 Open Access
Open Access
ARTICLE
A Surrogate Deep-Learning Super-Resolution Framework for Accelerating Finite Element Method-Based Fluid Simulations
1 Department of Innovative Divertor Development Group, Korea Institute of Fusion Energy, Daejeon, Republic of Korea
2 Department of Mechanical Engineering, Changwon National University, Changwon, Republic of Korea
3 Department of Neurosurgery, Sungkyunkwan University School of Medicine, Samsung Changwon Hospital, Changwon, Republic of Korea
* Corresponding Author: Jaemin Kim. Email:
(This article belongs to the Special Issue: Machine Learning, Data-Driven and Novel Approaches in Computational Mechanics)
Computer Modeling in Engineering & Sciences 2026, 146(3), 21 https://doi.org/10.32604/cmes.2026.079127
Received 15 January 2026; Accepted 15 January 2026; Issue published 30 March 2026
Abstract
This study develops a surrogate super-resolution (SR) framework that accelerates finite element method (FEM)-based computational fluid dynamics (CFD) using deep learning. High-resolution (HR) FEM-based CFD remains computationally prohibitive for time-sensitive applications, including patient-specific aneurysm hemodynamics where rapid turnaround is valuable. The proposed pipeline learns to reconstruct HR velocity-magnitude fields from low-resolution (LR) FEM solutions generated under the same governing equations and boundary conditions. It consists of three modules: (i) offline pre-training of a residual network on representative vascular geometries; (ii) lightweight fine-tuning to adapt the pretrained model to geometric variability, including patient-specific aneurysm morphologies; and (iii) an unstructured-to-structured sampling strategy with region-of-interest upsampling that concentrates resolution in flow-critical zones (e.g., the aneurysm sac) rather than the full domain. This targeted reconstruction substantially reduces inference and post-processing cost while preserving key HR flow features. Experiments on cerebral aneurysm models show that HR velocity-magnitude fields can be recovered with accuracy comparable to direct HR simulations at less than 1% of the direct HR simulation cost per analysis (LR simulation and SR inference), while adaptation to new geometries requires only lightweight fine-tuning with limited target-specific HR data. While clinical endpoints and additional variables (e.g., pressure or wall-based metrics) are left for future work, the results indicate that the proposed surrogate SR approach can streamline FEM-based CFD workflows toward near real-time hemodynamic analysis across morphologically similar vascular models.Keywords
Finite element method (FEM)–based CFD simulations play a central role in quantifying hemodynamics in vascular diseases such as cerebral aneurysms [1–4]. Clinically meaningful predictions often require highly refined meshes to resolve complex intra-aneurysmal flow structures, resulting in substantial computational cost [5–7]. This bottleneck becomes more pronounced in patient-specific pipelines, where multiple geometries and operating conditions may need to be evaluated within a limited turnaround time. Conventional acceleration strategies mitigate the cost–accuracy trade-off only partially. Adaptive mesh refinement [8–10] increases local resolution but depends on nontrivial error estimation and still expands the degrees of freedom. Parallel solvers [11–13] reduce wall-clock time, yet their benefit hinges on high-performance computing resources that are not always available in routine clinical or industrial workflows. Reduced-order modeling [14–17] can provide attractive speed-ups, but may smooth out fine-scale structures that are important for faithfully representing complex flow features. These limitations motivate alternative approaches that preserve high-fidelity information while remaining computationally and operationally efficient. Recent progress in deep learning (DL) has demonstrated strong potential for super-resolution (SR) reconstruction in fluid flows by learning to recover fine-scale information from coarse solutions [18–20]. In this context, coupling DL-based SR with physics-consistent FEM simulations provides a practical data-driven surrogate strategy: low-resolution (LR) FEM solutions supply boundary- and physics-consistent inputs, while the neural network approximates high-resolution (HR) flow features at a fraction of the cost. Such a surrogate SR paradigm offers a pathway to accelerate FEM-based CFD without relying solely on mesh refinement or large-scale computing infrastructure.
Super-resolution (SR) was originally developed in the computer vision community to recover fine-scale information from low-resolution image data. Convolutional neural networks (CNNs) [21]—including SRCNN [22], VDSR [23], and ESRGAN [24], as well as more recent transformer-based models [25,26], demonstrated strong performance in reconstructing high-frequency image details. However, these approaches are primarily tailored for RGB image data and do not explicitly incorporate the structure of scientific variables such as velocity or pressure fields. In fluid mechanics, physics-aware machine learning frameworks have therefore been introduced, including physics-informed neural networks (PINNs) [27,28] and operator-learning approaches [29], which embed governing equations into the learning process. While promising, these methods often require retraining across geometries or boundary conditions and can remain computationally demanding for high-resolution CFD on complex domains. In parallel, SR networks have been adapted for flow reconstruction by learning mappings between low- and high-resolution flow fields; representative examples include FlowSRNet [20] for turbulent flow reconstruction and the encoder–decoder framework proposed by Sofos and Drikakis [30] for turbulence and shock–boundary layer interactions. In biomedical applications, Habibi et al. [31] combined reduced-order modeling with multi-fidelity CFD to improve near-wall reconstruction, although super-resolution was not the primary focus of their work. Beyond fluids, SR concepts have been explored in computational mechanics, where deep learning surrogates have accelerated finite element simulations in additive manufacturing [32] and U-Net–based architectures have been applied to stress-field reconstruction under physical constraints [33]. Related studies, including PhySRNet [34] and CNN-based stress predictors [35], further support the feasibility of integrating SR with finite element method (FEM). Recent reviews on machine learning for CFD [36] and data-driven cardiovascular flow modeling [37] underscore the need for reliable and efficient surrogate strategies in hemodynamic applications. Despite these advances, SR workflows specifically tailored to finite element CFD remain limited, particularly when unstructured meshes must be interfaced with structured network inputs and when efficient reconstruction is required only in localized flow-critical regions.
The literature highlights two critical gaps for FEM-based CFD. First, many SR methods developed for images are not directly transferable to FEM outputs, as they are optimized for three-channel RGB data rather than physical quantities represented on unstructured meshes. Second, although SR has been applied in fluid and engineering contexts (e.g., turbulence reconstruction), dedicated studies on aneurysm hemodynamics remain limited, where irregular vascular geometries and strong near-wall gradients introduce additional challenges. PINNs provide another promising direction, but adapting them to complex vascular geometries is nontrivial and their training can become prohibitively expensive for high-resolution CFD in time-constrained settings. These considerations motivate a pragmatic perspective: a network can be pre-trained on FEM simulations of representative geometries and subsequently fine-tuned for new patient-specific morphologies. This data-driven surrogate strategy leverages the strengths of FEM—robust handling of complex geometries and boundary conditions—while alleviating the computational burden of repeated HR analyses, especially when the target cases share broad morphological similarity.
Cerebral aneurysm modeling provides a representative and practically relevant testbed for accelerated FEM-based CFD, where timely, high-fidelity hemodynamic information is valuable for downstream clinical interpretation. This study focuses on SR reconstruction of the velocity magnitude field from LR FEM solutions. Although clinical endpoints and wall-based metrics are beyond the scope of the present work, this choice provides a well-defined scalar target for systematic evaluation of surrogate SR performance in complex patient-specific geometries.
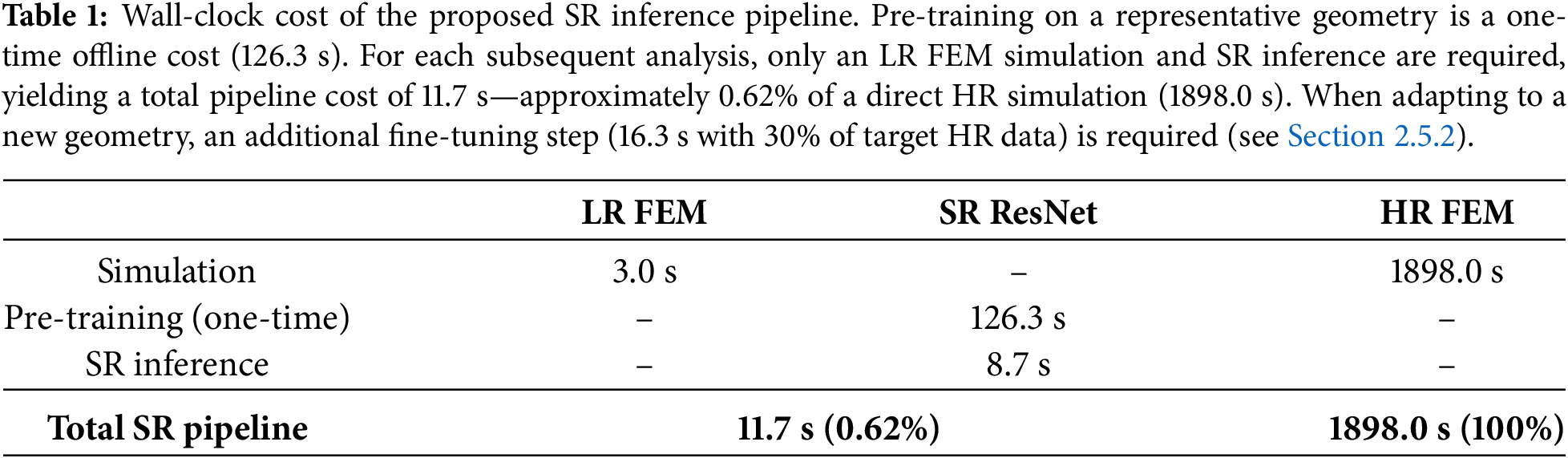
This study addresses these gaps by developing and validating a ResNet-based SR data-driven surrogate specifically adapted for FEM-based CFD. The framework is not intended as an end-to-end solver replacement, but rather as a data-driven enrichment module that complements conventional FEM solvers by recovering HR field detail from physics-consistent LR simulations. High-resolution data are required offline for pre-training on a representative geometry, while adaptation to new geometries primarily relies on a low-resolution FEM simulation, with only limited target-specific HR data employed for lightweight fine-tuning. Based on the current literature, this study is among the first to integrate a ResNet-based SR surrogate with FEM-based CFD for aneurysm hemodynamics through an end-to-end VTK probing pipeline and ROI-aware fine-tuning on unstructured meshes. By bridging the mismatch between unstructured FEM outputs and structured ResNet inputs, and by selectively refining flow-critical regions, the method achieves high-fidelity reconstructions at a fraction of the computational cost. In particular, the per-analysis cost of HR reconstruction—comprising LR simulation and SR inference—is approximately 0.62% of a direct HR simulation (Table 1), while adaptation to new geometries requires only lightweight fine-tuning with limited target-specific HR data. The proposed surrogate enables rapid patient-specific analyses that can support time-sensitive workflows. More broadly, the present work contributes to computational modeling by demonstrating how SR surrogates can enhance FEM-based CFD through transfer learning, efficient sampling, and practical workflow integration.
The remainder of this paper is organized as follows. Section 2 details the methodology, including the overall framework, dataset generation, workflow integration, network architecture, and training strategy. Section 3 presents the performance evaluation, covering cross-domain inference, point-wise error analysis, and computational efficiency. Section 4 discusses the broader implications and limitations of the proposed framework. Finally, Section 5 concludes with a summary of the key findings and outlines directions for future work.
The proposed approach integrates FEM-based CFD simulations with a deep learning surrogate for super-resolution (SR). Independent simulations were conducted on coarse and fine meshes using the open-source finite element library FEniCS [38,39], producing low-resolution (LR) and high-resolution (HR) data, respectively. All outputs were written in VTK (Visualization Toolkit) format [40], ensuring compatibility with ParaView [41,42] for visualization and post-processing. The overall workflow is summarized in Fig. 1.
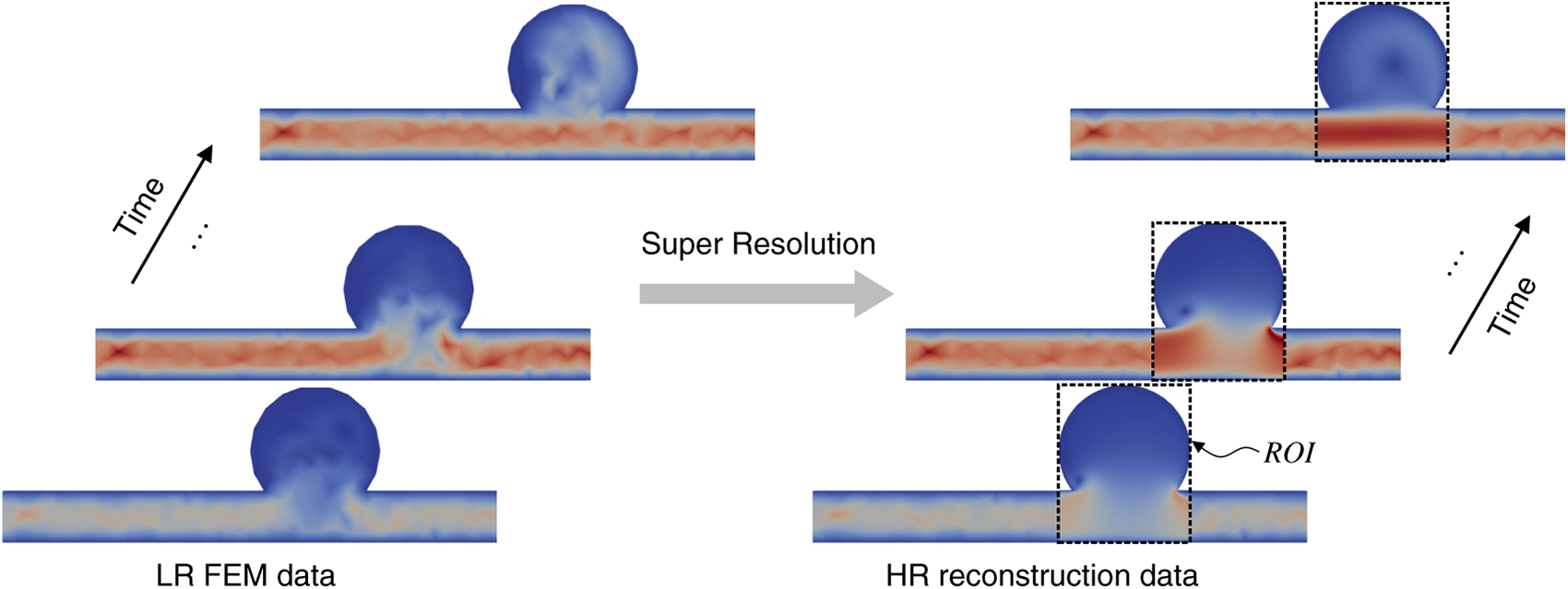
Figure 1: Overview of the data-driven super-resolution (SR) surrogate for accelerated high-fidelity flow reconstruction. Low-resolution (LR) CFD fields from coarse FEM simulation are mapped to a structured grid and super-resolved by a ResNet model to recover high-resolution (HR) fields within regions of interest (ROI). Because the LR FEM solution satisfies the prescribed governing equations and boundary conditions at the discrete level (e.g., inlet waveform and wall conditions), the surrogate operates on physics-consistent inputs while substantially reducing computational cost.
This study adopts a residual network (ResNet)—a convolutional neural network (CNN) architecture that employs skip (residual) connections to stabilize training and enable deeper models [43,44] (Fig. 2). In classical image and pattern recognition, data are sampled on a structured pixel grid where discrete points are uniformly distributed in the horizontal and vertical directions. Analogously, auxiliary Cartesian grids were constructed to span the computational domain, with locally refined resolution in regions of interest (ROI), such as the aneurysm bulge (Fig. 3). Since Cartesian grid nodes generally do not coincide with FEM mesh nodes, linear interpolation was applied to project the FEM solution (velocity magnitude) onto the grid points. This probing step produced paired LR–HR fields aligned on the same structured grid (i.e., velocity-magnitude values defined at identical nodes). Such alignment enabled one-to-one supervision during ResNet training while preserving the flexibility of the original unstructured meshes. The ResNet, implemented in PyTorch [45], was trained to map LR velocity-magnitude inputs to their HR counterparts. The architecture is illustrated in Figs. 4 and 5, and the PyTorch implementation is provided in Appendix A.
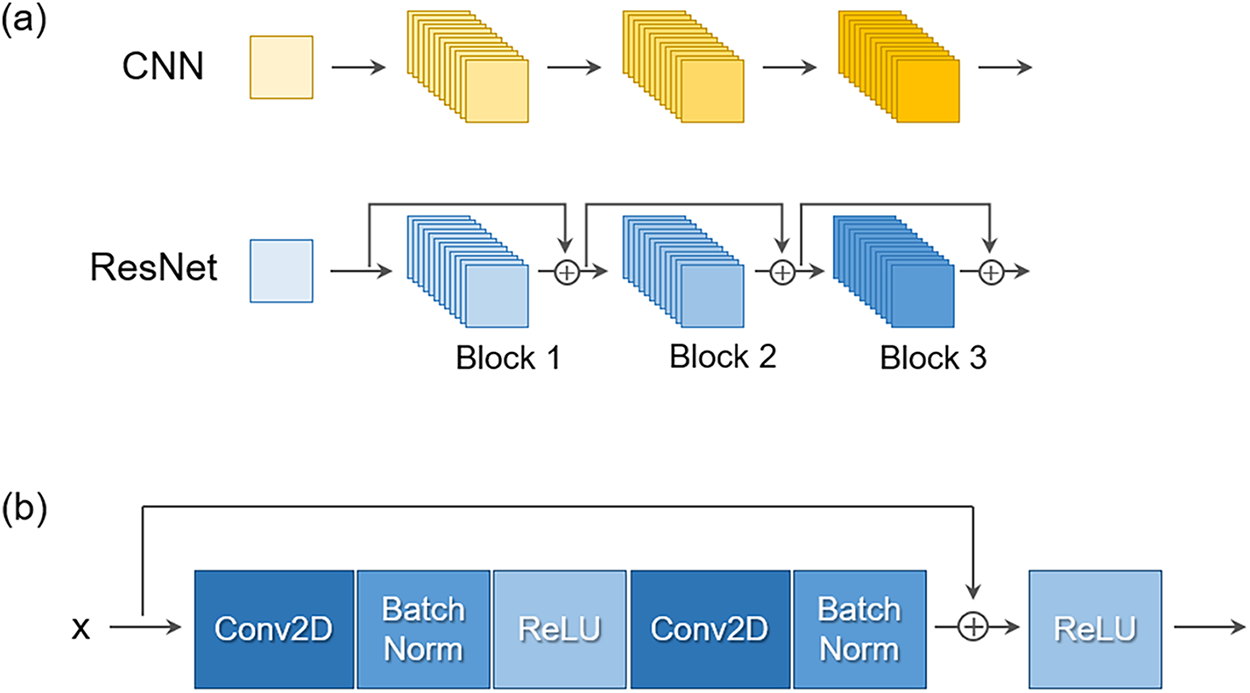
Figure 2: (a) Conventional CNN (top) vs. ResNet (bottom). (b) Structure of a residual block. Skip connections stabilize training and improve accuracy, making ResNet suitable for CFD super-resolution.
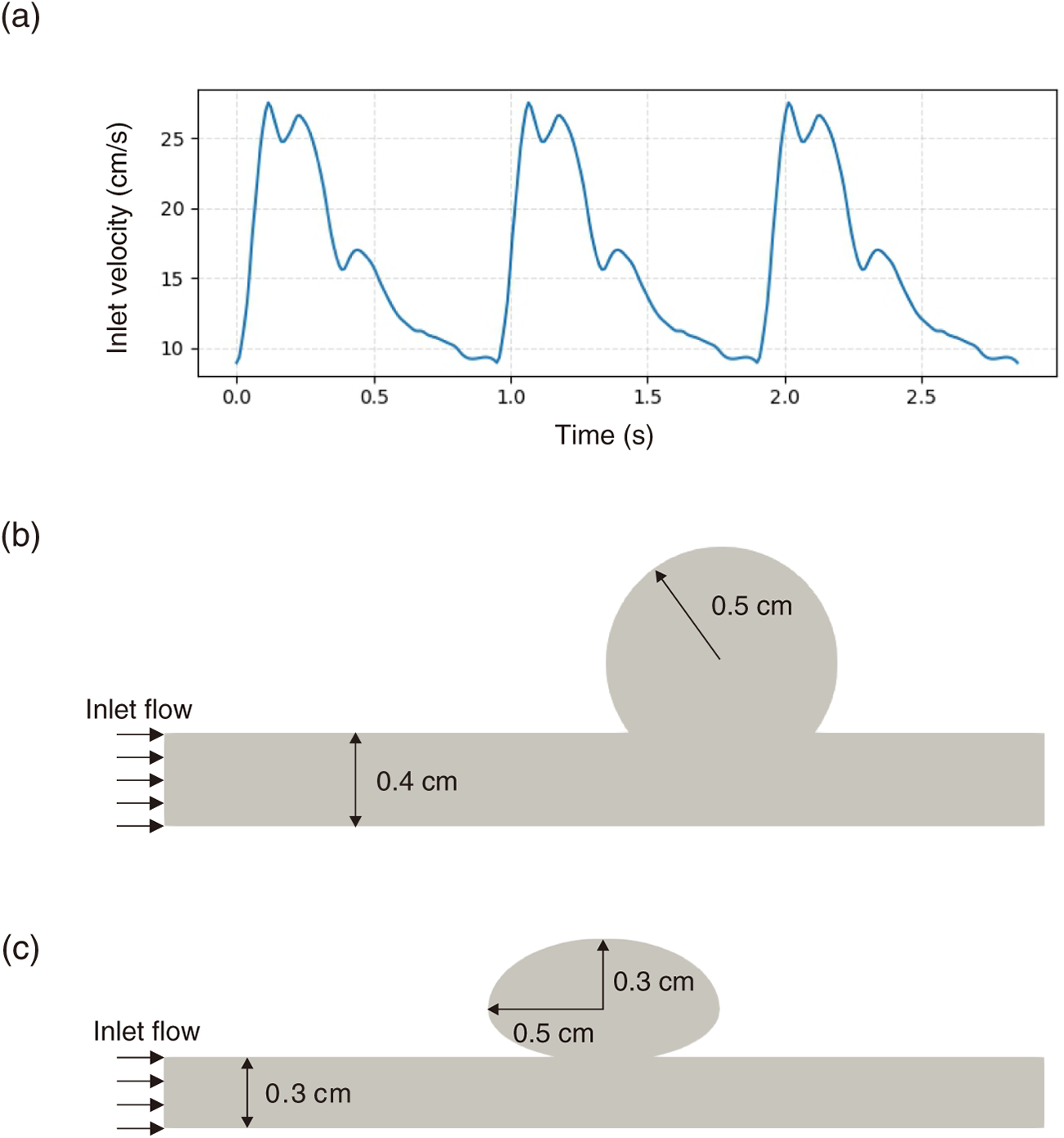
Figure 3: (a) Physiological pulsatile waveform prescribed as the inlet velocity boundary condition. (b) Baseline aneurysm domain employed for pre-training the super-resolution surrogate. (c) Variant aneurysm geometry used for cross-case validation to evaluate the generalization capability of the trained network.
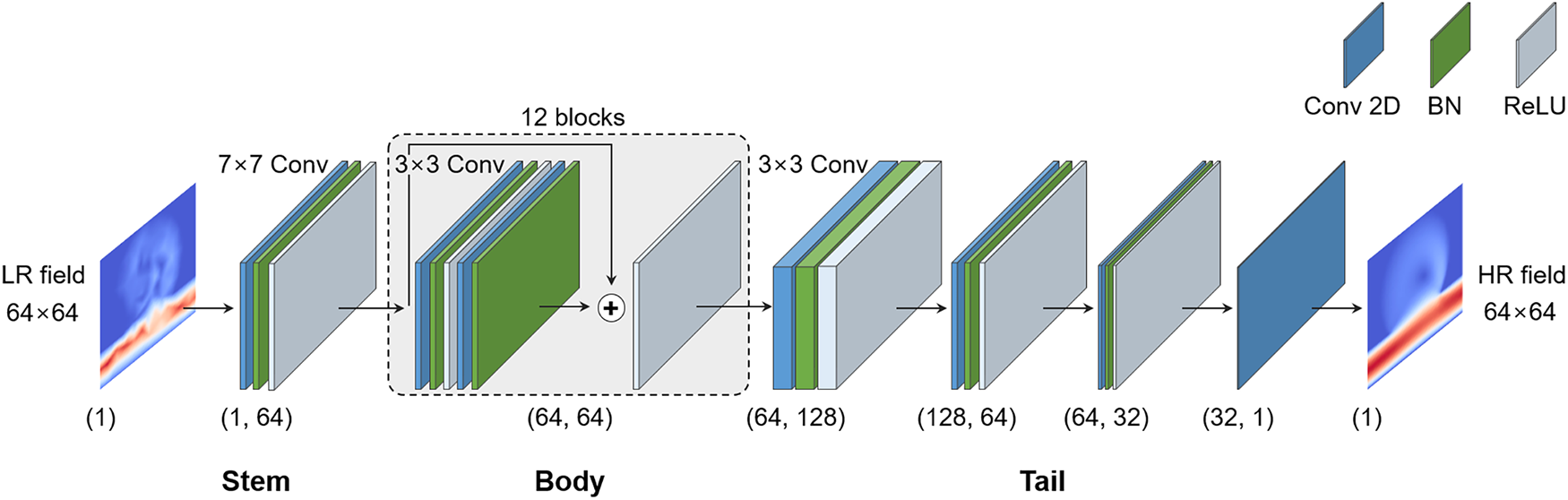
Figure 4: Architecture of the proposed ResNet-based super-resolution network. The pipeline consists of a stem layer (7
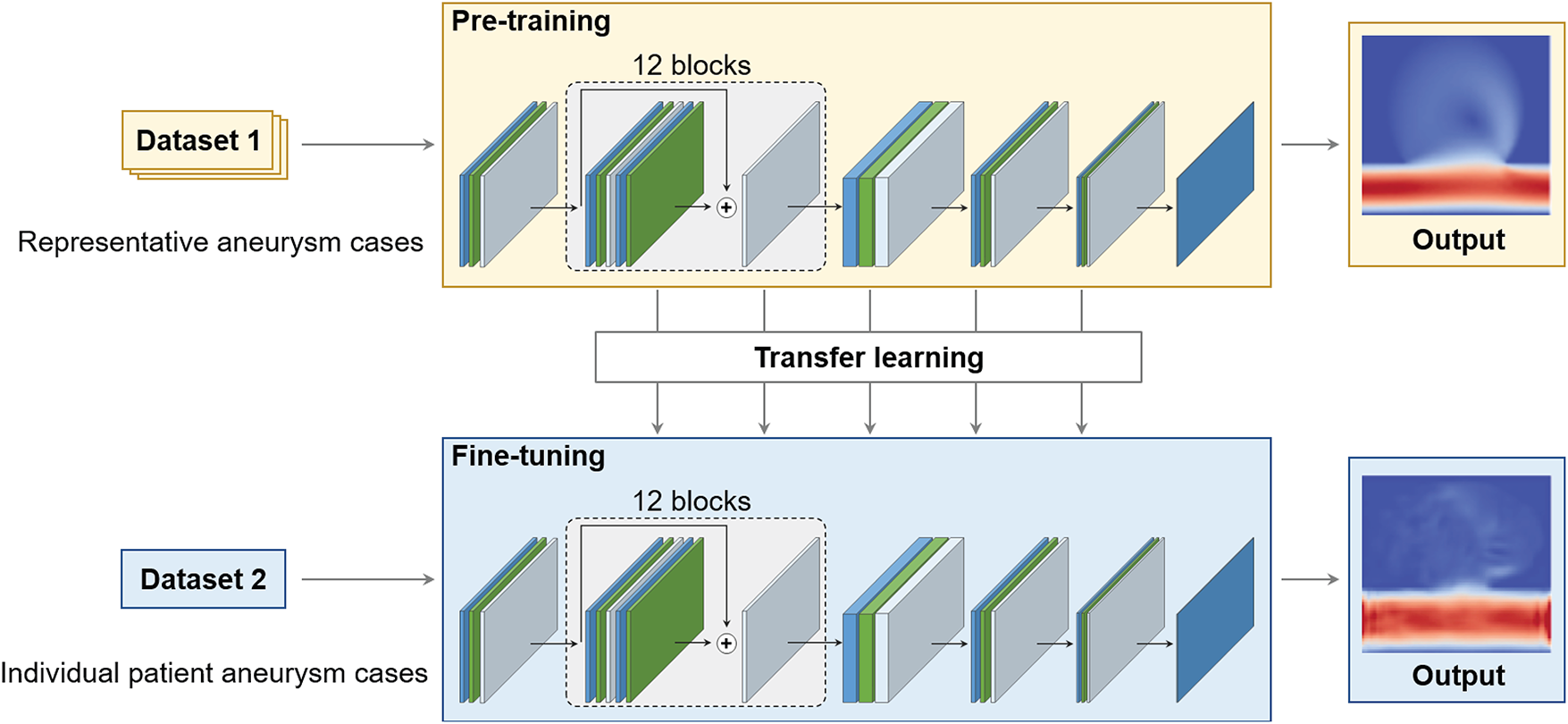
Figure 5: Pre-training (top) on representative aneurysm cases using Dataset 1, followed by fine-tuning (bottom) on an individual patient-specific case using Dataset 2. Transfer learning adapts the ResNet-based SR surrogate to accommodate geometric variations in aneurysm hemodynamics.
LR–HR pairs were first generated via FEM for a representative geometry (Fig. 3b) over three cardiac cycles (Fig. 3a) and used to pre-train the ResNet (Fig. 6). Next, LR data were produced for a variant geometry (Fig. 3c), and the pre-trained ResNet was fine-tuned to accommodate the geometric changes before predicting HR fields for the variant case. To reduce computational cost, fine-tuning was restricted to the ROI (Fig. 7a). The fine-tuned ResNet then predicted the HR fields (Fig. 7b). For validation, an HR FEM simulation was also performed for the variant geometry (Fig. 7c), and the SR predictions were quantitatively compared against this reference through error analysis (Figs. 8–10).
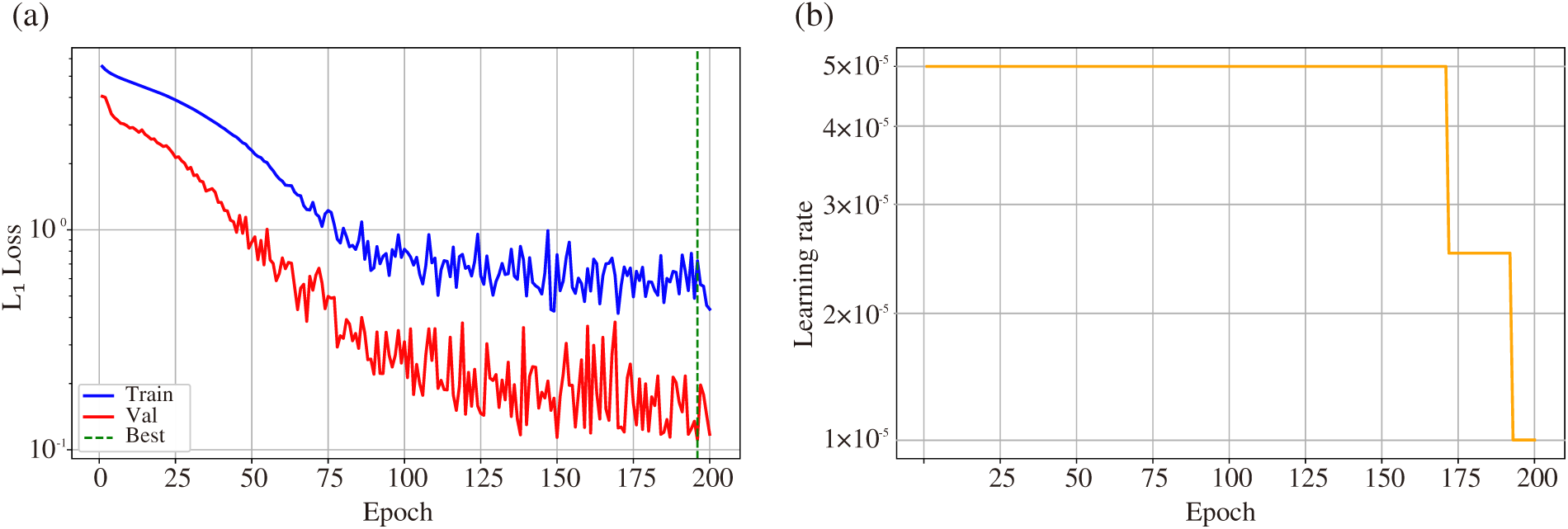
Figure 6: (a) Training (blue) and validation (red)
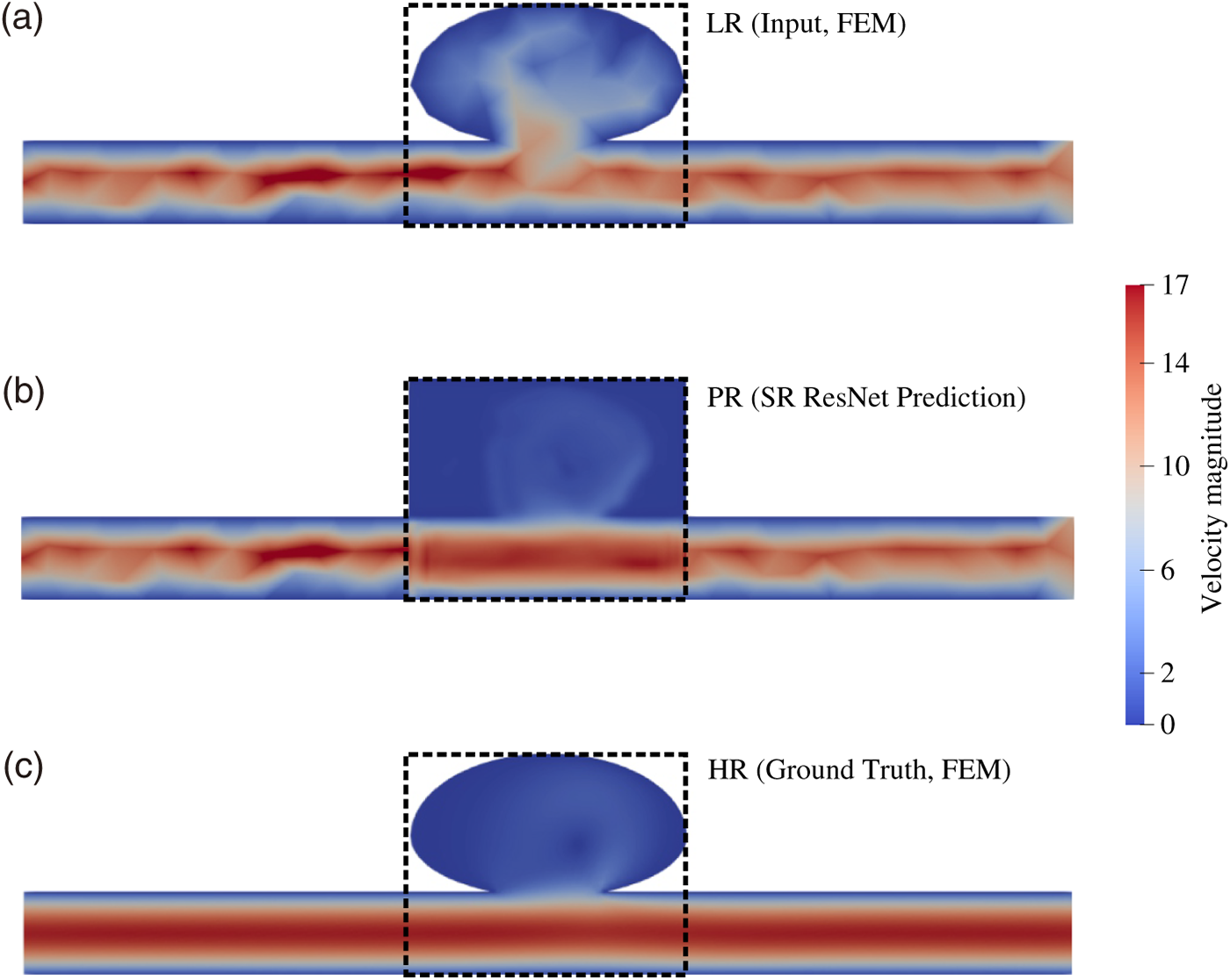
Figure 7: ROI-aware fine-tuning and inference. (a) Low-resolution (LR) input obtained from coarse FEM. (b) ResNet-based super-resolution (SR) prediction (PR) evaluated only within the dotted ROI to reduce cost. (c) High-resolution (HR) FEM reference.
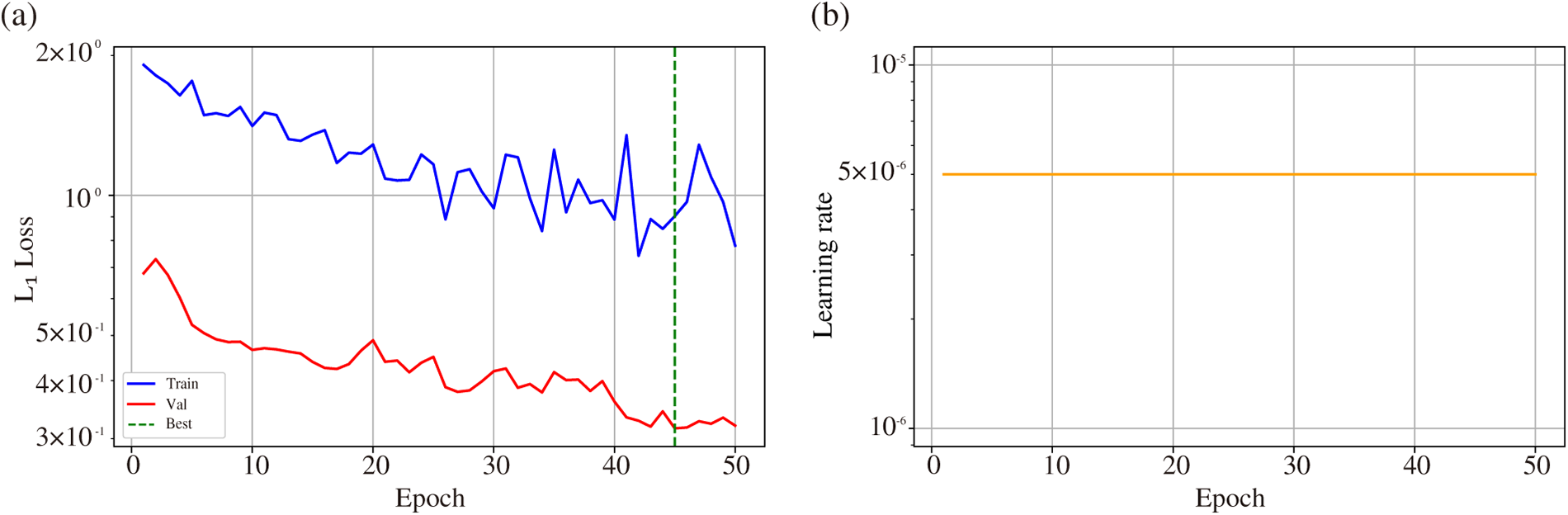
Figure 8: (a) Training (blue) and validation (red)
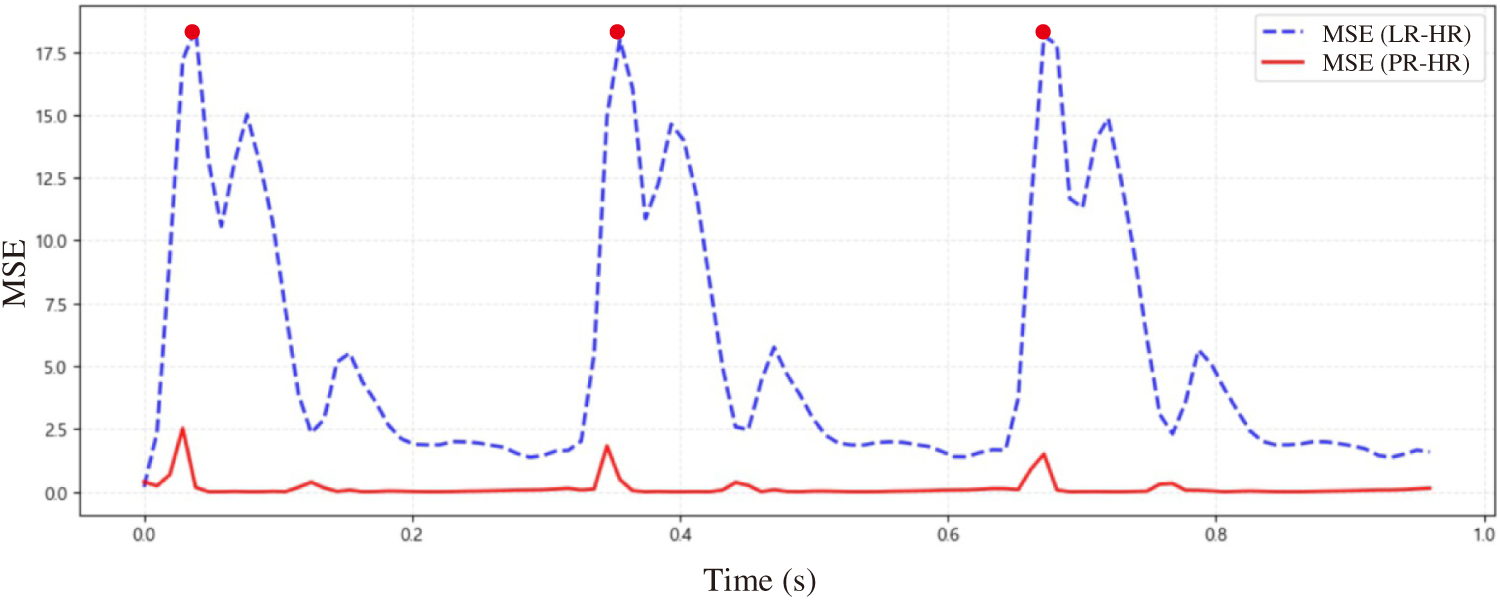
Figure 9: Temporal evolution of mean-squared error (MSE) at the aneurysm-center probe. The LR vs. HR curve (blue dashed) peaks near systole and remains elevated relative to diastole, whereas the ResNet SR prediction vs. HR (red solid) markedly suppresses error—achieving up to 96% reduction.
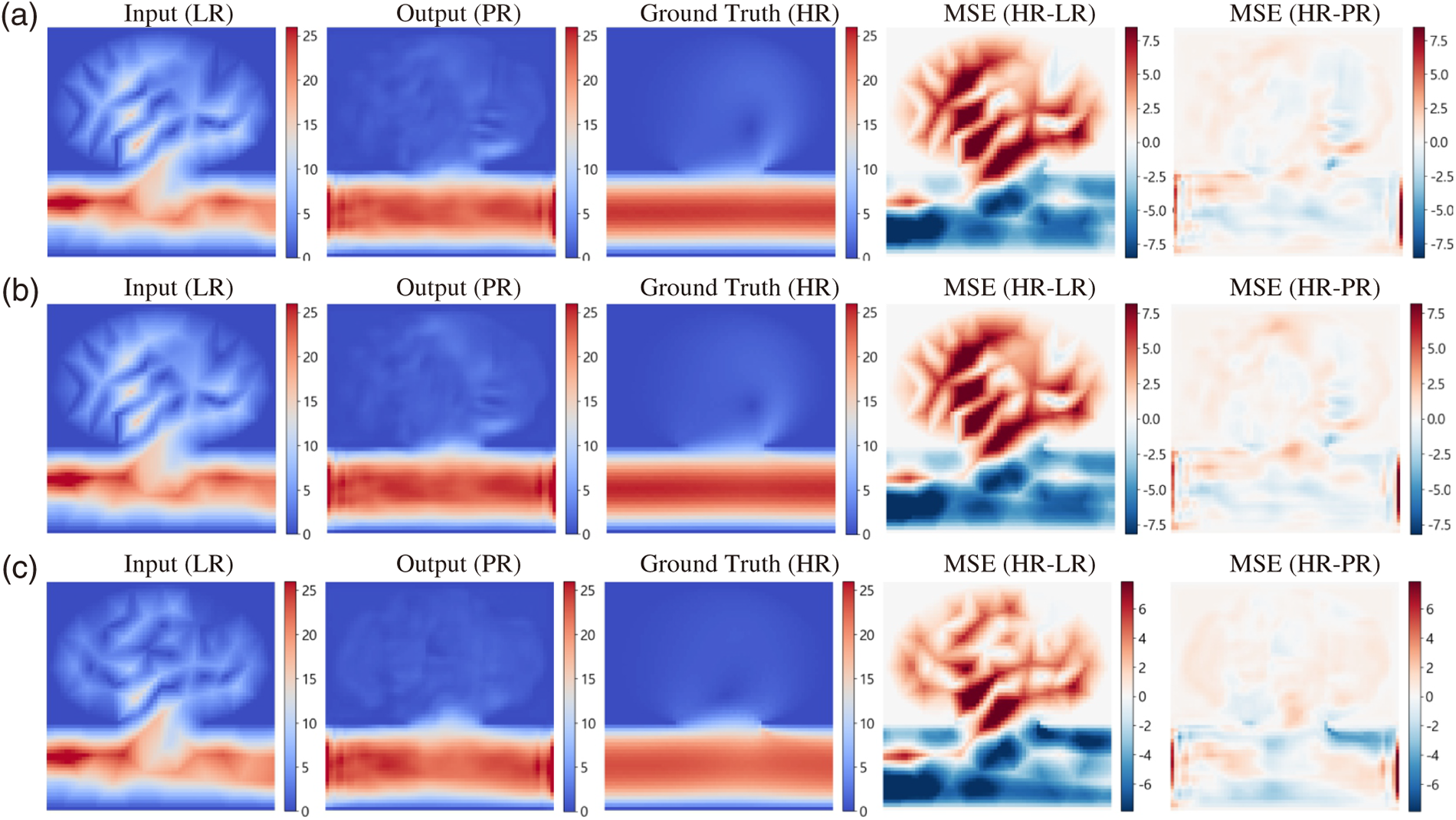
Figure 10: Cross-domain inference on the variant aneurysm geometry. Rows (a–c) correspond to three representative timesteps selected from the cardiac cycle (see red dots in Fig. 9). Columns show, left to right: low-resolution input (LR), ResNet super-resolution output (PR), high-resolution FEM reference (HR), and the mean-squared-error (MSE) maps comparing HR vs. LR and HR vs. PR. The SR prediction markedly suppresses error relative to the LR input.
2.2 Dataset Generation via FEM-Based CFD
The dataset was constructed from FEM-based CFD simulations of pulsatile blood flow in idealized aneurysm models (e.g., representative intracranial aneurysm configurations in the CFD literature) [2,6,31]. Two scenarios were considered: (i) a representative geometry for pre-training (Fig. 3b) and (ii) a variant geometry used to test the proposed framework (Fig. 3c). The HR mesh comprised 20,974 elements and 10,848 nodes, enabling the resolution of fine-scale shear layers and intra-aneurysmal vortices (wall-clock time: 1898.0 s). In contrast, the LR mesh contained 374 elements and 232 nodes, deliberately chosen as a strongly under-resolved setting that inevitably smooths fine-scale structures (wall-clock time: 3.0 s); the goal is not to obtain a mesh-converged LR solution, but to evaluate whether the pre-trained SR surrogate can recover flow features lost at coarse resolution. For consistency, the mesh densities (element and node counts) were kept identical across the two geometries, and the wall-clock times were comparable between corresponding HR and LR simulations for each case.
Both HR and LR simulations solved the unsteady incompressible Navier–Stokes equations:
where the blood density was set to
The total simulated time was
2.3 VTK Pipeline: From Unstructured FEM Meshes to ResNet-Ready Grids via Probe
The proposed workflow was integrated into a VTK-based pipeline [40], which is standard practice in CFD post-processing but is here repurposed as a data interface to bridge unstructured FEM outputs with the structured-grid input required by the deep-learning SR surrogate; to the best of the authors’ knowledge, this systematic integration has not been previously reported in the context of FEM–DL integration for flow-field super-resolution. LR and HR FEM simulations were performed on unstructured triangular meshes, and the resulting velocity fields were exported in VTK format. For pre-training, a
Following SR inference, the reconstructed HR fields were written back to VTK files, enabling direct visualization and comparison in ParaView (see Fig. 7b). Because the SR predictions and the HR reference are aligned on the same structured grid, quantitative training, fine-tuning, and post-processing can be carried out in a straightforward point-wise manner.
2.4 Neural Network Architecture
A ResNet [43,44] tailored for scalar CFD fields is adopted. Fig. 2a shows the difference between a conventional CNN and ResNet. ResNet’s skip connections stabilize training in deeper networks, enabling more accurate recovery of fine-scale flow features. Hence, ResNet was chosen as the backbone of the proposed SR framework. The basic residual block structure is illustrated in Fig. 2b. Building upon this design, the network extends ResNet into a three-stage stem–body–tail architecture, as shown in Fig. 4.
The stem employs a
The tail stage refines features and reduces the dimensionality back to the output channel through successive
Both LR and HR velocity-magnitude fields (single channel) were normalized using a fixed factor (=1.0) prior to training. The SR-ResNet was implemented in PyTorch [45] and optimized with AdamW (
Fig. 6a illustrates training dynamics: the
To prevent instability from large updates, gradient clipping with a maximum norm of 1.0 was applied. The loss function employed was the
2.5.2 Fine-Tuning Strategy (ROI-Aware)
To adapt the pre-trained SR-ResNet to patient-specific geometries, the network was fine-tuned from pre-training checkpoints (Fig. 5). Inputs were normalized identically to pre-training. The loss (L1Loss), optimizer (AdamW), and architecture were kept consistent to ensure a comparable optimization landscape. Fine-tuning was performed only within the ROI to reduce computational cost and concentrate learning on flow-critical areas. The protocol was:
• Initialization: start from pre-trained checkpoints.
• Reduced learning rate:
• Conservative training: fixed learning rate with 50 epochs and smaller batch sizes (minimum 4).
• Gradient stabilization: gradient clipping with maximum norm 0.5.
• Data efficiency: only 30% of the paired LR–HR variant-geometry data were used for fine-tuning.
As shown in Fig. 8a, the validation
Cross-domain generalization was evaluated by testing on a distinct aneurysm geometry unseen during pre-training. Performance was assessed by comparing SR predictions with HR reference fields on the target geometry (Fig. 7). Quantitative evaluation was performed at every saved timestep across three cardiac cycles; SR inference was conducted for all 101 temporal snapshots (Fig. 9). Improvement was quantified using an MSE-based metric:
where
Fig. 9 shows that the proposed framework improves LR data across all timesteps. The temporal profile of MSE(LR–HR) mirrors the inlet waveform: errors rise during systolic acceleration and peak near maximum flow speed (red markers), then decrease during diastole. MSE(PR–HR) exhibits a similar trend, indicating that SR performance remains partly bounded by the quality of the LR simulation. Nevertheless, the error magnitude is dramatically reduced—achieving up to 96% improvement of MSE(PR–HR) over MSE(LR–HR)—which supports the effectiveness of the pre-training plus ROI-aware fine-tuning strategy.
The unified-grid strategy places the LR (input), PR (SR prediction), and HR (reference) fields on the same Cartesian lattice, enabling strict point-wise evaluation. Fig. 10 examines three representative timesteps: columns show LR input, PR output, HR reference, and corresponding MSE maps comparing HR vs. LR and HR vs. PR. Across all snapshots, the PR fields suppress localized errors relative to LR—particularly within the aneurysm sac and along near-wall shear layers—while preserving the vessel boundaries present in the HR solution.
A point at the center of the aneurysm was further investigated to assess temporal fidelity. Fig. 11 plots the velocity magnitude time series for LR (input), PR (SR prediction), and HR (reference) over three cardiac cycles. While LR follows the overall waveform trend, it underestimates HR during both systolic peaks and diastolic levels. In contrast, the surrogate brings PR into close agreement with HR across the cycle, demonstrating improved time-resolved reconstruction.
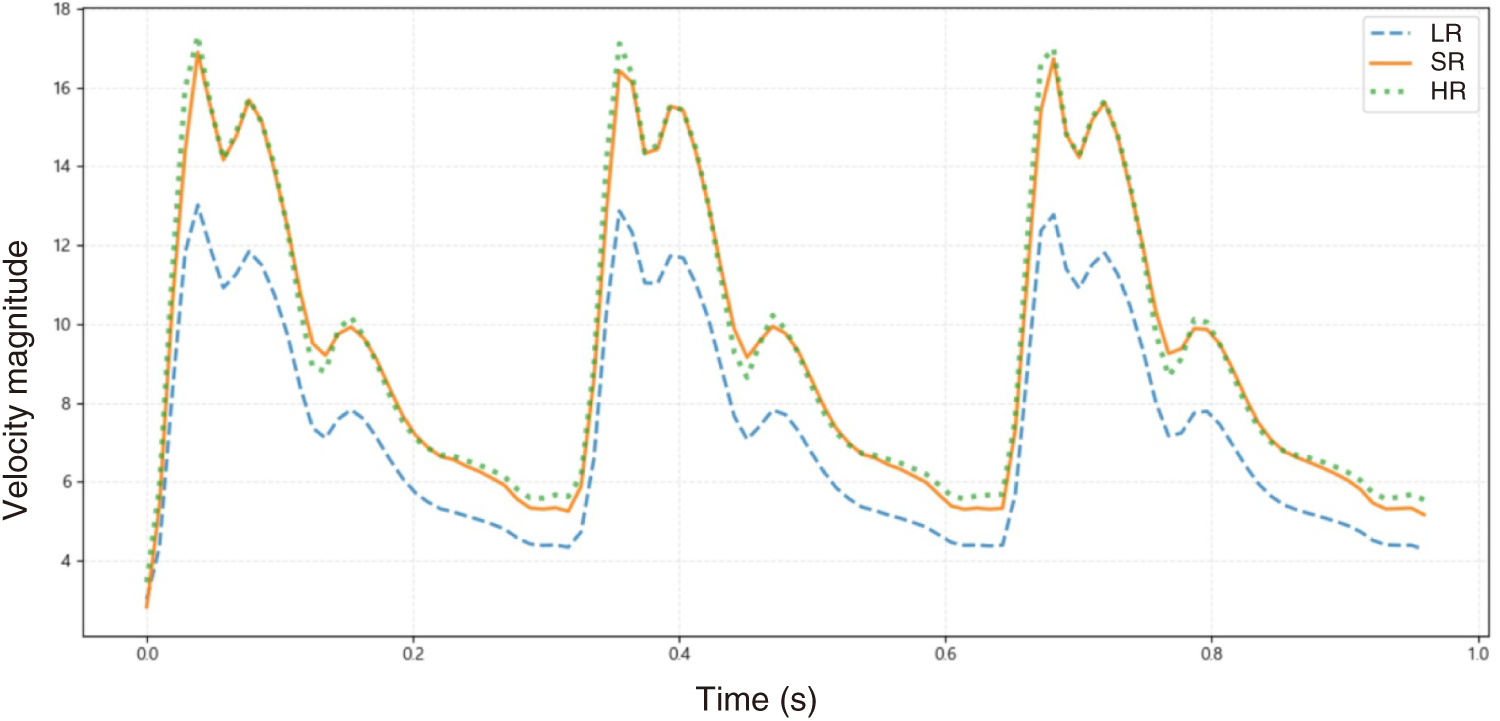
Figure 11: Temporal profile of velocity magnitude at the center of the aneurysm domain. The low-resolution (LR, blue dashed) signal underestimates both systolic peaks and diastolic levels, whereas the ResNet super-resolution (SR, red solid) closely tracks the high-resolution (HR, green dotted) reference. This indicates that the surrogate recovers both amplitude and phase characteristics of the unsteady pulsatile flow.
As reported in Table 1, the pre-trained SR surrogate produces HR predictions from a single LR FEM simulation (3.0 s) and SR inference (8.7 s), yielding a per-analysis pipeline cost of 11.7 s—approximately 0.62% of a direct HR FEM simulation (1898.0 s on the variant geometry, Fig. 3c). When the target geometry differs from the pre-training geometry (Fig. 3b), a one-time fine-tuning step is additionally required: paired LR–HR simulation data must be generated for the new geometry, of which 30% is used for fine-tuning (16.3 s). This raises the initial cost for a new geometry to 28.0 s (1.48% of HR); however, because fine-tuning is performed only once per geometry, all subsequent analyses on that geometry—such as parameter studies or variations in boundary conditions—require only LR simulation and SR inference (11.7 s, 0.62% of HR) with no additional HR simulation. The fine-tuning cost is thus amortized over repeated use, making the effective sper-analysis cost approximately 0.62% of the direct HR simulation. A summary is provided in Table 2. For point-wise comparison on a
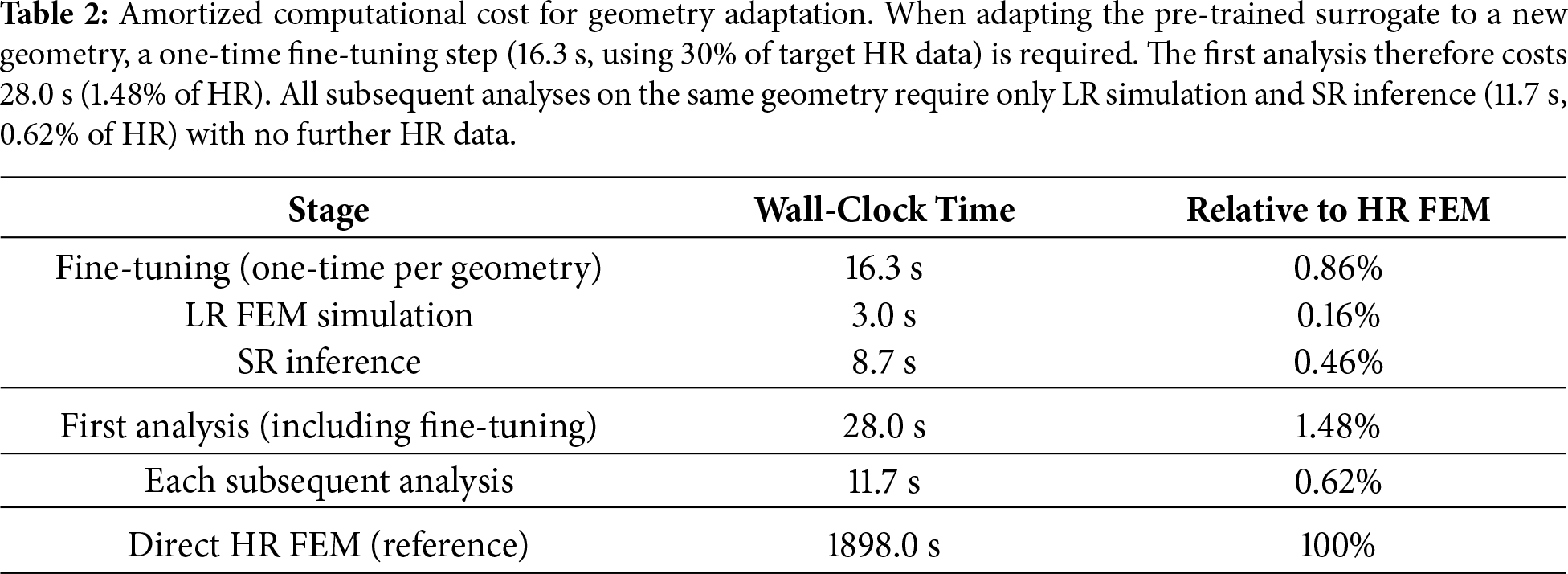
This study was motivated by a practical modeling question: can a deep learning model trained on a representative aneurysm configuration be efficiently adapted to a new, patient-specific geometry so that high-resolution (HR) finite element method (FEM)-based CFD fields can be obtained without repeatedly performing HR simulations? Despite steady advances in numerical methods and computing, HR FEM-CFD remains resource-intensive in many routine workflows, limiting rapid turnaround when multiple geometries or parameter variations must be evaluated. The results show that a pre-training with fine-tuning strategy with a residual neural network (ResNet) provides an effective data-driven surrogate for super-resolution (SR) enhancement of FEM-based CFD. Cross-domain tests indicate that the surrogate can be transferred to a variant aneurysm geometry with consistent gains. Using coarse-mesh, low-resolution (LR) velocity-magnitude fields as inputs, the surrogate reconstructs HR fields that closely match the HR FEM solution used as the reference in this study, while requiring less than 1% of the HR computational cost per analysis. Notably, the per-analysis SR pipeline (LR simulation plus SR inference) accounts for approximately 0.62% of the HR runtime (Table 1). When geometry adaptation is required, a one-time fine-tuning step using limited HR data raises the initial cost to 1.48%, but this cost is amortized over subsequent analyses. In terms of accuracy, the ResNet SR surrogate reduces the velocity-field mean-squared error (MSE) by up to 96% relative to the LR baseline. Residual learning stabilizes optimization and expedites convergence, enabling recovery of vortical patterns and near-wall structures that are resolved in the HR reference but are attenuated on the coarse mesh. Overall, these findings support SR surrogates as a practical computational strategy for accelerating FEM-based fluid simulations when repeated HR analyses are prohibitive.
To generate time-resolved intracranial flow data, a three-cardiac-cycle inlet waveform (Fig. 3a) was prescribed and a representative aneurysm geometry (Fig. 3b) was defined. Using FEniCS [38,39], paired LR and HR CFD datasets were produced. A ResNet model implemented in PyTorch [45] was pre-trained on the representative geometry (Fig. 4) and subsequently fine-tuned to accommodate geometric variations (Fig. 5). To further assess transferability, a distinct aneurysm configuration was considered (Fig. 3c). For computational efficiency, adaptation and inference were restricted to a region of interest (ROI)—e.g., the aneurysm sac—mapped onto a unified Cartesian grid (Fig. 7); the pretrained surrogate was then fine-tuned on the target domain (Fig. 8). This ROI-focused design is a key practical component because it avoids full-domain HR recovery when only localized flow features are of interest.
Performance and generalization were evaluated using complementary diagnostics: temporal MSE within the ROI (Fig. 9), spatial error maps at representative time steps (Fig. 10), and a probe-based velocity waveform at the aneurysm center (Fig. 11). Collectively, these results indicate that the surrogate (i) reduces error relative to LR throughout the cardiac cycle, (ii) reconstructs shear layers and intra-aneurysmal structures captured by the HR reference, and (iii) preserves phase behavior in time-resolved signals. From a computational modeling perspective, an important distinction is that the SR surrogate operates on LR FEM solutions that already satisfy the prescribed boundary conditions and the discrete form of the governing equations; the SR stage therefore functions as a data-driven reconstruction step operating on physics-consistent inputs, rather than a standalone physics solver; the physical fidelity of the reconstructed output should accordingly be interpreted as approximate, not physics-constrained. This contrasts with approaches that must enforce physics during training (e.g., PINNs) [27,28] or generic image SR models primarily designed for RGB data [22,24]. The achieved speed-ups—under 1% of the HR wall-clock cost per analysis (Table 1)—underscore the utility of the proposed surrogate workflow for accelerating FEM-based CFD in practice.
Several limitations should be noted to properly contextualize these results. First, the present workflow is restricted to 2D slice-based reconstruction; while this choice improves computational efficiency and simplifies training, it does not explicitly enforce inter-slice coherence in the reconstructed 3D field. Extending the surrogate to volumetric 3D architectures or incorporating cross-slice consistency mechanisms (e.g., 2.5D inputs or continuity regularization) is an important next step. Second, the surrogate is trained in a supervised manner using paired LR–HR datasets, which requires HR simulations during the offline stage. This cost is amortized when the model is reused across related geometries, but it may become substantial if the target distribution shifts significantly. Third, because the imaging-based formulation treats the velocity magnitude as a single-channel image on a structured grid, directional (vector) information is inherently discarded; wall-based clinical metrics such as wall shear stress (WSS), which require the tangential gradient of the velocity vector at the wall, therefore cannot be recovered and constitute an important direction for future work involving full vector-field super-resolution. Finally, generalization boundaries remain to be quantified more systematically, including sensitivity to geometric variability, inlet waveform changes, and boundary-condition specifications. Cross-domain generalization was evaluated on two unseen sidewall aneurysm geometries that differ in morphology and size (see Section C), each fine-tuned using LR inputs with limited HR supervision (30% of available HR data). The present study is limited to sidewall-type aneurysms; extending the framework to bifurcation aneurysms—which exhibit fundamentally different flow topology—will require volumetric 3D SR architectures and constitutes an important direction for future work. Developing a geometry-universal SR surrogate that generalizes to arbitrary unseen vascular morphologies without any target-domain adaptation remains a challenging open problem; the present transfer-learning pipeline represents a practical step toward that goal.
Despite these limitations, the proposed framework provides practical insights for computational modeling. Many learning-based approaches for fluid simulation struggle with simultaneously handling complex geometries, respecting boundary conditions, and achieving affordable training and inference costs. The results support an alternative pathway that integrates conventional FEM solvers with neural-network-based enrichment: physics-consistent LR simulations supply robust inputs, and the SR surrogate approximates localized HR detail within flow-critical regions such as the aneurysm sac. Beyond biomedical CFD, the demonstrated efficiency suggests broader engineering relevance, including rapid design-space exploration, optimization loops, and lightweight digital-twin updates under constrained computing budgets.
This study demonstrates that coupling the finite element method (FEM) with deep learning can substantially accelerate CFD through a practical data-driven super-resolution (SR) surrogate while recovering fine-scale flow features in regions of interest. The proposed framework reconstructs high-resolution (HR) velocity-magnitude fields from low-resolution (LR) FEM inputs with a per-analysis cost of approximately 0.62% of a direct HR simulation (Table 1), and it remains transferable across geometries via lightweight fine-tuning with limited target-specific HR data, particularly when aneurysm morphologies share broad topological similarity. By operating on physics-consistent LR solutions and applying a pre-training with fine-tuning strategy, the ResNet-based SR surrogate functions as a data-driven enrichment module that complements—rather than replaces—conventional FEM solvers, enabling repeated analyses without repeated HR meshing and solving.
Future work will extend the workflow to volumetric 3D aneurysm models and improve cross-slice coherence in reconstructed fields. Additional directions include incorporating physics-based regularization terms to further strengthen physical plausibility, exploring alternative surrogate architectures (e.g., transformer-based or adversarial refiners), and quantifying uncertainty under broader variations in geometry and boundary conditions. Ultimately, patient-specific validation and multiphysics extensions, including fluid–structure interaction, will be important steps toward broader deployment of SR surrogates in computational mechanics and biomedical engineering.
Acknowledgement: Not applicable.
Funding Statement: This research was funded by the ‘Changwon National University–Samsung Changwon Hospital joint Collaboration Research Support Project’ in 2025. This study was conducted as part of the Glocal University Project, supported by the RISE (Regional Innovation System & Education) program funded by the Ministry of Education.
Author Contributions: Sojin Shin: Validation; Visualization; Writing—original draft; Writing—review & editing; Funding acquisition; Project administration; Resources; Supervision. Guk Heon Kim: Formal analysis; Investigation; Data curation. Seung Hwan Kim: Writing—review & editing; Funding acquisition; Project administration; Resources; Supervision. Jaemin Kim: Conceptualization; Methodology; Software; Formal analysis; Investigation; Data curation; Validation; Visualization; Writingoriginal draft; Writing—review & editing; Funding acquisition; Project administration; Resources; Supervision. All authors reviewed and approved the final version of the manuscript.
Availability of Data and Materials: Data available upon reasonable request.
Ethics Approval: Not applicable.
Conflicts of Interest: The authors declare no conflicts of interest.
Nomenclature
| Symbols | |
| Velocity vector field | |
| Pressure field | |
| Fluid density | |
| Dynamic viscosity | |
| Body force vector | |
| Simulation timestep | |
| Mean absolute error loss function | |
| Abbreviations | |
| CFD | Computational Fluid Dynamics |
| CNN | Convolutional Neural Network |
| DL | Deep Learning |
| FEM | Finite Element Method |
| HR/LR | High-Resolution/Low-Resolution |
| MSE | Mean Squared Error |
| PSNR | Peak Signal-to-Noise Ratio |
| PR | Prediction (super-resolution output) |
| ResNet | Residual Network |
| ROI | Region of Interest |
| SR | Super-Resolution |
| VTK | Visualization Toolkit |
| WSS | Wall Shear Stress |
Appendix A PyTorch Script for ResNet Architecture
Listing A1: Python implementation of the ResNet-based super-resolution surrogate model architecture.
import torch
import torch.nn as nn
class BasicBlock(nn.Module):
"""
Basic residual block: Channel and spatial resolution are preserved.
"""
def __init__(self, ch, dropout_rate=0.1):
super().__init__()
self.net = nn.Sequential(
nn.Conv2d(ch, ch, kernel_size=3, padding=1),
nn.BatchNorm2d(ch),
nn.ReLU(inplace=True),
nn.Dropout2d(dropout_rate),
nn.Conv2d(ch, ch, kernel_size=3, padding=1),
nn.BatchNorm2d(ch)
)
self.relu = nn.ReLU(inplace=True)
def forward(self, x):
return self.relu(x + self.net(x))
class ResNetSR(nn.Module):
"""
Shallow ResNet-based super-resolution network.
- Stem: 1 (in_ch) -> 64 channels
- Body: 12 Residual blocks
- Tail: 64 -> 128 -> 64 -> 32 -> 1 (out_ch)
"""
def __init__(self, in_ch, out_ch, nPt = 64, blocks = 12):
super().__init__()
# Stem
self.stem = nn.Sequential(
nn.Conv2d(in_ch, 64, kernel_size=7, padding=3),
nn.BatchNorm2d(64),
nn.ReLU(inplace=True)
)
# Residual body
self.body = nn.Sequential(*[BasicBlock(64) for _ in range(blocks)])
# Tail
self.tail = nn.Sequential(
nn.Conv2d(64, 128, kernel_size=3, padding=1),
nn.BatchNorm2d(128),
nn.ReLU(inplace=True),
nn.Conv2d(128, 64, kernel_size=3, padding=1),
nn.BatchNorm2d(64),
nn.ReLU(inplace=True),
nn.Conv2d(64, 32, kernel_size=3, padding=1),
nn.BatchNorm2d(32),
nn.ReLU(inplace=True),
nn.Conv2d(32, out_ch, kernel_size=3, padding=1)
)
self.nPt = nPt
self.out_ch = out_ch
def forward(self, x):
x = self.stem(x)
x = self.body(x)
y = self.tail(x) # (N, out_ch, nPt, nPt)
return y.view(y.size(0), -1) # always flatten
Appendix B Ablation Study: Effect of Residual Block Depth
To justify the choice of 12 residual blocks, an ablation study was conducted by training three network variants—with 6, 12, and 18 residual blocks—under identical conditions (L1 loss, AdamW optimizer, learning rate
All three variants converge to comparable loss ranges (Fig. A1), but the cost-efficiency decreases with depth: the 6
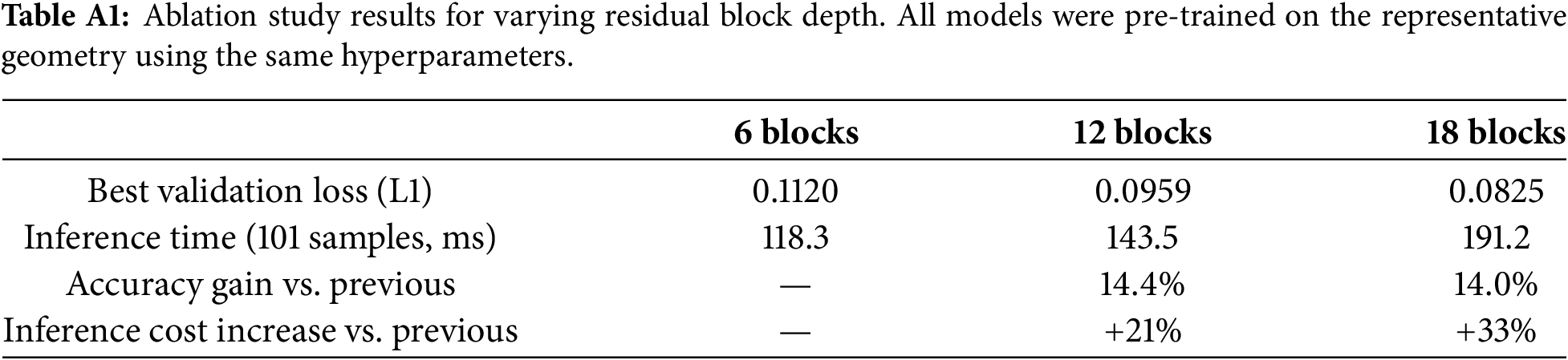
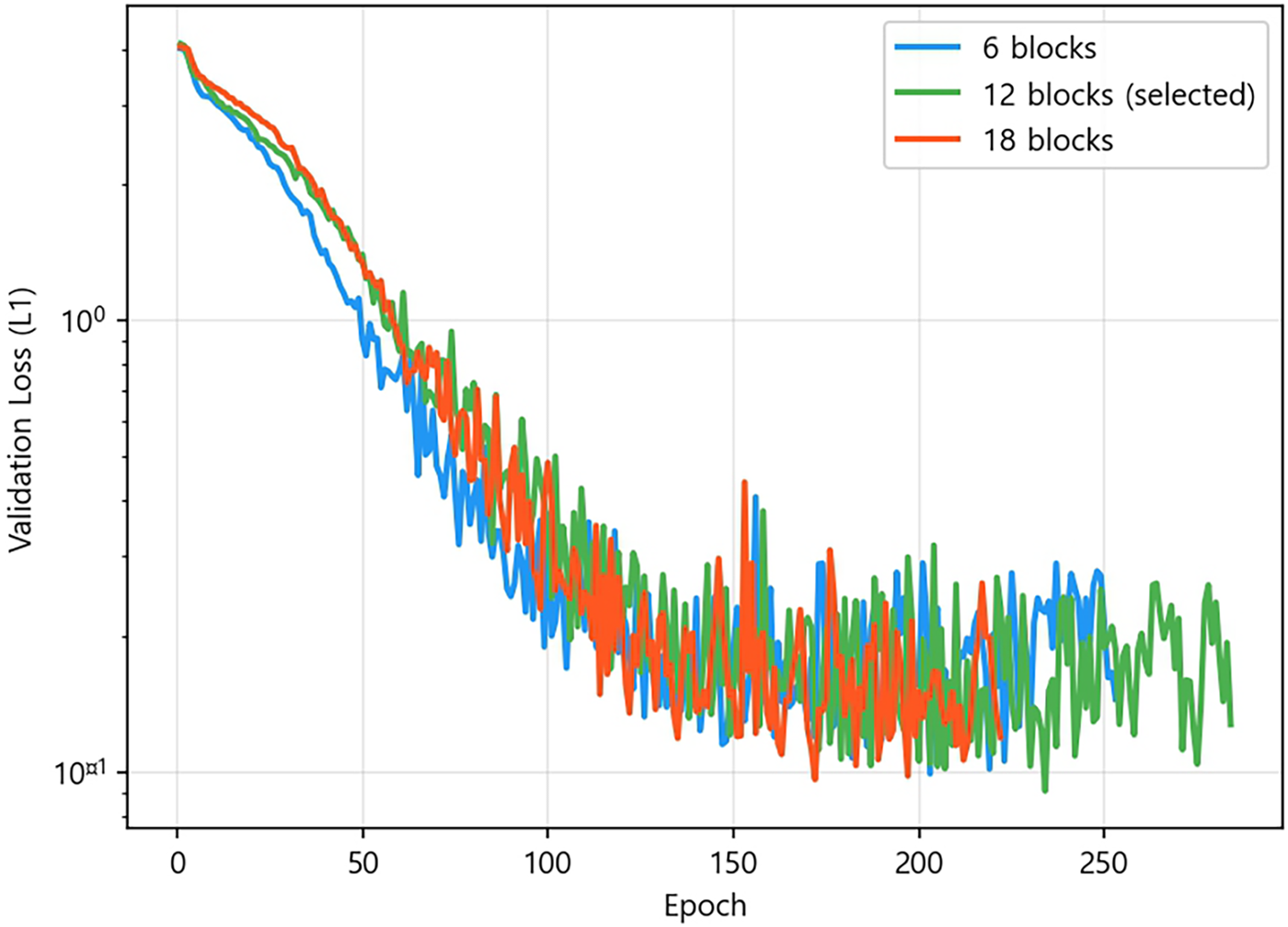
Figure A1: Validation loss (L1) vs. epoch for the 6-, 12-, and 18-block residual network variants during pre-training.
Appendix C Additional Cross-Domain Validation: Size Variation: To further assess the generalization capability of the proposed SR framework, the pre-trained model was fine-tuned and evaluated on a second unseen sidewall aneurysm geometry of different size, following the identical pipeline described in Section 2. While the main text evaluates transfer across morphological variation (different sac shape), this appendix examines transfer across size variation (small vs. large sidewall aneurysm). Fig. A2 compares the two sidewall aneurysm geometries, and Fig. A3 presents representative SR predictions alongside the HR reference and LR input.
The results confirm that the pre-training with fine-tuning strategy generalizes across size variation within the sidewall aneurysm category. Combined with the morphological-variation case presented in the main text, these two cross-domain evaluations indicate that the learned SR mapping is transferable across related vascular configurations. Extending the framework to bifurcation aneurysms—which exhibit fundamentally different 3D flow topology—will require volumetric SR architectures and is planned as future work.
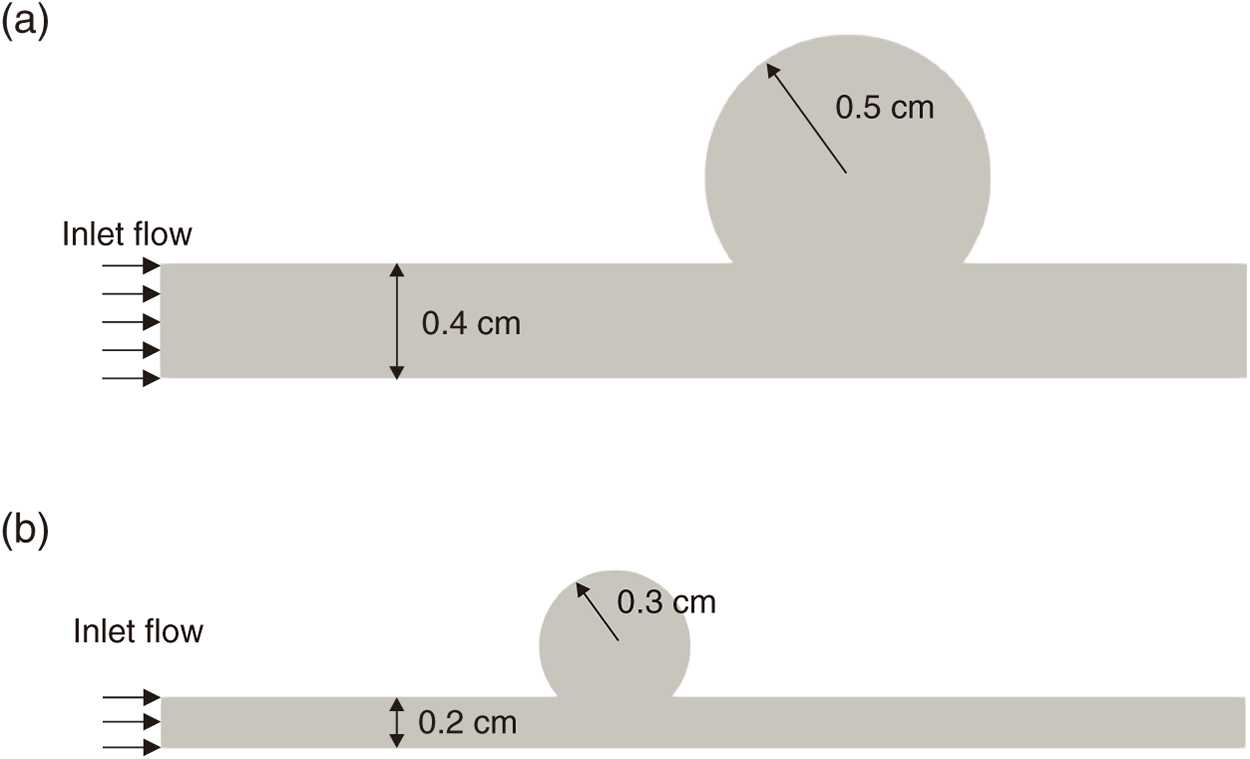
Figure A2: Sidewall aneurysm geometries used for cross-domain size-variation evaluation. (a) Large configuration: vessel diameter 0.4 cm, sac diameter 0.5 cm (used for pre-training in the main text). (b) Small configuration: vessel diameter 0.2 cm, sac diameter 0.3 cm (unseen target geometry for this appendix).
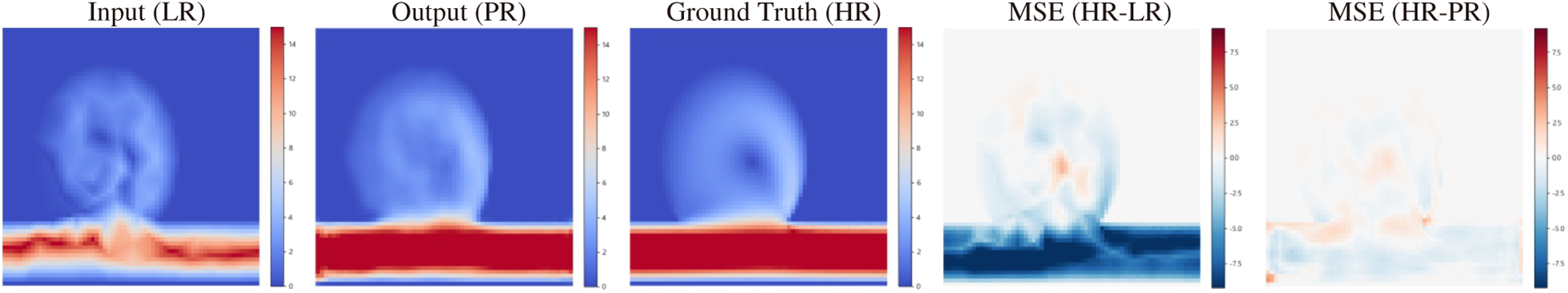
Figure A3: Cross-domain SR results for the size-variant sidewall aneurysm. Columns show, left to right: LR input, SR prediction, HR reference, and MSE error maps (HR vs. LR and HR vs. SR).
References
1. Walker JC, Ratcliffe MB, Zhang P, Wallace AW, Fata B, Hsu EW, et al. MRI-based finite-element analysis of left ventricular aneurysm. Am J Physiol Heart Circ Physiol. 2005;289(2):H692–700. doi:10.1152/ajpheart.01226.2004. [Google Scholar] [PubMed] [CrossRef]
2. Mut F, Löhner R, Chien A, Tateshima S, Viñuela F, Putman C, et al. Computational hemodynamics framework for the analysis of cerebral aneurysms. Int J Numer Methods Biomed Eng. 2011;27(6):822–39. doi:10.1002/cnm.1424. [Google Scholar] [PubMed] [CrossRef]
3. Zhang S, Laubrie JD, Mousavi SJ, Avril S. 3D finite-element modeling of vascular adaptation after endovascular aneurysm repair. Int J Numer Methods Biomed Eng. 2022;38(2):e3547. doi:10.1002/cnm.3547. [Google Scholar] [PubMed] [CrossRef]
4. Kim J, Zhang K, Canton G, Balu N, Meyer K, Saber R, et al. In vivo deformation of the human basilar artery. Ann Biomed Eng. 2025;53(1):83–98. doi:10.1007/s10439-024-03605-x. [Google Scholar] [PubMed] [CrossRef]
5. Fukushima T, Matsuzawa T, Homma T. Visualization and finite element analysis of pulsatile flow in models of the abdominal aortic aneurysm. Biorheology. 1989;26(2):109–30. doi:10.3233/bir-1989-26203. [Google Scholar] [PubMed] [CrossRef]
6. Steinman DA, Milner JS, Norley CJ, Lownie SP, Holdsworth DW. Image-based computational simulation of flow dynamics in a giant intracranial aneurysm. Am J Neuroradiol. 2003;24(4):559–66. [Google Scholar] [PubMed]
7. Fillingham P, Bhathal JR, Marsh LMM, Barbour MC, Kurt M, Ionita CN, et al. Improving the accuracy of computational fluid dynamics simulations of coiled cerebral aneurysms using finite element modeling. J Biomech. 2023;157(1):111733. doi:10.1101/2023.02.27.23286512. [Google Scholar] [CrossRef]
8. Ainsworth M, Oden JT. A posteriori error estimation in finite element analysis. Comput Methods Appl Mech Eng. 1997;142(1–2):1–88. doi:10.1016/s0045-7825(96)01107-3. [Google Scholar] [CrossRef]
9. Botti L, Piccinelli M, Ene-Iordache B, Remuzzi A, Antiga L. An adaptive mesh refinement solver for large-scale simulation of biological flows. Int J Numer Methods Biomed Eng. 2010;26(1):86–100. doi:10.1002/cnm.1257. [Google Scholar] [CrossRef]
10. Cant RS, Ahmed U, Fang J, Chakarborty N, Nivarti G, Moulinec C, et al. An unstructured adaptive mesh refinement approach for computational fluid dynamics of reacting flows. J Comput Phys. 2022;468:111480. doi:10.1016/j.jcp.2022.111480. [Google Scholar] [CrossRef]
11. Smith BF, Bjørstad PE, Gropp WD. Domain decomposition: parallel multilevel methods for elliptic partial differential equations. Cambridge, UK: Cambridge University Press; 2004. [Google Scholar]
12. Kačeniauskas A, Rutschmann P. Parallel FEM software for CFD problems. Informatica. 2004;15(3):363–78. [Google Scholar]
13. Moureau V, Domingo P, Vervisch L. Design of a massively parallel CFD code for complex geometries. C R Mécanique. 2011;339(2–3):141–8. doi:10.1016/j.crme.2010.12.001. [Google Scholar] [CrossRef]
14. Raveh DE. Reduced-order models for nonlinear unsteady aerodynamics. AIAA J. 2001;39(8):1417–29. doi:10.2514/2.1473. [Google Scholar] [CrossRef]
15. Lucia DJ, Beran PS, Silva WA. Reduced-order modeling: new approaches for computational physics. Prog Aerosp Sci. 2004;40(1–2):51–117. doi:10.1016/j.paerosci.2003.12.001. [Google Scholar] [CrossRef]
16. Lu Y, Li H, Saha S, Mojumder S, Al Amin A, Suarez D, et al. Reduced order machine learning finite element methods: concept, implementation, and future applications. Comput Model Eng Sci. 2021;129(3):1351–71. doi:10.32604/cmes.2021.017719. [Google Scholar] [CrossRef]
17. Schmid PJ. Dynamic mode decomposition and its variants. Annu Rev Fluid Mech. 2022;54(1):225–54. doi:10.1146/annurev-fluid-030121-015835. [Google Scholar] [CrossRef]
18. Fukami K, Fukagata K, Taira K. Super-resolution analysis via machine learning: a survey for fluid flows. arXiv:2301.10937. 2023. [Google Scholar]
19. Lino M, Fotiadis S, Bharath AA, Cantwell CD. Current and emerging deep-learning methods for the simulation of fluid dynamics. Proc R Soc A. 2023;479(2275):20230058. doi:10.1098/rspa.2023.0058. [Google Scholar] [CrossRef]
20. Bi X, Liu A, Fan Y, Yu C, Zhang Z. FlowSRNet: a multi-scale integration network for super-resolution reconstruction of fluid flows. Phys Fluids. 2022;34(12):127104. doi:10.1063/5.0128435. [Google Scholar] [CrossRef]
21. Li Z, Liu F, Yang W, Peng S, Zhou J. A survey of convolutional neural networks: analysis, applications, and prospects. IEEE Trans Neural Netw Learn Syst. 2021;33(12):6999–7019. doi:10.1109/tnnls.2021.3084827. [Google Scholar] [PubMed] [CrossRef]
22. Dong C, Loy CC, He K, Tang X. Image super-resolution using deep convolutional networks. IEEE Trans Pattern Anal Mach Intell. 2015;38(2):295–307. doi:10.1109/tpami.2015.2439281. [Google Scholar] [PubMed] [CrossRef]
23. Kim J, Lee JK, Lee KM. Accurate image super-resolution using very deep convolutional networks. In: Proceedings of the 2016 IEEE Conference on Computer Vision and Pattern Recognition; 2016 Jun 26–Jul 1; Las Vegas, NV, USA. Piscataway, NJ, USA: IEEE; 2016. p. 1646–54. [Google Scholar]
24. Wang X, Yu K, Wu S, Gu J, Liu Y, Dong C, et al. ESRGAN: enhanced super-resolution generative adversarial networks. In: Computer Vision—ECCV 2018 Workshops (ECCV 2018). Cham, Switzerland: Springer; 2018. p. 63–79. [Google Scholar]
25. Yan C, Shi G, Wu Z. SMIR: a transformer-based model for MRI super-resolution reconstruction. In: 2021 IEEE International Conference on Medical Imaging Physics and Engineering (ICMIPE). Piscataway, NJ, USA: IEEE; 2021. p. 1–6. [Google Scholar]
26. Xu Q, Zhuang Z, Pan Y, Wen B. Super-resolution reconstruction of turbulent flows with a transformer-based deep learning framework. Phys Fluids. 2023;35(5):055130. doi:10.1063/5.0149551. [Google Scholar] [CrossRef]
27. Cai S, Mao Z, Wang Z, Yin M, Karniadakis GE. Physics-informed neural networks (PINNs) for fluid mechanics: a review. Acta Mech Sin. 2021;37(12):1727–38. doi:10.1007/s10409-021-01148-1. [Google Scholar] [CrossRef]
28. Arzani A, Wang JX, D’Souza RM. Uncovering near-wall blood flow from sparse data with physics-informed neural networks. Phys Fluids. 2021;33(7):071905. doi:10.1063/5.0055600. [Google Scholar] [CrossRef]
29. Han R, Yang X, Liu W. Unsteady flow prediction based on a hybrid network-operator learning model. Phys Fluids. 2025;37(2):027150. doi:10.1063/5.0249234. [Google Scholar] [CrossRef]
30. Sofos F, Drikakis D. A review of deep learning for super-resolution in fluid flows. Phys Fluids. 2025;37(4):041303. doi:10.1063/5.0265738. [Google Scholar] [CrossRef]
31. Habibi M, D’Souza RM, Dawson STM, Arzani A. Integrating multi-fidelity blood flow data with reduced-order data assimilation. Comput Biol Med. 2021;135:104566. doi:10.1016/j.compbiomed.2021.104566. [Google Scholar] [PubMed] [CrossRef]
32. Zhang Y, Freeman EL, Stinson JT, Riveros GA. Deep learning-based super resolution applied to finite element analysis of fused deposition modeling 3D printing. 3D Print Addit Manuf. 2025;12(4):315–24. doi:10.1089/3dp.2023.0191. [Google Scholar] [PubMed] [CrossRef]
33. Yonekura K, Maruoka K, Tyou K, Suzuki K. Super-resolving 2D stress tensor field conserving equilibrium constraints using physics-informed U-Net. Finite Elem Anal Des. 2023;213(4):103852. doi:10.1016/j.finel.2022.103852. [Google Scholar] [CrossRef]
34. Arora R. PhySRNet: physics informed super-resolution network for application in computational solid mechanics. In: 2022 IEEE/ACM International Workshop on Artificial Intelligence and Machine Learning for Scientific Applications (AI4S). Piscataway, NJ, USA: IEEE; 2022. p. 13–8. doi:10.1109/ai4s56813.2022.00008. [Google Scholar] [CrossRef]
35. Bolandi H, Li X, Salem T, Boddeti VN, Lajnef N. Bridging finite element and deep learning: high-resolution stress distribution prediction in structural components. Front Struct Civ Eng. 2022;16(11):1365–77. doi:10.1007/s11709-022-0882-5. [Google Scholar] [CrossRef]
36. Wang H, Cao Y, Huang Z, Liu Y, Hu P, Luo X, et al. Recent advances on machine learning for computational fluid dynamics: a survey. arXiv:2408.12171. 2024. [Google Scholar]
37. Arzani A, Dawson STM. Data-driven cardiovascular flow modelling: examples and opportunities. J R Soc Interface. 2021;18(175):20200802. doi:10.1098/rsif.2020.0802. [Google Scholar] [PubMed] [CrossRef]
38. Logg A, Mardal KA, Wells G. Automated solution of differential equations by the finite element method: The FEniCS book. Vol. 84. Heidelberg/Berlin, Germany: Springer Science & Business Media; 2012. [Google Scholar]
39. Alnæs M, Blechta J, Hake J, Johansson A, Kehlet B, Logg A, et al. The FEniCS project version 1.5. Arch Numer Softw. 2015;3(100):9–23. [Google Scholar]
40. Schroeder W, Martin KM, Lorensen WE. The visualization toolkit: an object-oriented approach to 3D graphics. Saddle River, NJ, USA: Prentice-Hall, Inc.; 1998. [Google Scholar]
41. Ahrens J, Geveci B, Law C. Paraview: An End-User Tool for Large Data Visualization [Internet]. 2005 [cited 2026 Jan 1]. Available from: https://www.researchgate.net/publication/247111133_ParaView_An_End-User_Tool_for_Large_Data_Visualization. [Google Scholar]
42. Ayachit U. The paraview guide: a parallel visualization application. New York, NY, USA: Kitware, Inc.; 2015. [Google Scholar]
43. He K, Zhang X, Ren S, Sun J. Deep residual learning for image recognition. In: Proceedings of the 2016 IEEE Conference on Computer Vision and Pattern Recognition; 2016 Jun 26–Jul 1; Las Vegas, NV, USA. Piscataway, NJ, USA: IEEE. p. 770–8. [Google Scholar]
44. Wu Z, Shen C, Van Den Hengel A. Wider or deeper: revisiting the ResNet model for visual recognition. Pattern Recognit. 2019;90:119–33. [Google Scholar]
45. Paszke A, Gross S, Massa F, Lerer A, Bradbury J, Chanan G, et al. PyTorch: an imperative style, high-performance deep learning library. In: Advances in Neural Information Processing Systems. Vol. 32. Red Hook, NY, USA: Curran Associates, Inc.; 2019. [Google Scholar]
46. Bell JB, Colella P, Glaz HM. A second-order projection method for the incompressible Navier-Stokes equations. J Comput Phys. 1989;85(2):257–83. doi:10.1016/0021-9991(89)90151-4. [Google Scholar] [CrossRef]
47. Turek S. On discrete projection methods for the incompressible Navier-Stokes equations: an algorithmical approach. Comput Methods Appl Mech Eng. 1997;143(3–4):271–88. doi:10.1016/s0045-7825(96)01155-3. [Google Scholar] [CrossRef]
48. Lim B, Son S, Kim H, Nah S, Lee KM. Enhanced deep residual networks for single image super-resolution. In: Proceedings of the 2017 IEEE Conference on Computer Vision and Pattern Recognition (CVPR) Workshops; 2017 Jul 21–26; Honolulu, HI, USA. Piscataway, NJ, USA: IEEE. p. 136–44. [Google Scholar]
Cite This Article
 Copyright © 2026 The Author(s). Published by Tech Science Press.
Copyright © 2026 The Author(s). Published by Tech Science Press.This work is licensed under a Creative Commons Attribution 4.0 International License , which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.


 Submit a Paper
Submit a Paper Propose a Special lssue
Propose a Special lssue View Full Text
View Full Text Download PDF
Download PDF Downloads
Downloads
 Citation Tools
Citation Tools