 Open Access
Open Access
ARTICLE
Evaluating Spanish Medical Entity Recognition: Large Language Models with Prompting versus Fine-Tuning
1 Departamento de Informática y Sistemas, Universidad de Murcia, Campus de Espinardo, Murcia, Murcia, Spain
2 Centro de Tecnología Biomédica, Universidad Politécnica de Madrid Campus de Montegancedo, Pozuelo de Alarcón, Madrid, Spain
3 Escuela Técnica Superior de Ingenieros Informáticos, Universidad Politécnica de Madrid, Campus de Montegancedo, Pozuelo de Alarcón, Madrid, Spain
* Corresponding Author: Rafael Valencia-García. Email:
Computers, Materials & Continua 2026, 87(3), 105 https://doi.org/10.32604/cmc.2026.077501
Received 10 December 2025; Accepted 26 February 2026; Issue published 09 April 2026
Abstract
The digitization of healthcare has resulted in the production of large amounts of structured and unstructured clinical data, creating the need for accurate and efficient named entity recognition (NER) to support medical procedures. This study evaluates and compares three approaches to NER in the medical domain in Spanish: using Large Language Models (LLMs) with In-Context Learning techniques (Zero-Shot, Few-Shot, and Chain-of-Thought); fine-tuning of LLMs; and fine-tuning of encoder-only models. Experiments were conducted on the Meddocan, Meddoprof, Meddoplace and Symptemist benchmark datasets. Fine-tuned encoder-only models achieve the best performance across all datasets, reaching macro-F1 scores of up to 76.71 on Meddocan, 71.51 on Meddoplace, 66.07 on Meddoprof and 63.50 on Symptemist. While LLMs with prompting offer flexibility and require no task-specific training, their performance varies significantly depending on the entity type. In addition, we evaluated fine-tuning of LLMs using QLoRA, but the improvements were limited due to the small amount of training data available per entity type, which made model adaptation less effective.Keywords
The rapid advancement of medical digitization is generating a large amount of structured and unstructured documentation, including electronic medical records, reports, consultations and other clinical texts. Recent studies show that electronic health records increasingly contain extensive unstructured narrative information that requires automated methods for efficient processing [1]. Automating the extraction of medical data reduces the administrative burden on healthcare professionals as Large Language Models (LLMs) and Natural Language Processing (NLP) systems can accurately identify diagnoses, medications and clinical events from lengthy documents [2–4]. This automation also minimizes the likelihood of human errors by standardizing the extraction process and improving the consistency of clinical data [5]. In addition, automated extraction enables faster access to critical information and supports more efficient integration of data into healthcare systems, enhancing clinical workflow and decision-making [6]. Consequently, there is a growing need to efficiently and accurately extract the relevant information from these documents.
Information extraction is a text mining process similar to preprocessing in classical data processing. It is mainly based on Machine Learning (ML) and NLP techniques, such as Named Entity Recognition (NER) and Relation Extraction (RE). According to [7], medical information extraction in clinics settings typically consists of three main steps: (1) extraction of medical problems, tests and treatments from discharge summaries and progress notes; (2) NER-based classification of statements about medical problems; and (3) classification of relationships between medical concepts.
In this context, NER in the medical domain involves identifying clinical events, such as diseases, symptoms or treatments, and their temporal expressions. A wide range of approaches has been proposed to effectively address this task, from traditional ML techniques relying on handcrafted features and domain-specific knowledge, such as rule-based systems [8], dictionaries [9], and conditional random fields (CRFs) [10], to advanced Deep Learning (DL) approaches based on transformer architectures (e.g., BERT and BioBERT) [11]. Advances in NLP, including contextual embeddings and pre-trained language models, together with the availability of various annotated medical corpora, such as Symptemist [12] and Distemist [13], have led to substantial improvements in medical NER performance.
More recently, the increasing number of NER-related studies and the emergence of generative models, particularly LLMs, have opened new research directions for medical NER. Open-source LLMs such as LLaMa-3 [14], Gemma-2 [15] and Phi [16] have demonstrated strong language understanding capabilities and the ability to generalize across tasks, raising the question of whether prompt-based or fine-tuned LLMs can reduce the dependency on large labeled datasets traditionally required by transformer-based NER models. More broadly, this shift reflects a transition from task-specific NLP pipelines towards general-purpose clinical language models capable of supporting multiple downstream tasks within a single framework.
Beyond NER, LLMs offer additional advantages that are particularly relevant in the clinical domain. Their instruction-following and generative capabilities enable a single model to support multiple information extraction tasks, such as relation extraction, entity normalization, clinical summarization and question answering, without requiring task-specific architectures or retraining [4,17]. This versatility facilitates the integration of NLP systems into real clinical workflows where heterogeneous tasks must be addressed jointly rather than in isolation. Furthermore, LLMs can be swiftly adapted to new annotation guidelines or entity inventories via prompting alone, making them ideal for dynamic clinical environments where requirements frequently evolve [18]. Despite these broader capabilities, NER remains a foundational component of clinical NLP pipelines, as it underpins downstream tasks such as relation extraction, normalization and clinical reasoning.
One of the main limitations of transformer-based models for NER in the medical domain is their dependence on large labelled datasets. To perform well, these models require extensive corpora spanning multiple medical specialities [19] and a wide variety of entity types. In contrast, LLMs have a clear advantage as they have been trained on large amounts of unstructured data and can generalize more easily to different medical sub-domains [17] without requiring large amounts of data specific to each one. Furthermore, LLMs can leverage their context generation and understanding capabilities to identify medical entities, reducing their reliance on labelled corpora and facilitating adaptation to new clinical scenarios [18].
Despite their potential, the practical effectiveness of LLMs for medical NER remains an open question, especially in non-English clinical settings. Most existing benchmarks and analyses focus on English-language datasets and tasks, reflecting the broader English-centric nature of current LLM evaluation practices and the limited availability of multilingual clinical resources. Surveys [20,21] highlight persistent challenges in adapting LLMs to diverse languages, particularly lower-resource ones, and studies of medical LLM datasets similarly reveal a concentration of resources in English, with less comprehensive evaluation in other languages such as Spanish. This leaves a limited understanding of how prompting-based LLMs, fine-tuned generative LLMs, and fine-tuned encoder-only models compare in other languages and under heterogeneous clinical conditions. In particular, Spanish medical NER has been underexplored, despite the availability of several high-quality annotated corpora covering diverse entity types and document structures.
Against this backdrop, the aim of this paper is to provide a systematic evaluation of medical NER approaches in Spanish by comparing multiple modeling paradigms under a unified experimental framework. Specifically, we analyze the performance of LLMs using different prompt engineering strategies, fine-tuned generative LLMs and fine-tuned encoder-only transformer models across four public Spanish medical datasets: Symptemist, Meddocan, Meddoprof, and Meddoplace.
To guide this study, we address the following research questions:
• RQ1. Can LLMs, when combined with various prompt engineering techniques such as Zero-Shot Learning (ZSL), Few-Shot Learning (FSL) and Chain of Thought (CoT), identify different medical entities?
• RQ2. Does fine-tuning pre-trained language models (such as BERT, RoBERTa, ALBERT, DistilBERT and XLM-RoBERTa) improve performance compared to LLMs?
• RQ3. Does fine-tuning generative LLMs for NER improve their performance compared to prompt-based use, and do these fine-tuned LLMs outperform fine-tuned encoder-only models?
In addition to the research questions, this work offers the following contributions:
• A unified benchmark evaluating prompting-based LLMs (ZSL, FSL, CoT), fine-tuned generative LLMs, and fine-tuned encoder-only models across four Spanish medical NER datasets (Meddocan, Meddoplace, Meddoprof, Symptemist).
• A standardized evaluation protocol, including strict entity-level macro-F1 scoring under IOB2, output-format validation for prompting approaches, and consistent BIO-based token-classification fine-tuning across architectures.
• A systematic comparison of modeling paradigms, revealing when prompting is viable, when generative LLM fine-tuning deteriorates, and why encoder-only models remain the most reliable option for Spanish clinical NER.
• Insights into model behaviour, including the impact of data sparsity, fine-grained entity inventories, and prompt sensitivity, as well as practical recommendations for clinical deployment.
In the healthcare domain, advances in digitization have generated large amounts of unstructured and structured textual data [22], such as clinical notes, medical records, conversations between medical professionals and patients and more [23]. These documents contain essential information that can drive advances in diagnosis, treatment and medical research, which has led to numerous studies and techniques related to information extraction in recent years. One of the key tasks in information extraction is NER, which involves automating the identification and recognition of medical entities, such as diseases, drugs and procedures, from large amounts of information.
Thus, the performance of these models has a direct impact on the quality of information retrieval and data analysis, as medical texts often contain complex and diverse terminologies. Accuracy in the identification and categorization of entities such as diseases, treatments, anatomical structures and drugs is critical for many healthcare applications, making NER an indispensable tool for the advancement of medical informatics [24].
NER in healthcare presents several challenges that affect performance and make it difficult to recognize certain medical entities. The complexity and specificity of medical language, which includes synonyms, acronyms and contextual meanings, requires continuous training of NER models [25]. In addition, recent studies have shown a large variability in the effectiveness of NER models on different medical datasets, which limits their applicability in the real world. Most of these models are based on advanced DL approaches that use pre-trained transformer-based language models and apply a fine-tuning process [26]. For example, in [27], they proposed a new formulation for the biomedical NER problem, treating the task as a reading comprehension problem instead of a classical sequence labelling problem. This improved approach used the BERT model for biomedical text mining and achieved state-of-the-art results on several biomedical NER datasets, such as BC5CDR and NCI-Disease, with F1-scores above 90%. On the other hand, reference [28] focused on the extraction of breast cancer information from Spanish clinical notes. They used pre-trained BERT and RoBERTa models and showed that transformer models, especially RoBERTa biomedical, achieved excellent accuracy with an F1-score of 95.01%. This study shows how fine-tuning pre-trained models, such as RoBERTa, can significantly improve the performance of medical entity extraction in Spanish clinical texts. Finally, reference [26] addressed the challenge of applying NER for the detection of eligibility criteria in clinical trials using transformer models. They compared different variants of BERT, ALBERT, RoBERTa and ELECTRA and showed that RoBERTa, when pre-trained with clinical notes from the MIMIC-III dataset, achieves the best results, with F1-scores up to 0.916 in clinical trial eligibility data.
One of the main limitations of transformer-based models for NER in the medical domain is their heavy reliance on large labelled datasets. These models typically achieve high performance by training on large, diverse corpora that span multiple medical specialities and a wide range of entity types, including diseases, treatments, medications, symptoms and medical procedures. There are several reasons for this limitation. First, collecting and annotating such large datasets is both time-consuming and costly, requiring significant human resources and domain expertise [29]. Second, the medical field is vast and constantly evolving, with new diseases, treatments and terminologies emerging regularly. This constant influx of new information requires models to be frequently retrained or fine-tuned to stay current. Third, medical terminology is highly specialized and often varies across specialities, regions or languages, further complicating the task of creating a comprehensive dataset that captures all possible entities and contexts [30,31]. Fourth, medical texts are often written in highly specialized, domain-specific language that can differ significantly from general language usage. This includes the use of abbreviations, acronyms, jargon and complex sentence structures. For example, a single term might have different meanings depending on the context, or different specialists may use different terms to refer to the same entity.
Generative models based on LLMs can produce human-like responses from textual input. As interest in these models has grown due to their remarkable ability to understand and process natural language, a variety of applications have been explored in fields such as healthcare, education, customer service and content creation. These models have also been trained on large amounts of data, enabling them to understand complex patterns in medical language, accurately identify medical entities and improve the consistency of their responses in different contexts.
Recent work on LLM-based NER has progressively shifted from framing NER strictly as token classification to treating it as an instruction-following or text-generation task [21]. Early efforts primarily explored Zero-Shot (ZS) and Few-Shot (FS) prompting to assess whether general-purpose LLMs could extract entities without task-specific training, often relying on carefully designed output schemas and task guidelines. More recent studies have moved toward stronger supervision signals, including instruction tuning and task-specific adaptation, showing that model behaviour can be significantly improved when the training objective and the prompting interface explicitly encode entity definitions and labeling conventions.
In the biomedical and clinical domain, this line of research has mostly been validated on English-language benchmarks, where both prompting and instruction-tuning approaches have shown promising results under controlled evaluation settings [32]. However, extending these findings to non-English clinical texts remains underexplored. Languages such as Spanish introduce additional challenges, including domain-specific terminology, regional lexical variation, and fewer publicly available annotated corpora [33]. Consequently, there is still limited evidence on how prompting-based LLMs, fine-tuned LLMs, and fine-tuned encoder-only models compare systematically for Spanish medical NER across heterogeneous entity types and datasets.
Against this backdrop, LLMs have been proposed as a promising alternative to mitigate some limitations of traditional medical NER, particularly in data-scarce settings. LLMs offer a significant advantage because they have been trained on large amounts of unstructured data, allowing them to better generalize to different medical subdomains without relying on large amounts of domain-specific data. In addition, these models can leverage their advanced generation and context understanding capabilities to identify medical entities with less reliance on labelled corpora. For example, a study by [34] explored the potential of LLMs such as GPT-3.5 [35] and GPT-4 [36] for clinical NER tasks, demonstrating that task-specific prompt engineering can significantly improve model performance. The study evaluated GPT models on extracting medical problems, treatments and tests from clinical notes, as well as identifying adverse events related to nervous system disorders from safety reports. Researchers achieved notable improvements in the models’ ability to recognize clinical entities by integrating structured prompts, including task descriptions, annotation guidelines and error analysis-based instructions [37].
Similarly, reference [38] introduced an instruction-tuning approach to improve biomedical NER using LLMs. Instead of treating NER as a sequence labelling task, their proposed paradigm transformed it into a text-generation problem, leveraging instruction-based learning. They developed BioNER-LLaMA, an LLM trained on biomedical entity recognition tasks, and tested it across multiple datasets containing disease, chemical and gene entities. The results showed that BioNER-LLaMA consistently achieved higher F1 scores compared to GPT-4 in FSL settings and, in some cases, matched the performance of domain-specific models like PubMedBERT and PMC-LLaMA.
Despite these advantages, the use of LLMs in medical settings is not without its challenges. The accuracy and reliability of their outputs remain critical concerns, particularly in clinical environments where errors may have serious consequences. Moreover, most existing studies on LLM-based medical NER have focused on English-language datasets, often under controlled experimental conditions, leaving open questions regarding their effectiveness in other languages and real-world clinical scenarios.
In this context, there is still a lack of systematic benchmarks that jointly evaluate prompting-based LLMs, fine-tuned generative LLMs and fine-tuned encoder-only models for medical NER in non-English settings. In particular, the Spanish language has been underrepresented in prior work, despite the availability of several high-quality annotated clinical corpora with heterogeneous entity types and document structures.
To address this gap, this paper presents a comprehensive empirical evaluation of medical NER in Spanish across multiple modeling paradigms. We compare LLMs using different prompting strategies (ZSL, FSL and CoT), fine-tuned generative LLMs using QLoRA, and fine-tuned encoder-only transformer models on four publicly available Spanish medical datasets: Symptemist [12], Meddocan [39], Meddoprof [40], and Meddoplace [41]. Beyond reporting performance metrics, our study analyzes cross-dataset patterns, identifies consistent strengths and limitations of each approach, and provides practical insights into when each paradigm is most suitable for Spanish medical NER.
This paper explores several approaches to medical entity recognition using LLMs and encoder-only models. First, the baseline performance of LLMs was evaluated using prompt engineering techniques such as ZSL, FSL and CoT, as illustrated in Fig. 1. ZSL involves providing the model with a single instruction or question, without any prior examples, and relies on its existing knowledge and general reasoning ability. In contrast, the FSL technique introduces the model to input and output examples within the same prompt to guide its behaviour and improve its accuracy in specific tasks. Finally, the CoT technique aims to make the model reveal its reasoning step by step before providing a final answer. This can improve comprehension in complex tasks such as the extraction of medical entities in different documents.
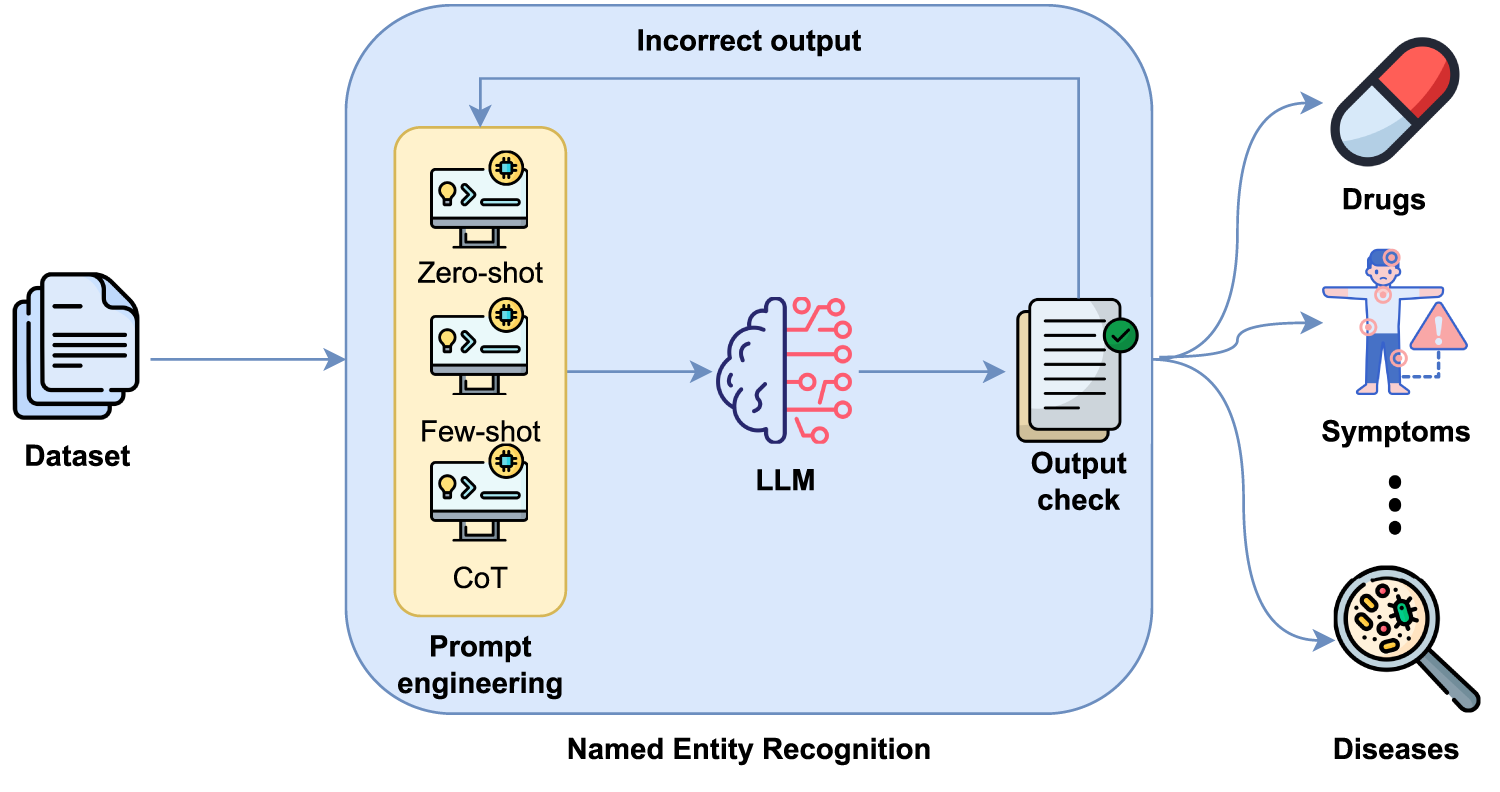
Figure 1: Entity detection based on LLMs and prompt engineering.
However, it should be noted that when LLMs are used with prompting strategies, the output generated by these models is not always consistent with the format structure explicitly defined in the prompt. This lack of consistency can negatively affect subsequent processing and performance evaluation. For this reason, an output check module has been included to validate that the generated response conforms to the expected format. If the required structure is not met, a new inference is performed that incorporates additional instructions explicitly indicating to the model that its previous response did not respect the required format, requesting strict adherence to the output format. This process can be repeated up to three times to ensure that the final output strictly conforms to the defined schema. This improves the robustness and reliability of the model’s responses.
In order to evaluate LLMs in the context of Spanish NER, we developed prompts for three strategies: ZSL, FSL and CoT. All prompts follow a consistent structure to ensure reliable output and enable automated scoring.
The ZSL prompt includes a task description and output format specifying how entities should be extracted, separated (e.g., using hash symbols “#”) and how missing types should be handled (e.g., by returning “null”). The FSL prompt builds on this by providing input-output examples to help the model recognise patterns, particularly in ambiguous or domain-specific cases.
For FSL, five examples are randomly selected per dataset. Each example includes the full clinical text and the corresponding gold-standard entity annotations, providing both the entity type and the extracted surface form. Consequently, the model is exposed to both entity labels and realistic textual mentions. The five examples are sampled from the training split using a fixed random seed to ensure reproducibility, and the same example set is used acroos all evaluated LLMs for a given dataset. This controlled sampling procedure guarantees that performance differences between models are not attributed to different few-shot inputs.
The CoT prompt provides a step-by-step explanation of the reasoning behind the extraction. It guides the model through the processes of understanding, classification and formatting, which is useful when a deeper contextual understanding is required. Additional details and illustrative examples of the ZSL, FSL and CoT prompting strategies can be found in Appendices A–C.
All LLM outputs follow a strictly defined structured format, which is explicitly specified in the prompt and involves grouping entities by type using a fixed schema. To ensure the validity of the format and enable automatic evaluation, an output verification process is applied to validate that the generated response conforms to the expected format. If the generated output does not match the expected structure, the model is automatically re-prompted, indicating that the previous response did not respect the required format and providing additional constraints to correct it. This validation–re-prompting cycle can be repeated up to three times to guarantee that the final output adheres precisely to the defined schema. This process ensures robust scoring across all models and minimises inconsistencies in the evaluation pipeline.
We also evaluated the use of fine-tuning for NER using supervised training (see Fig. 2), where entities are annotated following the BIO tagging scheme.
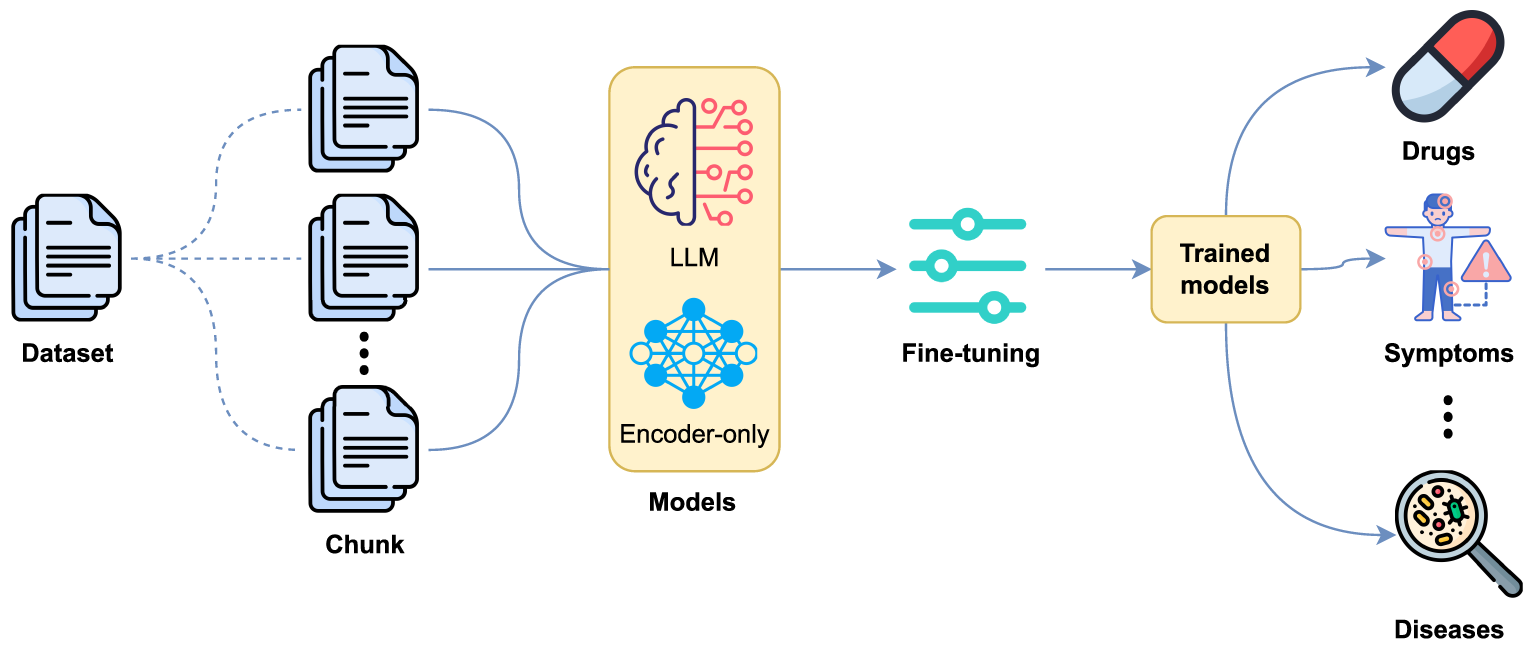
Figure 2: Entity detection based on fine-tuning with LLMs and encoder-only models.
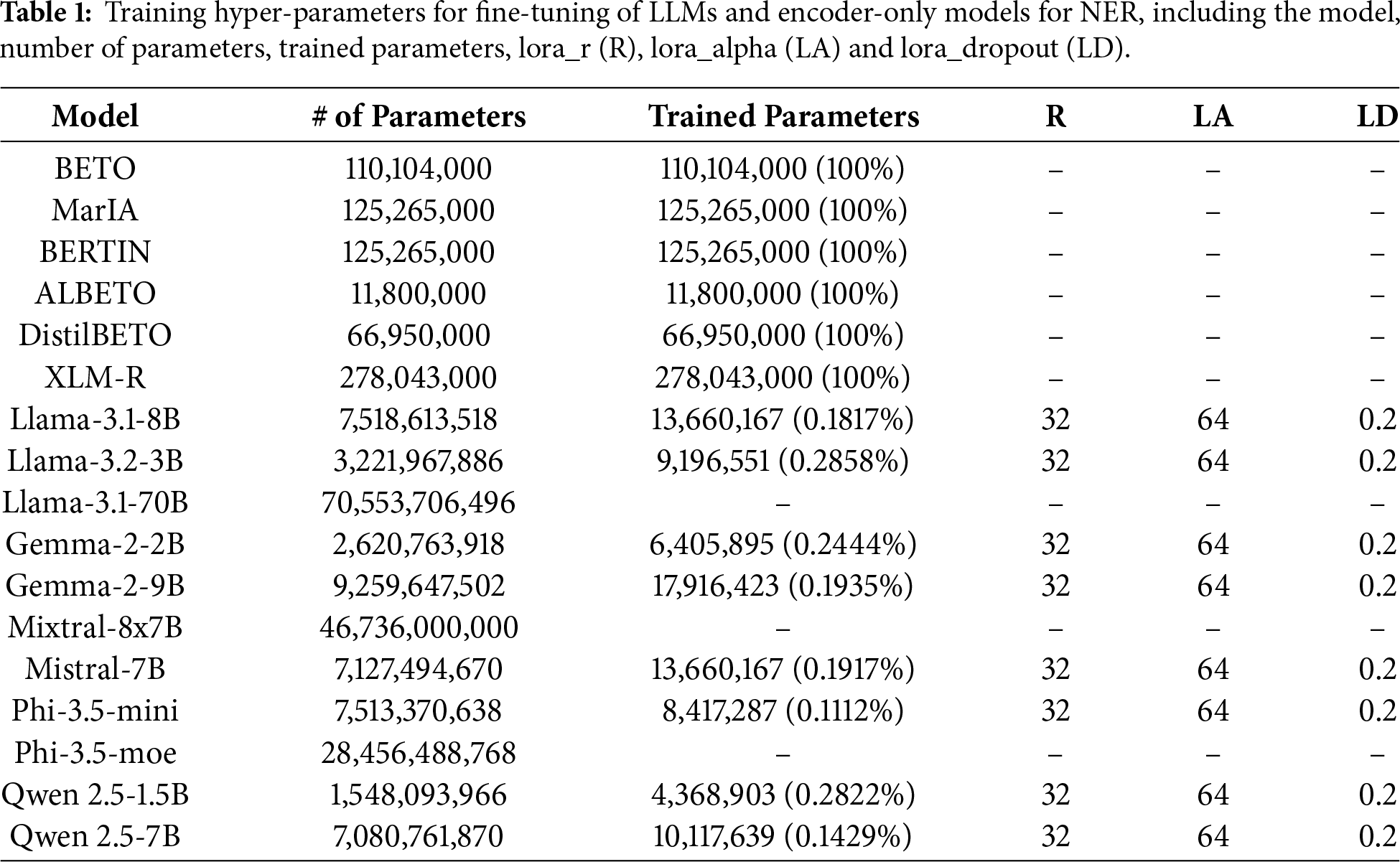
Two types of models were considered: decoder-based LLMs, including LLaMA (3.1 [14] and 3.2 [42] versions), Gemma 2 [15], Mixtral [43], Mistral [44], Phi 3.5 [16] and Qwen 2.5 [45]; and encoder-only models, including BETO (cased and uncased) [46], MarIA [47], BERTIN [48], ALBETO [49], DistilBETO [49] and XLM-RoBERTa [50]. These models are optimized for classification and token-level labelling, making them particularly suitable for NER. Table 1 shows the percentage of parameters trained on each model.
Encoder-only models were trained in a token-level classification setting, making them particularly suitable for BIO-based NER. Furthermore, when working with encoder-only models, clinical documents were segmented into fixed-length chunks prior to fine-tuning in order to address input length limitations. Each document was split into contiguous segments of up to 200 tokens, without overlap. Special care was taken to preserve entity integrity under the BIO tagging scheme: chunk boundaries were adjusted to avoid splitting entities, ensuring that no segment ended with an inside tag (I–XXX). If a potential boundary fell within an entity span, the cut was shifted backward until a valid boundary was found.
Each resulting chunk retained a reference to its original document identifier, allowing evaluation to be performed consistently at the chunk level. This strategy ensures compatibility with model input constraints while minimizing annotation noise caused by entity fragmentation.
For LLMs, the NER task is formulated as a token-level classification problem using the same BIO tagging scheme applied to the encoder-only models. Each model is adapted by adding a small classification layer on top of the final hidden representation of every token, enabling the system to assign a BIO label to each input token rather than generating text in free-form. All parameters of the original decoder-only model remain frozen during fine-tuning. The NER adaptation is performed by adding two trainable components: (1) a token-classification head, and (2) the low-rank LoRA/QLoRA adapter matrices injected into selected projection layers. Only these components are updated during training, while all base model weights remain unchanged. This setup enables efficient fine-tuning under low memory conditions while preserving the original generative model. Input sequences are first tokenized into subword units. Special tokens and padding positions are ignored by the loss function. For words that split into multiple subword tokens, the BIO label assigned to the first subword is propagated to the remaining ones in order to maintain consistency with the gold annotations. The model then predicts one label per token from the shared BIO tag set. Unlike encoder-only models, decoder-based LLMs can process longer clinical documents directly thanks to their larger context window, which avoids the need for document chunking. This setup ensures that both model families are trained under a comparable BIO-based token-classification formulation, differing mainly in their architectural structure rather than in the labeling scheme.
The LLMs were chosen for their multilingual capabilities and strong contextual understanding. All models were tuned with a learning rate of 2e–5, a batch size of 16 and were trained for 10 epochs. The evaluation metric used was the macro F1 score.
3.3 Output Validation and Structured Entity Extraction for Prompting-Based LLMs
When LLMs are used with prompting strategies (ZS, FS and CoT), they generate free-form textual outputs that must be converted into structured entity predictions in order to perform NER evaluation. In contrast, fine-tuned LLMs and encoder-only models produce structured predictions directly and therefore do not require additional output validation.
To ensure compatibility with standard NER evaluation, we implemented a strict output validation and post-processing pipeline specifically for prompting-based LLMs. Models are instructed through system-level prompts to return entities following an explicit and fixed schema, where each entity type is listed separately and entity mentions are separated using predefined delimiters.
After generation, each model output is automatically checked to verify whether all expected entity types are present and correctly formatted. Outputs that do not comply with the required format are considered invalid. To improve robustness, an automatic retry mechanism is applied: if an invalid output is detected, the model is prompted again with an additional reflexive instruction explicitly requesting strict adherence to the output format. This process is repeated for up to three attempts.
Once a valid response is obtained, the output text is parsed using regular expressions to extract entity mentions for each entity type. These mentions are normalized into a structured JSON representation, which is subsequently used for evaluation. This procedure ensures a fair and consistent evaluation of prompting-based LLMs alongside fine-tuned LLMs and encoder-only models.
In order to accurately measure the ability of these models to correctly identify and classify entities such as the names of diseases, drugs, locations and healthcare professionals, a variety of datasets are used to address the diversity and complexity of medical texts.
• Symptemist [12] A dataset consisting of 1000 Spanish clinical reports, manually annotated for mentions of symptoms, signs and clinical findings following SNOMED CT guidelines. The texts are sourced from the SPACCC corpus and cover various medical specialities. Annotation was performed by clinical experts and validated through iterative quality controls. The corpus contains a single entity type.
• Meddocan [39] This corpus is designed for the de-identification of clinical documents and contains 1000 cases annotated with Protected Health Information (PHI), including names, dates, institutions and personal identifiers. The annotations were produced by clinical and linguistic experts according to strict guidelines, achieving a 98% inter-annotator agreement rate. The dataset includes 22 entity types.
• Meddoprof [40] It contains 1844 clinical cases from over 20 medical specialities, annotated with mentions of professions and occupational statuses. The aim is to support the development of systems for recognizing and standardizing professions in Spanish medical texts. Annotations were harmonized using the ESCO classification and SNOMED CT, and validated through clinician–linguist cross-checking. The corpus includes 3 entity types.
• Meddoplace [41] It contains 1000 clinical reports selected for their high density of geographic and location mentions. More than 10,000 entities were annotated and normalized to GeoNames, PlusCodes and SNOMED CT, covering five classes of clinical interest. The dataset defines 10 entity types.
This section analyses the results obtained by different approaches using LLMs and encoder-only models applied to the task of medical entity recognition with the Meddocan, Meddoplace, Meddoprof and Symptemist datasets.
All models are evaluated on the corresponding test split using a unified NER evaluation protocol: predictions are obtained at token level under the IOB2 tagging scheme and converted to word-level labels, and we report macro-averaged F1 scores at entity level using the seqeval library with strict span matching.
First, three prompting configurations were evaluated: ZSL, FSL and CoT, the results of which are shown in Table 2. In general, it is observed that model performance improves substantially when moving from a ZSL to a FSL approach, which confirms the importance of including examples in the prompt for specialized tasks such as clinical entity extraction. Also, the use of CoT strategies allows for additional improvement in some models, although this is not uniform across datasets or models.
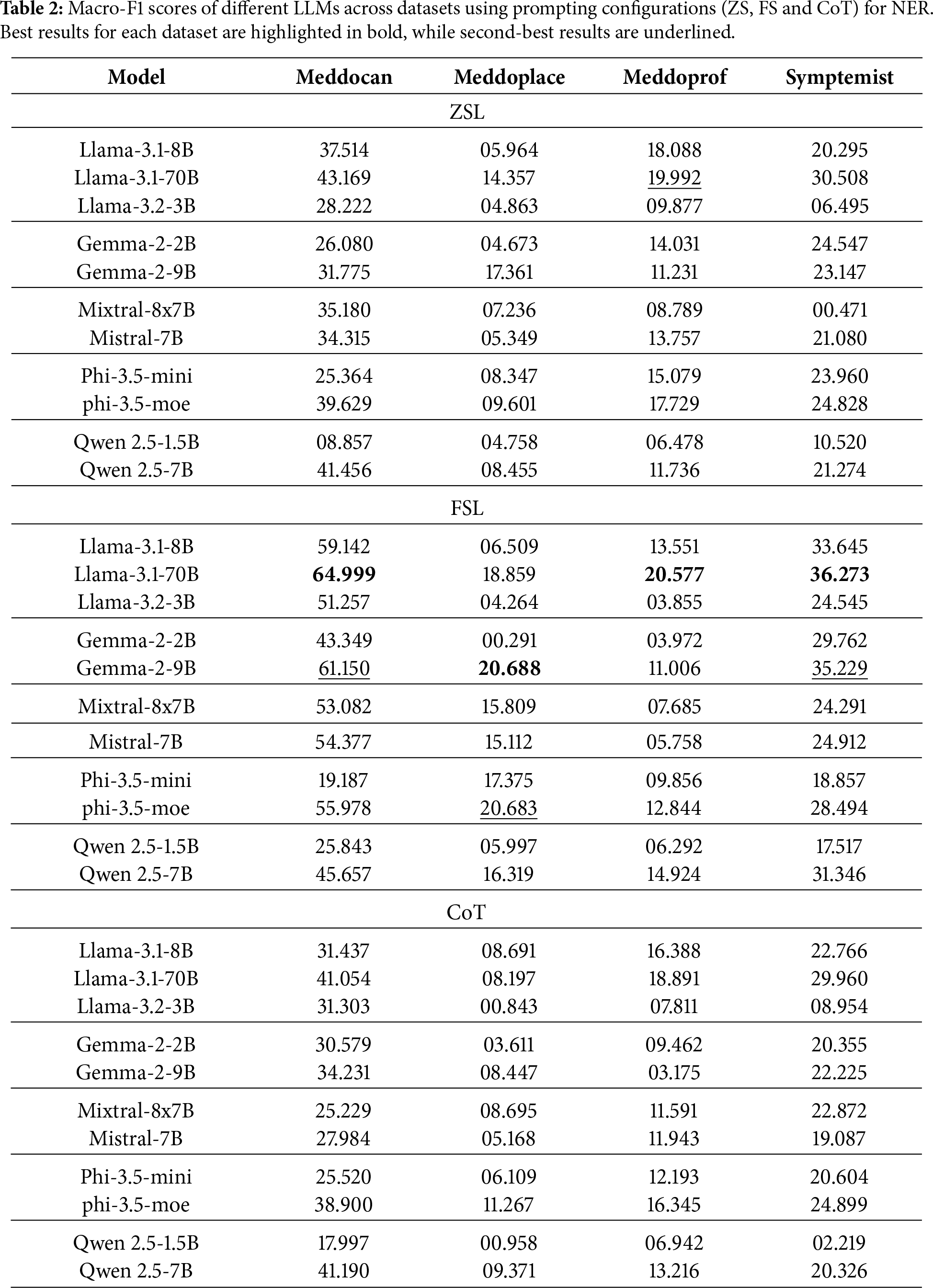
In the case of Meddocan, which focuses on sensitive entity de-identification, the best performing models are Llama-3.1-70B in FSL configuration achieving up to an F1-score of 64.999 and Gemma-2-9B in FSL configuration achieving an F1-score of 61.150. These results show that larger models tend to better capture entity structure in complex clinical texts, although smaller models such as Llama-3.1-8B also achieve competitive results with F1-score of 59.142 in the FSL configuration.
For Meddoplace, oriented to geographic location recognition, a lower overall performance is seen in all models. Despite this, Gemma-2-9B stands out with an F1-score of 17.361 in ZSL configuration, while the same model obtains a better result in FSL with a F1-score of 20.688.
As for Meddoprof, which attempts to identify occupations and occupational status, performance is generally moderate. Llama-3.1-70B obtains F1-scores of around 20% in both ZSL and FSL configurations. For Symptemist, which focuses on the detection of symptoms and clinical findings, Llama-3.1-70B stands out with an F1-score of around 30% in both ZSL and CoT configurations, while the same model obtains a better result with FSL, with a F1-score of 36.273.
Table 3 shows the best results obtained for each dataset using prompting techniques and LLMs. For the Meddocan dataset, the FS configuration achieves the best performance with an F1-score of 64.99, while the second-best result is also obtained with FS prompting, reaching an F1-score of 61.15. This strong performance can be attributed to the structured nature of Meddocan reports and the dataset’s focus on PHI entities such as names, dates, instructions and personnel identifiers, which are generally easier to identify compared to other datasets.
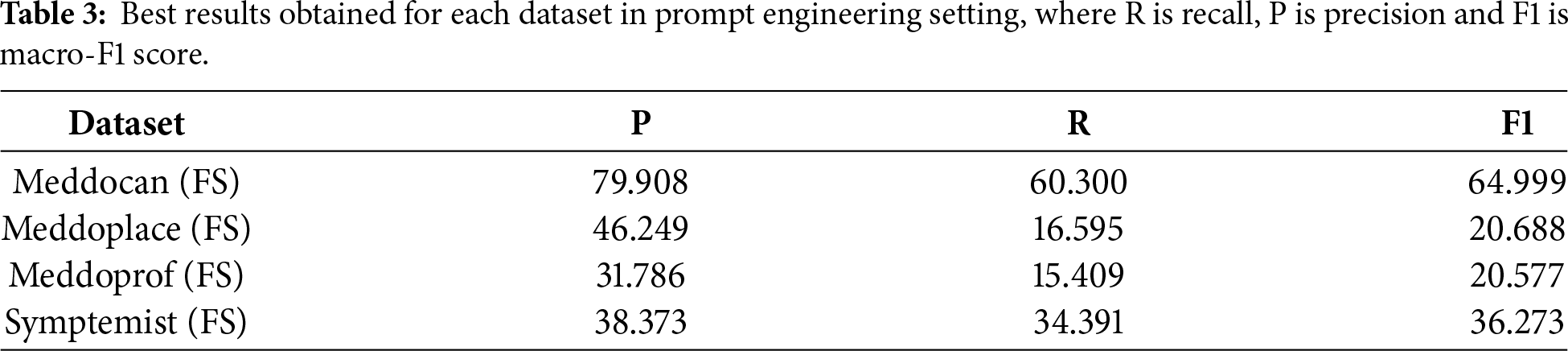
For the Meddoprof dataset, which focuses on the identification of professions within medical texts, prompting techniques with LLMs achieve their best performance under FS configuration, with an F1-score of 20.688. The second-best result is also obtained with FS prompting, reaching an F1-score of 20.683. The relatively low performance on this dataset can be attributed to the length and complexity of the clinical documents, compared to other datasets such as Meddocan, which makes the identification of professional roles particularly challenging.
As for the Meddoplace dataset, which focuses on the identification of geographical and location mentions within medical texts, the best performance obtained with prompting techniques corresponds to the FS configuration, achieving an F1-score of 20.577. The second-best result is obtained with the ZS configuration, with an F1-score of 19.992. Overall performance on this dataset remains lower than on others, which can be attributed to the fact that many hospital and clinic names are not part of the prior knowledge of LLMs, as they were not explicitly trained on this type of location-specific information.
Finally, for the Symptemist dataset, which focuses on the identification of symptoms, signs and clinical findings within medical texts, the best performance obtained with prompting techniques corresponds to the FS configuration, achieving an F1-score of 36.273. The second-best result is also obtained with FS prompting, with an F1-score of 35.229. This relatively higher performance compared to other datasets reflects the fact that symptom-related entities are more explicitly expressed in clinical narratives and are better represented in the prior knowledge of LLMs.
Beyond individual dataset results, several consistent patterns emerge across models and prompting strategies. First, FSL clearly outperforms ZSL and CoT prompting in all evaluated datasets. This suggests that, for Spanish medical NER, providing task-specific examples is more beneficial than relying solely on model priors or explicit reasoning steps. In particular, FSL appears to help LLMs better internalize entity boundaries and dataset-specific annotation conventions, which are critical for NER tasks.
Second, the effectiveness of prompting strategies strongly depends on the nature of the target entities. Datasets involving well-defined and surface-level entities, such as PHI elements in Meddocan or symptom mentions in Symptemist, consistently achieve higher performance. In contrast, datasets such as Meddoprof and Meddoplace, which involve semantically ambiguous entities (e.g., professions or locations) and longer clinical documents, remain challenging for prompting-based LLMs. In these cases, LLMs tend to over-generate entity mentions, leading to lower precision.
Third, model scale has a noticeable but secondary impact compared to prompting strategy and dataset characteristics. Larger models, such as Llama-3.1-70B, generally achieve better results, particularly under FSL configurations. However, size alone is insufficient to overcome limitations related to domain specificity, long document structure, and ambiguous entity definitions.
From a practical perspective, these findings indicate that prompting-based LLMs can be a viable solution for Spanish medical NER in scenarios where annotated data is scarce and entity definitions are relatively explicit. However, their performance degrades significantly for complex entity types and long clinical narratives, suggesting that prompting alone is insufficient for robust, high-precision medical NER in real-world settings.
To maximize the performance of LLMs in medical entity recognition, two fine-tuning based approaches were tested.
First, LLMs were trained for detection purposes only. In general, LLMs are generative models designed to produce text or responses based on user input. In this case, the model was fine-tuned to focus its output solely on identifying specific medical entities, formulated as a token-level classification task rather than free-form text generation.
To this end, LLMs have been adapted to token classification tasks using dedicated architectures and QLoRA-based training. This fine-tuning technique allows LLMs to be quantized to lower precision (e.g., 4-bits or 8-bits) and, instead of directly modifying the entire model, introduces small low-rank matrices that are trained to adjust the output without changing the original parameters. In our implementation, QLoRA adds trainable low-rank adapter matrices on top of the frozen decoder-only model. Together with these adapters, the token-classification head is also trained. All other parameters of the base model remain frozen. This means that only a percentage of the model is trained, reducing hardware requirements without significantly compromising performance.
Finally, another approach is based on the fine-tuning of Spanish-only encoder models, as well as multilingual models such as XLM-RoBERTa. These models are widely used for NER tasks in the biomedical domain. However, unlike generative LLMs, these models are lighter and specifically designed for contextual representation of input tokens, which makes them more efficient in token classification tasks.
Table 1 shows the percentage of parameters trained on each model and Table 4 details the results obtained by each model fine-tuned using fine-tuning configurations (LLMs fine-tuned with QLoRA and fine-tuned encoder-only models) on different datasets.
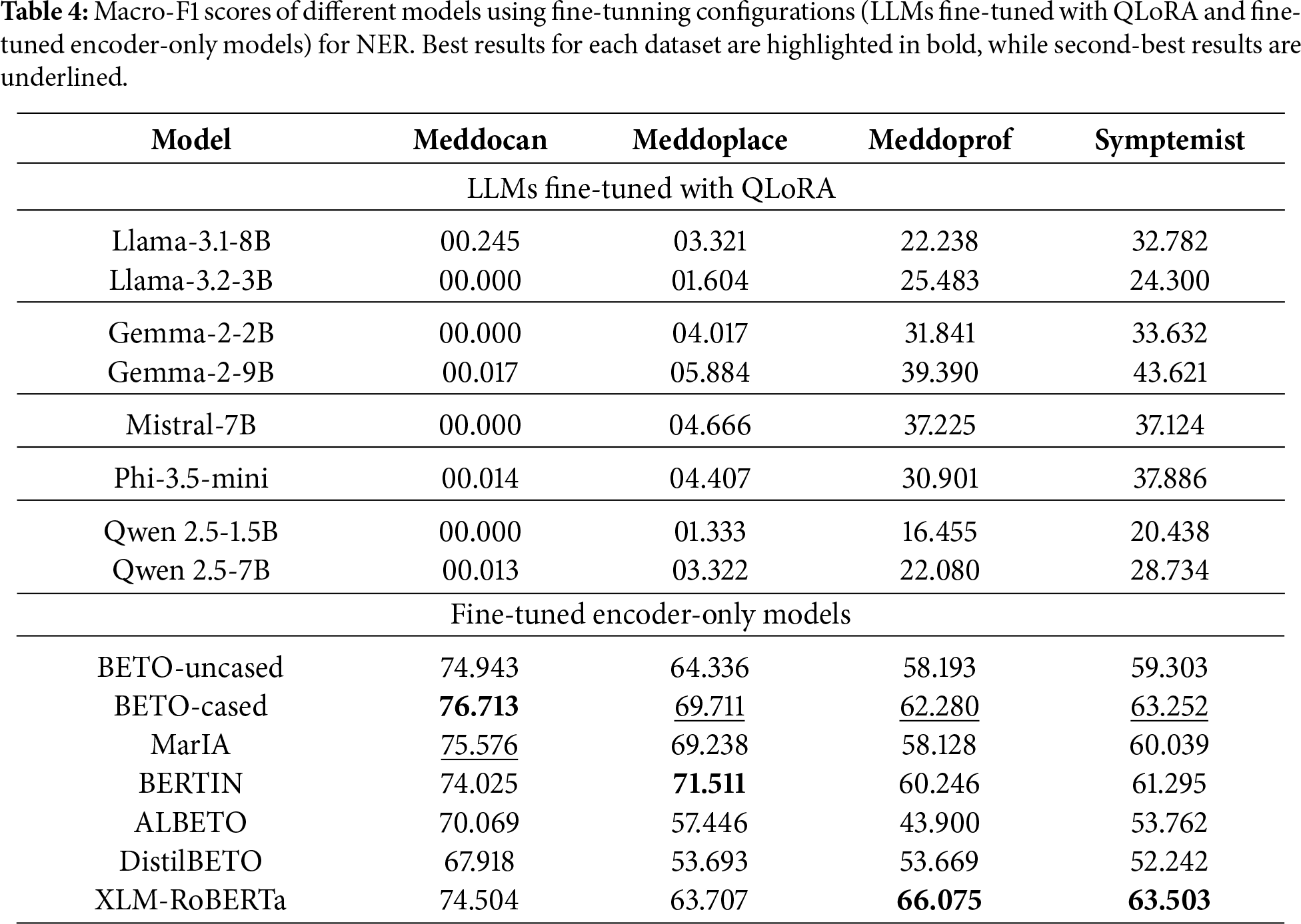
The fine-tuning of generative LLMs shows irregular behaviour. In some cases, such as Symptemist (F1-score of 43.62) and Meddoprof (F1-score of 39.39), it outperforms prompting, indicating that fine-tuning can improve the detection of certain medical entities. However, in other datasets, such as Meddocan and Meddoplace, performance is significantly lower (F1-scores of 0.24 and 5.88, respectively). This is partly due to the fact that these datasets contain large medical documents and the number of examples available for fine-tuning was insufficient, which introduced more noise rather than improving the baseline performance. Moreover, due to GPU memory and computational constraints, only a small proportion (less than one percent) of the generative LLMs’ parameters were trained with QLoRA in fine-tuning experiments, as shown in Table 1.
To better understand these extremely low values, we performed an additional error analysis. The near-zero performance (
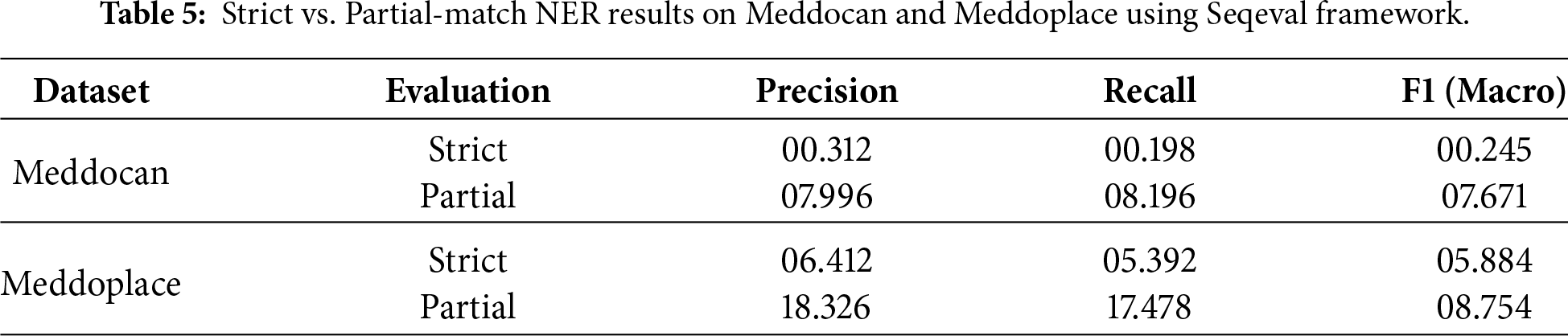
Therefore, to better characterize the actual behaviour of the models and disentangle span detection errors from type classification errors, we conducted a complementary error analysis focused specifically on the Meddocan and Meddoplace datasets. In this additional study, we removed the strict constraint by evaluating the predictions under “partial” matching mode of the best model (Llama-3.1-8B and Gemma-2-9B), as shown in 5, while keeping the rest of the experimental setup unchanged. This allows us to assess whether the models can identify the correct regions of text corresponding to named entities, even when the predicted entity type does not exactly match the gold annotation.
The comparison between strict and partial evaluation is shown in Table 5 reveals a consistent and substantial performance gap across both Meddocan and Meddoplace. While strict evaluation yields near-zero macro-F1 scores, partial matching increases the score by more than an order of magnitude in both datasets. This suggests that, while the models can often identify the correct textual regions corresponding to named entities, they struggle to assign the correct fine-grained entity type. Consequently, the degradation observed under strict evaluation is primarily driven by type misclassification rather than errors in detecting entity boundaries.
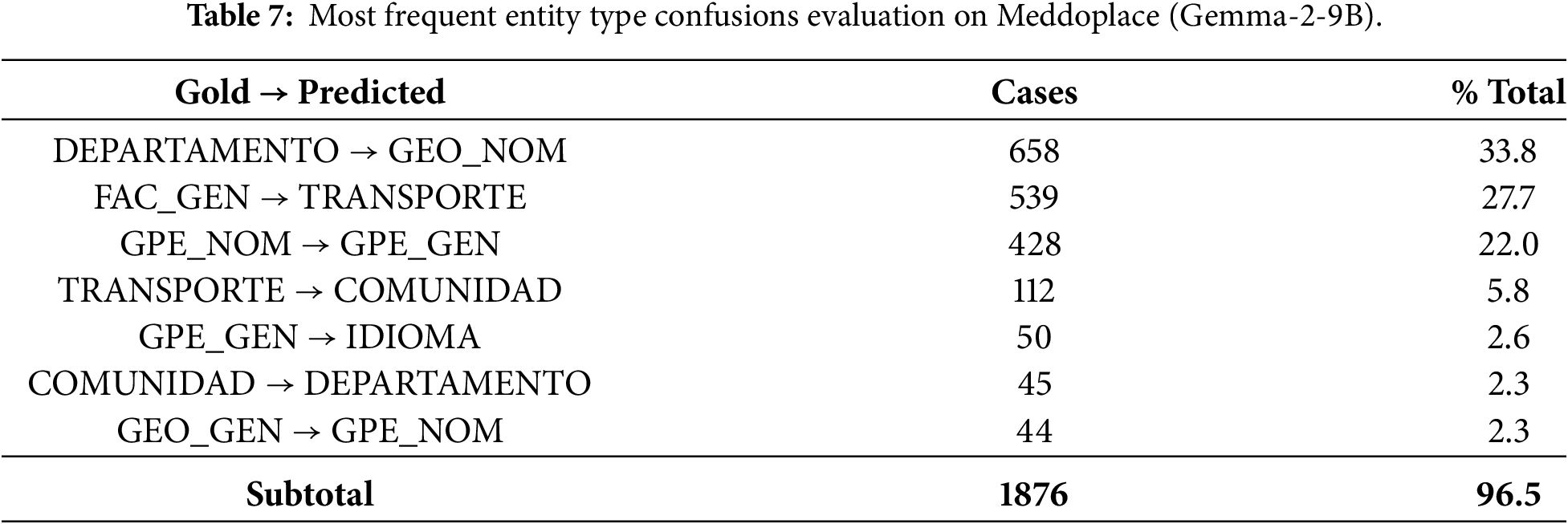
To further investigate the origin of the strict evaluation failures, we conducted a detailed, entity-level confusion analysis. This analysis focuses exclusively on entities whose span boundaries exactly match the gold annotation, but for which the predicted entity type differs. This isolates pure type classification errors under strict evaluation. Tables 6 and 7 report the most frequent Gold
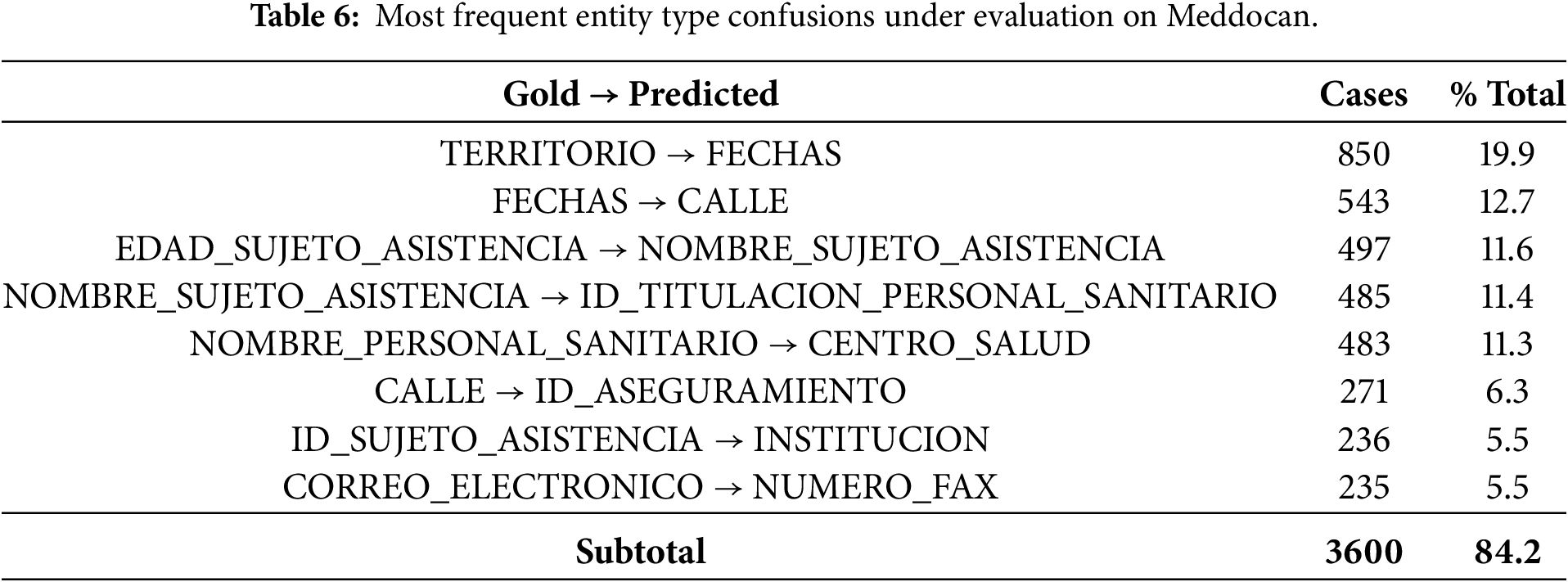
The results in Table 6 show that type errors in Meddocan are highly concentrated. The eight most frequent errors account for over 84% of all type mismatches, suggesting systematic error patterns rather than random occurrences. These errors predominantly occur between semantically related categories, such as territorial entities, temporal expressions, personal identifiers and institutional roles. Notably, these errors arise despite correct entity boundary detection, suggesting that while the model can identify entity spans, it struggles to distinguish between closely related entity subtypes reliably.
A similar, but even more pronounced, pattern is observed in Meddoplace, where the seven most common confusions explain more than 96% of all type errors, these are reported in Table 7. The most frequent confusions involve medical departments, geographic entities and transportation-related categories, reflecting the semantic overlap inherent in the dataset annotation scheme. This extreme concentration of errors results in a particularly low macro-F1 score under strict evaluation, highlighting the sensitivity of strict span–type matching to fine-grained label distinctions in low-resource settings.
These confusion patterns provide a direct explanation for the collapse of macro-F1 score under strict evaluation. Since macro-F1 score assigns equal weight to each entity type, the failure to correctly classify many categories, many of which receive a score of zero, dominates the final score, even when the model successfully detects entity spans. Consequently, a small number of frequent type confusions is sufficient to bring the overall macro-F1 score close to zero.
Taken together, the strict vs. partial comparison and the confusion-based error analysis demonstrate that the low strict macro-F1 scores should not be interpreted as an inability of the models to perform NER or to generate valid BIO sequences. Rather, they reflect the interaction between highly granular annotation schemes, extreme class imbalance, limited fine-tuning capacity under QLoRA constraints and an evaluation protocol requiring exact span–type matches.
On the other hand, the fine-tuned encoder-only models show clearly superior performance on all evaluated datasets. In Meddocan, Meddoplace and Meddoprof, these models achieve F1-scores of 76.71, 71.51 and 66.07, respectively, which is a significant improvement over the other methods. In Symptemist, where even the fine-tuning of the LLMs showed a slight improvement, the encoder-only models achieve the best result, specifically XLM-RoBERTa, with an F1-score of 63.50, demonstrating their robustness under different conditions.
A cross-dataset analysis of the fine-tuning results reveals several consistent patterns that provide insight into the strengths and limitations of each approach. First, fine-tuned encoder-only models clearly and consistently outperform both prompting-based LLMs and fine-tuned generative LLMs across all datasets. This trend holds regardless of entity type, document length or dataset size, highlighting the strong inductive bias of encoder-only architectures for token-level NER tasks.
Table 8 presents a comparison of the best macro-F1 scores obtained with the three evaluated strategies across the four datasets. This is complemented by a statistical significance analysis based on an approximate randomization test (ART) with 10,000 iterations. The strategies correspond to: prompting with LLMs (S1); fine-tuning generative LLMs using QLoRA (S2); and fine-tuning encoder-only transformer models (S3). In addition to reporting absolute performance differences (
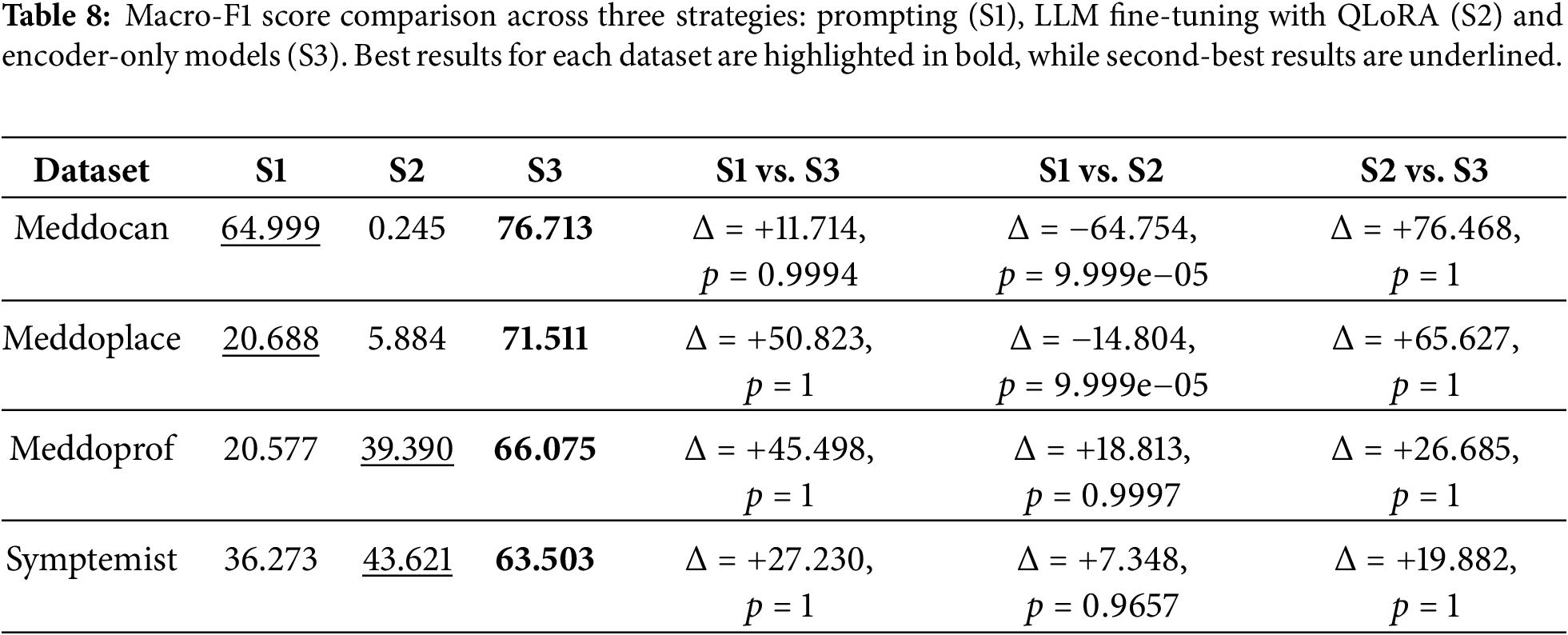
Overall, the results reveal a clear and consistent performance hierarchy across all datasets. Fine-tuned encoder-only models (S3) consistently achieve the highest macro-F1 scores, significantly outperforming prompting with LLMs (S1) and fine-tuned LLMs using QLoRA (S2). ART analysis confirms that these improvements are statistically significant in all S3-related comparisons, with p-values close to 1 indicating consistent superiority at document level. These findings are important because they show that the large absolute differences observed in macro-F1 score correspond to statistically reliable gains. This emphasises the value of complementing point estimates with significance testing in NER evaluation.
Firstly, prompting-based approaches (S1) demonstrate moderate but highly variable performance, depending on the dataset. For Meddocan, prompting achieves a relatively strong macro-F1 score of 64.99, which partially narrows the gap with fine-tuned encoder-only models. However, the ART comparison between S1 and S3 for this dataset yields a p-value close to 1, indicating that the performance difference is statistically significant and consistently favours encoder-only models at the document level. In contrast, prompting performs poorly on Meddoplace and Meddoprof, with macro-F1 scores around 20, confirming its limited robustness in datasets with more complex entity distributions or stricter boundary requirements.
Secondly, fine-tuned generative LLMs with QLoRA (S2) exhibits mixed and strongly dataset-dependent behaviour. Although S2 achieves a higher absolute macro-F1 score than prompting in Meddoprof and Symptemist, the ART comparisons between S1 and S2 yield p-values close to 1, suggesting that these improvements are statistically significant. Conversely, in Meddocan and Meddoplace, fine-tuned generative LLMs perform substantially worse than prompting. The corresponding ART comparisons yield very small p-values (p < 0.001), confirming that the degradation introduced by fine-tuning is statistically significant. These results highlight the instability of fine-tuned generative LLMs for NER and its sensitivity to dataset characteristics.
Thirdly, the ART analysis clearly confirms the superiority of fine-tuned encoder-only models (S3) over fine-tuned generative LLMs (S2) across all datasets. All S2 vs. S3 comparisons yield p-values close to 1, indicating statistically significant and consistent improvements in favour of encoder-only architectures. This suggests that the large absolute performance gaps observed in macro-F1 scores are not sporadic effects, but rather reflect systematic advantages at the document level. Encoder-only models are particularly effective in datasets such as Meddocan, which require precise span boundary detection and strict adherence to annotation guidelines, properties that align naturally with token-level classification objectives.
Fourthly, the observed trends are also shaped by computational constraints and experimental design. Due to hardware limitations, fine-tuning experiments were restricted to small and medium-sized generative LLMs, while larger models were evaluated exclusively under a prompting approach. While this restricts direct architectural comparability across all model scales, the consistent statistical dominance of encoder-only models, confirmed by ART, suggests that architectural suitability for token-level prediction, rather than model size alone, is the primary factor driving effective Spanish medical NER performance.
From a practical perspective, these findings suggest that fine-tuned encoder-only models is the most reliable and effective approach to Spanish medical NER when annotated data is available. While prompting can yield competitive results in specific datasets, it lacks consistency. Fine-tuned generative LLMs exhibit unstable behaviour and are not a robust alternative for high-precision clinical entity recognition. Importantly, the inclusion of ART-based significance testing shows that the observed differences in macro-F1 scores are statistically reliable and should be interpreted as systematic performance trends rather than random variation.
The analysis of the results confirms the initial doubts. With regard to RQ1, which asks whether LLMs combined with different query techniques can identify various medical entities, Table 3 shows that basic LLMs perform well in detecting PHI entities (e.g., names, dates, institutions and personal identifiers) in the Meddocan dataset, even without external knowledge.
Furthermore, it is observed that these models are able to identify certain medical entities in the Symptemist dataset, achieving an F1-score of 36.27. However, the LLMs perform less well in the Meddoprof and Meddoplace datasets, achieving F1-scores of 20.58 and 20.69, respectively. This lower performance is due to these datasets containing longer clinical documents, for which the LLMs have not been sufficiently trained.
Regarding RQ2, which asks whether an approach based on fine-tuning pre-trained encoder-only language models (such as BERT, RoBERTa, ALBERT, DistilBERT and XLM-RoBERTa) is superior to basic LLMs applying prompting techniques, Table 8 shows that fine-tuning encoder-only models remains the optimal choice. This is because these models are specifically trained on a given dataset, allowing their performance to be optimized. However, this strategy requires a different model to be trained for each dataset. In this case, four fine-tuned models would be required for the four evaluation datasets. Conversely, an advantage of LLMs is that they do not require specific training for each dataset. With a single LLM, it is possible to adapt the model for the NER task on different types of entities, making it more flexible in environments where insufficient data is available for individual fine-tuning.
Finally, with respect to RQ3, which asks whether adapting LLMs specifically for NER detection through fine-tuning would improve performance compared to prompting and fine-tuning encoder-only models, Table 8 allows us to analyse this. It is observed that fine-tuning LLMs specifically for NER can improve performance compared to prompting, as seen in the Meddoprof and Symptemist datasets. However, performance deteriorates in the in Meddocan and Meddoplace datasets due to the lack of a sufficiently large training set, which introduces more “noise” into the model instead of optimizing it. In this context, the term “noise” does not refer to annotation errors or deficiencies in the datasets, but rather to the weak and highly uneven supervision signal available during fine-tuning. Both Meddocan and Meddoplace contain a large number of fine-grained entity types, many of which appear only a few times in the training split. As a result, the model receives very limited and non-representative gradient updates for these rare classes, especially when applied to long clinical documents that yield few independent training instances. Although the models learn to produce valid BIO labels, the scarcity of examples prevents them from learning reliable boundaries between entity types, leading to systematic confusions and an almost total absence of correct span–type matches. Therefore, the poor performance observed in these datasets reflects the difficulty of learning highly granular and low-frequency entity inventories from limited supervision, rather than formatting issues or structural degradation of the models.
The comparison between prompt-based and fine-tuning approaches with generative LLMs is affected by computational constraints. Due to GPU memory limitations, the QLoRA fine-tuning experiments only included small and medium-sized LLMs, whereas larger models (e.g., those with 70B parameters) were exclusively evaluated under prompting settings. This asymmetry in model scale introduces a potential bias into the comparison and limits the validity of the conclusions drawn for RQ3. Consequently, any differences observed between prompting and fine-tuned LLMs should be interpreted with caution, as they may partly reflect model size rather than the inherent effectiveness of the adaptation strategy.
Furthermore, it is confirmed that fine-tuning LLMs does not outperform fine-tuning encoder-only models. These findings supports the idea that pre-trained encoder-only language models remain the most effective approach for NER in the medical domain.
It is important to note that only small- and medium-scale LLMs were fine-tuned using QLoRA due to hardware limitations, while large-scale models were only evaluated using the prompting approach. Consequently, this asymmetry in model scale may have negatively affected the absolute performance of the fine-tuned LLMs compared to both prompting and encoder-only models.
Although our evaluation primarily focuses on F1 scores, it is important to consider the substantial differences in inference cost and latency among the compared approaches. Prompting large generative LLMs (e.g., 70B parameters) requires considerably more memory, results in higher latency, and incurs higher computational costs—factors that limit their practicality in real clinical settings where responsiveness and efficiency are crucial. In contrast, fine-tuned encoder-only models are lightweight and offer fast, low-cost inference, making them more suitable for large-scale processing of clinical narratives and real-time integration into clinical workflows. These cost–latency advantages reinforce the practical value of encoder-based models beyond their superior F1 performance.
It is also worth noting that datasets such as Meddoprof contain considerably longer and more complex clinical documents, which pose specific challenges for generative LLMs. In these settings, LLMs tend to over-generate entity mentions, leading to a notable drop in precision, particularly for entity types that are semantically ambiguous or weakly delimited. Unlike encoder-only models, which benefit from explicit token-level supervision, LLMs rely on sequence generation and broader contextual cues, making them more prone to false positives in long clinical narratives. While chunking may limit long-range contextual information for encoder-only models, this effect does not appear to be the main performance bottleneck in Meddoprof, where encoder-only models achieve substantially higher F1-scores than LLM-based approaches.
6 Conclusions and Further Work
The widespread digitization of healthcare has generated vast amounts of clinical data, underscoring the need for effective methods for extracting medical information. This study compares LLMs using prompting strategies (ZSL, FSL and CoT) with fine-tuned encoder-only models for the NER task. Although prompting can produce acceptable results, performance is inconsistent. Fine-tuned encoder-only models, however, consistently outperform both prompted and fine-tuned LLMs across all datasets. Although fine-tuning LLMs has led to improvements in some cases, it has also led to training instability and degradation in some datasets and failed to outperform encoder-based models. One of the main advantages of LLMs is their ability to adapt to different domains without additional training, making them more flexible in scenarios with limited data. However, when sufficient labelled corpora are available, encoder-only models are still the best choice, as their specific training allows for better performance in medical entity detection.
A future area of research will be to explore instruct-tuning of LLMs for the NER task. The aim will be to assess whether this fine-tuning technique improves performance compared to untuned LLMs and the fine-tuning of encoder-only models. Additionally, we will examine whether self-learning and synthetic data generation techniques can reduce reliance on large annotated datasets and improve the applicability of these models in real clinical settings. Furthermore, future work will examine the impact of few-shot example variability, long-context architectures, and more efficient parameter-efficient tuning methods to better understand the trade-offs between accuracy, generalization and computational cost in medical NER.
Acknowledgement: Not applicable.
Funding Statement: This research was funded by the project “Data-driven drug repositioning applying graph neural networks (3DR-GNN)” (PID2021-122659OB-I00) and LaTe4PoliticES (PID2022-138099OB-I00) funded by MICIU/AEI/10.13039/501100011033 and the European Fund for Regional Development (ERDF)-a way of making Europe and by ELADAIS (https://eladais.org/), funded by the Spanish Ministry of Economic Affairs and Digital Transformation under UNICO I+D Cloud programme. Mr. Tomás Bernal-Beltrán is supported by University of Murcia through the predoctoral programme.
Author Contributions: The authors confirm contribution to the paper as follows: Conceptualization, Rafael Valencia-García and Ernestina Menasalvas-Ruíz; methodology, Alejandro Rodríguez-González; software, Ronghao Pan and Tomás Bernal-Beltrán; validation, Ronghao Pan, Tomás Bernal-Beltrán and Ernestina Menasalvas-Ruíz; formal analysis, Alejandro Rodríguez-González; investigation, Ronghao Pan; resources, Ernestina Menasalvas-Ruíz; data curation, Ronghao Pan and Tomás Bernal-Beltrán; writing—original draft preparation, Ronghao Pan; writing—review and editing, Alejandro Rodríguez-González and Tomás Bernal-Beltrán; visualization, Ernestina Menasalvas-Ruíz; supervision, Rafael Valencia-García; project administration, Rafael Valencia-García; funding acquisition, Rafael Valencia-García. All authors reviewed and approved the final version of the manuscript.
Availability of Data and Materials: Source code of the fine-tuning, zero-, few-shot and CoT evaluation approaches is available in https://github.com/NLP-UMUTeam/evaluating-spanish-medical-ner (accessed on 11 December 2025). No new data were created in this research. Therefore, it is necessary to request the evaluated dataset from the original authors.
Ethics Approval: Not applicable.
Conflicts of Interest: The authors declare no conflicts of interest.
Appendix A Base Prompt for the ZSL: In Listing A1, an example of a base prompt for the ZSL approach is shown.
Listing A1: ZSL prompt example As an expert in Named-Entity Recognition (NER) in Spanish for medical documents, your task is to accurately extract specified entities from the given text.
Extract the following medical entities from the given text and make sure that:
- Each entity is separated by a hash (#).
- Each entity type is separated by a line break.
- If no entity of a type is extracted, return “null” for that entity type.
Extract the following medical entities from the given text and ensure that each entity is separated by a hash (#) in the output:
{list_of_entities}
**Output format:**
Your response should strictly follow this format. Group the extracted entities based on the provided output format. Ensure entities are separated by a hash (#):
{output_format}
Apendix B Base Prompt for the FSL: In Listing A2, an example of a base prompt for the FSL approach is shown.
Listing A2: FSL prompt example As an expert in Named-Entity Recognition (NER) in Spanish for medical documents, your task is to accurately extract specified entities from the given text.
Extract the following medical entities from the given text and make sure that:
- Each entity is separated by a hash (#).
- Each entity type is separated by a line break.
- If no entity of a type is extracted, return “null” for that entity type.
Extract the following medical entities from the given text and ensure that each entity is separated by a hash (#) in the output:
{list_of_entities}
**Examples:**
{few_shot_examples}
**Output format:**
Your response should strictly follow this format. Group the extracted entities based on the provided output format. Ensure entities are separated by a hash (#):
{output_format}
Appendix C Base Prompt for the CoT: In Listing A3, an example of a base prompt for the CoT approach is shown.
Listing A3: CoT prompt example You are tasked with analyzing the given input text to extract specific pieces of information. Please follow a step-by-step reasoning process to identify and extract the relevant entities or details from the text. Ensure that you extract only the required entities and follow the defined output format exactly as specified below.
**Process:**
1. Understand the context: Begin by reading through the entire text to comprehend the subject and overall structure. Pay attention to headings, subheadings, and specific terminology that may indicate important information.
2. Identify key entities or details: As you analyze the text, look for specific items that match the categories or types of information you are required to extract.
3. Classify the information: Based on the type of entity or detail you’ve identified, categorize it correctly. Make sure you understand the specific definition of each category you’re working with.
4. Extract the information: Carefully extract the identified entities, ensuring the extracted text is complete and matches the intended category without including unnecessary details.
5. Organize the output: Group the extracted entities based on the provided output format.
Extract the following medical entities from the given text and make sure that:
- Each entity is separated by a hash (#).
- Each entity type is separated by a line break.
- If no entity of a type is extracted, return “null” for that entity type.
Extract the following medical entities from the following text:
{list_of_entities}
**Output format (Always use this format for your responses):**
Your response should strictly follow this format. Group the extracted entities based on the provided output format. Ensure entities are separated by a hash (#):
{output_format}
References
1. Clay B, Bergman HI, Salim S, Pergola G, Shalhoub J, Davies AH. Natural language processing techniques applied to the electronic health record in clinical research and practice-an introduction to methodologies. Comput Biol Med. 2025;188(13):109808. doi:10.1016/j.compbiomed.2025.109808. [Google Scholar] [PubMed] [CrossRef]
2. Zhou B, Yang G, Shi Z, Ma S. Natural language processing for smart healthcare. IEEE Rev Biomed Eng. 2022;17(4):4–18. doi:10.1109/rbme.2022.3210270. [Google Scholar] [PubMed] [CrossRef]
3. Sumner J, Wang Y, Tan SY, Chew EHH, Wenjun Yip A. Perspectives and experiences with large language models in health care: survey study. J Med Internet Res. 2025;27:e67383. doi:10.2196/67383. [Google Scholar] [PubMed] [CrossRef]
4. Bedi S, Liu Y, Orr-Ewing L, Dash D, Koyejo S, Callahan A, et al. A systematic review of testing and evaluation of healthcare applications of large language models (LLMs). medRxiv. 2024 [Google Scholar]
5. Fornasiere R, Brunello N, Scotti V, Carman M. Medical information extraction with large language models. In: Proceedings of the 7th International Conference on Natural Language and Speech Processing (ICNLSP 2024); 2024 Oct 19–20; Trento, Italy. p. 456–66. [Google Scholar]
6. Eguia H, Sánchez-Bocanegra CL, Vinciarelli F, Alvarez-Lopez F, Saigí-Rubió F. Clinical decision support and natural language processing in medicine: systematic literature review. J Med Internet Res. 2024;26(2):e55315. doi:10.2196/55315. [Google Scholar] [PubMed] [CrossRef]
7. Landolsi MY, Hlaoua L, Ben Romdhane L. Information extraction from electronic medical documents: state of the art and future research directions. Knowl Inf Syst. 2022;65(2):463–516. doi:10.1007/s10115-022-01779-1. [Google Scholar] [PubMed] [CrossRef]
8. Soomro PD, Kumar S, Banbhrani AAS, Shaikh AA, Raj H. Bio-NER: biomedical named entity recognition using rule-based and statistical learners. Int J Adv Comput Sci Appl. 2017;8(12):163–70. doi:10.14569/ijacsa.2017.081220. [Google Scholar] [CrossRef]
9. Wen C, Chen T, Jia X, Zhu J. Medical named entity recognition from un-labelled medical records based on pre-trained language models and domain dictionary. Data Intell. 2021;3(3):402–17. doi:10.1162/dint_a_00105. [Google Scholar] [CrossRef]
10. Xu K, Zhou Z, Hao T, Liu W. A bidirectional LSTM and conditional random fields approach to medical named entity recognition. In: International Conference on Advanced Intelligent Systems and Informatics. Cham, Switzerland: Springer; 2017. p. 355–65. [Google Scholar]
11. Sun C, Yang Z. Transfer learning in biomedical named entity recognition: an evaluation of BERT in the PharmaCoNER task. In: Proceedings of the 5th Workshop on BioNLP Open Shared Tasks; 2019 Nov 4; Hong Kong, China. p. 100–4. [Google Scholar]
12. Lima-López S, Farré-Maduell E, Gasco-Sánchez L, Rodríguez-Miret J, Krallinger M. Overview of SympTEMIST at BioCreative VIII: corpus, guidelines and evaluation of systems for the detection and normalization of symptoms, signs and findings from text. In: Proceedings of the BioCreative VIII Challenge and Workshop: Curation and Evaluation in the Era of Generative Models; 2023 Nov 12; New Orleans, LA, USA. [Google Scholar]
13. Miranda-Escalada A, Gascó L, Lima-López S, Farré-Maduell E, Estrada D, Nentidis A, et al. Overview of DisTEMIST at BioASQ: automatic detection and normalization of diseases from clinical texts: results, methods, evaluation and multilingual resources. CLEF Working Notes. 2022;3180:179–203. [Google Scholar]
14. Grattafiori A, Dubey A, Jauhri A, Pandey A, Kadian A, Al-Dahle A, et al. The llama 3 herd of models. arXiv:2407.21783. 2024. [Google Scholar]
15. Team G, Riviere M, Pathak S, Sessa PG, Hardin C, Bhupatiraju S, et al. Gemma 2: improving open language models at a practical size. arXiv:2408.00118. 2024. [Google Scholar]
16. Abdin M, Aneja J, Awadalla H, Awadallah A, Awan A, Bach N, et al. Phi-3 technical report: a highly capable language model locally on your phone. arXiv:2404.14219. 2024. [Google Scholar]
17. Singhal K, Azizi S, Tu T, Mahdavi SS, Wei J, Chung HW, et al. Large language models encode clinical knowledge. Nature. 2023;620(7972):172–80. doi:10.1038/s41586-023-06291-2. [Google Scholar] [PubMed] [CrossRef]
18. Agrawal M, Hegselmann S, Lang H, Kim Y, Sontag D. Large language models are few-shot clinical information extractors. arXiv:2205.12689. 2022. [Google Scholar]
19. Hu Y, Ameer I, Zuo X, Peng X, Zhou Y, Li Z, et al. Zero-shot clinical entity recognition using chatgpt. arXiv:2303.16416. 2023. [Google Scholar]
20. Zhang D, Xue X, Gao P, Jin Z, Hu M, Wu Y, et al. A survey of datasets in medicine for large language models. Intell Robot. 2024;4(4):457–78. doi:10.20517/ir.2024.27. [Google Scholar] [CrossRef]
21. Pagad NS, Pradeep N. Clinical named entity recognition methods: an overview. In: International Conference on Innovative Computing and Communications: Proceedings of ICICC 2021. Vol. 2. Cham, Switzerland: Springer; 2021. p. 151–65. [Google Scholar]
22. Wu Y, Jiang M, Xu J, Zhi D, Xu H. Clinical named entity recognition using deep learning models. AMIA Annu Symp Proc. 2018;2017:1812–9. doi:10.3233/978-1-61499-564-7-624. [Google Scholar] [CrossRef]
23. Perera N, Dehmer M, Emmert-Streib F. Named entity recognition and relation detection for biomedical information extraction. Front Cell Dev Biol. 2020;8:673. doi:10.3389/fcell.2020.00673. [Google Scholar] [PubMed] [CrossRef]
24. Liu S, Wang A, Xiu X, Zhong M, Wu S. Evaluating medical entity recognition in health care: entity model quantitative study. JMIR Med Inform. 2024;12:e59782. doi:10.2196/59782. [Google Scholar] [PubMed] [CrossRef]
25. Song B, Li F, Liu Y, Zeng X. Deep learning methods for biomedical named entity recognition: a survey and qualitative comparison. Brief Bioinform. 2021;22(6):bbab282. doi:10.1093/bib/bbab282. [Google Scholar] [PubMed] [CrossRef]
26. Tian S, Erdengasileng A, Yang X, Guo Y, Wu Y, Zhang J, et al. Transformer-based named entity recognition for parsing clinical trial eligibility criteria. In: Proceedings of the 12th ACM International Conference on Bioinformatics, Computational Biology, and Health Informatics. New York, NY, USA: ACM; 2021. p. 1–6. doi:10.1145/3459930.3469560. [Google Scholar] [PubMed] [CrossRef]
27. Sun C, Yang Z, Wang L, Zhang Y, Lin H, Wang J. Biomedical named entity recognition using BERT in the machine reading comprehension framework. J Biomed Inform. 2021;118:103799. doi:10.1016/j.jbi.2021.103799. [Google Scholar] [PubMed] [CrossRef]
28. Solarte-Pabón O, Montenegro O, García-Barragán A, Torrente M, Provencio M, Menasalvas E, et al. Transformers for extracting breast cancer information from Spanish clinical narratives. Artif Intell Med. 2023;143(4):102625. doi:10.1016/j.artmed.2023.102625. [Google Scholar] [PubMed] [CrossRef]
29. Spasic I, Nenadic G. Clinical text data in machine learning: systematic review. JMIR Med Inform. 2020;8(3):e17984. [Google Scholar] [PubMed]
30. Huang K, Altosaar J, Ranganath R. ClinicalBERT: modeling clinical notes and predicting hospital readmission. arXiv:1904.05342. 2019. [Google Scholar]
31. Dodge J, Ilharco G, Schwartz R, Farhadi A, Hajishirzi H, Smith N. Fine-tuning pretrained language models: weight initializations, data orders, and early stopping. arXiv:2002.06305. 2020. [Google Scholar]
32. Monajatipoor M, Yang J, Stremmel J, Emami M, Mohaghegh F, Rouhsedaghat M, et al. LLMS in biomedicine: a study on clinical named entity recognition. arXiv:2404.07376. 2024. [Google Scholar]
33. Névéol A, Dalianis H, Velupillai S, Savova G, Zweigenbaum P. Clinical natural language processing in languages other than English: opportunities and challenges. J Biomed Semantics. 2018;9(1):12. doi:10.1186/s13326-018-0179-8. [Google Scholar] [PubMed] [CrossRef]
34. Hu Y, Chen Q, Du J, Peng X, Keloth VK, Zuo X, et al. Improving large language models for clinical named entity recognition via prompt engineering. J American Med Inform Associat. 2024;31(9):1812–20. doi:10.1093/jamia/ocad259. [Google Scholar] [PubMed] [CrossRef]
35. Radford A, Narasimhan K. Improving language understanding by generative pre-training. 2018 [cited 2026 Feb 2]. Available from: https://api.semanticscholar.org/CorpusID:49313245. [Google Scholar]
36. Achiam J, Adler S, Agarwal S, Ahmad L, Akkaya I, Aleman FL, et al. Gpt-4 technical report. arXiv:2303.08774. 2023. [Google Scholar]
37. Tang Y, Xiao Z, Li X, Fang Q, Zhang Q, Fong DYT, et al. Large language model in medical information extraction from titles and abstracts with prompt engineering strategies: a comparative study of GPT-3.5 and GPT-4. MedRxiv. 2025. doi:10.1101/2024.03.20.24304572. [Google Scholar] [CrossRef]
38. Keloth VK, Hu Y, Xie Q, Peng X, Wang Y, Zheng A, et al. Advancing entity recognition in biomedicine via instruction tuning of large language models. Bioinformatics. 2024;40(4):btae163. doi:10.1093/bioinformatics/btae163. [Google Scholar] [PubMed] [CrossRef]
39. Marimon M, Gonzalez-Agirre A, Intxaurrondo A, Rodriguez H, Martin JL, Villegas M, et al. Automatic de-identification of medical texts in Spanish: the MEDDOCAN track, corpus, guidelines, methods and evaluation of results. In: Proceedings of the Iberian Languages Evaluation Forum (IberLEF 2019); 2019 Sep 24; Bilbao, Spain. p. 618–38. [Google Scholar]
40. Lima-López S, Farré-Maduell E, Miranda-Escalada A, Brivá-Iglesias V, Krallinger M. NLP applied to occupational health: MEDDOPROF shared task at IberLEF, 2021 on automatic recognition, classification and normalization of professions and occupations from medical texts. Procesamiento Del Lenguaje Natural. 2021;67:243–56. [Google Scholar]
41. Lima-López S, Farré-Maduell E, Briva-Iglesias V, Gasco-Sanchez L, Krallinger M. MEDDOPLACE shared task overview: recognition, normalization and classification of locations and patient movement in clinical texts. Procesamiento Del Lenguaje Natural. 2023;71:301–11. [Google Scholar]
42. Meta AI. Llama 3.2: revolutionizing edge AI and vision with open, customizable models. 2024 [cited 2026 Feb 2]. Available from: https://ai.meta.com/blog/llama-3-2-connect-2024-vision-edge-mobile-devices. [Google Scholar]
43. Jiang AQ, Sablayrolles A, Roux A, Mensch A, Savary B, Bamford C, et al. Mixtral of experts. arXiv:2401.04088. 2024. [Google Scholar]
44. Jiang AQ, Sablayrolles A, Mensch A, Bamford C, Chaplot DS, de las Casas D, et al. Mistral 7B. arXiv:2310.06825. 2023. [Google Scholar]
45. Yang QA, Yang B, Zhang B, Hui B, Zheng B, Yu B, et al. Qwen2.5 technical report. arXiv:2412.15115. 2024. [Google Scholar]
46. Cañete J, Chaperon G, Fuentes R, Ho JH, Kang H, Pérez J. Spanish pre-trained BERT model and evaluation data. arXiv:2308.02976. 2023. [Google Scholar]
47. Gutiérrez-Fandiño A, Armengol-Estapé J, Pàmies M, Llop-Palao J, Silveira-Ocampo J, Carrino CP, et al. MarIA: spanish language models. Procesamiento Del Lenguaje Natural. 2022;68:39–60. [Google Scholar]
48. de la Rosa J, Ponferrada EG, Villegas P, de Prado Salas PG, Romero M, Grandury M. BERTIN: efficient pre-training of a Spanish language model using perplexity sampling. Procesamiento Del Lenguaje Natural. 2022;68:13–23. [Google Scholar]
49. Cañete J, Donoso S, Bravo-Marquez F, Carvallo A, Araujo V. ALBETO and DistilBETO: lightweight spanish language models. In: Proceedings of the Thirteenth Language Resources and Evaluation Conference; 2022 Jun 20–25; Marseille, France. p. 4291–8. [Google Scholar]
50. Devlin J, Chang MW, Lee K, Toutanova K. BERT: pre-training of deep bidirectional transformers for language understanding. In: Proceedings of the 2019 Conference of the North American Chapter of the Association for Computational Linguistics: Human Language Technologies. Vol. 1. Stroudsburg, PA, USA: ACL; 2019. p. 4171–86. [Google Scholar]
Cite This Article
 Copyright © 2026 The Author(s). Published by Tech Science Press.
Copyright © 2026 The Author(s). Published by Tech Science Press.This work is licensed under a Creative Commons Attribution 4.0 International License , which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.


 Submit a Paper
Submit a Paper Propose a Special lssue
Propose a Special lssue View Full Text
View Full Text Download PDF
Download PDF Downloads
Downloads
 Citation Tools
Citation Tools