

 | Phyton-International Journal of Experimental Botany |  |
DOI: 10.32604/phyton.2022.016972
ARTICLE
First Report of Fusarium striatum Causing Root Rot Disease of Panax notoginseng in Yunnan, China
1College of Plant Protection, Yunnan Agricultural University, Kunming, 650201, China
2Key Laboratory of Agro-Biodiversity and Pest Management of Education Ministry of China, Yunnan Agricultural University, Kunming, 650201, China
3State Key Laboratory for Conservation and Utilization of Bio-Resources in Yunnan, Yunnan Agricultural University, Kunming, 650201, China
*Corresponding Authors: Xiahong He. Email: hexiahong@googlemail.com; Yunlong Du. Email: yunlongdu@aliyun.com
#These authors have contributed equally to this work
Received: 15 April 2021; Accepted: 21 June 2021
Abstract: Panax notoginseng is a traditional Chinese medicinal plant. Root rot of P. notoginseng is one of the most serious diseases affecting P. notoginseng growth and causes wilted leaves, fewer lateral roots and rotten roots. Root rot is a soil-borne disease, and mainly occurs from June to August in Yunnan Province when the temperatures are high and the air is humid. In this study, the endophytic fungal genus Fusarium isolate E-2018.1.22-#3.2 was obtained from a P. notoginseng embryo. Fusarium isolate E-2018.1.22-#3.2 was identified as Fusarium striatum based on morphological characteristics and molecular analysis. The fungus was found to have conidiophores and macroconidia, and its ITS, LSU and TEF-1α genes shared 100%, 99.2% and 99% identities with the homologous genes of Fusarium striatum, respectively. Isolate F. striatum E-2018.1.22-#3.2 can cause root rot symptoms, including black, soft roots, fewer lateral roots and leaf wilt, in 93% of the experimental P. notoginseng plants, and could be re-isolated, fulfilling Koch’s postulates. When the P. notoginseng plants were treated with the fungicide pyraclostrobin, isolate F. striatum E-2018.1.22-#3.2 was unable to cause root rot. We have therefore demonstrated that F. striatum E-2018.1.22-#3.2 is able to cause root rot disease in P. notoginseng. This is the first report of root rot disease caused by F. striatum on P. notoginseng in China.
Keywords: Panax notoginseng; Fusarium striatum; root rot disease; endophytic fungus
Panax notoginseng (Burk.) F. H. Chen belongs to the family Araliaceae. The roots of P. notoginseng are used as traditional medicine in China, and dried flower buds of P. notoginseng also function as a medicine or tea [1]. Root rot caused by pathogenic fungi is a serious disease of P. notoginseng, and affects growth [2]. The P. notoginseng seedlings grow in shade, with soft, permeable soil. However, the weather in summer is hot, humid and rainy in most of the regions in Yunnan Province where P. notoginseng is cultivated, which include Wenshan and Kunming. These conditions are ideal for the growth of the pathogenic fungus responsible for root rot, and the disease normally occurs in P. notoginseng between June and August and seriously reduces the yield of this plant species in Yunnan. The P. notoginseng seedlings with root rot symptom show wilted leaves, few lateral roots and rotten roots [3]. The root rot disease in P. notoginseng is known to be caused by fungi of the genus Fusarium, specifically F. oxysporum and F. solani [4]. The jasmonic acid (JA) and ethylene signaling pathways may act synergistically to positively regulate the defence responses of P. notoginseng inoculated with F. solani [5]. Transient expression of the PnSN1 gene has been shown to enhance tobacco resistance to F. solani [6]. Where F. oxysporum has caused root rot in P. notoginseng, its growth can be antagonized by fungi from the family Herpotrichiellaceae, as well as Coniosporium [7]. The methanol extract from Artemisia annua leaves, which major ingredients were camphor, camphene, β-caryophyllene, and germacrene D, shows strong antifungal effects on the growth of F. oxysporum and F. solani [8]. The fungus Fusarium striatum has been reported to cause stem rot disease in tomato [9], however, the role of F. striatum remains unclear in root rot in P. notoginseng. In this study, an endophytic fungus F. striatum E-2018.1.22-#3.2 was isolated from a P. notoginseng embryo. Our results demonstrate that the endophytic fungus F. striatum E-2018.1.22-#3.2 can cause root rot disease on a new host P. notoginseng in Yunnan Province, China.
2.1 Plant Materials and Pathogen Isolation
Seeds of P. notoginseng were collected from Wenshan (104°16’34’’E, 23°17’4’’N), Yunnan, China, in December 2017. The P. notoginseng seeds were surface sterilized with 70% ethanol for 1.5 min, followed by a 2% sodium hypochlorite solution for 17 min, and were then washed five times with sterilized water. Embryos that developed from the P. notoginseng seeds were cultured on Murashige-Skoog (MS) medium at 28°C. Fungal colonies developed on the medium, and a single fungal spore was selected from these colonies, picked out and cultured on potato dextrose agar (PDA) medium at 28°C in the dark for 7 days. The fungal morphology was observed using a light microscope.
2.2 Gene Sequences and Phylogeny Analysis of Fungus Isolates
Fungal genomic DNA was isolated using the cetyltrimethylammonium bromide (CTAB) method. To clone the internal transcribed spacer (ITS) sequence and nuclear large subunit (LSU) of the ribosomal DNA, we referred to the previously published report [10,11]. Meanwhile, the transcription elongation factor (TEF-1α) gene was cloned using the primer pair TEF-F/TEF-R also following previous research [12]. The PCR products of the ITS, LSU and TEF-1α genes were sequenced at the Tsingke Company. Molecular Evolutionary Genetics Analysis version 5.1 (MEGA 5.1) was used to construct a phylogenetic tree using the TEF-1α gene sequences.
2.3 Determination of Pathogenicity
A pathogenicity assay was conducted following previous work [3]. The fungus was grown on PDA medium at 28°C in the dark for 30 days. Afterwards, the fungus was rinsed into sterilized water to reach a conidial suspension of 1 × 106 spores/mL. A volume of 100 mL conidial suspension was inoculated onto a healthy, two-year-old P. notoginseng seedlings (n = 21). The seedlings showed root rot symptoms after inoculation with the fungal spores for 45 days, and the rotten roots were collected to fulfil Koch’s postulates.
2.4 Fungicide Treatment of P. notoginseng
In order to find out whether treatment of the P. notoginseng seedlings with fungicide would prevent the disease after inoculation with the isolated fungal strain, we treated the seedlings with the fungicide pyraclostrobin as described previously [13], with few a minor modifications. Briefly, the roots of two-year-old P. notoginseng seedlings (n = 3) were washed with water, surface disinfected with 75% alcohol for 3 min, and then washed three times with sterilized water. The roots were grown on wet filter paper in a glass culture dish of 15 cm diameter; they were then pricked with a needle every 0.5 cm and sprayed with a fungal solution (2.14 * 106~4.7 * 106 spores/mL) one time every day, together with or without fungicide (25% pyraclostrobin in a suspension agent, CAS No. 175013-18-0, Hebei Chengyue Chemical Co., Ltd., China) at 1,000 times dilution. The roots were totally treated for 3 times. The roots (n = 3) treated with sterilized water or pyraclostrobin acted as a control. The root phenotypes were observed after 6 days of treatment with fungicide and fungal spores.
3.1 Morphological Characteristics of Isolate E-2018.1.22-#3.2
An endophytic fungus was isolated from the P. notoginseng embryos cultured on Murashige-Skoog (MS) medium. The fungus (Fig. 1A), named E-2018.1.22-#3.2, had a grey mycelium and grew fast; the grey-white colony reached 5.7–6.7 cm in diameter on PDA medium after 7 days (Figs. 1B and 1C).
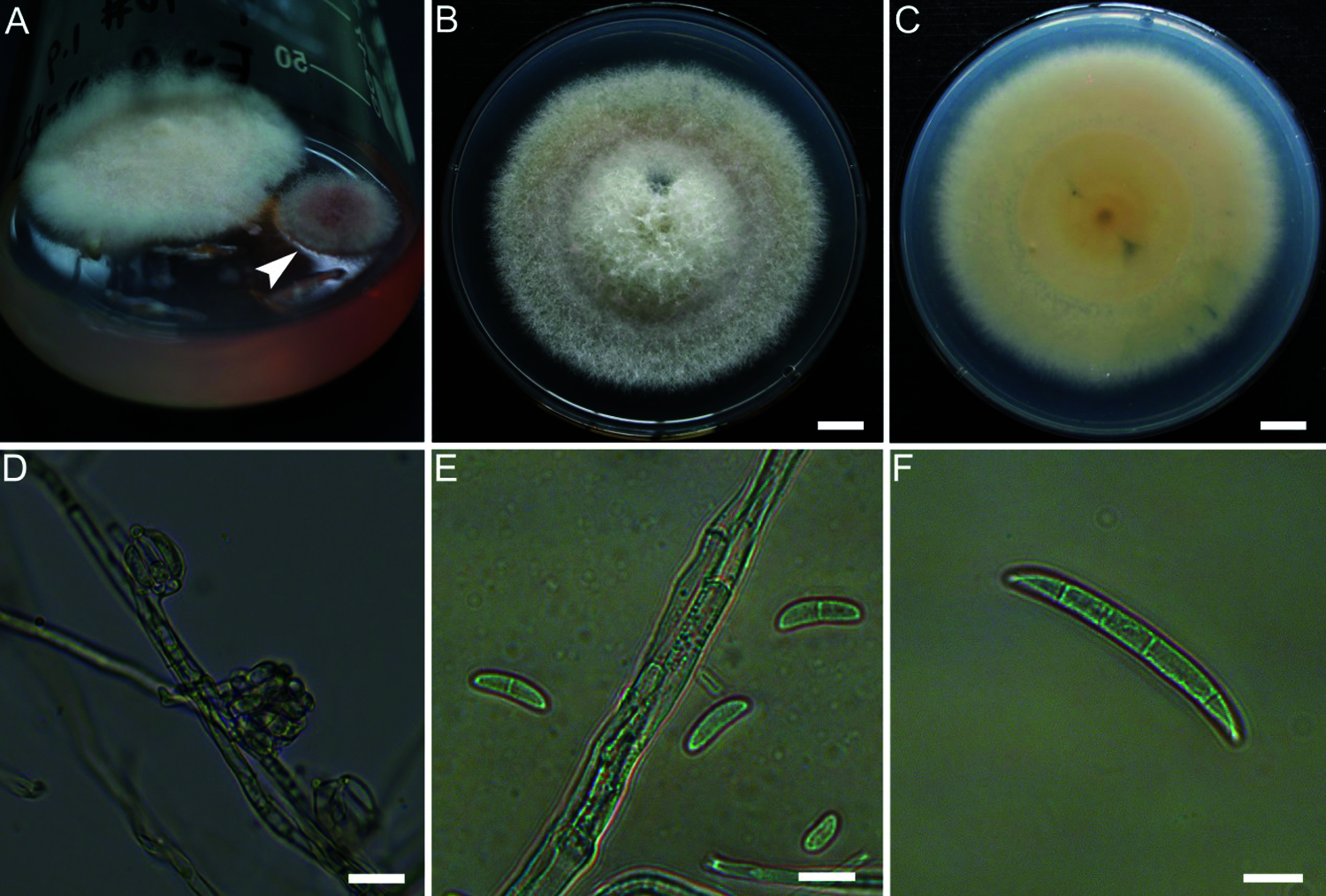
Figure 1: Morphology of the fungus isolate E-2018.1.22-#3.2. The fungi isolated from the P. notoginseng embryos (A). Morphologies of colony (B, C), microconidia (D, E) and macroconidia (F) of isolate E-2018.1.22-#3.2. Bar = 1 cm (B, C), or 10 μm (D–F). The arrowhead points to the isolate E-2018.1.22-#3.2
The hyphae were white and filamentous. The conidia were oval-, kidney- and sickle-shaped with 1 to 4 septa, and their length and width were 7.2 to 17.2 × 2.7 to 5.2 μm (mean ± SD = 12.2 ± 3.07 × 3.8 ± 0.74 μm, n = 14) (Figs. 1D–1F). After the isolate E-2018.1.22-#3.2 had been grown on PDA medium for 30 days, it produced typical conidiophores and macroconidia (Figs. 1D–1F) of the genus Fusarium [9].
3.2 Characterization of Gene Sequences and Phylogenetic Tree
To identify the isolate E-2018.1.22-#3.2, the sequences of the ITS (Genbank No. MK372369), LSU (Genbank No. MK463991) and TEF-1α (Genbank No. MW553277) genes were analyzed. The ITS, LSU and TEF-1α genes from isolate E-2018.1.22-#3.2 shared 100%, 99.2% and 99% identities, respectively, with the ITS (Genbank No. KY436003), LSU (Genbank No. MH876354) and TEF-1α (Genbank No. MF344651) genes from Fusarium striatum. The phylogenetic tree constructed from the sequenced TEF-1α genes from the species in the genus Fusarium showed isolate E-2018.1.22-#3.2 grouping together with F. striatum (Fig. 2).
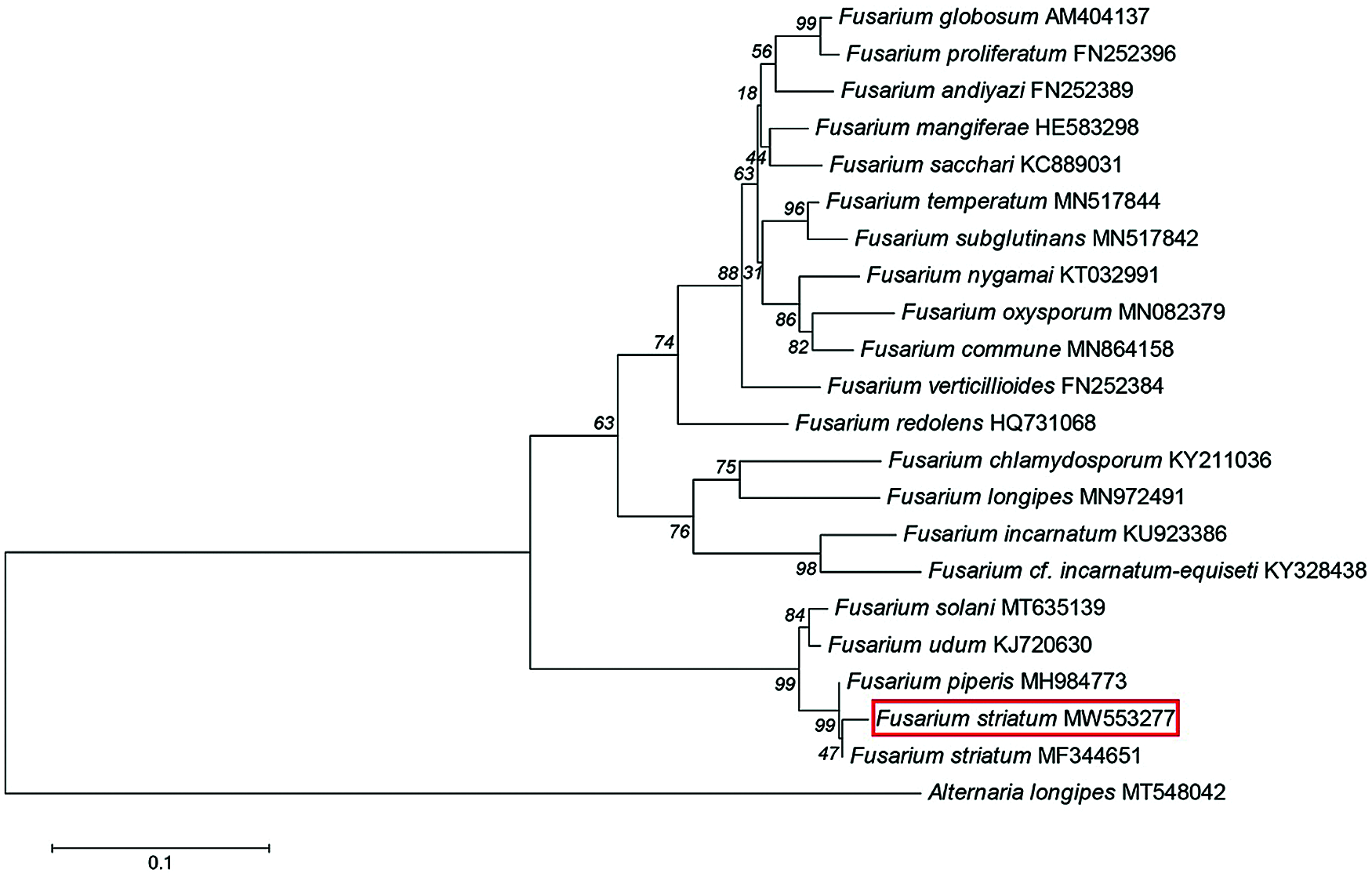
Figure 2: Phylogenetic tree of the genus Fusarium isolate E-2018.1.22-#3.2. A Phylogenetic tree was reconstructed from the sequences of the TEF-1α genes from the genus Fusarium, based on a neighbour-joining analysis. The red box indicates isolate E-2018.1.22-#3.2. The fungus Alternaria longipes was used as an out-group. The percentage values at the nodes display the accuracy of the dendrogram clades. Scale = 0.1
Analysis of morphological characteristics (Figs. 1D–1F) suggested that isolate E-2018.1.22-#3.2 belonged to the genus Fusarium, and the sequence analysis also confirmed this. Therefore, the isolate E-2018.1.22-#3.2 was identified as F. striatum based on morphological features and sequence analysis. Isolate E-2018.1.22-#3.2 was then deposited at the China General Microbiological Culture Collection Center (CGMCC 3.19871).
3.3 Fusarium striatum E-2018.1.22-#3.2 Causes Root rot Disease in P. notoginseng
The P. notoginseng seedlings were inoculated with either a conidial suspension from isolate E-2018.1.22-#3.2, or with water (Figs. 3A and 3B), After 45 days, compared with the seedlings treated with water (Fig. 3C), 93% of P. notoginseng seedlings inoculated with the conidial suspension showed symptoms of root rot (Fig. 3D), including soft, black roots, few lateral roots, and wilted leaves. However, none of the seedlings treated with sterilized water showed these symptoms. F. striatum E-2018.1.22-#3.2 could be subsequently re-isolated from the rotten roots to fufil Koch’s postulates (Figs. 3E and 3F).
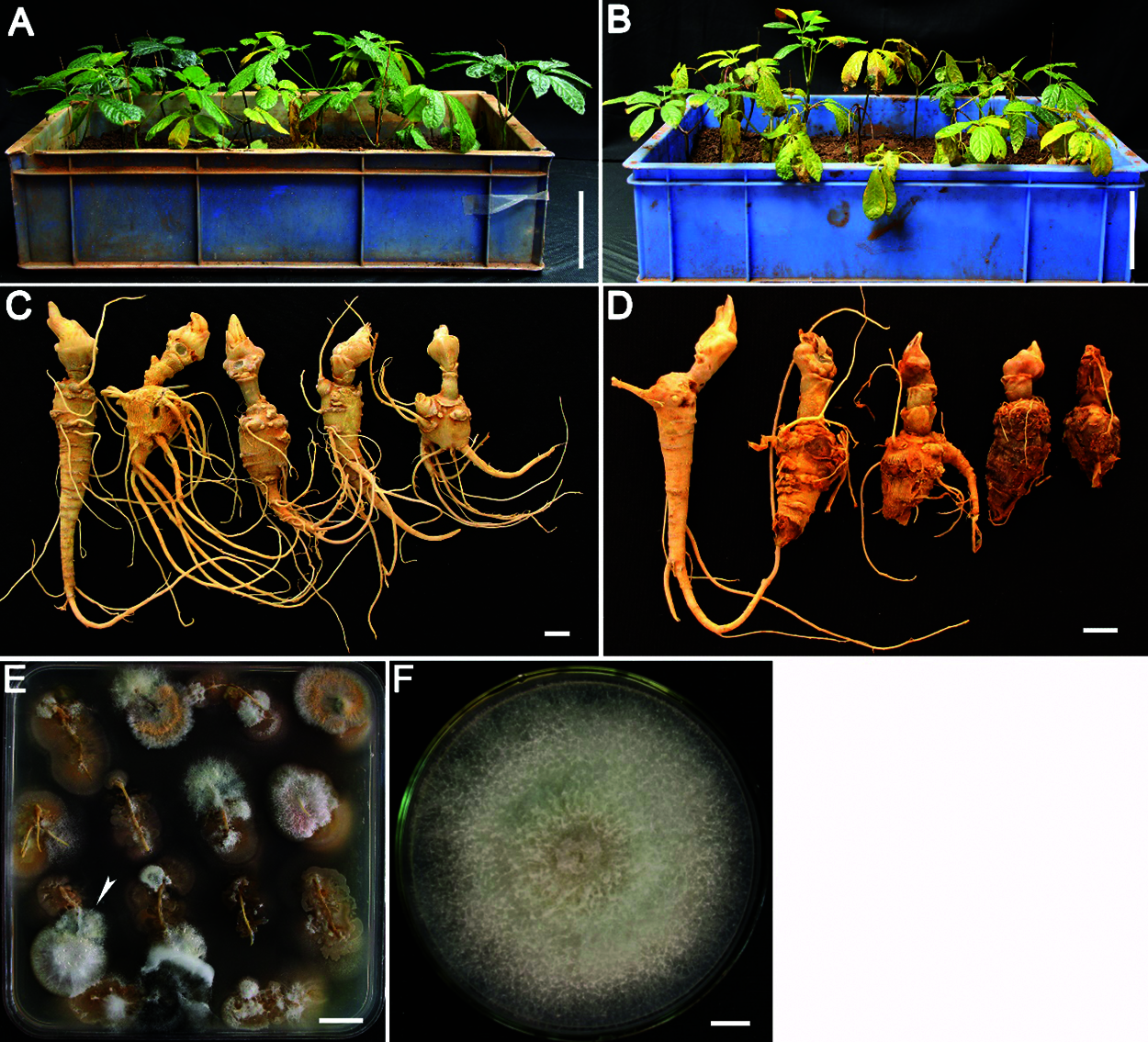
Figure 3: Phenotype of P. notoginseng seedlings inoculated with isolate E-2018.1.22-#3.2. The P. notoginseng seedlings were treated with sterilized water (A) or with isolate E-2018.1.22-#3.2 (B) for 45 days. Root phenotypes of the P. notoginseng seedlings treated with sterilized water (C) and isolate E-2018.1.22-#3.2 (D). A re-isolated colony from the rotten roots was grown on PDA medium for 4 days (E). Fusarium striatum E-2018.1.22-#3.2 was re-isolated from the rotten roots (E) and grown on PDA medium for 10 days (F). The arrowhead points to the colony from which isolate E-2018.1.22-#3.2 was taken and grown in image F. Bar = 10 cm (A, B), 1 cm (C, D, F), 2 cm (E)
To further test that the F. striatum can cause root rot in P. notoginseng, the roots of P. notoginseng seedlings were inoculated with F. striatum spores and co-treated with the fungicide pyraclostrobin (Fig. 4). Compared with the control treated with water (Figs. 4A and 4E), the roots treated either with pyraclostrobin alone (Figs. 4B and 4F) or co-treated with pyraclostrobin together with F. striatum E-2018.1.22-#3.2 (Figs. 4C and 4G) retained normal growth phenotypes. However, the roots inoculated only with F. striatum E-2018.1.22-#3.2 showed obvious root rot symptoms, including fewer lateral roots, and soft and rotten roots (Figs. 4D and 4H). These results demonstrate that F. striatum is able to cause root rot disease in P. notoginseng.

Figure 4: Phenotype of P. notoginseng roots treated with isolate E-2018.1.22-#3.2 and pyraclostrobin. Two-year-old P. notoginseng seedling roots treated with sterilized water (A, E), pyraclostrobin (B, F), isolate E-2018.1.22-#3.2 and pyraclostrobin (C, G) and isolate E-2018.1.22-#3.2 (D, H) for 6 days. Images E, F, G and H show the transverse phenotypes of the roots shown in images A, B, C and D, respectively. Bar = 2 cm
The genus Fusarium is an important pathogenic fungus and can cause rot disease [14–16] and wilt disease [17,18] in different host plants. Previous report shows that the F. striatum is also able to cause stem rot disease in tomato [9]. In this study, we found that the F. striatum E-2018.1.22-#3.2 was able to cause root rot disease in P. notoginseng in Yunnan, China. Interestingly, the F. striatum E-2018.1.22-#3.2 was isolated from the embryos of P. notoginseng seedlings and could be an endophytic fungus.
The P. notoginseng seedlings contain various endophytic fungi in different tissues [19], and the role of some endophytic fungi, such as Penicillium crustosum, are beneficial to P. notoginseng and can provide some protection against root rot disease [19]. However, certain endophytic fungi identified from P. notoginseng were pathogenic fungi, including some from the genus Alternaria and Fusarium [19]. Plant-endophyte interactions have been extensively investigated. Most endophytes are interchangeable [20]. Endophytic asymptomatic infection of plants may derive from common plant pathogens, which can asymptomatically colonize host plants and function as nonpathogens in certain lifestyles [21]. In this study, the F. striatum E-2018.1.22-#3.2 was acting as an endophytic fungus in P. notoginseng embryos and was able to cause root rot disease in P. notoginseng. It is behaves similary to the fungus Verticillium dahlia, which can colonize in plants as either an endophyte or a pathogenic fungus [22]. This implies that F. striatum E-2018.1.22-#3.2 has functional diversity during P. notoginseng development.
Root rot disease is one of the most serious diseases in P. notoginseng growth, and it cannot be effectively prevented in cultivation. F. striatum strain E-2018.1.22-#3.2 was able to colonize in the P. notoginseng embryo during its life cycle, suggesting that such strain is a seedborne pathogen. The colonization of P. notoginseng seeds by pathogens is a primary way by which the root rot disease can be transmitted. It reminds us that certain hygienic measures, including blocking of the pathogen spread and colonization of seeds, should be taken into account in P. notoginseng cultivation.
Acknowledgement: We thank Li Han and Qijing Fu (Yunnan Agricultural University) for their support of this article.
Funding Statement: This work was supported by grants from the National Natural Science Foundation of China (Grant Nos. 31660501 and 31860064), the Major Special Program for Scientific Research, Education Department of Yunnan Province (Grant No. ZD2015005), the Major Science and Technique Programs in Yunnan Province (Grant No. 2016ZF001), and the Earmarked Fund for Modern Agro-industry Technology Research System (Grant No. CARS-21).
Conflicts of Interest: The authors declare that they have no conflicts of interest to report regarding the present study.
1. Zhang, S., Chen, C., Lu, W. (2018). Phytochemistry, pharmacology, and clinical use of Panax notoginseng flowers buds. Phytotherapy Research: PTR, 32(11), 2155–2163. DOI 10.1002/ptr.6167. [Google Scholar] [CrossRef]
2. Sun, W., Ma, Y., Yin, Y., Chen, C., Cheng, Y. (2018). Effects of essential oils from zingiberaceae plants on root-rot disease of Panax notoginseng. Molecules, 23(5), 1021. DOI 10.3390/molecules23051021. [Google Scholar] [CrossRef]
3. Han, L., Zhou, X., Zhao, Y., Wu, L., Ping, X. et al. (2020). First report of Plectosphaerella plurivora causing root rot disease in Panax notoginseng in China. Journal of Phytopathology, 168(7–8), 375–379. DOI 10.1111/jph.12901. [Google Scholar] [CrossRef]
4. Ma, Y., Chen, C., Li, Q., Xu, F., Cheng, Y. et al. (2019). Monitoring antifungal agents of Artemisia annua against Fusarium oxysporum and Fusarium solani, associated with Panax notoginseng root-rot disease. Molecules, 24(1), 213. DOI 10.3390/molecules24010213. [Google Scholar] [CrossRef]
5. Liu, D., Zhao, Q., Cui, X., Chen, R., Li, X. et al. (2019). A transcriptome analysis uncovers Panax notoginseng resistance to Fusarium solani induced by methyl jasmonate. Genes & Genomics, 41, 1383–1396. DOI 10.1007/s13258-019-00865-z. [Google Scholar] [CrossRef]
6. Qiu, B., Zhang, Y., Wang, Q., Wang, Z., Liu, D. (2020). Panax notoginseng snakin gene increases resistance to Fusarium solani in transgenic tobacco. Industrial Crops and Products, 157, 112902. DOI 10.1016/j.indcrop.2020.112902. [Google Scholar] [CrossRef]
7. Dong, L., Xu, J., Feng, G., Li, X., Chen, S. (2016). Soil bacterial and fungal community dynamics in relation to Panax notoginseng death rate in a continuous cropping system. Scientific Reports, 6(1), 31802. DOI 10.1038/srep31802. [Google Scholar] [CrossRef]
8. Ma, Y. N., Chen, C. J., Li, Q. Q., Xu, F. R., Cheng, Y. X. et al. (2019). Monitoring antifungal agents of artemisia annua against Fusarium oxysporum and Fusarium solani, associated with Panax notoginseng root-rot disease. Molecules, 24(1), 213. DOI 10.3390/molecules24010213. [Google Scholar] [CrossRef]
9. Moine, L. M., Labbé, C., Louis-Seize, G., Seifert, K. A., Bélanger, R. R. (2014). Identification and detection of Fusarium striatum as a new record of pathogen to greenhouse tomato in northeastern america. Plant Disease, 98, 292–298. DOI 10.1094/PDIS-08-13-0844-RE. [Google Scholar] [CrossRef]
10. White, T. J., Bruns, T., Lee, S., Taylor, J. (1990). Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. in Innis M. A., Gelfand D. H., Sninsky J. J., White T. J. (eds.PCR protocols: A guide to methods and applications, pp. 315–322. San Diego, Academic Press. [Google Scholar]
11. Giraldo, A., Gene, J., Cano, J., de Hoog, S., Guarro, J. (2012). Two new species of Acremonium from Spanish soils. Mycologia, 104, 1456–1465. DOI 10.3852/11-402. [Google Scholar] [CrossRef]
12. Carbone, I., Kohn, L. (1999). A method for designing primer sets for speciation studies in filamentous ascomycetes. Mycologia, 91(3), 553–556. DOI 10.1080/00275514.1999.12061051. [Google Scholar] [CrossRef]
13. Mi, C., Yang, R., Rao, J., Yang, S., Hu, X. (2017). Unveiling of dominant fungal pathogens associated with rusty root rot of Panax notoginseng based on multiple methods. Plant Disease, 101(12), 2046–2052. DOI 10.1094/PDIS-01-17-0135-RE. [Google Scholar] [CrossRef]
14. Alabouvette, C., Lemanceau, P., Steinberg, C. (1993). Recent advances in the biological control of Fusarium wilts. Pest Management Science, 37(4), 365–373. DOI 10.1002/ps.2780370409. [Google Scholar] [CrossRef]
15. Ibrahim, N. F., Mohd, M. H., Mohamed Nor, N. M. I., Zakaria, L. (2017). Characterization of Fusarium spp. associated with pineapple fruit rot and leaf spot in peninsular Malaysia. Journal of Phytopathology, 165(11–12), 718–726. DOI 10.1111/jph.12611. [Google Scholar] [CrossRef]
16. Esfahani, M. N. (2018). Genetic and virulence variation in Fusarium oxysporum f. sp. cepae causing root and basal rot of common onion in Iran. Journal of Phytopathology, 166(7–8), 572–580. DOI 10.1111/jph.12720. [Google Scholar] [CrossRef]
17. Dubey, S. C., Suresh, M., Singh, B. (2007). Evaluation of trichoderma species against Fusarium oxysporum f. sp. ciceris for integrated management of chickpea wilt. Biological Control, 40(1), 118–127. DOI 10.1016/j.biocontrol.2006.06.006. [Google Scholar] [CrossRef]
18. Rojo, F. G., Reynoso, M. M., Ferez, M., Chulze, S. N., Torres, A. M. (2007). Biological control by trichoderma species of Fusarium solani causing peanut brown root rot under field conditions. Crop Protection, 26(4), 549–555. DOI 10.1016/j.cropro.2006.05.006. [Google Scholar] [CrossRef]
19. Zheng, Y., Miao, C., Chen, H., Huang, F., Xia, Y. et al. (2017). Endophytic fungi harbored in Panax notoginseng: diversity and potential as biological control agents against host plant pathogens of root-rot disease. Journal of Ginseng Research, 41(3), 353–360. DOI 10.1016/j.jgr.2016.07.005. [Google Scholar] [CrossRef]
20. Álvarez-Loayza, P., White, J. F., Torres, M. S., Balslev, H., Kristiansen, T. et al. (2011). Light converts endosymbiotic fungus to pathogen, influencing seedling survival and niche-space filling of a common tropical tree, iriartea deltoidea. PLoS One, 6(1), e16386. DOI 10.1371/journal.pone.0016386. [Google Scholar] [CrossRef]
21. Lana, T. G., Azevedo, J. L., Pomella, A. W. V., Monteiro, R. T. R., Silva, C. B. et al. (2011). Endophytic and pathogenic isolates of the cacao fungal pathogen Moniliophthora perniciosa (tricholomataceae) are indistinguishable based on genetic and physiological analysis. Genetics and Molecular Research, 10, 326–334. DOI 10.4238/vol10-1gmr895. [Google Scholar] [CrossRef]
22. Wheeler, D. L., Dung, J. K. S., Johnson, D. A. (2018). From pathogen to endophyte: an endophytic population of Verticillium dahliae evolved from a sympatric pathogenic population. New Phytologist, 222(1), 497–510. DOI 10.1111/nph.15567. [Google Scholar]
 | This work is licensed under a Creative Commons Attribution 4.0 International License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. |