 Open Access
Open Access
ARTICLE
Optimizing YOLOv11 for Rice Disease Detection: Integrating RepViT Backbone, BiFPN, and CBAM Attention
Department of Computer Engineering, Honam University, Gwangsangu, Gwangju, Republic of Korea
* Corresponding Author: Sang-Hyun Lee. Email:
Computers, Materials & Continua 2026, 87(3), 69 https://doi.org/10.32604/cmc.2026.077207
Received 04 December 2025; Accepted 17 February 2026; Issue published 09 April 2026
Abstract
Accurate and timely detection of rice leaf diseases is critical for ensuring global food security and maximizing agricultural yields. However, existing deep learning methods often struggle to balance the high accuracy required for detecting multi-scale lesions in complex field environments with the computational efficiency necessary for edge device deployment. This paper proposes You Only Look Once for Lightweight Detection (YOLOv11-LD), a lightweight object detection model for multi-scale rice leaf disease detection in real paddy field environments. The model is built on YOLOv11n and integrates a Re-parameterized Vision Transformer (RepViT) backbone, a Bidirectional Feature Pyramid Network (BiFPN) based neck, and a Convolutional Block Attention Module (CBAM) to enhance multi-scale feature representation to enhance multi-scale feature representation while maintaining a lightweight architecture suitable for edge deployment. A dataset of 3234 images captured in actual rice paddies was constructed, containing three major rice leaf diseases: bacterial blight, rice blast, and brown spot. and was split into 2241 training images and 993 validation images. Ablation experiments show that the full YOLOv11-LD configuration achieves 95.2% mAP_0.5 with 7.8 Giga Floating-Point Operations (GFLOPs) and 3.5M parameters, outperforming the baseline YOLOv11n (91.4% mAP_0.5) under the same input resolution of 640 × 640. Additional comparisons with Faster Region-based Convolutional Neural Network (Faster R-CNN), Single Shot MultiBox Detector (SSD), YOLOv5n, YOLOv8n, and YOLOv11n further confirm that YOLOv11-LD provides the best overall trade-off between detection accuracy and computational efficiency. These results demonstrate that YOLOv11-LD offers superior operational efficiency suitable for resource-constrained smart rice disease monitoring systems.Keywords
Supplementary Material
Supplementary Material FileRice is a staple crop consumed by a significant portion of the global population, and its productivity and quality are directly linked to food security and the stability of the agricultural economy [1]. However, various diseases occurring throughout the cultivation process simultaneously affect multiple organs, such as leaves, stems, and panicles. In severe cases, these infestations lead to yield reduction and quality deterioration [2]. In particular, failure to accurately identify the timing and density of Disease occurrences can result in dual negative outcomes: environmental pollution and the emergence of resistant Diseases due to excessive pesticide application, or yield losses caused by insufficient control. Consequently, precise monitoring of the location and distribution of Disease at the field level, followed by the establishment of a precision control system based on timely intervention, has emerged as a critical task for the implementation of smart agriculture [3,4].
Traditional surveys of rice Disease have relied heavily on skilled surveyors patrolling fields to visually identify lesions and count individuals. This method is limited by significant variations in results depending on the surveyor’s experience and subjectivity, as well as the considerable time and labor required to repeatedly observe large paddy fields [5]. Furthermore, rice leaf lesions initially appear as minute spots that gradually spread to surrounding tissues, and Diseases range from small individuals to relatively large insects, varying widely in size and shape. These factors make visual inspection structurally disadvantaged for early detection and quantitative population estimation. Recently, automated detection technologies based on imaging sensors such as drones, fixed cameras, and autonomous robots have been introduced. However, complex backgrounds in real-world environments, overlapping leaves, varying shooting distances and angles, and illumination changes due to natural light remain significant factors that degrade recognition performance [6–8].
To overcome these limitations, research on the automatic detection and classification of Diseases using computer vision and deep learning techniques has recently been actively conducted.
Against this background, the primary objective of this study is to systematically verify the performance of a lightweight YOLO-LD (YOLOv11-LD) model applied to rice disease detection. The YOLO-LD model is designed as a lightweight object detector based on YOLOv11n, integrating a Re-parameterized Vision Transformer (RepViT) backbone, a Bidirectional Feature Pyramid Network (BiFPN)-based neck, and a Convolutional Block Attention Module (CBAM). This design aims to efficiently fuse multi-scale features while simultaneously reducing computational complexity and model size.
Current constraints in agricultural AI require a delicate balance between detection accuracy for minute lesions and computational efficiency for low-power devices. To address this, the main contributions of this study are summarized as follows:
Systematic Architecture Optimization: We propose YOLOv11-LD, a specialized architecture that systematically integrates a RepViT backbone and a RepBiFPN neck. This design is not a mere combination of modules but a mathematically optimized structure that maximizes gradient flow and feature re-parameterization, achieving a superior trade-off between inference speed and accuracy compared to generic lightweight models.
Enhanced Multi-Scale Feature Representation: To tackle the extreme scale variation of rice lesions (from tiny initial spots to large blights), we designed a fusion mechanism coupling RepBiFPN with CBAM. This effectively suppresses complex background noise in paddy fields—a major failure point for standard lightweight detectors—while preserving semantic information for small objects.
Empirical Validation of Efficiency: Through rigorous benchmarking against state-of-the-art lightweight models, including YOLOv9-t and YOLOv10-n, we demonstrate that YOLOv11-LD achieves the highest parameter efficiency (95.2% mAP at 52 FPS), verifying its operational suitability for resource-constrained agricultural environments.
Early studies on plant disease detection attempted to identify specific diseases by combining manual feature engineering (based on color, texture, and shape) with traditional classifiers (e.g., SVM, Decision Trees, Probabilistic Neural Networks) [9]. However, this approach fails to sufficiently account for complex backgrounds, diverse shooting conditions, and the multi-scale characteristics of lesions. Moreover, it suffers from the structural limitation that features must be redesigned whenever the target object or environment changes.
Accordingly, object detection models utilizing Convolutional Neural Networks (CNNs) are being actively adopted for rice disease recognition. Studies utilizing not only two-stage (Faster R-CNN) and one-stage (SSD, RetinaNet) detection frameworks but also YOLO-series models, which simultaneously predict location and class in a single stage, have been reported [10–13]. In particular, the YOLO series is attracting attention as a suitable model for agricultural applications based on drones, mobile devices, and edge devices, as it offers both high inference speeds and excellent detection accuracy [14].
Recent research has focused on optimizing YOLO architectures for agricultural scenarios [15–17]. For instance, T-YOLO-rice introduced an enhanced tiny network for paddy agronomy, while other works have integrated attention mechanisms into MobileNet-based YOLO architectures to improve pest and disease detection. Similarly, bidirectional feature attention pyramid networks have been explored with YOLOv5 to enhance feature fusion.
Nevertheless, many high-precision YOLO models possess a large number of parameters and high computational volume, measured in floating-point operations (FLOPs), making them burdensome for real-time operation in embedded environments. Conversely, extremely lightweight models often struggle to secure sufficient expressive power to reliably recognize minute lesions and small Diseases in complex environments [18,19]. In other words, while the rice Disease detection problem requires an object detection model that satisfies both the expressive power to handle multi-scale and morphological diversity and the lightweight characteristics operable in edge environments, research systematically verifying these requirements remains insufficient.
Despite these advancements, many high-precision YOLO models possess a large number of parameters and high computational volume (FLOPs), making them burdensome for real-time operation in embedded environments. Conversely, extremely lightweight models often struggle to secure sufficient expressive power to reliably recognize minute lesions in complex environments. To address these issues, the YOLOv11-LD proposed in this study adopts YOLOv11n as the baseline structure but enhances it by integrating the RepViT backbone, BiFPN neck, and CBAM attention module to create a specialized lightweight disease detection model.
3.1 Overall Architecture of YOLOv11-LD Model
Fig. 1 illustrates the overall architecture of the proposed YOLOv11-LD. The model consists of three main parts: Backbone, Neck, and Head.
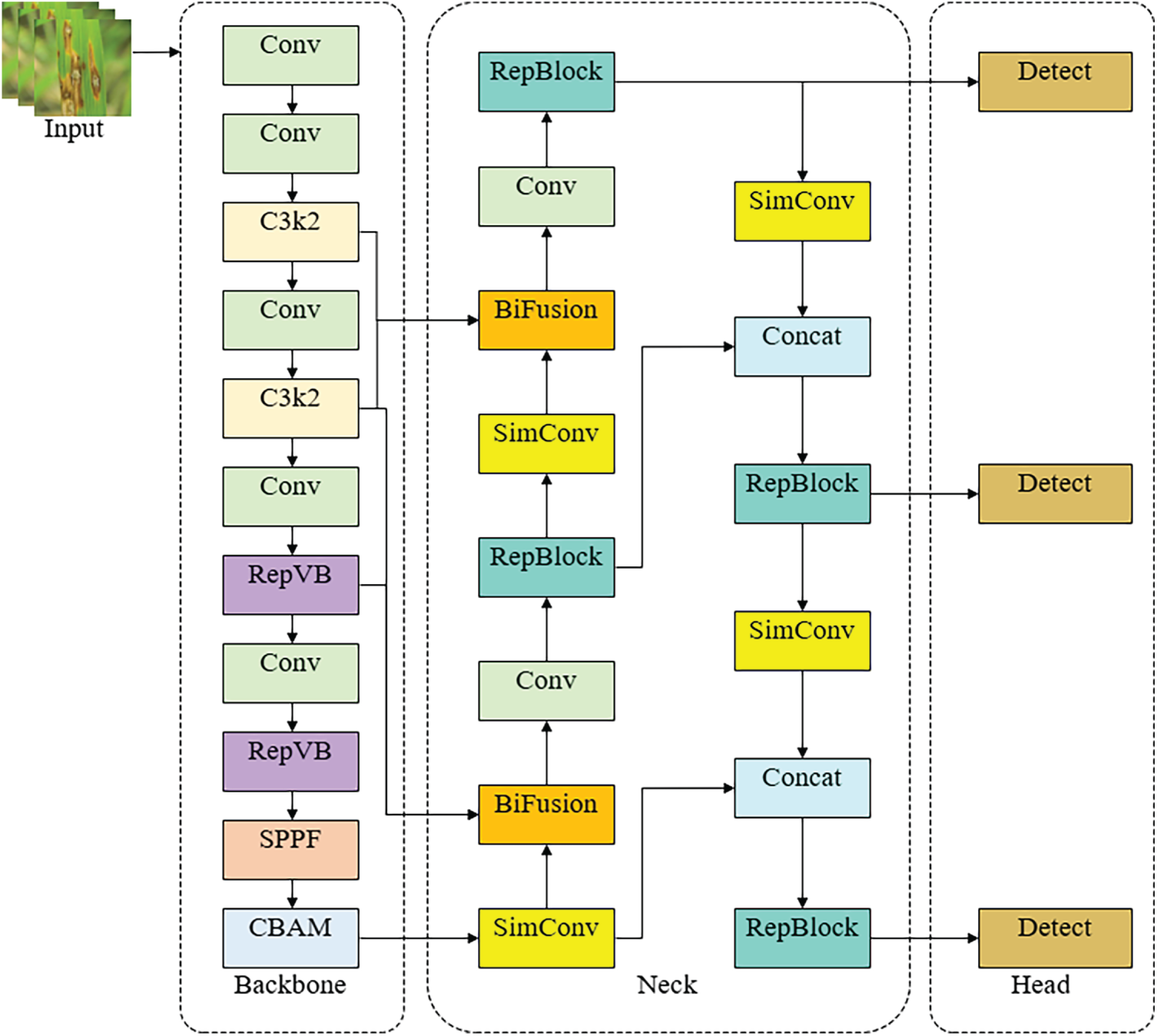
Figure 1: Structure of YOLOv11-LD model.
First, the Backbone utilizes a RepViT-based feature extraction network to extract features ranging from low-level to high-level from the input image. In the final stage, a Spatial Pyramid Pooling-Fast (SPPF) module is used to aggregate information from various receptive fields. Subsequently, a CBAM module is integrated to selectively weight important Disease regions across channel and spatial dimensions, thereby suppressing unnecessary features such as leaf backgrounds or noise while emphasizing candidate regions for lesions and Diseases.
In the Neck section, the multi-scale feature maps extracted from the backbone are fused using a BiFPN-based feature pyramid structure. Specifically, a bidirectional feature pyramid that considers both top-down and bottom-up paths allows for repetitive information exchange between feature maps of different resolutions. At each node, features from paths of varying depths are integrated using a fast normalized fusion method (weighted sum). During this process, Conv blocks, C3k2 blocks, and Upsample operations are appropriately arranged to ensure that small, medium, and large objects all retain sufficient spatial resolution and semantic information.
The Head is composed of a Path Aggregation Network (PANet)-based path aggregation structure and a Decoupled Detect Head. The multi-scale feature maps transmitted from the Neck are aggregated again along top-down and bottom-up paths. Finally, detection heads corresponding to three resolutions tailored for small, medium, and large objects predict the location and class of Disease, respectively. By consistently designing the Backbone–Neck–Head with a lightweight structure, YOLOv11-LD aims to maintain a model complexity comparable to the existing YOLOv11n while improving multi-scale detection performance in complex environments.
Implementation Details of Key Modules:
To ensure reproducibility, we explicitly define the specific modules denoted in our architecture:
SimConv (Standard Convolution Block): Represents a standard convolution unit consisting of a 2D Convolution layer, followed by Batch Normalization (BN) and a Sigmoid Linear Unit (SiLU) activation function.
C3k2 (CSP Bottleneck): Refers to a Cross Stage Partial (CSP) Bottleneck block optimized with a kernel size of
RepVB (RepViT Block): It corresponds to the re-parameterized building block from the RepViT architecture. During training, it utilizes a multi-branch topology (comprising 3 × 3 convolution, 1 × 1 convolution, and identity connection) which is fused into a single 3 × 3 convolution during inference.
BiFusion (Weighted Fusion Node): Refers to the weighted feature fusion operation within the BiFPN neck, utilizing the Fast Normalized Fusion method.
3.2 Re-Parameterized Vision Transformer Backbone
Fig. 2 shows the structure of the RepViT backbone used in the proposed model. RepViT is a lightweight feature extraction network designed to combine the global context extraction capability of Vision Transformers with the efficient computational structure of mobile-friendly CNNs [20]. The entire backbone consists of a Stem stage and Stages 1–4, where each stage comprises a Downsample block and multiple RepViTBlocks.
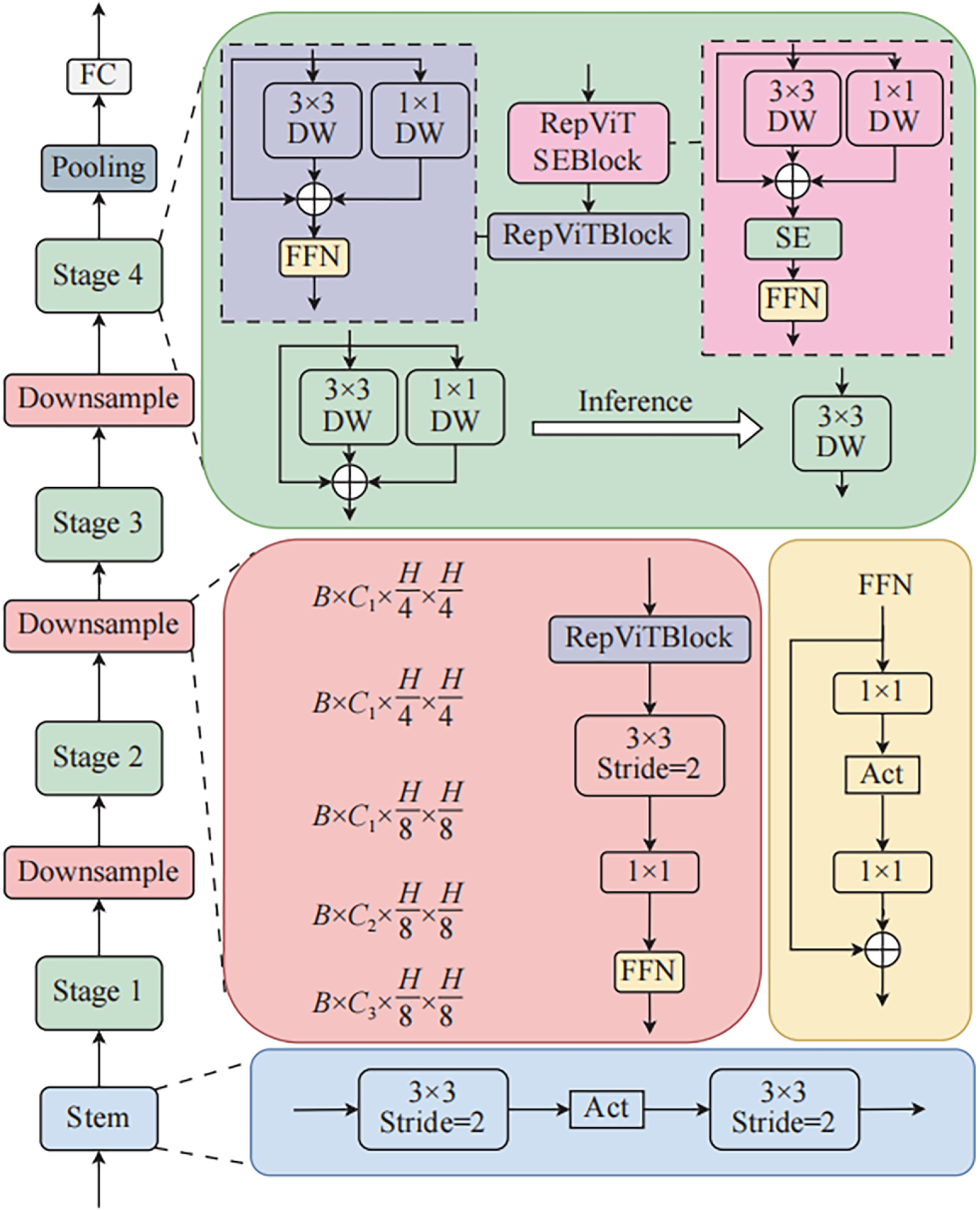
Figure 2: Structure of RepVit backbone.
Inside each RepViTBlock, a 3 × 3 depthwise convolution and a 1 × 1 pointwise convolution are combined to efficiently extract local spatial features, while a Feed-Forward Network (FFN) arranged in parallel or series supplements channel-wise expressiveness. During the training phase, multi-path convolution blocks with branch structures are used to increase expressiveness; during inference, these are converted into a single 3 × 3 convolution via structural re-parameterization, significantly reducing the number of calculations and parameters. Additionally, Squeeze-and-Excitation (SE) blocks are placed in some stages to model inter-channel dependencies and assign higher weights to important channels, thereby making Disease and disease candidate regions more distinct.
By utilizing this RepViT backbone instead of the standard CNN backbone of YOLOv11n, the proposed model can simultaneously reflect global and local features while maintaining lightweight characteristics, effectively representing multi-scale objects in high-resolution rice Disease images.
Fig. 3 compares the BiFPN (Bidirectional Feature Pyramid Network) structure [21] used in the neck of the proposed model with existing Feature Pyramid Network (FPN), Path Aggregation Network (PANet), and Neural Architecture Search Feature Pyramid Network (NAS-FPN) structures. Traditional FPN propagates high-level semantic information from low-resolution to high-resolution feature maps via a unidirectional (top-down) pyramid structure, but it has the limitation that fine spatial information from lower layers is not sufficiently fed back in a bottom-up manner. PANet added a bottom-up path on top of FPN to implement bidirectional information flow, but it is noted that efficient feature fusion is difficult because equal weights are assigned to all paths. NAS-FPN automatically designs more complex feature pyramid structures through neural architecture search, but its highly complex structure and high computational cost make it unsuitable for lightweight models.
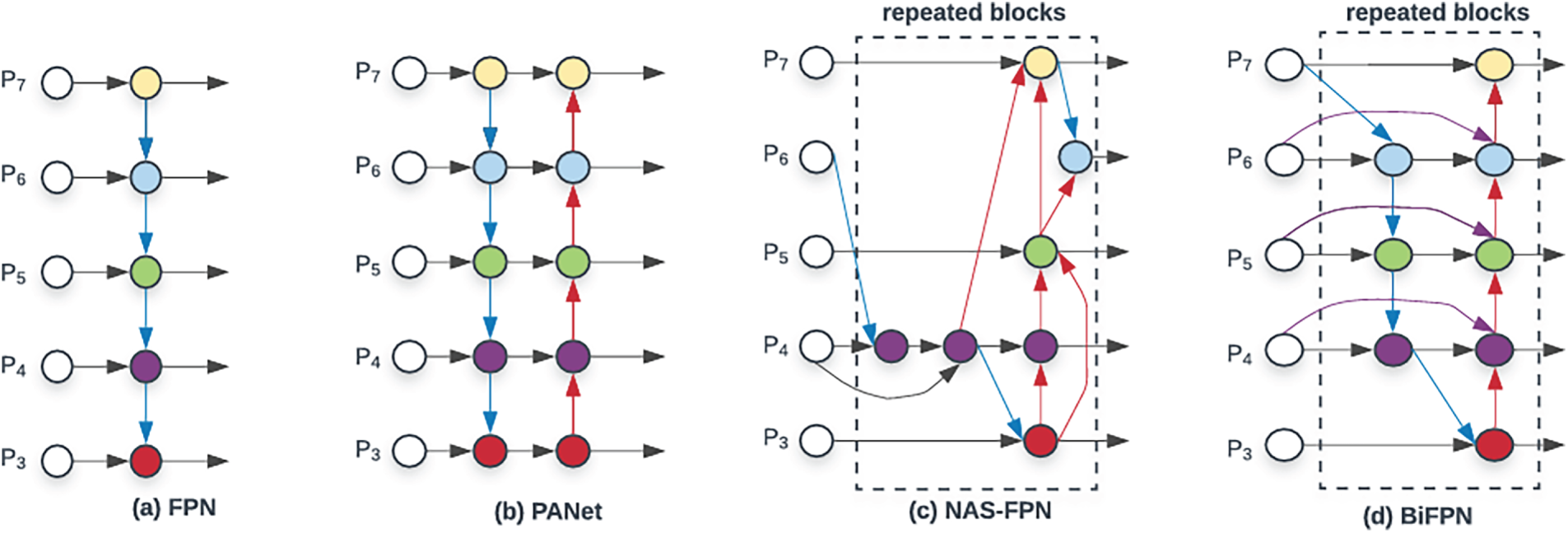
Figure 3: Structure of BiFPN.
BiFPN addresses these issues by introducing Weighted Feature Fusion on top of a simplified pyramid structure where top-down and bottom-up paths are repeatedly connected. Each node receives feature maps from two or more resolutions, assigns a learnable scalar weight
here,
In YOLOv11-LD, the BiFPN-based neck repeatedly fuses feature maps of different scales extracted from the backbone, helping objects of various sizes from small lesions and Diseases to medium or large ones maintain both sufficient semantic information and spatial resolution. This improves the detection performance of small Disease objects in complex backgrounds while minimizing the increase in computational volume compared to existing FPN or PANet, thus preserving the advantages of a lightweight model.
Table 1 presents the forward propagation process of the YOLOv11-LD model in pseudocode. It details the step-by-step procedure of generating multi-scale predictions
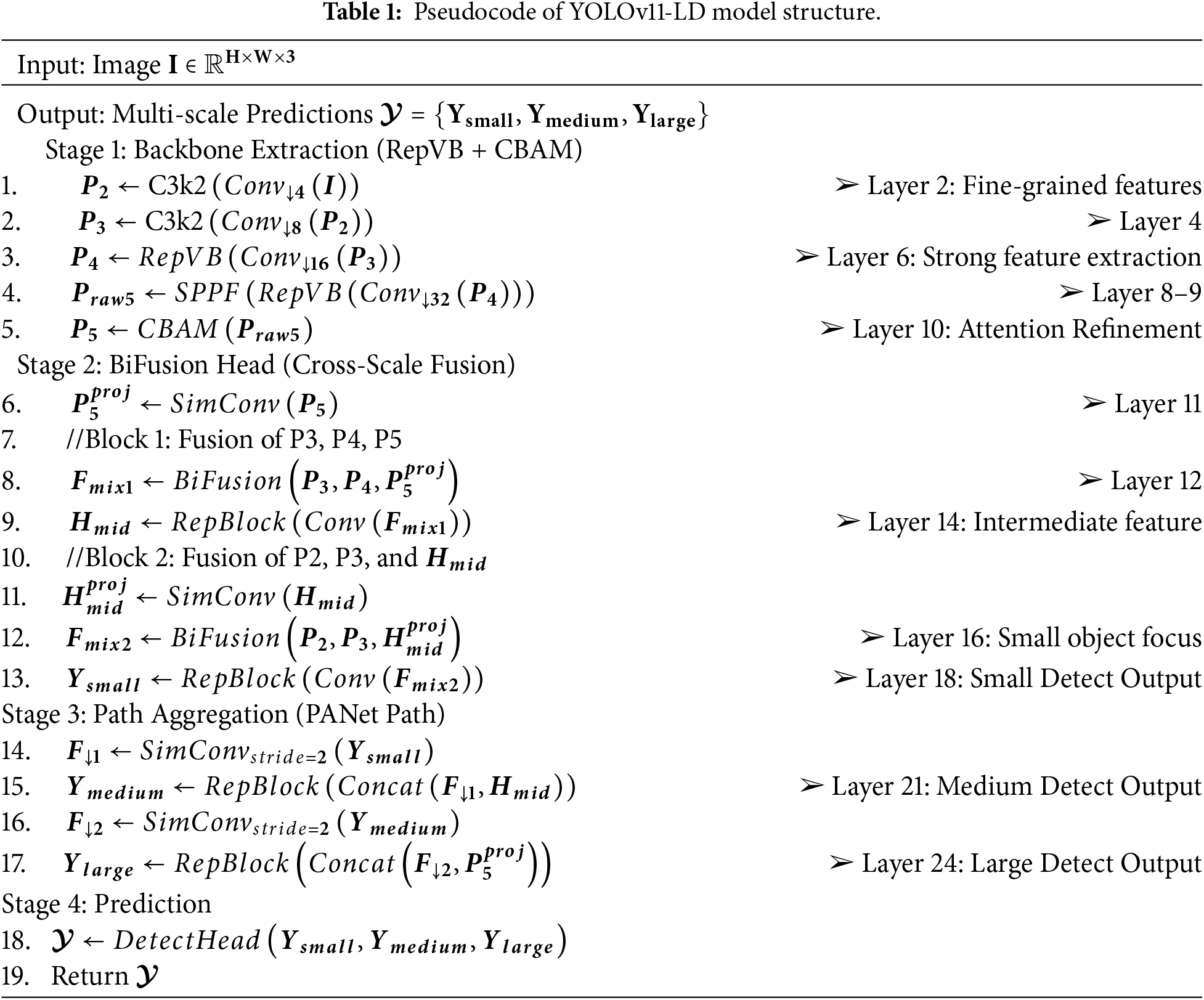

In Stage 1, the RepViT-based backbone (RepVB) and CBAM module are used to progressively extract rich features. The input image
Stage 2 fuses multi-scale features from the backbone using the BiFusion module.
Stage 3 reinforces representations for medium and large objects using a PANet-based structure.
Finally, Stage 4 inputs these three scale-specific features into the Decoupled Detect Head to predict candidate boxes, calculating position, size, objectness, and class probability to return the final prediction set
The training objective of the proposed YOLOv11-LD model is to minimize the total loss function (
here,
Box Regression Loss (
Classification Loss (
Distribution Focal Loss (
Fig. 4 shows the Train Class Loss curves for the baseline YOLOv11n and the proposed YOLOv11n-LD. YOLOv11n-LD consistently maintains lower loss values throughout the 300 epochs, demonstrating faster convergence and better fitting ability.
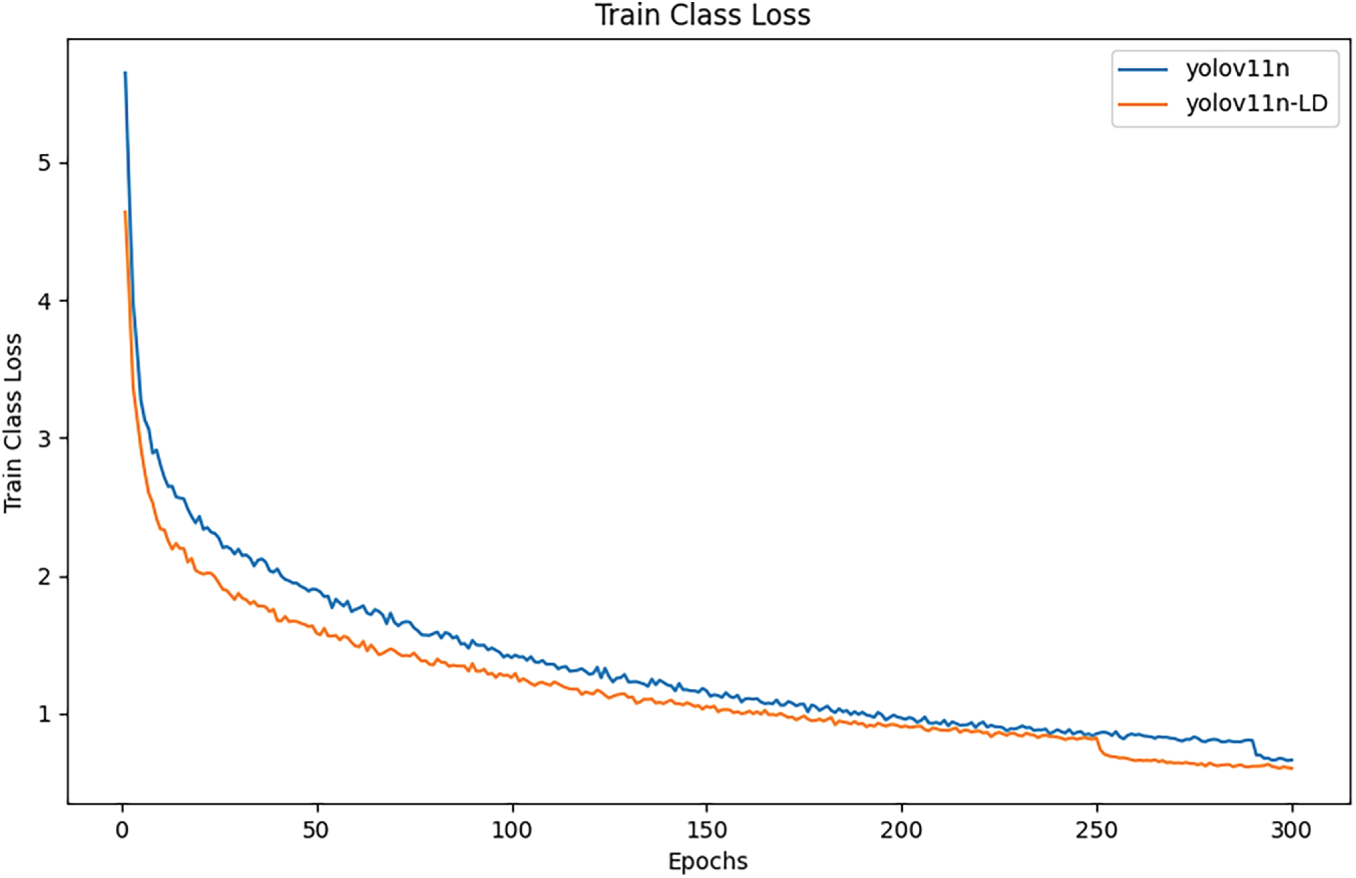
Figure 4: Train class loss compared.
In this study, to quantitatively evaluate the detection performance and computational efficiency of the proposed YOLOv11-LD model, Precision (
Precision refers to the proportion of candidates predicted by the model as diseases or Diseases that are actually positive, indicating how effectively the model suppresses false alarms. On the other hand, Recall refers to the proportion of actual positive samples correctly detected by the model, reflecting the degree of missed detections. mAP is a metric that synthesizes the average precision for all classes and is used to compare and evaluate the model’s overall detection accuracy at once when detecting various types of diseases simultaneously. Additionally, the complexity of each model was analyzed by reporting the number of parameters and the number of floating-point operations (FLOPs) together to analyze the balance between accuracy and computational cost.
Precision and Recall are calculated using Eqs. (3) and (4) based on the number of True Positives (TP), False Positives (FP), and False Negatives (FN).
here, TP is the number of diseases or Diseases correctly detected by the model, FP is the number of background or other objects incorrectly detected as Diseases, and FN is the number of Diseases that actually exist but were not detected by the model. High precision means fewer FPs, while high recall means fewer FNs. Since these two metrics generally have a trade-off relationship, it is important to analyze the P-R (Precision-Recall) curve together.
The Average Precision (AP) for a single class is defined by integrating the area under the P-R curve for that class, as shown in Eq. (5).
here,
The mean Average Precision (mAP) for all classes is defined as the average of the APs of each class, representing the model’s overall detection performance for all rice Disease as shown in Eq. (6).
here,
This study established a benchmark for rice disease and Disease detection using real-world imagery. We optimized the YOLO-LD model for agricultural contexts by refining data augmentation and scale configurations, verifying its robustness in complex, multi-scale environments. Through a holistic comparison with existing state-of-the-art models considering accuracy, computational load, and speed we demonstrated the quantitative viability of deploying lightweight models like YOLO-LD on edge devices for efficient agricultural monitoring.
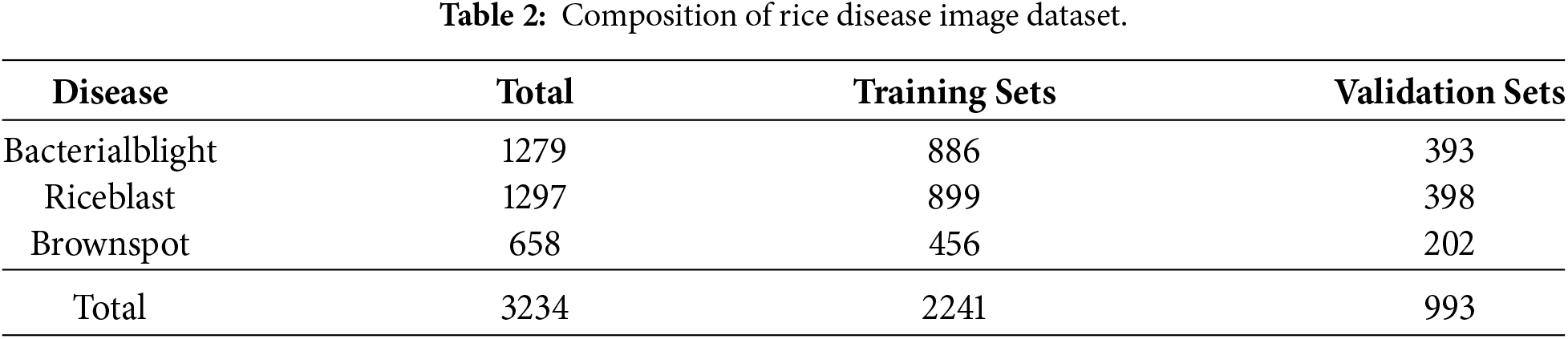
The rice disease image dataset utilized in this study consists of images captured in real-world environments, comprising a total of 3234 images. The dataset targets three representative diseases frequently reported in rice cultivation: Bacterial blight, Rice blast, and Brown spot. The class distribution includes 1279 images of Bacterial blight, 1297 of Rice blast, and 658 of Brown spot, as summarized in Table 2.
Images were collected using a handheld camera equipped with optical zoom to reflect actual field conditions, capturing subjects from various distances and angles. To incorporate diverse visual factors occurring in real-world environments such as complex backgrounds, occlusion, and lighting variations the frames were composed to include rice leaves, stems, and ears. Although the original images varied in resolution and aspect ratio, all images were resized to 640 × 640 pixels to ensure consistency during network training.
For annotation, experienced evaluators manually designated lesion areas for each image, assigning bounding boxes and class labels in the YOLO format. Since Bacterial blight, Rice blast, and Brown spot all form lesions of varying sizes and shapes on leaf surfaces, this dataset encompasses multi-scale rice diseases ranging from small spots to widely spread lesions, making it suitable for evaluating the robustness of detection models.
To facilitate model training and validation, the entire dataset was divided into training and validation sets. Stratified sampling was applied to maintain the distribution of disease classes. Approximately 69.3% of the total data, amounting to 2241 images, was allocated to the training set, while the remaining 993 images were used for the validation set. Specifically, the training and validation splits were configured as follows: 886 and 393 images for Bacterial blight, 899 and 398 for Rice blast, and 456 and 202 for Brown spot, respectively, as detailed in Table 2.
Note that in this study, the validation set was utilized to evaluate the model’s generalization performance and for the final comparative analysis. All ‘inference tests’ mentioned in subsequent sections refer to evaluations performed on this held-out validation dataset.
The dataset was specifically constructed to evaluate the model’s ability to distinguish between three major rice leaf diseases (Bacterial blight, Rice blast, and Brown spot). Note that this study focuses on the classification accuracy among these specific disease categories; therefore, healthy leaves or other background objects were excluded from the target classes during annotation. Additionally, a subset of sample images and annotation specifications is provided in the Supplementary Materials.
The experiments in this study were conducted on a standalone GPU workstation running a Windows operating system. The hardware configuration consisted of an Intel Core i9-10900K CPU and an NVIDIA Quadro RTX 6000 GPU equipped with 24 GB of GDDR6 VRAM. The software environment utilized the PyTorch 2.7.0 deep learning framework with CUDA 12.1.0, and the algorithms were implemented using Python 3.10.
All models were trained under identical experimental conditions to ensure a fair comparison. The input resolution was fixed at 640 × 640 pixels, the batch size was set to 16, and the training duration was set to 300 epochs. Detailed hyperparameters, including learning rate and optimizer settings, were primarily based on the default configuration of YOLOv11n, with fine-tuning applied to accommodate the specific characteristics of the rice disease dataset.
In this section, we conducted an ablation study to analyze the impact of the three key modules constituting YOLOv11-LD on rice disease detection performance. Using YOLOv11n as the baseline, we evaluated performance by stepwise integration of the RepViT backbone, BiFPN neck, and CBAM. The comparative models include the original YOLOv11n, YOLOv11n + RepViT, YOLOv11n + BiFPN, YOLOv11n + CBAM, and the final YOLOv11-LD integrating all three modules. Performance metrics, including Precision, mAP_0.5, computational cost (GFLOPs), and the number of parameters, are summarized in Table 3.
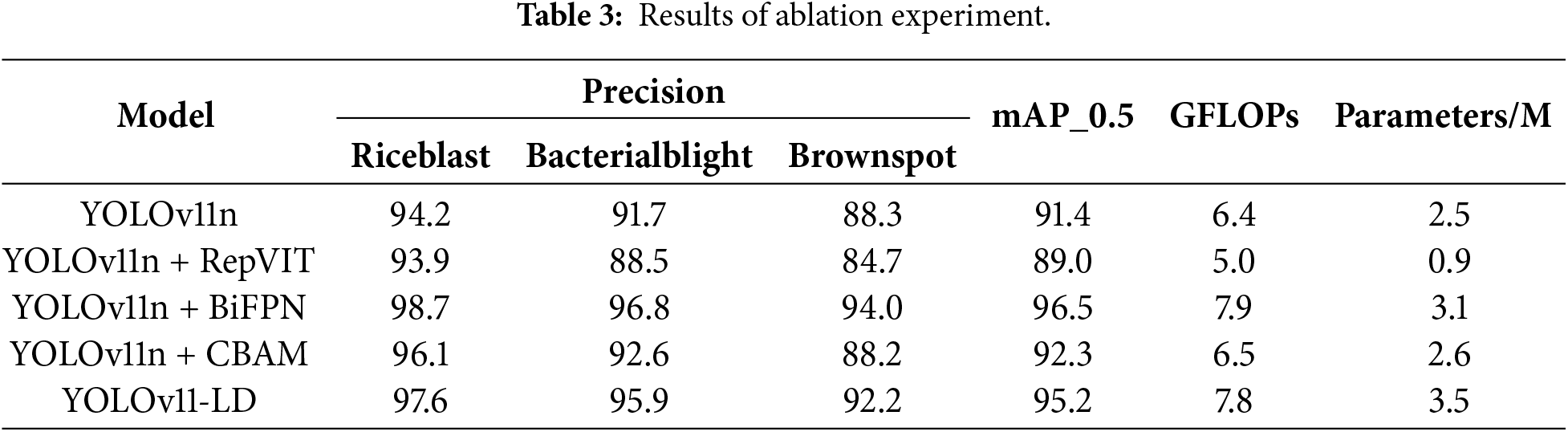
First, as shown in Fig. 5, the YOLOv11n + RepViT model, which incorporates the RepViT backbone, demonstrated significantly improved computational efficiency compared to the baseline. GFLOPs decreased by approximately 18.8% (from 6.4 to 5.0G), and the number of parameters was reduced by roughly 62.0% (from 2.5 to 0.9M). However, mAP_0.5 declined by 2.4%p (from 91.4% to 89.0%), suggesting that while RepViT offers advantages in lightweighting, it faces limitations in securing sufficient representational power for high-resolution rice disease images when applied in isolation.

Figure 5: YOLOv11n + RepViT.
Next, as shown in Fig. 6, the YOLOv11n + BiFPN model, applied to the neck architecture, enhanced the fusion of multi-scale information across small, medium, and large lesions. This resulted in a substantial increase in mAP_0.5 to 96.5%, a 5.1%p improvement over the baseline. However, this accuracy gain came with a trade-off in computational complexity, as GFLOPs increased to 7.9G and the number of parameters rose to 3.1M.

Figure 6: YOLOv11n + BiFPN.
As shown in Fig. 7, The YOLOv11n + CBAM model, which applies channel and spatial attention to highlight disease regions, raised mAP_0.5 to 92.3%, a 0.9%p improvement over the baseline. In this configuration, the increase in computational cost was relatively marginal compared to the accuracy gain, with GFLOPs at 6.5G and parameters at 2.6M.

Figure 7: YOLOv11n + CBAM.
Finally, as shown in Fig. 8, the YOLOv11-LD model, integrating the RepViT backbone, BiFPN neck, and CBAM, recorded per-class precision scores of 97.6% (Rice blast), 95.9% (Bacterial blight), and 92.2% (Brown spot). The overall mAP_0.5 reached 95.2%, a 3.8%p improvement over the YOLOv11n baseline. With 7.8 GFLOPs and 3.5M parameters, the proposed model demonstrated a balanced performance, maintaining high accuracy while reducing computational costs compared to the standalone BiFPN model. These results validate that the proposed architectural combination effectively addresses the multi-scale and complex background characteristics of rice diseases while successfully maintaining lightweight properties.

Figure 8: YOLOv11-LD.
Fig. 9 illustrates the visualization results of the ablation study. As observed from the figure, YOLO-LD demonstrates the best performance. Notably, after incorporating the CBAM attention mechanism, there is a significant improvement in the detection of dense small objects.
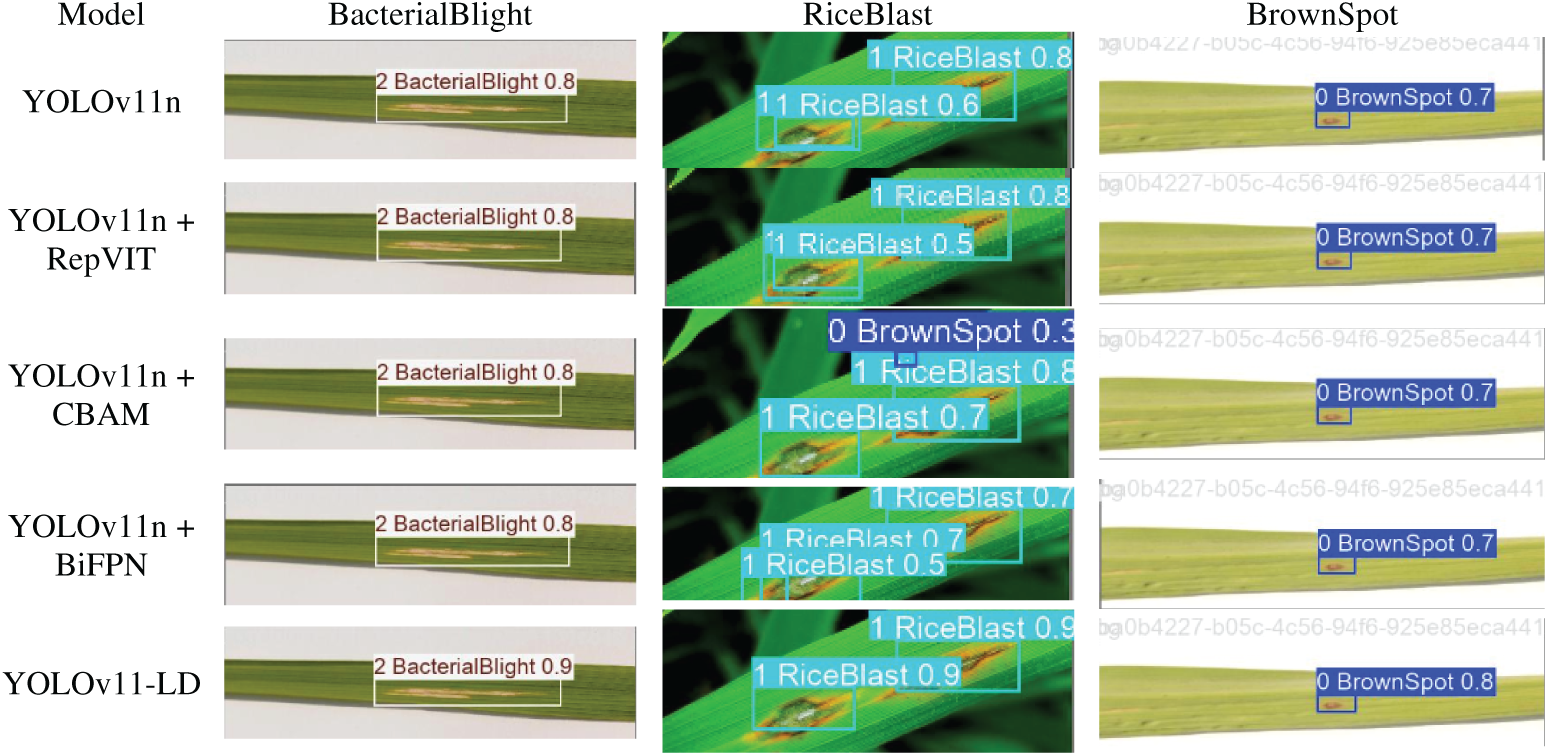
Figure 9: Results of ablation experiment.
4.4 Comparison with Existing Detection Models
In this section, we evaluated the relative performance and efficiency of the proposed YOLOv11-LD model in the context of rice disease detection by comparing it with existing representative object detectors. The comparison models included the two-stage detector Faster Region-based Convolutional Neural Network (Faster R-CNN), the one-stage detector SSD, and widely used lightweight YOLO variants: YOLOv5n, YOLOv8n, and YOLOv11n. All models were trained and evaluated under identical dataset and experimental conditions. Performance metrics, including mAP_0.5, Recall, GFLOPs, model size (Weights in MB), and Frames Per Second (FPS), are summarized in Table 4.
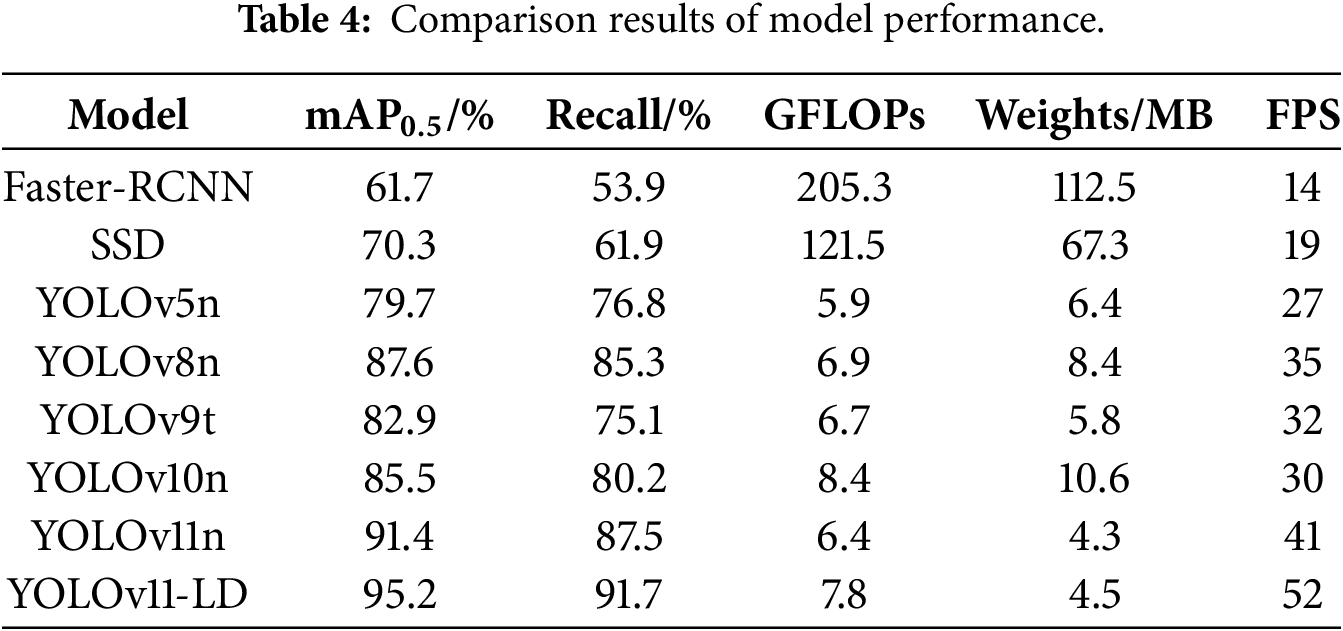
Faster-RCNN, a traditional two-stage architecture, demonstrated the lowest accuracy with an mAP_0.5 of 61.7% and a Recall of 53.9%. Furthermore, its high computational cost (205.3 GFLOPs, 112.5 MB) and slow inference speed (14 FPS) rendered it unsuitable for real-time rice disease detection. SSD similarly showed limited accuracy (mAP_0.5 70.3%, Recall 61.9%) and possessed a model size (67.3 MB) and computational load (121.5 GFLOPs) that imposed significant burdens for deployment on edge devices.
Among the YOLO series, YOLOv5n offered a relatively lightweight structure (5.9 GFLOPs, 6.4 MB, 27 FPS) with an mAP_0.5 of 79.7% and Recall of 69.5%; however, its accuracy was somewhat insufficient to serve as a standard for effective rice disease detection. YOLOv8n improved accuracy to 87.6% (mAP@0.5) and 78.1% (Recall) but still required a trade-off between performance and efficiency.
To ensure a comprehensive evaluation against state-of-the-art object detectors, we further benchmarked the model against the recently released YOLOv9-t and YOLOv10-n. YOLOv9-t, utilizing the Generalized Efficient Layer Aggregation Network (GELAN) architecture, achieved an mAP@0.5 of 82.9% and a Recall of 75.1%. While it demonstrated lower computational cost (6.7 GFLOPs) compared to YOLOv10-n, its inference speed (32 FPS) and detection accuracy were suboptimal for this specific task compared to the proposed method. Similarly, YOLOv10-n recorded an mAP@0.5 of 85.5% and a Recall of 80.2%. However, it incurred a higher computational load (8.4 GFLOPs) and a larger model size (10.6 MB), with a relatively lower inference speed of 30 FPS. In contrast, the proposed YOLOv11-LD significantly outperformed both competitors, achieving 95.2% mAP and 52 FPS, proving that the specialized integration of RepViT and BiFPN provides a distinct advantage for multi-scale rice disease detection.
The baseline model, YOLOv11n, exhibited superior comprehensive performance compared to YOLOv5n and YOLOv8n, recording an mAP_0.5 of 91.4%, Recall of 84.5%, 6.4 GFLOPs, 3.9 MB, and 41 FPS. However, the proposed YOLOv11-LD achieved the highest detection performance with an mAP_0.5 of 94.5% and Recall of 91.7%. Despite these gains, the increase in computational load and parameter count compared to YOLOv11n was marginal (7.8 GFLOPs, 4.5 MB). Notably, the FPS reached 52, representing an approximate 26.8% improvement over the baseline YOLOv11n (41 FPS), demonstrating the best results for real-time processing.
In summary, when compared to existing two-stage and one-stage object detection models, YOLOv11-LD offers the most outstanding balance in terms of accuracy, efficiency, and real-time capability, confirming it as the most suitable candidate for edge device-based rice disease detection systems.
4.5 Qualitative Analysis of Detection Performance
To visually evaluate performance, we conducted inference tests using identical test images across all models, as illustrated in Fig. 10. The results demonstrate that YOLO-LD achieved significantly higher detection accuracy and confidence compared to the other methods. Conversely, the YOLOv5n model exhibited a critical limitation, with frequent instances of missed detections. Similar issues regarding missed detections were also observed in the YOLOv8n and YOLOv11n models, highlighting the superior robustness of the proposed YOLO-LD.
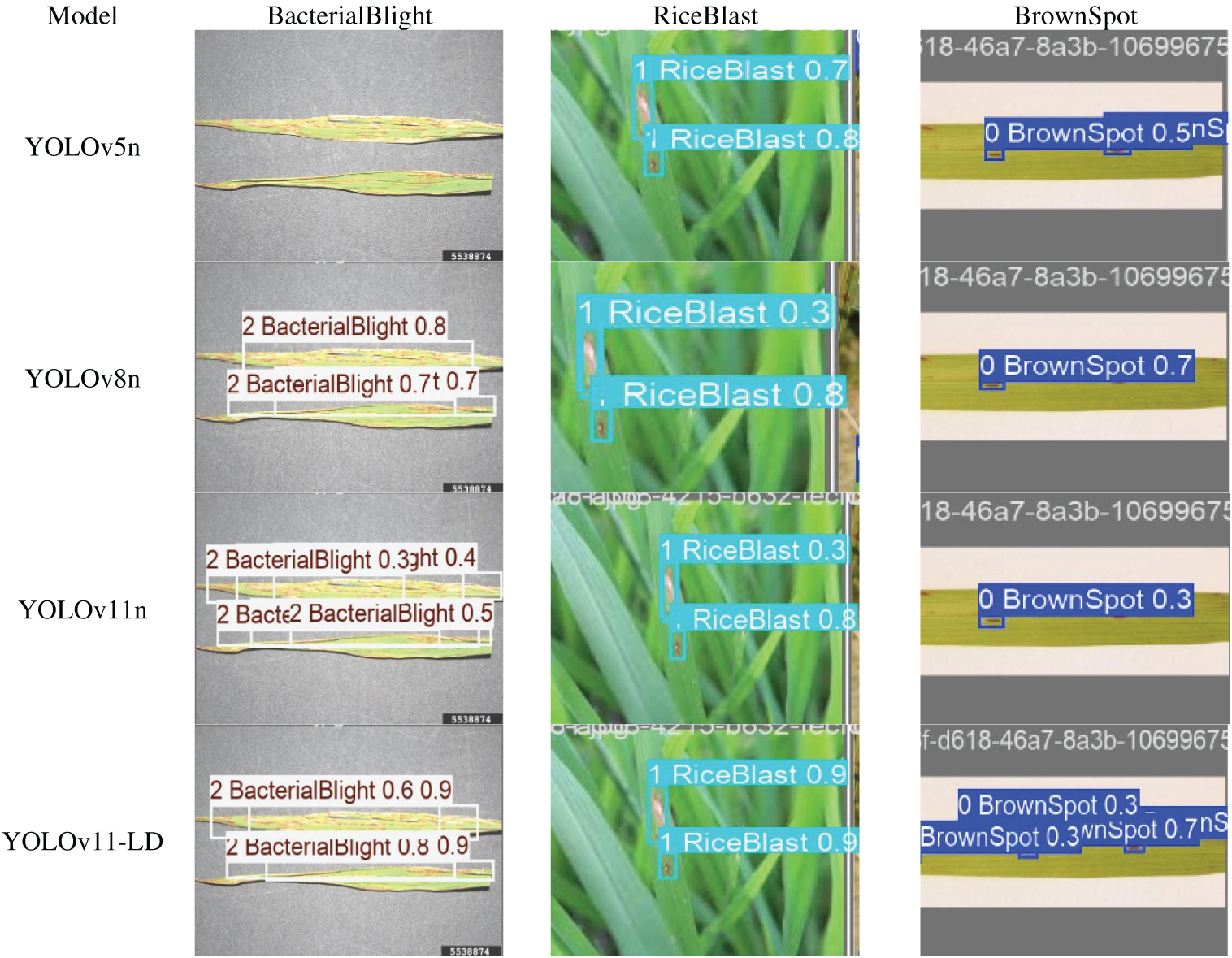

Figure 10: Comparison results of model performance.
In conclusion, this study proposed YOLO-LD, a lightweight and enhanced model based on YOLOv11n, developed to address the diverse types of rice diseases and the significant scale variations among targets. To support this research, we constructed a dataset comprising three multi-scale targets Bacterial blight, Rice blast, and Brown spot and secured model robustness through data augmentation techniques.
In terms of network architecture, we redesigned the backbone by incorporating the RepViT module and optimized detection performance by integrating BiFPN and CBAM. Experimental results demonstrated that YOLO-LD achieved an mAP of 94.5% and a Recall of 91.7%. While the proposed model exhibits a marginal increase in parameters (2.5M to 3.5M) and computational cost (6.4G to 7.8 GFLOPs) compared to the baseline, it achieves a substantial accuracy improvement of 3.8% (mAP@0.5). Furthermore, compared to other state-of-the-art lightweight models such as YOLOv10-n (10.6M), YOLOv11-LD remains significantly more compact and efficient, proving its superior operational feasibility in resource-constrained embedded environments.
Limitations and Future Work: Although the proposed YOLOv11-LD demonstrates superior performance in identifying specific rice diseases, the current dataset is limited to three disease classes and does not explicitly include a ‘healthy’ or ‘other’ class. In real-world paddy fields, this limitation could potentially lead to false positives where healthy leaves or unrelated background objects are misclassified as diseases. Future research will address this by expanding the dataset to include healthy samples and a wider variety of environmental distractors to enhance the model’s robustness and practical applicability in open-field scenarios.
Acknowledgement: Not applicable.
Funding Statement: This study was supported by research fund from Honam University, 2025.
Author Contributions: The authors confirm contribution to the paper as follows: writing—original draft preparation, Qingtao Meng; writing—review and editing, Sang-Hyun Lee; supervision, Sang-Hyun Lee. All authors reviewed and approved the final version of the manuscript.
Availability of Data and Materials: The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request. To facilitate understanding of the dataset structure, a subset of sample images and the annotation specifications have been provided as Supplementary Materials.
Ethics Approval: Not applicable.
Conflicts of Interest: The authors declare no conflicts of interest.
Supplementary Materials: The supplementary material is available online at https://www.techscience.com/doi/10.32604/cmc.2026.077207/s1.
References
1. Zhao M, Lin Y, Chen H. Improving nutritional quality of rice for human health. Theor Appl Genet. 2020;133(5):1397–413. doi:10.1007/s00122-019-03530-x. [Google Scholar] [PubMed] [CrossRef]
2. Sharma OP, Bambawale OM. Integrated management of key diseases of cotton and rice. In: Integrated management of diseases caused by fungi, phytoplasma and bacteria. Dordrecht, The Netherlands: Springer Netherlands; 2008. p. 271–302. doi:10.1007/978-1-4020-8571-0_14. [Google Scholar] [CrossRef]
3. Meng Q, Lee SH. Lightweight YOLOv5 with ShuffleNetV2 for rice disease detection in edge computing. Comput Mater Contin. 2026;86(1):1–15. doi:10.32604/cmc.2025.069970. [Google Scholar] [CrossRef]
4. Yang Y, Di J, Liu G, Wang J. Rice pest recognition method based on improved YOLOv8. In: 2024 4th International Conference on Consumer Electronics and Computer Engineering (ICCECE); 2024 Jan 12–14; Guangzhou, China. p. 418–22. doi:10.1109/iccece61317.2024.10504248. [Google Scholar] [CrossRef]
5. Arnal Barbedo JG. Digital image processing techniques for detecting, quantifying and classifying plant diseases. SpringerPlus. 2013;2(1):660. doi:10.1186/2193-1801-2-660. [Google Scholar] [PubMed] [CrossRef]
6. Mohanty SP, Hughes DP, Salathé M. Using deep learning for image-based plant disease detection. Front Plant Sci. 2016;7:1419. doi:10.3389/fpls.2016.01419. [Google Scholar] [PubMed] [CrossRef]
7. Ferentinos KP. Deep learning models for plant disease detection and diagnosis. Comput Electron Agric. 2018;145(6):311–8. doi:10.1016/j.compag.2018.01.009. [Google Scholar] [CrossRef]
8. Barbedo JGA. Factors influencing the use of deep learning for plant disease recognition. Biosyst Eng. 2018;172(660):84–91. doi:10.1016/j.biosystemseng.2018.05.013. [Google Scholar] [CrossRef]
9. Phadikar S, Sil J. Rice disease identification using pattern recognition techniques. In: 2008 11th International Conference on Computer and Information Technology; 2008 Dec 24–27; Khulna, Bangladesh. p. 420–3. doi:10.1109/ICCITECHN.2008.4803079. [Google Scholar] [CrossRef]
10. Zhou G, Zhang W, Chen A, He M, Ma X. Rapid detection of rice disease based on FCM-KM and faster R-CNN fusion. IEEE Access. 2019;7:143190–206. doi:10.1109/ACCESS.2019.2943454. [Google Scholar] [CrossRef]
11. Rahman CR, Arko PS, Ali ME, Iqbal Khan MA, Apon SH, Nowrin F, et al. Identification and recognition of rice diseases and pests using convolutional neural networks. Biosyst Eng. 2020;194(1):112–20. doi:10.1016/j.biosystemseng.2020.03.020. [Google Scholar] [CrossRef]
12. Li D, Wang R, Xie C, Liu L, Zhang J, Li R, et al. A recognition method for rice plant diseases and pests video detection based on deep convolutional neural network. Sensors. 2020;20(3):578. doi:10.3390/s20030578. [Google Scholar] [PubMed] [CrossRef]
13. Guo L, Wu Y, Zhao J, Yang Z, Tian Z, Yin Y, et al. Rice disease detection based on improved YOLOv8n. In: 2025 6th International Conference on Computer Vision, Image and Deep Learning (CVIDL); 2025 May 23–25; Ningbo, China. p. 123–32. doi:10.1109/CVIDL65390.2025.11085630. [Google Scholar] [CrossRef]
14. Jiang P, Chen Y, Liu B, He D, Liang C. Real-time detection of apple leaf diseases using deep learning approach based on improved convolutional neural networks. IEEE Access. 2019;7:59069–80. doi:10.1109/ACCESS.2019.2914929. [Google Scholar] [CrossRef]
15. Sangaiah AK, Yu FN, Lin YB, Shen WC, Sharma A. UAV T-YOLO-rice: an enhanced tiny yolo networks for rice leaves diseases detection in paddy agronomy. IEEE Trans Netw Sci Eng. 2024;11(6):5201–16. doi:10.1109/tnse.2024.3350640. [Google Scholar] [CrossRef]
16. Jia L, Wang T, Chen Y, Zang Y, Li X, Shi H, et al. MobileNet-CA-YOLO: an improved YOLOv7 based on the MobileNetV3 and attention mechanism for rice pests and diseases detection. Agriculture. 2023;13(7):1285. doi:10.3390/agriculture13071285. [Google Scholar] [CrossRef]
17. Senthil Kumar V, Jaganathan M, Viswanathan A, Umamaheswari M, Vignesh J. Rice leaf disease detection based on bidirectional feature attention pyramid network with YOLO v5 model. Environ Res Commun. 2023;5(6):065014. doi:10.1088/2515-7620/acdece. [Google Scholar] [CrossRef]
18. Saleem MH, Potgieter J, Mahmood Arif K. Plant disease detection and classification by deep learning. Plants. 2019;8(11):468. doi:10.3390/plants8110468. [Google Scholar] [PubMed] [CrossRef]
19. Liu J, Wang X. Early recognition of tomato gray leaf spot disease based on MobileNetv2-YOLOv3 model. Plant Methods. 2020;16(1):83. doi:10.1186/s13007-020-00624-2. [Google Scholar] [PubMed] [CrossRef]
20. Wang A, Chen H, Lin Z, Han J, Ding G. Rep ViT: revisiting mobile CNN from ViT perspective. In: 2024 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR); 2024 Jun 16–22; Seattle, WA, USA. p. 15909–20. doi:10.1109/CVPR52733.2024.01506. [Google Scholar] [CrossRef]
21. Tan M, Pang R, Le QV. EfficientDet: scalable and efficient object detection. In: IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR); 2020 Jun 13–19; Seattle, WA, USA. p. 10778–87. doi:10.1109/CVPR42600.2020.01079. [Google Scholar] [CrossRef]
Cite This Article
 Copyright © 2026 The Author(s). Published by Tech Science Press.
Copyright © 2026 The Author(s). Published by Tech Science Press.This work is licensed under a Creative Commons Attribution 4.0 International License , which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.


 Submit a Paper
Submit a Paper Propose a Special lssue
Propose a Special lssue View Full Text
View Full Text Download PDF
Download PDF Downloads
Downloads
 Citation Tools
Citation Tools