 Open Access
Open Access
ARTICLE
Proposed Framework for Detection of Breast Tumors
1 Department of Computer Science, Faculty of Computers and Informatics, Kafrelsheikh University, Kafrelsheikh, Egypt
2 Department of Computer Science, Faculty of Computers and Informatics, Zagazig University, Zagazig, Egypt
* Corresponding Author: Mostafa Elbaz. Email:
Computers, Materials & Continua 2023, 74(2), 2927-2944. https://doi.org/10.32604/cmc.2023.033111
Received 08 June 2022; Accepted 01 August 2022; Issue published 31 October 2022
Abstract
Computer vision is one of the significant trends in computer science. It plays as a vital role in many applications, especially in the medical field. Early detection and segmentation of different tumors is a big challenge in the medical world. The proposed framework uses ultrasound images from Kaggle, applying five diverse models to denoise the images, using the best possible noise-free image as input to the U-Net model for segmentation of the tumor, and then using the Convolution Neural Network (CNN) model to classify whether the tumor is benign, malignant, or normal. The main challenge faced by the framework in the segmentation is the speckle noise. It’s is a multiplicative and negative issue in breast ultrasound imaging, because of this noise, the image resolution and contrast become reduced, which affects the diagnostic value of this imaging modality. As result, speckle noise reduction is very vital for the segmentation process. The framework uses five models such as Generative Adversarial Denoising Network (DGAN-Net), Denoising U-Shaped Net (D-U-NET), Batch Renormalization U-Net (Br-U-NET), Generative Adversarial Network (GAN), and Nonlocal Neutrosophic of Wiener Filtering (NLNWF) for reducing the speckle noise from the breast ultrasound images then choose the best image according to peak signal to noise ratio (PSNR) for each level of speckle-noise. The five used methods have been compared with classical filters such as Bilateral, Frost, Kuan, and Lee and they proved their efficiency according to PSNR in different levels of noise. The five diverse models are achieved PSNR results for speckle noise at level (0.1, 0.25, 0.5, 0.75), (33.354, 29.415, 27.218, 24.115), (31.424, 28.353, 27.246, 24.244), (32.243, 28.42, 27.744, 24.893), (31.234, 28.212, 26.983, 23.234) and (33.013, 29.491, 28.556, 25.011) for DGAN, Br-U-NET, D-U-NET, GAN and NLNWF respectively. According to the value of PSNR and level of speckle noise, the best image passed for segmentation using U-Net and classification using CNN to detect tumor type. The experiments proved the quality of U-Net and CNN in segmentation and classification respectively, since they achieved 95.11 and 95.13 in segmentation and 95.55 and 95.67 in classification as dice score and accuracy respectively.Keywords
Recently, due to the rapid development of using computer vision technologies in medical imaging, a wide range of clinicians depend on computer vision and technology to make examination, analysis, and provide treatment based on medical images such as Magnetic Resonance Imaging (MRI), computed tomography (CT), and ultrasounds [1,2]. Medical images can be used for the treatment of different organ tumors such as tumors in the brain, breast, liver, cardiac, etc. Early detection of tumors in these organs is a very vital task for clinicians in the process of examination and treatment.
Deep learning (DL) is a significant trend in the medical field for examination in different organs and medical field such as radiology, pathology, cardiology, pharmacology, etc. [3,4]. DL has been presented good model in detecting breast cancer with high degree of accuracy [4]. DL has been presented many models to address the problem of detecting starting from the process of image enhancement, segmentation and classification the tumors such CNN, Fully Convolutional Network (FCN), GAN, neural network, etc.
The framework of detection of breast cancer is composed of a set of stages such as image enhancement, segmentation, feature, and classification. The result and accuracy in each stage affect the next stage, for example, the accuracy of the process of enhancement can change the results of segmentation positively or negatively. Image segmentation means dividing the medical image into a set of disjoint parts to simplify the process of examination and detection. The segmentation process helps clinicians detect objects such as the different types of tumors [5]. Segmentation of breast tumors is one of the most famous tasks during the recent increase in the number of cases [6]. Ultrasound images are a perfect solution for the segmentation of breast tumors because it is cost-effective, compact, and portable [7], but like anything in life, Ultrasound has a set of shortages such as speckle noise due to the method [8]. This type of noise must be removed before the segmentation step. There are many types of algorithms are being developed to address the problem of speckle [9,10]
This paper presents a new framework for the detection of breast tumors from ultrasound images that have speckle noise. The proposed framework contains a set of stages such as image enhancement, image segmentation, and classification. The Framework starts with image enhancement to remove the unnecessary parts in the ultrasound image. Removing the noise step helps to reduce loss of highly important information in ultrasound images, especially in small images [11–14]. There are many algorithms was developed to handle this problem such as Frost [14], Bilateral [15], Kuan [16], Lee [17], Mean, and Median Filters, wide inference parch residual patch (WIN5-RB) [18], Denoising Prior Driven Deep Neural Network (DPDNN) [19], Convolution Autoencoder Denoising Network (Di-Conv-AE-Net), Br-U-NET [7] and etc. The framework uses five different modified models such as DGAN-Net, D-U-NET, Br-U-NET to remove the speckle noise in the stage of image enhancement and then compare between the results to pass the most efficient image to segmentation stage.
The second stage in the framework is the segmentation stage and classification stage. The segmentation and classification stage depends on the previous enhancement stage. Recently, DL becomes a trend in dealing with medical segmentation for different tumors [20–23]. In DL, there are many architectures have been developed to deal with breast tumors and other tumors such as CNNs FCN, autoencoder, U-Net, etc. Ultrasound segmentation using DL models has a set of disadvantages such as, in FCN architecture, there are some drawbacks such as the low accuracy in small and irregular objects. U-Net was introduced to overcome the disadvantages of FCN architecture. The u-Net architecture was developed to combine the high-level features from the decoder with the low-level features from the encoder. U-Net can achieve high accuracy in small datasets [24,25]. For the previous reasons, the framework uses U-Net architecture to handle the segmentation process in ultrasound after removing the speckle noise and CNN for classifying the results into malignant, begin, and normal classes.
The contributions can be summarized as follows:
1. The framework uses new different five methods for image enhancement such as DGAN-Net, D-U-NET, Br-U-NET, GAN, and a new neutrosophic filter to reduce the speckle noise from the ultrasound images and the experiments have proved the efficiency of choosing these models against classical methods.
2. Reducing the noise and choosing the best free-noise images depending on PSNR values in different levels of noise helps many classifications and segmentation models to maximize their accuracies.
3. The framework achieves good accuracy in segmentation and classification of different level of noise
The remaining part of paper organized as follows: Section 2 describes the methodology of framework parts such as image enhancement using different methods, image segmentation and classification, Section 3 discussion and results shows the results of each stage in the methodology. Section 4 conclusion and future work.
The methodology steps organized as follow:
Step 1: image acquisition: The framework has been used 600 ultrasound images for women in age from 18 to 75.
Step 2: Image preprocessing step: in this step we use different filter, and DL models such as DGAN-Net, D-U-NET, Br-U-NET and neutrosophic for image enhancement such as removing speckle noise the choose the best outputted image for be used in the stage of segmentation.
Step 3: Feature Extraction and Image segmentation step: The final step in methodology is using modified U-Net model to segment the image after removing the noise using the neutrosophic filter.
Step 4: The final step is used the CNN to classify the output to malignant or begin class. Fig. 1 shows the flowchart of the framework steps.
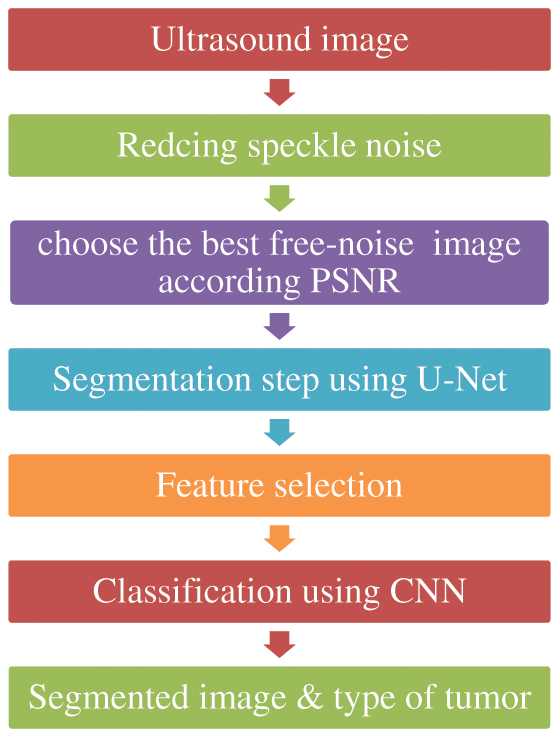
Figure 1: Methodology diagram
The pseudocode of the methodology as the follow:
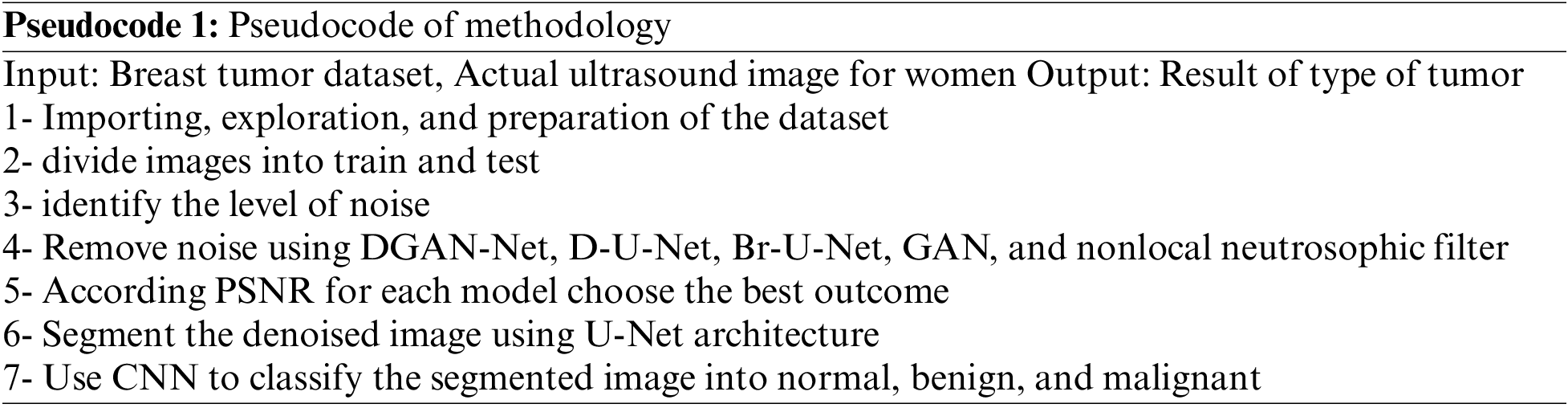

The paper used a dataset from Kaggle called breast ultrasound image dataset, which is being used for segmentation and detection. The image in the used dataset is collected for women of different and various ages in the range between 18 and 75. The dataset contains three types of ultrasound images benign tumor, malignant tumor, and normal. The dataset contains around 750 images in portable network graphics (PNG) format with size 500 * 500 [25]. The images are divided into two parts images for tumors and the masks Fig. 2 shows some results from the exploration of the five images from the dataset.
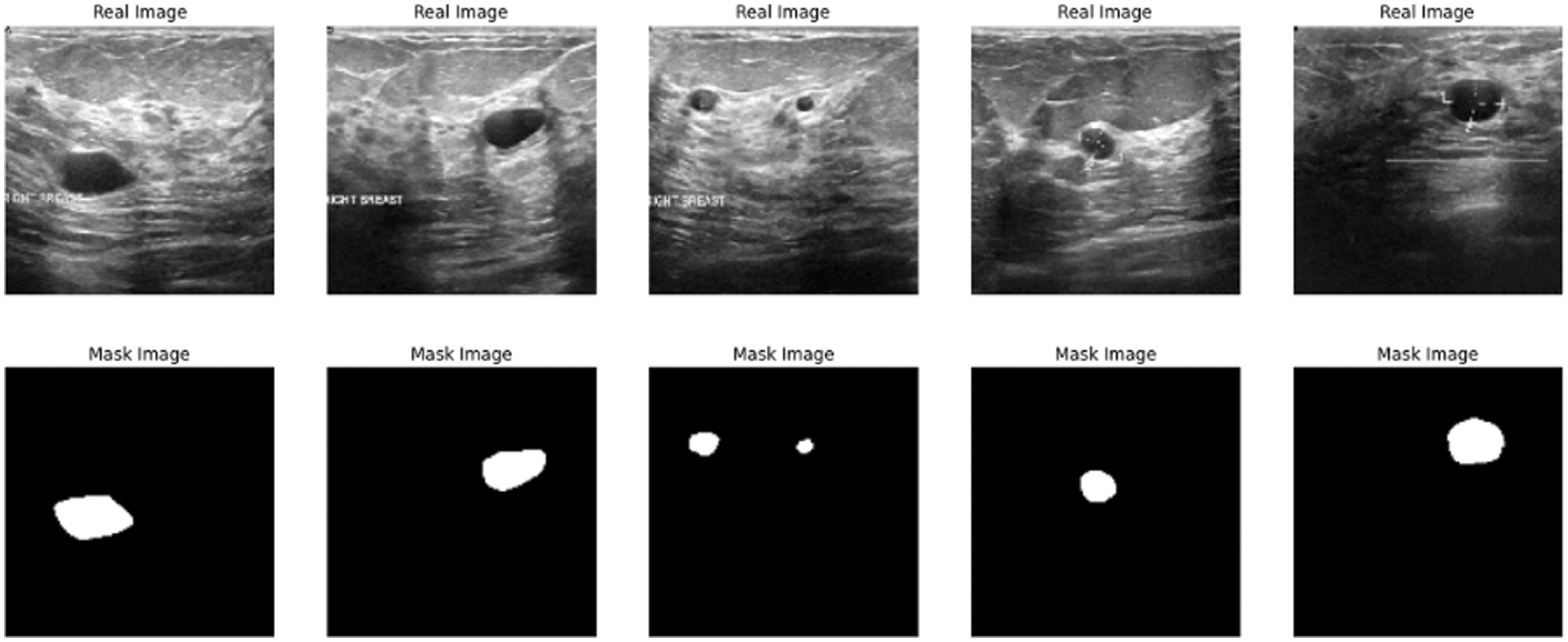
Figure 2: Data exploration of dataset
This step shows the algorithms used in the process of image enhancement especially in removing speckle noise steps such as NLNWF, U-Net enhanced model, Br-U-Net, and DGAN-Net. The five models are being used to enhance the ultrasound image. This framework uses these models to deal with the noise at a different level. The framework compares the results of these models and the classical filter and chooses the best outcome according to the results of the evaluation.
In this step we add speckle noise to the previous dataset to be used in testing the efficiency of the different method. The image without noise expressed by G(x,y), f(x,y) express the image with noise and the multiplicative and additive noises are ηm(x,y) and ηa(x,y), respectively as mentioned in Eq. (1) [26].
2.2.2 Nonlocal Neutrosophic of Wiener Filtering
For neutrosophic-based ultrasound breast image removing speckle noise, the noisy ultrasound breast image is converted to neutrosophic set which is defined by degree of true, degree of indeterminacy, and degree of false, Like T, I, and F, respectively. The intermediacy degree is used to represent the degree of speckle noise in ultrasound images. In this paper, the operator is used on true set and false set to minimize the indeterminacy set and removing speckle noise from the image, as shown in Fig. 3.
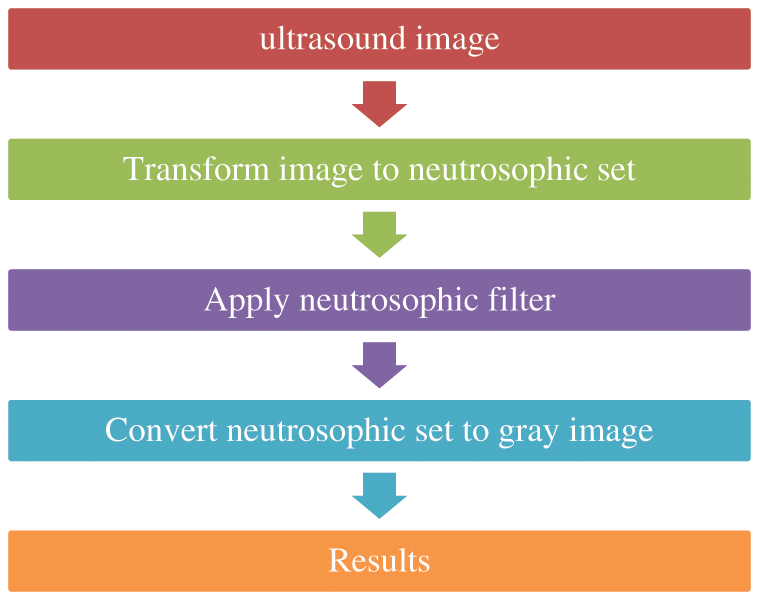
Figure 3: Block diagram of removing speckle noise using neutrosophic filter
The noisy image in ultrasound converted into neutrosophic sets where the image is defined by T, I, and F since T is the true set, I is the intermediate set and F is the false set, so the neutrosophic domain for image G(X,Y) = {T(x,y),I(X,Y),F(X,Y)} [27,28] which represented for image G as:
Since G’(x,y) is a local mean and
The entropy of the image calculated for true set, intermediacy set, false set and total entropy by the next equations respectively. The function p used to express the probabilities for element since entropy (ENT) is the total entropy of true set, ENI is the total entropy of intermediate set, ENF is the total entropy and En is the summation of the different types of entropies.
The next pseudocode shows the process of removing speckle noise from the breast ultrasound using the next pseudocode
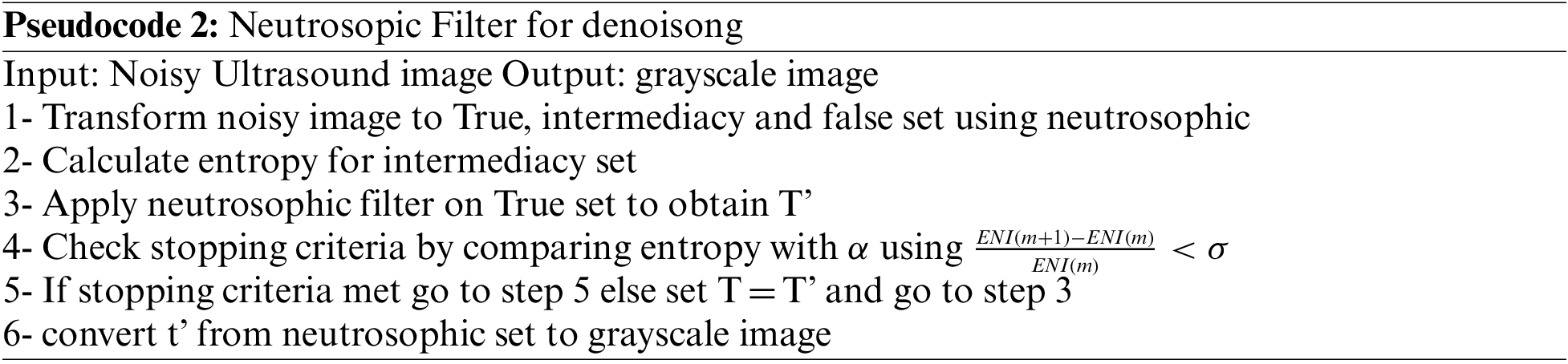

2.2.3 Generative Adversarial Network
Recent GAN applications can produce impressive results. The denoised network of GAN is composed of two parts, a generator, and a discriminator. The generator network used to generate new samples simulate the real samples, and the second network which is called the discriminator is used to differentiate between the noisy images and the free-noise ultrasound images [29]. The two networks combine to operate as adversaries. The generator part is used to increase errors and the discriminator to reduce them. It starts by reshaping the ultrasound image and using a set of upsamples layers then using flatten and the activation function. The discriminator uses convolutional layers to build a discriminative network. It is composed of an input convolutional layer and nine convolutional layers followed by batch normalization (BN) and Rectified Linear (ReLU). The output channels of consecutive convolutional layers are 64, 128, 256, 512, and 1. Therefore, when the input image is passed through each convolution block, the spatial dimension is decreased by a factor of two as shown in Fig. 4.
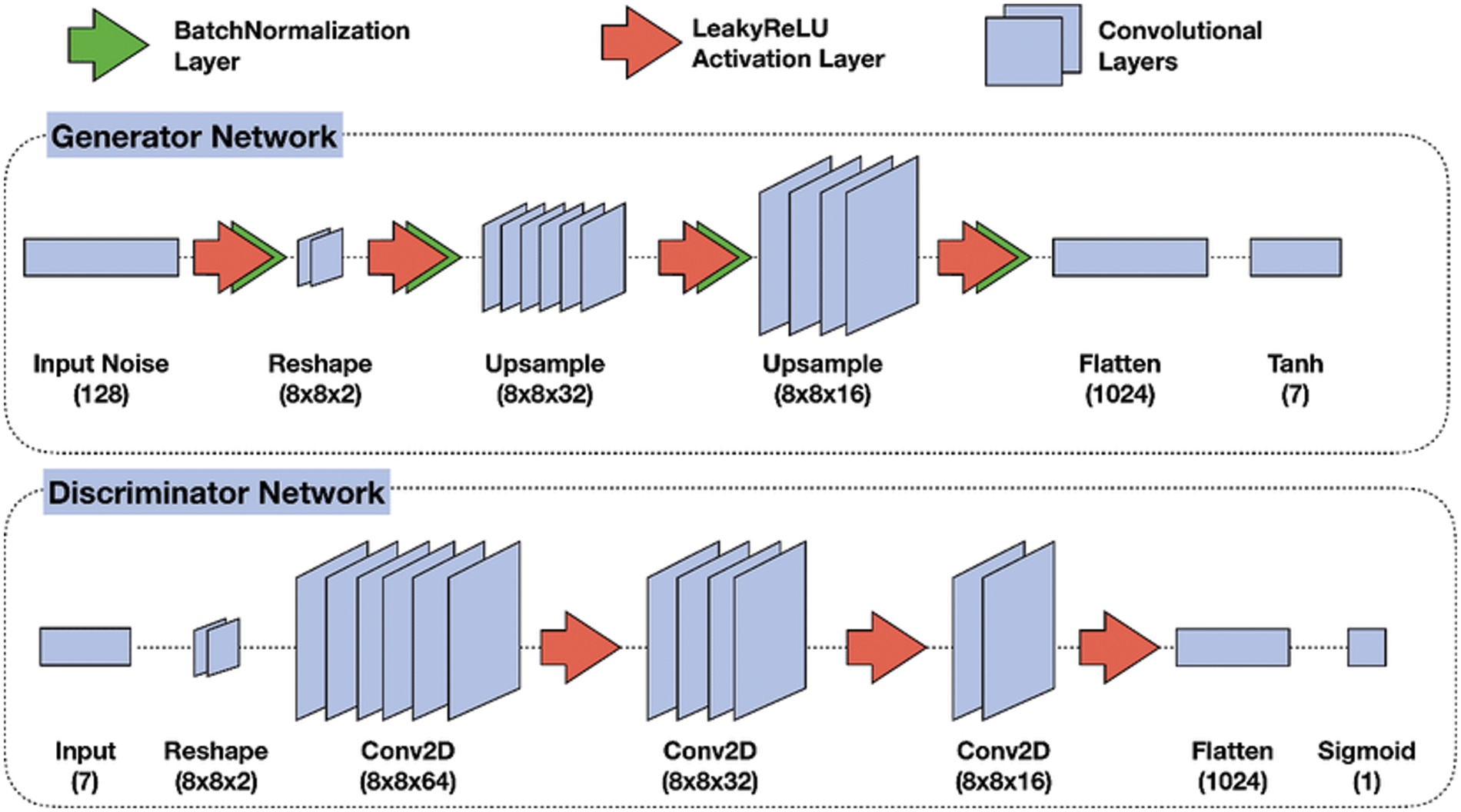
Figure 4: GAN model for removing speckle noise [30]
The structure of the U-Net model contains two separate parts the first one called contraction and the other part called expansion part as shown in the next figure. The two parts work separately but with connectivity between each other as shown in Fig. 5. The first part of U-Net is used for minimizing the speckle noise in the breast ultrasound using the convolution and carpooling layer to the minimized image. The other part of u-net used the minimized image from the previous part to the original image. In this modified U-Net or enhanced U-Net the, the model uses a convolution layer regarding using the upsampling layer [31].
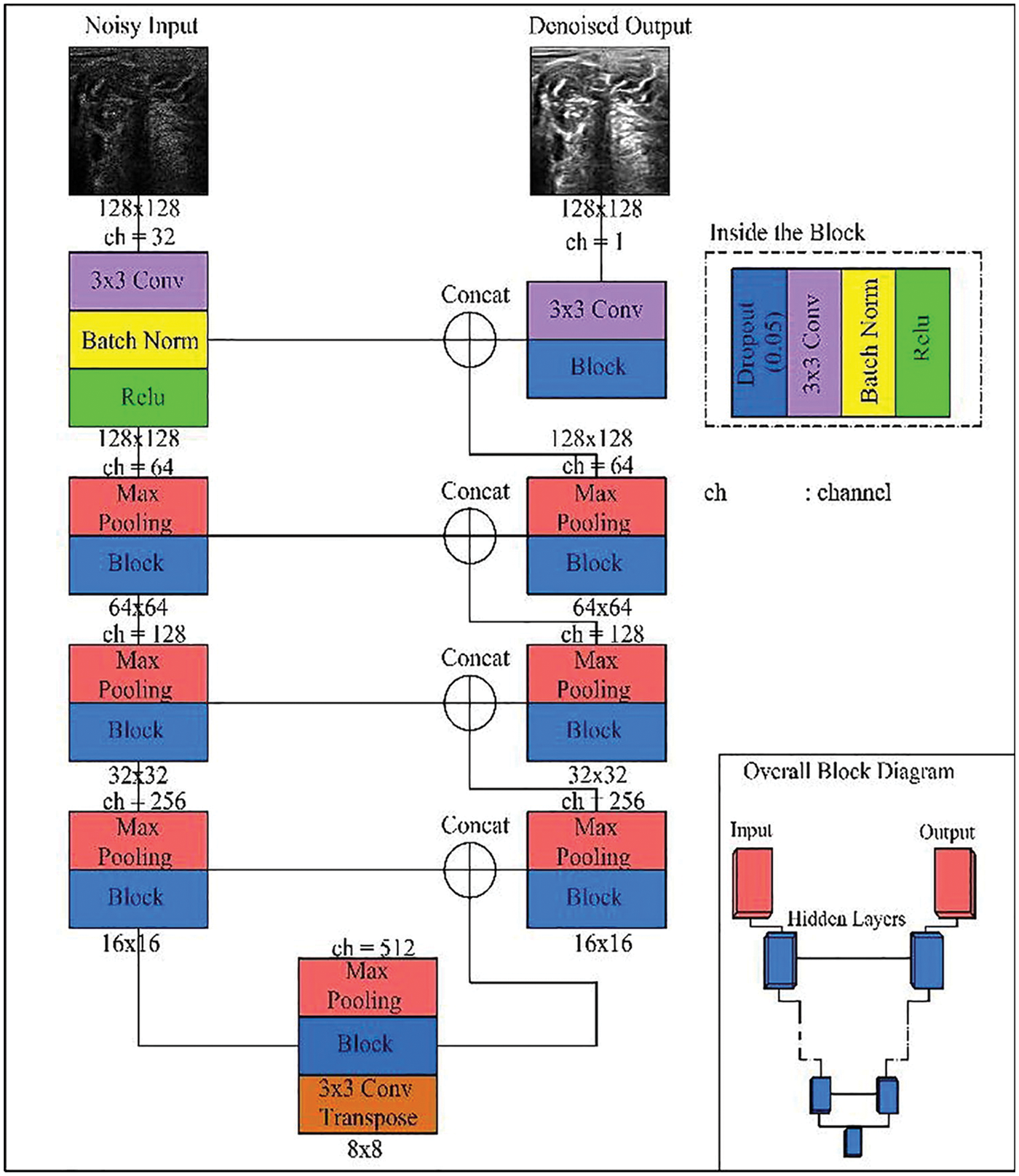
Figure 5: U-Net enhanced model [7]
2.2.5 Batch Renormalization U-Net
Br-U-Net model is the same structure as the U-Net model but with a renormalization, layer according to the use of batch normalization as shown in Fig. 6. In this model, the batch renormalization, the model uses fixed values for gamma and beta [32].
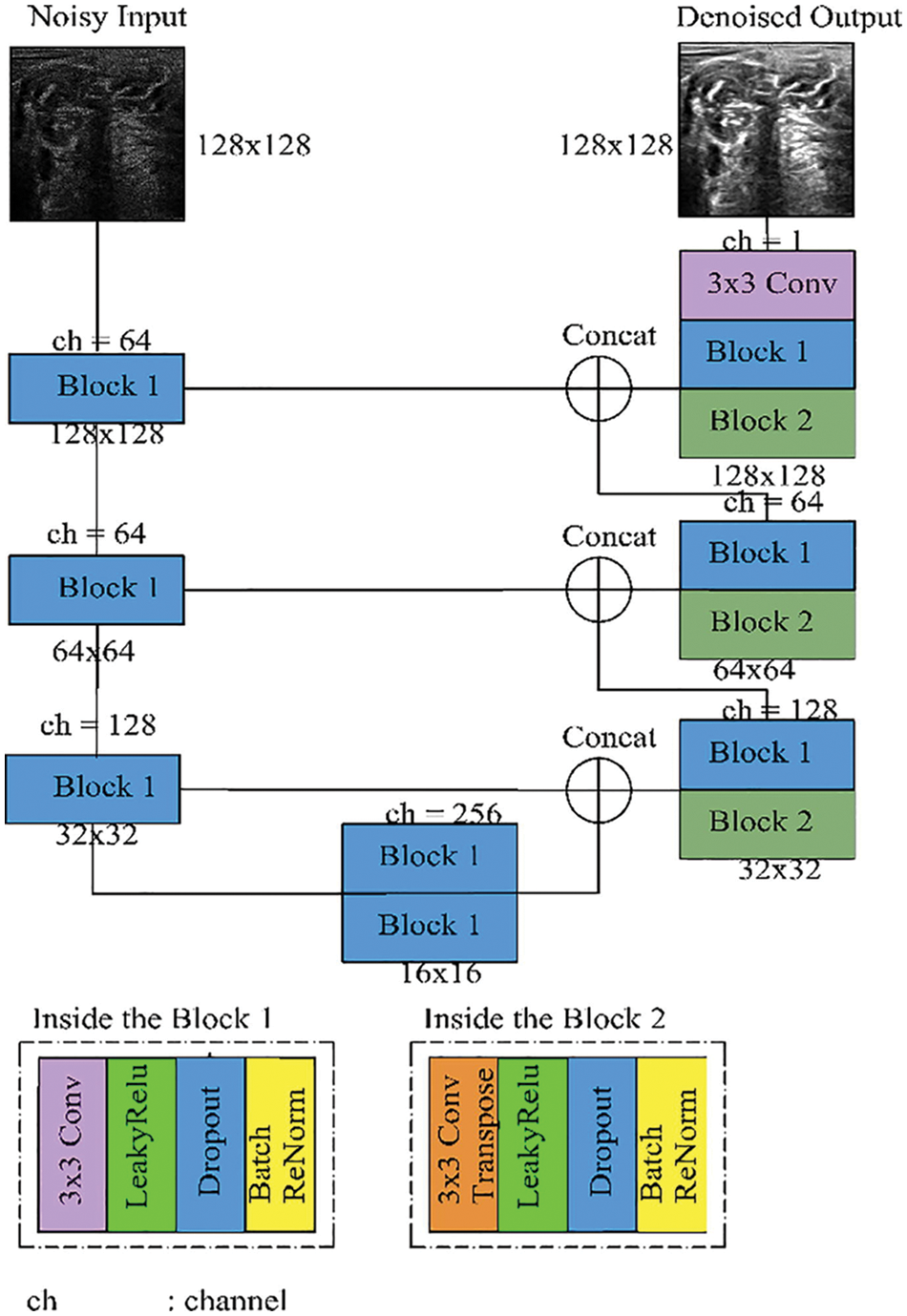
Figure 6: Batch renormalization U-Net model [7]
2.2.6 Generative Adversarial Denoising Network
DGAN-Net is a modified version of the classic GAN model. The structure of GAN contains two parts, The first part is called generator which is used to generate fake data, and the other part is called discriminator which is used to differentiate between the original and fake data [33] as shown in Fig. 7.
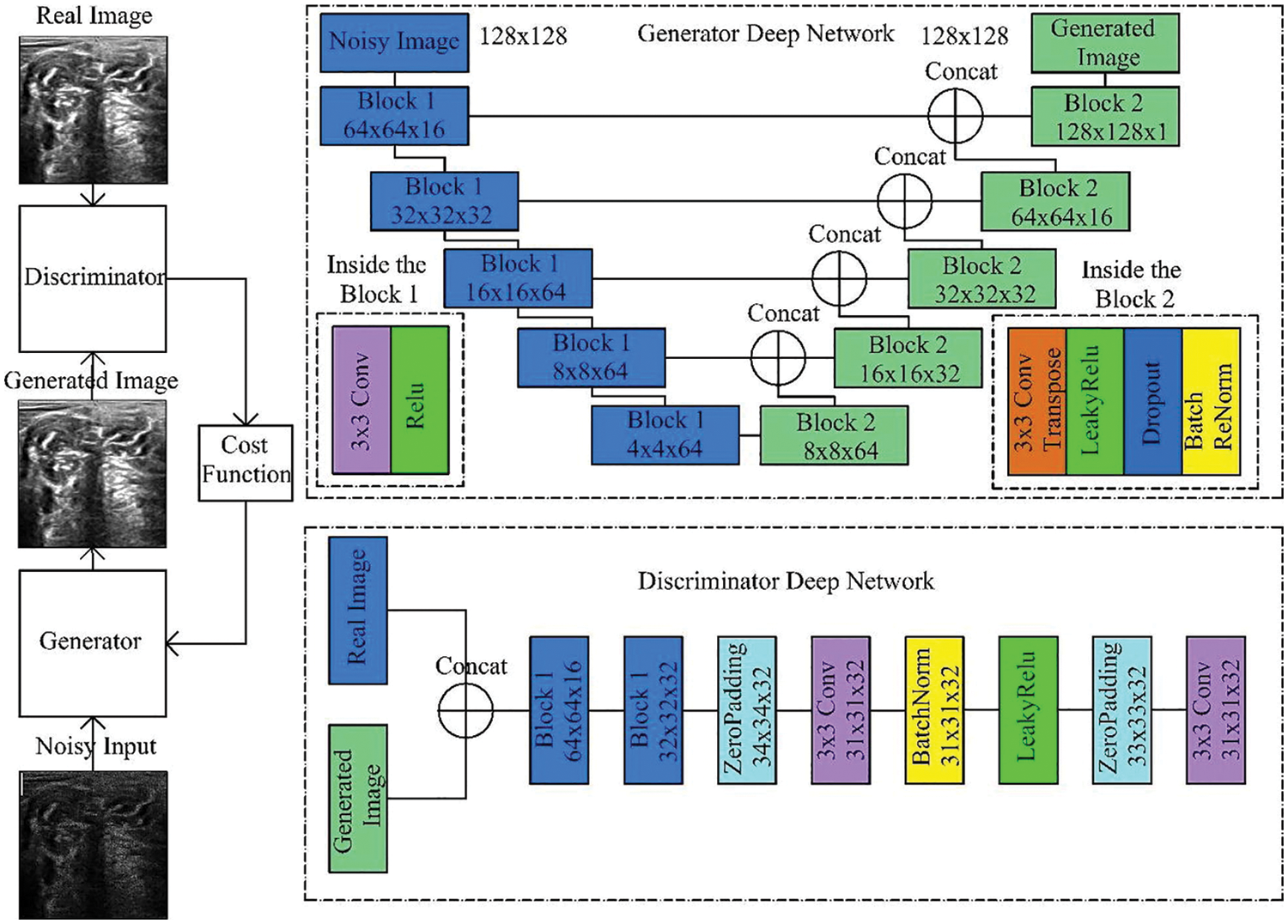
Figure 7: Generative adversarial denoising network [7]
2.3 Segmentation Step Using U-Net Model
In this step, the paper uses the U-Net for of segmentation of ultrasound images from the previous output of the neutrosophic denoising filter-Net takes the output from the neutrosophic filter (denoised ultrasound image) as input and produces a segmented image for the tumor. The architecture of the U-Net model consists of two parts encoder and decoder as shown in Fig. 8. The U-Net model can deal with medical images of any size and overlapping-tilling strategies are suitable for small pieces cutting using U-Net due to its network structure [34]. The first part which is called the encoder contains a stack of convolutional layers and maximum pooling layers and the second part is called the decoder and is used for localization. In the up-sampling portion, The model guarantees the architecture can spread background data through higher resolution layers across a wide range of feature channels [6]. The architecture of the U-Net model reduces ultrasound breast image 572 × 572 to 570 × 570 and finally to 568 × 568 to create unpadded convolution. The encoder block in the model multiplies the values by constant to minimize the size of ultrasound to help the max-pooling layers of strides 2. The skip connection part is used to preserve the loss from the previous layer.
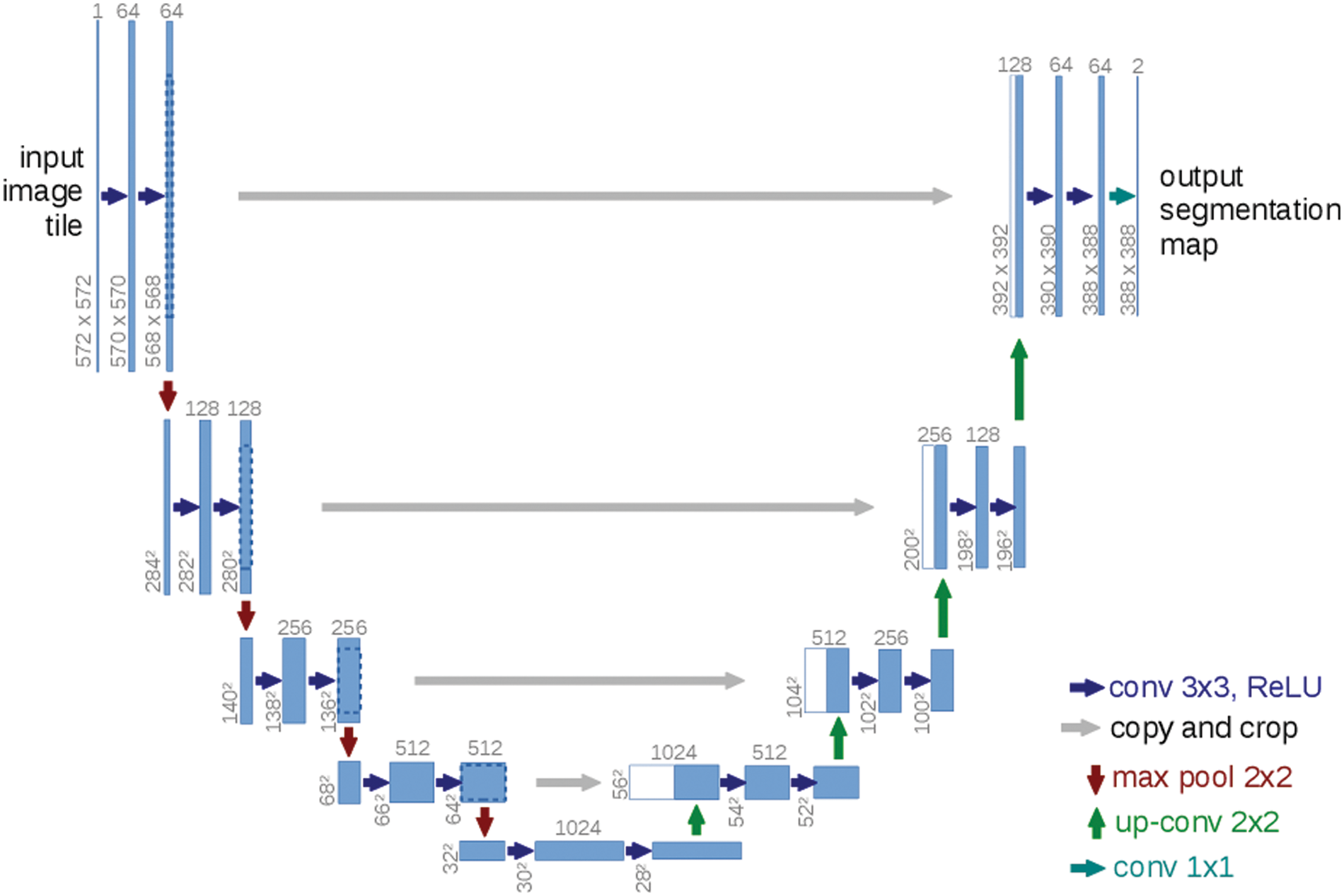
Figure 8: U-net architecture [20]
The final step in the methodology is using the segmented image from the previous step to divide the results into malignant and begin tumor. In this step the methodology uses the CNN to divide the tumors into three classes begin, normal and malignant.
2.5 Deep Learning Hyperparameters
Tab. 1 shows the hyperparameters which have been used in the framework.
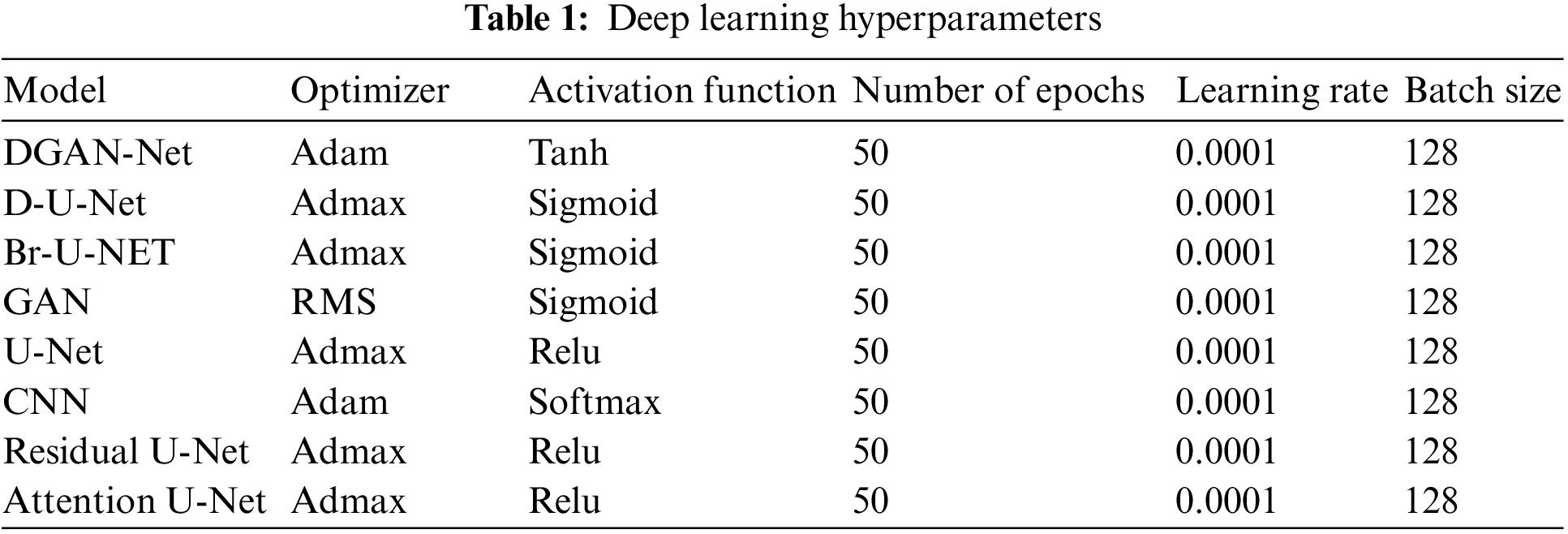

In this framework, 5 metrics are used to measure the performance of segmentation and classification models after reducing the noise using the five methods from the ultrasound images Dice Coefficient (DC), accuracy (ACC), Precision (PRE), Recall (REC), specificity (SPE), F1-score (F1).
This section shows the final results for each stage in the framework such as image enhancement using different modified method and choosing the most efficient for next stage, segmentation using U-Net model and the final classification process using CNN.
3.1 Results and Comparison of Removing Noise Step
In this subsection, the paper shows the results of comparison between different methods such as Frost [14], Bilateral [15], Kuan [16], Lee [17], Mean, and Median Filters, DGAN-Net, D-U-NET, GAN, Br-U-NET [7] and NLNWF. The comparison done using evaluation metrics PSNR and structural similarity index (SSIM). Tab. 2 shows the comparison done for five different level of noise .1, .25, .5 and .75 and other classical filters.
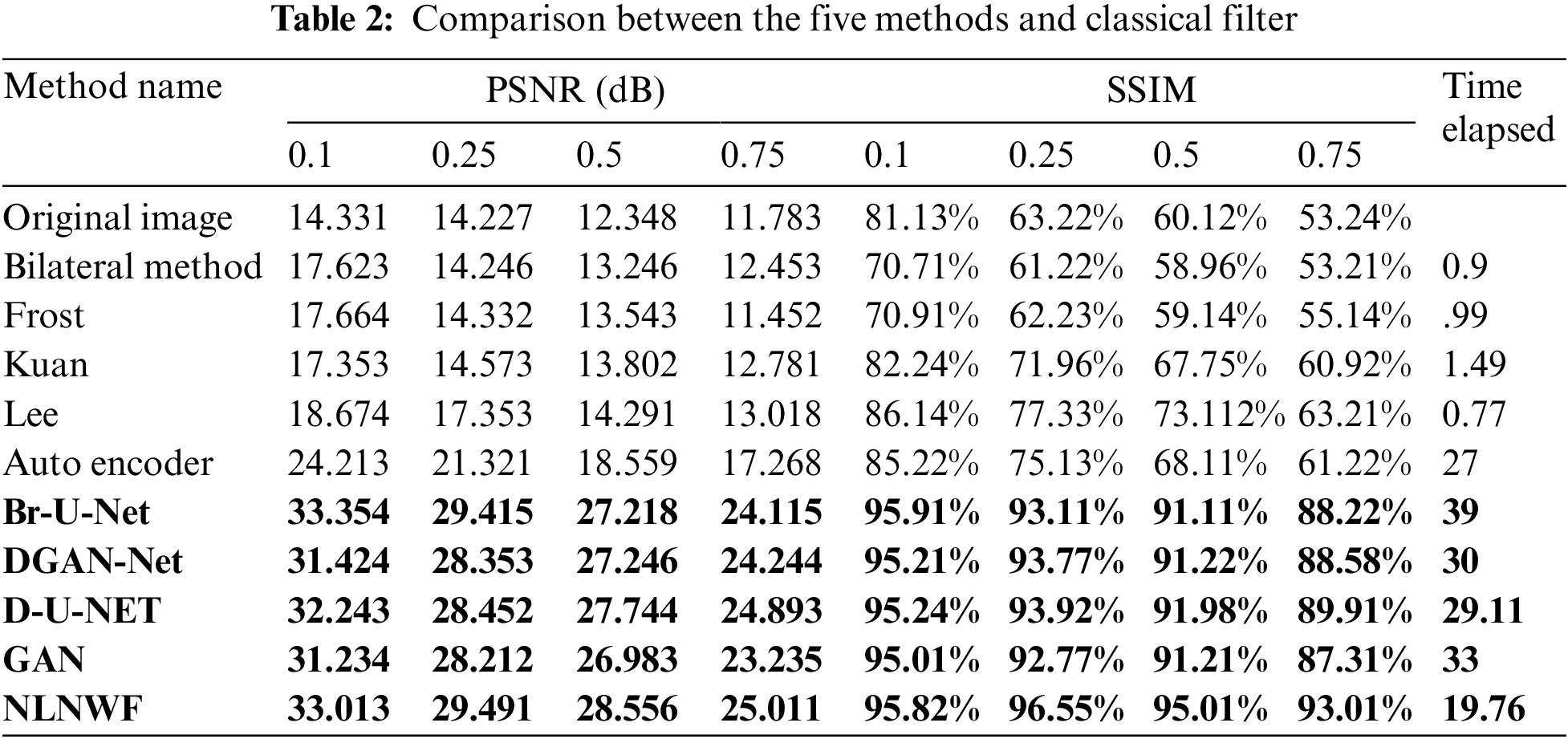
The bolded five models are the qualified five models for next stage, according to the level of noise and the value of PSNR the frame work choose the qualified ultrasound image for segmentation stage. Fig. 9 shows the comparison between the five modified models and the other available models filters according PSNR and SSIM. The results show the efficiency of the used five models in the framework in different level of noise.
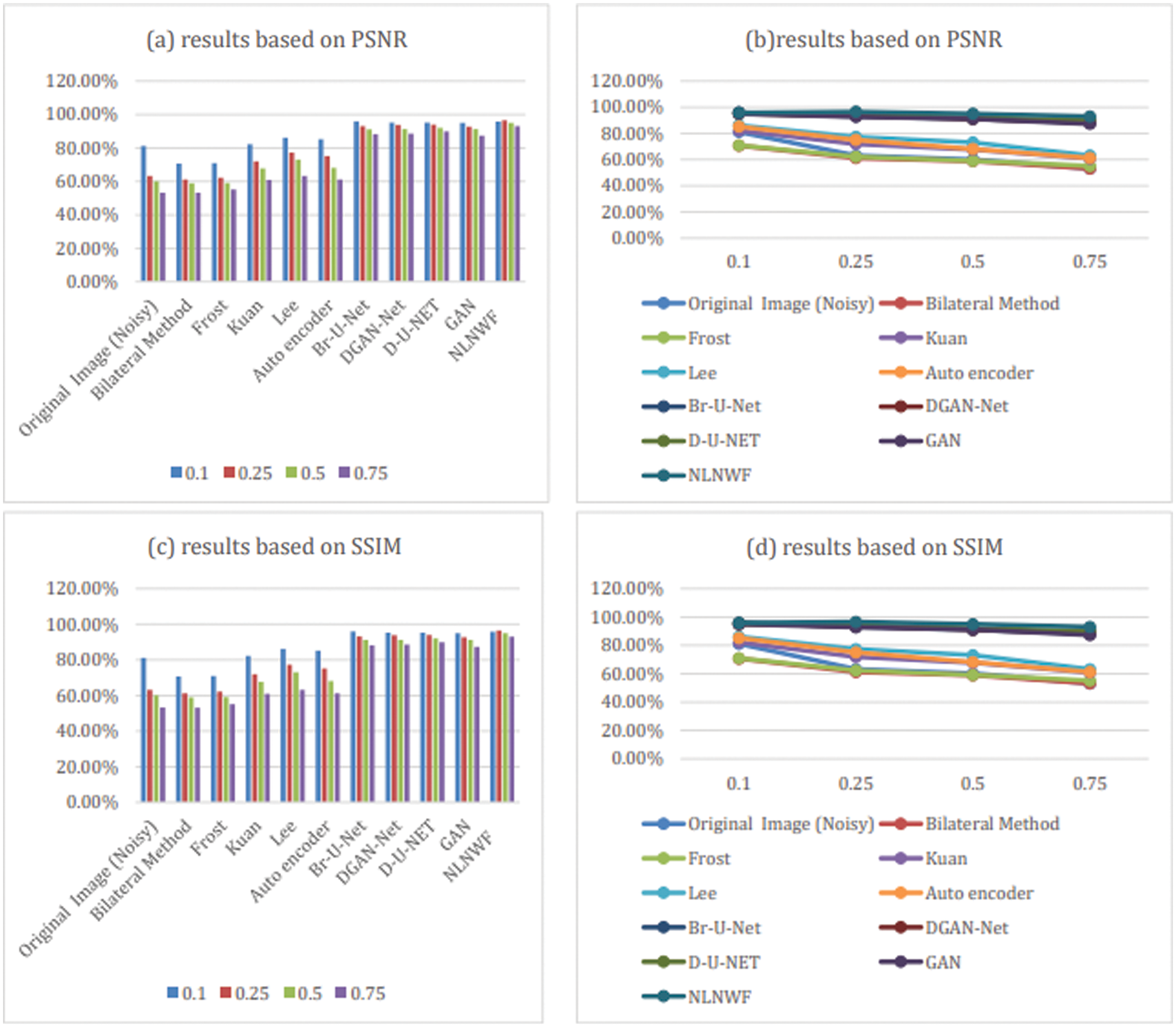
Figure 9: Comparison between models and classical filters based on PSNR and SSIM
3.2 Results of Segmentation Step
In this subsection, the framework shows the results after using U-Net architecture to segment the image after the process of denoising the image in the previous step. The U-Net model uses the result from the previous step after comparing the images of the five models and the level of noise. Tabs. 3 and 4 show the results after and before denoising the dataset of ultrasound, the results shows the enhancement of different metrics in the segmentation process after denoising the images, and the results also show the quality of u-net against the other methods, Fig. 10 shows the results of using U-Net in the segmentation process. The figure shows the ultrasound, mask, and predicted result using U-Net. Tabs. 5 and 6 show the results of classification of the segmented image into the malignant, tumor, and normal using different methods before and after removing noise, the results show the efficiency of methods after removing noise from images, and the results also show the efficiency of CNN against models. Fig. 11 shows the results of classifying the output into benign and malignant classes and Fig. 12 shows the results of removing noise in the enhancement stage.
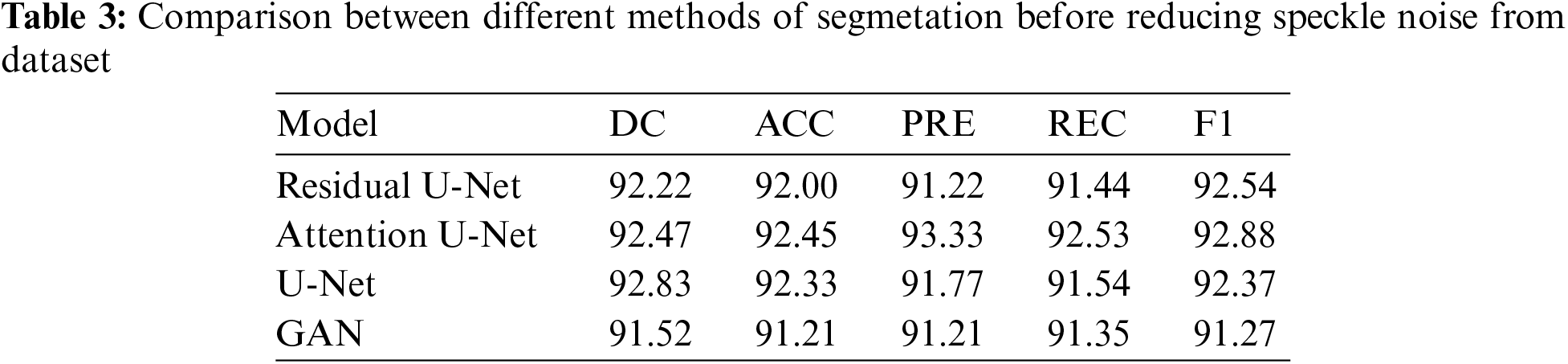
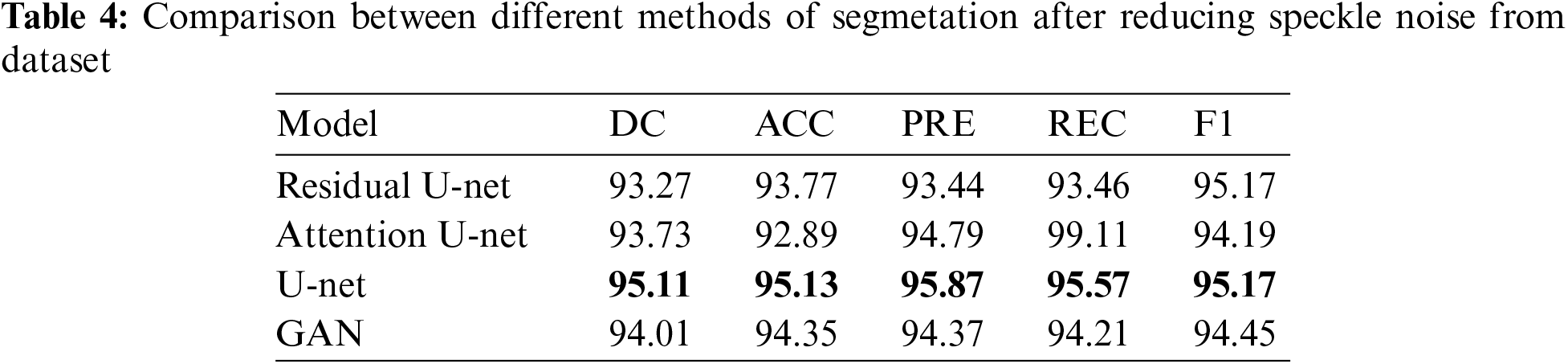
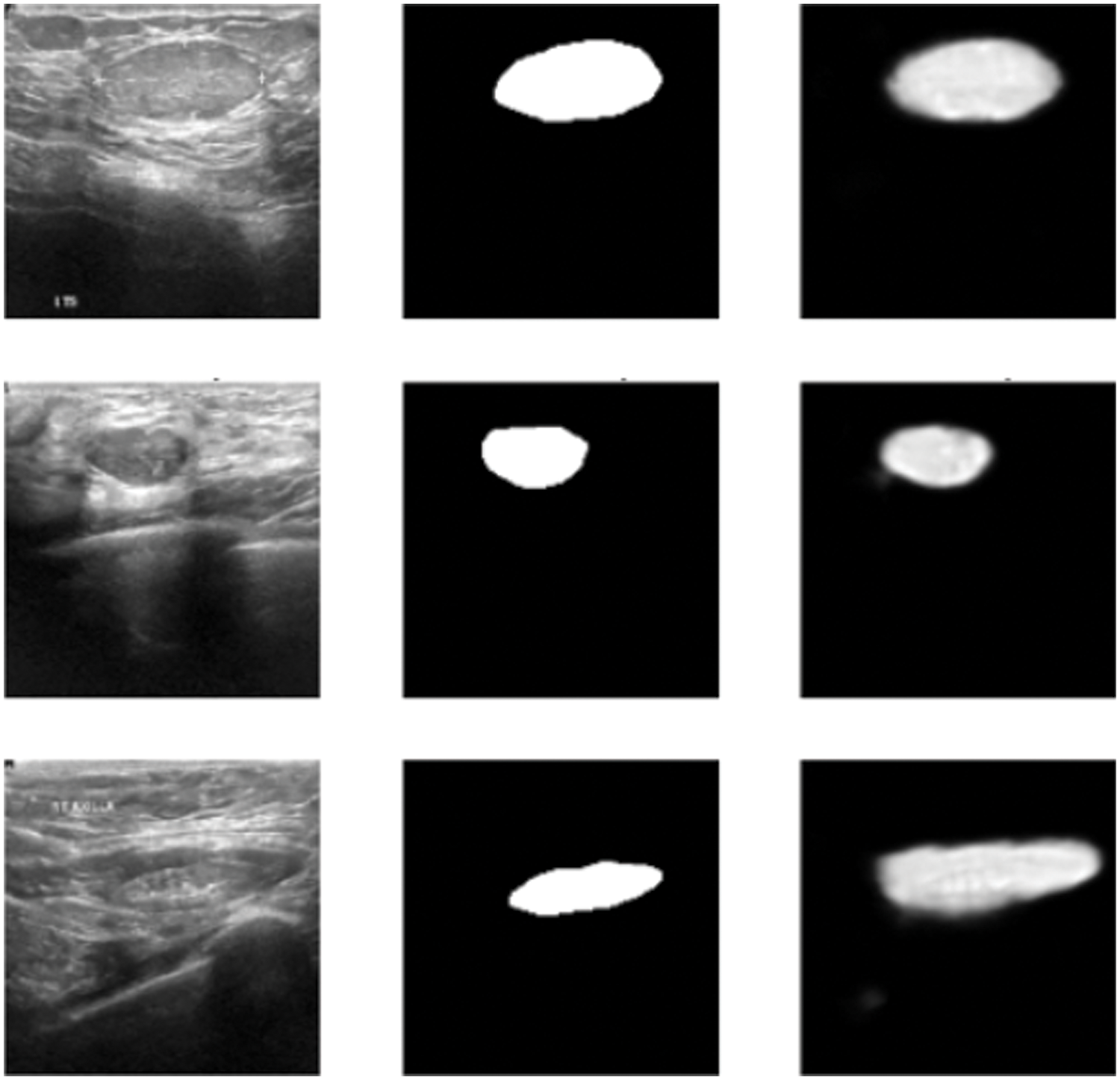
Figure 10: Original ultrasound, mask, predicted mask respectively
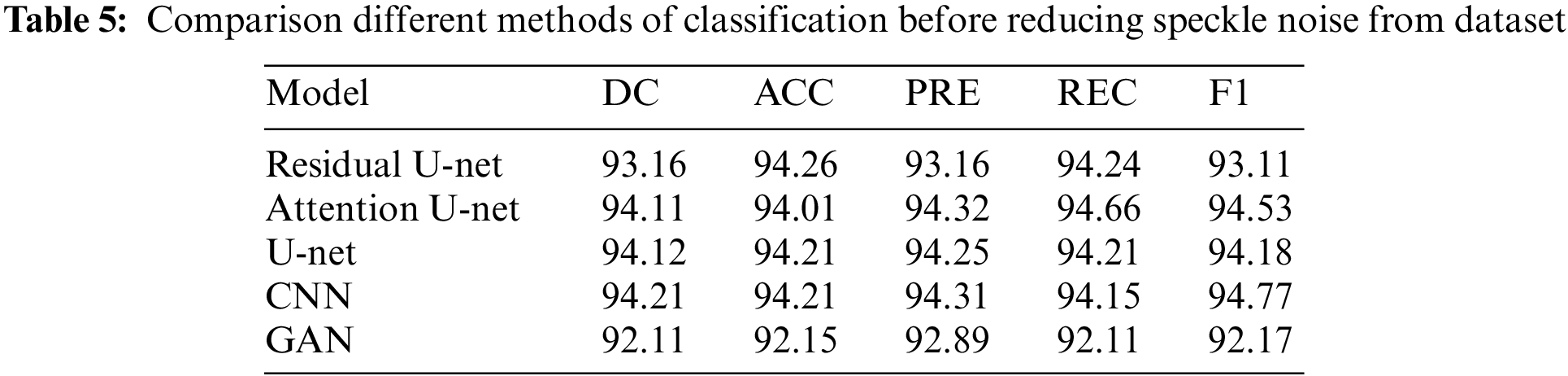
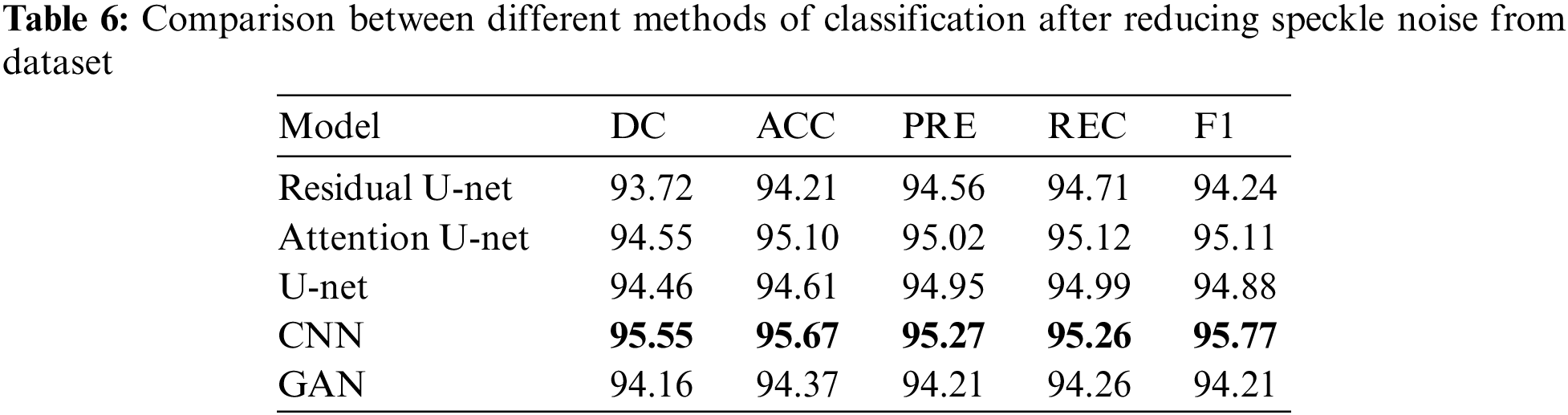
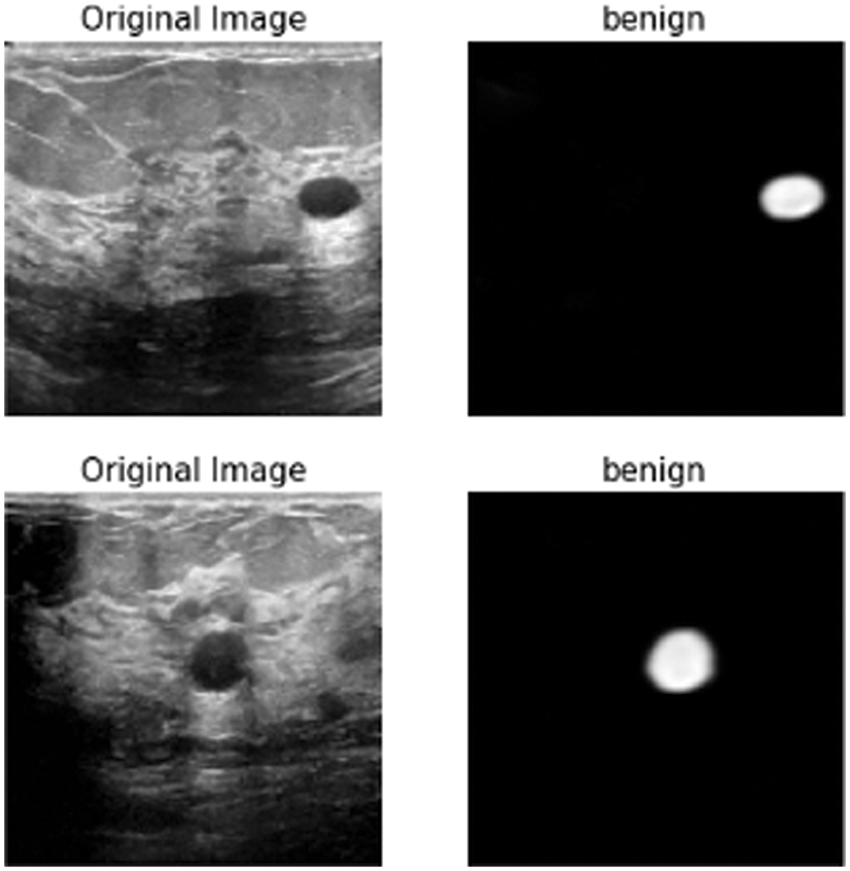
Figure 11: Class of tumor results
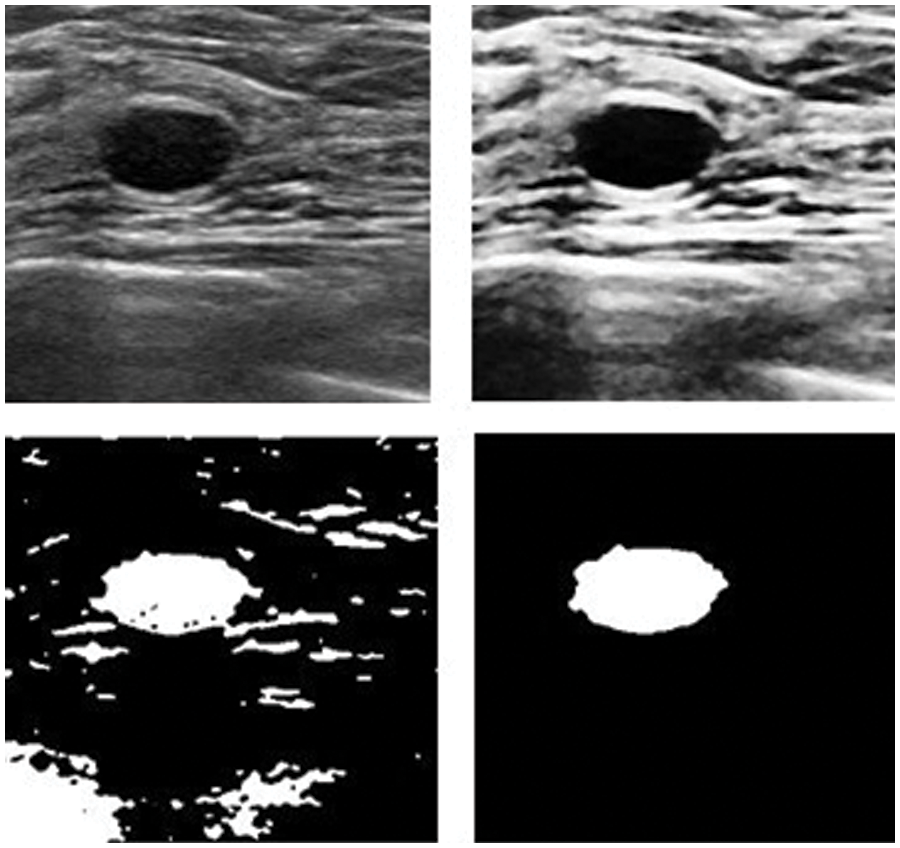
Figure 12: Results after and before removing noise
Early detection of breast tumors plays an important and urgent role in the process of examination and treatment for clinicians. The process of detection of breast tumors is divided into a set of stages starting from choosing and enhancing the image till the results and evaluation. Each stage in the process can affect the accuracy of the final results. The process of enhancement and the elimination of noise plays a vital role in the accuracy of segmentation. The accuracy of the segmentation process is also a challenge and the classification stage. The proposed framework presents good results in each stage, because it is compared good and modified algorithm in the stage of enhancement and choose the best free-noise one for the next stage. It uses many modified models in the stage of enhancement such as NLNWF, U-Net enhanced model, Br-U-Net, GAN, and DGAN-Net for speckle noise reduction. The stage of image enhancement made a good affection in the next stages such as segmentation using U-Net and classification stage using CNN in the accuracy of segmentation and classification. The experiments showed the efficiency of the five models in noise reduction and the efficiency of reducing the noise in the segmentation process. It also shows the difference in different accuracy matrices after and before reducing the noise. The drawback of U-Net is the number of parameters which has been used during the process of training, so in the future, reducing the number of the parameter must be used. Transfer learning will be used to enhance the process of training by additional pre-processed data.
Funding Statement: The authors received no specific funding for this study.
Conflicts of Interest: The authors declare that they have no conflicts of interest to report regarding the present study.
References
1. M. Shams, O. Elzeki, L. Abou El-Magd, A. E. Hassanien, M. Abdelfatah et al., “A healthy artificial nutrition analysis model during COVID-19 pandemic,” Computers in Biology and Medicine, vol. 135, no. 4, pp. 104606–104620, 2021. [Google Scholar]
2. M. F. Mridha, M. A. Hamed, M. M. Monor, A. G. Keya, A. Q. Ohi et al., “A comprehensive survey on deep learning-based breast cancer diagnosis,” Cancers Basel, vol. 13, no. 23, pp. 6116–6130, 2021. [Google Scholar]
3. L. Abou El-Magd, M. Shams, N. El-Attar and A. Hassanien, “Feature selection based coral reefs optimization for breast cancer classification,” Medical Informatics and Bio Imaging Using Artificial Intelligence, vol. 1004, no. 12, pp. 53–72, 2022. [Google Scholar]
4. R. Aggarwal, V. Sounderajah, G. Martin, D. S. Ting, A. Karthikesalingam et al., “Diagnostic accuracy of deep learning in medical imaging: A systematic review and meta-analysis,” Ithenticate Plagiarism Detection Digital Medicine, vol. 4, no. 1, pp. 65–80, 2021. [Google Scholar]
5. J. Li, M. Erdt, F. Janoos, T. Chang and J. Egger, “Medical image segmentation in oral-maxillofacial surgery,” Computer Aided Oral and Maxillofacial Surgery, vol. 12, no. 2, pp. 1–27, 2021. [Google Scholar]
6. M. Robin, J. John and A. Ravikumar, “Breast tumor segmentation using U-NET,” in Proc. of the 5th Int. Conf. on Computing Methodologies and Communication (ICCMM 2021), Erode, India, pp. 1164–1167, 2021. [Google Scholar]
7. O. Karaoglu, H. S. Bilge and I. Uluer, “Removal of speckle noises from ultrasound images using five different deep learning networks,” Engineering Science and Technology, an International Journal, vol. 29, no. 1, pp. 101030–101045, 2022. [Google Scholar]
8. M. Jayaraman, K. Vellingiri and Y. Guo, “Neutrosophic set in medical image denoising,” Neutrosophic Set in Medical Image Analysis, vol. 1, no. 2, pp. 77–100, 2019. [Google Scholar]
9. A. G. Rudnitskii and M. A. Rudnytska, “Segmentation and denoising of phase contrast MRI image of the aortic lumen via fractal and morphological processing,” in Proc. of the 37th Int. Conf. on Electronics and Nanotechnology (ELNANO 2017), Kiev, Ukraine, pp. 344–348, 2017. [Google Scholar]
10. S. Karthikeyan, T. Manikandan, V. Nandalal, J. L. Mazher Iqbal and J. J. Babu, “A survey on despeckling filters for speckle noise removal in ultrasound images,” in Proc. of the 3rd Int. Conf. on Electronics, Communication and Aerospace Technology (ICECA 2019), Tamil Nadu, India, pp. 605–609, 2019. [Google Scholar]
11. H. Wang, C. Wu, J. Chi, X. Yu and Q. Hu, “Speckle noise removal in ultrasound images with stationary wavelet transform and canny operator,” in Proc. of the Chinese Control Conf. (CCC 2019), Guangzhou, China, pp. 7822–7827, 2019. [Google Scholar]
12. S. Wang, T. Z. Huang, X. L. Zhao, J. Mei and J. Huang, “Speckle noise removal in ultrasound images by first-and second-order total variation,” Numerical Algorithms, vol. 78, no. 1, pp. 513–533, 2018. [Google Scholar]
13. N. Rahimizadeh, R. Hasanzadeh and F. J. Sharifi, “An optimized non-local LMMSE approach for speckle noise reduction of medical ultrasound images,” Multimedia Tools Applications, vol. 80, no. 1, pp. 1–23, 2021. [Google Scholar]
14. V. S. Frost, J. A. Stiles, K. S. Shanmugan and J. C. Holtzman, “A model for radar images and its application to adaptive digital filtering of multiplicative noise,” IEEE Transaction Pattern Analysis Machine Intelligence, vol. 4, no. 2, pp. 157–166, 1982. [Google Scholar]
15. M. Elad, “On the origin of the bilateral filter and ways to improve it,” IEEE Transction of Image Processing, vol. 11, no. 10, pp. 1141–1151, 2002. [Google Scholar]
16. D. T. Kuan, A. A. Sawchuk, T. C. Strand and P. Chavel, “Adaptive noise smoothing filter for images with signal dependent noise,” IEEE Transaction Pattern Analysis Machine Intelligence, vol. 7, no. 2, pp. 165–177, 1985. [Google Scholar]
17. J. -S. Lee, “Speckle analysis and smoothing of synthetic aperture radar images,” Computers Graphics and Image Procesing, vol. 17, no. 1, pp. 24–32, 1981. [Google Scholar]
18. P. Liu and R. Fang, “Learning pixel-distribution prior with wider convolution for image denoising,” arXiv Prepr.arXiv1707.09135,vol. 20, no. 2, pp. 1–21, 2017. [Google Scholar]
19. W. Dong, P. Wang, W. Yin, G. Shi, F. Wu et al., “Denoising prior driven deep neural network for image restoration,” IEEE Traniction on Pattern Analysis and Machine Intellegence, vol. 29, no. 12, pp. 27237–27253, 2018. [Google Scholar]
20. O. Ronneberger, P. Fischer and T. Brox, “U-Net: Convolutional networks for biomedical image segmentation,” in Proc. in the Int. Conf. on Medical Image Computing and Computer-Assisted Intervention (MICCAI 2017), Quebec, Canada, pp. 234–241, 2015. [Google Scholar]
21. X. Yao, X. Wang, S. H. Wang and Y. D. Zhang, “A comprehensive survey on convolutional neural network in medical image analysis,” Multimedia Tools and Application, vol. 25, no. 12, pp. 1–45, 2020. [Google Scholar]
22. L. Alzubaidi, J. Zhang, A. J. Humidi, Y. Duan, O. Alshamma et al., “Review of deep learning: Concepts, CNN architectures, challenges, applications, future directions,” Journal of Big Data, vol. 8, no. 1, pp. 53–69, 2021. [Google Scholar]
23. O. Ronneberger, P. Fischer and T. Brox, “U-Net convolutional networks for biomedical image segmentation,” in Proc. of the Medical Image Computing and Computer Assisted Intervention (MICCAI 2015), Springer, Cham, pp. 234–241, 2015. [Google Scholar]
24. A. Baccouche, B. G. Zapirain, C. C. Olea and A. S. Elmaghraby, “Connected-UNets: A deep learning architecture for breast mass segmentation,” Breast Cancer Research Journal, vol. 7, no. 1, pp. 151–166, 2021. [Google Scholar]
25. W. Dhabyani, M. Gomaa, H. Khaled and A. Fahmy, “Dataset of breast ultrasound images,” Data in Brief, vol. 28, no. 1, pp. 104863–104863, 2020. [Google Scholar]
26. X. Y. Song, S. Zhang, K. O. Song, W. Yang and Y. Z. Chen, “Speckle supression for medical ultrasound images based on modelling speckle with Rayleigh distribution in contourlet domain,” in Proc. of the Int. Conf. on Wavelet Analysis and Pattern Recognition (ICWAPR 2008), Hong Kong, China, vol. 1, pp. 194–199, 2008. [Google Scholar]
27. Y. Guo, H. D. Cheng and Y. Zhang, “A new neutrosophic approach to image denoising,” New Mathmatical and Natural Compution, vol. 5, no. 3, pp. 653–662, 2009. [Google Scholar]
28. J. Rajan, B. Jeurissen, M. Verhoye, J. Audekerke and J. Sijbers, “Maximum likelihood estimation-based denoising of magnetic resonance images using restricted local neighborhoods,” Physics in Medicine and Biology, vol. 56, no. 16, pp. 5221–5234, 2011. [Google Scholar]
29. L. D. Tran, S. M. Nguyen and M. Arai,“GAN based noise model for denoising real images,” in Proc. of the 15th Asian Conf. on Computer Vision (ACCV 2020), Tokyo, Japan, pp. 560–572, 2020. [Google Scholar]
30. R. Sipio, M. Giannelli, S. Haghighat and S. Palazzo, “DijetGAN: A generative-adversarial network approach for the simulation of QCD dijet events at the LHC,” Journal of High Energy Physics, vol. 2019, no. 8, pp. 113–130, 2019. [Google Scholar]
31. C. Tian, Y. Xu, and W. Zuo, “Image denoising using deep CNN with batch renormalization,” Neural Networks, vol. 121, no. 12, pp. 461–473, 2020. [Google Scholar]
32. S. Ioffe, “Batch renormalization: Towards reducing minibatch dependence in batch-normalized models,” in Proc. of the Int. Conf. on Neural Information Processing Systems (ICNIPS 2017), California USA, pp. 1942–1950, 2017. [Google Scholar]
33. Z. Chen, Z. Zeng, H. Shen, X. Zheng, P. Dai et al., “DN-GAN: Denoising generative adversarial networks for speckle noise reduction in optical coherence tomography images,” Biomedical Signal Processing and Control, vol. 55, no. 1, pp. 101632–101645, 2020. [Google Scholar]
34. X. Liu, L. Song, S. Liu and Y. Zhang, “A review of deep learning based medical image segmentation methods,” Sustainability, vol. 13, no. 3, pp. 117–130, 2021. [Google Scholar]
Cite This Article
 Copyright © 2023 The Author(s). Published by Tech Science Press.
Copyright © 2023 The Author(s). Published by Tech Science Press.This work is licensed under a Creative Commons Attribution 4.0 International License , which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.


 Submit a Paper
Submit a Paper Propose a Special lssue
Propose a Special lssue View Full Text
View Full Text Download PDF
Download PDF Downloads
Downloads
 Citation Tools
Citation Tools