 Open Access
Open Access
ARTICLE
FedPA: Federated Learning with Performance-Based Averaging for Efficient Medical Image Classification
1 Faculty of Data Science and Information Technology, INTI International Univeristy, Nilai, 71800, Malaysia
2 Prince Sattam bin Abdulaziz University, Al-Kharj, 16278, Saudi Arabia
3 Department of Computer Engineering and Information, College of Engineering in Wadi Alddawasir, Prince Sattam bin Abdulaziz University, Al-Kharj, 16273, Saudi Arabia
4 Faculty of Arts, Society and Professional Studies, Newman University, Birmingham, B32 3NU, UK
* Corresponding Authors: Atif Mahmood. Email: ,
Computer Modeling in Engineering & Sciences 2026, 146(3), 39 https://doi.org/10.32604/cmes.2025.073501
Received 19 September 2025; Accepted 12 December 2025; Issue published 30 March 2026
Abstract
Federated learning is a decentralized model training paradigm with significant potential. However, the quality of Federated Network’s client updates can vary due to non-IID data distributions, leading to suboptimal global models. To address this issue, we propose a novel client selection strategy called FedPA (Performance-Based Federated Averaging). This proposed model selectively aggregates client updates based on a predefined performance threshold. Only clients whose local models achieve an F1 score of 70% or higher after training are included in the aggregation process. Clients below this threshold receive the updated global model but do not contribute their parameters. In this way, the low-performance clients are still in the process of learning and, after some rounds, will be able to contribute. If no client meets the performance threshold in a given round, the system falls back to standard FedAvg aggregation. This ensures the global model continues to improve even when most clients perform poorly. We evaluate FedPA on a subset of the MURA dataset for abnormality detection in radiographs of four bone types. Compared to baseline federated learning algorithms such as Federated Averaging (FedAvg), Federated Proximal (FedProx), Federated Stochastic Gradient Descent (FedSGD), and Federated Batch Normalization (FedBN), FedPA consistently ranks first or second across key performance metrics, particularly in accuracy, F1 score, and recall. Moreover, FedPA demonstrates notable efficiency, achieving the lowest average round time (Keywords
Medical imaging has surfaced as a cardinal approach in modern-day healthcare, providing the roots for meticulous diagnosis, effectual monitoring, and targeted treatment of miscellaneous pathological conditions [1]. Computed tomography (CT), ultrasound, magnetic resonance imaging (MRI) and X-rays integrated with intelligent analysis are the imaging solutions that impart in-depth evaluation of the human body. This is intensifying the ability of healthcare professionals to spot early stage anomalies with greater accuracy and operational excellence [2]. Medical image analysis has seen a boom as deep learning and artificial intelligence (AI) [3] show substantial growth enhancing precision, speed and proficiency in diagnosing various clinical abnormalities namely cancer [4], pneumonia [5], neurological disorders [6] and cardiovascular ailments [7]. These AI-driven modalities lessen the strain on radiologists streamlining automated detection, segmentation, and classification of medical images and improving patient clinical outcomes [8,9].
Training deep learning models for medical imaging requires vast amounts of high-quality, annotated medical data to ensure accurate and reliable predictions [10]. Improving annotation quality enhances the performance of these models for medical image segmentation. Distributing crowdsourced annotations across more images is often more efficient than collecting multiple annotations per image [11]. Data privacy and compliance regulations lead to substantial glitches in the acquisition and handling of medical imaging datasets for AI tasks. Researchers have projected numerous strategies to tackle these concerns. Federated learning [12] is one such approach that assists reciprocal model training exclusive of revealing the raw data, preserving patient privacy (See Fig. 1) [13]. Synthetic data generation offers a promising approach to augment and anonymise real imaging data, though it requires careful consideration of realism, diversity, and ethical implications [14]. Privacy-preserving techniques such as differential privacy and encrypted computation can further protect sensitive information [15]. Federated averaging (FedAvg) is a common aggregation procedure in federated contexts. However, it experiences convergence issues, especially when there is significant diversity in the data distributions among clients [16].
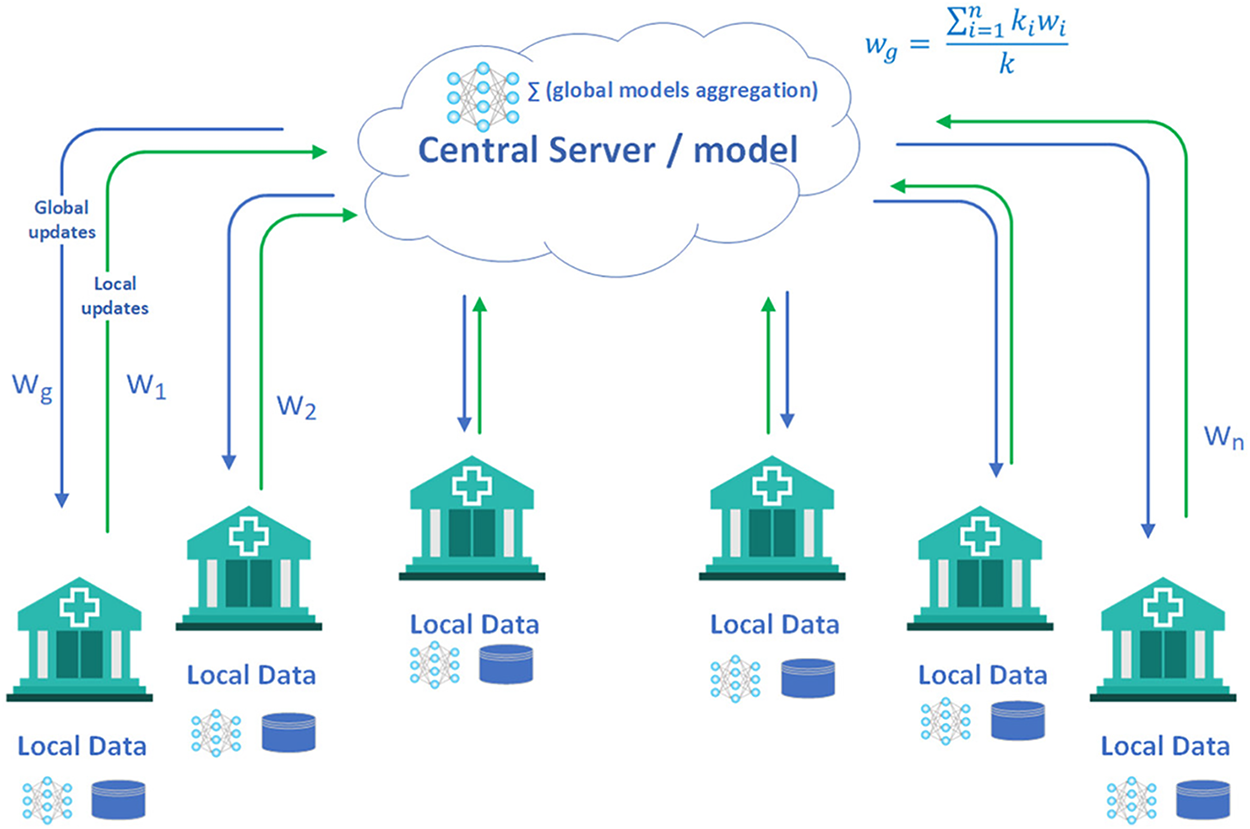
Figure 1: Federated learning in healthcare [17]
Diverse aggregation approaches have been proposed in recent study to enrich federated learning. Among all, Federated Averaging (FedAvg) is one such technique in the lead taken up for its minimalism and operational efficiency [18]. FedAvg counts on uniform participation from each client, making it susceptible to delays or unreliability amid contributors. In practical federated networks, clients may not be available all the time owing to network failures, power constraints, or computational limits. Such divergence may induce encounters like oscillating universal updates, as elevated client dropout rates can make model updates erratic and reduce overall productivity. It also stems into ineffective optimization as the server deficits influence over client participation, hampering the ability to restructure the learning process. Moreover, model performance may deteriorate when optimal clients are missing during particular rounds, yielding lower-quality updates.
An array of advanced aggregation techniques has been put forward to tackle these challenges. FedAvg is enhanced with Proximal Optimization (FedProx) [19] by minimizing the detrimental impacts of client data non-uniformity. However, its additional hyperparameter is strenuous to tune and its default values may not generalize well amongst different datasets. Federated Multi-Task Learning via Multi-Task Association (FedMA) [20] boosts global and local models conjointly, refining convergence; however, its density makes its execution challenging in some federated learning systems. Quantization-based Federated Learning (QFFL) [21] cuts down communication expenses by squeezing model updates through quantization, but it incurs quantization errors potentially impairing convergence and final model accuracy.
Regardless of notable progress, federated learning (FL) continues to encounter critical challenges that limit its effectiveness in real-world medical imaging applications [22]. Persistent issues such as high communication overhead, data heterogeneity, inefficient client participation, and unresolved privacy and security vulnerabilities undermine its practical deployment. Standard aggregation strategies, including Federated Averaging (FedAvg) and Federated Stochastic Gradient Descent (FedSGD) [23], rely on the assumption of equal client contributions. This assumption rarely holds in heterogeneous environments, where imbalanced data distributions and unequal computational capacities across clients often result in biased, unstable, or suboptimal global model updates [24,25].
In the healthcare domain, these limitations become even more pronounced, as FL must collaboratively train diagnostic models across diverse hospital datasets while safeguarding privacy and ensuring scalability. Variations in patient demographics, imaging modalities, and institutional protocols generate highly non-IID data distributions that hinder model generalization. At the same time, frequent client-server communication places heavy demands on network bandwidth, which is often constrained in hospital environments. Noisy updates from clients with poor-quality data or limited computational capacity further compromise stability and slow convergence. While the wide disparity in hardware resources across hospitals makes uniform deployment challenging. Compounding these issues, FL remains susceptible to privacy and security threats such as model inversion, adversarial data poisoning, unreliable client updates, robust aggregation and client-selection mechanisms [26].
In response to these challenges, we advocate Performance-Based Federated Averaging (FedPA) [27], a pioneering aggregation and client selection strategy tailored for medical imaging. Unlike FedAvg, which aggregates updates from all clients without bias, FedPA strategically integrates contributions solely from best-performing clients, thereby reducing noise and improving model stability. After each local training round, clients are evaluated on validation data, and only those exceeding a performance threshold (e.g., F1 score >= 0.70) contribute to the global update, while others are synchronized with the latest global model and remain eligible for future rounds. This dynamic selection not only filters out unreliable updates but also mitigates data imbalance effects, since contributions from clients with severely skewed or underrepresented samples are down-weighted unless they demonstrate sufficient predictive quality. We validate FedPA on a subset of the Stanford MURA dataset under non-IID conditions, where each client is restricted to a single bone type category. Experimental results, evaluated across convergence time, accuracy, loss, scalability, efficiency, and class balance, demonstrate that FedPA accelerates convergence. Also enhances stability, effectively addresses data imbalance, and improves the practicality of FL for large-scale medical imaging while preserving data privacy.
2.1 Medical Imaging and Federated Learning
Healthcare imaging techniques possess distinct data and characteristics. As depicted in Fig. 2, various electromagnetic (EM) scanning methods are employed for diagnosing and monitoring medical conditions covering the full EM spectrum. Each technique functions at a specific frequency and wavelength, exhibiting unique properties [2]. When EM waves interact with an object, they may be scattered, reflected, or absorbed. Magnetic Resonance Imaging (MRI) utilizes a magnetic field to align the body’s natural protons. When subjected to a radio-frequency signal, these protons gain energy and shift their orientation relative to the magnetic field. Computed Tomography (CT) and X-ray imaging operate in the high-frequency range of
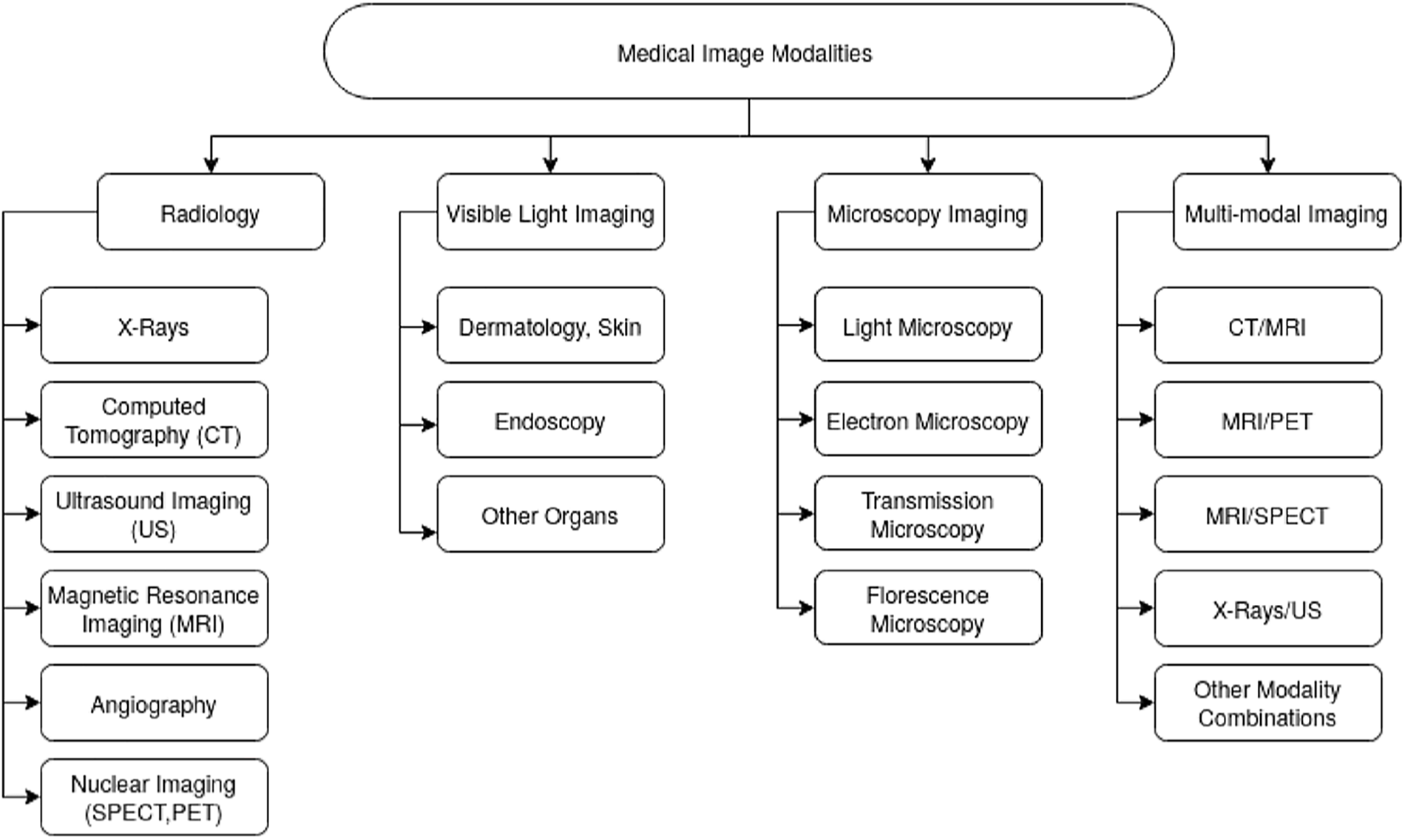
Figure 2: Medical imaging modalities classification [2]
2.2 Medical Imaging and Federated Learning
Medical imaging datasets are often distributed across multiple hospitals, research institutions, and diagnostic centers. Studies have demonstrated that FL can enhance diagnostic accuracy by leveraging data from multiple institutions without transferring sensitive patient data. For instance, Sheller et al. [29] applied FL to brain tumor segmentation using MRI scans, showing that FL achieved performance comparable to centralized training while ensuring data privacy. Similarly, the authors of [30] explored FL for chest X-ray classification and demonstrated that training across multiple institutions improved generalization to unseen data. A systematic review by [31] on federated learning in healthcare highlights important research areas, model aggregation improvements, and case studies on EHR data. It emphasizes privacy concerns and proposes a general architecture based on existing studies. Federated learning (FL) in healthcare, highlighting its role in collaborative model training while preserving data privacy. FL enables the use of diverse healthcare datasets without compromising patient confidentiality [32].
Federated Learning (FL) has emerged as a promising paradigm for training machine learning models across decentralized datasets while preserving data privacy [33]. FL is particularly relevant in the healthcare sector, where sensitive patient data is often kept across multiple institutions. FL applications in healthcare include disease diagnosis, personalized treatment recommendations, and drug discovery [34]. Integration with IoT devices and wearables enhances real-time, decentralized data processing for predictive analytics [35]. Federated Learning (FL) is a decentralized machine learning technique that enables multiple clients (e.g., hospitals, medical research centers, or edge devices) to collaboratively train a shared model while keeping their data private and localized. Unlike traditional centralized learning, where data is transferred to a central server, FL ensures that model updates (rather than raw data) are exchanged among participating nodes, reducing privacy risks and regulatory concerns.
This study presents a PRISMA-based meta-analysis of recent surveys on Federated Learning in Medical Imaging (FL-MI). It synthesizes key trends, challenges, and strategies for addressing data heterogeneity, privacy concerns, and non-IID performance. The review compares commonly used datasets, machine learning models, and FL frameworks in applications such as tumor detection, organ segmentation, and disease classification. Research gaps are identified, particularly the need for stronger privacy protection, offering directions for future FL-MI development [36].
In a conventional FL setup, every single client trains a local model employing its own dataset and transfers model updates to a central server, which sums these updates to amend the global model. The model is trained sequentially until convergence. (see Fig. 3). That said, traditional FL algorithms such as FedAvg sweat with non-IID data distributions, client heterogeneity, and communication overhead, acutely in healthcare domains where data availability and quality differ radically around institutions.
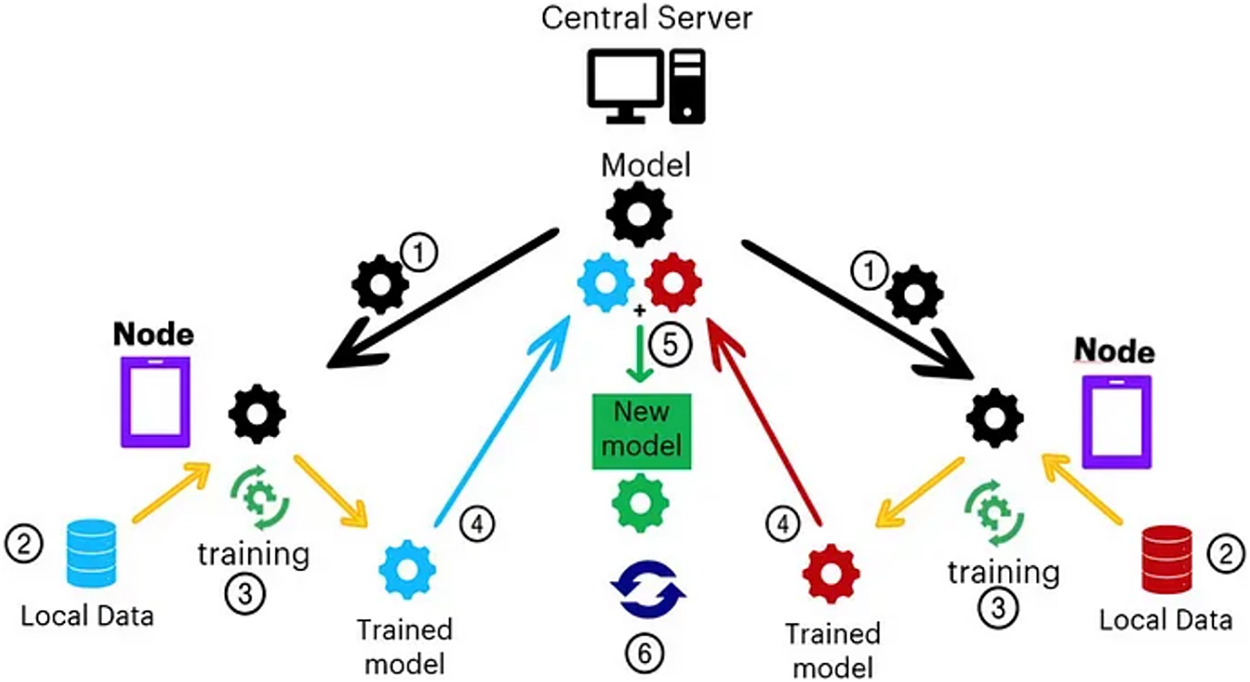
Figure 3: Federated learning step by step process [17]
However, several challenges, including aggregation inefficiencies, performance variations among clients, and non-IID data distributions, hinder its effectiveness [37]. Privacy-preserving techniques like differential privacy and secure multiparty computation are being explored to address these issues. Despite these challenges, FL offers revolutionary potential for secure, data-driven healthcare systems, promising improved patient outcomes and operational efficiency. Future research directions include refining FL implementations and expanding its use to broader healthcare applications [38]. Federated Learning is increasingly adopted in healthcare because it enables shared model training while meeting data privacy requirements like HIPAA and GDPR. It is especially effective for medical imaging, predictive diagnostics, personalized treatment planning, and disease-progression monitoring.
Federated learning (FL) [39] has proved to be an appealing framework for decentralized model training across distributed sources of data, particularly in domains such as healthcare, where privacy regulations against the aggregation of central data prohibit data centralization. The standard FL process of server-side model averaging following local training was established with the initial FedAvg algorithm [23]. Although extensively employed, FedAvg presumes all client updates have the same informativeness, something that commonly collapses under non-IID scenarios. Heterogeneity in FL data may lead to biased, unstable updates that hurt overall model performance.
Several techniques have been proposed to address the challenges introduced by non-IID data. FedProx [24] adjusts the local objective function by including something known as the proximal term, thereby stabilizing local updates and limiting client drift. However, it still averages updates from all clients indiscriminately regardless of quality. FedSGD [23], on the other hand, employs a more sophisticated technique by averaging gradients more frequently but at the cost of excessive communication overhead, making it less scalable in practice.
FedBN [20] addresses distributional heterogeneity by disentangling batch normalization layers in a way that clients are able to maintain locally adapted statistics. While this guarantees improved personalization and overall accuracy, it does so at the expense of higher memory costs and does not directly check the credibility of client updates. Outside of algorithmic modifications, client choice schemes have also been studied and explored to improve robustness and efficiency. For example, Wang et al. [40] proposed a system-aware selection mechanism that takes device availability into account, while Nakakita et al. [41] focused on fairness in participation by using a resource-aware scheduling. Such methods generally rely on system-level metrics and do not involve explicit measurement of model performance as a selection criterion.
Existing aggregation and selection mechanisms are often insensitive to the quality of individual client models. In high-stakes applications like medical imaging, with very heterogeneous data and significant emphasis on model accuracy, these limitations can become issues. This was the inspiration behind our work, which advances previous research by proposing FedPA. A performance-sensitive aggregation method, that leverages local F1 score for filtering and weighing client responses. F1-score is a performance metric mainly used for evaluating imbalanced classification tasks. Thus, F1 was determined to be most suitable to act as a threshold metric along with accuracy. In comparison to existing methods, it incorporates model-level performance metrics at the aggregation phase itself, aiming to enhance reliability, reduce communication overhead, and increase global accuracy over heterogeneous environments.
2.2.1 FedAvg (Federated Averaging)
FedAvg is one of the first federated learning algorithms, introduced by McMahan et al. [23]. The algorithm facilitates decentralized model training, where clients train models locally on their local data and send their updates (model weights) to a central server. The server aggregates these updates by averaging them with weights proportional to the size of the client’s local dataset. This approach allows the global model to take advantage of the heterogenous data of various clients even when the data is non-IID (non-independent and identically distributed) without exchanging raw data. The mathematical expression for FedAvg is given below:
where:
•
•
•
•
FL with EfficientNet-B0 and FedAvg [42] enhances privacy and diagnostic accuracy in brain tumor MRI. EfficientNet-B0 outperforms ResNet under data heterogeneity, achieving approx. 99% average client accuracy and low loss across diverse datasets.
2.2.2 FedSGD (Federated Stochastic Gradient Descent)
FedSGD is a basic version of FedAvg, which applies the stochastic gradient descent (SGD) optimization algorithm. In FedSGD, clients calculate one step of the gradient on their local data and transmit gradients to the central server. The server aggregates these gradients, with a weighted update to the global model. Compared to FedAvg, which aggregates model weights, FedSGD aggregates gradients, which can potentially achieve faster convergence in some cases but might be more susceptible to client data heterogeneity. The mathematical expression for FedSGD is:
where:
•
•
•
•
•
In [43], the brain tumor segmentation using federated learning shows that increasing the number of clients negatively impacts model performance. Skewed data distributions yield better results than equal splits. FedAvg consistently outperforms FedSGD in handling such non-IID scenarios.
2.2.3 FedProx (with Proximal Term)
FedProx enhances FedAvg by adding a proximal term to counteract the issue of client drift in federated learning, particularly when clients hold non-IID data. The term compels local models to stay close to the global model, thereby evading the phenomenon of divergent updates caused by heterogeneous client data distributions. FL enables privacy-preserving cancer diagnosis using deep learning across institutions. Using heterogeneous cancer datasets (cervical, lung, colon), FedAvg and FedProx were compared under non-IID conditions. FedAvg and FedProx [44] match centralized training under IID data. With increasing heterogeneity, both degrade, but FedProx remains superior. FL struggles with many clients and non-uniform labels, highlighting the need for advanced methods in such settings. FedProx [45] outperformed FedAvg in highly heterogeneous settings. Bayesian optimization improved hyperparameter tuning for both local and global models.
where:
•
•
•
•
2.2.4 FedBN (Federated Batch Normalization)
FedBN [46] is constructed to address federated learning issues due to the presence of Batch Normalization (BN) layers in models. BN layers operate on the statistics (mean, variance) that are calculated on the entire set of data, but in the federated case, these might be extremely different across clients due to the fact that the data is non-IID. To mitigate this, FedBN prevents aggregation of clients’ BN layers’ parameters but instead aggregates the non-BN parameters only. This serves to mitigate the domain shift of clients. The new local batch statistics and global model of every client are represented as:
FedBN Local BatchNorm Statistics:
where:
•
•
•
•
•
FedAvg showed comparable internal validation to single-site models but improved generalizability. Personalized FL algorithms outperformed FedAvg in internal validation, though FedAvg retained better generalization. FedBN achieved superior performance on both internal and external validation [47].
We here present FedPA (Federated Learning Performance-based Averaging), a novel client selection strategy committed to improving global model update quality in federated learning. FedPA is designed to surmount challenges posed by non-IID data and heterogeneous client performance, thereby making sure high-quality clients with trustable updates build up the model. The key idea of FedPA is to introduce a performance-aware client selection mechanism by the local model performance, as measured by the F1 score, to filter low-quality updates before aggregation.
FedPA operates in two primary phases: client selection and aggregation. During the client selection phase, clients are locally evaluated after training, and only those whose local F1-score exceeds a predefined threshold are allowed to participate in the aggregation process. F1 was selected as the evaluation metric because it effectively handles imbalanced and non-IID data distributions, which characterize our task. In our experiments, we initiated the adaptive threshold tuning process from an initial value of 0.60, incrementally increasing it by +0.02 after each complete training cycle (i.e., after all clients completed local updates). This progressive approach allowed us to observe the stability and performance gain at different F1 cut-offs. The experiments therefore started with a minimum F1 threshold of 0.60 and continued up to a maximum observed F1 of 0.77. Based on this exploration, we finalized the threshold value at 0.75 for the MURA dataset, as it provided the optimal trade-off between participation rate and global model accuracy. Clients whose F1-scores fell below the threshold are allow to take the global model parameters to maintain consistency across rounds.
3.2 Client Selection and Performance Threshold
The client selection in FedPA is based on local F1 scores, a common metric for measuring classification performance, particularly in datasets with imbalances. This performance metric is particularly appealing in federated learning since it estimates both precision and recall and provides a more robust measure of client model quality than accuracy alone. The F1 score threshold can then serve to filter out only those clients generating sufficiently strong updates, so that model aggregation is not contaminated by low-quality contributions. In order to make clients dynamically available, a fallback mechanism is introduced: if no client achieves the threshold in a round, the aggregation defaults to a regular FedAvg-style aggregation. This fallback provides resilience such that the global model can still be updated even when most clients are performing poorly.
Once the best-performing clients are selected, their local models are then aggregated into the global model. Also, FedPA uses a weighted average combination method, with each of the selected clients’ contribution as a fraction of the performance of its local model (F1 score). This ensures that clients with higher F1 scores contribute more to the global model, also making the process of combination optimal.
In summary, FedPA introduces a performance-aware mechanism to federated learning to make the global model more efficient and reliable by preferring well-performing clients and curbing the impact of bad updates. The experimental configuration and the results of applying FedPA to a real-world dataset are introduced in the next section.
3.4 FedPA (Federated Performance-Based Averaging)
FedPA (Federated Performance-based Aggregation) is a novel technique in federated learning that prioritizes client contributions based on their F1 scores, which represent the balance between precision and recall in classification tasks. Unlike conventional approaches where all client updates are treated equally, FedPA assigns greater influence to clients with higher F1 scores, allowing the global model to be updated primarily by the most reliable and well-performing participants.
A key component of FedPA is its adaptive threshold mechanism, which dynamically filters out clients whose F1 scores fall below a predefined threshold. This ensures that only top-performing clients are included in the aggregation process, thereby enhancing the overall stability and convergence of the global model. Clients excluded from aggregation are synchronized with the latest global model parameters, enabling them to continue local training and potentially meet the threshold in subsequent rounds.
Through this selective and performance-driven strategy, FedPA effectively reduces the impact of noisy or underperforming updates, leading to faster convergence and improved model generalization. The following Algorithm 1 presents the algorithmic workflow of FedPA:
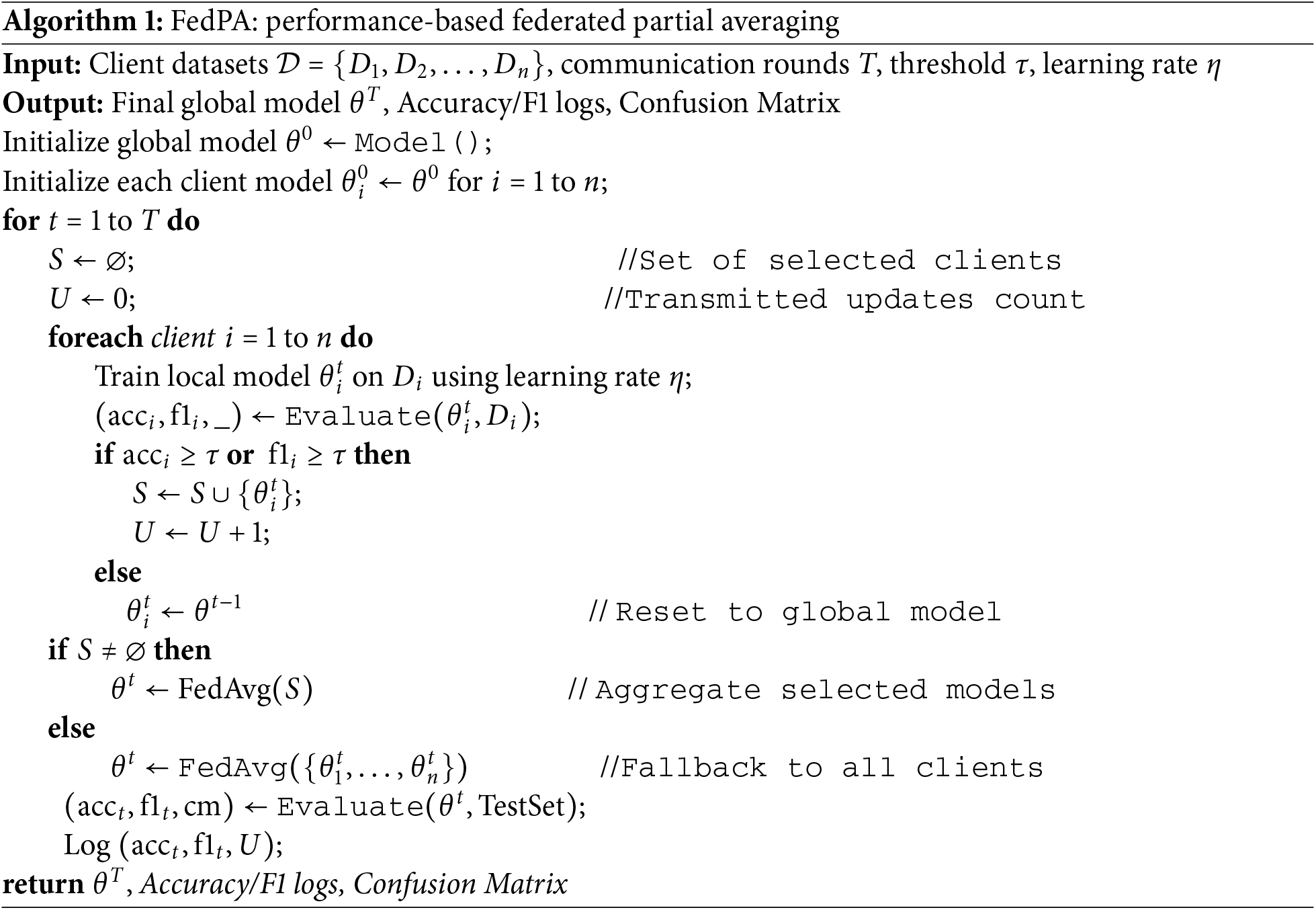

4 Experimental Setup and Evaluation
We use a pre-trained and fine-tuned DenseNet121 as our base model architecture for binary classification. Local training is performed for 5 rounds of communication, and each round has 5 epochs. Experimental findings show that FedPA is as good as or even outperforms state-of-the-art approaches such as FedAvg, FedProx, FedBN, and FedSGD [22,19,46] on the important measures of Accuracy, F1 Score, and Recall. In addition, it achieves similar performance with less round time and memory overhead, showing its feasibility in resource-limited federated learning environments [19,48].
The MURA (Musculoskeletal Radiographs) dataset, originally assembled by the Stanford ML Group, comprises 40,561 X-ray images from 14,863 studies (12,173 patients) across seven upper-extremity regions, including the finger, hand, wrist, forearm, elbow, humerus, and shoulder. All images were extracted from Stanford Hospital’s PACS system, and the authors report that the dataset is Institutional Review Board (IRB) approved, with all radiographs fully de-identified and HIPAA-compliant to ensure patient privacy. The dataset is publicly available for research via its canonical webpage [49]. In this study, we utilized a clinically relevant subset of MURA, specifically radiographs of the hand, finger, shoulder, and wrist (see Table 1 and Fig. 4), as these regions are central to diagnosing fractures and related abnormalities. The problem is formulated as a binary classification task—distinguishing normal from abnormal cases—reflecting realistic clinical workflows where X-rays from multiple healthcare centers are aggregated to support diagnostic decision-making.
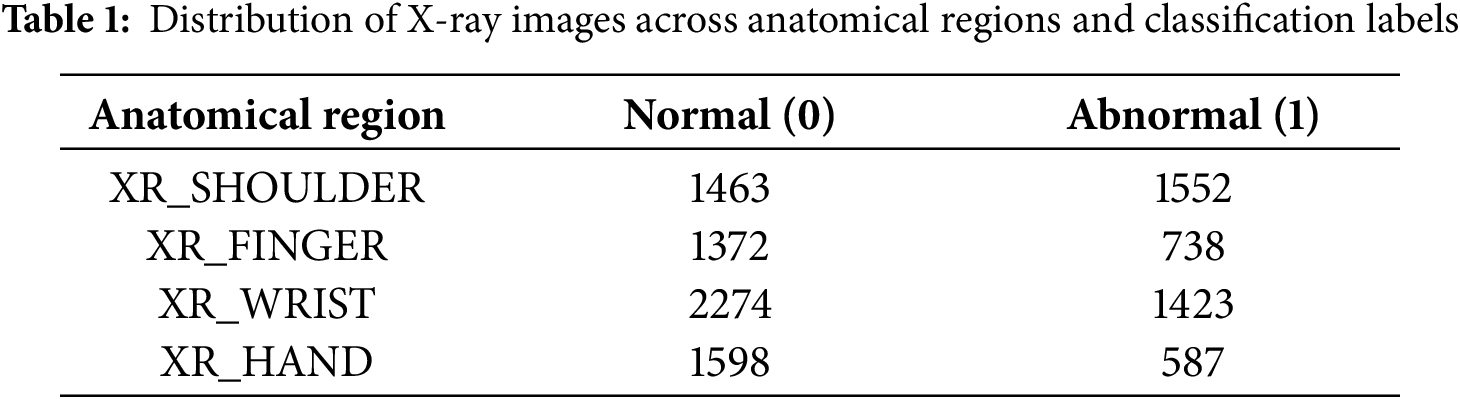
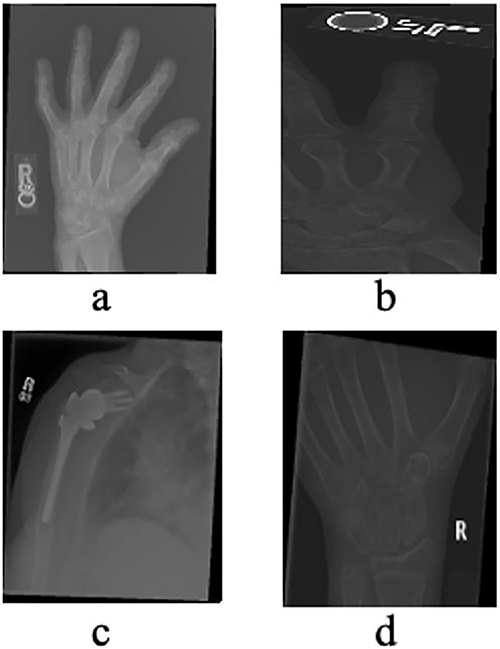
Figure 4: X-rays of Hand (a), Finger (b), Shoulder (c) and Wrist (d) taken from the validation dataset
The dataset was created with the intention to simulate real-world non-IID (Non-Independent and Identically Distributed) environments. The dataset was especially distributed across different simulated hospital clients, with each specializing in a particular anatomical body part. This segmentation is evocative of real-world healthcare systems’ specialization, where different hospitals can specialize in the treatment of specific body parts or diseases, and this segmentation can also pick up on patient population differences. Prior work has noted the usefulness of this approach, with diverse, non-IID data shown to enhance model robustness and generalizability [46,50]. To train the model, the data was preprocessed in several ways to feed high-quality input to the machine learning model. All images were resized to 224 × 224 pixels, a common size for deep learning models [51], and then converted to grayscale to simplify the amount of information being processed, as color may not provide much more detail here. Normalization to normalize pixel values to a standard range to increase the model’s learning ability was utilized. Histogram equalization was also utilized to boost image contrast so that details can be more easily picked up by the model, as in medical imaging, small abnormalities or fractures need to be detected. Local data augmentation techniques like random rotation, flipping, and zooming were utilized for model robustness and to prevent overfitting. These improvements are fairly standard for image processing towards getting a more generalized model that will have fewer tendencies to memorize some properties of the training set. The data was split into three parts: 70% for local training, 10% for local validation (for calculating each client’s local F1 score), and 20% for final testing of the global model.
For this purpose, we employed the DenseNet121 model, a 121-layer deep convolutional neural network that is state-of-the-art, efficient, and effective. It is especially so in settings where data may be limited or fragmented. DenseNet121 has a unique feature of dense connectivity, where each layer is fed by all the previous layers, allowing for more efficient gradient flow and feature reuse. This property renders DenseNet121 extremely appropriate for medical image classification, as it can learn from relatively smaller datasets without overfitting, which is an essential feature for medical applications where labeled data may be scarce.
To adapt DenseNet121 for binary fracture classification, the model’s original classification head was discarded and substituted with one fully connected layer and a sigmoid activation function. This change thus allows the model to give a probability score of whether there is a fracture in an image or not. The use of a sigmoid function is common in binary classification issues since it provides values between 0 and 1, which are probabilities. The model was initialized with ImageNet pre-trained weights, a common practice in transfer learning. This allowed the DenseNet121 architecture to make use of prior knowledge from a vast diversity of general image datasets and specialize it for the medical domain.
We did not include dropout layers or explicit max-pooling operations. This is because DenseNet121 employs efficient downsampling and regularization methods in its architecture. This decision is supported by literature [52], where DenseNet’s robust regularization power is highlighted, and the inclusion of dropout or pooling layers would thus be redundant and maybe even detrimental.
The federated learning setting in this study simulates a distributed training procedure among multiple healthcare organizations, thus Federated learning makes it viable to shape a global model by merging updates from local models, where the data remains at the clients (hospitals) and is not shared directly with the server, ensuring compliance with data protection law and confidentiality [53].
Clients
The federated learning was carried out on four clients, each representing a distinct hospital or medical center. Each client trained on a particular subset of data for a different type of bone (hand, finger, shoulder, wrist). This setup exposed the model to a wide variety of X-ray images from different anatomical sites, which enhanced the model’s robustness and generalizability (See Fig. 5 and Table 2).
Figure 5: Federated learning workflow across hospitals with local specialist models for different body parts. Each hospital trains a local model on private data, and the models are coordinated via a central cloud server
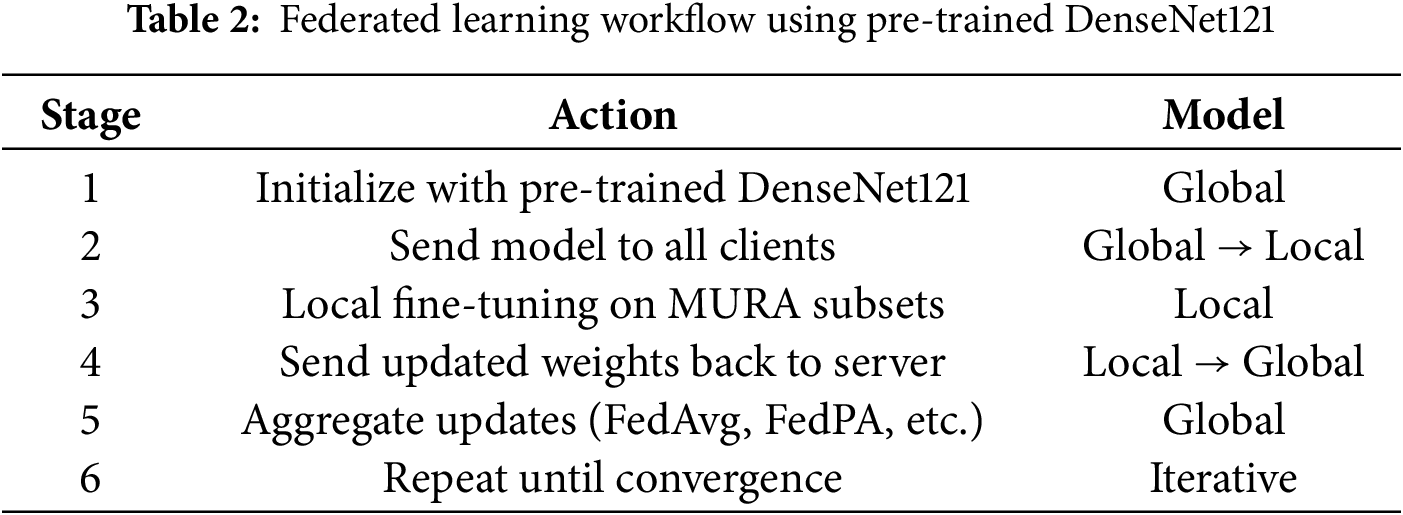

This federated setup follows the principles laid out in previous work on multi-center healthcare datasets [54].
The global model was initialized using DenseNet121, pre-trained on the ImageNet dataset to utilize its strong feature extraction ability from large-scale natural images [55]. In the proposed federated learning setup (see Table 2), the pre-trained parameters were distributed to all participating clients. Each client locally fine-tuned the model on its subset of the MURA radiographic dataset, enabling domain adaptation without sharing raw medical data. After local training, clients sent the updated model parameters to the global server for aggregation using the Federated Averaging (FedAvg) algorithm. This process was repeated over multiple communication rounds, allowing the global model to gradually improve its representation and diagnostic accuracy. The combined use of transfer learning and federated optimization enhanced model generalization while preserving data privacy. Such collaboration has proven effective in improving performance across decentralized healthcare systems [52].
4.5 Evaluation Metrics and Protocol
All binary classification metrics were computed using standard implementations from the scikit-learn library to ensure validated and reproducible results. The following metrics were used in model evaluation:
• Accuracy: Measures the proportion of correctly classified samples among all predictions.
where TP (True Positives), TN (True Negatives), FP (False Positives), and FN (False Negatives) denote the respective classification outcomes.
• Precision: Represents the proportion of true positives among all positive predictions.
• Recall (Sensitivity): Represents the proportion of actual positives correctly identified by the model.
• F1 Score: Harmonic mean of Precision and Recall, balancing both measures.
Evaluation Protocol: Model evaluation was performed on validation sets maintaining identical anatomical distributions as the training data. Each client model was tested on both its target anatomical region and cross-domain samples to evaluate specialization and generalization performance. Random seed control (random_state = 42) ensured consistency and reproducibility across experimental runs.
4.6 Federated Learning Parameters
For all federated training experiments, we fixed a random seed of 42 to assure replicability and consistent outcomes. The model was trained for 5 global rounds, with each client performing 5 local epochs before transmitting updates to the server. A batch size of 32 was chosen to balance computational efficiency with convergence stability. The Adam optimizer was employed with a learning rate of 0.0001, a value determined experimentally to be effective for fine-tuning the DenseNet121 model. Performance was evaluated using F1 score, accuracy, precision, and recall as primary metrics. To standardize preprocessing across clients, we applied consistent data augmentation, including random horizontal flipping and normalization. Batch Normalization layers were not frozen but updated during local training to adapt to client-specific data distributions. These parameters were carefully selected in line with best practices for federated learning in medical image classification [56].
4.6.1 Client Participation and Selection
Every client, in the federated learning setup, trained its local model and evaluated it using the F1 score. The updates from each client were dynamically weighted based on their F1 scores, where clients with better F1 scores contributed more to the global model. This weighted averaging technique ensures that all clients with good performing models contribute more to the global model, which thus improves overall accuracy and fairness. This approach has been shown to enhance model performance and minimize biases (see Fig. 6) that may be incurred due to imbalanced or heterogeneous data distributions across clients [56].
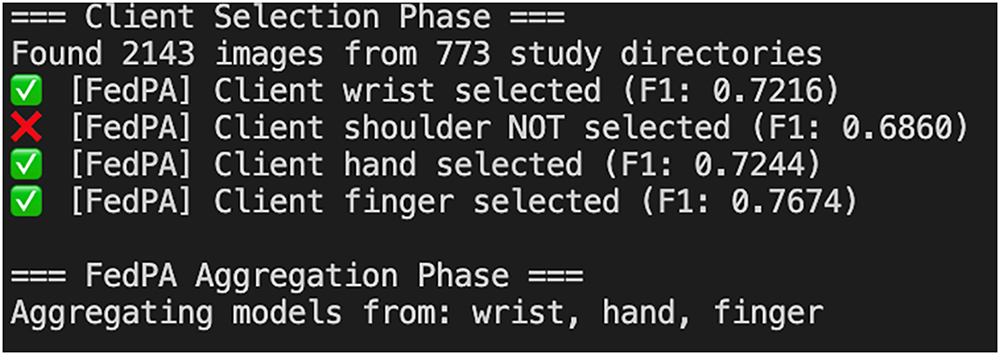
Figure 6: Client selection process based on F1 threshold in FedPA
Following each round of local training, the updates from all the clients were aggregated on the basis of a weighted averaging method. Clients with higher F1 scores contributed more to the global model, reflecting their higher accuracy during local testing. This performance-based aggregation method ensures that the overall model performance is maximized and that it also prevents underperforming models from decreasing the accuracy of the global model. Aggregation is a critical component in federated learning as it facilitates client collaboration without compromising data privacy and confidentiality, which is a key consideration in healthcare settings [54].
The learning rate was 0.0001, experimentally found to be quite suitable to fine-tune the DenseNet121 model. The batch size was chosen to trade off computation overhead against model performance, and the local epochs parameter was 5, indicating that every client trained the model five times before uploading the model updates to the server.
5.1 Final Performance Comparison
Table 3 for global model performance and, Table 4 and Fig. 7, shows the overall performance of the final model at round 5 for all five federated learning approaches. FedProx consistently had the highest overall accuracy and F1 score, followed closely by FedPA, particularly in recall where FedPA outperformed all other methods. Precision was highest under FedProx, followed by FedSGD.
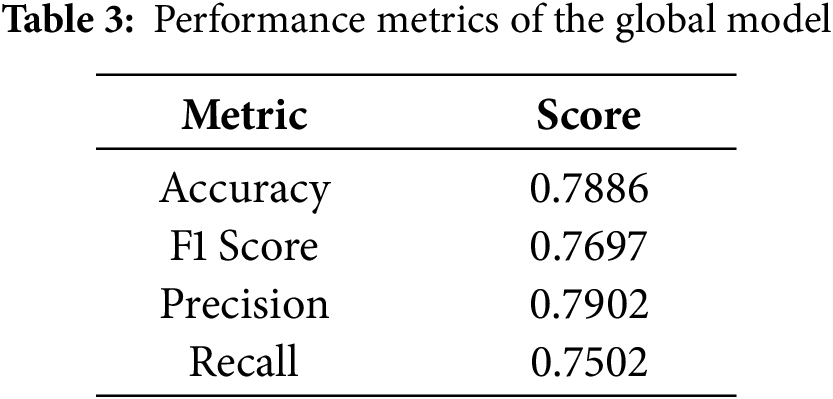
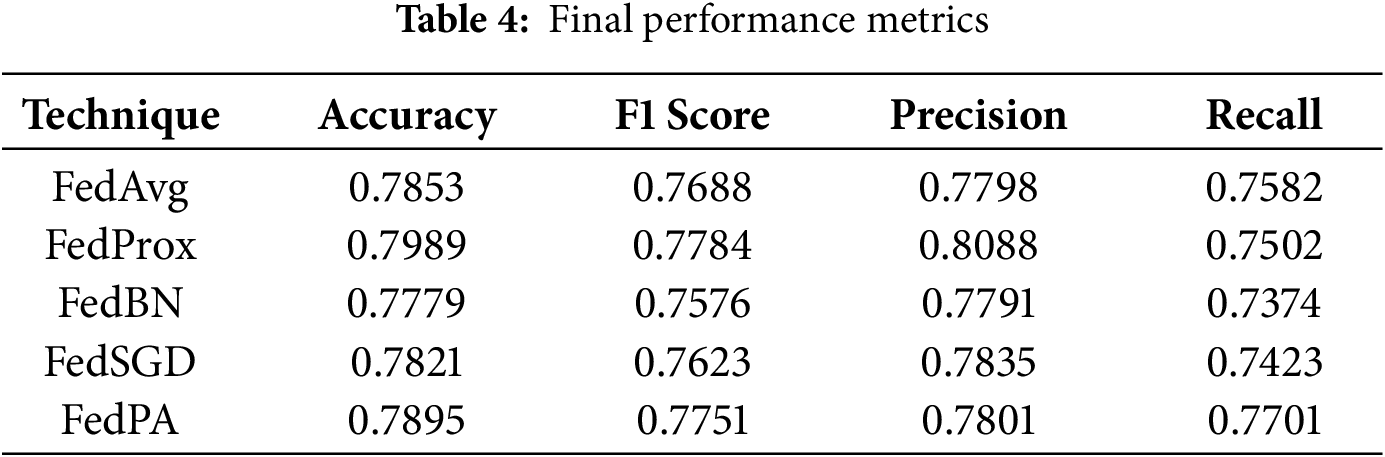
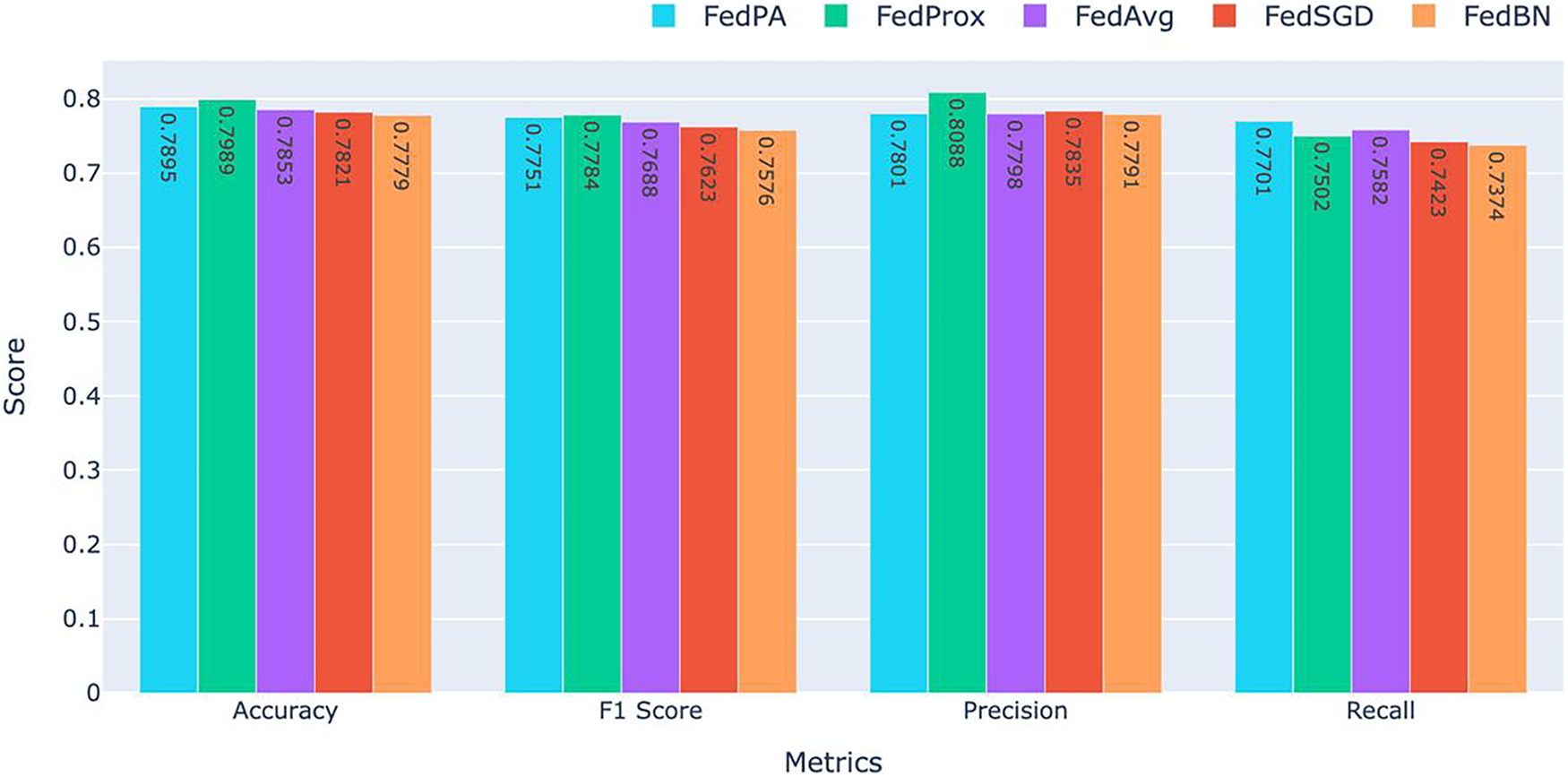
Figure 7: Comparison of Accuracy, F1 Score, Precision, and Recall for FedAvg, FedProx, FedBN, and FedSGD. FedPA achieved the highest performance in Recall with the competitive results in the other metrics
As shown in Fig. 7, FedProx achieved the best accuracy (0.7989) and F1 score (0.7784), which indicates its ability to balancing precision and recall consistently. The high accuracy and F1 score of FedProx demonstrate that this method is robust in maintaining model performance for all clients and has the ability to generalize well even in the presence of non-IID settings prevalent in federated learning.
FedPA, though slightly behind FedProx when it comes to accuracy (0.7895), was the top performer when it came to recall (0.7701) among all the other methods, beating all the other techniques. This indicates that FedPA is particularly adept at capturing true positive instances critical in medical applications where it’s important to catch all potential instances of abnormalities. The increased recall clearly indicates that FedPA very sensitive to abnormalities in the data and would therefore be a great choice in situations where a failure to identify a positive case would be catastrophic.
For precision, FedProx again ranked first with the highest precision (0.8088), indicating its ability to avoid false positives. Precision is a valuable metric when the cost of false positives is high, such as in the healthcare industry. FedSGD (0.7835) ranked second in precision, but its overall performance in other metrics was a bit lower than that of FedProx and FedPA.
FedBN had the worst results overall across all the metrics, with an accuracy of 0.7779, F1 score of 0.7576, precision of 0.7791, and recall of 0.7374. The worse performance of FedBN can be attributed to the challenge of normalization across non-IID data distributions, which must have impacted its ability to generalize poorly. It is especially evident when comparing the recall and F1 score to those of FedProx and FedPA, which reported higher results.
FedProx achieved the highest accuracy (0.7989) and F1 score (0.7784), showcasing its ability to provide balanced performance across metrics. FedPA, with its highest recall (0.7701), outperformed all other methods in detecting positive instances and is therefore the most suitable for applications where it is of utmost importance to detect every potential abnormality, such as in medical diagnosis. Meanwhile, FedBN performed worse across all the metrics and particularly on recall, which indicates the difficulty of normalizing data in federated learning when dealing with non-IID data distributions.
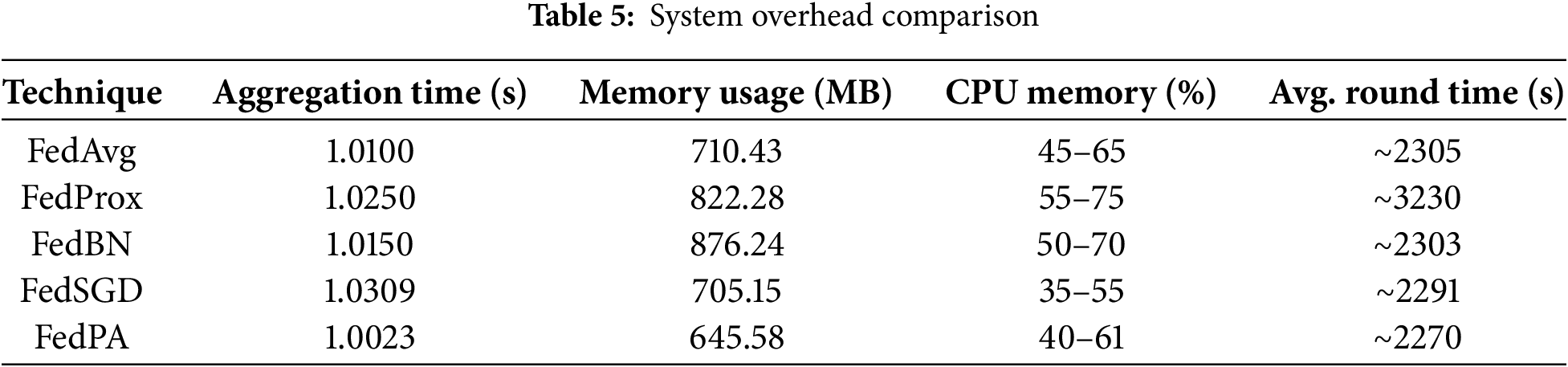
Computational Overhead Metrics (see Table 5) were analyzed to assess the system-level efficiency of the proposed federated learning framework.
• Memory Usage: Memory consumption for each process was monitored using the Python psutil library:
where the Resident Set Size (RSS) represents the portion of memory occupied in RAM. Both peak and average memory usage were recorded across all federated training rounds to measure the memory footprint of each algorithm.
• CPU Utilization: CPU load was sampled every 0.3 s using psutil.Process.cpu_percent (interval = 0.3). Peak and average CPU utilization were computed for each training phase to evaluate computational intensity and efficiency.
• Communication Overhead: The size of each model (
The total communication cost (
where
Training Effectiveness Metrics (see Table 5) were used to evaluate time-based performance. All timing measurements were obtained using Python’s time.time() function with microsecond precision. The total training duration included client-side model training, global aggregation, and evaluation steps across all federated rounds to provide a complete view of the system’s end-to-end efficiency.
Table 5, Figs. 8 and 9 summarize the observed system-level resource utilization and training performance. Among all methods, FedPA demonstrated the shortest average round time and the lowest memory consumption, while FedBN exhibited significantly higher memory usage. Fig. 8 illustrates the total training time (left) and aggregation time (right) for each algorithm. Notably, FedPA achieved the best overall efficiency, owing to its dynamic client weighting and performance-driven aggregation mechanism, which promote faster convergence without increasing either local training or global aggregation overhead.
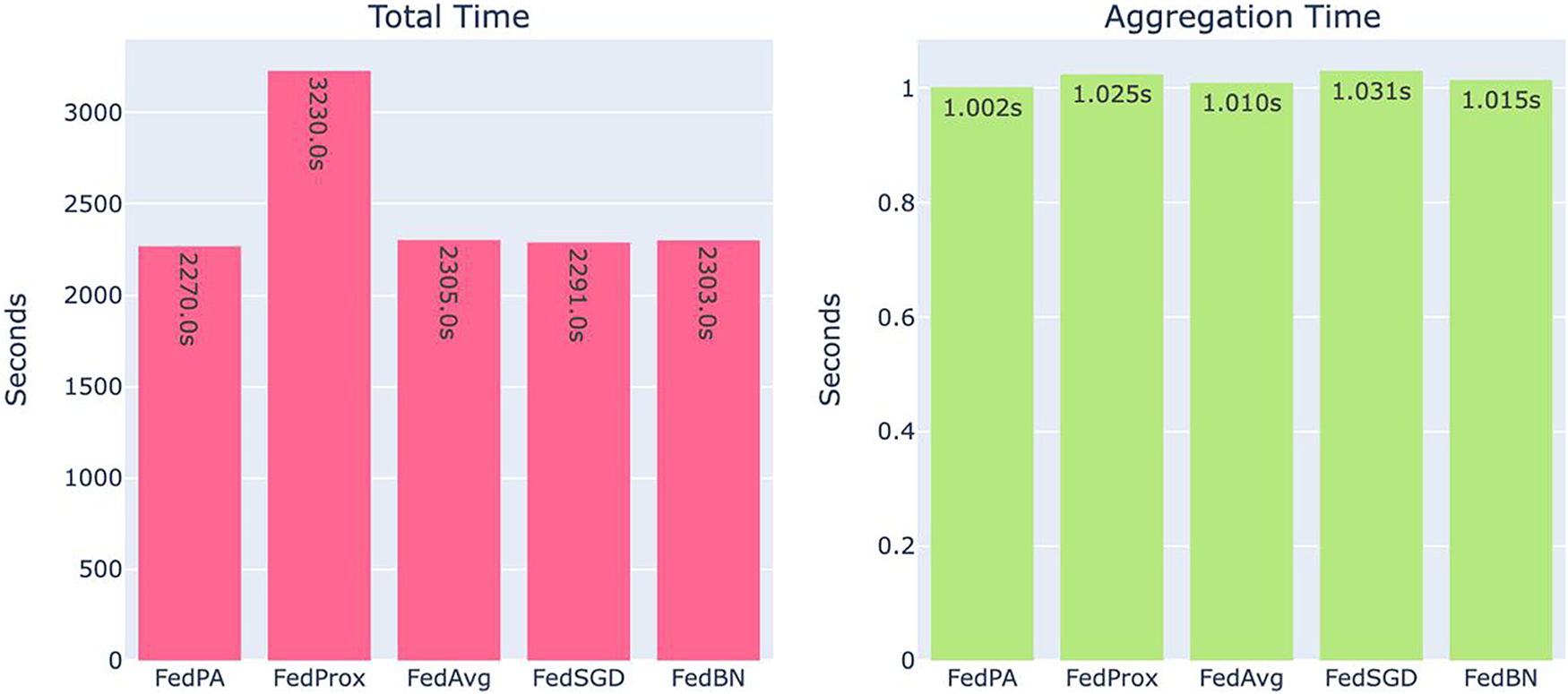
Figure 8: System overhead comparison: total training time (left) and aggregation time (right). FedPA achieves the least in both metrics
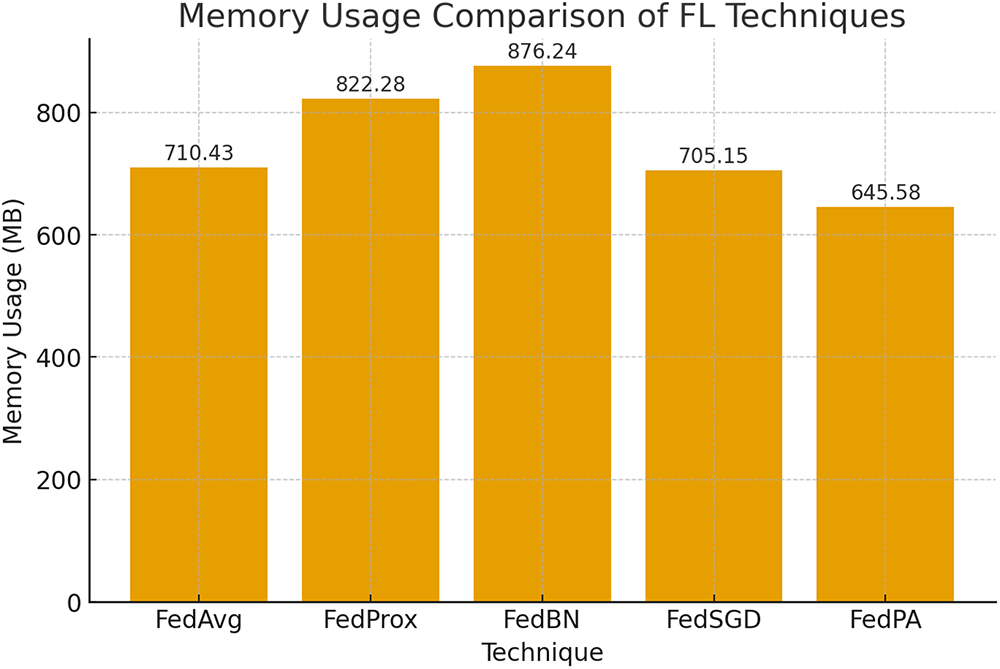
Figure 9: Usage of memory in the different federated learning algorithms
The light-weight computation resource distribution of FedPA pays off most in applications where prompt training and system response are of maximum importance, e.g., sparse resource-enabled edge devices or real-time applications in healthcare settings. Conversely, FedBN incurs the highest system overhead both in total training time and aggregation time, as depicted in Fig. 8. This is due to the increased complexity of aggregating BatchNorm statistics from non-IID client data, which leads to increased processing time for each global update. Computational complexity arises due to the requirement of addressing variations in BatchNorm parameters among clients, so FedBN is less preferable in resource-constrained settings compared to FedPA.
Fig. 9 once more compares the usage of memory in the different federated learning algorithms. FedPA and FedProx, as shown, have a minimum usage with only 645.58 MB a significant advantage in federated learning scenarios where memory usage is one of the most important parameters. The relatively minimal memory overhead of FedPA facilitates better scalability on a larger set of clients or even edge devices with limited memory capacity, and hence is more practical for use in distributed healthcare systems or mobile applications.
As a comparison, FedBN requires far more memory compared to the others, something that can be attributed to the increased storage space needed to deal with BatchNorm parameters from all clients during aggregation. Increased memory consumption could pose to be the bottleneck in scenarios of scale deployment or where there are numerous clients, further limiting its applicability in situations where memory is a precious commodity.
Overall, the findings in Table 6, Figs. 8 and 9 show that FedPA strikes the best balance between efficiency and usage of resources and is hence a strong candidate for federated learning in healthcare settings and other contexts requiring high model accuracy and low resource consumption.
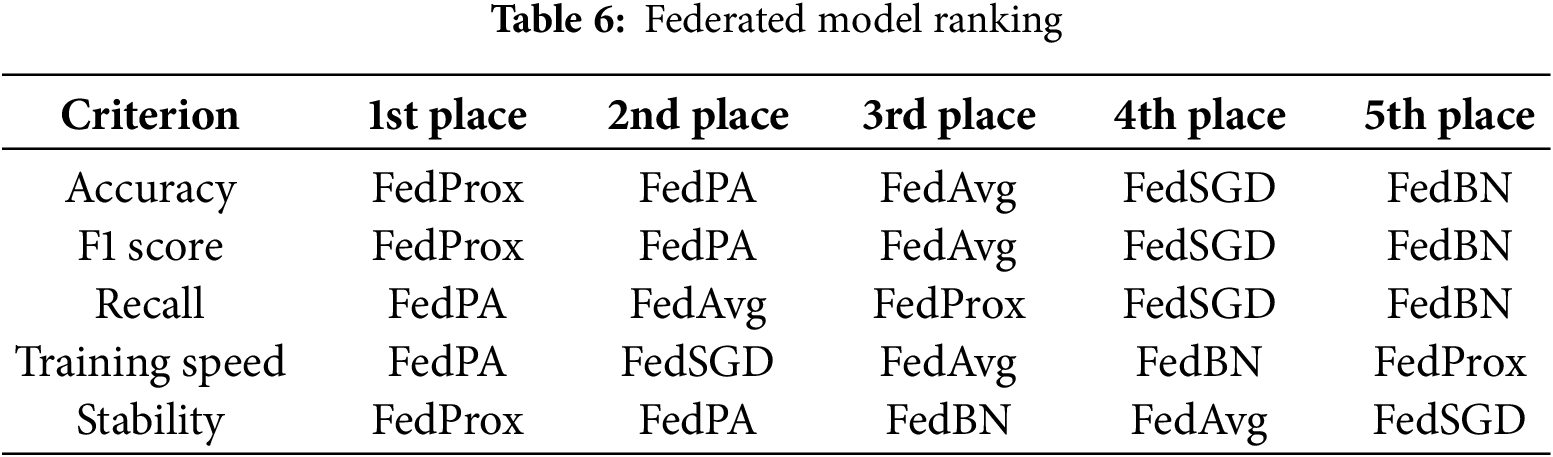
5.3 Overall Ranking by Evaluation Criterion
FedProx was the top scorer in model quality and reliability. It finished first in accuracy and F1 score and also finished first in stability. These rankings thus highlight FedProx’s advantage in handling heterogeneous data among clients, with stable outcomes and very minimal performance variance throughout the rounds. Its stability across these metrics means it is a trustworthy choice when model reliability is of the highest priority.
FedPA, on the other hand, performed best in training sensitivity and efficiency, ranking first in both recall and training efficiency. Its dynamic performance-aware aggregation method managed to drive the usage of resources to an absolute minimum while identifying more true positives than existing methods a desirable feature where responsiveness and detection sensitivity are paramount. FedPA also came in second for accuracy, F1 score, and stability, demonstrating that it can deliver nearly-competitive model performance at significantly lower system overhead, which makes it extremely well-suited for resource-limited environments.
FedAvg performed middle-of-the-pack on most measures. It had decent recall and training efficiency but did not outperform FedProx or FedPA on any single measure. FedSGD, while slightly quicker to train than some others, lagged behind both in model efficacy and in stability, ranking last in stability and near the bottom in most others.
FedBN, as expected, ranked last on nearly all measures, reaffirming its inappropriateness for the experimental setting. Having been designed to rely on batch normalization, FedBN was unable to deal with the non-IID data distributions used in this study, leading to nongeneralization and unstable training. Its performance highlights the challenge of using normalization-dependent techniques for federated learning with highly heterogeneous client data distributions.
In general, FedProx was the most resilient to attain high and stable model performance, and FedPA was the most efficient in training and best in detecting positive cases. This compromise makes FedPA a viable option in speed- and recall-oriented settings where absolute accuracy can be compromised to a certain degree. Conversely, FedBN’s low ranking in all the metrics suggests that it is not suitable for federated settings with heterogeneous clients without further modifications.
5.4 FedPA Aggregation Analysis
The proposed Federated Performance-Aware (FedPA) method demonstrates a very good balance between model performance and computational expense, primarily because of its dynamic client selection and weighted aggregation technique. Unlike baseline methods such as FedAvg or FedSGD, which uniformly take into account all client contributions. FedPA not only allocates aggregation weights based on the local F1 score of a client but also disregards clients with performance lower than some threshold value (t). This ensures that just those clients with sufficiently reliable updates contribute to the global model. Empirical results validate this strategy: FedPA is the second best in accuracy (0.7895) and F1 score (0.7751), while outperforming all methods in average round time (2270 s) and memory usage (645.45 MB). These results show that FedPA can be a promising choice for federated deployment on edge devices or bandwidth-limited environments, by eliminating low-performing clients.
FedPA avoids aggregating noisy updates, thereby improving convergence stability and model generalization as a whole. Moreover, FedPA is runner-up in recall, indicating its strength in capturing positive examples—a quality much in demand for high-stakes tasks such as medical diagnosis. FedPA achieves competitive performance with its lean footprint, closely trailing the deeper and more computationally expensive FedProx. Its ratio of performance to cost demonstrates FedPA’s merit in real-world federated learning environments where clients hold heterogeneous computational resources and data qualities.
For the CIFAR-10 dataset, the experimental setup followed a similar adaptive threshold tuning approach. The initial F1-score threshold was set at 0.60, and incrementally increased by +0.02 after each full training round. During this process, the maximum achieved F1-score reached 0.89, leading to the final threshold being set at 0.85 for optimal performance and stability. FedPA demonstrated strong and consistent improvements across all evaluation metrics, even under the challenging non-IID data distribution of CIFAR-10.
The model was trained for five global rounds, with each client performing ten local epochs per round. FedPA consistently outperformed the baseline methods, showing faster convergence and higher accuracy. In contrast, FedSGD performed significantly worse due to the multiclass nature of CIFAR-10, which rendered stochastic gradient descent ineffective in estimating accurate global gradients under non-IID conditions. This instability resulted in poor convergence and overall degraded model performance.
In conclusion, FedPA achieved the best overall results across all metrics on the CIFAR-10 benchmark, clearly demonstrating its robustness, scalability, and suitability for real-world Federated Learning scenarios. The detailed performance comparison is presented in Table 7 and Fig. 10.
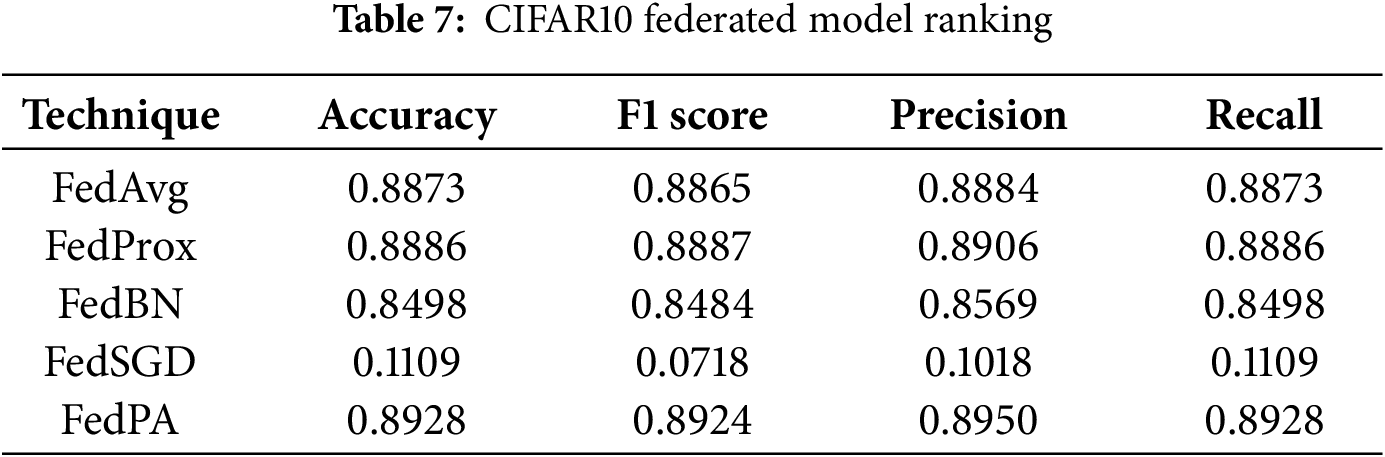
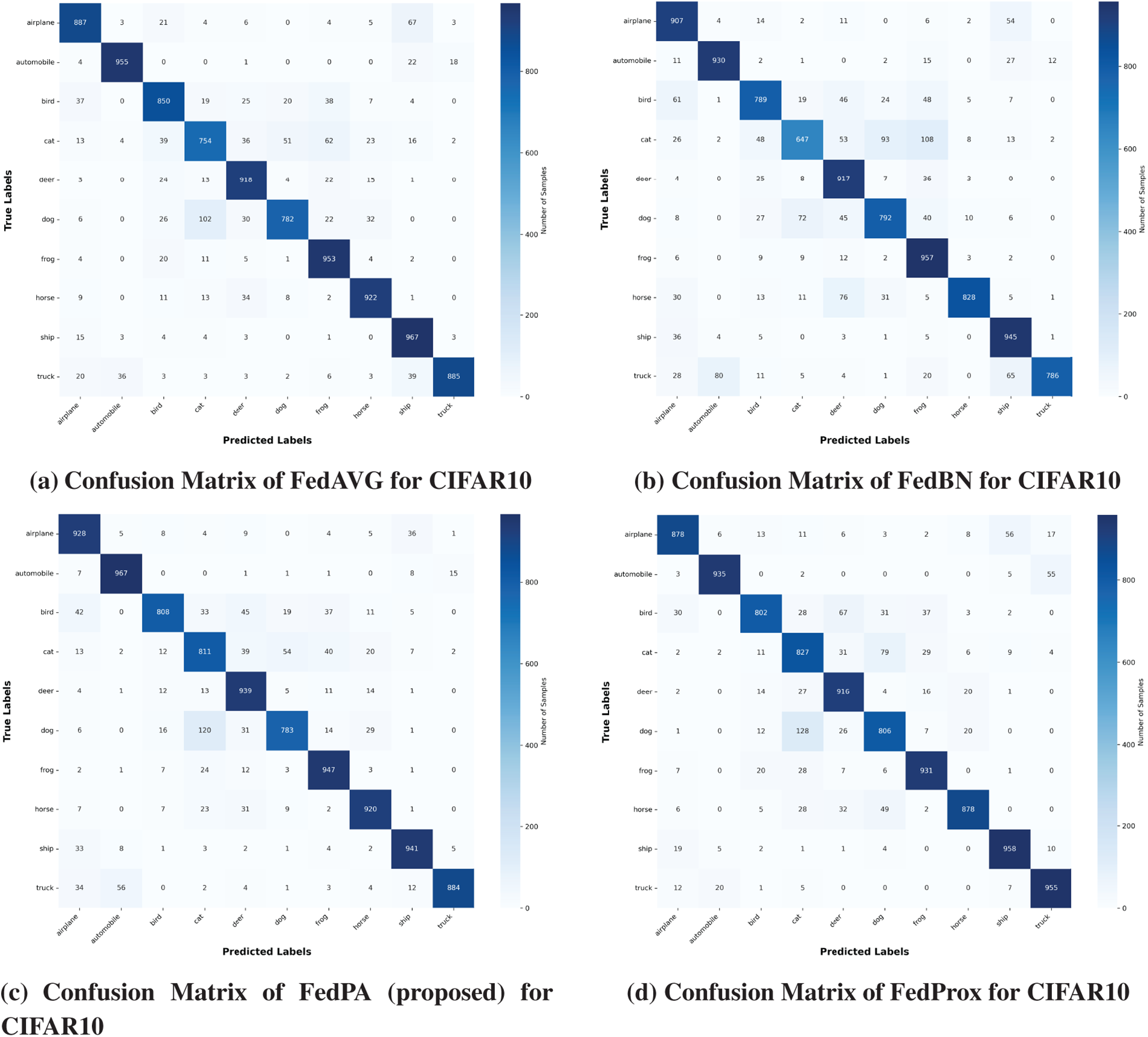
Figure 10: Confusion matrix of CIFAR10 dataset
This paper presents a comparative evaluation of five federated learning methods FedPA-Federated Performance-Based Averaging (proposed), FedProx, FedAvg, FedSGD, and FedBN. Used in a binary classification task with a DenseNet121-based deep neural network for fracture diagnosis on medical X-ray images. The data was partitioned into four heterogeneous clients, each possessing experience in a different body part (elbow, finger, hand, and shoulder), simulating real-world scenarios in healthcare environments.
Out of the methods experimented with, FedProx demonstrated the highest all-around performance across accuracy, F1 score, and stability. FedPA, our suggested methodology, though, gave a competitively comparable performance with significant reduction in system overhead. This places FedPA in an ideal position to be employed in cases of low computational capability, i.e., small health centers or edge devices. FedBN, on the other hand, was behind on all metrics, presumably because BatchNorm layers are difficult to aggregate across non-IID data distributions, a widely documented limitation in federated learning with healthcare data.
Our work, FedPA, introduces a dynamic client weighting and selection mechanism via local F1 scores. The approach allows for improved aggregation of model updates, where more performing clients contribute more to the global model update. The approach is a good balance between model accuracy and computational cost, especially useful in real-world healthcare federated learning scenarios.
There will be continuous efforts to improve FedPA in the context of asynchronous Federated Learning, where clients can choose to participate or skip training rounds based on their availability and performance. This will make the decentralized learning process more efficient and flexible. Adaptive, real-time client involvement will also be explored, where the system learns how to adjust to varying client availability and optimize the overall training procedure.
These improvements will be complemented further in FedPA through adaptive threshold tuning, so that the thresholds for weighting and client selection can be adaptively tuned based on real-time performance feedback. This would enable even greater control over the precision of aggregation, improving both efficiency and robustness of the federated learning system.
Moreover, FedPA will be evaluated on large-scale and diverse data, including multi-class and multi-modal clinical data, to investigate its scalability and performance in more complex and diverse real-world healthcare scenarios. This will encompass datasets with various medical imaging modalities (e.g., MRI, CT scans) and patient populations, enabling us to test the generalizability and reliability of FedPA across various medical contexts.
By addressing these domains, subsequent research will continue to make FedPA more appropriate for healthcare federated learning systems so that models can continue to remain accurate, scalable, and privacy-preserving while being computationally optimized.
Acknowledgement: None.
Funding Statement: The authors received no specific funding for this study.
Author Contributions: The authors confirm their contributions to the paper as follows: study conception and design were conducted by Atif Mahmood and Yasin Saleem; data collection, analysis, and interpretation of results were carried out by Usman Tariq, Yousef Ibrahim Daradkeh and Adnan N. Qureshi. The initial manuscript draft was prepared accordingly. All authors reviewed the results and approved the final version of the manuscript.
Availability of Data and Materials: The dataset is publicly available for research via its canonical webpage: https://stanfordmlgroup.github.io/competitions/mura/.
Ethics Approval: Not applicable.
Conflicts of Interest: The authors declare no conflicts of interest to report regarding the present study.
References
1. Ting DS, Liu Y, Burlina P, Xu X, Bressler NM, Wong TY. AI for medical imaging goes deep. Nat Med. 2018;24(5):539–40. doi:10.1038/s41591-018-0029-3. [Google Scholar] [PubMed] [CrossRef]
2. Iqbal S, Qureshi AN, Li J, Mahmood T. On the analyses of medical images using traditional machine learning techniques and convolutional neural networks. Arch Comput Methods Eng. 2023;30(5):3173–233. doi:10.1007/s11831-023-09899-9. [Google Scholar] [PubMed] [CrossRef]
3. Panayides AS, Amini A, Filipovic ND, Sharma A, Tsaftaris SA, Young A, et al. AI in medical imaging informatics: current challenges and future directions. IEEE J Biomed Health Inform. 2020;24(7):1837–57. doi:10.1109/jbhi.2020.2991043. [Google Scholar] [PubMed] [CrossRef]
4. Kourou K, Exarchos TP, Exarchos KP, Karamouzis MV, Fotiadis DI. Machine learning applications in cancer prognosis and prediction. Comput Struct Biotechnol J. 2015;13:8–17. doi:10.1016/j.csbj.2014.11.005. [Google Scholar] [PubMed] [CrossRef]
5. Gao CA, Markov NS, Stoeger T, Pawlowski A, Kang M, Nannapaneni P, et al. Machine learning links unresolving secondary pneumonia to mortality in patients with severe pneumonia, including COVID-19. J Clin Investig. 2023;133(12):e170682. doi:10.1172/jci170682. [Google Scholar] [PubMed] [CrossRef]
6. Khan P, Kader MF, Islam SR, Rahman AB, Kamal MS, Toha MU, et al. Machine learning and deep learning approaches for brain disease diagnosis: principles and recent advances. IEEE Access. 2021;9:37622–55. doi:10.1109/access.2021.3062484. [Google Scholar] [CrossRef]
7. Kilic A. Artificial intelligence and machine learning in cardiovascular health care. Ann Thorac Surg. 2020;109(5):1323–9. doi:10.1016/j.athoracsur.2019.09.042. [Google Scholar] [PubMed] [CrossRef]
8. Barragán-Montero A, Javaid U, Valdés G, Nguyen D, Desbordes P, Macq B, et al. Artificial intelligence and machine learning for medical imaging: a technology review. Phys Medica. 2021;83:242–56. doi:10.1016/j.ejmp.2021.04.016. [Google Scholar] [PubMed] [CrossRef]
9. Salah Eldin W, Kaboudan A. AI-driven medical imaging platform: advancements in image analysis and healthcare diagnosis. J ACS Adv Comput Sci. 2023;14(1):47–63. doi:10.21608/asc.2023.328064. [Google Scholar] [CrossRef]
10. Willemink MJ, Koszek WA, Hardell C, Wu J, Fleischmann D, Harvey H, et al. Preparing medical imaging data for machine learning. Radiology. 2020;295(1):4–15. doi:10.1148/radiol.2020192224. [Google Scholar] [PubMed] [CrossRef]
11. Wesemeyer T, Jauer ML, Deserno TM. Annotation quality vs. quantity for deep-learned medical image segmentation. In: Medical imaging 2021: imaging informatics for healthcare, research, and applications. Vol. 11601. Bellingham, WA, USA: SPIE; 2021. p. 63–76. [Google Scholar]
12. Kaissis GA, Makowski MR, Rückert D, Braren RF. Secure, privacy-preserving and federated machine learning in medical imaging. Nat Mach Intell. 2020;2(6):305–11. doi:10.1038/s42256-020-0186-1. [Google Scholar] [CrossRef]
13. Shah U, Dave I, Malde J, Mehta J, Kodeboyina S. Maintaining privacy in medical imaging with federated learning, deep learning, differential privacy, and encrypted computation. In: 2021 6th International Conference for Convergence in Technology (I2CT). Piscataway, NJ, USA: IEEE; 2021. p. 1–6. [Google Scholar]
14. Koetzier LR, Wu J, Mastrodicasa D, Lutz A, Chung M, Koszek WA, et al. Generating synthetic data for medical imaging. Radiology. 2024;312(3):e232471. doi:10.1148/radiol.232471. [Google Scholar] [PubMed] [CrossRef]
15. Adnan M, Kalra S, Cresswell JC, Taylor GW, Tizhoosh HR. Federated learning and differential privacy for medical image analysis. Sci Rep. 2022;12(1):1953. doi:10.21203/rs.3.rs-1005694/v1. [Google Scholar] [CrossRef]
16. Hossen MN, Ahmed K, Bui FM, Chen L. FedRSMax: an effective aggregation technique for federated learning with medical images. In: 2023 IEEE Canadian Conference on Electrical and Computer Engineering (CCECE). Piscataway, NJ, USA: IEEE; 2023. p. 229–34. [Google Scholar]
17. Mahmood A, Azizul ZH, Zakariah M, Belhaouari SB, Altameem A, Ramli R, et al. Implementing federated learning over VPN-based wireless backhaul networks for healthcare systems. PeerJ Comput Sci. 2024;10(1):e2422. doi:10.7717/peerj-cs.2422. [Google Scholar] [PubMed] [CrossRef]
18. Li T, Sanjabi M, Beirami A, Smith V. Fair resource allocation in federated learning. arXiv:1905.10497. 2019. [Google Scholar]
19. Li T, Sahu AK, Zaheer M, Sanjabi M, Talwalkar A, Smith V. Federated optimization in heterogeneous networks. Proc Mach Learn Syst. 2020;2:429–50. [Google Scholar]
20. Smith V, Chiang CK, Sanjabi M, Talwalkar AS. Federated multi-task learning. In: NIPS’17: Proceedings of the 31st International Conference on Neural Information Processing Systems. Red Hook, NY, USA: Curran Associates Inc; 2017. p. 4427–37. [Google Scholar]
21. Zhao H. Non-IID quantum federated learning with one-shot communication complexity. Quantum Mach Intell. 2023;5(1):3. doi:10.1007/s42484-022-00091-z. [Google Scholar] [CrossRef]
22. Kairouz P, McMahan HB, Avent B, Bellet A, Bennis M, Bhagoji AN, et al. Advances and open problems in federated learning. Found Trends® Mach Learn. 2021;14(1–2):1–210. doi:10.1561/2200000083. [Google Scholar] [CrossRef]
23. McMahan B, Moore E, Ramage D, Hampson S, Arcas BA. Communication-efficient learning of deep networks from decentralized data. In: Artificial intelligence and statistics. London, UK: PMLR; 2017. p. 1273–82. [Google Scholar]
24. Zhao Y, Li M, Lai L, Suda N, Civin D, Chandra V. Federated learning with non-iid data. arXiv:1806.00582. 2018. [Google Scholar]
25. Ye M, Fang X, Du B, Yuen PC, Tao D. Heterogeneous federated learning: state-of-the-art and research challenges. ACM Comput Surv. 2023;56(3):1–44. doi:10.1145/3625558. [Google Scholar] [CrossRef]
26. Koutsoubis N, Yilmaz Y, Ramachandran RP, Schabath M, Rasool G. Privacy preserving federated learning in medical imaging with uncertainty estimation. arXiv:2406.12815. 2024. [Google Scholar]
27. Mahmood A, Sadique TK, Azzuhri SR, Ramli R, Ismail L. Federated performance-based averaging (FedPAa robust and selective learning framework for chest X-ray classification in heterogeneous data environments. Int J Adv Comput Sci Appl. 2025;16(10):940–52. doi:10.14569/ijacsa.2025.0161092. [Google Scholar] [CrossRef]
28. Azam MA, Khan KB, Salahuddin S, Rehman E, Khan SA, Khan MA, et al. A review on multimodal medical image fusion: compendious analysis of medical modalities, multimodal databases, fusion techniques and quality metrics. Comput Biol Med. 2022;144(3):105253. doi:10.1016/j.compbiomed.2022.105253. [Google Scholar] [PubMed] [CrossRef]
29. Sheller MJ, Edwards B, Reina GA, Martin J, Pati S, Kotrotsou A, et al. Federated learning in medicine: facilitating multi-institutional collaborations without sharing patient data. Sci Rep. 2020;10(1):12598. doi:10.1038/s41598-020-69250-1. [Google Scholar] [PubMed] [CrossRef]
30. Rieke N, Hancox J, Li W, Milletari F, Roth HR, Albarqouni S, et al. The future of digital health with federated learning. npj Digit Med. 2020;3(1):119. doi:10.1038/s41746-020-00323-1. [Google Scholar] [PubMed] [CrossRef]
31. Antunes RS, André da Costa C, Küderle A, Yari IA, Eskofier B. Federated learning for healthcare: systematic review and architecture proposal. ACM Trans Intell Syst Technol. 2022;13(4):1–23. doi:10.1145/3501813. [Google Scholar] [CrossRef]
32. Stamatis V, Radoglou-Grammatikis P, Sarigiannidis A, Pitropakis N, Lagkas T, Argyriou V, et al. Advancements in federated learning for health applications: a concise survey. In: 2024 20th International Conference on Distributed Computing in Smart Systems and the Internet of Things (DCOSS-IoT). Piscataway, NJ, USA: IEEE; 2024. p. 503–8. [Google Scholar]
33. Zhang C, Xie Y, Bai H, Yu B, Li W, Gao Y. A survey on federated learning. Knowl Based Syst. 2021;216(1):106775. doi:10.1016/j.knosys.2021.106775. [Google Scholar] [CrossRef]
34. Mishra S, Tondon R, Rathore NPS. Revolutionizing healthcare with federated learning: a comprehensive review. In: 2024 International Conference on Advances in Computing Research on Science Engineering and Technology (ACROSET). Cham, Switzerland: Springer; 2024. p. 1–5. [Google Scholar]
35. Abbas SR, Abbas Z, Zahir A, Lee SW. Federated learning in smart healthcare: a comprehensive review on privacy, security, and predictive analytics with IoT integration. Healthcare. 2024;12(24):2587. doi:10.3390/healthcare12242587. [Google Scholar] [PubMed] [CrossRef]
36. Raza A, Guzzo A, Ianni M, Lappano R, Zanolini A, Maggiolini M, et al. Federated learning in radiomics: a comprehensive meta-survey on medical image analysis. Comput Methods Programs Biomed. 2025;267(C):108768. doi:10.1016/j.cmpb.2025.108768. [Google Scholar] [PubMed] [CrossRef]
37. Bharati S, Rubaiyat M, Mondal H, Podder P, Prasath VBS. Federated learning: applications, challenges and future scopes. arXiv:2205.09513. 2022. [Google Scholar]
38. Ali MS, Ahsan MM, Tasnim L, Afrin S, Biswas K, Hossain MMS, et al. Federated learning in healthcare: model misconducts, security, challenges, applications, and future research directions—a systematic review. arXiv:2405.13832. 2024. [Google Scholar]
39. Liu B, Lv N, Guo Y, Li Y. Recent advances on federated learning: a systematic survey. Neurocomputing. 2024;597(4):128019. doi:10.1016/j.neucom.2024.128019. [Google Scholar] [CrossRef]
40. Wang S, Tuor T, Salonidis T, Leung KK, Makaya C, He T, et al. Adaptive federated learning in resource constrained edge computing systems. IEEE J Sel Areas Commun. 2019;37(6):1205–21. doi:10.1109/jsac.2019.2904348. [Google Scholar] [CrossRef]
41. Nakakita S, Kaneko T, Takamaeda-Yamazaki S, Imaizumi M. Federated learning with relative fairness. arXiv:2411.01161. 2024. [Google Scholar]
42. Zhou L, Wang M, Zhou N. Distributed federated learning-based deep learning model for privacy MRI brain tumor detection. arXiv:2404.10026. 2023. [Google Scholar]
43. Le TT, Pham NT, Tran PN, Dang DNM. Federated learning with U-net for brain tumor segmentation: impact of client numbers and data distribution. In: 2024 15th International Conference on Information and Communication Technology Convergence (ICTC). Piscataway, NJ, USA: IEEE; 2024. p. 2048–53. [Google Scholar]
44. Amgain S, Shrestha P, Bano S, Torres IV, Cunniffe M, Hernandez V, et al. Investigation of federated learning algorithms for retinal optical coherence tomography image classification with statistical heterogeneity. arXiv:2402.10035. 2024. [Google Scholar]
45. Subramanian M, Rajasekar V, VE S, Shanmugavadivel K, Nandhini P. Effectiveness of decentralized federated learning algorithms in healthcare: a case study on cancer classification. Electronics. 2022;11(24):4117. doi:10.3390/electronics11244117. [Google Scholar] [CrossRef]
46. Li X, Jiang M, Zhang X, Kamp M, Dou Q. Fedbn: federated learning on non-iid features via local batch normalization. arXiv:2102.07623. 2021. [Google Scholar]
47. Peng L, Luo G, Walker A, Zaiman Z, Jones EK, Gupta H, et al. Evaluation of federated learning variations for COVID-19 diagnosis using chest radiographs from 42 US and European hospitals. J Am Med Inform Assoc. 2023;30(1):54–63. doi:10.1093/jamia/ocac188. [Google Scholar] [PubMed] [CrossRef]
48. Park J, Lim H. Privacy-preserving federated learning using homomorphic encryption. Appl Sci. 2022;12(2):734. doi:10.3390/app12020734. [Google Scholar] [CrossRef]
49. MURA dataset: towards radiologist-level abnormality detection in musculoskeletal radiographs—stanfordmlgroup.github.io [Internet]. [cited 2025 Dec 9]. Available from: https://stanfordmlgroup.github.io/competitions/mura/. [Google Scholar]
50. Xie C, Koyejo S, Gupta I. Asynchronous federated optimization. arXiv:1903.03934. 2019. [Google Scholar]
51. Fadly F, Kurniawan TB, Dewi DA, Zakaria MZ, binti Hisham PAA. Deep learning based face mask detection system using MobileNetV2 for enhanced health protocol compliance. J Appl Data Sci. 2024;5(4):2067–78. [Google Scholar]
52. Zhang S, Zhao Z, Liu D, Cao Y, Tang H, You S. Edge-assisted U-shaped split federated learning with privacy-preserving for Internet of Things. Expert Syst Appl. 2025;262(10):125494. doi:10.1016/j.eswa.2024.125494. [Google Scholar] [CrossRef]
53. Nasajpour M, Pouriyeh S, Parizi RM, Han M, Mosaiyebzadeh F, Liu L, et al. Federated learning in smart healthcare: a survey of applications, challenges, and future directions. Electronics. 2025;14(9):1750. doi:10.3390/electronics14091750. [Google Scholar] [CrossRef]
54. Liu H, Zhou H, Chen H, Yan Y, Huang J, Xiong A, et al. A federated learning multi-task scheduling mechanism based on trusted computing sandbox. Sensors. 2023;23(4):2093. doi:10.3390/s23042093. [Google Scholar] [PubMed] [CrossRef]
55. Huang G, Liu Z, Van Der Maaten L, Weinberger KQ. Densely connected convolutional networks. In: Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition. Piscataway, NJ, USA: IEEE; 2017. p. 4700–8. [Google Scholar]
56. Mohammadi M, Shrestha R, Sinaei S. Integrating federated learning and differential privacy for secure anomaly detection in smart grids. In: Proceedings of the 2024 8th International Conference on Cloud and Big Data Computing. New York, NY, USA: ACM; 2024. p. 60–6. [Google Scholar]
Cite This Article
 Copyright © 2026 The Author(s). Published by Tech Science Press.
Copyright © 2026 The Author(s). Published by Tech Science Press.This work is licensed under a Creative Commons Attribution 4.0 International License , which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.


 Submit a Paper
Submit a Paper Propose a Special lssue
Propose a Special lssue View Full Text
View Full Text Download PDF
Download PDF Downloads
Downloads
 Citation Tools
Citation Tools