 Open Access
Open Access
ARTICLE
Multiple Point MedSAM Prompting for Enhanced Medical Image Segmentation
1 Departamento de Lenguajes y Ciencias de la Computación, Universidad de Málaga, Bulevar Louis Pasteur, 35, Málaga, Spain
2 ITIS Software, Universidad de Málaga, C/Arquitecto Francisco Peñalosa 18, Málaga, Spain
3 Biomedic Research Institute of Málaga, IBIMA Plataforma BIONAND. C/Doctor Miguel Díaz Recio, 28, Málaga, Spain
* Corresponding Author: Karl Thurnhofer-Hemsi. Email:
Computers, Materials & Continua 2026, 87(2), 92 https://doi.org/10.32604/cmc.2026.077561
Received 11 December 2025; Accepted 26 January 2026; Issue published 12 March 2026
Abstract
Automatic and accurate medical image segmentation remains a fundamental task in computer-aided diagnosis and treatment planning. Recent advances in foundation models, such as the medical-focused Segment Anything Model (MedSAM), have demonstrated strong performance but face challenges in many medical applications due to anatomical complexity and a limited domain-specific prompt. This work introduces a methodology that enhances segmentation robustness and precision by automatically generating multiple informative point prompts, rather than relying on single inputs. The proposed approach randomly samples sets of spatially distributed point prompts based on image features, enabling MedSAM to better capture fine-grained anatomical structures and boundaries. During inference, probability maps are aggregated to reduce local misclassifications without additional model training. Extensive experiments on various computed tomography (CT) and magnetic resonance imaging (MRI) datasets demonstrate improvements in Dice Similarity Coefficient (DSC) and Normalized Surface Dice (NSD) metrics compared to baseline SAM and Scribble Prompt models. A semi-automatic point sampling version based on the ground truth segmentations yielded enhanced results, achieving up to 92.1% DSC and 86.6% NSD, with significant gains in delineating complex organs such as the pancreas, colon, kidney, and brain tumours. The main novelty of our method consists of effectively combining the results of multiple point prompts into the medical segmentation pipeline so that single-point prompt methods are outperformed. Overall, the proposed model offers a straightforward yet effective approach to improve medical image segmentation performance while maintaining computational efficiency.Keywords
Supplementary Material
Supplementary Material FileMedical image segmentation is now branching into two main areas. Some continue to research specific designs, like SwinHCAD [1] and the Med-ReLU activation function [2]. These results indicate that task-specific work is still required. However, the release of the Segment Anything Model (SAM) [3] and MedSAM [4] has shifted much of the focus to foundation models. MedSAM, which was trained on lots of biomedical data [4], aims to be a universal tool. It can segment different body parts using simple cues, possibly removing the need to retrain models. This paradigm represents a fundamental change from conventional task-specific deep learning approaches exemplified by architectures like nnU-Net [5].
Moving from focused models to general ones has resulted in a trade-off between ease of interaction and stability of results. Tools like Scribble Prompt have shown that drawing inputs can cut down marking time [6]. However, using simple point-based prompts is still a risk. Recent work suggests a potential issue with prompting in clinical use. Chang et al. [7] found that MedSAM performs well with bounding boxes, but its correctness drops significantly with point prompts. It does not do as well as specialized models like nnU-Net. Saha and van der Pol [8] observed that MedSAM undergoes significant changes based on the size and clarity of lesions. It often makes smooth borders that hide irregular issues. This shakiness comes from depending on a fixed process driven by unclear input. Li et al. [9] note that different user clicks introduce user-caused uncertainty, resulting in mixed results. While Test-Time Augmentation (TTA) has helped address data uncertainty in images [10], there is currently no standard way to handle the uncertainty arising from the placement of prompts. However, point prompts are especially useful for inference because they offer a cheap and precise way to steer the predicted mask without changing the model: a few clicks inject local information that helps correct under- or over-segmentation due to domain shift or ambiguous appearance. Unlike boxes, which only provide a rough region prior, points can be placed exactly on uncertain boundaries or tiny scattered lesions, yielding better refinements while remaining fast and accessible to non-experts. Compared to scribbles, which demand more time and suitable interfaces, point prompting keeps interactions simple, making it ideal for inference-time use, such as clinical review, where the aim is to adjust an existing mask with minimal effort rather than produce dense annotations.
To match the speed of points with the safety of boxes, we propose a new method for inference: multi-point MedSAM prompting through aggregation. Instead of using one click from the user, we see the prompt spot as random. By generating N random point prompts in a key area and grouping the predictions based on the ensemble of the probability maps, our method eliminates unusual results and improves the segmentation mask. This method leverages test-time augmentation [10] in the prompt area, ensuring robust performance even with minor changes in user input.
To the best of our knowledge, the combination of the results of multiple point prompts has not been explored in the existing literature. The MedSAM model has a multiple prompt mode, but it tries to enforce all point prompts at the same time. Therefore, if just one point is noisy, the whole process is likely to fail. Our method takes the opposite strategy because it is a consensus mechanism that filters out such errors. Our approach is not related to ensembles because the components of an ensemble are different models that receive the same inputs, in this case, the same point prompt. Furthermore, our proposal is not linked to Monte Carlo sampling either because Monte Carlo would require a previous tentative segmented region as input, while we only require several point prompts. Also, this proposal differs from our previous work on segmentation stability and robust interactive segmentation [11] because our previous work does not consider multiple point prompts as an input to the segmentation system.
The main contributions of this work are:
• A segmentation method based on the adaptation of test-time augmentation from multiple versions of the same test image to multiple point-based prompts. Prompt placement variation is used to enhance segmentation strength without retraining the model.
• A deep optimization of the hyperparameters using segmentation measures: the number of prompts and the aggregation function.
• A set of experiments with benchmark datasets indicating similar or better gains than direct or bounding-box based segmentation, specifically in the conflictive cases.
The rest of the paper is organized as follows: Section 2 summarizes the recent techniques. Then, our methodology and experiments are described in Section 3. The results and the discussion are depicted in Sections 4 and 5. Finally, conclusions and future work are presented in Section 6.
Generating effective prompts for foundation models is now a key area of research. Prompts in segmentation models usually have semantic prompts (text) or spatial prompts (points, boxes, etc.). Bounding boxes provide good spatial information and are not overly sensitive to user changes, but point prompts are easier to use, albeit very sensitive to the location of the click [12].
Researchers have explored automatic prompt generation strategies. Zhang et al. [13] created UR-SAM, which fixes SAM outputs that are not certain by using estimated uncertainty maps. This greatly improved things for many organs without requiring manual intervention. This suggests that understanding the uncertainty of a model can help guide the improvement of automatic prompts. Using learning to improve prompts has also shown promise. Some ways train models to make the best prompts for specific tasks, but these often need to be adjusted for each task [14,15]. Finding the right balance between automating tasks and giving users control remains a challenge.
Recent work on calibration and uncertainty in medical imaging has also highlighted the importance of ensemble voting. Aldhahi and Sull [16] developed Uncertain-CAM, which uses machine learning and confidence calibration to make predictions more reliable. This is particularly useful for methods that combine predictions, where you need to integrate multiple predictions effectively. While considerable progress has been made in quantifying data and model uncertainty [17], the exploration of prompt-related uncertainty is necessary in medical image segmentation. Iterative methods have been proposed to mitigate user-induced uncertainty. PRISM [12] enables users to refine segmentations over time with additional prompts.
Test-Time Augmentation (TTA) is a fast method for reducing data-related uncertainty. Wang et al. [10] demonstrated that TTA can address data issues without requiring additional training. By adjusting the input images slightly and combining the predictions, TTA improves the accuracy and reliability across various medical imaging tasks. Using ensembles, where multiple models are combined, is another effective way to estimate uncertainty. Deep Ensembles, where models are trained independently, provide better uncertainty estimates than single models [18]. However, they are expensive to train and operate. Knowledge distillation is a viable option, demonstrating that uncertainty estimates from deep ensembles can be incorporated into single models, achieving comparable performance while reducing costs [19].
Combining multiple predictions is a common practice in machine learning, using approaches such as majority voting for classification and probability averaging for dense predictions [20]. In medical image segmentation, aggregation methods must maintain spatial consistency, and recent work has explored more effective strategies like confidence-weighted averaging, for example, calibration-based ensembles that weigh predictions by model confidence to improve segmentation and uncertainty estimates [21].
3.1 Proposed Method: Multi-Point MedSAM Prompting and Aggregation
The proposed Multi-Point Aggregation method enhances the baseline MedSAM model by generating multiple prompts and aggregating the resulting predictions. Let us consider the baseline MedSAM model. Given a 2D input image
Our proposal consists of building a consensus of the segmentations yielded by the MedSAM for multiple relevant point prompts. Let R be a suitable region of interest within the input image:
Then, a set of N query points
Finally, the N probability maps are combined by a suitable aggregation function
In the particular case that
The overall workflow is illustrated in Fig. 1. The process branches into two strategies for defining the sampling region R: Semi-Automatic Ground Truth (GT) sampling or Automatic sampling based on a baseline MedSAM prediction. Then, inference is performed using multiple random prompts, and the outputs are fused via an aggregation function
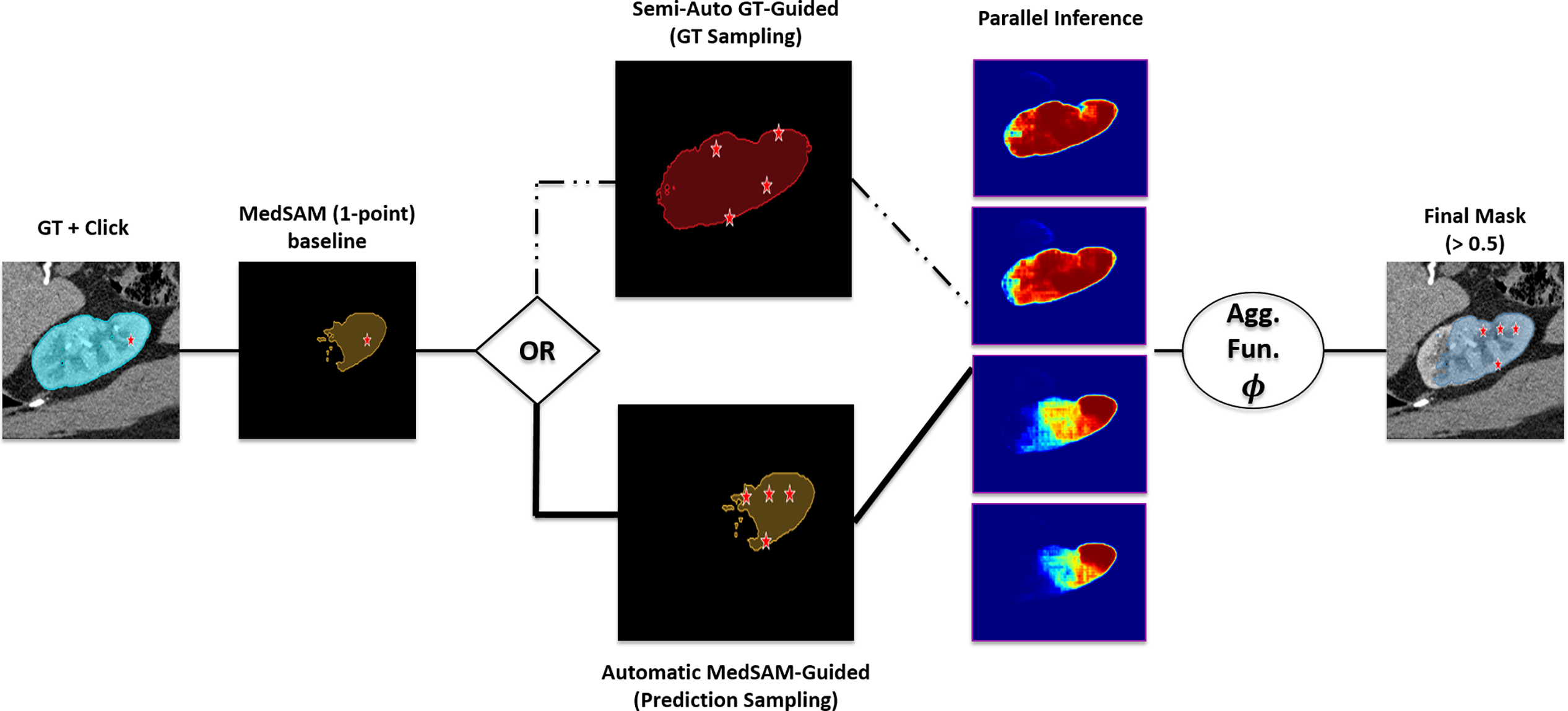
Figure 1: Schematic overview of the proposed Multi-Point MedSAM Prompting framework.
The specific procedure for a given 2D slice is as follows: The specific procedure for a given 2D slice is as follows: (1) Generate N random points from a defined region of interest (e.g., ground truth mask or prior segmentation); (2) Execute MedSAM N times, using each point as an individual prompt, yielding N probability maps; (3) Combine the probability maps via a pixel-wise mean or median operation to produce an aggregated map; and (4) Threshold the aggregated map at 0.5 to generate the final binary segmentation mask.
We compared the proposed test-time augmentation strategy with three existing segmentation models: SAM [3], Scribble Prompt [6], and MedSAM [4]. Comparison was centered on the point-prompt performance of each model to establish a benchmark of minimum user input performance. SAM is a general-purpose segmentation model that uses a lightweight decoder and a Vision Transformer encoder to produce real-time segmentation masks on various visual scenes. Scribble Prompt is a biomedical imaging interactive resource that modifies the SAM architecture by tuning the decoder with a training policy based on synthetic interactions and artificial labels. MedSAM is a baseline model trained on more than 1.57 million biomedical image pairs that transforms the generalist architecture for medical segmentation. Although the original version focuses on bounding boxes, we trained the point-prompt checkpoint on the FLARE2022 dataset [22] in order to make a direct comparison.
3.3 Datasets and Preprocessing
The number of selected public datasets to evaluate the proposed method is nine: FLARE2022_Abdominal [22], the MSD datasets (Spleen, Liver, Pancreas, Colon, and BrainTumor) [23], KiTS_Kidney [24], MsLesSeg_MSLesion [25], and LASC_Heart [26]. A detailed summary of these datasets is presented in the Supplementary Material. The dataset used to analyze the hyperparameters of the model, specifically the optimal choice of the number of queries, N, and the aggregation function, was the FLARE2022_Abdominal dataset. It is composed of 50 abdominal CT images with standardized annotations of 13 different organs. Nonetheless, during preprocessing, one label of organs was excluded due to its scattered nature, leaving 12 organs to be used for tuning the hyperparameters. The remaining datasets comprised the validation suite, which was selected to test the generalization of the model to various imaging modalities and anatomical complexities.
The preprocessing pipeline that was applied in preparation of the data was based on the MedSAM methodology to normalize the diverse nature of the selected datasets. The pipeline was used to process control NIfTI files of both CT and MRI modalities of the training sets. Preprocessing began with cleaning the ground truth label mask. All volumes were filtered by removing small, unconnected volumes with a 3D connected-components threshold (1000 voxels) and a 2D threshold (100 pixels) on each slide. Thereafter, there was intensity normalization based on modality. Normalization of CT scans was done by windowing (Window Level 40, Window Width 400). Using MRI, there was a normalization of the scan by cutting off the scans between 0.5 and 99.5 percentiles to remove outliers. Every volume was trimmed to the annotations’ Z-axis range and stored as .nii.gz files. For inference, the 2D slices were resized to
Experiments were conducted on a workstation with an Intel i7 CPU, 32 GB RAM, and an NVIDIA Titan X. The implementation used Python 3.10.13, PyTorch 2.2.0, CUDA 11.8, and standard libraries (NumPy, SciPy, SimpleITK, segment_anything). The Dice Similarity Coefficient (DSC) and Normalized Surface Dice (NSD) were used to determine the performance of the methods, as they are the gold standard in segmentation accuracy [27].
The FLARE2022 dataset was used to determine the optimal aggregation function (mean vs median) and the optimal number of query points (N, ranging from 2 to 25) to use. The initial data point was kept separate to assess the stability of the consensus strategy. The study evaluated success by considering both the accuracy of the image segmentations and the time required for the process. To ensure distinct test configurations, the analysis excluded anatomical components smaller than 26 pixels, avoiding sampling with repetition.
To quantify the variance introduced by the stochastic nature of the prompts, a repeatability study was performed. For every test slice,
Two types of experiments were then conducted: the first compared the improved multi-point combination method with a single-point MedSAM baseline in a semi-automatic setup. This experiment pretends to emulate a real-world scenario in which a doctor makes the choices of the point prompts within the GT region. The model does not use any GT information itself. Here,
4.1 Hyperparameter Setup: Optimal N and Aggregation Function
Results for the mean aggregation function are displayed in the first row of Fig. 2. The horizontal dashed line represents the fixed MedSAM baseline using the geometric center of the ROI (mean DSC 0.7116). The solid curve illustrates the performance of the proposed multi-point aggregation method. The solid curve begins at
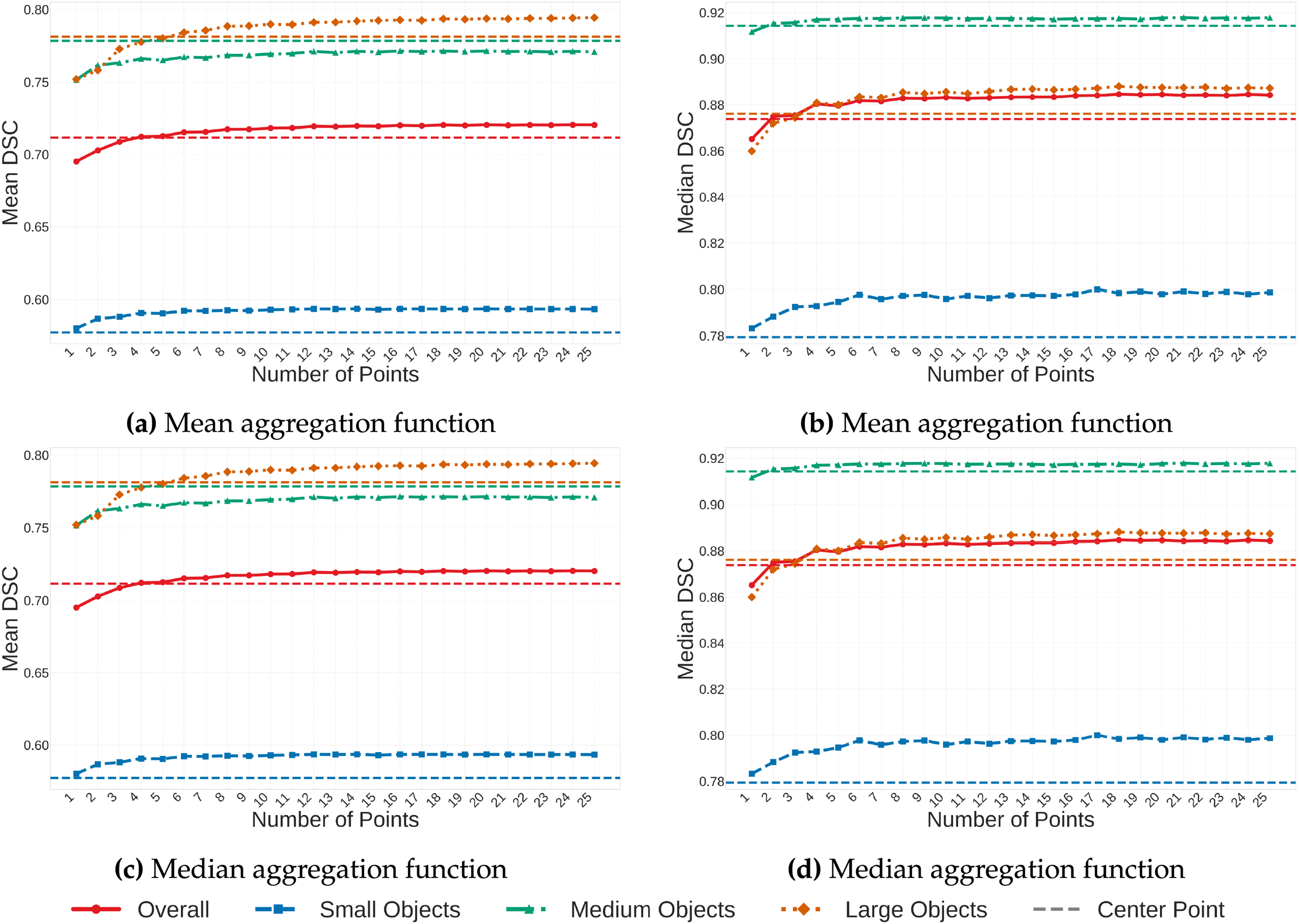
Figure 2: Parameter analysis on FLARE2022. Mean and median DSC values are displayed for the mean and median aggregation functions. Solid lines: multi-point method; dashed lines: MedSAM (1-point) baseline.
The second row of Fig. 2 presents results for the median aggregation function. The random point methodology achieves a mean DSC of 0.7181 at
Since performance gains were minimal after
4.2 Repeatability and Stability Analysis
Before comparing the proposed method against state-of-the-art models, we validated the intrinsic stability of the aggregation strategy. This analysis ensures that the stochastic nature of the prompt placement does not introduce significant variance into the final segmentation.
A statistical breakdown of the repeatability trials (
Fig. 3 plots the relationship between the Across-Slice STD (
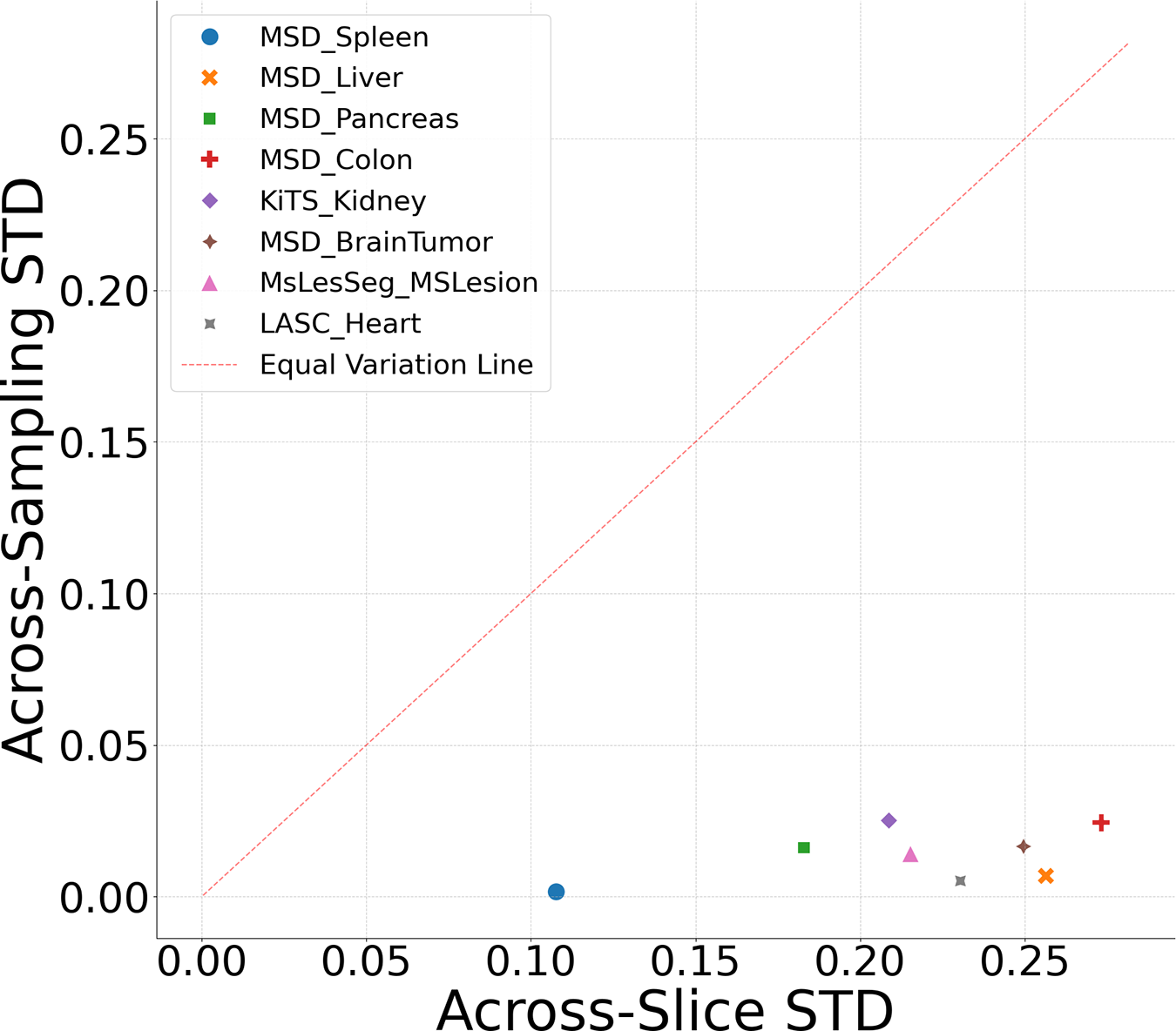
Figure 3: Repeatability analysis: dataset variability vs. instability.
4.3 Experiment 1: Semi-Automatic GT-Guided Prompt
Using
4.3.1 Quantitative Analysis: Volumetric Accuracy and Boundary Adherence
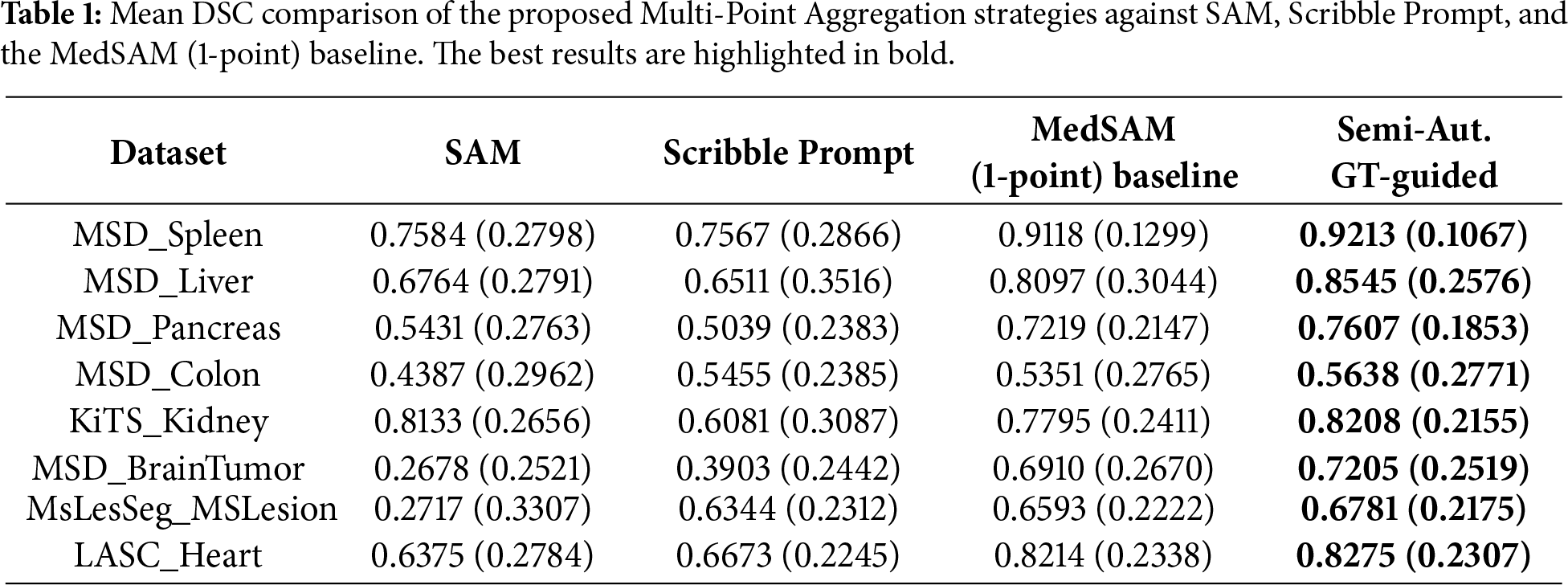
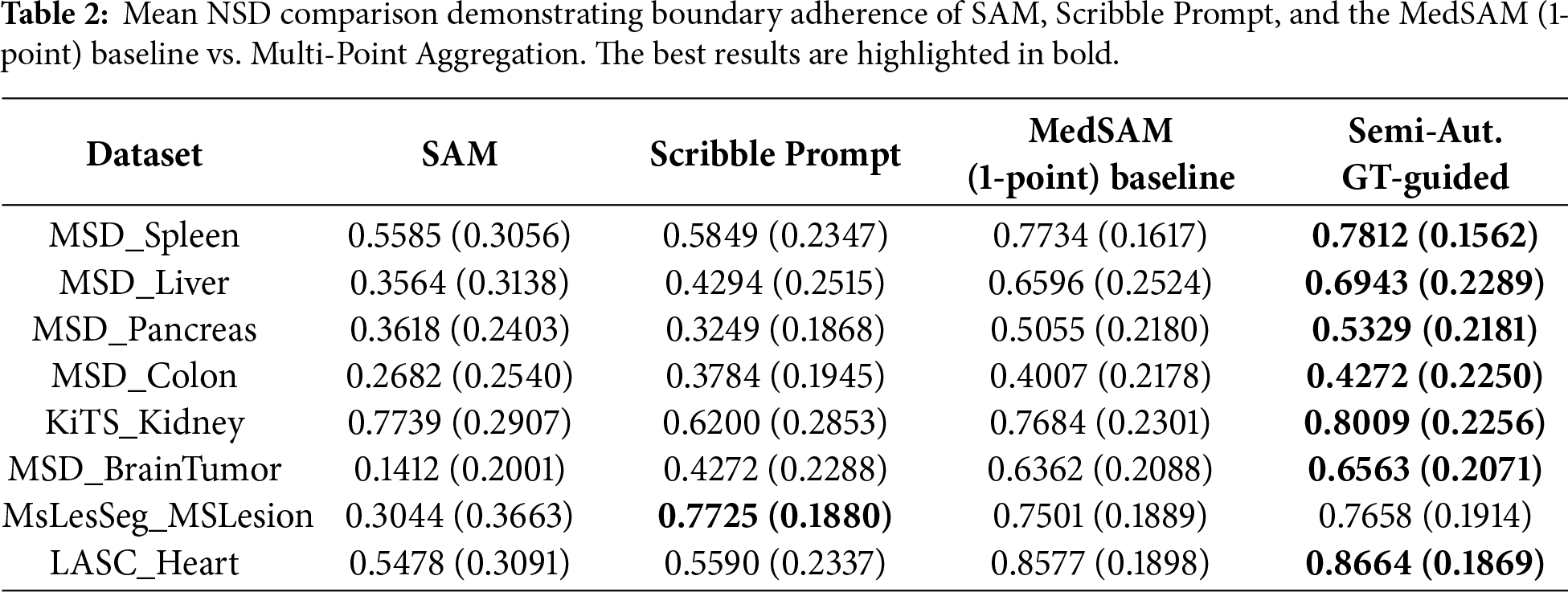
Table 1 presents the mean and standard deviation DSC scores. In MSD_Liver, the Semi-Automatic GT-guided method achieved a mean DSC of 0.8545 compared to 0.8097 for the MedSAM baseline. For MSD_Pancreas, the mean DSC was 0.7607 vs. 0.7219 for the baseline. In MSD_BrainTumor, the method yielded 0.7205 compared to 0.6910. Table 2 shows the Normalized Surface Dice metric. For KiTS_Kidney, the mean NSD was 0.8009 with the proposed method, compared to 0.7684 for the baseline. LASC_Heart showed a mean NSD of 0.8664 vs. 0.8577. In general, the proposed method outperforms between 0.01–0.04 of the original MedSAM metric values, which is a great improvement for medical purposes.
Inference time results are reported in the Supplementary material, Section S5. Although our proposal slightly increases the time it takes for a particular computation compared to the single-point MedSAM baseline (e.g., 0.4697 vs. 0.1337 s to complete the MSD_Spleen step), it is still much faster than the Scribble Prompt method, which takes 0.5614 s to perform the same operation. This indicates that the proposed strategy of aggregation is advantageous in terms of performance, as it provides better segmentation and incurs a minimal cost in inference latency.
4.3.2 Visual Qualitative Assessment
Fig. 4 displays segmentation masks for the compared methods. In the KiTS_Kidney example, the single-point baseline masks cover a portion of the region of interest. The Semi-Automatic GT-guided Prompt masks cover the full extent of the organ structures in these examples. In the MSD_Liver case, the baseline mask includes areas outside the organ boundary, whereas the proposed method’s mask is contained within the organ boundary.
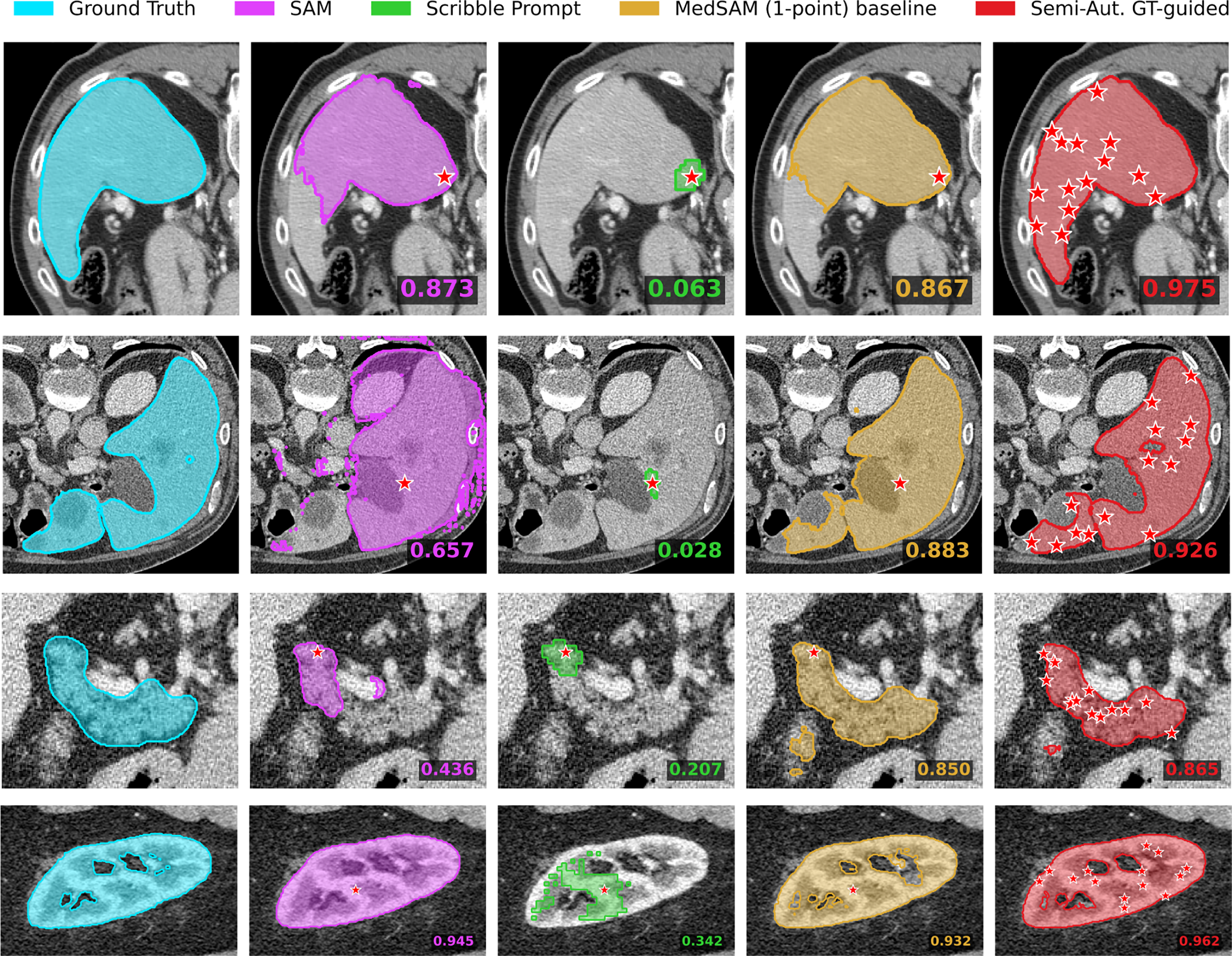
Figure 4: Qualitative results for Experiment 1. Rows correspond to (top to bottom): MSD_Spleen, MSD_Liver, MSD_Pancreas, and KiTS_Kidney.
In the MSD_Pancreas example, the MedSAM (1-point) baseline mask occupies the central body of the organ but does not extend to the lateral tail region. The Semi-Automatic GT-guided mask covers this lateral area, overlapping with the ground truth geometry. Similarly, in the MSD_Spleen case, the baseline prediction is strictly central and smaller than the ground truth area, whereas the proposed method yields a segmentation mask that extends to the outer boundaries of the structure.
4.4 Experiment 2: Automatic MedSAM-Guided Prompt
We evaluated the fully automatic pipeline using
The DSC distributions are presented in the Supplementary Material (Fig. S19). To evaluate the method’s ability to capture fine-grained boundaries, we analyzed the Normalized Surface Dice (NSD) in Fig. 5. The median NSD for the Automatic method is higher than the baseline in the MSD_Pancreas and KiTS_Kidney datasets. In MSD_BrainTumor, the 75th percentile values for the Automatic method are higher than those of the baseline, while in MSD_Colon, the method achieves a higher maximum NSD.
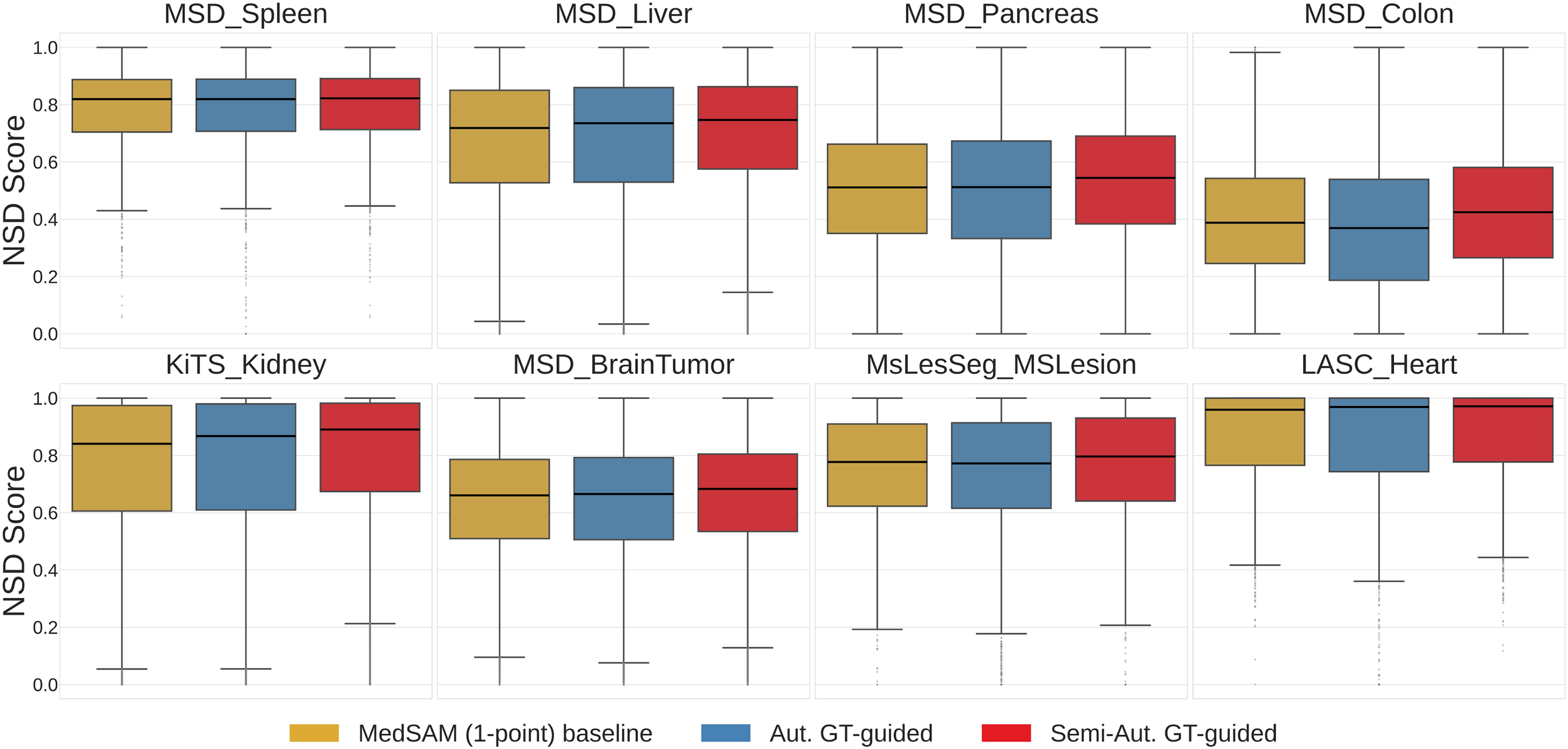
Figure 5: Normalized Surface Dice (NSD) distributions comparing the MedSAM (1-point) baseline, Automatic MedSAM-guided Prompt, and Semi-Automatic GT-guided Prompt across all datasets.
Conflictive Cases Analysis
“Conflictive cases” were defined as instances where the initial MedSAM (1-point) baseline yielded a DSC
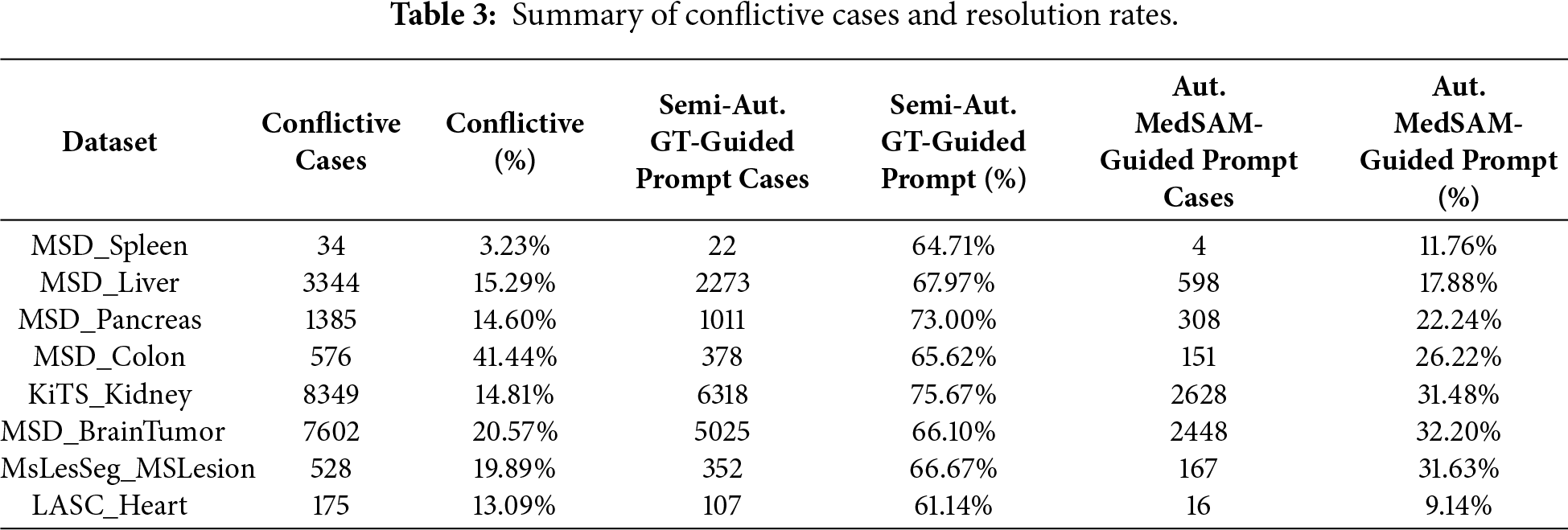
Fig. 6 displays conflictive cases from the MSD_Colon, MSD_BrainTumor, and LASC_Heart datasets. In the MSD_Colon example (top row), SAM and Scribble Prompt yielded DSC scores of 0.207 and 0.270, respectively. The MedSAM (1-point) baseline produced a mask with a DSC of 0.465, whereas the Automatic MedSAM-guided method achieved a DSC of 0.632. In the MSD_BrainTumor example (middle row), SAM showed a DSC of 0.183, and Scribble Prompt showed 0.511. The baseline mask resulted in a DSC of 0.468, while the Automatic method produced a mask with a DSC of 0.842. In the LASC_Heart example (bottom row), the DSC values for SAM and Scribble Prompt were 0.116 and 0.582, respectively. The baseline model resulted in a DSC of 0.276, indicating a conflictive case, while the Automatic MedSAM-guided method yielded a DSC of 0.747.
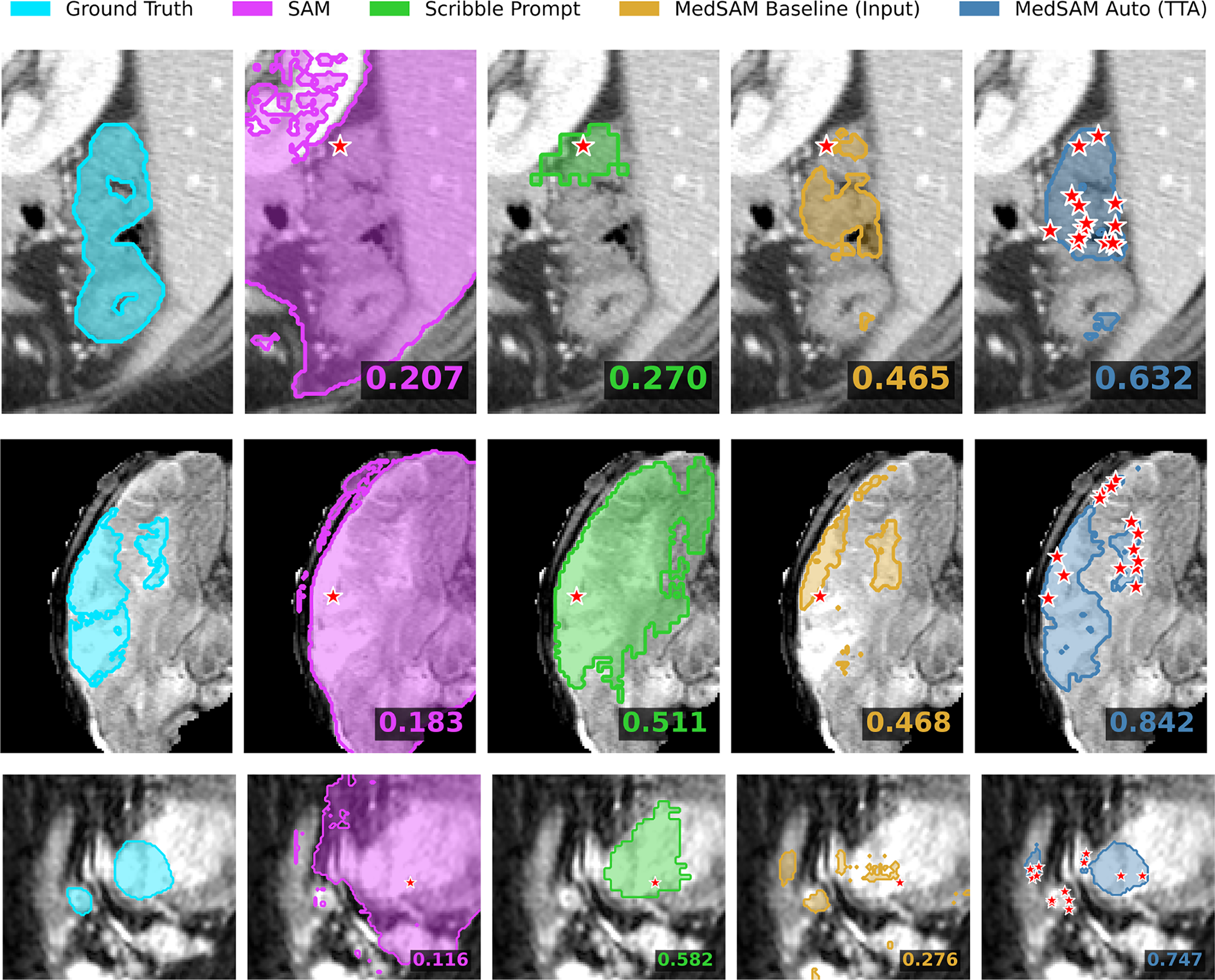
Figure 6: Visual analysis of conflictive cases (rows: MSD_Colon, MSD_brain, LASC_Heart). These examples illustrate instances where the single-point MedSAM baseline fails (
During the parameter setup stage, the decision to use mean aggregation with
The findings of the Semi-Automatic GT-guided experiment prove that multiple random prompts are beneficial in achieving segmentation consistency without loss of clinical usefulness. As demonstrated by Table 1 (mean and standard deviation DSC), the proposed aggregation method performed significantly better than the MedSAM (1-point) baseline, showing notable improvements in complex tasks, such as MSD_Liver (mean DSC improvement of
The proposed aggregation strategy has the potential to act as a self-correcting mechanism in a fully automated workflow, a fact emphasized by the Automatic MedSAM-guided experiment. As demonstrated in Fig. 5 and Fig. S19, using the preliminary model prediction to create another set of random prompts produced noticeable enhancements in median DSC and NSD on challenging datasets such as MSD_Pancreas and KiTS_Kidney. More importantly, the “Conflictive Cases” analysis shows that this iterative procedure is able to restore a large fraction of failures where the single-point baseline fails (DSC
To overcome the crucial problem of variability in interactive segmentation of medical images, this work presented a Multi-Point Aggregation framework for MedSAM. The proposed approach eliminates the unpredictability of inconsistent user prompts by replacing individual and deterministic prompts with a stochastic aggregate of N random points. We concluded that Mean Aggregation with
After parameter optimization on the multi-organ FLARE2022 dataset, all eight diverse CT and MRI benchmarks were experimentally validated, demonstrating that this approach consistently outperforms the standard MedSAM baseline, particularly in experiments involving complex anatomical structures. Boundary precision and overlap were significantly enhanced in semi-automatic tests, with a Mean DSC increase of about 4.5% on liver segmentation, while being much faster than other tools, such as Scribble Prompt. Besides, the fully automatic pipeline was shown to have a strong self-correction ability; it recovered more than 30% of the cases that the baseline initially failed on (DSC
Nevertheless, this enhanced strength comes at the cost of higher computational demand, because the inference procedure is repeated several times on each slice. Although the latency of the current system is acceptable for interactive applications, future efforts will be made to optimize the aggregation algorithm to further reduce the inference time. Also, as the existing system is based on 2D slices, 3D extensions of this stochastic prompting approach may utilize volumetric spatial context more effectively, potentially improving organ-specific 3D geometry performance.
Acknowledgement: The authors thankfully acknowledge the computer resources, technical expertise and assistance provided by the SCBI (Supercomputing and Bioinformatics) center of the University of Málaga, as well as acknowledge the support of NVIDIA Corporation with the donation of an RTX A6000 GPU with 48 GB. The authors also thankfully acknowledge the grant of the Universidad de Málaga and the Instituto de Investigación Biomédica de Málaga y Plataforma en Nanomedicina-IBIMA Plataforma BIONAND.
Funding Statement: This work is partially supported by the Autonomous Government of Andalusia (Spain) under project UMA20-FEDERJA-108, and also by the Ministry of Science and Innovation of Spain, grant number PID2022-136764OA-I00. It includes funds from the European Regional Development Fund (ERDF). It is also partially supported by the Fundación Unicaja (PUNI-003_2023), and the Instituto de Investigación Biomédica de Málaga y Plataforma en Nanomedicina-IBIMA Plataforma BIONAND (ATECH-25-02).
Author Contributions: Conceptualization, Karl Thurnhofer-Hemsi; methodology, Ezequiel López-Rubio; software, Wasfieh Nazzal; validation, Wasfieh Nazzal and Karl Thurnhofer-Hemsi; formal analysis, Karl Thurnhofer-Hemsi and Miguel A. Molina-Cabello; investigation, Wasfieh Nazzal; resources, Wasfieh Nazzal and Miguel A. Molina-Cabello; data curation, Wasfieh Nazzal; writing—original draft preparation, Wasfieh Nazzal; writing—review and editing, Karl Thurnhofer-Hemsi, Miguel A. Molina-Cabello and Ezequiel López-Rubio; visualization, Wasfieh Nazzal; supervision, Ezequiel López-Rubio and Karl Thurnhofer-Hemsi; project administration, Ezequiel López-Rubio; funding acquisition, Miguel A. Molina-Cabello. All authors reviewed and approved the final version of the manuscript.
Availability of Data and Materials: The data that support the findings of this study are openly available in public repositories. The MSD datasets (Spleen, Liver, Pancreas, Colon, and BrainTumor) are available at http://medicaldecathlon.com/. The KiTS dataset is available at https://kits-challenge.org/kits23/. The FLARE2022 dataset is available at https://flare.grand-challenge.org/. The MsLesSeg dataset is available at https://github.com/alessiarondinella/MSLesSeg-2024. The LASC Heart dataset is available at https://www.kaggle.com/datasets/adarshsng/heart-mri-image-dataset-left-atrial-segmentation.
Ethics Approval: Not applicable.
Conflicts of Interest: The authors declare no conflicts of interest.
Supplementary Materials: The supplementary material is available online at https://www.techscience.com/doi/10.32604/cmc.2026.077561/s1. Additional experimental results are provided as Supplementary Material, including experiments showing the evolution of the performance on each dataset varying the number of points, as well as other quantitative and qualitative results, including inference time.
References
1. Jin S, Fayaz M, Dang LM, Song HK, Moon H. SwinHCAD: a robust multi-modality segmentation model for brain tumors using transformer and channel-wise attention. Comput Mater Contin. 2025;86(1):1–23. doi:10.32604/cmc.2025.070667. [Google Scholar] [CrossRef]
2. Waqas N, Islam M, Yahya M, Habib S, Aloraini M, Khan S. Med-ReLU: a parameter-free hybrid activation function for deep artificial neural network used in medical image segmentation. Comput Mater Contin. 2025;84(2):3029–51. doi:10.32604/cmc.2025.064660. [Google Scholar] [CrossRef]
3. Kirillov A, Mintun E, Ravi N, Mao H, Rolland C, Gustafson L, et al. Segment anything. In: Proceedings of the 2023 IEEE/CVF International Conference on Computer Vision (ICCV); 2023 Oct 1–6; Paris, France. p. 4015–26. [Google Scholar]
4. Ma J, He Y, Li F, Han L, You C, Wang B. Segment anything in medical images. Nat Commun. 2024;15(1):654. doi:10.1038/s41467-024-44824-z. [Google Scholar] [PubMed] [CrossRef]
5. Isensee F, Jaeger PF, Kohl SAA, Petersen J, Maier-Hein KH. nnU-Net: a self-configuring method for deep learning-based biomedical image segmentation. Nat Methods. 2021;18(2):203–11. doi:10.1038/s41592-020-01008-z. [Google Scholar] [PubMed] [CrossRef]
6. Wong HE, Rakic M, Guttag J, Dalca AV. Scribbleprompt: fast and flexible interactive segmentation for any biomedical image. In: Proceedings of the 18th European Conference on Computer Vision (ECCV); 2024 Sep 29–Oct 4; Milan, Italy. Cham, Switzerland: Springer Nature Switzerland; 2024. p. 207–29. [Google Scholar]
7. Chang C, Law H, Poon C, Yen S, Lall K, Jamshidi A, et al. Segment anything model (SAM) and medical SAM (MedSAM) for lumbar spine MRI. Sensors. 2024;24(9):2826. doi:10.3390/s25123596. [Google Scholar] [PubMed] [CrossRef]
8. Saha A, van der Pol CB. Liver observation segmentation on contrast-enhanced MRI: SAM and MedSAM performance in patients with probable or definite hepatocellular carcinoma. Can Assoc Radiol J. 2024;75(4):771–9. doi:10.1177/08465371241250215. [Google Scholar] [PubMed] [CrossRef]
9. Li H, Liu H, Hu D, Wang J, Oguz I. Assessing test-time variability for interactive 3D medical image segmentation with diverse point prompts. In: Proceedings of the 2024 IEEE International Symposium on Biomedical Imaging (ISBI); 2024 May 27–30; Athens, Greece. p. 1–5. [Google Scholar]
10. Wang G, Li W, Aertsen M, Deprest J, Ourselin S, Vercauteren T. Aleatoric uncertainty estimation with test-time augmentation for medical image segmentation with convolutional neural networks. Neurocomputing. 2019;338(1):34–45. doi:10.1016/j.neucom.2019.01.103. [Google Scholar] [PubMed] [CrossRef]
11. Nazzal W, Thurnhofer-Hemsi K, López-Rubio E. Improving medical image segmentation using test-time augmentation with MedSAM. Mathematics. 2024;12(24):4003. [Google Scholar]
12. Li H, Liu H, Hu D, Wang J, Oguz I. PRISM: a promptable and robust interactive segmentation model with visual prompts. In: International Conference on Medical Image Computing and Computer-Assisted Intervention (MICCAI); 2024 Oct 6–10; Marrakesh, Morocco. Cham, Switzerland: Springer; 2024. p. 389–99. [Google Scholar]
13. Zhang Y, Hu S, Xue L, Ren S, Hu Z, Cheng Y, et al. Enhancing the reliability of auto-prompting SAM for medical image segmentation with uncertainty estimation and rectification. In: Proceedings of the IEEE/CVF International Conference on Computer Vision Workshops (ICCVW); 2025 Oct 19–23; Honolulu, HI, USA. p. 1282–91. [Google Scholar]
14. Li C, Sultan RI, Khanduri P, Qiang Y, Indrin C, Zhu D. AutoProSAM: automated prompting SAM for 3D multi-organ segmentation. In: Proceedings of the 2025 IEEE/CVF Winter Conference on Applications of Computer Vision (WACV); 2025 Feb 26–Mar 6; Tucson, AZ, USA; 2025. p. 3570–80. [Google Scholar]
15. Yin D, Zheng Q, Chen L, Hu Y, Wang Q. APG-SAM: automatic prompt generation for SAM-based breast lesion segmentation with boundary-aware optimization. Expert Syst Appl. 2025;276(5):127048. doi:10.1016/j.eswa.2025.127048. [Google Scholar] [CrossRef]
16. Aldhahi W, Sull S. Uncertain-CAM: uncertainty-based ensemble machine voting for improved COVID-19 CXR classification and explainability. Diagnostics. 2023;13(3):441. doi:10.3390/diagnostics13030441. [Google Scholar] [PubMed] [CrossRef]
17. Lê M, Unkelbach J, Ayache N, Delingette H. Sampling image segmentations for uncertainty quantification. Med Image Anal. 2016;34(11):42–51. doi:10.1016/j.media.2016.04.005. [Google Scholar] [PubMed] [CrossRef]
18. Li HB, Navarro F, Ezhov I, Bayat A, Das D, Kofler F, et al. QUBIQ: uncertainty quantification for biomedical image segmentation challenge. arXiv:2405.18435. 2024. [Google Scholar]
19. Noothout JM, Lessmann N, Van Eede MC, Van Harten LD, Sogancioglu E, Heslinga FG, et al. Knowledge distillation with ensembles of convolutional neural networks for medical image segmentation. J Med Imaging. 2022;9(5):052407. doi:10.1117/1.jmi.9.5.052407. [Google Scholar] [PubMed] [CrossRef]
20. Alotaibi A. Ensemble deep learning approaches in health care: a review. Comput Mater Contin. 2025;82(3):3741–71. doi:10.32604/cmc.2025.061998. [Google Scholar] [CrossRef]
21. Buddenkotte T, Sanchez LE, Crispin-Ortuzar M, Woitek R, McCague C, Brenton JD, et al. Calibrating ensembles for scalable uncertainty quantification in deep learning-based medical image segmentation. Comput Biol Med. 2023;163(1):107096. doi:10.1016/j.compbiomed.2023.107096. [Google Scholar] [PubMed] [CrossRef]
22. Ma J, Zhang Y, Gu S, Ge C, Ma S, Young A, et al. Unleashing the strengths of unlabelled data in deep learning-assisted pan-cancer abdominal organ quantification: the FLARE22 challenge. Lancet Digit Health. 2024;6(2):e113–24. doi:10.1016/s2589-7500(24)00154-7. [Google Scholar] [PubMed] [CrossRef]
23. Antonelli M, Reinke A, Bakas S, Farahani K, Kopp-Schneider A, Landman BA, et al. The medical segmentation decathlon. Nat Commun. 2022;13(1):4128. [Google Scholar] [PubMed]
24. Heller N, Isensee F, Maier-Hein KH, Hou X, Xie C, Li F, et al. The state of the art in kidney and kidney tumor segmentation in contrast-enhanced CT imaging: results of the KiTS19 challenge. Med Image Anal. 2021;67(1):101821. doi:10.1016/j.media.2020.101821. [Google Scholar] [PubMed] [CrossRef]
25. Guarnera F, Rondinella A, Crispino E, Russo G, Di Lorenzo C, Maimone D, et al. MSLesSeg: baseline and benchmarking of a new multiple sclerosis lesion segmentation dataset. Sci Data. 2025;12(1):920. doi:10.1038/s41597-025-05250-y. [Google Scholar] [PubMed] [CrossRef]
26. Simpson AL, Antonelli M, Bakas S, Bilello M, Farahani K, Van Ginneken B, et al. A large annotated medical image dataset for the development and evaluation of segmentation algorithms. arXiv:1902.09063. 2019. [Google Scholar]
27. Wang Z, Wang E, Zhu Y. Image segmentation evaluation: a survey of methods. Artif Intell Rev. 2020;53(8):5637–74. doi:10.1007/s10462-020-09830-9. [Google Scholar] [CrossRef]
Cite This Article
 Copyright © 2026 The Author(s). Published by Tech Science Press.
Copyright © 2026 The Author(s). Published by Tech Science Press.This work is licensed under a Creative Commons Attribution 4.0 International License , which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.


 Submit a Paper
Submit a Paper Propose a Special lssue
Propose a Special lssue View Full Text
View Full Text Download PDF
Download PDF Downloads
Downloads
 Citation Tools
Citation Tools