DOI:10.32604/cmc.2022.025750

| Computers, Materials & Continua DOI:10.32604/cmc.2022.025750 |  |
| Article |
Pulmonary Diseases Decision Support System Using Deep Learning Approach
1Department of Computer Engineering, Yarmouk University, Irbid, 21163, Jordan
2Department of Biomedical Systems and Informatics Engineering, Yarmouk University, Irbid, 21163, Jordan
3Department of Biomedical Engineering, Jordan University of Science and Technology, Irbid 22110, Jordan
4Pediatrician, Mediclinic Hospital, Al Ain, 14444, UAE
*Corresponding Author: Ali Mohammad Alqudah. Email: ali_qudah@hotmail.com
Received: 03 December 2021; Accepted: 17 March 2022
Abstract: Pulmonary diseases are common throughout the world, especially in developing countries. These diseases include chronic obstructive pulmonary diseases, pneumonia, asthma, tuberculosis, fibrosis, and recently COVID-19. In general, pulmonary diseases have a similar footprint on chest radiographs which makes them difficult to discriminate even for expert radiologists. In recent years, many image processing techniques and artificial intelligence models have been developed to quickly and accurately diagnose lung diseases. In this paper, the performance of four popular pretrained models (namely VGG16, DenseNet201, DarkNet19, and XceptionNet) in distinguishing between different pulmonary diseases was analyzed. To the best of our knowledge, this is the first published study to ever attempt to distinguish all four cases normal, pneumonia, COVID-19 and lung opacity from Chest-X-Ray (CXR) images. All models were trained using Chest-X-Ray (CXR) images, and statistically tested using 5-fold cross validation. Using individual models, XceptionNet outperformed all other models with a 94.775% accuracy and Area Under the Curve (AUC) of Receiver Operating Characteristic (ROC) of 99.84%. On the other hand, DarkNet19 represents a good compromise between accuracy, fast convergence, resource utilization, and near real time detection (0.33 s). Using a collection of models, the 97.79% accuracy achieved by Ensemble Features was the highest among all surveyed methods, but it takes the longest time to predict an image (5.68 s). An efficient effective decision support system can be developed using one of those approaches to assist radiologists in the field make the right assessment in terms of accuracy and prediction time, such a dependable system can be used in rural areas and various healthcare sectors.
Keywords: Pulmonary diseases; deep learning; lung opacity; classification; majority voting; ensemble features
The likelihood of illnesses is elevating as a result of environmental changes, climate change, lifestyle, and other reasons. In 2016, over 3.4 million people died from Chronic Obstructive Pulmonary Disease (COPD), which is primarily caused by smoking and pollution, while 400,000 people perished because of asthma [1]. A new coronavirus illness (COVID-19) has been causing major lung damage and breathing issues since late December 2019 [2,3]. The impact of lung illness is substantial, particularly in emerging and low-middle-income nations where millions of people are impoverished and exposed to air pollution. According to the World Health Organization (WHO) estimates, nearly 4 million premature deaths occur each year as a result of home air pollution-related illnesses such as asthma and pneumonia. As a result, it is critical to take the required actions to minimize air pollution and carbon emissions. It is also essential to put in place effective diagnostic methods that can aid in the detection of lung problems.
Early, quick, and reliable detection of pulmonary diseases patients is critical in helping them receive the proper treatment and assist health authorities identify and isolate infected patients which is going to help contain the disease. Traditional Polymerase Chain Reaction (PCR) has been the primary tool for detecting COVID-19 and other pulmonary diseases, but the problem with PCR and similar laboratory tests is that they are expensive, and the results might take more than 1–2 working days to appear. Delays and long processing times of traditional methods can worsen the overall health situation [4,5]. Thus, various researchers tried to employ contemporary computer vision techniques, those techniques use a labelled dataset to train the models, eventually the models will learn, and be able to make predictions on unseen radiographs with high accuracy.
Machine learning techniques have been used for years to detect and diagnose various illnesses and diseases. Deep learning techniques a subset of machine learning techniques proved to be effective and efficient in analyzing medical images [6]. Since the beginning of the pandemic, Convolutional Neural Networks (CNNs) have been employed to detect COVID-19 in Chest-X-Ray (CXR) and Computed Tomography (CT) images [7] and to differentiate between COVID19 and other pulmonary diseases. The problem with CT images is that CT scanners are expensive and cannot be easily found in remote areas and developing countries. Despite their weak resolution, Chest-X-Ray (CXR) radiographs are inexpensive and easier to obtain. For all of that, detecting the presence or absence of COVID-19 using Chest-X-Ray (CXR) images might be a quick, easy, and reliable fix. The problem is that few annotated chest radiographs are publicly available which makes such techniques difficult to apply.
The pandemic overwhelmed radiologists with patients and radiographs, making the process of reading and interpreting radiographs a highly variable one. To make things worser, some COVID-19 patients do not show any symptoms (asymptomatic patients), they themselves are not aware of them being infected which makes identifying and isolating them a challenging task. The aim of this study is to compare the performance of four pretrained deep learning-based models (DenseNet201, Darknet, Xception, and VGG16) in distinguishing between normal, pneumonia, and COVID-19 (multi class classification) using publicly available Anterior Posterior (AP) Chest-X-Ray (CXR) radiographs. Additionally, a collection of models like the Features Extraction and Majority Voting techniques will be employed. Eventually, the most effective method will be used towards building a reliable system that will help healthcare providers diagnose COVID-19 patients in masses quickly and accurately.
The organization of the paper is as follows. Section 2 describes the relevant literature, and Section 3 describes the data used in this study. Section 4 describes the pretrained models, our proposed methods, and training procedure. Section 5 addresses the experimental results obtained, and finally Section 6 concludes the article, and outlines the future research direction.
In the past couple of years and as a consequence of the pandemic, a lot of research was conducted to early detect different pulmonary diseases mainly COVID-19 with high accuracy. In this section we survey the most recent and relevant literature. To the best of our knowledge, nobody previously attempted to classify all four pulmonary diseases (normal, pneumonia, COVID-19 and lung opacity) from Chest-X-Ray (CXR) images.
In 2020, Yasin Yari et al. used transfer learning particularly ResNet50 and DenseNet121 combined with a deep classifier to discriminate between normal, viral pneumonia, and COVID-19 radiographs. The proposed architecture achieved 97.83% accuracy using a small dataset with augmentation [8]. Jothi Pranav et al. compared the performance of pretrained models (VGG19, ResNet50, and DenseNet121) in discriminating between normal, COVID-19, and viral pneumonia. DenseNet121 outperformed all other models with a 95% sensitivity and 97% specificity [9]. Irfan Ullah Khan et al. used four deep learning models namely VGG16, VGG19, DenseNet121, and ResNet50 to detect the presence or absence of COVID-19. They merged multiple open-source datasets, and VGG16/VGG19 achieved the highest accuracy of 99.38% with respect to the other models [10]. Lacruz et al. compared the performance of six different pretrained architectures namely VGG16, Xception, ResNet50, ResNet101, DenseNet201, and InceptionResNetV2. They used the Kaggle dataset, they augmented the data because of the unbalanced nature of the dataset. The goal was to discriminate between healthy people, COVID-19, and viral pneumonia. The best performance was obtained by Xception with an accuracy of 97.34% [11].
In 2021, Weihan Zhang et al. developed a novel architecture to distinguish between healthy, pneumonia, and COVID-19 in chest radiographs. They developed a Support Vector Machine (SVM) that was trained with a combination of 308 handcrafted and 1000 VGG16/ResNet50 deep features. The proposed framework achieved a classification accuracy of 98.8% [12]. Matthias Fontanellaz et al. developed a deep learning diagnostic support system to discriminate between normal, COVID-19 pneumonia and other types of pneumonia in Chest-X-Rays (CXR). The suggested model uses a light architecture and achieves an overall diagnostic accuracy of 94.3% surpassing the performance of eleven expert radiologists with different expertise (61.4%) [13]. Oyelade et al. developed a deep learning framework for COVID-19 detection named CovFrameNet. The suggested model uses an enhanced image preprocessing technique combined with a Convolutionsal Neural Network (CNN). They also used radiographs from the COVID-19 radiography database and the NIH Chest-X-Ray (CXR) datasets. They claim to have achieved 100% accuracy in detecting the presence or absence of COVID-19 [14]. Tuan Pham et al. used transfer learning to compare the performance of three popular pretrained models particularly SqueezeNet, GoogleNet, and AlexNet to perform two classes and three classes discrimination tasks. They used subsets from three publicly available databases to train the models in short time for rapid deployment [15]. Alquran et al. utilized the texture features from the enhanced Chest-X-Ray images to distinguish between two pulmonary disease classes alongside the normal case with 93.2% accuracy [16]. Finally, Alsharif et al. focused their research on pediatric Chest-X-Ray images and employed deep learning techniques, they created a light CNN to discriminate between causes of pulmonary diseases whether it is viral or bacterial, their model achieved a near 100% accuracy [17].
The methodology in this paper consists of 3 main stages using four different deep learning models trained with image augmentation. Fig. 1 illustrates the overall system diagram for the four-class image classification problem. All experiments were written in MATLAB R2021, they ran on a desktop computer that uses Windows 10 operating system, the computer used up to 16 GB of RAM, 500 GB Hard Disk Drive (HDD), and an Intel core i7–6700/3.4 GHz microprocessor.

Figure 1: Block diagram of the proposed system
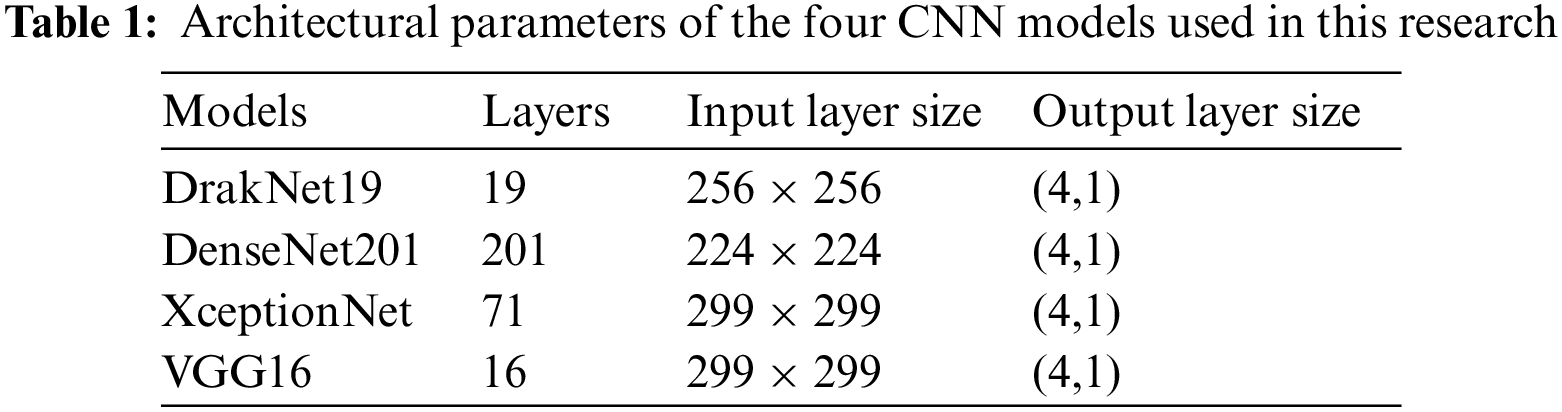
In this section we will briefly discuss some of the popular pretrained models used in visual recognition tasks. Of particular interest are the frameworks used to conduct the comparative performance study. All models were trained on the ImageNet dataset, using some 1.28 million images with 1000 object categories [18]. Here, the transfer learning property is utilized to be compatible with the new task of clas8sifying pulmonary diseases. Tab. 1 summarizes the deep learning models that were employed.
Dense Convolutional Network (DenseNet) is a Convolutional Neural Network that resembles ResNet but uses dense connections between layers termed dense blocks, all layers are directly connected to each other in a feed forward pattern. Shorter connections closer to the input and output layers were introduced and resulted in a more precise network. DenseNets are good feature extractors, they require few parameters and less computation time compared to other traditional models [19].
Developed in 2017 by Francois Chollet at Google, it is inspired by and based upon the Inception model. It is a deep learning structure with depth separable convolutions instead of Inception modules, it also uses bypass links between convolution blocks. It consists of 71 tiers and contains the same number of parameters (23 million parameters) used in the development of Inception V3. All convolution and depth separable convolution layers are followed by batch normalization [20].
It is a recently developed pre-trained Convolutional Neural network (CNN) that is used as the spine of the You Only Look Once (YOLOv2) model. It exhibited a remarkable performance on the real time object detection tasks. This model comprises of 5 maxpooling layers, and 19 convolutional layers, analogous to the VGG model, it usually utilizes 3 x 3 filters. The model employs a max and average pooling layer and takes 256 x 256 sized images as input to the network. It increases the number of channels by two folds after each max pooling step, it also uses 1 x 1 filters to squeeze the feature representations [21].
Introduced in 2015 by Visual Geometry Group (VGG) at Oxford, it uses a simple, very deep Convolutional Neural Network architecture that exploits very small 3 x 3 filters with small receptive fields. Other components include five max pooling layers, and the stack of convolutional layers is followed by three fully connected layers. By fixating all other model parameters, advancing the network depth up to 16 weight layers significantly improved the model large scale image classification accuracy [22]. Tab. 1 explains the architectural parameters of the four CNN models used in this research.
After generating the classification results using the four deep learning models, we will have four decisions for each image. The output class of each image are stacked, then the most frequent decision is selected. This technique is known as majority voting technique and will provide us with a strong prediction (low error) for each image since it is not dependent on a single model decision. Fig. 2 illustrates the proposed majority voting technique.

Figure 2: Block diagram of the proposed majority voting system
Ensemble learning improves the ability of the artificial intelligence learning system to produce a generalized model. Ensemble learning methods mainly include bagging, boosting, and stacking. These days, the most common tactic for generating base classifiers using deep learning models is to train different types of deep learning models using the same dataset, then stack the extracted deep features from these models together [23]. Usually, the base classifiers obtained by this method are different. The combination technique of integrated learning mainly includes stacking features or average or majority voting of classes. According to the different uses of integrated learning, different combination methods are usually selected. For example, if the purpose of integrated learning is classification, the features of each individual deep learning model are stacked together to obtain the final combined features space that will be used for classification [24,25]. In this research the deep features extracted from the four models are taken as input for the softmax classifier. Fig. 3 explains the proposed ensemble learning technique.

Figure 3: Block diagram of the proposed ensemble features system
This study utilized the COVID-19 radiography database which was published recently online by Chowdhury et al. [26,27]. The dataset contains a total of 14, 777 Posterior-to-Anterior (PA) and Anterior-to-Posterior (AP) Chest-X-Rays (CXRs). The images in this database are real radiographs used by radiologists in clinical diagnosis. The dataset was created using six different sub-databases. The 2,493 COVID-19 images were collected from publicly available databases, while normal and viral pneumonia databases were created from publicly available Kaggle databases. The collected dataset was labelled by specialists in the field of pulmonary diseases. Therefore, the proposed supervised approach is built and validated based on the ground truth of the labeled images. Tab. 2 shows the distribution of the dataset into four different types of diseases Normal, COVID-19, Viral Pneumonia, and Lung Opacity (Non-COVID lung infection). Fig. 4 shows sample images from the used dataset.
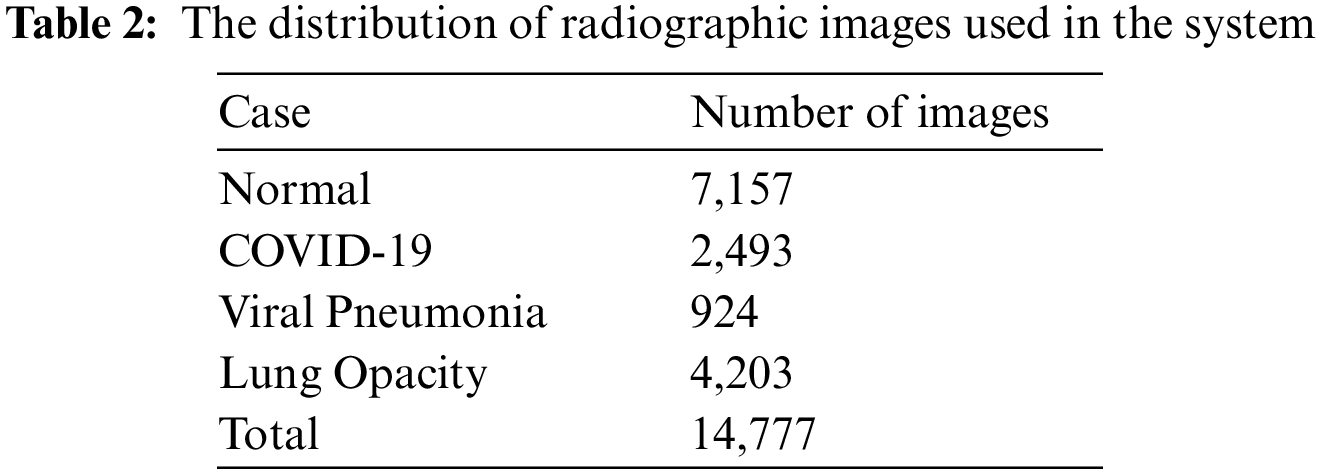



Figure 4: Sample images of the used dataset; (a) Normal; (b) COVID-19; (c) Viral Pneumonia; (d) Lung Opacity
Various data augmentation methods were applied to solve the scarcity of COVID-19 radiographs. In this study, three different image augmentation techniques (rotation, reflection, and shear) were utilized to generate the training images. The rotation operation used for image augmentation was done by rotating the images in the clockwise and counterclockwise direction with an angle of -5 and 5 degrees. Image reflection was employed by flipping the image horizontally and vertically, and finally the shear was done by shearing the images randomly by 5% in both x and y directions [17]. Fig. 5 shows sample-augmented radiographs.

Figure 5: (A) Original chest X-ray image; (B) Image after rotation by 5 degrees clockwise; (C) Image after horizontal flip; (D) Image after vertical flip; (E) Image after 5% horizontal shear; (F) Image after 5% horizontal shear
In general, evaluating any artificial intelligence model is a crucial and complex endeavor due to many factors like variations on the size of the used dataset and difference in the architecture of the model used. Traditionally, researchers used a simple method to evaluate the system that basically depends on splitting the used dataset into training, and testing sets using different ratios, usually 70% for training and 30% for testing. Following that they use the training set to train the model, later the testing set is used to test the ability of the model to deal with unseen data. Finally, the performance metrics of the model are collected and evaluated [17,28]. Nevertheless, this method is not suitable to build a generalized model and to test it is performance on different testing sets since the obtained accuracy using one test set can be very different from the one obtained from the other testing sets. As a result, the best method to evaluate model performance is using the K-fold Cross Validation, which provides a super solution to this problem. The data is split into different folds defined by the user and each fold is used as a testing set at some point while the other folds are used for training. Using this method, we ensure that the models are reliable, and they generalize properly. Fig. 6 shows the general form of the K-fold cross-validation technique [29].

Figure 6: Block diagram of k-fold cross-validation
All experiments were written in MATLAB R2021, they ran on a desktop computer that uses Windows 10 operating system, the computer used up to 16 GB of RAM, 500 GB Hard Disk Drive (HDD), and an Intel core i7–6700/3.4 GHz microprocessor. For training models, a 5-fold cross-validation methodology was employed with the RMSProp optimizer. The initial learning rate was 0.0001, the mini-batch size was 8, the max epochs was 20, and different models were trained for 5910 iterations per fold. The selection of these parameters were done based on many experiments and these were the best combinations.
The first part of Figs. 7–10 show the multiclass confusion matrix for all four inspected models. The columns represent the actual class, while the rows represent the predicted class. It is clear from the figures that all models successfully extracted the hidden features that are correlated with each class category. DarkNet19 managed to discriminate all four classes with a 93.6% accuracy, and the error represented by the False Positives and the False Negatives is nearly 6.4%. DenseNet201 managed to distinguish all four classes with a 93.8% accuracy, and the error represented by the False Positives and the False Negatives is nearly 6.2%. Xception managed to discriminate all four classes with a 94.8% accuracy, and the error represented by the False Positives and the False Negatives is nearly 6.2%. VGG16 managed to distinguish all four classes with a 94.5% accuracy, and the error represented by the False Positives and the False Negatives is nearly 5.5%. The first part of Figs. 11 and 12 show the confusion matrix for a collection of models particularly majority voting and feature ensemble methods. The majority voting managed to discriminate all four classes with a 96.69% accuracy, and the error represented by the False Positives and the False Negatives is nearly 3.3%. On the other hand, the featured ensemble managed to distinguish all four classes with a 97.79% accuracy, and the error represented by the False Positives and the False Negatives is nearly 2.2%. In the second part of Figs. 7–12, the Receiver Operating Characteristic (ROC) curve for all surveyed architectures is presented; the curve shows the balance between sensitivity and specificity. The curve in all figures is very close to the upper left corner, Area Under the Curve (AUC) of the Receiver Operating Characteristic curve (ROC) is nearly one (0.998) indicating high performance in discriminating between all four classes.

Figure 7: (A) DarkNet19 confusion matrix (B) Receiver Operating Characteristic (ROC) curve

Figure 8: (A) DenseNet201 confusion matrix (B) Receiver Operating Characteristic (ROC) curve

Figure 9: (A) Xception confusion matrix (B) Receiver Operating Characteristic (ROC) curve

Figure 10: (A) VGG16 confusion matrix (B) Receiver Operating Characteristic (ROC) curve

Figure 11: (A) Majority Voting confusion matrix (B) Receiver Operating Characteristic (ROC) curve

Figure 12: (A) Ensemble Features confusion matrix (B) Receiver Operating Characteristic (ROC) curve
Discriminating between the four lung diseases can be a challenging task even for the expert radiologist because they have a similar or comparable “footprint” on the chest radiograph. As shown in Tab. 3, the Xception model 94.78% accuracy was the highest among all individual models, DarkNet19 exhibited the weakest model performance in distinguishing between the four diseases with a 93.61% accuracy, it can be seen from the table that the performance of all models with respect to the accuracy metric is nearly equivalent. Sensitivity is an important metric in medical applications, it measures the ability of the test to accurately recognize patients with the illness. The Xception model also showed the highest sensitivity of 94.13%, while the worst sensitivity was that of the of the DenseNet201. Specificity on the other hand measures the ability of the test to correctly detect patients without the infection, VGG16 displayed the highest specificity of 97.896%, and DarkNet19 showed the lowest specificity of 97.288%. The Xception model showed the highest precision of 95.734%, F1 Score of 94.973, whereas the VGG16 precision was 93.631, F1 score was 93.876. Finally, as revealed by Tab. 3, the performance of a cluster of models far exceeded that of distinct models with respect to all performance metrics. Among all procedures used, the Ensemble Features technique achieved the highest overall accuracy of 97.79%, 3% higher than that of 94.491% achieved by the top individual Xception model.
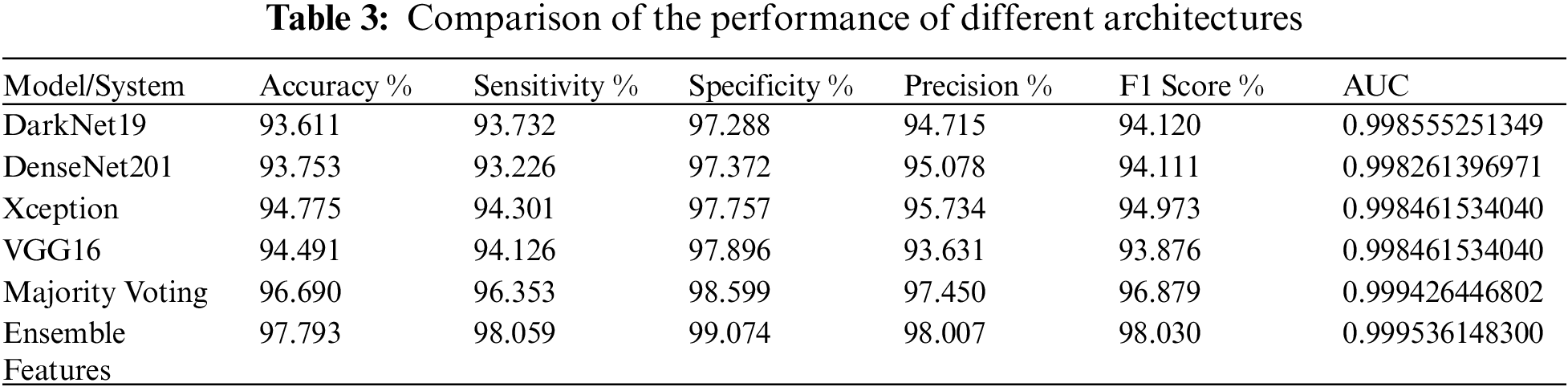
Overall, the highest accuracy of 97.793% is achieved by Ensemble Features, and the best prediction time of 0.33 s is achieved by DarkNet19. As shown in Tab. 4, among individual models, the worst average prediction time per image is displayed by the Xception model that takes 2.31 s whereas the DarkNet19 average prediction time per image is the best since it takes 0.33 s. Nonetheless, it is clear from the table that a collection of models takes more time than individual models in predicting the image, majority voting average prediction time per image is 1.12 s whereas ensemble features take 5.68 s. The performance of all methods in predicting the disease far exceeds that of the Traditional Polymerase Chain Reaction (PCR). The PCR is complex, time consuming, and can produce a lot of False Negatives. In such times, delays, and False Negatives (FN) can be problematic, telling a patient that he is not infected while him being infected can result in worsening of the public health conditions.

For distinct models, if accuracy is the goal, the authors recommend using the Xception model since it displayed the best accuracy of 94.775% among all individual models analyzed while DarkNet19 performance was the worst at 93.611%. As shown the difference in performance between the Xception and DarkNet19 models is in the range of 1.164% which is not substantial. Instead, we recommend using the DarkNet19 architecture because it gives a good tradeoff between accuracy, training time, lightweight, and average prediction time per image. Tab. 1 reveals that DarkNet19 uses only 19 layers meaning it is light, and does not take a lot of training time, it only takes 6.5 h to train per fold, it also exhibits the fastest average prediction time per image of 0.33 s. On the other hand, although the Xception model accuracy is slightly better by 1.164%, it is made of 71 convolutional layers (extra 52 layers), takes nearly 36 h to train per fold (extra 29.5 h), and average prediction time per image is 2.31 s (extra ∼2 s).
In summary, for individual models, opting for the DarkNet19 architecture seems to be more practical since it is overall performance seems to be better than that of the other probed models. DarkNet19 offers an acceptable accuracy, and a near real time prediction as measured by it is average prediction time per image. Alternatively, for a group of models, if resources, prediction time are not an issue, and the target is accuracy, the authors recommend using the Ensemble Features method since it displayed the highest accuracy among all approaches employed (4.182% higher than DarkNet19). Bear in mind that Ensemble Features average prediction time per image is 5.68 s (5.35 s worser than that of DarkNet19). In the future, a decision support system that utilizes either DarkNet19 or Ensemble Features can be developed to help the beginner radiologists in areas that lack resources particularly radiologist expertise.
The results in this study can be used to build a reliable and dependable system that can help novice radiologists in remote areas detect pulmonary diseases rapidly, and cost effectively. Such a system can help lessen interpretation variability and subjectivity thus reducing False Negatives (FN) and providing the accurate diagnosis with high accuracy.
Rapid and precise detection of pulmonary diseases is vital for the containment of the deteriorating health situation. Currently, the complex nature of the PCR and other tests makes it time consuming and susceptible to False Negatives (FN). In this study, we used transfer learning to compare the effectiveness of four popular pretrained models in distinguishing between four different pulmonary diseases normal, pneumonia, lung opacity and COVID-19 in chest radiographs. For individual models, the Xception architecture outperformed DenseNet201, VGG16, and DarkNet19 with a 94.78% accuracy. Alternatively, this research advocates the utilization of DarkNet19 because it is lightweight, it demonstrated a near real time detection capability (0.33 s), and acceptable training time. For a group of models, the 97.79% accuracy achieved by Ensemble Features was the highest among all surveyed methods, nonetheless it takes 5.68 s to predict an image. In the future, instead of building a proprietary model from scratch, an intelligent decision support system can be constructed using either DarkNet19 or Ensemble Features to assist radiologists in the field make quick decisions to distinguish all four lung diseases with high accuracy and few false negatives. Obtaining such an automated system for the classification of Chest-X-Ray (CXR) images is important for health care organizations. It will reduce the False Negatives (FN) rate and makes the accurate diagnosis a simple task. It will help the physicians and the clinics in COVID-19 situation and reduce the burden on the health care sector. This highly dependable software will save patients and physicians time and effort, it will reduce the mortality rate that originates from misdiagnosis of CXR images, in addition it will be a dependable software that can be deployed in rural regions that are lacking specialist’s expertise.
Acknowledgement: The authors would like to thank the anonymous reviewers for their valuable comments. Also, the authors would like to thank the authors of the used dataset to make their dataset available online.
Funding Statement: The authors received no specific funding for this study.
Conflicts of Interest: The authors declare that they have no conflicts of interest to report regarding the present study.
1. C. Li, C. Zhao, J. Bao, B. Tang, Y. Wang and B. Gu, “Laboratory diagnosis of coronavirus disease-2019 (COVID-19),” Clinica Chimica Acta; International Journal of Clinical Chemistry, vol. 51, pp. 35–46, 2020. [Google Scholar]
2. C. Huang, Y. Wang, X. Li, L. Ren, J. Zhao et al., “Clinical features of patients infected with 2019 novel coronavirus in wuhan, China,” The Lancet, vol. 395, pp. 497–506, 2020. [Google Scholar]
3. COVID-19Worldwide Statistics. 2021. [Online]. Available: https://www.worldometers.info/coronavirus/? (Accessed on 04 November 2021). [Google Scholar]
4. C. P. West, V. M. Montori and P. Sampathkumar, “COVID-19 testing: The threat of false-negative results,” Mayo Clinic Proceedings, vol. 95, no. 6, pp. 1127–1129, 2020. [Google Scholar]
5. G. Guyatt, D. Rennie, M. Meade and D. Cook, “Users’ guides to the medical literature: A manual for evidence-based clinical practice,” in IEEE Conf. on Computer Vision and Pattern Recognition, Chicago: AMA press, vol. 706, pp. 6517–6525, 2002. [Google Scholar]
6. N. Liu, L. Wan, Y. Zhang, T. Zhou, H. Huo et al., “Exploiting convolutional neural networks with deeply local description for remote sensing image classification,” IEEE Access, vol. 6, pp. 11215–11228, 2018. [Google Scholar]
7. Z. Y. Zu, M. D. Jiang, P. P. Xu, W. Chen, W. Ni, et al., “Coronavirus disease 2019 “(COVID-19A perspective from China,” Radiology, vol. 296, no. 2, pp. 15–25, pp. 2020. [Google Scholar]
8. Y. Yari, T. V. Nguyen and H. Nguyen, “Accuracy improvement in detection of COVID-19 in chest radiography,”. in 2020 14th Int. Conf. on Signal Processing and Communication Systems (ICSPCS), Adelaide Australia, pp. 1–6, 2020. [Google Scholar]
9. J. V. Pranav, R. Anand, T. Shanthi, K. Manju, S. Veni et al., “Detection and identification of COVID-19 based on chest medical image by using convolutional neural networks,” International Journal of Intelligent Networks, vol. 1, pp. 112–118, 2020. [Google Scholar]
10. I. U. Khan and N. Aslam, “A Deep-learning-based framework for automated diagnosis of COVID-19 using X-ray images,” Information, vol. 11, no. 9, pp. 419–432, 2020. [Google Scholar]
11. F. Lacruz and R. Vidarte, “Analysis of deep learning models for COVID-19 diagnosis from X-ray chest images,” Researchgate, 2020. [Google Scholar]
12. W. Zhang, B. Pogorelsky, M. Loveland and T. Wolf, “Classification of COVID-19 X-ray images using a combination of deep and handcrafted features,” arXiv preprint, arXiv:2101.07866, 2021. [Google Scholar]
13. M. Fontanellaz, L. Ebner, A. Huber, A. Peters, L. Löbelenz et al., “A Deep-learning diagnostic support system for the detection of COVID-19 using chest radiographs: A multireader validation study,” Investigative Radiology, vol. 56, no. 6, pp. 348–356, 2021. [Google Scholar]
14. O. N. Oyelade, A. E. Ezugwu and H. Chiroma, “Covframenet: An enhanced deep learning framework for COVID-19 detection,” IEEE Access, vol. 9, pp. 77905–77919, 2021. [Google Scholar]
15. T. D. Pham, “Classification of COVID-19 chest X-rays with deep learning: New models or fine tuning?” Health Information Science and Systems, vol. 9, no. 1, pp. 1–11, 2021. [Google Scholar]
16. H. Alquran, M. Alsleti, R. Alsharif, I. A. Qasmieh, A. M. Alqudah et al., “Employing texture features of chest X-ray images and machine learning in COVID-19 detection and classification,” Mendel, vol. 27, no. 1, pp. 9–17, 2021. [Google Scholar]
17. R. Alsharif, Y. Al-Issa, A. M. Alqudah, I. A. Qasmieh, W. A. Mustafa et al., “PneumoniaNet: Automated detection and classification of pediatric pneumonia using chest x-ray images and cnn approach,” Electronics, vol. 10, no. 23, pp. 2949–2962, 2021. [Google Scholar]
18. J. Deng, W. Dong, R. Socher, L. J. Li, K. Li et al., “Imagenet: A large-scale hierarchical image database,” in 2009 IEEE Conf. on Computer Vision and Pattern Recognition, Miami, FL, USA, pp. 248–255, 2009. [Google Scholar]
19. G. Huang, Z. Liu, L. Van Der Maaten and K. Q. Weinberger, “Densely connected convolutional networks,” in Proc. of the IEEE Conf. on Computer Vision and Pattern Recognition, Honolulu, Hawaii, USA, pp. 4700–4708, 2017. [Google Scholar]
20. F. Chollet, “Xception: Deep learning with depthwise separable convolutions,” in Proc. of the IEEE Conf. on Computer Vision and Pattern Recognition, Honolulu, Hawaii, USA, pp. 1251–1258, 2019. [Google Scholar]
21. J. Redmon and A. Farhadi, “YOLO9000: Better, faster, stronger,” in Proc. of the IEEE Conf. on Computer Vision and Pattern Recognition, Honolulu, Hawaii, USA, pp. 7263–727, 2017. [Google Scholar]
22. K. Simonyan and A. Zisserman, “Very deep convolutional networks for large-scale image recognition,” arXiv preprint, arXiv:1409.1556, 2014. [Google Scholar]
23. A. Alqudah and A. M. Alqudah, “Sliding window based deep ensemble system for breast cancer classification,” Journal of Medical Engineering and Technology, vol. 45, no. 4, pp. 313–323, 2021. [Google Scholar]
24. A. Alqudah and A. M. Alqudah, “Sliding window based support vector machine system for classification of breast cancer using histopathological microscopic images,” IETE Journal of Research, 2019. [Google Scholar]
25. A. M. Alqudah, S. Qazan, H. Alquran, I. A. Qasmieh and A. Alqudah, “Covid-19 detection from x-ray images using different artificial intelligence hybrid models,” Jordan Journal of Electrical Engineering, vol. 6, no. 2, pp. 168–178, 2020. [Google Scholar]
26. M. E. H. Chowdhury, T. Rahman, A. Khandakar, R. Mazhar, M. A. Kadir et al., “Can AI help in screening viral and COVID-19 pneumonia?,” IEEE Access, vol. 8, pp. 132665–132676, 2020. [Google Scholar]
27. T. Rahman, A. Khandakar, Y. Qiblawey, A. Tahir, S. Kiranyaz et al., “Exploring the effect of image enhancement techniques on COVID-19 detection using chest X-ray images,” Computers in Biology and Medicine, vol. 132, pp. 104319–104319, 2020. [Google Scholar]
28. A. M. Alqudah, “Towards classifying non-segmented heart sound records using instantaneous frequency based features,” Journal of Medical Engineering and Technology, vol. 43, no. 7, pp. 418–430, 2019. [Google Scholar]
29. T. T. Wong, “Performance evaluation of classification algorithms by k-fold and leave-one-out cross validation,” Pattern Recognition, vol. 48, no. 9, pp. 2839–2846, 2015. [Google Scholar]
 | This work is licensed under a Creative Commons Attribution 4.0 International License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. |