 Open Access
Open Access
ARTICLE
Trametinib boosts palbociclib’s efficacy in breast cancer via autophagy inhibition
1 Sichuan Key Medical Laboratory of New Drug Discovery and Druggability Evaluation, Luzhou Key Laboratory of Activity Screening and Druggability Evaluation for Chinese Materia Medica, Key Laboratory of Medical Electrophysiology of Ministry of Education, School of Pharmacy, Southwest Medical University, Luzhou, 646000, China
2 The Department of Pathology, The Affiliated Hospital of Southwest Medical University, Luzhou, 646000, China
3 The Department of Pathology, Mianyang Central Hospital, School of Medicine, University of Electronic Science and Technology of China, Mianyang, 621000, China
4 The Department of Pathology, Yongchuan Hospital of Chongqing Medical University, Chongqing, 402160, China
5 School of Basic Medical Sciences, Southwest Medical University, Luzhou, 646000, China
6 School of Clinical Medical Sciences, Southwest Medical University, Luzhou, 646000, China
* Corresponding Authors: JIANMING WU. Email: ; XIULI XIAO. Email:
(This article belongs to the Special Issue: Signaling Pathway Crosstalk in Malignant Tumors: Molecular Targets and Combinatorial Therapeutics)
Oncology Research 2024, 32(7), 1197-1207. https://doi.org/10.32604/or.2024.046139
Received 20 September 2023; Accepted 09 January 2024; Issue published 20 June 2024
Abstract
Breast cancer, a predominant global health issue, requires ongoing exploration of new therapeutic strategies. Palbociclib (PAL), a well-known cyclin-dependent kinase (CDK) inhibitor, plays a critical role in breast cancer treatment. While its efficacy is recognized, the interplay between PAL and cellular autophagy, particularly in the context of the RAF/MEK/ERK signaling pathway, remains insufficiently explored. This study investigates PAL’s inhibitory effects on breast cancer using both in vitro (MCF7 and MDA-MB-468 cells) and in vivo (tumor-bearing nude mice) models. Aimed at elucidating the impact of PAL on autophagic processes and exploring the potential of combining it with trametinib (TRA), an MEK inhibitor, our research seeks to address the challenge of PAL-induced drug resistance. Our findings reveal that PAL significantly decreases the viability of MCF7 and MDA-MB-468 cells and reduces tumor size in mice while showing minimal cytotoxicity in MCF10A cells. However, PAL also induces protective autophagy, potentially leading to drug resistance via the RAF/MEK/ERK pathway activation. Introducing TRA effectively neutralized this autophagy, enhancing PAL’s anti-tumor efficacy. A combination of PAL and TRA synergistically reduced cell viability and proliferation, and in vivo studies showed notable tumor size reduction. In conclusion, the PAL and TRA combination emerges as a promising strategy for overcoming PAL-induced resistance, offering a new horizon in breast cancer treatment.Graphic Abstract
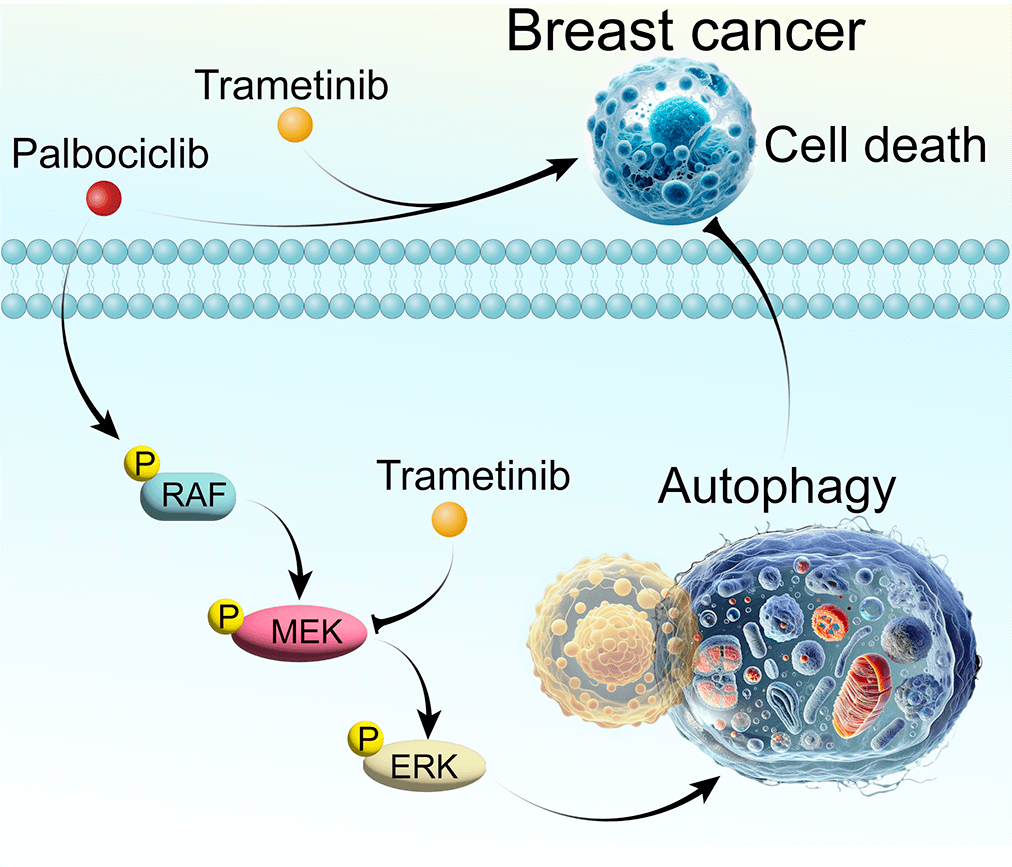
Keywords
Supplementary Material
Supplementary Material FileCite This Article
 Copyright © 2024 The Author(s). Published by Tech Science Press.
Copyright © 2024 The Author(s). Published by Tech Science Press.This work is licensed under a Creative Commons Attribution 4.0 International License , which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.


 Submit a Paper
Submit a Paper Propose a Special lssue
Propose a Special lssue View Full Text
View Full Text Download PDF
Download PDF Downloads
Downloads
 Citation Tools
Citation Tools