 Open Access
Open Access
ARTICLE
Model Agnostic Meta Learning Ensemble Based Prediction of Motor Imagery Tasks Using EEG Signals
1 Department of Electrical and Biomedical Engineering, Khwaja Fareed University of Engineering and Information Technology, Rahim Yar Khan, Pakistan
2 Department of Electrical Engineering, Imam Mohammad Ibn Saud Islamic University (IMSIU), Riyadh, Saudi Arabia
3 Department of Computer Science, Bahria School Engineering and Applied Sciences (BSEAS), Bahria University Islamabad, Islamabad, Pakistan
* Corresponding Author: Muhammad Irfan. Email:
(This article belongs to the Special Issue: Machine Learning and Deep Learning-Based Pattern Recognition)
Computer Modeling in Engineering & Sciences 2026, 146(2), 36 https://doi.org/10.32604/cmes.2026.076332
Received 18 November 2025; Accepted 14 January 2026; Issue published 26 February 2026
Abstract
Automated detection of Motor Imagery (MI) tasks is extremely useful for prosthetic arms and legs of stroke patients for their rehabilitation. Prediction of MI tasks can be performed with the help of Electroencephalogram (EEG) signals recorded by placing electrodes on the scalp of subjects; however, accurate prediction of MI tasks remains a challenge due to noise that is incurred during the EEG signal recording process, the extraction of a feature vector with high interclass variance, and accurate classification. The proposed method consists of preprocessing, feature extraction, and classification. First, EEG signals are denoised using a bandpass filter followed by Independent Component Analysis (ICA). Multiple channels are combined to form a single surrogate channel. Short Time Fourier Transform (STFT) is then applied to convert time domain EEG signals into the frequency domain. Handcrafted and automated features are extracted from EEG signals and then concatenated to form a single feature vector. We propose a customized two-dimensional Convolutional Neural Network (CNN) for automated feature extraction with high interclass variance. Feature selection is performed using Particle Swarm Optimization (PSO) to obtain optimal features. The final feature vector is passed to three different classifiers: Support Vector Machine (SVM), Random Forest (RF), and Long Short-Term Memory (LSTM). The final decision is made using the Model-Agnostic Meta Learning (MAML). The Proposed method has been tested on two datasets, including PhysioNet and BCI Competition IV-2a, and it achieved better results in terms of accuracy and F1 score than existing state-of-the-art methods. The proposed framework achieved an accuracy and F1 score of 96% on the PhysioNet dataset and 95.5% on the BCI Competition IV, dataset 2a. We also present SHapley Additive exPlanations (SHAP) and Gradient-weighted Class Activation Mapping (Grad-CAM) explainable techniques to enhance model interpretability in a clinical setting.Keywords
For over a billion individuals worldwide living with disabilities due to stroke, Brain Computer Interfaces (BCIs) offer a groundbreaking means of restoring interaction and independence by translating brain activity directly into device control [1]. BCIs monitor and analyze brain signals, converting them into commands for output devices to perform specified tasks for individuals with motor impairment [2]. The quality of life of patients affected by stroke can be improved. It facilitates interaction with the external environment by interpreting brain activity, offering valuable insights into the relationship between an individual’s motor movements and their brain’s electrical signals [3]. Brain–computer interfaces (BCIs) [4] can be categorized into different paradigms based on the elicitation mechanism, including Motor Imagery (MI), Slow Cortical Potentials (SCP), P300 event-related potentials [5], and steady-state visual evoked potentials (SSVEP) [6]. Among these, MI-based BCIs rely on self-generated neural activity and are particularly attractive for rehabilitation applications due to their intuitive and stimulus-independent nature. MI-based BCIs function by interpreting the user’s intended actions through their brain activity, which is commonly captured using Electroencephalogram (EEG) signals [7]. Multiple electrodes are placed on the scalp of the subjects to record EEG signals using the 10-10 and 10-20 systems [8]. MI involves the mental simulation or rehearsal of physical movements, enabling the transformation of motor intent into control signals and facilitating the detection of real-time motor imagery conditions, such as imagined movements of the left and right hands [9].
Researchers have proposed multiple machine/deep learning algorithms to predict MI tasks. All these algorithms consist of common steps, including EEG data recording, preprocessing of these signals for noise removal, followed by feature extraction/selection and classification. In the preprocessing of EEG signals, various methods such as bandpass filter [10], Finite Impulse Response (FIR) filter [11], Notch filter [12], Butterworth filter [13], and Independent Component Analysis (ICA) [14] have been used to remove noise. These signals are susceptible to multiple sources of noise, including power line interference at 50–60 Hz [15], baseline noise caused by electrode interference [16], and artifacts from physiological activities such as eye movements and cardiac pulsations [17]. Alazrai et al. [18] utilized a bandpass filter (0.5–32.5 Hz) to improve the Signal to Noise Ratio (SNR), followed by Automated Artifact Removal (AAR) and downsampling to 256 Hz. To eliminate power line interference, Barmpas et al. [3] implemented notch filtering at 60 Hz and its harmonics, with an additional filter at 460 Hz to address unidentified artifacts. Similarly, Tiwari et al. [19] applied a low-pass Butterworth filter
After preprocessing, feature extraction is performed. Multiple feature extraction methods have been proposed by researchers for the automated classification of MI tasks. Feature extraction reduces the dimensionality of EEG signals by extracting useful characteristics from preprocessed signals [22]. Feature extraction methods can be categorized into two types: handcrafted and automated features. Handcrafted features are further classified into time-domain and frequency-domain features, which have demonstrated good sensitivity and specificity. To extract discriminative features from EEG signals, traditional signal processing methods, including Fourier transforms, DWT, and CSP [23], have been used by researchers. Chen et al. [23] utilized CSP as one of the feature extraction techniques in their study. Lazcano-Herrera et al. [24] extracted five features, including Power Spectral Density (PSD), Spectral Entropy, and Shannon Entropy, from time-frequency representations generated using DWT and spectrograms. Principal Component Analysis (PCA) and Deep Belief Networks (DBN) [25] have also been applied by researchers. Musallam et al. [26] proposed CNNs for feature extraction. Sun et al. [27] applied Short-Time Fourier Transform (STFT) to extract time frequency features from the EEG signals within certain frequency bands (6–13 Hz, 17–30 Hz).
Recent developments in Deep Learning (DL) have resulted in significant improvements over traditional EEG analysis methods. Some of the models have gained increasing attention (CNNs [20], RNNs [24], LSTM [24], autoencoder [21]) due to their ability to recognize complex patterns and tackling the variability of different classes in data. Figures are presented by Zhang et al. [28] presented a Graph-based Convolutional Recurrent Attention Model (GCRAM) to better capture individual differences in motor imagery EEG signals. Building upon this work, Sun et al. [10] then came up with a Time-Frequency Feature Extraction Method (TFEM) that extracts features directly from time domain EEG recordings using CNN. Hwaidi et al. [21] also explored improving input representations by combining a deep autoencoder with a CNN, designing an architecture with five feature extraction layers and four max-pooling layers for dimensionality reduction.
Predicting motor imagery tasks requires accurate classification after feature extraction from EEG signals [29]. Alazrai et al. [18] utilized (SVM) for EEG signal classification. Sun et al. [10] proposed EEG-ARNN, a novel hybrid deep framework integrating convolutional and graph convolutional networks. Geng et al. [30] evaluated four classifiers: Bagging Tree (BT), SVM, Bayesian Linear Discriminant Analysis (BLDA), and Linear Discriminant Analysis (LDA) using within-subject residual one cross-validation. Taran et al. [31] proposed SVM for the classification of right-foot and right-hand MI tasks. Classification is performed after feature selection using Machine Learning (ML) classifiers and DL methods. SVM [30], Random Forest [32],
Current motor imagery prediction methods often suffer from limitations, including a lack of comprehensive inter-subject evaluation, sensitivity to class imbalance, insufficient noise reduction, and a lack of comparative analysis regarding DL classifier parameter optimization. Addressing these issues, such as ensuring thorough evaluation across all subjects, mitigating the impact of class imbalance, implementing robust noise removal techniques, and optimizing classifier parameters, is crucial for improving motor imagery prediction’s sensitivity, specificity, and accuracy. In this research, we propose a novel method for the classification of MI tasks using EEG signals. The proposed method consists of three steps, i.e., preprocessing, feature extraction, and classification. EEG signals are first preprocessed for noise removal and combining of multichannel EEG signals into a single surrogate channel using the (CSP) filter. A Butterworth bandpass filter followed by ICA is used for noise removal. We propose a customized 2D-CNN architecture for automated feature extraction. Both handcrafted and automated features are concatenated to form a single feature vector, and (PSO) is applied to get an optimal feature set. The final feature vector is passed to three classifiers, including RF, SVM, and LSTM, and the final decision is made using (MAML) as an ensemble classifier. The proposed model provides superior performance in terms of both accuracy and F1 score on two publicly available benchmark datasets. The main contributions of this work are summarized as follows:
• A customized 2D-CNN architecture designed to effectively learn discriminative time frequency representations from STFT-based EEG signals for multi-class motor imagery classification.
• A hybrid feature fusion framework that combines handcrafted temporal–spectral features with deep CNN features, followed by PSO to select an optimal and compact feature subset.
• A robust ensemble classification strategy based on MAML that integrates heterogeneous classifiers (SVM, RF, and LSTM) to improve generalization across subjects.
• An explainable artificial intelligence (XAI) analysis using Grad-CAM and SHAP to provide insight into both deep and handcrafted feature contributions, enhancing the interpretability of the proposed framework.
The proposed approach for MI EEG classification consists of three major components: (1) Preprocessing to denoise the raw EEG signals, (2) Feature Extraction (including both hand-crafted feature calculation and deep feature learning via a CNN), and (3) Classification using MAML-based ensemble classifier. The general flow of the methodology is shown in Fig. 1. The raw multichannel EEG signals are first filtered and cleansed to eliminate undesired artifacts. This is followed by CSP and STFT to prepare the data in order to extract features. A set of hand-crafted features is computed from the denoised signals, and in parallel, a custom 2D-CNN extracts deep features from the STFT representation. These sets of features are then combined into one feature set. PSO is thereafter utilized to identify an optimal subset of the fused features that will provide the maximum discriminative power. Lastly, the features selected are passed into three classifiers (SVM, RF, LSTM), and MAML is used as an ensemble classifier for the final prediction of the MI task. We describe each step of the pipeline in the subsections below.
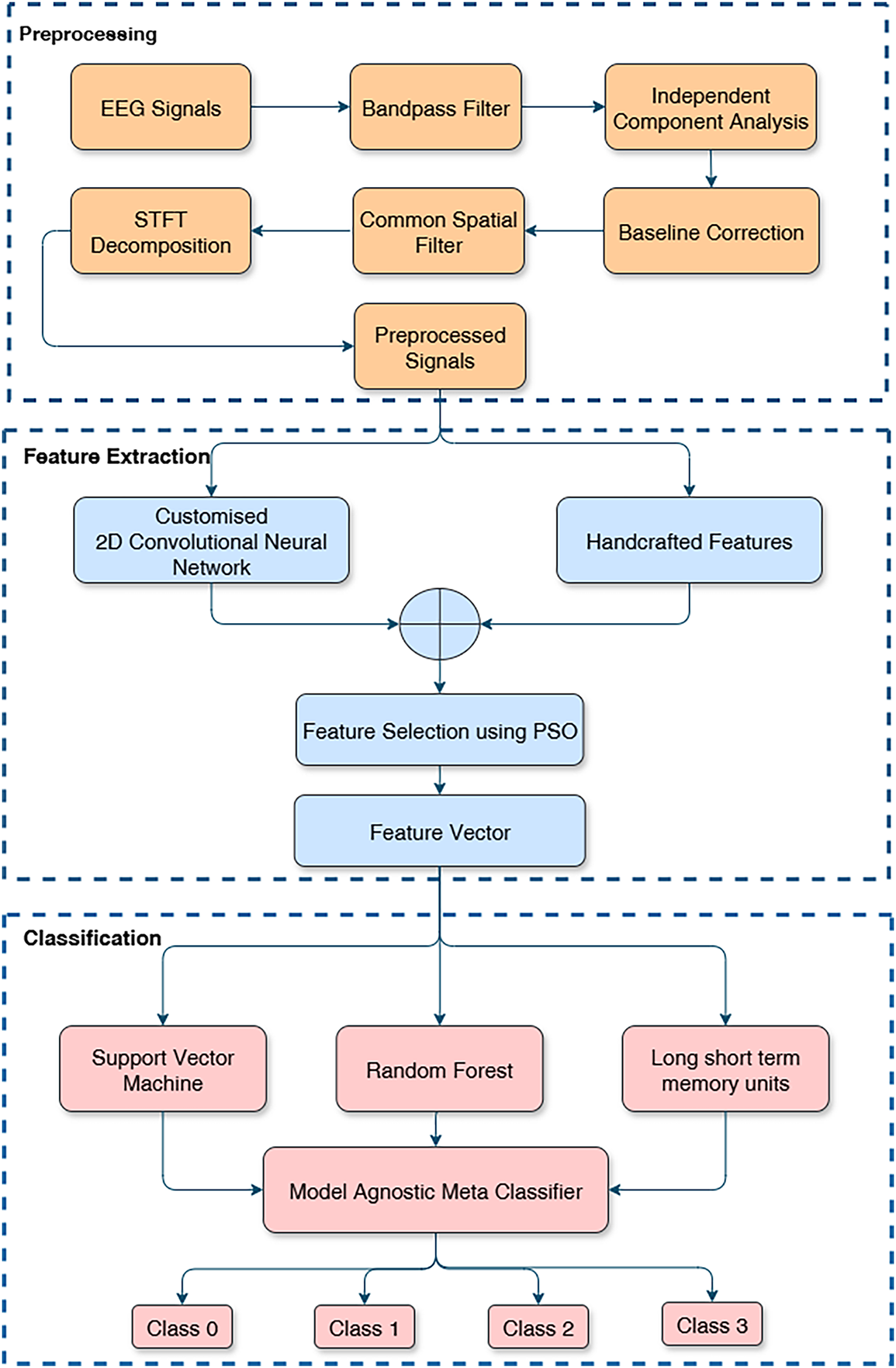
Figure 1: Flow diagram of the proposed methodology.
We design a preprocessing pipeline to reduce different types of noise in the EEG signals without loss of features useful in distinguishing MI. The digital band-pass filter (0.5–50 Hz) is used in the first step to filter each channel to draw the frequency components that are likely to contain MI information and eliminate drifts and high-frequency noise. This band covers significant EEG activity in the
where
In order to separate the ocular, muscular, and cardiac artifacts from the EEG recordings, ICA is used to distinguish the signals into independent components. Components related to the artifacts corresponding to eye movements or muscle activity are identified and equated to zero, and hence their effect is removed. The EEG signals are then reconstructed from the remaining components, resulting in a cleaner dataset.
In order to standardize the data and eliminate low-frequency drifts, baseline correction is carried out by deducting the mean signal value for each trial or epoch from a pre-stimulus baseline period. This step helps reduce variations that are not related to the motor imagery task.
Baseline correction is then applied to normalize the signal:
To further amplify the motor imagery signals relevant to the task and suppress noise for improved class distinction (e.g., left-hand vs. right-hand movements), we apply a CSP filter to the preprocessed EEG signals. CSP enhances class separability by projecting the multichannel EEG into spatial filters that maximize variance differences between classes:
where
where
As illustrated in Fig. 2, the original Channel 1 signal (yellow) exhibits elevated power at very low (<0.5 Hz) and high-frequency bands (>50 Hz), indicating residual drifts and noise. After ICA-based denoising (orange), the PSD tightly conforms to the 0.5–50 Hz band while power outside this range is markedly attenuated, confirming the effectiveness of our cleaning pipeline.
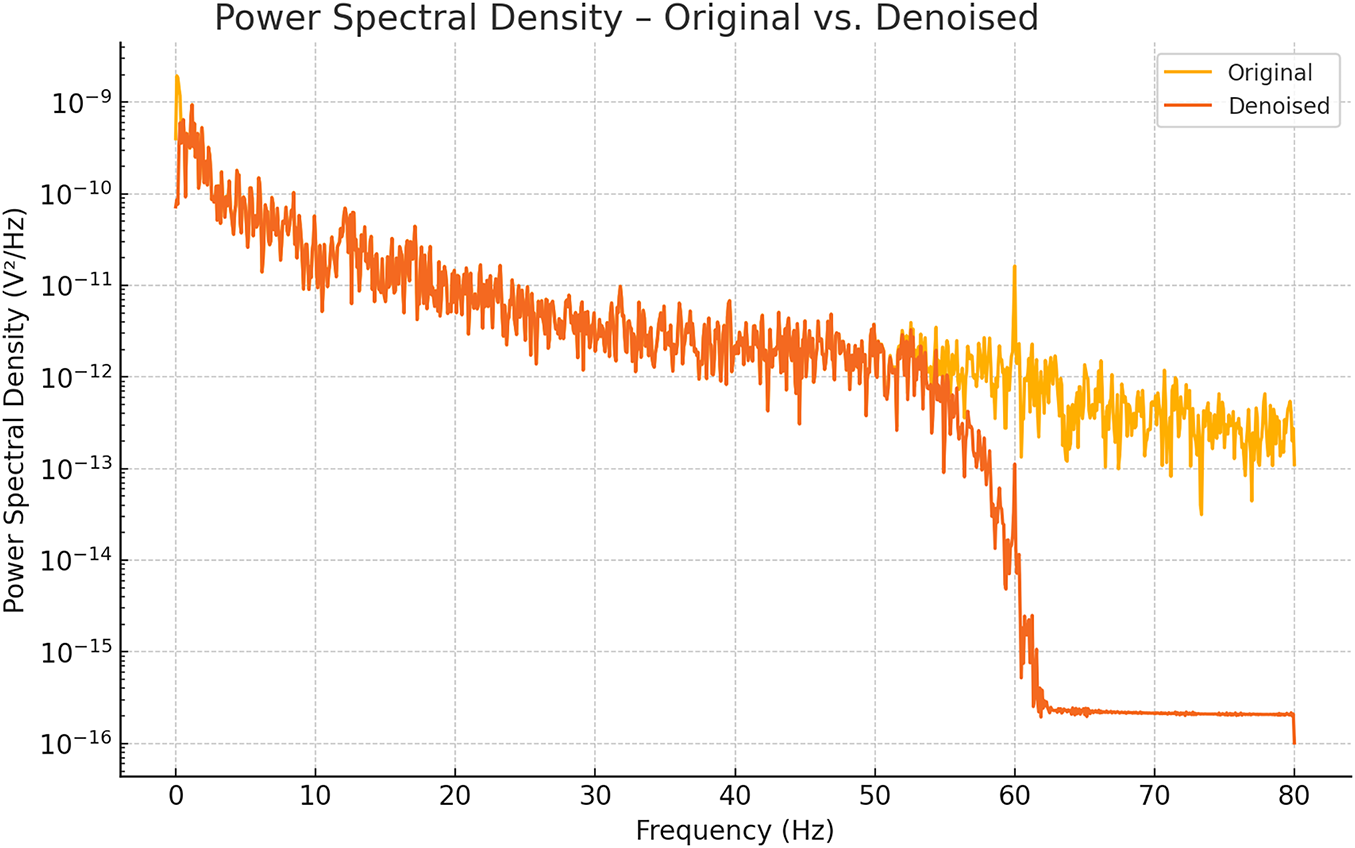
Figure 2: PSD plot after denoising of EEG signal.
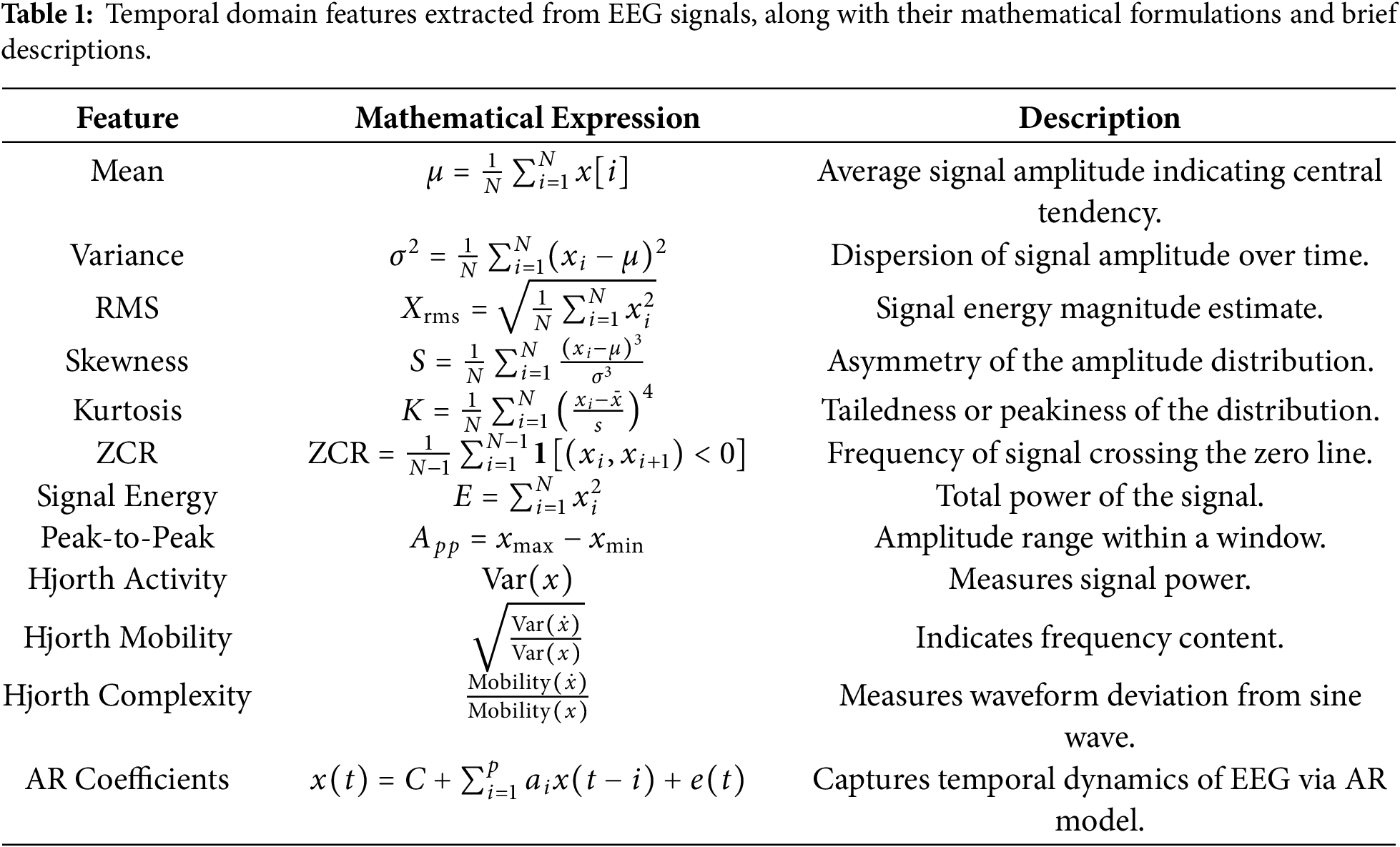
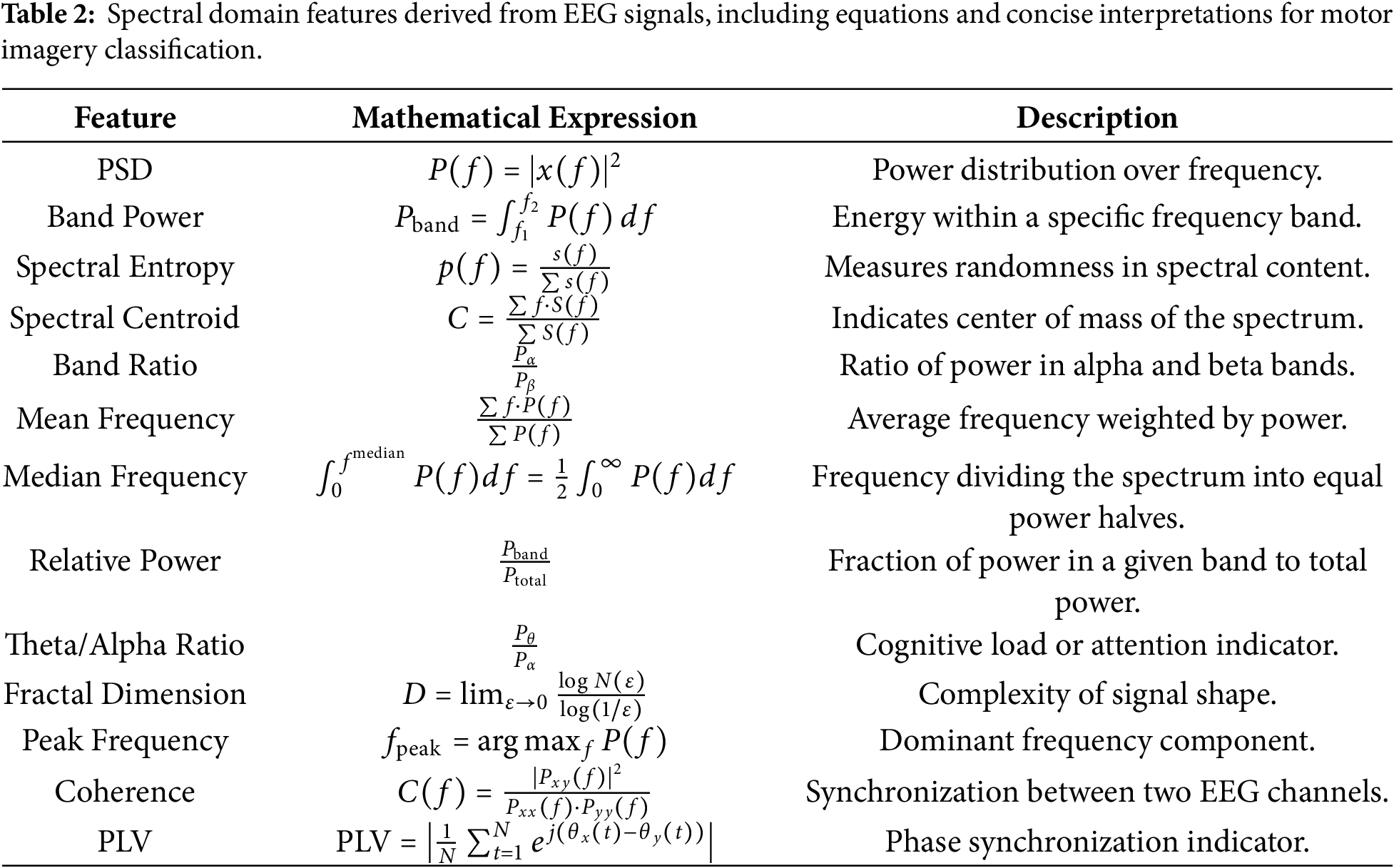
To capture time-domain and frequency-domain aspects pertinent to motor imagery tasks, we meticulously built a set of features from the EEG signals. Temporal characteristics extracted from the EEG signals include mean, variance, RMS, skewness, kurtosis, ZCR, energy, peak-to-peak amplitude, Hjorth metrics (activity, mobility, complexity), and AR coefficients. These features effectively quantify the signal’s amplitude dynamics, statistical distribution, and temporal dependencies. In addition to temporal descriptors, we compute several spectral features such as power spectral density (PSD), band power, spectral entropy, spectral centroid, spectral band ratios, mean frequency, median frequency, relative power, theta/alpha ratio (TAR), fractal dimension, peak frequency, coherence, and phase locking value (PLV). These spectral features provide insights into the energy distribution and frequency-specific behavior of EEG signals. A detailed summary of the extracted features, along with their corresponding mathematical expressions and descriptions, is provided in Tables 1 and 2.

The STFT is computed using a 128-sample (0.8 s at 160 Hz) Hamming window with 50% overlap. This window length is selected as a compromise between time and frequency resolution for non-stationary EEG signals. Specifically, it provides a frequency resolution of approximately 1.25 Hz, which is sufficient to discriminate motor imagery–related
We propose a customized architecture of 2D-CNN aimed at effective extraction and processing of spatial information from the image data. The model consists of several convolutional, normalization, pooling, and fully connected layers in such a way to optimize feature extraction and classification performance, as shown in Fig. 3. The proposed 2D-CNN treats the STFT time–frequency representation as a single-channel 2D input, enabling joint learning of spectral and temporal patterns across EEG channels. Small
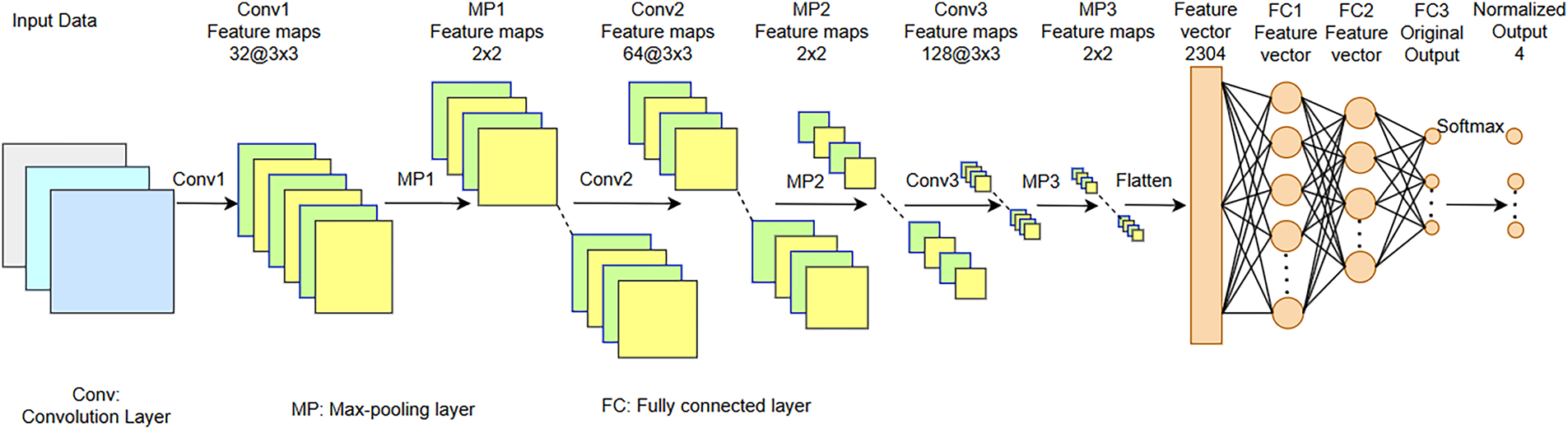
Figure 3: Architecture of customized 2D-CNN proposed for MI classification tasks.
The second convolutional block enhances the network’s capacity with a Conv2D layer that uses 64 filters (
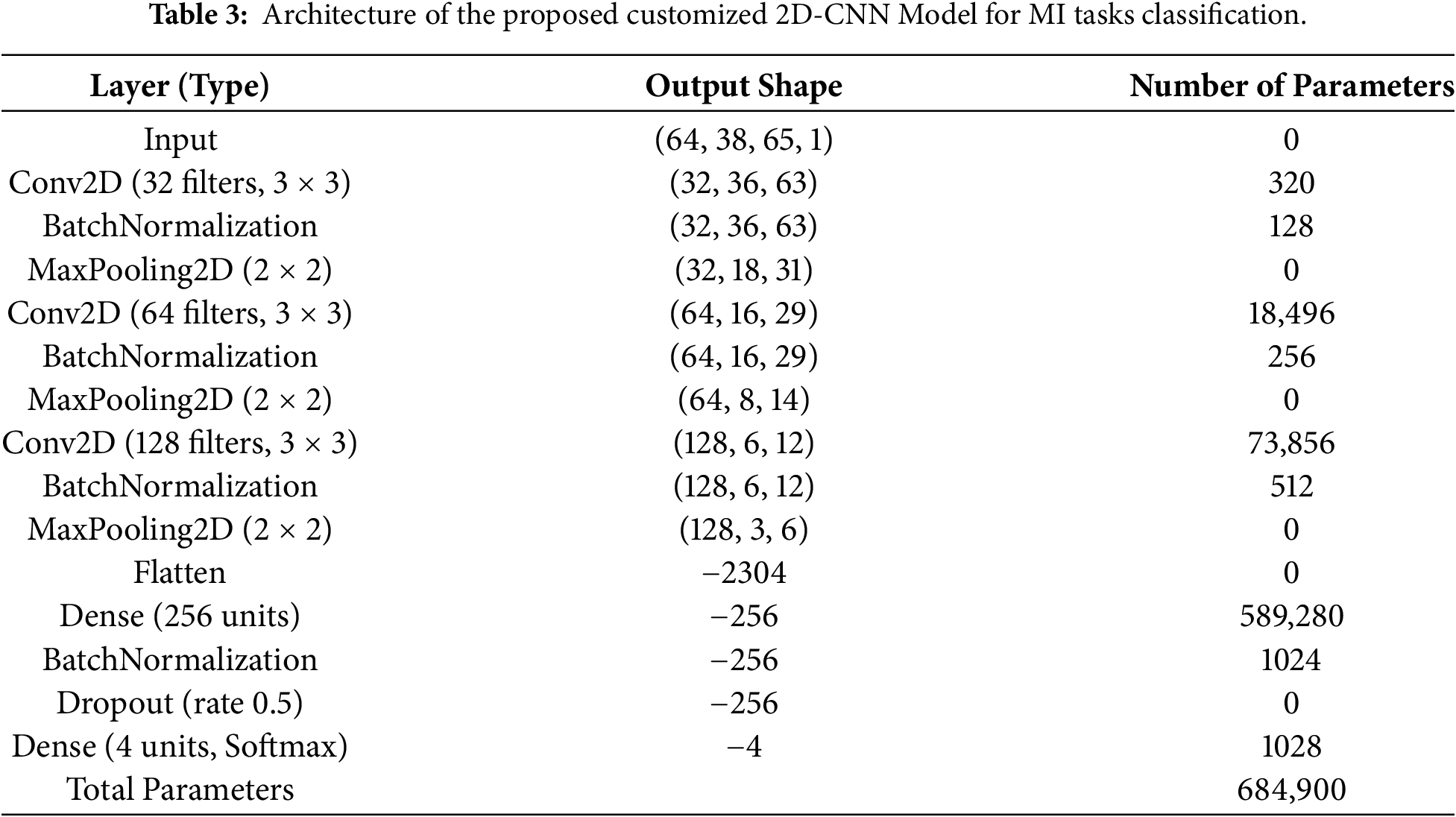
The proposed 2D-CNN is trained using the Adam optimizer with a learning rate of 0.001 and categorical cross-entropy loss. Training is performed for 100 epochs with a batch size of 32. Batch normalization and dropout (0.5) are used to improve generalization, and the model with the best validation performance is selected. To create a single feature vector, a concatenation of a hand-crafted feature and an automatically retrieved one from a specially built 2D-CNN is used. To select a suitable feature subset, PSO is then employed. It aims at optimizing the feature subset by maximizing the fitness function. This fitness function evaluates how well a classifier (for example, a Support Vector Machine or condensed DL model) does using a feature subset represented by a particle. To optimize classification, PSO selects the most discriminative subset of features:
Velocity Update:
Position Update:
Fitness Function:
In PSO-based feature selection, a swarm size of 30 particles is used with inertia weight w = 0.7, acceleration coefficients
Three classifiers, i.e, LSTM, SVM, and RF, are subsequently provided with the feature vector selected by PSO. MAML is used as an ensemble classifier that combines the outputs of these classifiers to produce the final decision of classification. RF Classifier is a technique used for ensemble learning, which trains multiple decision trees and provides the result in a classification problem based on the mode of classes or with a mean prediction in case of regression problems. By introducing randomization and averaging each tree’s predictions, RF reduces the risk of overfitting that the usual decision trees have. In addition to this, the averaging procedure helps increase the generalization capability of the model by reducing the variance. SVMs seek to determine an optimum separating hyperplane, which maximizes the margin between the classes, which allows high-dimensional feature space performance. Besides, SVMs process non-linearly separable data by using kernel functions, by definition mapping input features to a space where linear separation is possible. By using the MAML, the ensemble approach combines the predictions given by the RF, SVM, and LSTM classifiers. In this way, each classifier votes for the class it forecasts, and the most voted-for class determines the final prediction. By combining their complementary advantages, RF’s ensemble averaging for strong decision boundaries, SVM’s margin maximization for distinct class separation, and LSTMs perform exceptionally well in MI task prediction by successfully capturing the important temporal dependencies in EEG signals over long time periods, improving classification accuracy. MAML is then applied as an ensemble classifier for the final prediction with the base classifier as LSTM. Algorithm 1 describes the ensemble classification of the proposed method.
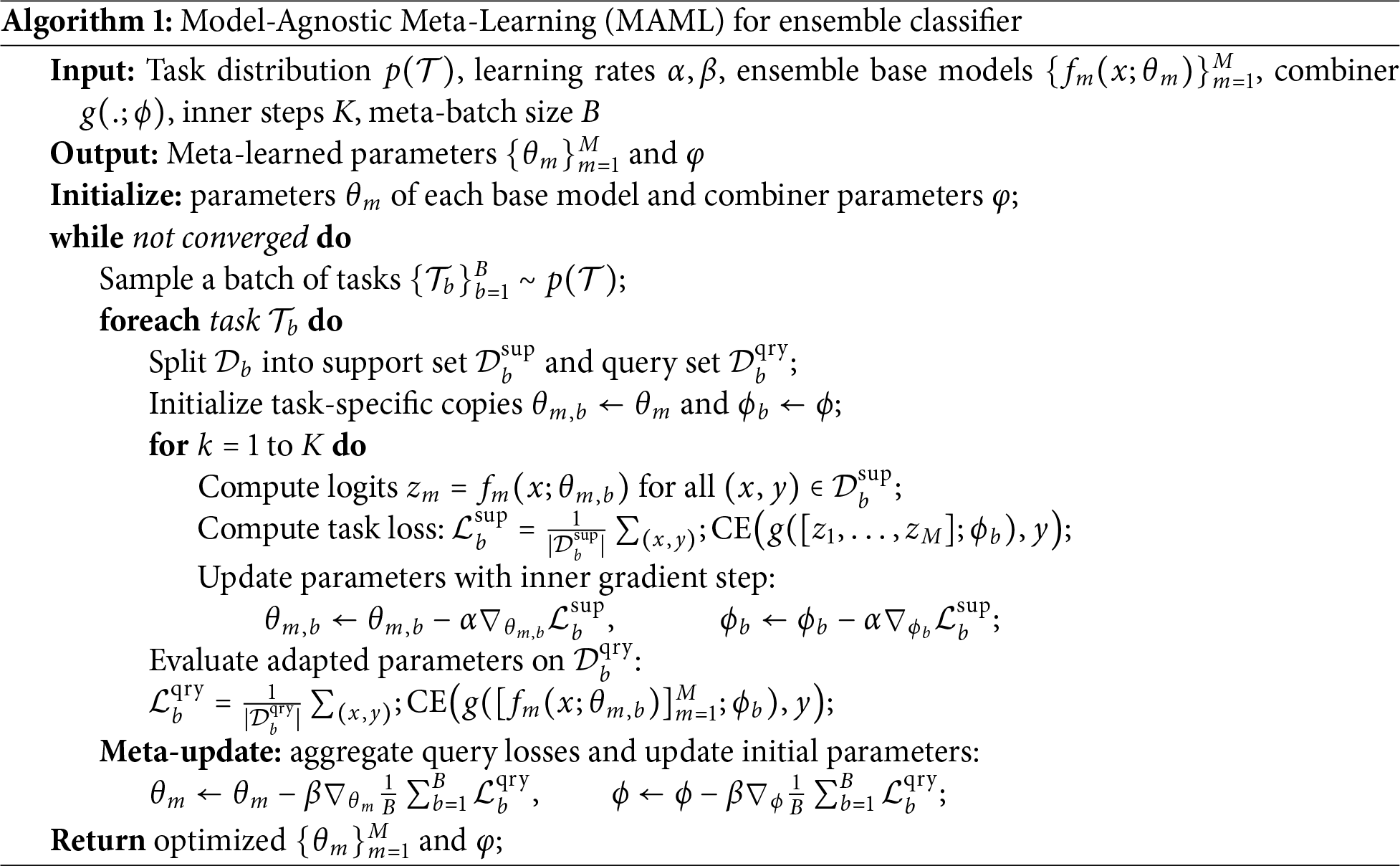

Overfitting during meta-learning is mitigated by applying MAML at the decision-level rather than directly on raw features, by restricting the number of inner-loop adaptation steps, and by leveraging the diversity of heterogeneous base classifiers (SVM, RF, and LSTM). Furthermore, PSO-based feature selection and regularization strategies within individual classifiers (e.g., dropout in LSTM) contribute to improved generalization across subjects. The ablation results demonstrate the incremental benefit of each processing stage. Basic bandpass filtering improves performance by suppressing out-of-band noise, while baseline correction further stabilizes the signal by reducing low-frequency drifts. Incorporating ICA leads to a noticeable gain by removing ocular and muscular artifacts. The transition from handcrafted features to STFT-based time–frequency representations with CNN learning significantly enhances class discrimination, highlighting the advantage of automated feature learning. Feature fusion further boosts performance by combining complementary handcrafted and deep features, and PSO-based feature selection refines this representation by retaining only the most discriminative components. Finally, the MAML-based ensemble consistently yields the highest accuracy and F1 score by leveraging the complementary strengths of heterogeneous classifiers. The complete dataset is divided into 80% for training and validation and 20% for testing at the trial level.
An open-access resource for motor imagery-enabled BCI research, the PhysioNet and BCI competition IV, Dataset 2a is used in this research. The PhysioNet dataset consists of EEG recordings from 109 subjects, recorded using a 64-channel EEG system, according to the 10-10 electrode placement system, with certain exclusions. The dataset has four motor tasks. Movements of the left hand, right hand, both hands, and both feet movements performed under both motor execution and motor imagery conditions. EEG signals are acquired at a 160 Hz sampling rate and bandpass filtered with 0.5–100 Hz. The large subject pool and uniform protocol of the dataset make it useful for the BCI algorithm development. To evaluate generalization performance, the proposed framework is assessed in a subject-independent setting on the PhysioNet dataset, where training and is performed on data from disjoint subjects, ensuring that no subject overlap occurred between the two sets.
Fig. 4 shows the confusion matrices of first six experiments whereas Fig. 5 shows the next five experiment results on PhysioNet dataset. Similarly, Table 4 shows the relative performance of different methods. First, a baseline experiment is carried out, whereby the handcrafted traits are revealed with the assistance of a Support Vector Machine from the raw EEG data. Overall Accuracy, as delivered by this strategy, is moderate, i.e., 71%, F1 Score (71%), and task-specific accuracy scores ranging from 68% (T3) to 70% (T1). Bandpass filtering improved the performance much better, increasing the Overall Accuracy and F1 Score to 75%. This speaks to the fact that even basic filtering helps to isolate major frequency bands, stressing the importance of carrying it out for MI works. Once baseline correction is implemented and exerted, the findings are even more improved to produce an F1 Score of 78% and an Overall Accuracy of 78%. This suggests that drifts and low-frequency noise are mitigated when baseline correction is used.
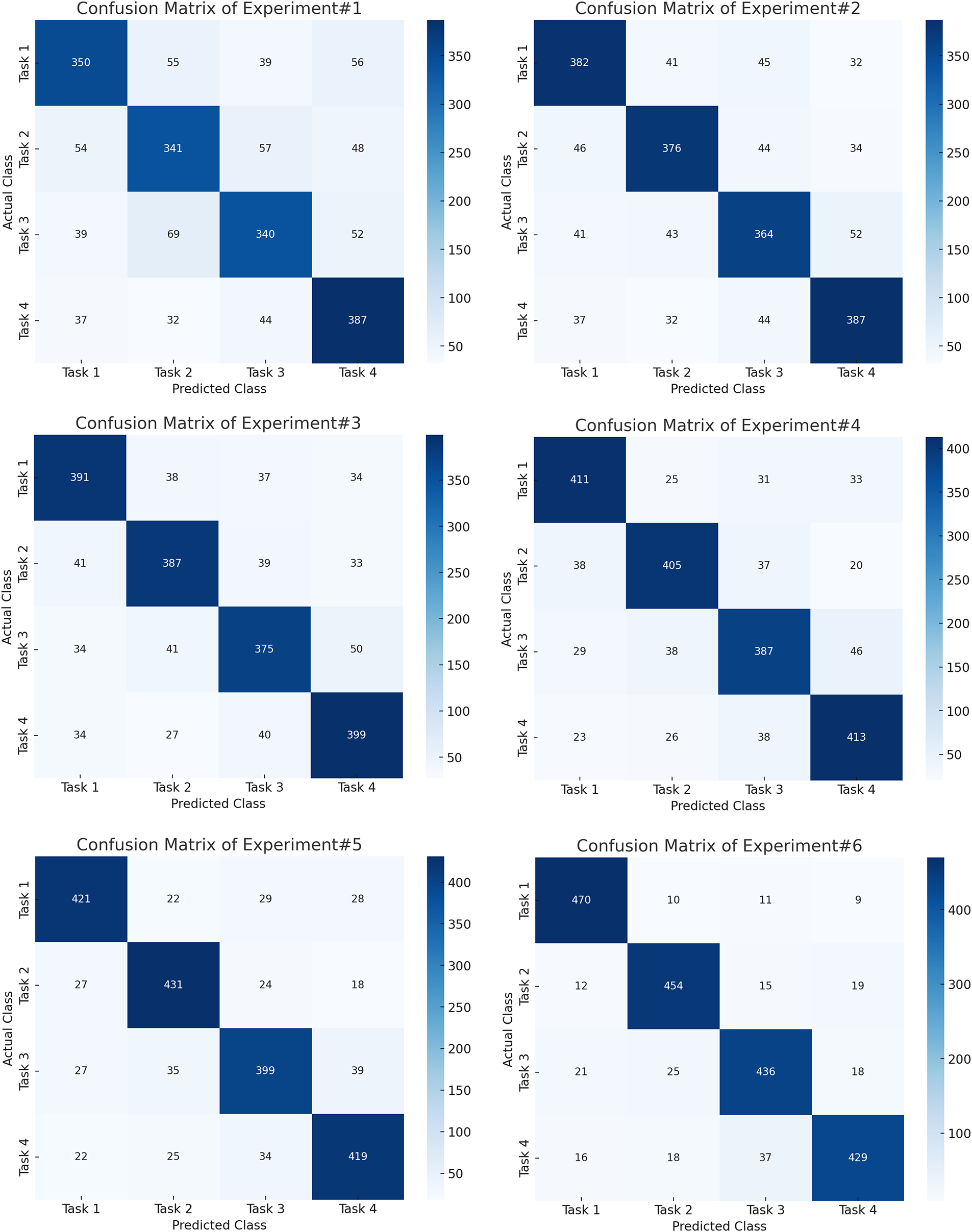
Figure 4: Confusion matrices for the first six experiments on the PhysioNet dataset, showing progressive improvement in class-wise accuracy with successive preprocessing and CNN-based feature extraction.
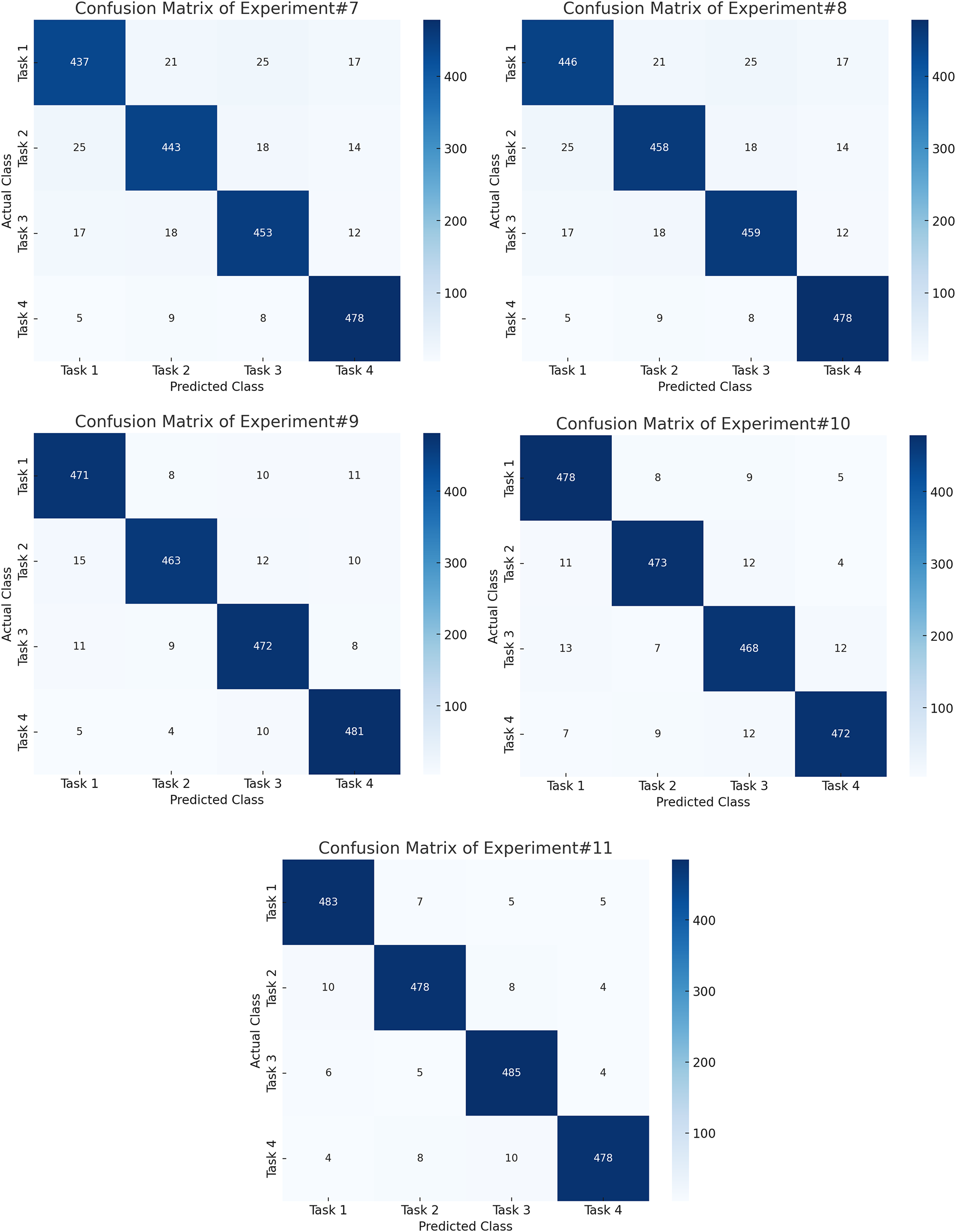
Figure 5: Confusion matrices for the final five experiments on the PhysioNet dataset, demonstrating enhanced class discrimination achieved by feature fusion, PSO optimization, and the ensemble classifier.
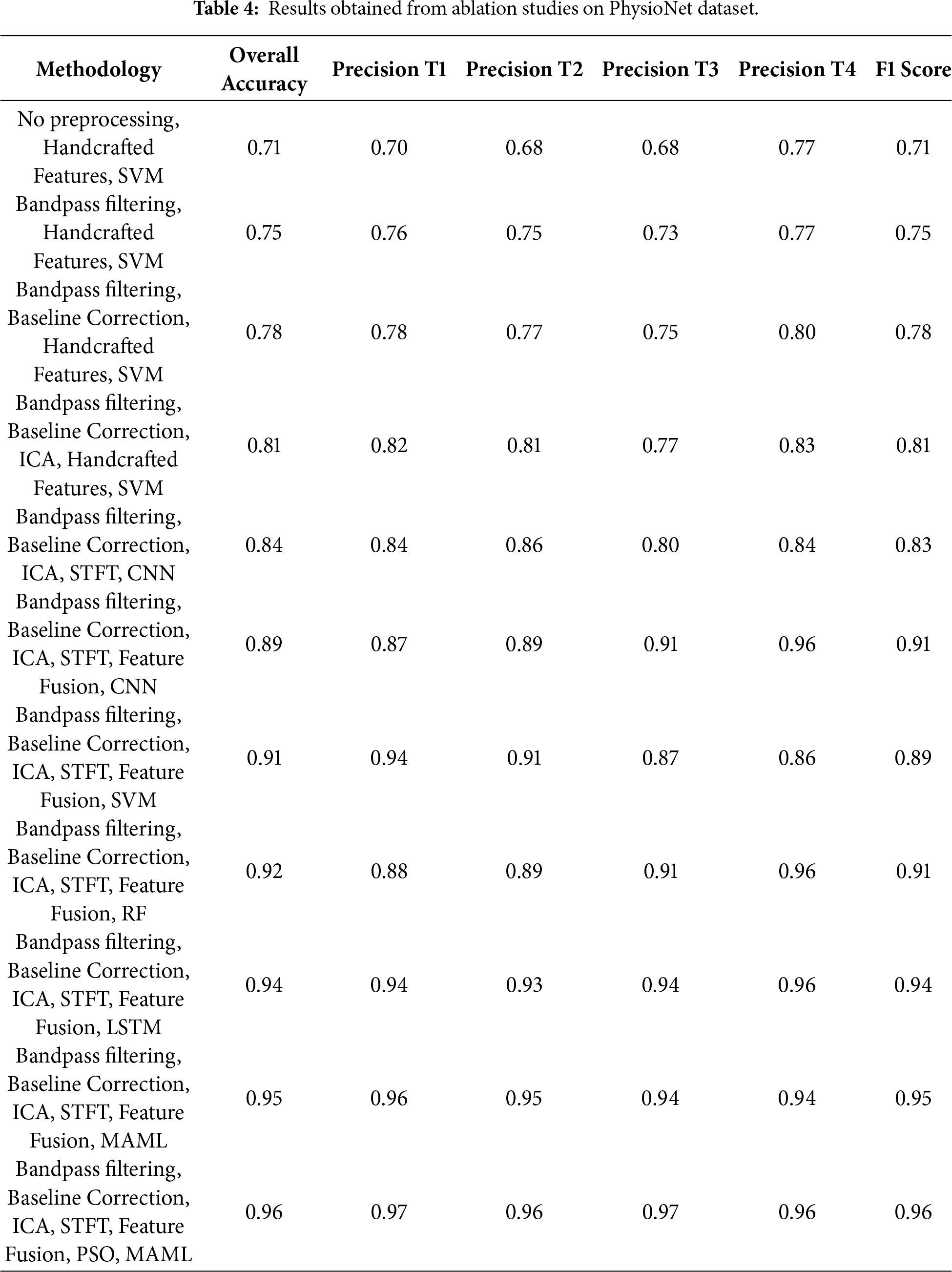
Another notable improvement is the use of ICA for artifact removal, which raised the F1 Score and Overall Accuracy to 81%. This emphasizes the benefits of isolating brain signals from sources of noise. A move toward DL-based feature learning is signaled by the switch from handcrafted features with SVM to employing a STFT representation as input to a CNN. Replacing handcrafted features with STFT-based time-frequency representations and a CNN improved accuracy to 84%, demonstrating the superiority of learned features over manual ones, and also showed that CNNs can automatically extract pertinent time-frequency patterns, even in the absence of explicit feature fusion at this step.
The classifier’s performance improved uniformly across models when feature fusion is implemented (by merging STFT features with other pertinent information) with CNN, the overall accuracy increased to 89%, and the F1 score rose to 91%, with SVM, the accuracy reached 91%, and the F1 score is 89%. While on RF, it got good accuracy (92%) and strong precision F1 score is 91% likely because it can merge non-linear relations in fused features (96%). By fusing the fused features and feeding them into an LSTM network, a large improvement in performance is achieved due to the ability of LSTM networks to capture temporal dependencies. This network achieved an Overall Accuracy of 94% and an F1 Score of 94%. The MI EEG signals, it is shown through this, are highly dependent on the sequential information. By using an ensemble classifier MAML to aggregate predictions of several models, the findings are improved marginally and led to an F1 score of 95% and overall accuracy of 95%. Last but not least, PSO, most likely for feature selection or weighting inside the MAML framework, produced the greatest results across all tests. The significance of model optimization in EEG analysis is highlighted by PSO’s capacity to improve the MAML classifier (95%
The Grad-CAM visualizations show that our proposed customized architecture of CNN focuses on regions within the
In this research, we have also used another benchmark dataset for motor imagery task classification, i.e., BCI Competition IV, dataset 2a. It consists of EEG signal recordings of 09 subjects who performed four tasks, including left hand, right hand, both feet, and tongue movements. A cue-based experimental paradigm is adopted, and all 09 subjects provided instructions to imagine the particular movement for 6 sec following a visual arrow cue. Each subject recorded two sessions on multiple days with six runs in each session and 48 trials that include 12 trials per class; therefore,a total of 288 trials are recorded for every session. EEG signals are recorded using the international 10–20 system, followed by sampling at a rate of 250 Hz. A band-pass filter of 0.5–100 Hz, and a 50 Hz notch filter are then applied for noise removal. In addition, three monopolar EOG channels are recorded to capture ocular activity, enabling subsequent artifact rejection. Trials with significant artifacts are identified by experts and flagged for exclusion.
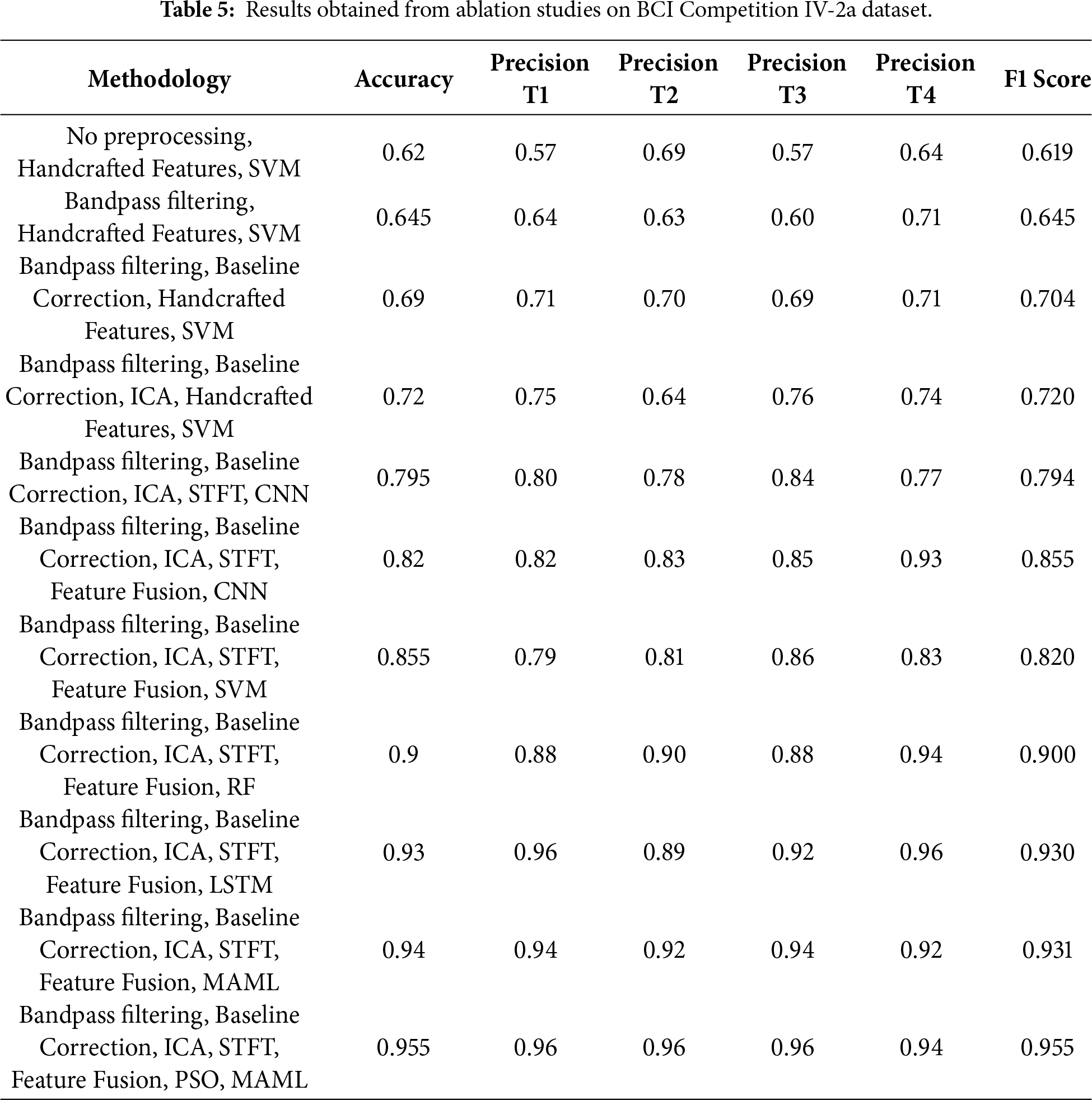
Table 5 presents the ablation study conducted during this research on the BCI Competition IV-2a. Results show that the performance of experiments is increased in terms of accuracy when more processing is performed on EEG signals. Similarly, automated features, when combined with the handcrafted features, performed well. Only handcrafted features with SVM could achieve an accuracy of 62%, whereas when ICA and CNN is applied, the performance improved to 79.5%. The best performance is achieved when PSO-based feature selection is combined with the ensemble classifier. An accuracy of 95.5% and 95.5% F1 score is achieved in the final experiment. Hyperparameters are varied for PSO-based feature selection, STFT windowing, and CNN architecture. Around the selected configuration (PSO: swarm size
To further interpret the decisions made by the proposed framework, we employed two complementary explainability techniques: Grad-CAM for the CNN-based spectral features and SHAP for the handcrafted temporal–spectral features. The Grad-CAM heatmap shown in the Fig. 6 is superimposed on the STFT time–frequency representation of a representative EEG channel from the PhysioNet dataset. The highlighted regions indicate frequency bands and temporal segments that most strongly influenced the model’s decision. To explain the role of handcrafted features in the ensemble model, we generated SHAP values across temporal and spectral descriptors such as RMS amplitude, variance, Hjorth parameters, band powers, entropy, and frequency statistics on the PhysioNet dataset as presented in Fig. 7. To explain the contribution of handcrafted temporal and spectral EEG features, we employed KernelSHAP, a model-agnostic SHAP variant, which is suitable for interpreting heterogeneous classifiers within the MAML-based ensemble framework. Each point represents an individual trial, with its horizontal position showing the magnitude and direction of feature contribution. The color encodes the raw feature value. The results indicate that band power features (alpha and beta ranges), Hjorth mobility/complexity, and spectral entropy had the strongest and most consistent influence on classification outcomes. This suggests that traditional handcrafted descriptors remain highly informative and, when combined with deep features, contribute to the model’s robustness and generalization. Figs. 8 and 9 show the confusion matrices of all the experiments performed on the BCI dataset, whereas Figs. 10 and 11 present the GradCam and SHAP visualizations on the BCI Competition IV-2a dataset.
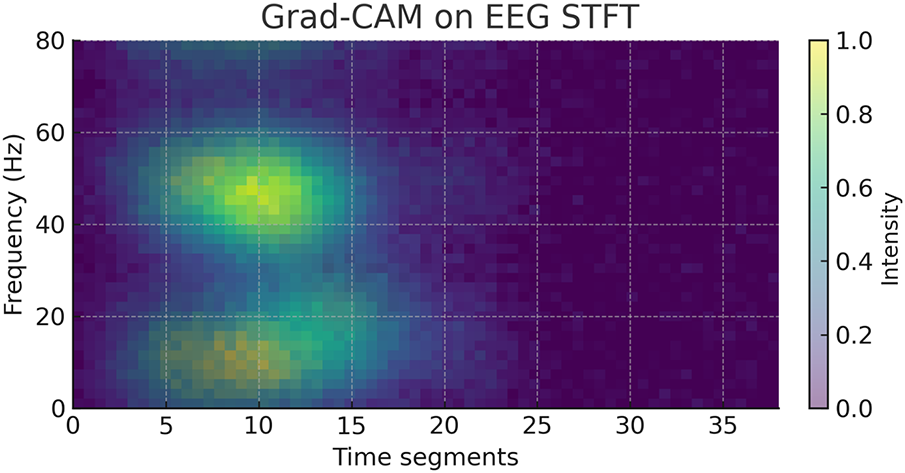
Figure 6: Grad-CAM heatmap on the STFT representation of an EEG channel from the PhysioNet motor imagery dataset.
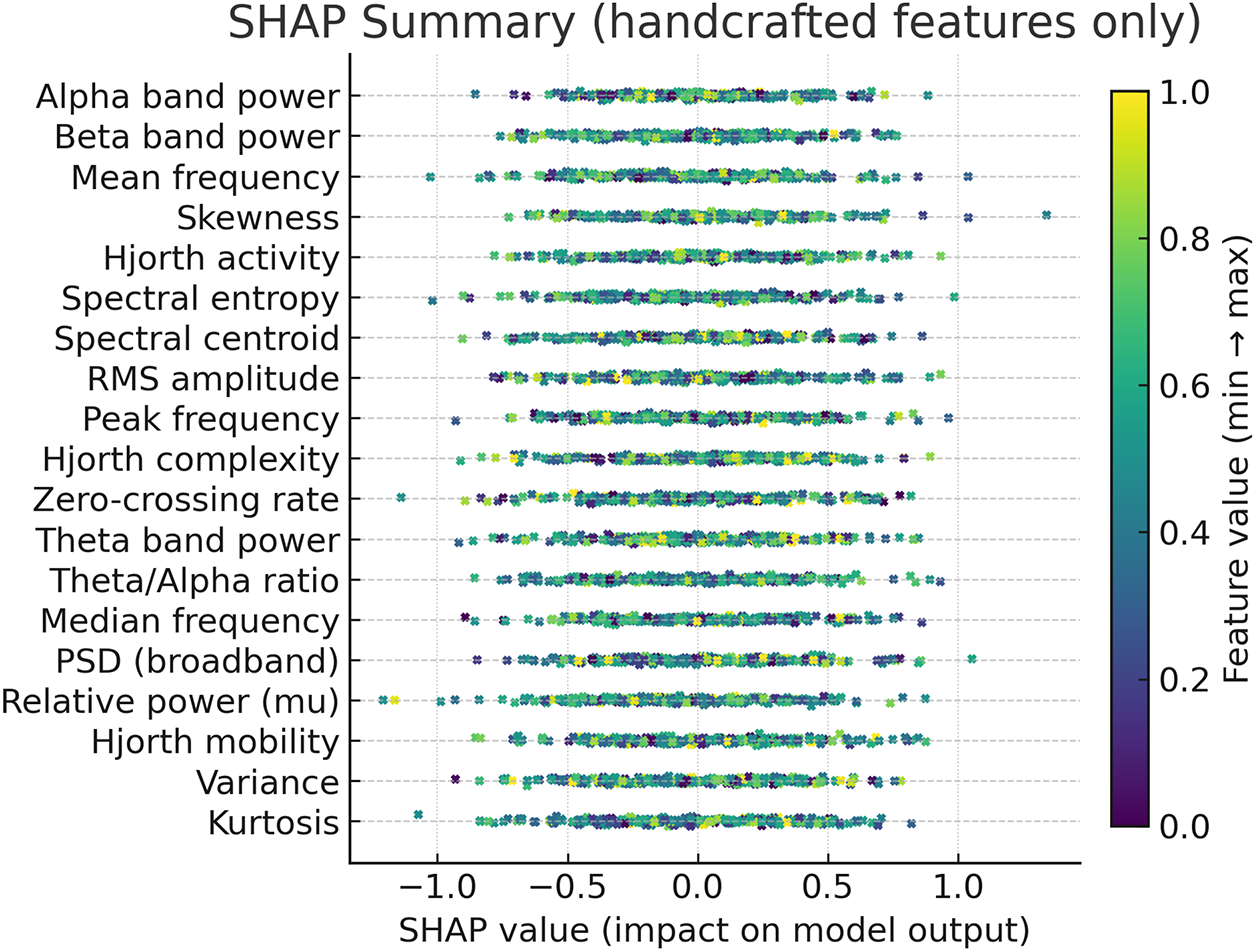
Figure 7: SHAP summary of handcrafted EEG features from the PhysioNet dataset.
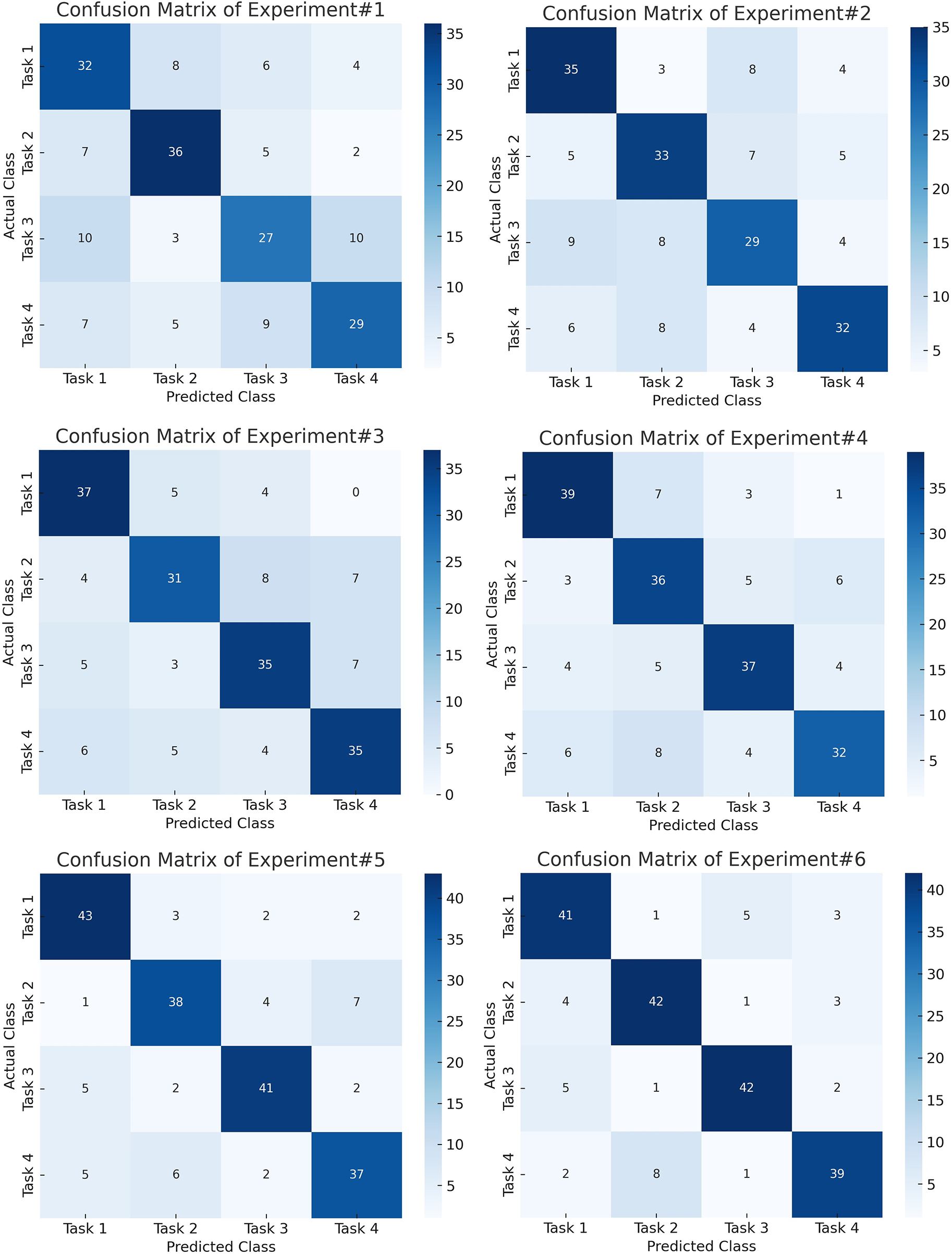
Figure 8: Confusion matrices for the initial experiments on the BCI Competition IV-2a dataset, highlighting reduced inter-class confusion after STFT-based CNN feature learning.
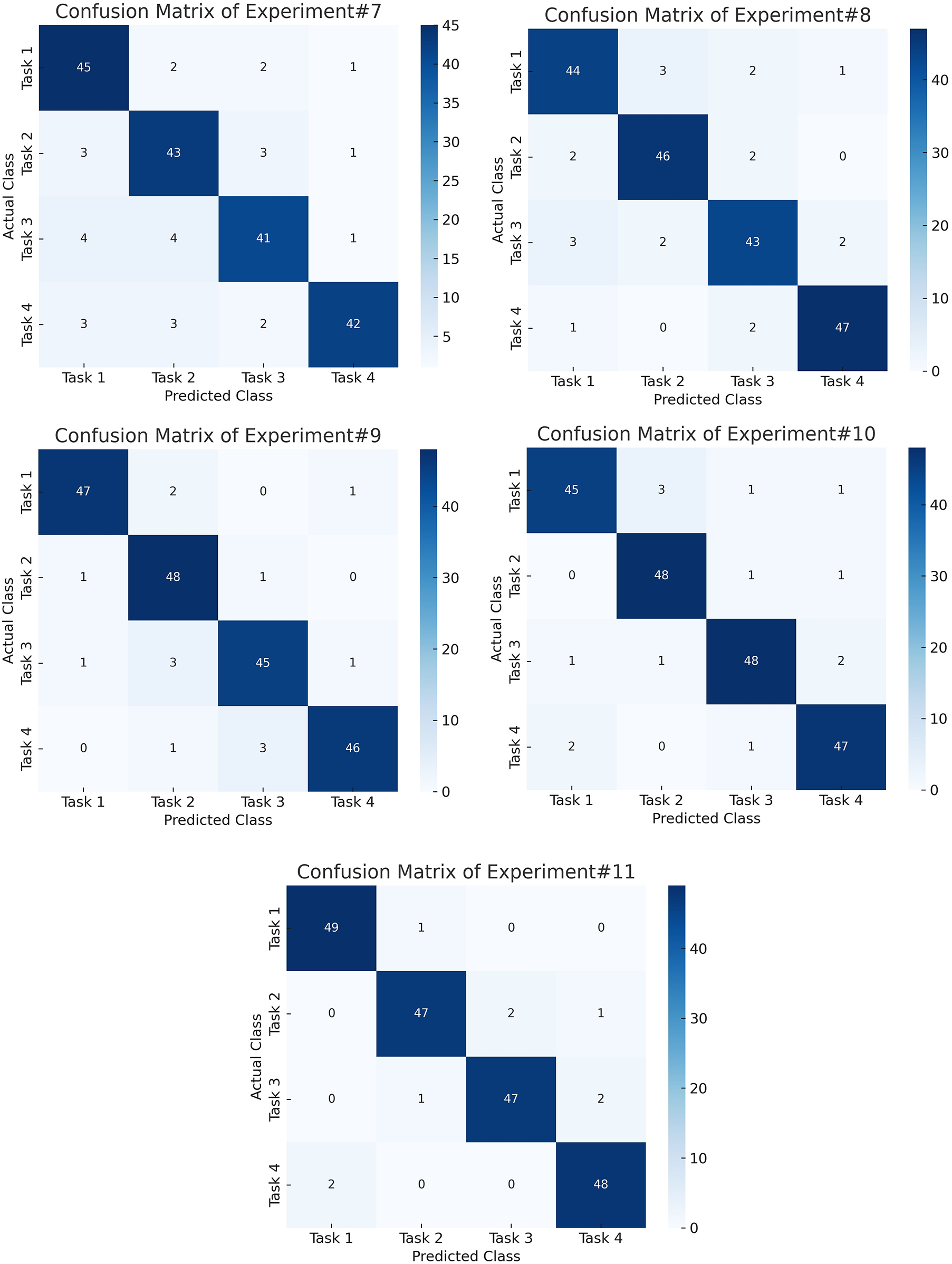
Figure 9: Confusion matrices for the final experiments on the BCI Competition IV-2a dataset, showing robust and balanced performance across all motor imagery classes using the proposed hybrid framework.
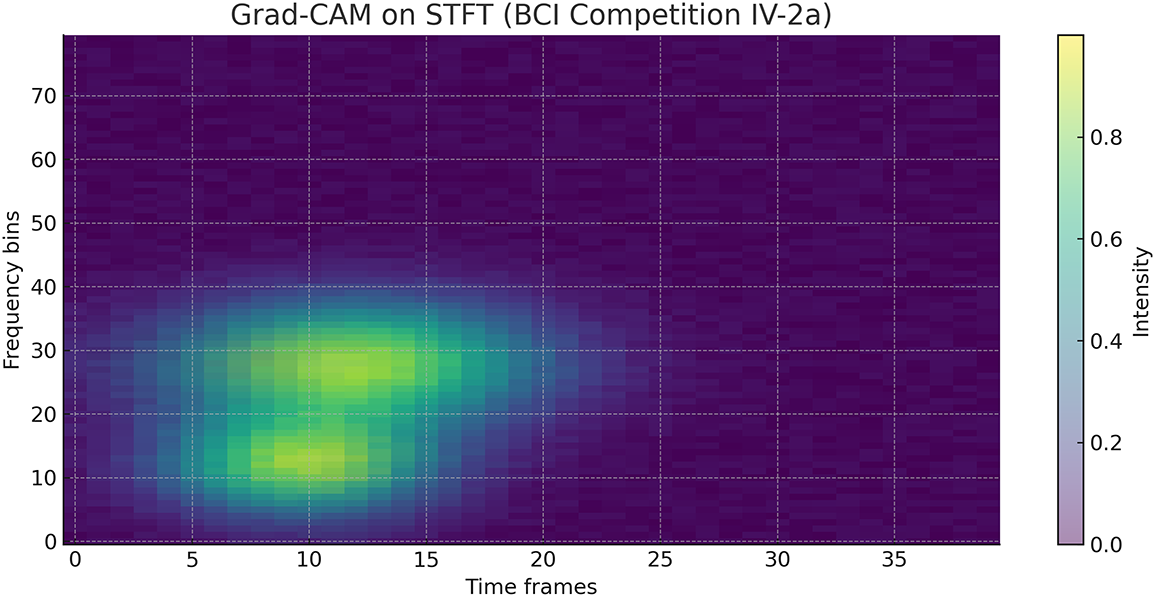
Figure 10: Grad-CAM heatmap on the STFT representation of an EEG channel from the BCI Competition IV-2a dataset, highlighting frequency–time regions most relevant to CNN classification.
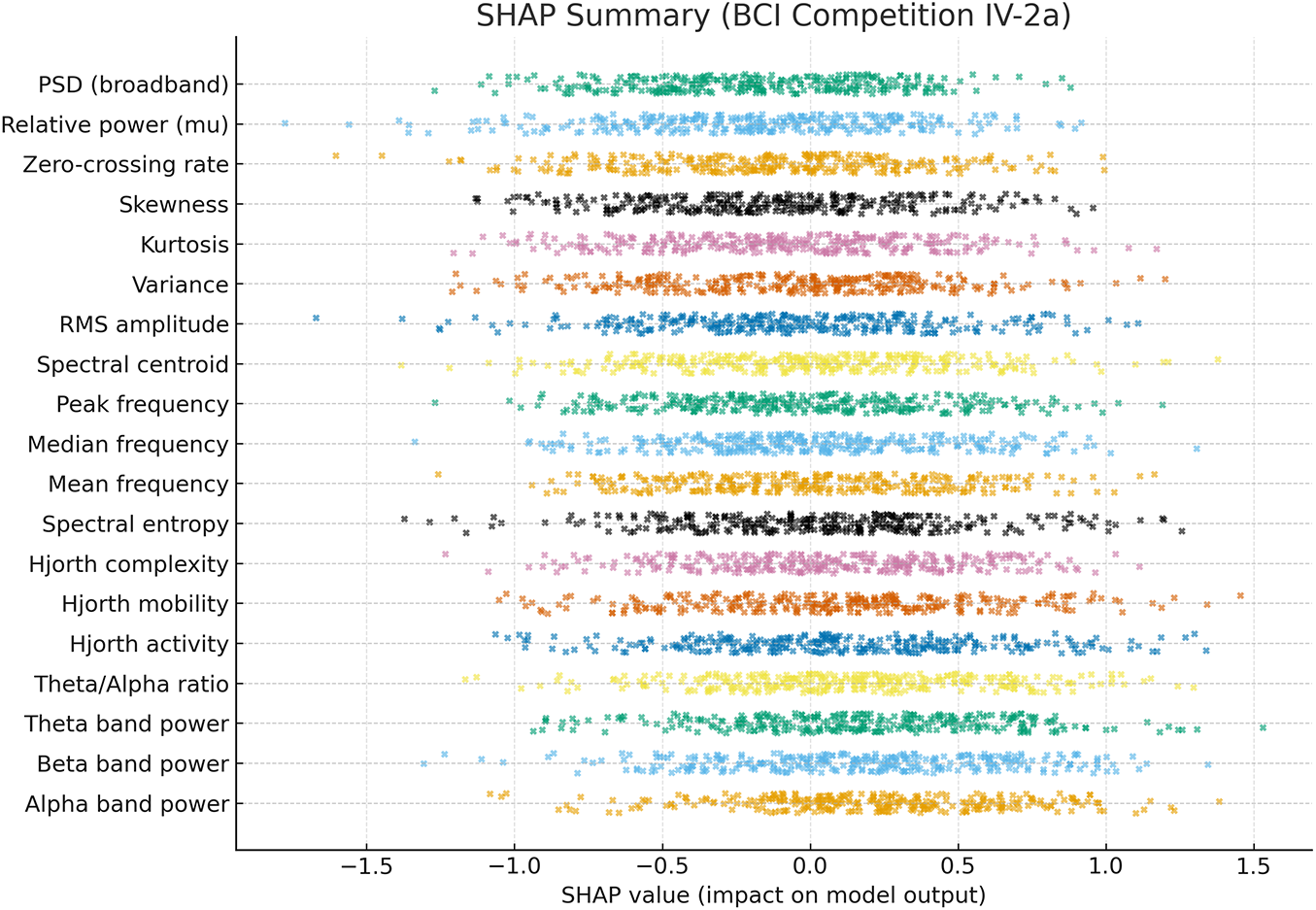
Figure 11: SHAP summary of handcrafted EEG features from the BCI Competition IV-2a dataset, showing feature contributions (SHAP values) to classification, with alpha/beta band power, Hjorth parameters, and spectral entropy as the most influential.
Comparison of the accuracy and F1 score of the proposed method with state-of-the-art methods on the PhsioNet dataset is presented in Table 6. The proposed approach beats most of the existing approaches with 96% accuracy and 96% F1-score. Despite the perfect accuracy of 100% reported by Taghizadeh et al. [36], the lack of the corresponding F1 score does not allow making a balanced evaluation of the effectiveness of this model. Other similar studies, such as Alnaanah et al. [37], Hwaidi et al. [21], Herrera et al. [24], Yu et al. [38], and Huang et al. [39], only reported accuracy, making it hard to measure their performance in other classes. Compared to our model, other studies reporting both metrics also have competitive performance, Sartipi et al. [40] and Mitul et al. [41] demonstrated competitive performance, yet still fall short compared to our model. For instance, Lotey et al. [42] achieve an accuracy of 95.05% and an F1 score of 95%, which is closest to our results. Sartipi et al. [40] and Mitul et al. [41] demonstrated competitive performance, yet still fall short compared to our model. For instance, Lotey et al. [42] achieved an accuracy of 95.05% and an F1 score of 95%, which is closest to our results. To statistically demonstrate the superiority of the proposed method, a paired two-tailed t-test is performed on the classification outcomes received over 10 independent runs on the PhysioNet dataset. The results demonstrate that both the improvements in accuracy and F1 score are statistically significant on comparison with the best performing baseline (Lotey et al. [42]), p-values being 0.0042 and 0.0037, respectively. Such values are far below the traditional threshold of significance of 0.05. This level of performance and statistical validation reveals the robustness and generalizability of the presented model for practical utility in real-world diagnostic scenarios.
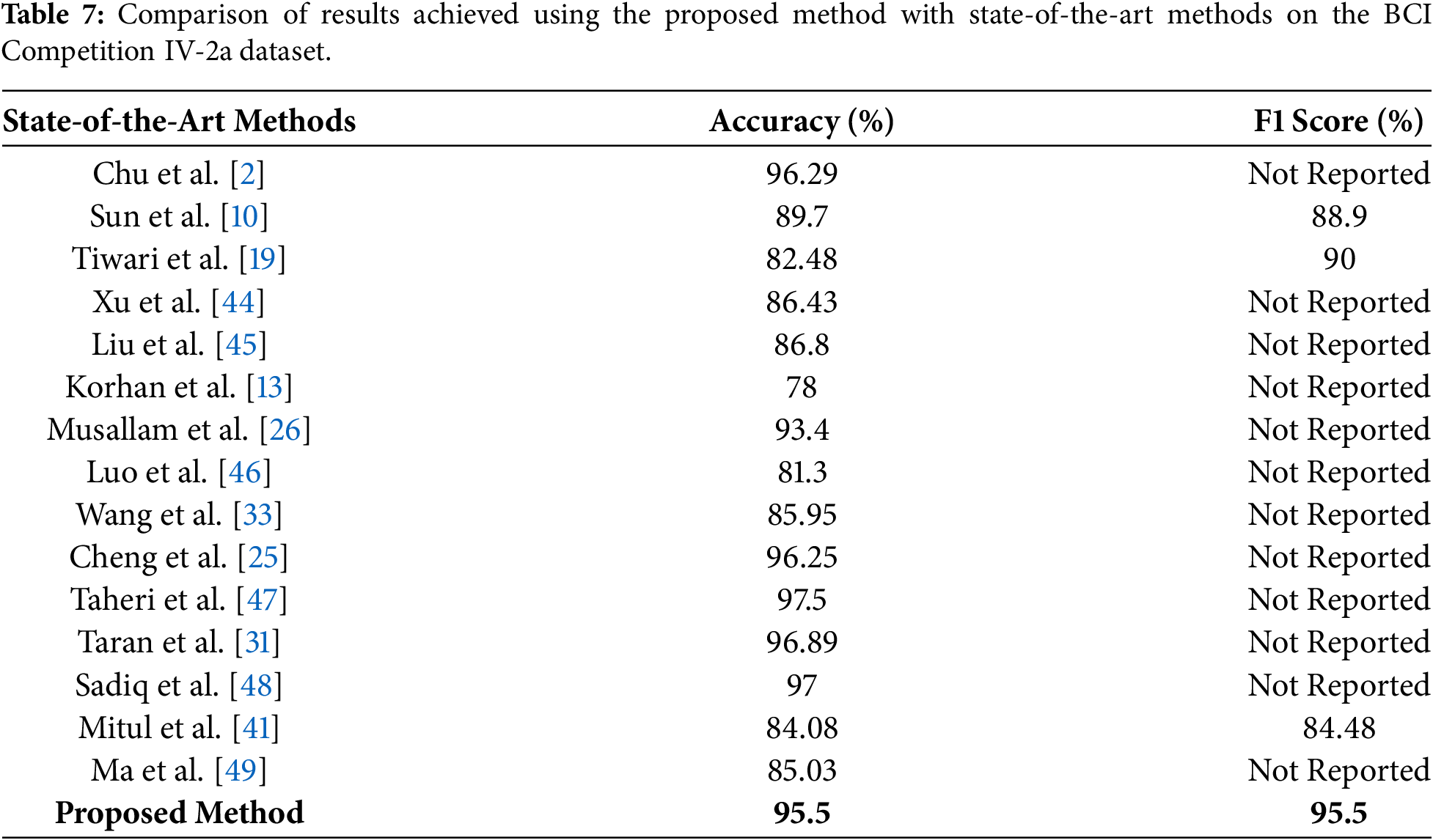
Table 7 compares the proposed method with state-of-the-art approaches on the BCI Competition IV-2a dataset. Our proposed method achieved an accuracy and F1 score of 95.5% which shows the improved performance and generalizability across all classes. We have also conducted a statistical significance test to validate the generalization of our proposed method on the BCI dataset. A paired two-tailed t-test is performed over 10 independent experimental runs that compared the proposed MAML-based ensemble method for MI task classification. Classification accuracy and F1 score are used as evaluation metrics. Results of this statistical analysis show the p-values of 0.0061 for accuracy and 0.0048 for F1 score, which show the improved performance. These findings confirm that the observed gains are not due to random variation and further reinforce the generalizability and reliability of the proposed approach across different motor imagery EEG benchmarks.
We have also computed the computational cost of the proposed method to assess its performance in real-time BCI applications. For a single EEG trial, bandpass filtering and baseline correction did not include any major overhead, while ICA decomposition requires approximately 20–30 ms per trial on a CPU-only setup. STFT computation using a 128-sample window with 50% overlap requires approximately 25–35 ms, depending on signal length. CNN-based feature extraction constitutes the dominant cost during inference, requiring approximately 60–80 ms per trial. Importantly, PSO-based feature selection and MAML meta-learning are performed offline during training and therefore do not contribute to online inference latency. The ensemble inference stage (SVM, RF, and LSTM with decision-level fusion) adds less than 10 ms. Overall, the end-to-end online inference latency remains below 150 ms per trial on a CPU-only platform, indicating that the proposed framework is suitable for near real-time, window-based BCI applications, provided that task decoding is performed at the trial or sliding-window level.
4 Conclusion and Future Directions
In this research, we propose a novel ensemble method for the classification of MI tasks using EEG signals. The proposed method consists of a bandpass filter and ICA for denoising EEG signals and conversion of multichannel EEG signals into a single surrogate channel using a CSP filter. We propose a customized 2D-CNN for automated feature extraction and concatenated both handcrafted and automated features to form a single feature vector. PSO is applied for the selection of an optimal feature vector. An ensemble classifier, MAML, is used, which receives the outputs of three classifiers, including SVM, RF, and LSTM. We trained and tested the proposed method on two benchmark datasets of MI task classification, including PhysioNet and the BCI Competition IV-2a. The proposed method outperforms the state-of-the-art methods proposed by researchers in recent years in terms of both accuracy and F1 score. The proposed framework is suitable for real-time deployment, with inference latency below 150 ms per trial on a CPU-only setup and negligible overhead from the PSO-optimized MAML ensemble after training. The customized 2D-CNN contains 684,900 trainable parameters, indicating a favorable balance between accuracy and computational efficiency for practical BCI applications. In the future, we intend to further expand our work in multiple directions. Proposed work can be applied in multiple domains, including wheelchair automation, prosthetic arms for stroke patients, robotics, and Gaming. Real-time implementation of the proposed method can also be done in the future with optimization of computational resources and reduced latency in wearable devices. Further, inclusion of different modalities like EMG, EEG, and fMRI can also be utilized to further enhance the robustness of the MI tasks classification method.
Acknowledgement: Not applicable.
Funding Statement: This work was supported and funded by the Deanship of Scientific Research at Imam Mohammad Ibn Saud Islamic University (IMSIU) (grant number IMSIU-DDRSP2601).
Author Contributions: Conceptualization, Fazal Ur Rehman, Yazeed Alkhrijah, Syed Muhammad Usman and Muhammad Irfan; methodology, Fazal Ur Rehman, Yazeed Alkhrijah, Syed Muhammad Usman and Muhammad Irfan; software, Fazal Ur Rehman, Yazeed Alkhrijah, Syed Muhammad Usman and Muhammad Irfan; validation, Fazal Ur Rehman, Yazeed Alkhrijah, Syed Muhammad Usman and Muhammad Irfan; formal analysis, Fazal Ur Rehman, Yazeed Alkhrijah, Syed Muhammad Usman and Muhammad Irfan; investigation, Fazal Ur Rehman, Yazeed Alkhrijah, Syed Muhammad Usman and Muhammad Irfan; resources, Fazal Ur Rehmanr, Yazeed Alkhrijah, Syed Muhammad Usman and Muhammad Irfan; data curation, Fazal Ur Rehman, Yazeed Alkhrijah, Syed Muhammad Usman and Muhammad Irfan; writing—original draft preparation, Fazal Ur Rehman, Yazeed Alkhrijah, Syed Muhammad Usman and Muhammad Irfan; writing—review and editing, Fazal Ur Rehman, Yazeed Alkhrijah, Syed Muhammad Usman and Muhammad Irfan; visualization, Fazal Ur Rehman, Yazeed Alkhrijah, Syed Muhammad Usman and Muhammad Irfan; supervision, Syed Muhammad Usman and Muhammad Irfan; project administration, Syed Muhammad Usman and Muhammad Irfan. All authors reviewed and approved the final version of the manuscript.
Availability of Data and Materials: The datasets generated and/or analyzed during this study are available in the PhysioNet Cardiology Challenge 2017 repository at https://archive.physionet.org/challenge/2017/, https://www.bbci.de/competition/iv/#dataset2a.
Ethics Approval: Not applicable.
Conflicts of Interest: The authors declare no conflicts of interest.
References
1. McFarland DJ, Wolpaw JR. Brain-computer interfaces for communication and control. Commun ACM. 2011;54(5):60–6. doi:10.1145/1941487.1941506. [Google Scholar] [PubMed] [CrossRef]
2. Chu C, Xiao Q, Chang L, Shen J, Zhang N, Du Y, et al. EEG temporal information-based 1-D convolutional neural network for motor imagery classification. Multimed Tools Appl. 2023;82(29):45747–67. doi:10.1007/s11042-023-16536-x. [Google Scholar] [CrossRef]
3. Barmpas K, Panagakis Y, Adamos DA, Laskaris N, Zafeiriou S. BrainWave-Scattering Net: a lightweight network for EEG-based motor imagery recognition. J Neural Eng. 2023;20(5):056014. doi:10.1088/1741-2552/acf78a. [Google Scholar] [PubMed] [CrossRef]
4. Padfield N, Zabalza J, Zhao H, Masero V, Ren J. EEG-based brain-computer interfaces using motor-imagery: techniques and challenges. Sensors. 2019;19(6):1423. doi:10.3390/s19061423. [Google Scholar] [PubMed] [CrossRef]
5. Roy G, Bhoi AK, Das S, Bhaumik S. Cross-correlated spectral entropy-based classification of EEG motor imagery signal for triggering lower limb exoskeleton. Signal Image Video Process. 2022;16(7):1831–9. doi:10.1007/s11760-022-02142-1. [Google Scholar] [CrossRef]
6. Orban M, Elsamanty M, Guo K, Zhang S, Yang H. A review of brain activity and EEG-based brain-computer interfaces for rehabilitation application. Bioengineering. 2022;9(12):768. doi:10.3390/bioengineering9120768. [Google Scholar] [PubMed] [CrossRef]
7. Wang J, Bi L, Fei W. EEG-Based motor BCIs for upper limb movement: current techniques and future insights. IEEE Trans Neural Syst Rehabil Eng. 2023;31:4413–27. doi:10.1109/tnsre.2023.3330500. [Google Scholar] [PubMed] [CrossRef]
8. Iyama Y, Hori J, Baba K. High-resolution cortical dipole imaging using EEG data measured with a small number of electrodes based on the international 10-20 system. IEEJ Trans Electr Electron Eng. 2024;20(3):422–30. doi:10.1002/tee.24215. [Google Scholar] [CrossRef]
9. Park S, Ha J, Kim L. Improving performance of motor imagery-based brain-computer interface in poorly performing subjects using a hybrid-imagery method utilizing combined motor and somatosensory activity. IEEE Trans Neural Syst Rehabil Eng. 2023;31:1064–74. doi:10.1109/tnsre.2023.3237583. [Google Scholar] [PubMed] [CrossRef]
10. Sun B, Liu Z, Wu Z, Mu C, Li T. Graph convolution neural network based end-to-end channel selection and classification for motor imagery brain-computer interfaces. IEEE Trans Ind Inform. 2022;19(9):9314–24. doi:10.1109/tii.2022.3227736. [Google Scholar] [CrossRef]
11. Annaby MH, Said M, Eldeib A, Rushdi MA. EEG-based motor imagery classification using digraph Fourier transforms and extreme learning machines. Biomed Signal Process Control. 2021;69:102831. doi:10.1016/j.bspc.2021.102831. [Google Scholar] [CrossRef]
12. Barmpas K, Panagakis Y, Bakas S, Adamos DA, Laskaris N, Zafeiriou S. Improving generalization of CNN-based motor-imagery EEG decoders via dynamic convolutions. IEEE Trans Neural Syst Rehabil Eng. 2023;31:1997–2005. doi:10.1109/tnsre.2023.3265304. [Google Scholar] [PubMed] [CrossRef]
13. Korhan N, Abilzade L, Ölmez T, Ölmez ZD. Classification of left and right hand motor imagery EEG signals by using deep neural networks. Int J Appl Math Electron Comput. 2021;9(4):85–90. doi:10.18100/ijamec.995022. [Google Scholar] [CrossRef]
14. Ferracuti F, Iarlori S, Mansour Z, Monteriù A, Porcaro C. Comparing between different sets of preprocessing, classifiers, and channels selection techniques to optimise motor imagery pattern classification system from EEG pattern recognition. Brain Sci. 2021;12(1):57. doi:10.3390/brainsci12010057. [Google Scholar] [PubMed] [CrossRef]
15. Leske S, Dalal SS. Reducing power line noise in EEG and MEG data via spectrum interpolation. Neuroimage. 2019;189:763–76. doi:10.1016/j.neuroimage.2019.01.026. [Google Scholar] [PubMed] [CrossRef]
16. Yang SY, Lin YP. Movement artifact suppression in wearable low-density and dry EEG recordings using active electrodes and artifact subspace reconstruction. IEEE Trans Neural Syst Rehabil Eng. 2023;31:3844–53. doi:10.1109/tnsre.2023.3319355. [Google Scholar] [PubMed] [CrossRef]
17. Seok D, Lee S, Kim M, Cho J, Kim C. Motion artifact removal techniques for wearable EEG and PPG sensor systems. Front Electron. 2021;2:685513. doi:10.3389/felec.2021.685513. [Google Scholar] [CrossRef]
18. Alazrai R, Abuhijleh M, Alwanni H, Daoud MI. A deep learning framework for decoding motor imagery tasks of the same hand using EEG signals. IEEE Access. 2019;7:109612–27. doi:10.1109/access.2019.2934018. [Google Scholar] [CrossRef]
19. Tiwari S, Goel S, Bhardwaj A. MIDNN-a classification approach for the EEG based motor imagery tasks using deep neural network. Appl Intell. 2022;52(5):4824–43. doi:10.1007/s10489-021-02622-w. [Google Scholar] [CrossRef]
20. Xie J, Zhang J, Sun J, Ma Z, Qin L, Li G, et al. A transformer-based approach combining deep learning network and spatial-temporal information for raw EEG classification. IEEE Trans Neural Syst Rehabil Eng. 2022;30:2126–36. doi:10.1109/tnsre.2022.3194600. [Google Scholar] [PubMed] [CrossRef]
21. Hwaidi JF, Chen TM. Classification of motor imagery EEG signals based on deep autoencoder and convolutional neural network approach. IEEE Access. 2022;10:48071–81. doi:10.1109/access.2022.3171906. [Google Scholar] [CrossRef]
22. Nawaz R, Cheah KH, Nisar H, Yap VV. Comparison of different feature extraction methods for EEG-based emotion recognition. Biocybern Biomed Eng. 2020;40(3):910–26. doi:10.1016/j.bbe.2020.04.005. [Google Scholar] [CrossRef]
23. Chen J, Yu Z, Gu Z, Li Y. Deep temporal-spatial feature learning for motor imagery-based brain-computer interfaces. IEEE Trans Neural Syst Rehabil Eng. 2020;28(11):2356–66. doi:10.1109/tnsre.2020.3023417. [Google Scholar] [PubMed] [CrossRef]
24. Lazcano-Herrera AG, Fuentes-Aguilar RQ, Ramirez-Morales A, Alfaro-Ponce M. BiLSTM and SqueezeNet with transfer learning for EEG motor imagery classification: validation with own dataset. IEEE Access. 2023;11:136422–36. doi:10.1109/access.2023.3328254. [Google Scholar] [CrossRef]
25. Cheng L, Li D, Yu G, Zhang Z, Li X, Yu S. A motor imagery EEG feature extraction method based on energy principal component analysis and deep belief networks. IEEE Access. 2020;8:21453–72. doi:10.1109/access.2020.2969054. [Google Scholar] [CrossRef]
26. Musallam YK, AlFassam NI, Muhammad G, Amin SU, Alsulaiman M, Abdul W, et al. Electroencephalography-based motor imagery classification using temporal convolutional network fusion. Biomed Signal Process Control. 2021;69:102826. doi:10.1016/j.bspc.2021.102826. [Google Scholar] [CrossRef]
27. Sun B, Zhao X, Zhang H, Bai R, Li T. EEG motor imagery classification with sparse spectrotemporal decomposition and deep learning. IEEE Trans Autom Sci Eng. 2020;18(2):541–51. doi:10.1109/tase.2020.3021456. [Google Scholar] [CrossRef]
28. Zhang D, Chen K, Jian D, Yao L. Motor imagery classification via temporal attention cues of graph embedded EEG signals. IEEE J Biomed Health Inform. 2020;24(9):2570–9. doi:10.1109/jbhi.2020.2967128. [Google Scholar] [PubMed] [CrossRef]
29. Venkatachalam K, Devipriya A, Maniraj J, Sivaram M, Ambikapathy A, Iraj SA. A novel method of motor imagery classification using eeg signal. Artif Intell Med. 2020;103:101787. doi:10.1016/j.artmed.2019.101787. [Google Scholar] [PubMed] [CrossRef]
30. Geng X, Li D, Chen H, Yu P, Yan H, Yue M. An improved feature extraction algorithms of EEG signals based on motor imagery brain-computer interface. Alex Eng J. 2022;61(6):4807–20. doi:10.1016/j.aej.2021.10.034. [Google Scholar] [CrossRef]
31. Taran S, Bajaj V. Motor imagery tasks-based EEG signals classification using tunable-Q wavelet transform. Neural Comput Appl. 2019;31(11):6925–32. doi:10.1007/s00521-018-3531-0. [Google Scholar] [CrossRef]
32. Subasi A, Mian Qaisar S. The ensemble machine learning-based classification of motor imagery tasks in brain-computer interface. J Healthc Eng. 2021;2021:1970769–12. doi:10.1155/2021/1970769. [Google Scholar] [PubMed] [CrossRef]
33. Wang J, Feng Z, Ren X, Lu N, Luo J, Sun L. Feature subset and time segment selection for the classification of EEG data based motor imagery. Biomed Signal Process Control. 2020;61:102026. doi:10.1016/j.bspc.2020.102026. [Google Scholar] [CrossRef]
34. Chae MS, Rehman A, Kim Y, Kim J, Han Y, Park S, et al. Eye-blink and SSVEP-based selective attention interface for XR user authentication: an explainable neural decoding and machine learning approach to reducing visual fatigue. IEEE Access. 2025;13:176998–7018. doi:10.1109/access.2025.3613355. [Google Scholar] [CrossRef]
35. Rehman A, Lee M, Kim Y, Chae MS, Mun S. Machine learning-driven XR interface using ERP decoding. Electronics. 2025;14(19):3773. doi:10.3390/electronics14193773. [Google Scholar] [CrossRef]
36. Taghizadeh M, Vaez F, Faezipour M. EEG motor imagery classification by feature extracted deep 1D-CNN and semi-deep fine-tuning. IEEE Access. 2024;12(1):111265–79. doi:10.1109/access.2024.3430838. [Google Scholar] [CrossRef]
37. Alnaanah M, Wahdow M, Alrashdan M. CNN models for EEG motor imagery signal classification. Signal Image Video Process. 2023;17(3):825–30. doi:10.1007/s11760-022-02293-1. [Google Scholar] [CrossRef]
38. Yu H, Baek S, Lee J, Sohn I, Hwang B, Park C. Deep neural network-based empirical mode decomposition for motor imagery EEG classification. IEEE Trans Neural Syst Rehabil Eng. 2024;32:3647–56. doi:10.1109/tnsre.2024.3432102. [Google Scholar] [PubMed] [CrossRef]
39. Huang Y, Zheng J, Xu B, Li X, Liu Y, Wang Z, et al. An improved model using convolutional sliding window-attention network for motor imagery EEG classification. Front Neurosci. 2023;17:1204385. doi:10.3389/fnins.2023.1204385. [Google Scholar] [PubMed] [CrossRef]
40. Sartipi S, Cetin M. Subject-independent deep architecture for EEG-based motor imagery classification. IEEE Trans Neural Syst Rehabil Eng. 2024;32:718–27. doi:10.1109/tnsre.2024.3360194. [Google Scholar] [PubMed] [CrossRef]
41. Mitul MKH, Ferdous MJ. Real-time BCI scheme to assisting disabled persons using motor imagery EEG signal classification. Khulna Univ Stud. 2025;22(1):49–61. doi:10.53808/kus.2025.22.01.1204-se. [Google Scholar] [CrossRef]
42. Lotey T, Keserwani P, Dogra DP, Roy PP. Feature reweighting for EEG-based motor imagery classification. arXiv: 230802515. 2023. [Google Scholar]
43. Fan C, Yang B, Li X, Zan P. Temporal-frequency-phase feature classification using 3D-convolutional neural networks for motor imagery and movement. Front Neurosci. 2023;17:1250991. doi:10.3389/fnins.2023.1250991. [Google Scholar] [PubMed] [CrossRef]
44. Xu S, Zhu L, Kong W, Peng Y, Hu H, Cao J. A novel classification method for EEG-based motor imagery with narrow band spatial filters and deep convolutional neural network. Cogn Neurodynamics. 2022;16(2):379–89. doi:10.1007/s11571-021-09721-x. [Google Scholar] [PubMed] [CrossRef]
45. Liu X, Shi R, Hui Q, Xu S, Wang S, Na R, et al. TCACNet: temporal and channel attention convolutional network for motor imagery classification of EEG-based BCI. Inf Process Manag. 2022;59(5):103001. doi:10.1016/j.ipm.2022.103001. [Google Scholar] [CrossRef]
46. Luo J, Gao X, Zhu X, Wang B, Lu N, Wang J. Motor imagery EEG classification based on ensemble support vector learning. Comput Methods Programs Biomed. 2020;193:105464. doi:10.1016/j.cmpb.2020.105464. [Google Scholar] [PubMed] [CrossRef]
47. Taheri S, Ezoji M, Sakhaei SM. Convolutional neural network based features for motor imagery EEG signals classification in brain-computer interface system. SN Appl Sci. 2020;2(4):555. doi:10.1007/s42452-020-2378-z. [Google Scholar] [CrossRef]
48. Sadiq MT, Yu X, Yuan Z, Zeming F, Rehman AU, Ullah I, et al. Motor imagery EEG signals decoding by multivariate empirical wavelet transform-based framework for robust brain-computer interfaces. IEEE Access. 2019;7:171431–51. doi:10.1109/access.2019.2956018. [Google Scholar] [CrossRef]
49. Ma X, Chen W, Pei Z, Zhang Y, Chen J. Attention-based convolutional neural network with multi-modal temporal information fusion for motor imagery EEG decoding. Comput Biol Med. 2024;175:108504. doi:10.1016/j.compbiomed.2024.108504. [Google Scholar] [PubMed] [CrossRef]
Cite This Article
 Copyright © 2026 The Author(s). Published by Tech Science Press.
Copyright © 2026 The Author(s). Published by Tech Science Press.This work is licensed under a Creative Commons Attribution 4.0 International License , which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.


 Submit a Paper
Submit a Paper Propose a Special lssue
Propose a Special lssue View Full Text
View Full Text Download PDF
Download PDF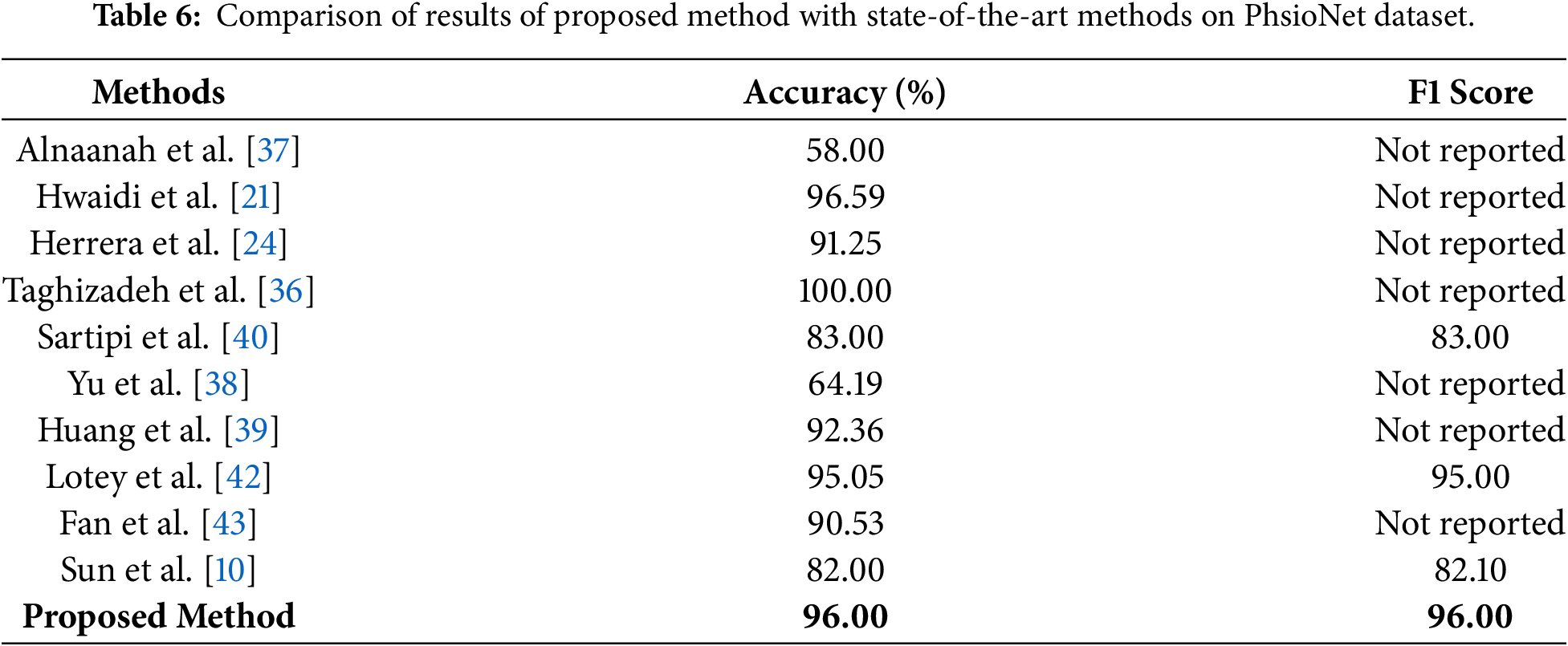
 Downloads
Downloads
 Citation Tools
Citation Tools