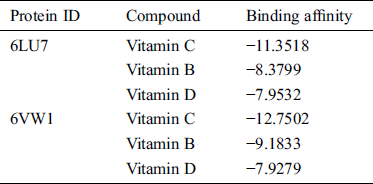
Table 2: List of H bonds between the ligand and the target protein
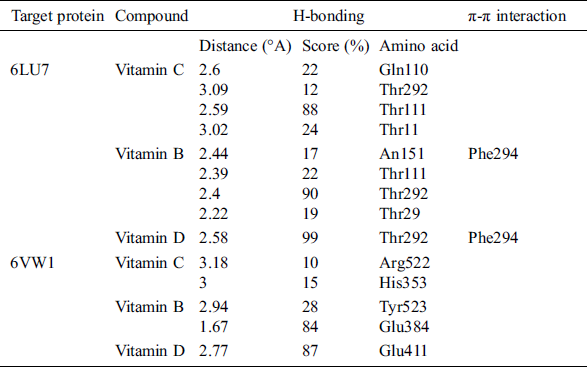
Vitamin C exhibited binding affinity with Gln110, Thr292, and Thr111 (−11.3518 kcal/mol). These amino acid residues are located in the binding pocket of 6LU7 (Fig. 1). Recently, viricidal activities of vitamin C have been reported against retroviruses and the influenza virus [9,10,11]. The proposed antiviral mechanism is based on the production of H2O2 and other free radical species [11,12]. Moreover, vitamin C also showed binding affinity with the ACE2 receptor protein (6VW1) (−12.7502 kcal/mol). Vitamin C interacts with Arg522, Glu411, His387, His383, and His353 (Fig. 2). One of the most important interactions between vitamin C and the ACE2 receptor within the zinc-binding domain is interesting, as all the previously reported inhibitors of ACE2 bind with the zinc ion located in the active site of ACE2.
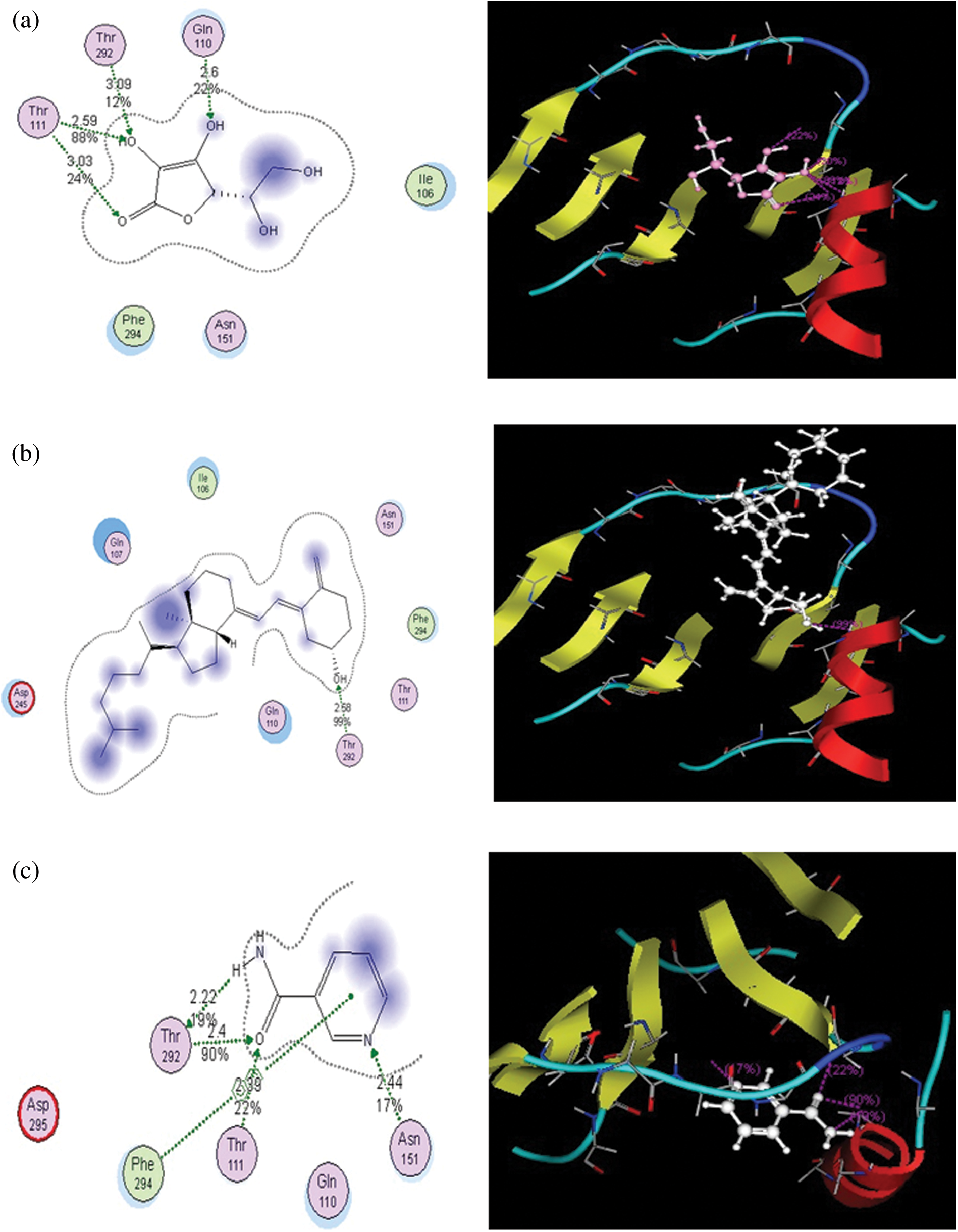
Figure 1: Putative binding modes of vitamins C (a), D (b), and B (c) in the binding pocket of the main protease. Metal or ion contact is depicted via a purple dotted line, and side-chain proton acceptor/donors via a green dotted line The amino acids circled in blue and red represent basic and acidic amino acids. The blue background behind certain amino acids denotes their solvent exposure. Solvent exposure is also depicted via the blue shading behind certain ligand atoms
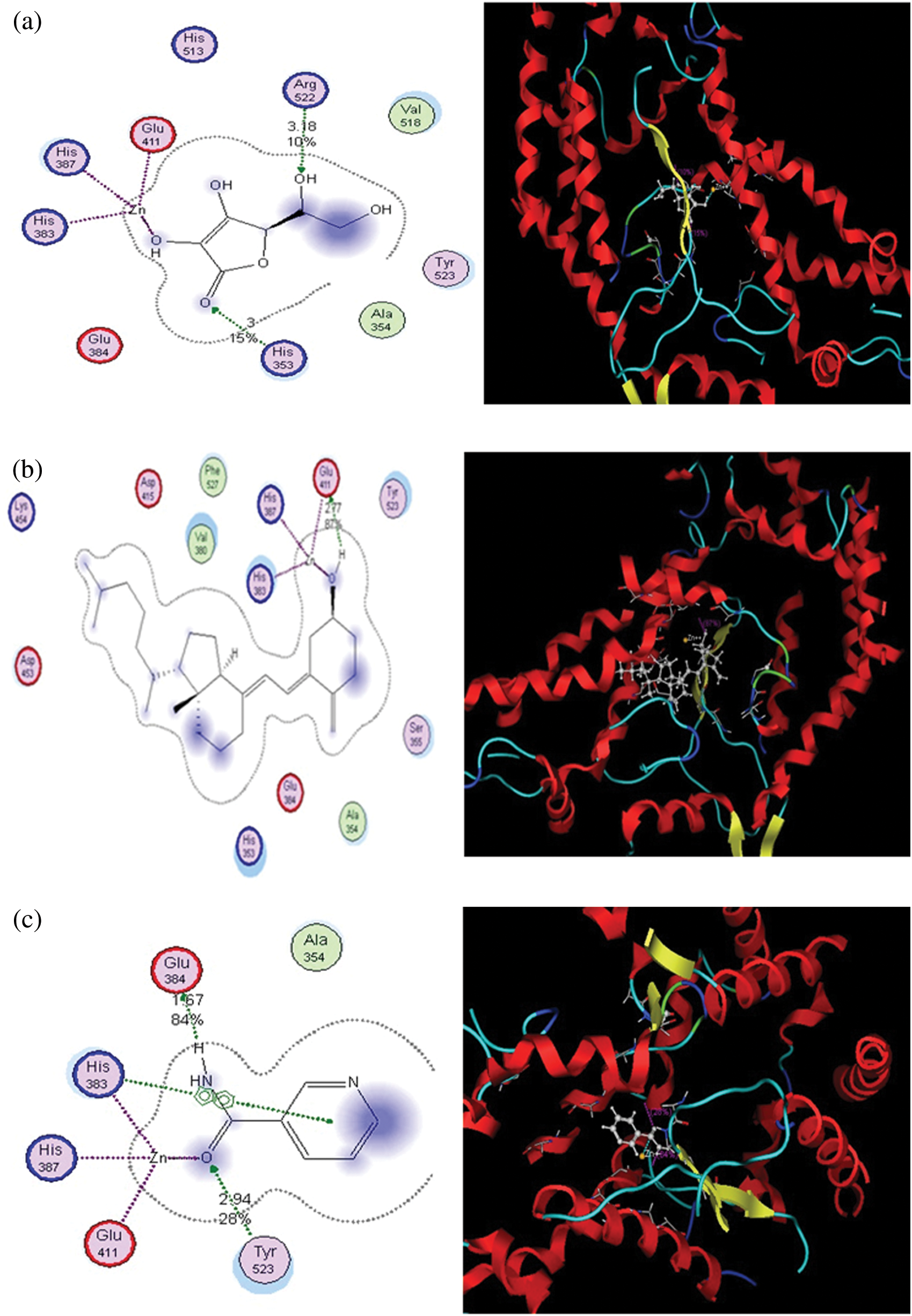
Figure 2: Putative binding modes of vitamins C (a), D (b), and B (c) in the binding pocket of ACE2. Metal or ion contact is depicted via a purple dotted line, and side-chain proton acceptor/donors via a green dotted line. The amino acids circled in blue and red represent basic and acidic amino acids. The blue background behind certain amino acids denotes their solvent exposure. Solvent exposure is also depicted via the blue shading behind certain ligand atoms
Different studies recommend vitamin C as a potent immunomodulator because it enhances interferon production, which helps fight against viral infection. Further, it also downregulates pro-inflammatory cytokines, which could prevent the development of a “cytokine storm”, the main contributor to the mortality of COVID-19 [11]. Vitamin C is an essential component of the cellular antioxidant defense system, and thus offers a good option in the critical care of COVID-19 patients. High doses of this vitamin could minimize oxidative stress in patients and ultimately prevent the progression of the disease to an acute respiratory distress syndrome [13]. In China, a clinical trial of a high-dose intervention with vitamin C administered to SARS-CoV-2 infected individuals showed that all the patients had a better oxygenation index, and surprisingly, all the patients who took vitamin C recovered [13]. Based on these findings, it seems plausible that vitamin C has the potential to fight this the newly emerged viral infection.
Likewise, vitamin B contains several active moieties against Mpro and the ACE2 receptor protein of the virus. This vitamin interacts with Asn141, Thr111, Thr292, and Phe 294 residues of the main protease (Fig. 1). The binding energy score for vitamin B is −8.3799 kcal/mol. Vitamin B also interacts with Glu384, His383, His387, Glu411, and Tyr523 of the ACE2 receptor protein (Fig. 2), with an excellent binding score of −9.1833 kcal/mol. Vitamin B was used in the treatment of MERS-CoV, and a significant decrease in the viral load was obtained [14]. Vitamin B has already been reported as being useful in many viral infections, including influenza, HIV, and MERS [15]. Based on these docking results, it is possible that apart from inhibiting the main protease of SARS-CoV-2, this vitamin also has the potential to block the S protein of the virus.
Several epidemiological and experimental studies have highlighted the relationship between vitamin D and pulmonary diseases. Vitamin D has a strong effect in inflammatory cells [16]. Evidence exists to show that this vitamin stimulates the immune system. The mechanism of action of vitamin D involves the release two potent molecules; cathelicidin and defensin, which have significant antibacterial and antiviral activities [17,18].
Based on the docking results from this study, vitamin D has comparatively less activity against both the main protease and ACE2 receptors of COVID-19. This vitamin interacts with the Thr292 residue of Mpro, with a binding score of −7.9532 kcal/mol (Fig. 1). It also interacts with Glu411 and forms two bonds with the His387 and His383 residues of the ACE2 receptor protein, with a binding score of −7.9279 kcal/mol (Fig. 2). Overall, the binding score for vitamin D is less than those of the other two vitamins studied here. However, these results are still significant when compared to the other drugs that are under clinical trials for the treatment of COVID-19.
Chloroquine has been tested in a randomized controlled trial and showed a beneficial effect on in COVID-19 patients with pneumonia [19]. However, the results of this molecular docking study showed that the docking score is −6.2930 kcal/mol less than those obtained for vitamins C, B, and D. Apart from this drug, researchers are exploring other antivirals in an effort to combat COVID-19. A similar study conducted on 16 well-known antiviral drugs observed that six—nelfinavir (−8.4 kcal/mol), rhein (−8.1 kcal/mol), withanolide D (−7.8 kcal/mol), withaferin A (−7.7 kcal/mol), enoxacin (−7.4 kcal/mol), and aloe-emodin (−7.4 kcal/mol)—were potential inhibitors of SARS-CoV-2. The results of the present study indicate that vitamins C and B have a greater inhibitory effect against SARS-CoV-2 than these antiviral drugs [20]. Moreover, these drugs have several side effects, so they must undergo comprehensive clinical trials prior to their use.
The COVID-19 pandemic is currently causing havoc across many regions of the world it is necessary to explore all the possible potential compounds that are safe to use and possess an excellent binding scores, as predicted in this simulation-based study, as potential alternative treatments for this infection.
Extensive research exploring the potential drugs available for the treatment of this novel coronavirus is in progress, along with ongoing clinical trials for the development of vaccines. Because the development of drugs and vaccines are time-consuming processes, these vitamins may be an excellent option to mitigate the adverse effects of COVID-19. The findings of this study may potentially make an important contribution to the management of this viral disease. The results suggest that the use of vitamins C and B is crucial; however, well-designed clinical trials are needed to develop standard protocols. The maintenance of vitamins at a normal physiological level might significantly reduce the chances of COVID-19 infection within the community.
Funding Statement: The author(s) received no specific funding for this study.
Conflicts of Interest: The authors declare that they have no conflicts of interest to report regarding the present study.
References
1. M. Hussain, N. Jabeen, F. Raza, S. Shabbir, A. A.Baig et al. (2020). , “Structural variations in human ACE2 may influence its binding with SARS-CoV-2 spike protein,” Journal of Medical Virology, vol. 10, no. 1002, pp. 1–7.
2. J. Lung, Y. S. Lin, Y. H. Yang, Y. L. Chou, L. H. Shu et al. (2020). , “The potential chemical structure of anti-SARS-CoV-2 RNA-dependent RNA polymerase,” Journal of Medical Virology, vol. 92, no. 6, pp. 693–697.
3. K. M. Kapoor and A. Kapoor. (2020). “Role of chloroquine and hydroxychloroquine in the treatment of COVID-19 infection-a systematic literature review,” medRxiv.
4. L. Zhang and R. Zhou. (2020). “Structural basis of the potential binding mechanism of remdesivir to SARS-CoV-2 RNA-dependent RNA polymerase,” Journal of Physical Chemistry, vol. 124, no. 32, pp. 6955–6962.
5. F. Hu, J. Jiang and P. Yin. (2020). “Prediction of potential commercially inhibitors against SARS-CoV-2 by multi-task deep model,” arXiv preprint arXiv: 2003.00728.
6. R. Yan, Y. Zhang, Y. Li, L. Xia, Y.Guo et al. (2020). , “Structural basis for the recognition of SARS-CoV-2 by full-length human ACE2,” Science, vol. 367, no. 6485, pp. 1444–1448.
7. M. Stoermer. (2020). “Homology models of coronavirus 3CLpro protease,” ChemRxiv. Preprint.
8. G. Onder, G. Rezza and S. Brusaferro. (2020). “Case-fatality rate and characteristics of patients dying in relation to COVID-19 in Italy,” Journal of American Medical Association, vol. 323, no. 18, pp. 1775–1776.
9. Y. Cai, Y. F. Li, L. P. Tang, B. Tsoi, M.Chen et al. (2015). , “A new mechanism of vitamin C effects on A/FM/1/47(H1N1) virus-induced pneumonia in restraint-stressed mice,” BioMed Research International, vol. 2015, 675149.
10. D. Slain, J. R. Amsden, R. A. Khakoo, M. A. Fisher, D. Lalka et al. (2005). , “Effect of high-dose vitamin C on the steady-state pharmacokinetics of the protease inhibitor indinavir in healthy volunteers,” Pharmacotherapy, vol. 25, no. 2, pp. 165–170.
11. R. M. L. Colunga Biancatelli, M. Berrill and P. E. Marik. (2020). “The antiviral properties of vitamin C,” Expert Review of Anti-infective Therapy, vol. 18, no. 2, pp. 99–101.
12. A. C. Carr. (2020). “A new clinical trial to test high-dose vitamin C in patients with COVID-19,” Critical Care, vol. 24, no. 1, pp. 1–2.
13. R. Z. Cheng. (2020). “Can early and high intravenous dose of vitamin C prevent and treat coronavirus disease 2019 (COVID-19)?,” Medicine in Drug Discovery, vol. 5, pp. 100028.
14. S. D. Keil, R. Bowen and S. Marschner. (2016). “Inactivation of Middle East respiratory syndrome coronavirus (MERS-CoV) in plasma products using a riboflavin-based and ultraviolet light-based photochemical treatment,” Transfusion, vol. 56, no. 12, pp. 2948–2952.
15. M. C. Chan, R. W. Chan, M. M. Ng, R. H. Ching and J. Peiris. (2017). “Regulation of proinflammatory cytokine by vitmain B2 in Influenza A/h5n1 virus infected human alveolar epithelial cells and macrophages, ” in Host-Pathogen Interactions. Washington, USA: American Thoracic Society, pp. A4045.
16. C. Herr, T. Greulich, R. A. Koczulla, S. Meyer, T. Zakharkina et al. (2011). , “The role of vitamin D in pulmonary disease: COPD, asthma, infection, and cancer,” Respiratory Research, vol. 12, no. 1, pp. 31.
17. G. Telcian, M. T. Zdrenghea, M. R. Edwards, V. Laza-Stanca, P. Mallia et al. (2017). , “Vitamin D increases the antiviral activity of bronchial epithelial cells in vitro,” Antiviral Research, vol. 137, pp. 93–101.
18. S. M. Currie, E. Gwyer Findlay, A. J. Mcfarlane, P. M. Fitch, B. Böttcher et al. (2016). , “Cathelicidins have direct antiviral activity against respiratory syncytial virus in vitro and protective function in vivo in mice and humans,” Journal of Immunology, vol. 196, no. 6, pp. 2699–2710.
19. J. Gao, Z. Tian and X. Yang. (2020). “Breakthrough: Chloroquine phosphate has shown apparent efficacy in treatment of COVID-19 associated pneumonia in clinical studies,” Bioscience Trends, vol. 14, no. 1, pp. 72–73.
20. V. Chandel, S. Raj, B. Rathi and D. Kumar. (2020). “In silico identification of potent COVID-19 main protease inhibitors from FDA approved antiviral compounds and active phytochemicals through molecular docking: A drug repurposing approach,” Preprints.
 | This work is licensed under a Creative Commons Attribution 4.0 International License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. |