DOI:10.32604/cmc.2022.022177

| Computers, Materials & Continua DOI:10.32604/cmc.2022.022177 |  |
| Article |
Plant Identification Using Fitness-Based Position Update in Whale Optimization Algorithm
1Department of Computer Science, College of Applied Studies, King Saud University, Riyadh, 11495, Saudi Arabia
2Department of Computer Science and Engineering, CHRIST (Deemed to be University), Bangalore, 560074, India
3Department of Computer Science, CHRIST (Deemed to be University), Bangalore, 560029, India
4Information Systems Department, Imam Mohammad Ibn Saud Islamic University (IMSIU), Riyadh, 11432, Saudi Arabia
*Corresponding Author: Abdul Khader Jilani Saudagar. Email: aksaudagar@imamu.edu.sa
Received: 30 July 2021; Accepted: 22 September 2021
Abstract: Since the beginning of time, humans have relied on plants for food, energy, and medicine. Plants are recognized by leaf, flower, or fruit and linked to their suitable cluster. Classification methods are used to extract and select traits that are helpful in identifying a plant. In plant leaf image categorization, each plant is assigned a label according to its classification. The purpose of classifying plant leaf images is to enable farmers to recognize plants, leading to the management of plants in several aspects. This study aims to present a modified whale optimization algorithm and categorizes plant leaf images into classes. This modified algorithm works on different sets of plant leaves. The proposed algorithm examines several benchmark functions with adequate performance. On ten plant leaf images, this classification method was validated. The proposed model calculates precision, recall, F-measurement, and accuracy for ten different plant leaf image datasets and compares these parameters with other existing algorithms. Based on experimental data, it is observed that the accuracy of the proposed method outperforms the accuracy of different algorithms under consideration and improves accuracy by 5%.
Keywords: Bag-of-features; feature optimization; plant leaf classification; swarm intelligence; nature-inspired algorithm
In the agricultural sector, plants play a vital role for all living things. Plant identification with greater precision is a complex issue that requires interdisciplinary research. The multidisciplinary approach combines computer vision and plant/botanical taxonomy. This approach helps in the automated classification and identification of plants. These techniques use leaf images as an input and perform classification/identification using machine learning and deep learning approaches. The main obstacle to automated identification is the lack of a suitable dataset. With the advancement in computer vision, many sophisticated models have been proposed to automate agricultural activities, including plant identification, soil classification, disease identification, weeds plant identification, and crop row detection. The timely and efficient identification of plants helps in management of crop related activities, including timely irrigation, supply of fertilizer, and removal of weed plants.
The process of plant identification includes feature extraction, optimization, and classification. Generally, feature extraction is performed by speed-up robust features (SURF) [1], scale-invariant feature transform (SIFT) [2] etc. The feature optimization phase is an important step where an individual can deploy an algorithm to select the best feasible set of features. Recent research used spider monkey optimization (SMO) [3], whale optimization algorithm (WOA) [4], bat algorithm [5], differential evolution (DE) [6], salp swarm algorithm (SSA) [7], sine cosine algorithm (SCA) [8], and many other swarms and evolutionary algorithms for solving this combinatorial optimization problem. Most of the recent research in the domain of plant identification used artificial neural network (ANN) and support vector machine (SVM) [9], deep residual network [10], convolutional neural network (CNN) [11], deep learning models (AlexNet, VGGNet, and GoogLeNet) [12] CNN models (Xception, ResNet50, InceptionV3, and MobileNet) and recurrent neural networks (RNN) including simple RNN, gated recurrent unit (GRU), and long short term memory (LSTM) models [13], and convolutional siamese network (CSN) [14] for efficient classification.
The automated identification of plants is based on the analysis of leaf images. Therefore, leaves are critical sources of information about the plant. However, this task is challenged by many hurdles like similarity in plant leaves, background variation, and the colour of leaves. Moreover, natural images require an efficient segmentation approach for further processing. Thus, developing machine learning and deep learning approaches to identify plants with higher accuracy is highly desirable. This study presents a new method for plant identification with improved WOA-based feature selection, leading to efficient classification. The significant research contribution of this paper is as follows:
1. A fitness-based WOA (FWOA) was proposed, and its performance was evaluated over a set of benchmark functions.
2. Feature extraction was performed using SIFT and SVM classifier used for classification.
Following is a breakdown of the rest of the paper. Section 2 discusses some recent developments in plant identification and WOA. Feature extraction, feature selection, and classification are discussed in Section 3. Section 4 discusses the experimental results of FWOA and its application to plant leaf identification. Section 5 concludes the paper.
Computational intelligence-based techniques can solve a complex optimization problem with fewer resources. These techniques are classified into different classes based on their source of inspiration, like swarm-based, evolutionary, and bio-inspired. These algorithms start with a set of randomly generated populations. Subsequently, each update their position shares details with other individuals and selects the best one for the next iteration. This section discusses recent development in plant identification and the basics of WOA [15].
Machine learning and deep learning are becoming more popular nowadays for the identification of plants. Some of the recent research contributions for the identification of plants using these techniques are discussed here. Pankaja and Suma deployed WOA to reduce dimensions and classified using Random Forest (RF) [15]. The author extracted texture, shape, and color features from the leaf image dataset. The WOA-based approach selects a set of optimal features. They used Flavia and Swedish leaf datasets for this experiment. Results reported that the WOA-based strategy outperformed other considered algorithms for feature extraction, feature selection, and plant identification. Sun et al. deployed a deep residual network with 26 layers on the BJFU100 dataset collected from their university campus [16]. The new approach was first validated on the Flavia leaf dataset with a 99.65% recognition rate. The main feature of this work is that Sun et al. acquired this data set on a mobile device [16]. Ghazi et al. employed transfer learning with a deep neural network [12]. Here, the author performed fine-tuning of the pre-trained model and used AlexNet, VGGNet, and GoogLeNet. The new model gives significantly improved results. Zhu et al. deployed a deep CNN with a set of five max-pooling layers, five soft-max layers, three fully connected layers, and sixteen convolutional layers [11]. This study concluded that the use of ReLUs along with these layers improved overall performance. Finally, Rzanny et al. studied various image acquisition and preprocessing techniques to identify plants with varying backgrounds [17]. Kho et al. focused on intact leaves [9] and used ANN and SVM to identify Ficus species plants. Original images were preprocessed by detecting edges and segmentation. While extracting shape and texture features, they reported 83% accuracy. Bodhwani et al. deployed a 50-layer deep residual network for plant identification and achieved 93% accuracy [10]. Liu et al. combined pre-trained CNN (Exception, ResNet50, InceptionV3, and Mobile Net) and RNN (simple RNN, GRU, and LSTM) models [13]. Some of the combinations archived very high accuracy. A summary of some of the recent development in plant identification is illustrated in Tab. 1.

2.2 Whale Optimization Algorithm
The WOA is a new nature-inspired algorithm developed by Mirjalili and Lewis in 2016 [4]. This nature-inspired optimization algorithm is used to solve many complex real-world optimization problems. WOA is inspired by the bubble-net hunting approach used by humpback whales during foraging. This method mimics the hunting style by using the fittest search agent to hunt the prey, and the spiral method is used to model the bubble-net attacking mechanism. The hunting method is an exciting mechanism for humpback whales. This approach of hunting is recognized as the bubble-net feeding strategy [20]. The mathematical model of this optimization algorithm majorly consists of three steps. The first step is encircling prey, the second step is a bubble net attacking method (exploitation phase), and the last step is the search for prey (exploration phase) [4]. Each phase is illustrated in subsequent sections.
Humpback whales locate the target and encircle it. Initially, the optimal design is unknown; hence, the WOA method assumes the target prey as the present ideal candidate solution, or it can be close to the optimum. Once the optimal search agent is well-defined, some other agent will update the location of the existing best search agent.
where
where
2.2.2 Bubble-Net Attacking Method
The exploitation phase in WOA is simulated by the bubble-net behaviour of humpback whales with two steps.
1. Shrinking encircling mechanism: The behaviour of the humpback whale is accomplished by decreasing the value in Eq. (3) from 2 to 0. As the value of
2. Spiral updating position: In this step, compute the distance between the whale situated at (
where d is the distance of prey from ithwhale to take the best solution obtained so far, l is a random number, b is a constant, which defines the shape of the spiral. Thus, humpback whales continuously swim in a spiral-shaped path within a decreasing circle around the prey. To model this synchronized behaviour, assume a 50% probability of selecting either a shrinking circle or a spiral model to update the whale's location. The calculated model is shown in Eq. (6).
where p is a random number within the range [0, 1].
In the bubble net technique, humpback whales hunt prey randomly according to each other's location. Vector A is used to search for prey in the exploration phase, calculated in the first phase. In this step, update the location of the search agent by using the randomly selected search agent. This method sheds light on exploration and allows this algorithm to perform global searches. The calculated model is shown in Eqs. (7) and (8).
where Xrandis an|| arbitrarily chosen whale from the current population. The detailed pseudo code for WOA is given in Algorithm 1.
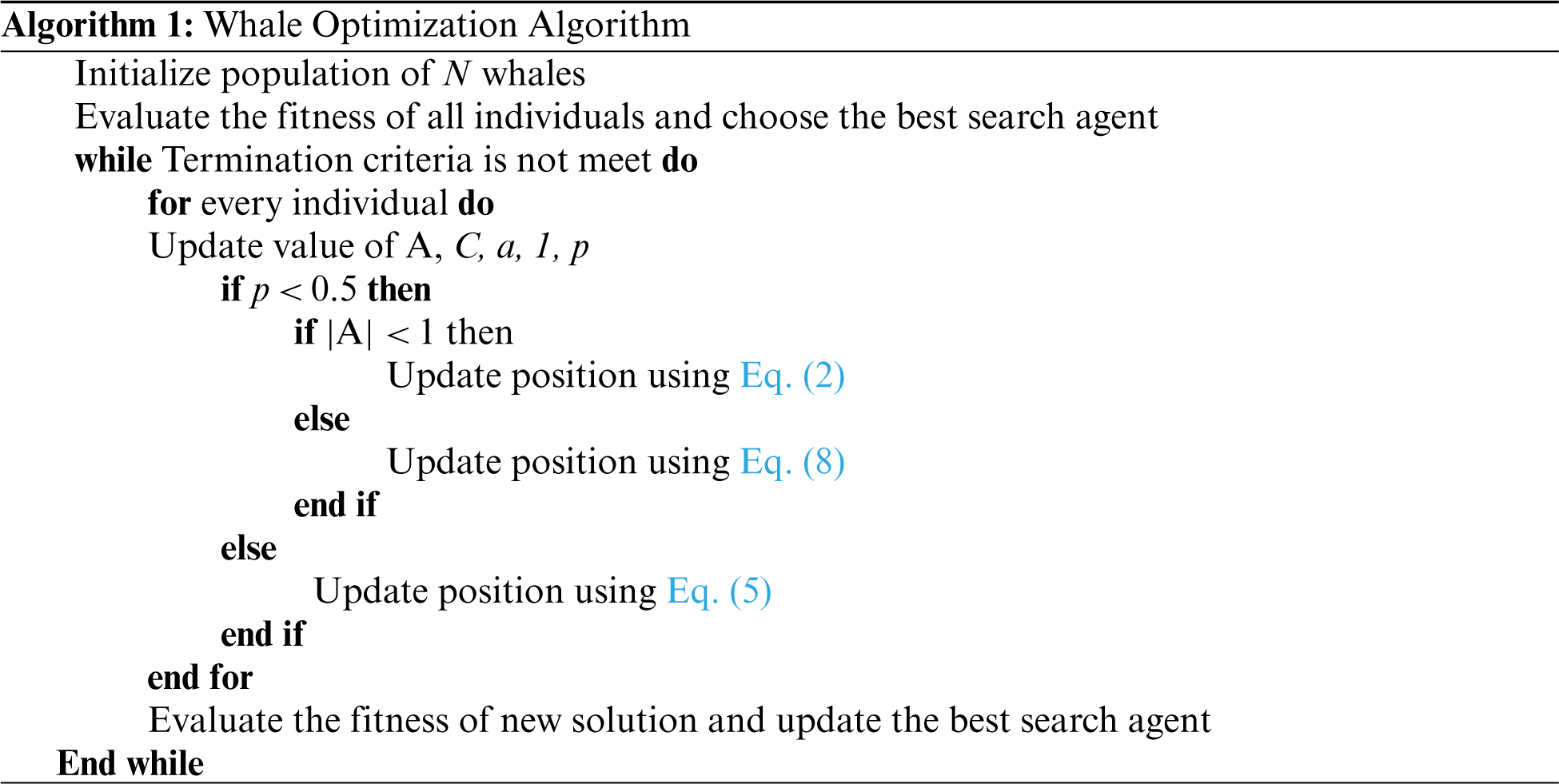

Initially, the WOA starts with some arbitrary solutions. Then, individuals update the positions using the best answer ever found on each iteration or an arbitrarily picked individual. Using vector,

3 Fitness-Based Whale Optimization Algorithm
Exploitation and exploration are two significant phases in all the meta-heuristic algorithms for accomplishing the precise solution. The performance of an algorithm is strongly reliant on balancing these two opposing processes. In the WOA algorithm, bubble-net attacking is responsible for exploitation and search for prey phase perform exploration. They are essential phases in the WOA algorithm and affect the convergence behavior of WOA. The exploration phase searches whales’ property for renovating position; this selection uses the random function for updating to recognize the best whale [4]. To improve the performance of WOA, a new version of WOA is proposed here and named fitness-based status update WOA (FWOA). The new variant update uses highly fitted solutions and explores the search space for a solution with low fitness. The introduced concept works on the principle that solutions in the proximity of higher fitness solutions are also highly fitted and try to exploit the best solution. In the case of low fitness, it updates its position according to the search for prey phase. Detailed pseudo-code for the new strategy is given in Algorithm 2.
Additionally, a fitness-based method is used to compute the value of
The vectors
The new approach takes advantage of a highly fitted solution. It assumes that the proximity of highly-fitted solutions may be a feasible solution for the considered problem. As a result, the swarm always moves in the direction of the solution with good fitness with self-organizing characteristics, and it improves the convergence speed and avoids skipping real solutions.
The performance of the newly proposed FWOA is evaluated over a set of thirteen benchmark problems [4]. The selected problems are uni-modal and multi-modal optimization problems with known solutions and search. Performance of FWOA and other competitive algorithms compared in terms of the average function value (Avg), standard deviation (SD), and optimal function value. All the algorithms are implemented in MATLAB R2020b on an Intel Core i7 machine with 16 GB RAM and 8 GB GeForce GTX1650Ti Graphics processor to measure these parameters. Tabs. 2 and 3 illustrate results for FWOA, WOA [4], SSA [16], SCA [8]. Tab. 2 illustrates the efficiency and robustness of FWOA in comparison to other algorithms. Graphical representations of results for functions F1, F8, and F12 are depicted in Fig. 1. These results proved that FWOA outperformed considered algorithms in terms of best function value, as shown in Tab. 3.

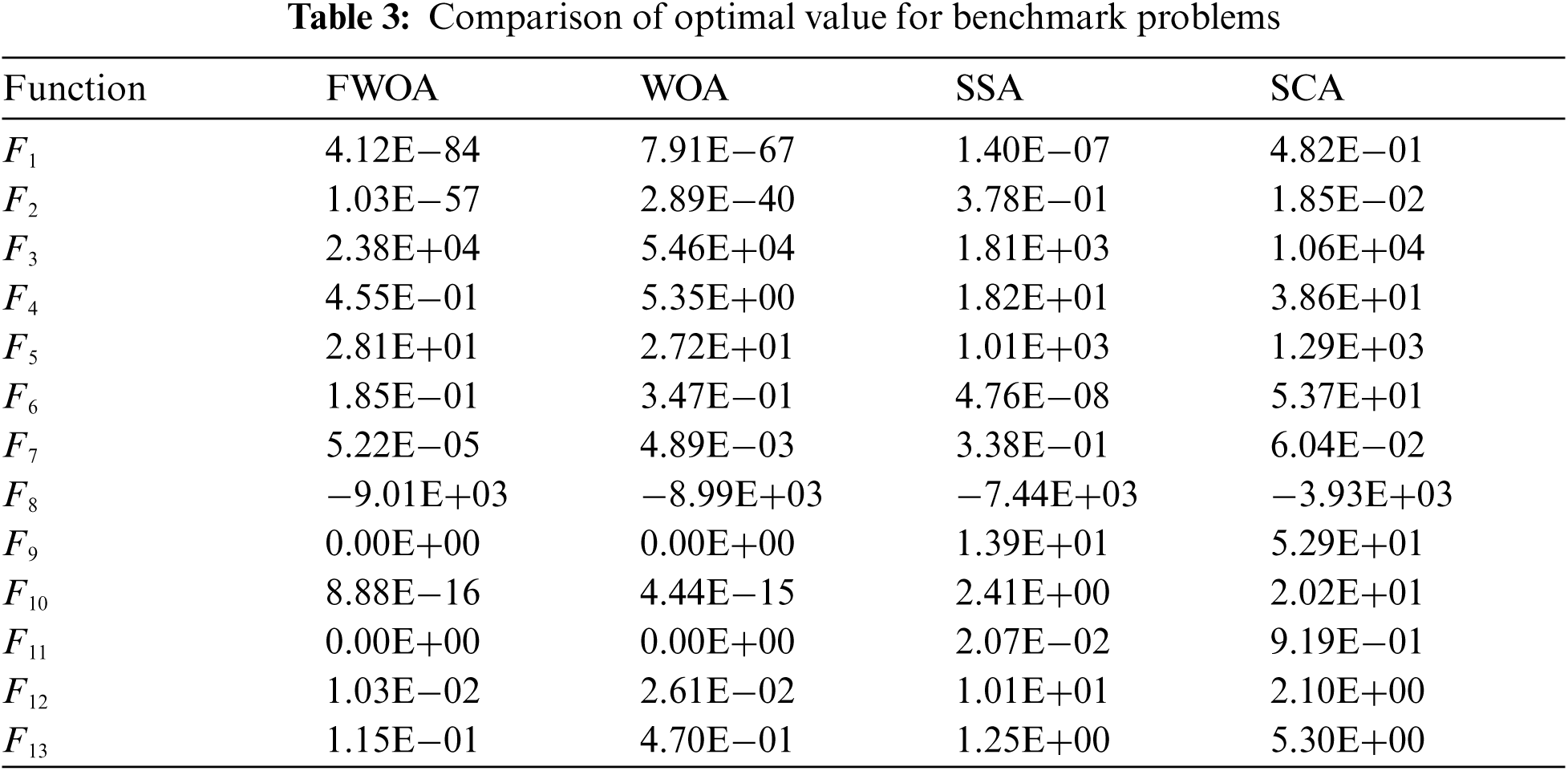


Figure 1: Performance assessment for selected functions. (a) F1 WOA (b) F1 SSA (c) F1 SCA (d) F1 FWOA (e) F8 WOA (f) F8 SSA (g) F8 SCA (h) F8 FWOA (i) F12 WOA (j) F12 SSA (k) F12 SCA (l) F12 FWOA
4 FWOA-Based Plant Identification System
The proposed model introduced a fitness-based WOA to classify plants based on the leaf image dataset. The suggested model has three significant steps: feature extraction using the SIFT algorithm, feature selection, histogram generation using the modified WOA, and classification of plants based on their leaf image using the SVM classifier, as shown in Fig. 2. A detailed description is given in later sections. The detailed process of the plant leaf classification model is shown in Fig. 3. To validate the proposed model, images of apple, banana, borages, maize, grapes, mint, orange, pepper, potato, and tomato leaf were used in this research. This dataset is used as a sample dataset for validating this model. Some sample images from each category are depicted in Fig. 4.

Figure 2: Process of the proposed model

Figure 3: Proposed plant leaf classification model

Figure 4: Sample plant leaf image set for considered plants. (a) Apple (b) Banana (c) Borages (d) Corn (e) Grapes (f) Mint (g) Orange (h) Pepper (i) Potato (j) Tomato
In image processing and computer vision, a feature is an information in a picture [21]. Objects, edges, and points, for example, have extraordinary quality and distinct structure. Feature extraction is a process of classifying essential features of an image, classifying common themes from a broad collection of images, and pattern recognition [22]. The proposed model's first step is to extract all image features and group them into corresponding groups. This extraction is one of the leading steps for image analysis relating to their features. Similar and different image features have to be extracted and stored in respective clusters for practical analysis. The SIFT algorithm is used to extract the features in the proposed method. SIFT is a feature detection method that detects and defines local features of plant leaf images. These local features are essential points in the image that aid in identifying the object of the image [23]. This method can rotate and select an image of a different scale and handle the noise points. Therefore, it is a practical algorithm for feature extraction.
Feature selection is a technique that significantly affects the performance of the proposed classification model. A combinatorial optimization problem is selecting an optimal collection of features from a vast set of extracted features. Thus, it is highly desired to solve this problem with a non-conventional optimization algorithm. This step chooses the most relevant elements that will aid in estimating the class of each leaf image. Next, extracted features from the previous step are used to select the optimal features and create clusters using selected features, increasing accuracy and decreasing overfitting. This paper used the modified WOA for clustering to select optimal features. Finally, a histogram is plotted using selected features by the proposed model. This histogram shows the fundamental frequency distribution of the selected features. In addition, the histogram allows the review of the selected features in terms of outliers and skewness. The graphical representation is depicted in Fig. 5.

Figure 5: Histogram of the proposed model
Classification of plants using a leaf image dataset is the final step of this proposed method. In the previous step, the histogram is generated based on the selected features and passed to SVM classifier along with their labels [24]. SVM is a high-performance binary classifier, which creates a hyperplane in ample feature space for separate leaf images into their respective classes [25]. In this step, the SVM classifier predicts the class labels of each plant leaf image based on training. Hence labelling, training and testing plant leaf image dataset confirm the accuracy of this model. Experimental results are discussed in the next section.
5 Experimental Results for Plant Identification
Three steps are used to analyze the proposed plant classification using a leaf image dataset based on FWOA. The first step represents plant leaf dataset description, the second step shows the performance of benchmark functions, and the third step analyses the result of FWOA based plant leaf classification.
This dataset consists of more than 10000 images; 200 images from each category are used for training and testing this model. This dataset is categorized into ten different classes named apple, banana, borages, corn, grapes, mint, orange, pepper, potato, and tomato. This dataset is taken from Plant Village [26] and Kaggle [27]. This dataset is used to measure the performance of the proposed method in terms of the accuracy of classification of each class using a leaf image dataset. These images are divided into a 70%–30% train cross-test split for each class.
5.2 Experimental Results for Plant Leaf Image Classification
The proposed model has been predicted outputs using Python programming. In this section, the proposed approach is described using experimental results based on the input image dataset. Tab. 4 shows some of the parameters and best fitness values. The value of these parameters is decided with exhaustive experiments. The proposed modified WOA has been compared with SCA, BAT, SSA, DE, and WOA. An equal number of image sets have been from each class for these algorithms. Create a confusion matrix concerning each class for performance analysis. The confusion matrix for each class is depicted in Fig. 6. These matrixes show the comparison of actual data and predicted data. The performance of all the considered algorithms for classification is illustrated in Fig. 6. It is important as the considered data set has ten classes. In the case of three or more categories, it is better to visualize results with confusion matric as accuracy can be misleading. The results are measured by calculating the F1 score, precision, recall, and accuracy.
where TP stands for true positive, TN for true negative, FP for false positive, and FN for false negative. The measured performance and comparisons are shown in Tab. 5. This table summarizes accuracy and other matrices for considered algorithms. The proposed method shows better performance when compared with another existing algorithm. For example, compare the accuracy of the modified WOA algorithm with some other algorithms, which is depicted in Fig. 7. Hence, it can be stated that the proposed modified WOA classification method is better than the existing algorithms.


Figure 6: Confusion matrix for considered algorithms. (a) SCA (b) BAT (c) SSA (d) DE (e) WOA (f) MWOA


Figure 7: Comparison of overall accuracy
Using a plant leaf image dataset, this study presents a new plant classification method. The new version of WOA uses a fitness-based status update method instead of random numbers. This method shows the effectiveness of the results by estimating the maximum accuracy value. In this study, we primarily used three steps: feature extraction using the SIFT method, feature selection using the modified WOA method, and classification using the SVM classifier. The proposed method achieves maximum recall, precision, F1 scores, and accuracy with 80.16%. We analyze the experimental results, and it was found that the WOA with the fitness function increased the efficiency of the proposed algorithm. WOA is employed to handle the problem of feature selection and clustering in this study. The proposed algorithm results are compared with well-known stochastic algorithms such as BAT, DE, WOA, SCA, and SSA.
Furthermore, when compared to other algorithms, the proposed method's results were effective, practical, and simple to implement. In the future, the proposed method can be applied to various plant classifications utilizing different plant leaf image datasets. Besides this, the WOA can be combined with another clustering approach to improve performance.
Acknowledgement: The authors extend their appreciation to the Deanship of Scientific Research at King Saud University for funding this research.
Funding Statement: This work was supported by the Deanship of Scientific Research, King Saud University, Saudi Arabia.
Conflicts of Interest: The authors declare that they have no conflicts of interest to report regarding the present study.
1. H. Bay, A. Ess, T. Tuytelaars and L. V. Gool, “Speeded-up robust features (SURF),” Computer Image and Vision Understanding, vol. 110, no. 3, pp. 346–359, 2008. [Google Scholar]
2. H. R. Kher and V. K. Thakar, “Scale invariant feature transform-based image matching and registration,” in Proc. 2014 Fifth Int. Conf. Signal Image Processing, Bangalore, India, pp. 50–55, 2014. [Google Scholar]
3. J. C. Bansal, H. Sharma, S. S. Jadon and M. Clerc, “Spider monkey optimization algorithm for numerical optimization,” Memetic Computing, vol. 6, no. 1, pp. 31–47, 2014. [Google Scholar]
4. S. Mirjalili and A. Lewis, “The whale optimization algorithm,” Advances in Engineering Software, vol. 95, pp. 51–67, 2016. [Google Scholar]
5. X. S. Yang and A. H. Gandomi, “Bat algorithm: A novel approach for global engineering optimization,” Engineering Computations, vol. 29, no. 5, pp. 464–483, 2012. [Google Scholar]
6. R. Storn and K. Price, “Differential evolution: A simple and efficient adaptive scheme for global optimization over continuous spaces,” Journal of Global Optimization, vol. 11, pp. 341–359, 1997. [Google Scholar]
7. S. Mirjalili, A. H. Gandomi, S. Z. Mirjalili, S. Saremi, H. Faris et al., “Salp swarm algorithm: A bio-inspired optimizer for engineering design problems,” Advanced Engineering Software, vol. 114, pp. 163–191, 2017. [Google Scholar]
8. S. Mirjalili, “SCA: A sine cosine algorithm for solving optimization problems,” Knowledge-Based Systems, vol. 96, pp. 120–133, 2016. [Google Scholar]
9. S. J. Kho, S. Manickam, S. Malek, M. Mosleh and S. K. Dhillon, “Automated plant identification using artificial neural network and support vector machine,” Frontiers in Life Sciences, vol. 10, pp. 98–107, 2017. [Google Scholar]
10. V. Bodhwani, D. P. Acharjya and U. Bodhwani, “Deep residual networks for plant identification,” Procedia Computer Science, vol. 152, pp. 186–194, 2019. [Google Scholar]
11. H. Zhu, Q. Liu, Y. Qi, X. Huang, F. Jiang et al., “Plant identification based on very deep convolutional neural networks,” Multimedia Tools and Applications, vol. 77, pp. 29779–29797, 2018. [Google Scholar]
12. M. M. Ghazi, B. Yanikoglu and E. Aptoula, “Plant identification using deep neural networks via optimization of transfer learning parameters,” Neurocomputing, vol. 235, pp. 228–235, 2017. [Google Scholar]
13. X. Liu, F. Xu, Y. Sun, H. Zhang and Z. Chen, “Convolutional recurrent neural networks for observation-centered plant identification,” Journal of Electrical and Computer Engineering, vol. 2018, pp. 1–7, 2018. [Google Scholar]
14. G. Figueroa-Mata and E. Mata-Montero, “Using a convolutional siamese network for image-based plant species identification with small datasets,” Biomimetics, vol. 5, no. 1, pp. 1–17, 2020. [Google Scholar]
15. K. Pankaja and V. Suma, “Plant leaf recognition and classification based on the whale optimization algorithm (WOA) and random forest (RF),” Journal of the Institution of Engineers (IndiaSeries B, vol. 101, pp. 597–607, 2020. [Google Scholar]
16. Y. Sun, Y. Liu, G. Wang and H. Zhang, “Deep learning for plant identification in natural environment,” Computer Intelligence and Neuroscience, vol. 2017, pp. 1–6, 2017. [Google Scholar]
17. M. Rzanny, M. Seeland, J. Wäldchen and P. Mäder, “Acquiring and preprocessing leaf images for automated plant identification: Understanding the tradeoff between effort and information gain,” Plant Methods, vol. 13, pp. 1–11, 2017. [Google Scholar]
18. J. Chen, J. Chen, D. Zhang, Y. Sun and Y. A. Nanehkaran, “Using deep transfer learning for image-based plant disease identification,” Computers and Electronics in Agriculture, vol. 173, pp. 1–11, 2020. [Google Scholar]
19. S. R. G. Reddy, G. P. S. Varma and R. L. Davuluri, “Optimized convolutional neural network model for plant species identification from leaf images using computer vision,” International Journal of Speech Technology, pp. 1–28, 2020. https://doi.org/10.1007/S10772-021-09843-X. [Google Scholar]
20. W. A. Watkins and W. E. Schevill, “Aerial observation of feeding behavior in four baleen whales: Eubalaena glacialis, balaenoptera Borealis, megaptera novaeangliae, and balaenoptera physalus,” Journal of Mammalia, vol. 60, no. 1, pp. 155–163, 1979. [Google Scholar]
21. J. Sklansky, “Image segmentation and feature extraction,” IEEE Transactions on Systems, Man, and Cybernetics, vol. 8, no. 4, pp. 237–247, 1978. [Google Scholar]
22. G. Kumar and P. K. Bhatia, “A detailed review of feature extraction in image processing systems,” in Proc. 2014 Fourth Int. Conf. on Advanced Computing & Communication Technologies, Rohtak, India, pp. 5–12, 2014. https://doi.org/10.1109/ACCT.2014.74. [Google Scholar]
23. Z. N. Khudhair, F. Mohamed and K. A. Kadhim, “A review on copy-move image forgery detection techniques,” Journal of Physics: Conference Series, vol. 1892, pp. 1–11, 2021. [Google Scholar]
24. M. A. Chandra and S. S. Bedi, “Survey on SVM and their application in image classification,” International Journal of Information Technology, 2018. https://doi.org/10.1007/s41870-017-0080-1. [Google Scholar]
25. X. Li, L. Wang and E. Sang, “Multi-label SVM active learning for image classification,” in Proc. 2004 Int. Conf. on Image Processing, vol. 4, pp. 2207–2210, 2004. [Google Scholar]
26. Plant village dataset, https://mavyn.in/webapp/pdf/googledrive/okjocta/3e3833-plant-village-dataset (accessed on July 26, 20212021. [Google Scholar]
27. S. P. Mohanty, D. P. Hughes and M. Salathé, “Using deep learning for image-based plant disease detection,” Frontiers in Plant Science, vol. 7, pp. 1–10, 2016. [Google Scholar]
 | This work is licensed under a Creative Commons Attribution 4.0 International License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. |