 Open Access
Open Access
ARTICLE
ABCC8 is correlated with immune cell infiltration and overall survival in lower grade glioma
1 Department of Academic Research, The Secondary Hospital, Cheeloo College of Medicine, Shandong University, Jinan, 250033, China
2 Department of Cancer Center, The Secondary Hospital, Cheeloo College of Medicine, Shandong University, Jinan, 250033, China
* Corresponding Authors: Ming Jia, ;
(This article belongs to the Special Issue: Bioinformatics Study of Diseases)
BIOCELL 2023, 47(1), 109-123. https://doi.org/10.32604/biocell.2023.024620
Received 02 June 2022; Accepted 14 July 2022; Issue published 26 September 2022
Abstract
ATP binding cassette subfamily C member 8 (ABCC8) encodes a protein regulating the ATP-sensitive potassium channel. Whether the level of ABCC8 mRNA in lower grade glioma (LGG) correlates with immune cell infiltration and patient outcomes has not been evaluated until now. Comparisons of ABCC8 expression between different tumors and normal tissues were evaluated by exploring publicly available datasets. The association between ABCC8 and tumor immune cell infiltration, diverse gene mutation characteristics, tumor mutation burden (TMB), and survival in LGG was also investigated in several independent datasets. Pathway enrichment analysis was conducted to search for ABCC8-associated signaling pathways. Through an online database, we found that ABCC8 expression in LGG was lower than in normal tissues. Then, the association of ABCC8 expression and immune cell infiltration in LGG was discussed. As we expected, the ABCC8 mRNA levels were negatively associated with non-T immune cell infiltration levels in all datasets. Consistently, TCGA_LGG RNA-seq data revealed that ABCC8 downregulated several non-T immune cell-associated signaling pathways in gene set enrichment analysis. Different ABCC8 expression groups showed diverse gene mutation characteristics and TMB. The high expression of ABCC8 was linked to improved survival of LGG patients. A pathway enrichment analysis of ABCC8-associated genes indicated that the GABAergic synapse signaling pathway might be involved in regulating immunity in LGG. Our findings show that ABCC8 reflects LGG tumor immunity and is an ideal prognostic biomarker for LGG.Keywords
Lower grade glioma (LGG) is a heterogeneous malignant brain tumor in humans that amounts to approximately 20% of intracranial tumors (Hoshide and Jandial, 2016). It was identified as grades II and III brain tumors by the World Health Organization. The incidence is estimated to be approximately 0.8 cases per 100,000 population (Nakasu and Nakasu, 2022). LGGs arise from supporting glial cells and usually affect young adults, and their main treatment involves surgical resection, followed by radiation and chemotherapy (Youssef and Miller, 2020). They are slowly growing tumors and often lack symptoms, except for seizures (Nakasu et al., 2021). While the pathogenesis mechanisms and risk factors for LGG are poorly understood, it seems plausible that an association of genetic susceptibility and biological, functional, and environmental factors influences the process (Darlix et al., 2017). The overall survival (OS) of LGG varies dramatically, as does the patient’s response to standard therapy (Brat and Pachter, 2015). For decades, there have been no significant improvements in the treatment of LGG; thus, the prognosis has not changed significantly (Claus et al., 2015).
Recently, immunotherapy has become an extremely promising strategy for many types of cancers and may help us improve the survival rate of patients with LGG. A dysfunctional immune response leads to tumor immune evasion in gliomas (Wang et al., 2018). Immunotherapy can block the dysfunction and kill cancer cells by activating the immune system (Chuah and Chew, 2020). Currently, immune checkpoint inhibitors (ICIs) are the most successful immunotherapy drugs in use, while programmed death-ligand 1 (PD-L1) expression, microsatellite instability, mismatch repair, and tumor mutation burden (TMB) are the most valuable biomarkers for predicting ICI efficiency (Hodges et al., 2017; Rizvi et al., 2018). Yin et al. (2020) found that TMB is negatively related to immune infiltration and OS in LGG. However, unlike other prevalent solid malignant tumors, the immune activity and immunotherapy efficiency are still largely unknown in LGG and need to be studied further.
ATP binding cassette subfamily C member 8 (ABCC8), an ATP-binding cassette (ABC) transporter, leads to multidrug resistance in many kinds of cancer cells by pumping anticancer agents out. ABCC8 variant was linked with diabetes and hypertension by numerous studies (Beltrand et al., 2020; de Franco et al., 2020; Flagg et al., 2007; Southgate et al., 2020). Recently, Rehman et al. (2022, 2020) also reported that the genetic variant of ABCC8 is associated with cardiac diseases and metabolic disorders. Loss-of-function mutations of ABCC8 were also reported by Bohnen et al. (2018) to be associated with pulmonary arterial hypertension (Bohnen et al., 2018). Its expression was found to promote cerebral edema after brain ischemia and injury (Alquisiras-Burgos et al., 2020). The expression of ABCC8 was also decreased in pancreatic, lung, and breast cancers. Interestingly, in these tumors, ABCC8 expression was linked to favorable survival (Hlavac et al., 2013; Mohelnikova-Duchonova et al., 2013; Wang et al., 2020). A recent study reported that ABCC8 mRNA levels are positively related to survival in patients with glioma (Zhou et al., 2020). ABC transporters regulate the development, differentiation, and maturation of immune cells and are involved in the migration of immune effector cells to sites of inflammation (van de Ven et al., 2009). They were also shown recently to regulate T-cell populations, such as thymocytes, natural killer T cells, CD8+ T cells, and regulatory T cells (Thurm et al., 2021). One family member, ABCC5, was shown by Chen et al. (2021) to be associated with immune infiltration of hepatocellular carcinoma. However, whether ABCC8 participates in immune responses and provides survival benefits in LGG is still not clear and needs to be investigated.
In this study, we systematically evaluated whether ABCC8 expression reflects the immune microenvironment of LGG tumor tissues. The results were verified in several independent datasets. We also attempted to identify the related signaling pathways. The association between ABCC8 gene expression and LGG patient survival was also investigated. This study might help identify an ideal biomarker for predicting survival and sensitivity to immunotherapy in patients with LGG.
Gene expression data and clinical information of LGG patients from The Cancer Genome Atlas (TCGA_LGG) database were obtained from the UCSC website (http://xena.ucsc.edu/public) on October 01, 2021. The RNA expression levels and survival information of LGG patients in the REMBRANDT dataset and CGGA datasets were downloaded from the CGGA website (http://www.cgga.org.cn/index.jsp) on the same day (Bao et al., 2014; Liu et al., 2018; Wang et al., 2015; Zhao et al., 2017).
If the same patient provided two or more tumor samples to those datasets, only the data of the primary lesion were selected according to the sample numbers. Gene expression data of fragments per kilobase per million were converted to transcripts per million and then log-transformed to provide more precise results. Gene symbols were extracted from the provided documents from the dataset websites.
ABCC8 expression levels between diverse cancer types and normal tissues were analyzed by Gene Expression Profiling Interactive Analysis (GEPIA) (http://gepia.cancerpku.cn/index.html) (Tang et al., 2017). The cutoff value of probability was 0.05.
Correlation between ABCC8 and tumor immune cell infiltration
We studied the relationship between ABCC8 and the infiltration of six immune cell types (CD4+ T cells, CD8+ T cells, B cells, macrophages, dendritic cells, and neutrophils) in LGG tumor tissues from the TCGA dataset through TIMER (https://cistrome.shinyapps.io/timer/). The relationship between gene expression levels and the degree of tumor purity is shown in the first panel of Fig. 2A (Aran et al., 2015). In addition, we also investigated the associations between the expression of ABCC8 and six highly researched immunotherapy-targeted genes by correlation modules. The hypothesis test at p < 0.05 was considered statistically significant.
Then, the CGGA datasets with the same type of RNA-seq expression levels as the TCGA dataset were chosen to validate the association of ABCC8 and immune cell infiltration in LGG. The infiltration levels of six immune cells were calculated using the same method as TIMER via the immunedeconv package in R (Sturm et al., 2019). The input data were converted to transcripts per million normalized without log transformation.
Correlation between ABCC8 and tumor mutation burden
The Mutation Annotation Format (MAF) file containing all gene mutation characteristics of tumor samples (workflow type: VarScan2) of the TCGA_LGG cohort was downloaded from the GDC database (https://portal.gdc.cancer.gov/). The gene mutation characteristics and TMB of different ABCC8 expression groups were analyzed by the maftools package in R.
Gene set enrichment analysis (GESA)
Pathways obviously related to ABCC8 mRNA levels were analyzed using GSEA through GSEA software 4.0.0 (Subramanian et al., 2005). The gene set database was C2.cp.kegg.v7.4. symbols.gmt. The pathways enriched with a PFWER < 0.1 were considered significant.
The top 50 ABCC8-associated genes were screened from the RNA-seq data of LGG samples on the Cancer Genomics website (cBioPortal: https://www.cbioportal.org). The resulting protein network was built on the STRING website (https://string-db.org/) (Szklarczyk et al., 2021). Moreover, the WEB-based gene set analysis toolkit (http://www.webgestalt.org/) was applied to launch the gene ontology (GO) annotation and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis (Liao et al., 2019).
We implemented all statistical analyses with R version 4.0.5 (R Foundation for Statistical Computing, Vienna, Austria) and GraphPad Prism 6.01 (GraphPad Software, Inc., San Diego, CA, USA). LGG patients were divided into two groups (ABCC8 low expression and high expression groups) by the median ABCC8 mRNA expression levels in the CGGA, REMBRANDT, and TCGA databases. Overall survival analysis between the two groups was conducted by Kaplan–Meier curves with the Wilcoxon log-rank test. Multiple factor analyses were carried out by the Cox regression model and are presented in forest plots. The correlation between ABCC8 gene expression and other genes was assessed by Spearman’s correlation analysis. Spearman’s correlation analysis was used to search for ABCC8-related genes. The entry criterion was a statistical p < 0.05.
The level of ABCC8 mRNA in multiple cancer types
The DiffExp module of TIMER displayed the ABCC8 mRNA levels in different tumors together with their adjacent normal tissues. As shown in Fig. 1A, ABCC8 expression in bladder cancer, colon cancer, esophageal cancer, head and neck cancer, kidney cancer, lung cancer, prostate cancer, rectal cancer, stomach cancer, thyroid cancer, and uterine corpus endometrial carcinoma was lower than that in adjacent normal tissues. However, it was significantly higher in cholangiocarcinoma and liver cancer than in adjacent normal tissue. For LGG, ABCC8 mRNA was higher in metastatic tumors than in primary tumors. However, there are no data comparing ABCC8 expression between tumors and adjacent normal tissues in LGG owing to the absence of a normal tissue sample.
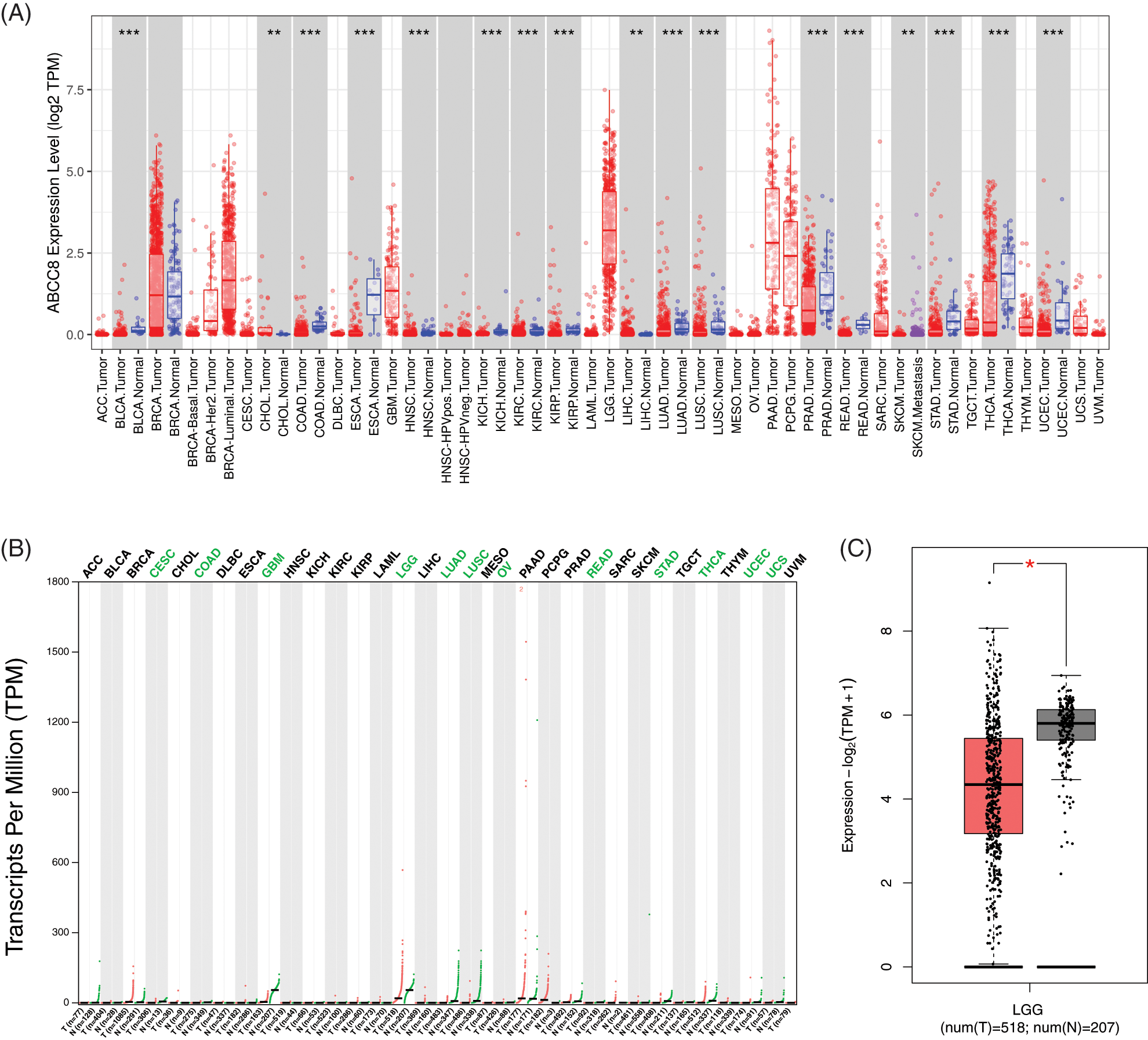
Figure 1: ABCC8 expression levels in different types of cancers. (A) The expression levels of ABCC8 in cancer and normal tissues in the TCGA database using TIMER. (B) The expression data of ABCC8 from the GEPIA database. (C) The expression of ABCC8 in LGG in cancer and normal tissues in the GEPIA database. *p < 0.05; **p < 0.01; ***p < 0.001.
The GEPIA database was also applied to evaluate the expression of ABCC8 in various human tumors (Fig. 1B). ABCC8 expression in cervical squamous cell carcinoma, endocervical adenocarcinoma, glioblastoma multiforme, colon cancer, lung cancer, rectal cancer, stomach cancer, thyroid cancer, ovarian serous cystadenocarcinoma, uterine corpus endometrial carcinoma, uterine carcinosarcoma, and LGG was lower than normal (Fig. 1B). Furthermore, most results from the above datasets were consistent. For LGG, the detailed expression levels between tumor tissue and normal tissues are shown in Fig. 1C, and the difference was significant (p < 0.05).
Association between ABCC8 and immune cells
The gene module of TIMER was then used to study the relationship between ABCC8 and immune cell infiltration in various tumor tissues. There were close relationships between ABCC8 mRNA expression and most immune cells in several types of cancers, such as head and neck squamous cell cancer, stomach cancer, thyroid cancer, and thymoma (Suppl. Figs. S1–S3). In LGG, the ABCC8 mRNA levels were negatively correlated with B cells, CD4+ T cells, CD8+ T cells, macrophages, neutrophils, and dendritic cells (Fig. 2A). Additionally, the infiltration levels of B cells, CD4+ T cells, macrophages, neutrophils, and dendritic cells seemed to be connected to altered ABCC8 gene copy numbers (Fig. 2B). However, there was no significant relationship between ABCC8 gene copy numbers, and CD8+ T-cell infiltration levels. We also tested the correlation of the infiltration levels of immune cells with the survival of LGG patients. As shown in Fig. 2C, the infiltration levels of six immune cells were significantly associated with the survival of LGG patients, which indicated that tumor immunity might influence the tumor phenotype of LGG and lead to different events.
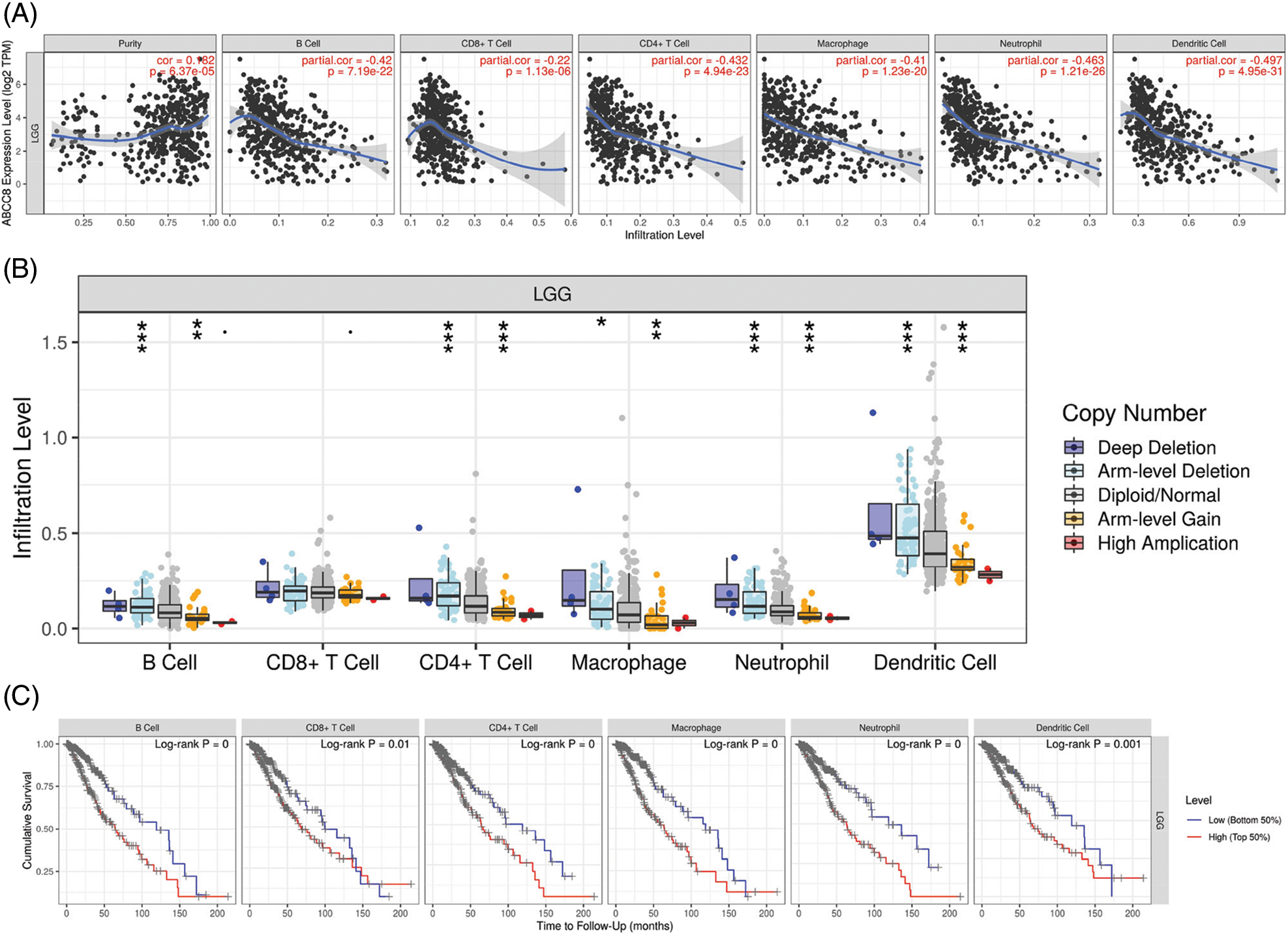
Figure 2: Correlation of ABCC8 expression, gene copy numbers and patient prognosis with immune infiltration levels in LGG. (A) The expression of ABCC8 is positively related to tumor purity and negatively correlated with the infiltrating levels of CD4+ T cells, CD8+ T cells, B cells, macrophages, neutrophils, and dendritic cells in LGG. The association between ABCC8 copy numbers and immune cell infiltration levels in LGG (B). Immune cell infiltration levels are significantly associated with the survival of LGG patients (C). *p < 0.05; **p < 0.01; ***p < 0.001.
To validate the observed association between ABCC8 and immune cell infiltration in LGG, the immune cell infiltration levels in the CGGA datasets were estimated by the same method as TIMER. As shown in Figs. 3A and 3B, ABCC8 mRNA levels had a clear negative correlation with the infiltration of B cells, macrophages, neutrophils, and dendritic cells. For CD8+ and CD4+ T cells, the results from different datasets were not consistent. Taken together, ABCC8 may affect the tumor immunity microenvironment by regulating non-T-cell immune cells.
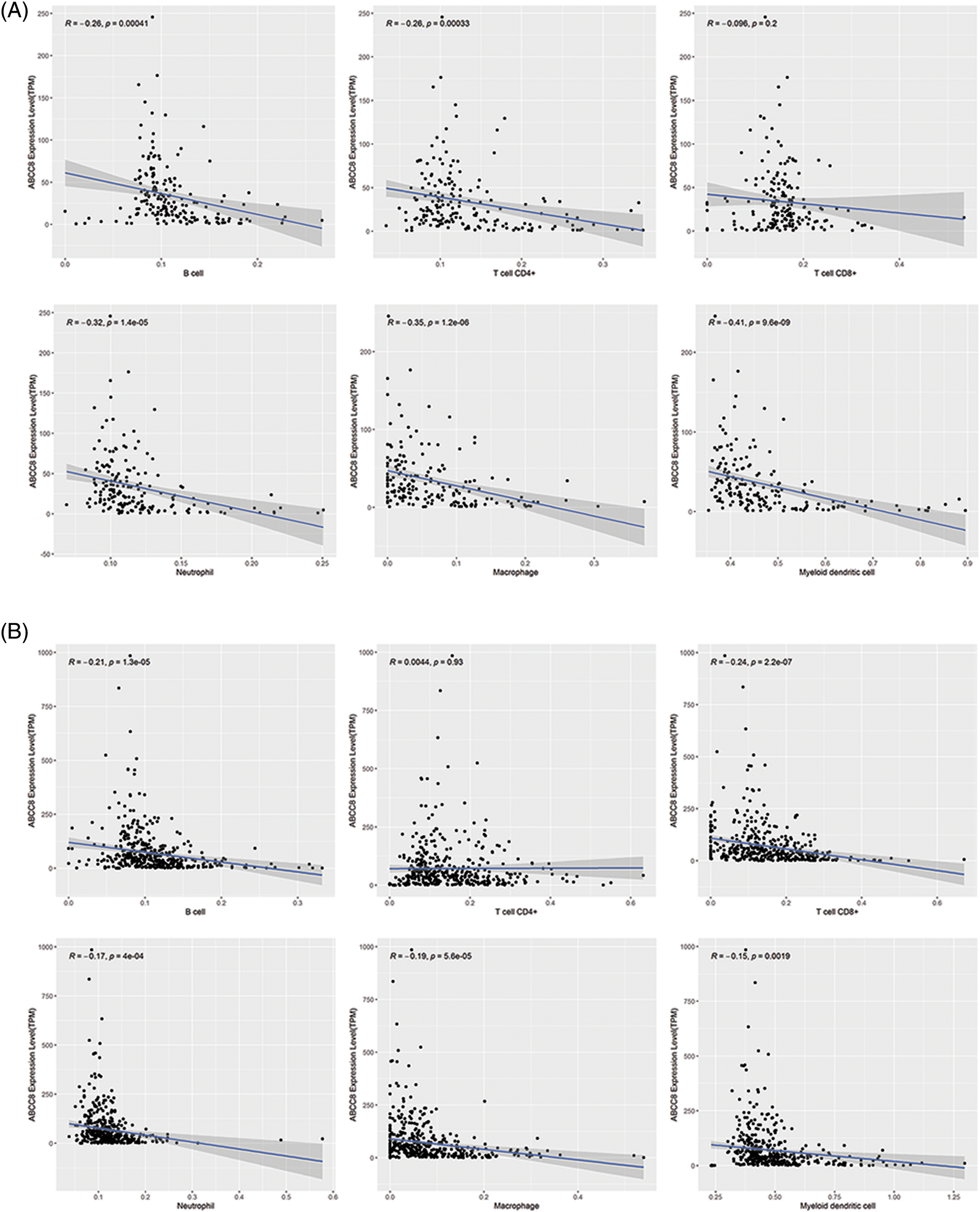
Figure 3: Validation of the association between ABCC8 expression levels and immune infiltration levels of LGG in CGGA datasets. (A) The expression of ABCC8 is negatively correlated with the infiltrating levels of B cells, CD4+ T cells, macrophages, neutrophils, and dendritic cells in the CGGA mRNAseq_325 dataset. The expression of ABCC8 is negatively correlated with the infiltrating levels of B cells, CD8+ T cells, macrophages, neutrophils, and dendritic cells in the CGGA mRNAseq_693 (B) dataset.
Association between ABCC8 and immunotherapy-targeted genes
Because ABCC8 reflects the tumor immune activity of LGG, we further investigated the expression relationship between ABCC8 and six highly researched immunotherapy-targeted genes in patients from the TCGA-LGG dataset. After adjusting for tumor purity, we found a negative association between the expression of ABCC8 and PDCD1 (correlation coefficient (Cor = −0.327, p < 0.001) (Fig. 4A), CD274 (Cor = −0.125, p = 0.006) (Fig. 4B), CTLA4 (Cor = −0.242, p < 0.001) (Fig. 4C), LAG3 (Cor = −0.178, p < 0.001) (Fig. 4D) and HAVCR2/TIM3 (Cor = −0.495, p < 0.001) (Fig. 4E). However, only the expression of TIGIT was weakly positively related to the expression of ABCC8 (Cor = 0.166, p < 0.001) (Fig. 4F). Owing to the higher expression of the five immunotherapy-targeted genes and the infiltration of cytotoxic lymphocytes, LGG patients with low ABCC8 expression may receive great survival improvement from combined ICI treatment. In summary, the expression of ABCC8 in LGG patients may help predict the sensitivity of immunotherapy.
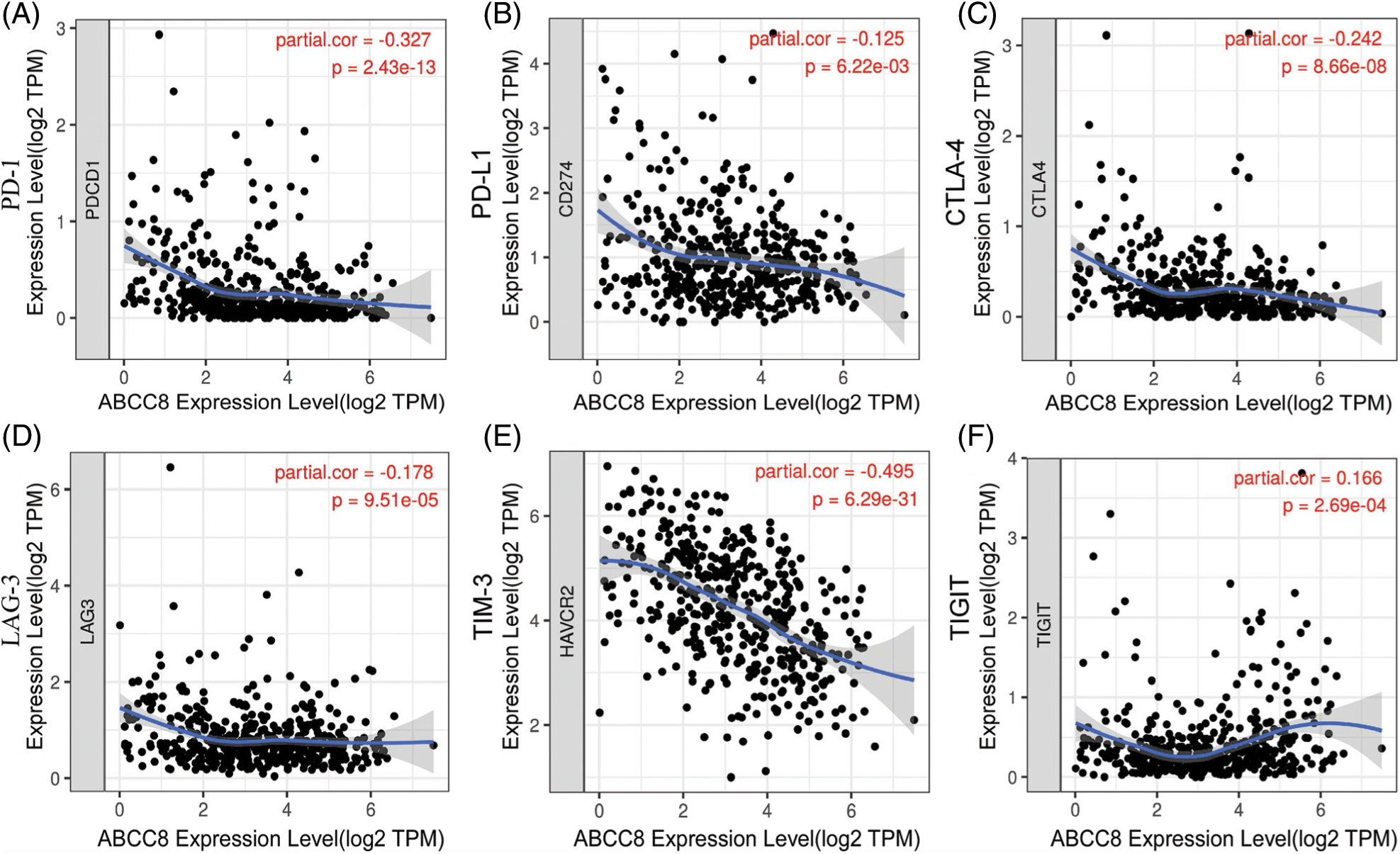
Figure 4: Correlation of ABCC8 expression and the expression of six hot immunotherapy-targeted genes from the TCGA_LGG dataset. (A) The expression of ABCC8 is negatively related to PDCD1 expression. (B) The expression of ABCC8 is negatively related to CD274 expression. (C) The expression of ABCC8 is negatively related to CTLA4 expression. (D) The expression of ABCC8 is negatively related to LAG3 expression. (E) The expression of ABCC8 is negatively related to HAVCR2/TIM3 expression. (F) The expression of ABCC8 is positively related to TIGIT expression.
The prognostic implications of ABCC8 in lower grade glioma
To further investigate the prognostic value of ABCC8 in LGG, we selected LGG samples from TCGA, CGGA (mRNAseq_325, mRNAseq_693), and REMBRANDT datasets. A high ABCC8 mRNA expression levels corresponded with a favorable prognosis in LGG patients in the TCGA dataset (OS hazard ratio (HR) = 0.260, 95% confidence interval (CI) = 0.131–0.514, p < 0.001, respectively) (Fig. 5A), CGGA mRNAseq_325 dataset (OS HR = 0.264, 95% CI = 0.171–0.407, p < 0.001, respectively) (Fig. 5B), CGGA mRNAseq_693 dataset (OS HR = 0.349, 95% CI = 0.262–0.465, p < 0.001, respectively) (Fig. 5C), and REMBRANDT dataset (OS HR = 0.485, 95% CI = 0.305–0.714, p = 0.0009, respectively) (Fig. 5D). Multiple factor analyses showed ABCC8 as an independent predictive marker for OS in LGG regardless of age, sex, IDH mutation status, 1p19q codeletion status or MGMT methylation status in the TCGA dataset (OS HR = 2.462, 95% CI = 1.674–3.520, p < 0.001, respectively) (Fig. 5E), CGGA mRNAseq_325 dataset (OS HR = 2.031, 95% CI = 1.237–3.333, p = 0.005, respectively) (Fig. 5F), CGGA mRNAseq_693 dataset (OS HR = 2.614, 95% CI = 1.785–3.828, p < 0.001, respectively) (Fig. 5G), and REMBRANDT dataset (OS HR = 1.997, 95% CI = 1.194–3.340, p = 0.008, respectively) (Fig. 5H). These results suggest that the expression of ABCC8 may be independently correlated with variations in prognoses of patients with LGG.
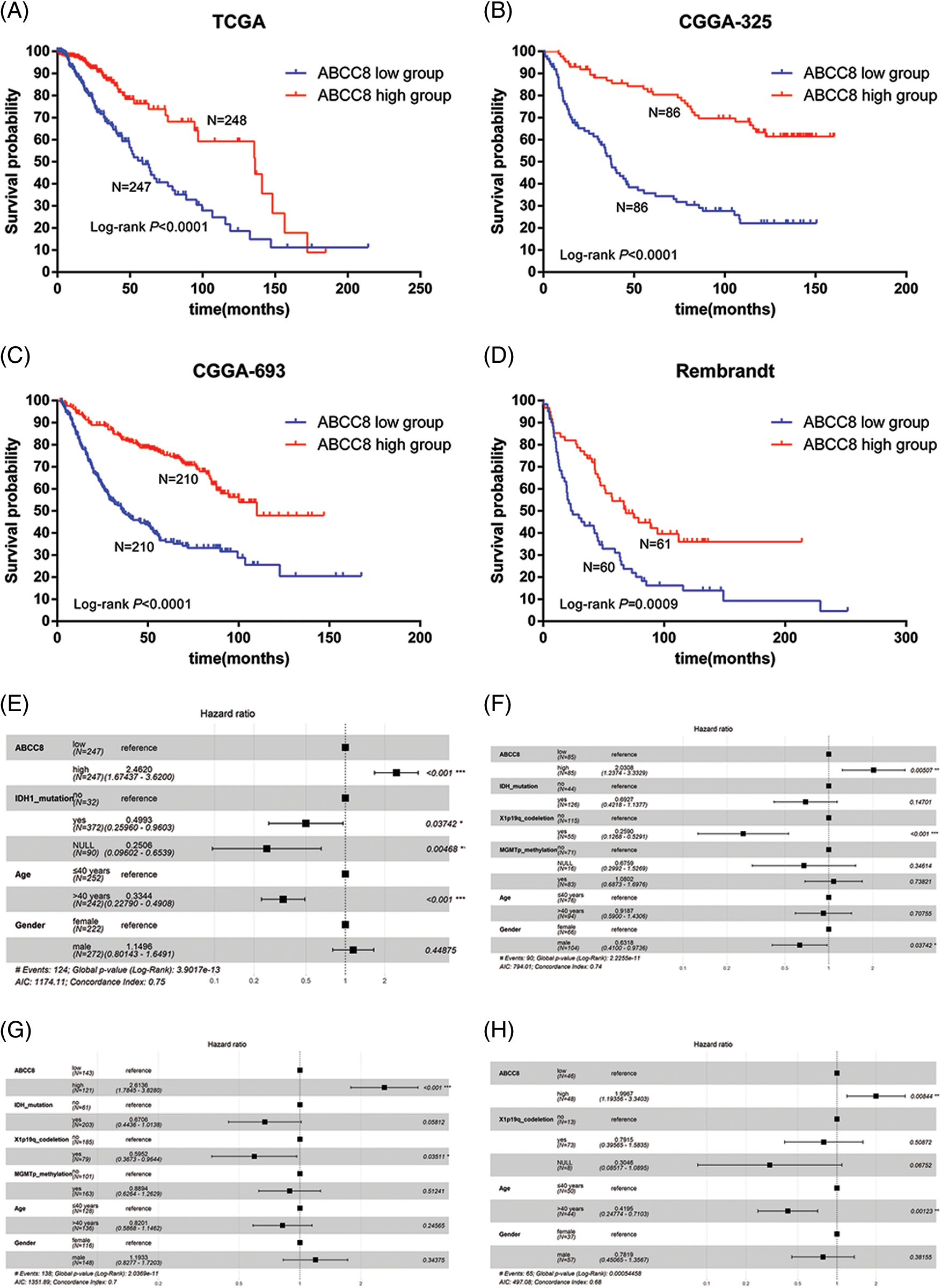
Figure 5: Kaplan-Meier survival analysis of ABCC8 in LGG from different public datasets. (A) Kaplan-Meier overall survival analysis of ABCC8 in the TCGA_LGG dataset. (B) Kaplan-Meier overall survival analysis of ABCC8 in the CGGA mRNAseq_325 dataset. Kaplan-Meier overall survival analysis of ABCC8 in the CGGA mRNAseq_693 (C) and REMBRANDT (D) datasets. Forest plot showing the results of multiple factor Cox regression analysis of ABCC8 in OS of TCGA_LGG (E), CGGA mRNAseq_325 (F), CGGA mRNAseq_693 (G), and REMBRANDT (H) datasets with other clinical factors. OS: overall survival.
Correlation between ABCC8 and tumor mutation burden
TMB has been used as an immunotherapy resistance biomarker in the majority of solid malignant tumors. We also wondered whether TMB varied between different ABCC8 expression groups and compared the mutation frequency of all genes between LGG tumor samples with different ABCC8 expression levels (Figs. 6A and 6B). Generally, the mutation frequency was extremely low in LGG tumors. Mutations of TP53, ATRX, and EGFR in the ABCC8 low expression group were more common than in the ABCC8 high expression group. In contrast, mutations in CIC, FUBP1, and IDH were less common in the low expression group (Fig. 6C). These mutation variances may also partly explain the different survival outcomes of the groups. Consequently, the ABCC8 high expression group had a significantly lower TMB than the ABCC8 low expression group (0.38 vs. 0.46/MB, p < 0.001) (Fig. 6D), which suggests that ABCC8 could reflect the level of TMB in LGG.
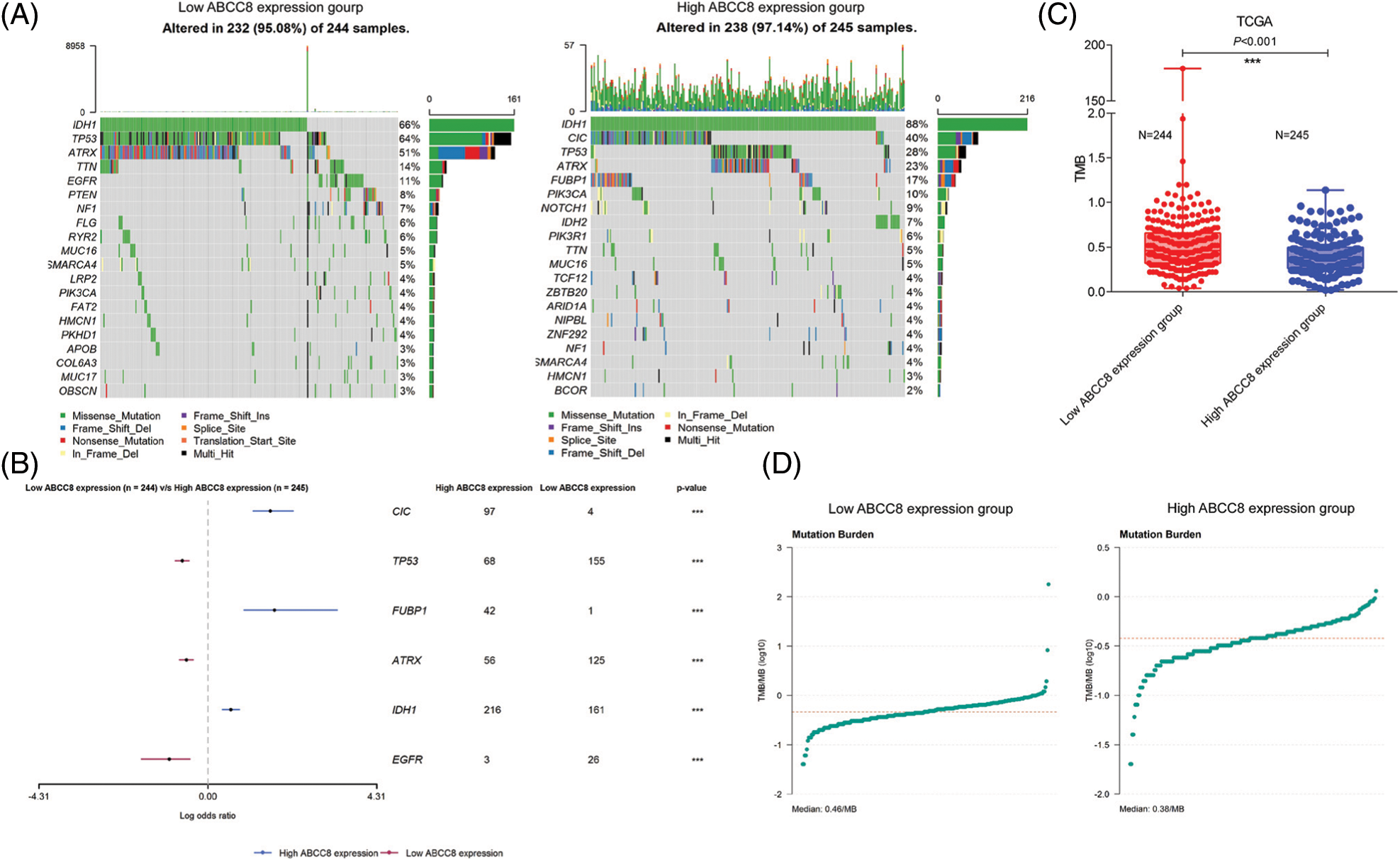
Figure 6: Analysis of mutation burden in different ABCC8 expression groups. Mutation landscape of LGG tumor samples between different ABCC8 mRNA expression groups (A). Forest plot showing the most differentially mutated genes with low ABCC8 and high ABCC8 expression (B). (C) Comparison of mutation frequency between the low ABCC8 expression group and high ABCC8 expression group. (D) Tumor mutation burden of the low ABCC8 expression group and the high ABCC8 expression group. *p < 0.05; **p < 0.01; ***p < 0.001.
Identification of ABCC8-associated key genes and pathways in the lower grade glioma immune response
The tumor sample tissues of TCGA-LGG datasets were dichotomized into ABCC8high and ABCC8low groups according to the median ABCC8 mRNA level. GSEA was applied to identify the ABCC8-associated immune signaling pathways. We found that the B-cell receptor signaling pathway (NES = −2.14, P FWER = 0.011) (Fig. 7A), antigen processing and presentation pathway (NES = −2.05, P FWER = 0.032) (Fig. 7B), leukocyte transendothelial migration pathway (NES = −1.93, P FWER = 0.074) (Fig. 7C), and FC gamma mediated phagocytosis pathway (NES = −1.92, P FWER = 0.084) (Fig. 7D) were all negatively correlated with ABCC8 expression. Interestingly, these immune pathways are matched to B cells, macrophages, neutrophils, and dendritic cells. These results further indicate that ABCC8 expression affects LGG immunity.
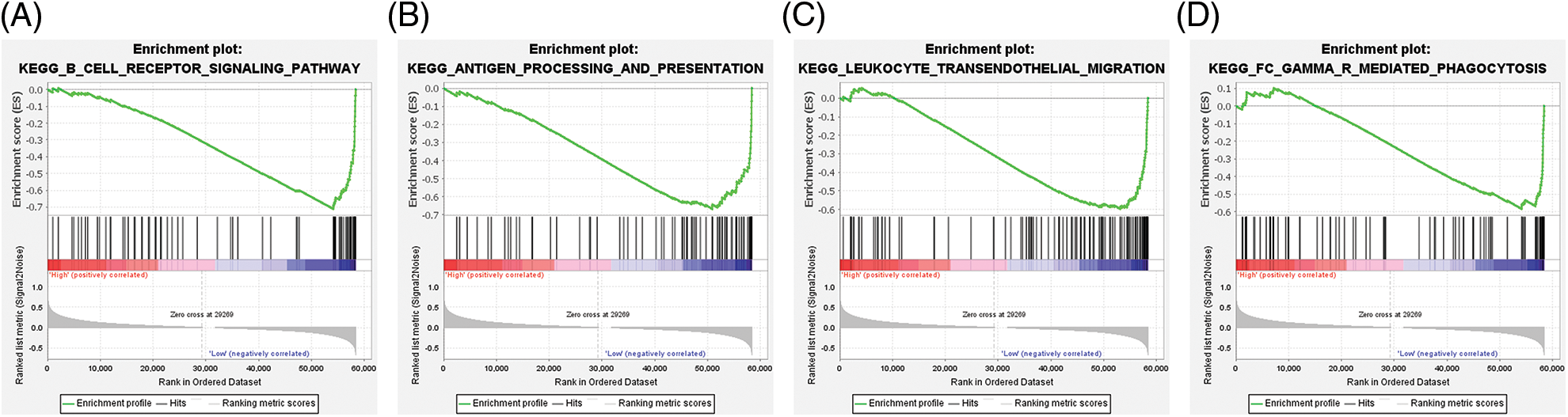
Figure 7: Gene set enrichment analysis of ABCC8-associated immune signaling pathways in LGG. A negative association was observed between ABCC8 mRNA levels and the B-cell receptor signaling pathway (A), antigen processing and presentation pathway (B), leukocyte transendothelial migration pathway (C), and FC gamma-mediated phagocytosis pathway (D).
We then explored the key molecular factors and signaling pathways by which ABCC8 might regulate the immune microenvironment in LGG. We built a network of the top 50 genes that were tightly correlated with ABCC8 using STRING (Fig. 8A). GO was used to annotate these genes (Fig. 8B), and the GABAergic synapse pathway was found to be related to ABCC8-mediated immune events (Fig. 8C).
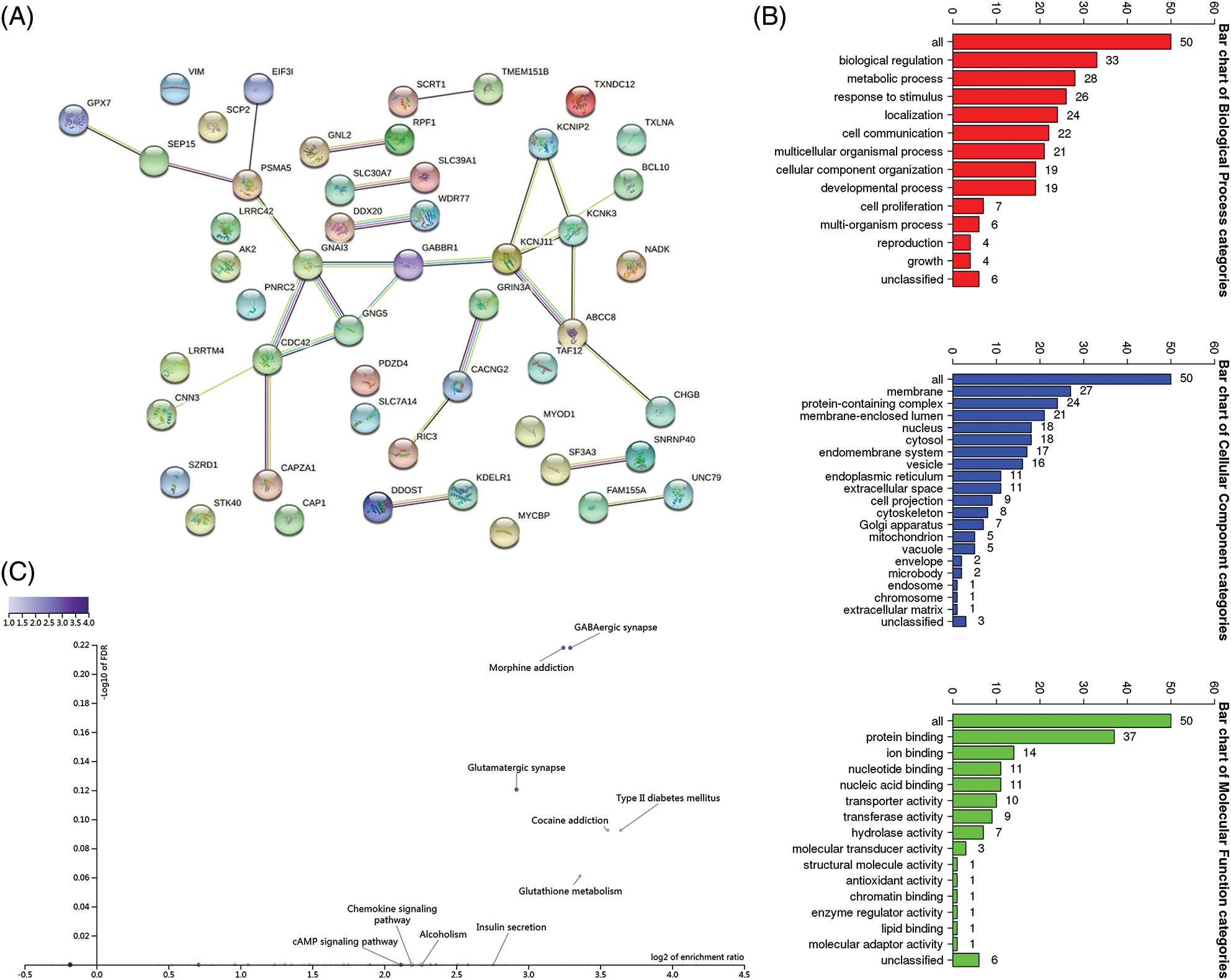
Figure 8: Integrated analysis of closely related genes associated with the ABCC8 gene. (A) The protein-protein network of ABCC8 and the top 50 ABCC8-correlated genes in LGG on the STRING website. (B) Gene Ontology annotation of ABCC8 and the top 50 ABCC8-correlated genes in LGG. (C) KEGG pathway analysis of the above-mentioned 51 genes.
ABCC8 encodes ATP binding cassette subfamily C member 8, also known as sulfonylurea receptor 1 (SUR1). It belongs to the ATP binding cassette transporter superfamily, which regulates multidrug resistance of cells by transporting various molecules across extra and intracellular membranes. Numerous studies have indicated the association between ABCC8 and diabetes, hypertension, cardiac diseases, and metabolic disorders (Rehman et al., 2022; Rehman et al., 2020). In the nerve system, ABCC8 has been found to control the activity of ATP-sensitive potassium channels (Martin et al., 2020). These channels are widely expressed in all cells of the neurovascular unit and induce the efflux of potassium to decrease neuronal excitability (Yamada and Inagaki, 2005; Zhang et al., 2018). ATP-sensitive potassium channels could also affect neurotransmitter release by the process mentioned above.
In general, ABCC8 was found to participate in the inflammatory process, resulting in cytotoxic edema and cell apoptosis in many neurological diseases, including ischemic stroke, spinal cord injury, subarachnoid hemorrhage, traumatic brain injury, and brain metastasis (Simard et al., 2006; Simard et al., 2009a; Simard et al., 2009b; Simard et al., 2007; Thompson et al., 2013). Glibenclamide, an ABCC8 inhibitor, was found to improve neurological functions and reduce mortality in patients with diabetic acute ischemic stroke (Gladstone et al., 2009; Kunte et al., 2012). Thompson et al. (2013) also reported that glibenclamide could reduce brain edema in a mouse model of brain metastasis (Thompson et al., 2013).
The putative role of ABCC8 in brain tumors has been shown earlier. Thompson et al. (2018) reported a higher expression of ABCC8 in medulloblastoma and supratentorial ependymoma than in glioblastoma, which suggests that the expression of ABCC8 is higher in benign brain tumors than in malignant tumors (Thompson et al., 2018). Recently, Zhou et al. (2020) found that ABCC8 expression levels are negatively related to the World Health Organization grade, 1p/19q noncodeletion, and IDH wild type in gliomas, while patients with high ABCC8 mRNA expression showed a longer survival period than others. However, the underlying mechanism has not been illustrated.
Our study found a significantly lowered ABCC8 mRNA expression in LGG than in normal tissues and was positively correlated with a favorable prognosis in LGG, consistent with previous studies. We also observed a negative correlation between ABCC8 mRNA expression and B cells, macrophages, neutrophils, and dendritic cells in LGG, which has not been discussed thus far. Varied ABCC8 gene copy numbers also seem to reflect different immune cell infiltration levels; however, owing to the relatively small mutation numbers, the results should be validated by a mouse model in vivo. ABCC8 expression was also found to be negatively correlated with markers of Treg and T-cell failure (PDCD1, CD274, CTLA4, LAG3, and HAVCR2/TIM3) and TMB, which may indicate that LGG patients without ABCC8 expression should have a good response to immune therapy. GSEA demonstrated a negative association between the expression of ABCC8 and the B-cell receptor signaling pathway, antigen processing and presentation pathway, FC gamma-mediated phagocytosis pathway, and leukocyte transendothelial migration pathway. Interestingly, these immune pathways matched only the B cells, macrophages, neutrophils, and dendritic cells that we found were related to ABCC8 expression. KEGG pathway analysis of ABCC8-related genes through TCGA_LGG data revealed that the GABAergic synapse signaling pathway is involved in the ABCC8-mediated immune response. Collectively, these findings imply that ABCC8 plays an important role in recruiting and governing non-T immune cells in LGG and may influence survival by regulating the antitumor immune response. To our knowledge, this is the first study to report a link between ABCC8 and tumor immunity in LGG.
There have been limited studies on the relationship between ABCC8 and immunity, with a focus on brain diseases. Thompson et al. (2018) reported that ABCC8 is a putative therapeutic target to reduce neuroinflammation in adult and pediatric brain tumors. Makar et al. (2015) found that silencing or inhibiting ABCC8 leads to a reduced inflammatory burden and correlates with better preservation of myelin, better preservation of axons, and more numerous mature and precursor oligodendrocytes in mouse models (Makar et al., 2015). In murine experimental autoimmune encephalomyelitis models, blockage of ABCC8-TRPM4 channels, expressed mostly by astrocytes, dramatically decreased the inflammatory response by downregulating TNF, BAFF, CCL2, and NOS2 mRNA (Gerzanich et al., 2017). Astrocytes only exist in the central neuron system, which may explain the results of our study. Regarding the studies focusing on ABCC8 and cancer immunity, there was only one study by Meng et al. (2022) was found on the PubMed website, which reported a positive correlation of ABCC8 with CD4+ T-cell and macrophage infiltration and a negative correlation with OS in liver hepatocellular carcinoma, totally opposite to our results, probably due to the different tumor backgrounds.
Recently, immunotherapy has revolutionized many kinds of solid malignant tumor treatments. Although its efficiency rate is not very high, it could bring great survival benefits to the responsive population. A major problem is finding the correct population. Luckily, ICIs have been proven to penetrate the blood-brain barrier and thus could be a promising immunotherapy strategy for LGG. Previous studies with glioma mouse models have proven that the inhibition of CTLA-4, IDO, or PD-L1 could significantly reduce tumor-infiltrating Treg cell numbers and increase survival, which indicates that ICIs may have wide application prospects in glioma treatment in the future (Wainwright et al., 2014). To date, some related clinical trials have indicated limited efficacy of ICIs, although most trials have not yet been completed (Xu et al., 2020). The mechanism underlying this paradox has not been illustrated. Our results showed that the TMB of LGG is quite low, so the anticancer immune response is probably inhibited because of a lack of antigen stimulation, which may partly explain the problem. Since the OS of LGG ranges from 1 to 15 years due to its large intrinsic biological and clinical heterogeneity, the detailed classification of LGG may help improve the efficiency of immunotherapy. Based on our present study, we found that ABCC8 may be a promising biomarker to predict the ICI treatment response.
We must acknowledge that there are several potential limitations in the present analysis. The study was a retrospective analysis, and all analyses were carried out based on public datasets. The results need to be validated in large, prospective studies. In addition, the mechanisms underpinning ABCC8-mediated antitumor immunity have not been fully illustrated. The GABAergic synapse signaling pathway, which has been proven to play a crucial role in immune cell immunomodulation by many studies, may be responsible for the phenomenon (Jin et al., 2013; Kim et al., 2018; Zheng et al., 2021). Finally, the average anticancer immune response is probably low in LGG, and whether modulators of ABCC8 or the GABAergic synapse signaling pathway can stimulate the immune response to provide survival benefits to LGG patients receiving ICIs remains to be elucidated. Subsequent experimental verification is required to test this hypothesis.
The results of this study suggest that ABCC8 might play a crucial role in regulating LGG tumor immunity. The mRNA expression levels of ABCC8 were independently predictive of OS in LGG, which indicated that ABCC8 could be a candidate biomarker for favorable survival. Non-T immune cells and the GABAergic synapse signaling pathway may participate in ABCC8-associated immune regulation. The potential role of ABCC8 inhibitors, such as glibenclamide, in interfering with immune cells should be evaluated. Whether a combination of ABCC8 inhibitors and ICIs could provide more survival benefits for LGG patients also needs to be discussed.
Acknowledgement: The results shown here are in part based upon data generated by the TCGA Research (https://www.cancer.gov/tcga), CGGA database (http://www.cgga.org.cn/index.jsp), and REMBRANDT database (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE108476).
Availability of Data and Materials: All datasets analyzed in the present study are open access. These data can be found on the following websites: Cancer Genome Atlas (https://portal.gdc.cancer.gov/), the GDC hub of UCSC Xena website (http://xena.ucsc.edu/public), and the CGGA database (http://www.cgga.org.cn/index.jsp).
Author Contribution: The authors confirm their contributions to the paper as follows: study conception and design: MJ; data collection: LG; analysis and interpretation of results: MJ, LG; draft manuscript preparation: MJ, LG. All authors reviewed the results and approved the final version of the manuscript.
Ethics Approval: Our study is based on open source data (TCGA, REMBRANDT, and CGGA). Ethical review and approval were not required for the study on human participants in accordance with the local legislation and institutional requirements. All methods were carried out in accordance with relevant guidelines and regulations.
Funding Statement: This work was supported by the Scientific and Technological Innovation Program for Clinical Medicine of Jinan (202019132) to LIPING GONG.
Conflicts of Interest: The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict with any competing interests.
References
Alquisiras-Burgos I, Ortiz-Plata A, Franco-Perez J, Millan A, Aguilera P (2020). Resveratrol reduces cerebral edema through inhibition of de novo SUR1 expression induced after focal ischemia. Experimental Neurology 330: 113353. DOI 10.1016/j.expneurol.2020.113353. [Google Scholar] [CrossRef]
Aran D, Sirota M, Butte AJ (2015). Systematic pan-cancer analysis of tumour purity. Nature Communications 6: 8971. DOI 10.1038/ncomms9971. [Google Scholar] [CrossRef]
Bao ZS, Chen HM, Yang MY, Zhang CB, Yu K et al. (2014). RNA-seq of 272 gliomas revealed a novel, recurrent PTPRZ1-MET fusion transcript in secondary glioblastomas. Genome Research 24: 1765–1773. DOI 10.1101/gr.165126.113. [Google Scholar] [CrossRef]
Beltrand J, Busiah K, Vaivre-Douret L, Fauret AL, Berdugo M, Cave H, Polak M (2020). Neonatal diabetes mellitus. Frontiers in Pediatrics 8: 540718. DOI 10.3389/fped.2020.540718. [Google Scholar] [CrossRef]
Bohnen MS, Ma L, Zhu N, Qi H, McClenaghan C et al. (2018). Loss-of-function ABCC8 mutations in pulmonary arterial hypertension. Circulation Genomic and Precision Medicine 11: e002087. DOI 10.1161/CIRCGEN.118.002087. [Google Scholar] [CrossRef]
Brat DJ, Pachter L (2015). Comprehensive, integrative genomic analysis of diffuse lower-grade gliomas. The New England Journal of Medicine 372: 2481–2498. DOI 10.1056/NEJMoa1402121. [Google Scholar] [CrossRef]
Chen L, Yang Z, Cao Y, Hu Y, Bao W, Wu D, Hu L, Xie J, Yu H (2021). Pan-cancer analysis and single-cell analysis revealed the role of ABCC5 transporter in hepatocellular carcinoma. Channels 15: 541–554. DOI 10.1080/19336950.2021.1968592. [Google Scholar] [CrossRef]
Chuah S, Chew V (2020). High-dimensional immune-profiling in cancer: Implications for immunotherapy. Journal for Immunotherapy of Cancer 8: e000363. DOI 10.1136/jitc-2019-000363. [Google Scholar] [CrossRef]
Claus EB, Walsh KM, Wiencke JK, Molinaro AM, Wiemels JL, Schildkraut JM, Bondy ML, Berger M, Jenkins R, Wrensch M (2015). Survival and low-grade glioma: The emergence of genetic information. Neurosurgical Focus 38: E6. DOI 10.3171/2014.10.FOCUS12367. [Google Scholar] [CrossRef]
Darlix A, Goze C, Rigau V, Bauchet L, Taillandier L, Duffau H (2017). The etiopathogenesis of diffuse low-grade gliomas. Critical Reviews in Oncology/Hematology 109: 51–62. DOI 10.1016/j.critrevonc.2016.11.014. [Google Scholar] [CrossRef]
de Franco E, Saint-Martin C, Brusgaard K, Knight Johnson AE, Aguilar-Bryan L et al. (2020). Update of variants identified in the pancreatic beta-cell KATP channel genes KCNJ11 and ABCC8 in individuals with congenital hyperinsulinism and diabetes. Human Mutation 41: 884–905. DOI 10.1002/humu.23995. [Google Scholar] [CrossRef]
Flagg TP, Patton B, Masia R, Mansfield C, Lopatin AN, Yamada KA, Nichols CG (2007). Arrhythmia susceptibility and premature death in transgenic mice overexpressing both SUR1 and Kir6.2[ΔN30,K185Q] in the heart. American Journal of Physiology Heart and Circulatory Physiology 293: H836–H845. DOI 10.1152/ajpheart.00011.2007. [Google Scholar] [CrossRef]
Gerzanich V, Makar TK, Guda PR, Kwon MS, Stokum JA et al. (2017). Salutary effects of glibenclamide during the chronic phase of murine experimental autoimmune encephalomyelitis. Journal of Neuroinflammation 14: 177. DOI 10.1186/s12974-017-0953-z. [Google Scholar] [CrossRef]
Gladstone DJ, Bui E, Fang J, Laupacis A, Lindsay MP, Tu JV, Silver FL, Kapral MK (2009). Potentially preventable strokes in high-risk patients with atrial fibrillation who are not adequately anticoagulated. Stroke 40: 235–240. DOI 10.1161/STROKEAHA.108.516344. [Google Scholar] [CrossRef]
Hlavac V, Brynychova V, Vaclavikova R, Ehrlichova M, Vrana D et al. (2013). The expression profile of ATP-binding cassette transporter genes in breast carcinoma. Pharmacogenomics 14: 515–529. DOI 10.2217/pgs.13.26. [Google Scholar] [CrossRef]
Hodges TR, Ott M, Xiu J, Gatalica Z, Swensen J et al. (2017). Mutational burden, immune checkpoint expression, and mismatch repair in glioma: Implications for immune checkpoint immunotherapy. Neuro-Oncology 19: 1047–1057. DOI 10.1093/neuonc/nox026. [Google Scholar] [CrossRef]
Hoshide R, Jandial R (2016). World health organization classification of central nervous system tumors: An era of molecular biology. World Neurosurgery 94: 561–562. DOI 10.1016/j.wneu.2016.07.082. [Google Scholar] [CrossRef]
Jin Z, Mendu SK, Birnir B (2013). GABA is an effective immunomodulatory molecule. Amino Acids 45: 87–94. DOI 10.1007/s00726-011-1193-7. [Google Scholar] [CrossRef]
Kim JK, Kim YS, Lee HM, Jin HS, Neupane C et al. (2018). GABAergic signaling linked to autophagy enhances host protection against intracellular bacterial infections. Nature Communications 9: 4184. DOI 10.1038/s41467-018-06487-5. [Google Scholar] [CrossRef]
Kunte H, Busch MA, Trostdorf K, Vollnberg B, Harms L, Mehta RI, Castellani RJ, Mandava P, Kent TA, Simard JM (2012). Hemorrhagic transformation of ischemic stroke in diabetics on sulfonylureas. Annals of Neurology 72: 799–806. DOI 10.1002/ana.23680. [Google Scholar] [CrossRef]
Liao Y, Wang J, Jaehnig EJ, Shi Z, Zhang B (2019). WebGestalt 2019: Gene set analysis toolkit with revamped UIs and APIs. Nucleic Acids Research 47: W199–W205. DOI 10.1093/nar/gkz401. [Google Scholar] [CrossRef]
Liu X, Li Y, Qian Z, Sun Z, Xu K et al. (2018). A radiomic signature as a non-invasive predictor of progression-free survival in patients with lower-grade gliomas. NeuroImage Clinical 20: 1070–1077. DOI 10.1016/j.nicl.2018.10.014. [Google Scholar] [CrossRef]
Makar TK, Gerzanich V, Nimmagadda VK, Jain R, Lam K et al. (2015). Silencing of Abcc8 or inhibition of newly upregulated Sur1-Trpm4 reduce inflammation and disease progression in experimental autoimmune encephalomyelitis. Journal of Neuroinflammation 12: 210. DOI 10.1186/s12974-015-0432-3. [Google Scholar] [CrossRef]
Martin GM, Sung MW, Shyng SL (2020). Pharmacological chaperones of ATP-sensitive potassium channels: Mechanistic insight from cryoEM structures. Molecular and Cellular Endocrinology 502: 110667. DOI 10.1016/j.mce.2019.110667. [Google Scholar] [CrossRef]
Meng X, Dong S, Yangyang L, Wang S, Xu X, Liu T, Zhuang X (2022). Adenosine triphosphate-binding cassette subfamily C members in liver hepatocellular carcinoma: Bioinformatics-driven prognostic value. Medicine 101: e28869. DOI 10.1097/MD.0000000000028869. [Google Scholar] [CrossRef]
Mohelnikova-Duchonova B, Brynychova V, Oliverius M, Honsova E, Kala Z, Muckova K, Soucek P (2013). Differences in transcript levels of ABC transporters between pancreatic adenocarcinoma and nonneoplastic tissues. Pancreas 42: 707–716. DOI 10.1097/MPA.0b013e318279b861. [Google Scholar] [CrossRef]
Nakasu S, Nakasu Y (2022). Malignant progression of diffuse low-grade gliomas: A systematic review and meta-analysis on incidence and related factors. Neurologia Medico-Chirurgica 62: 177–185. DOI 10.2176/jns-nmc.2021-0313. [Google Scholar] [CrossRef]
Nakasu S, Notsu A, Nakasu Y (2021). Prevalence of incidental meningiomas and gliomas on MRI: A meta-analysis and meta-regression analysis. Acta Neurochirurgica 163: 3401–3415. DOI 10.1007/s00701-021-04919-8. [Google Scholar] [CrossRef]
Rehman K, Niaz S, Tahir A, Jabeen K, Akash MSH (2022). FTO, PPAR-γ and ABCC8 gene variation and hypertension as determinants of cardiometabolic risk in CVD patients. Metabolism-Clinical and Experimental 128: 154972. DOI 10.1016/j.metabol.2021.154972. [Google Scholar] [CrossRef]
Rehman K, Tahir A, Niaz S, Shabbir S, Jabeen K, Faheem A, Akash MSH (2020). Frequency of PPAR-gamma, FTO and ABCC8 genetic variation in Pakistani cardiovascular smokers. Environmental Science and Pollution Research International 27: 42611–42620. DOI 10.1007/s11356-020-10226-z. [Google Scholar] [CrossRef]
Rizvi H, Sanchez-Vega F, La K, Chatila W, Jonsson P et al. (2018). Molecular determinants of response to anti-programmed cell death (PD)-1 and anti-programmed death-ligand 1 (PD-L1) blockade in patients with non-small-cell lung cancer profiled with targeted next-generation sequencing. Journal of Clinical Oncology 36: 633–641. DOI 10.1200/JCO.2017.75.3384. [Google Scholar] [CrossRef]
Simard JM, Chen M, Tarasov KV, Bhatta S, Ivanova S, Melnitchenko L, Tsymbalyuk N, West GA, Gerzanich V (2006). Newly expressed SUR1-regulated NC(Ca-ATP) channel mediates cerebral edema after ischemic stroke. Nature Medicine 12: 433–440. DOI 10.1038/nm1390. [Google Scholar] [CrossRef]
Simard JM, Geng Z, Woo SK, Ivanova S, Tosun C, Melnichenko L, Gerzanich V (2009a). Glibenclamide reduces inflammation, vasogenic edema, and caspase-3 activation after subarachnoid hemorrhage. Journal of Cerebral Blood Flow and Metabolism 29: 317–330. DOI 10.1038/jcbfm.2008.120. [Google Scholar] [CrossRef]
Simard JM, Kilbourne M, Tsymbalyuk O, Tosun C, Caridi J, Ivanova S, Keledjian K, Bochicchio G, Gerzanich V (2009b). Key role of sulfonylurea receptor 1 in progressive secondary hemorrhage after brain contusion. Journal of Neurotrauma 26: 2257–2267. DOI 10.1089/neu.2009.1021. [Google Scholar] [CrossRef]
Simard JM, Tsymbalyuk O, Ivanov A, Ivanova S, Bhatta S, Geng Z, Woo SK, Gerzanich V (2007). Endothelial sulfonylurea receptor 1-regulated NC Ca-ATP channels mediate progressive hemorrhagic necrosis following spinal cord injury. The Journal of Clinical Investigation 117: 2105–2113. DOI 10.1172/JCI32041. [Google Scholar] [CrossRef]
Southgate L, Machado RD, Graf S, Morrell NW (2020). Molecular genetic framework underlying pulmonary arterial hypertension. Nature Reviews Cardiology 17: 85–95. DOI 10.1038/s41569-019-0242-x. [Google Scholar] [CrossRef]
Sturm G, Finotello F, Petitprez F, Zhang JD, Baumbach J, Fridman WH, List M, Aneichyk T (2019). Comprehensive evaluation of transcriptome-based cell-type quantification methods for immuno-oncology. Bioinformatics 35: i436–i445. DOI 10.1093/bioinformatics/btz363. [Google Scholar] [CrossRef]
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL et al. (2005). Gene set enrichment analysis: A knowledge-based approach for interpreting genome-wide expression profiles. Proceedings of the National Academy of Sciences 102: 15545–15550. DOI 10.1073/pnas.0506580102. [Google Scholar] [CrossRef]
Szklarczyk D, Gable AL, Nastou KC, Lyon D, Kirsch R et al. (2021). The STRING database in 2021: Customizable protein-protein networks, and functional characterization of user-uploaded gene/measurement sets. Nucleic Acids Research 49: D605–D612. DOI 10.1093/nar/gkaa1074. [Google Scholar] [CrossRef]
Tang Z, Li C, Kang B, Gao G, Li C, Zhang Z (2017). GEPIA: A web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Research 45: W98–W102. DOI 10.1093/nar/gkx247. [Google Scholar] [CrossRef]
Thompson EM, Halvorson K, McLendon R (2018). Sulfonylurea receptor 1 expression is variable in adult and pediatric brain tumors. Clinical Neuropathology 37: 221–227. DOI 10.5414/NP301102. [Google Scholar] [CrossRef]
Thompson EM, Pishko GL, Muldoon LL, Neuwelt EA (2013). Inhibition of SUR1 decreases the vascular permeability of cerebral metastases. Neoplasia 15: 535–543. DOI 10.1593/neo.13164. [Google Scholar] [CrossRef]
Thurm C, Schraven B, Kahlfuss S (2021). ABC transporters in T cell-mediated physiological and pathological immune responses. International Journal of Molecular Sciences 22: 9186. DOI 10.3390/ijms22179186. [Google Scholar] [CrossRef]
van de Ven R, Oerlemans R, van der Heijden JW, Scheffer GL, de Gruijl TD, Jansen G, Scheper RJ (2009). ABC drug transporters and immunity: Novel therapeutic targets in autoimmunity and cancer. Journal of Leukocyte Biology 86: 1075–1087. DOI 10.1189/jlb.0309147. [Google Scholar] [CrossRef]
Wainwright DA, Chang AL, Dey M, Balyasnikova IV, Kim CK et al. (2014). Durable therapeutic efficacy utilizing combinatorial blockade against IDO, CTLA-4, and PD-L1 in mice with brain tumors. Clinical Cancer Research 20: 5290–5301. DOI 10.1158/1078-0432.CCR-14-0514. [Google Scholar] [CrossRef]
Wang Y, Qian T, You G, Peng X, Chen C et al. (2015). Localizing seizure-susceptible brain regions associated with low-grade gliomas using voxel-based lesion-symptom mapping. Neuro-Oncology 17: 282–288. DOI 10.1093/neuonc/nou130. [Google Scholar] [CrossRef]
Wang Z, Wang Z, Zhang C, Liu X, Li G et al. (2018). Genetic and clinical characterization of B7-H3 (CD276) expression and epigenetic regulation in diffuse brain glioma. Cancer Science 109: 2697–2705. DOI 10.1111/cas.13744. [Google Scholar] [CrossRef]
Wang Z, Xu H, Zhu L, He T, Lv W, Wu Z (2020). Establishment and evaluation of a 6-Gene survival risk assessment model related to lung adenocarcinoma microenvironment. BioMed Research International 2020: 6472153. DOI 10.1155/2020/6472153. [Google Scholar] [CrossRef]
Xu S, Tang L, Li X, Fan F, Liu Z (2020). Immunotherapy for glioma: Current management and future application. Cancer Letters 476: 1–12. DOI 10.1016/j.canlet.2020.02.002. [Google Scholar] [CrossRef]
Yamada K, Inagaki N (2005). Neuroprotection by KATP channels. Journal of Molecular and Cellular Cardiology 38: 945–949. DOI 10.1016/j.yjmcc.2004.11.020. [Google Scholar] [CrossRef]
Yin W, Jiang X, Tan J, Xin Z, Zhou Q et al. (2020). Development and validation of a tumor mutation burden-related immune prognostic model for lower-grade glioma. Frontiers in Oncology 10: 1409. DOI 10.3389/fonc.2020.01409. [Google Scholar] [CrossRef]
Youssef G, Miller JJ (2020). Lower grade gliomas. Current Neurology and Neuroscience Reports 20: 21. DOI 10.1007/s11910-020-01040-8. [Google Scholar] [CrossRef]
Zhang Q, Li C, Zhang T, Ge Y, Han X, Sun S, Ding J, Lu M, Hu G (2018). Deletion of Kir6.2/SUR1 potassium channels rescues diminishing of DA neurons via decreasing iron accumulation in PD. Molecular and Cellular Neurosciences 92: 164–176. DOI 10.1016/j.mcn.2018.08.006. [Google Scholar] [CrossRef]
Zhao Z, Meng F, Wang W, Wang Z, Zhang C, Jiang T (2017). Comprehensive RNA-seq transcriptomic profiling in the malignant progression of gliomas. Scientific Data 4: 170024. DOI 10.1038/sdata.2017.24. [Google Scholar] [CrossRef]
Zheng Z, Zhang X, Liu J, He P, Zhang S et al. (2021). GABAergic synapses suppress intestinal innate immunity via insulin signaling in Caenorhabditis elegans. Proceedings of the National Academy of Sciences 118: e2021063118. DOI 10.1073/pnas.2021063118. [Google Scholar] [CrossRef]
Zhou K, Liu Y, Zhao Z, Wang Y, Huang L, Chai R, Li G, Jiang T (2020). ABCC8 mRNA expression is an independent prognostic factor for glioma and can predict chemosensitivity. Scientific Reports 10: 12682. DOI 10.1038/s41598-020-69676-7. [Google Scholar] [CrossRef]
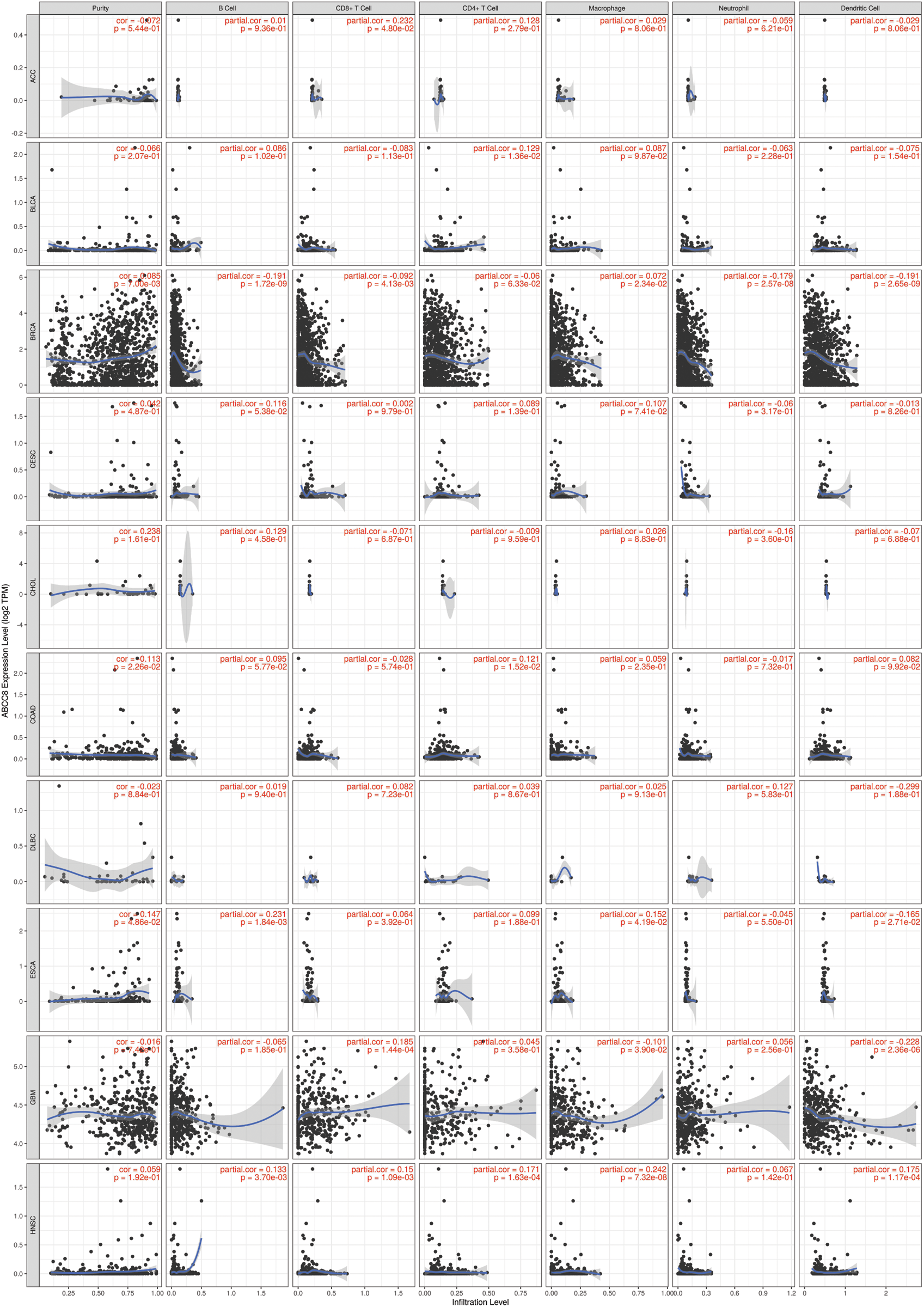
Figure S1: Association between ABCC8 expression levels and immune infiltration levels in ACC, BLCA, BRCA, CESC, CHOL, COAD, DLBC, ESCA, GBM, and HNSC.
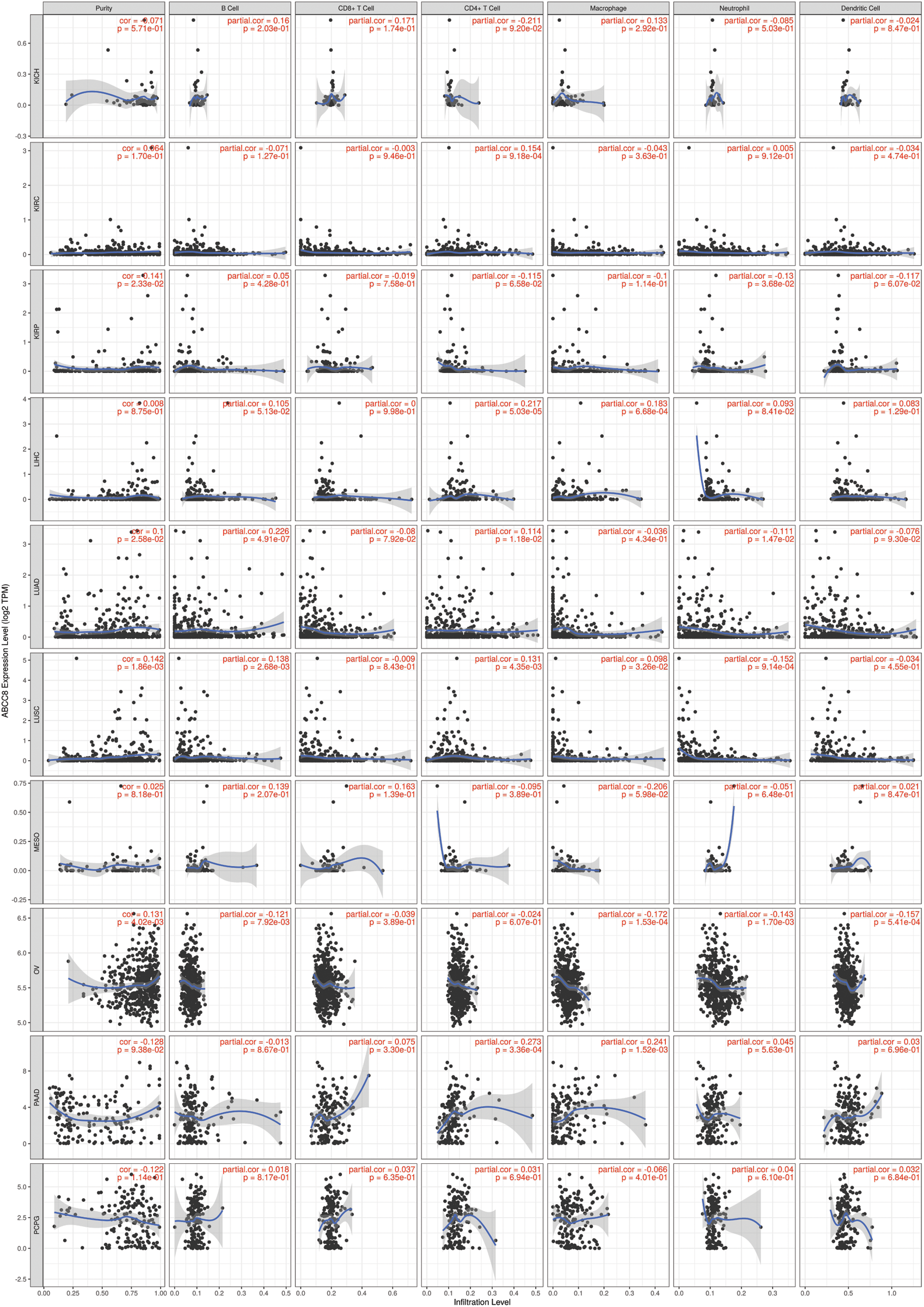
Figure S2: Association between ABCC8 expression levels and immune infiltration levels in KICH, KIRC, KIRP, LIHC, LUAD, LUSC, MESO, OV, PAAD, and PCPG.
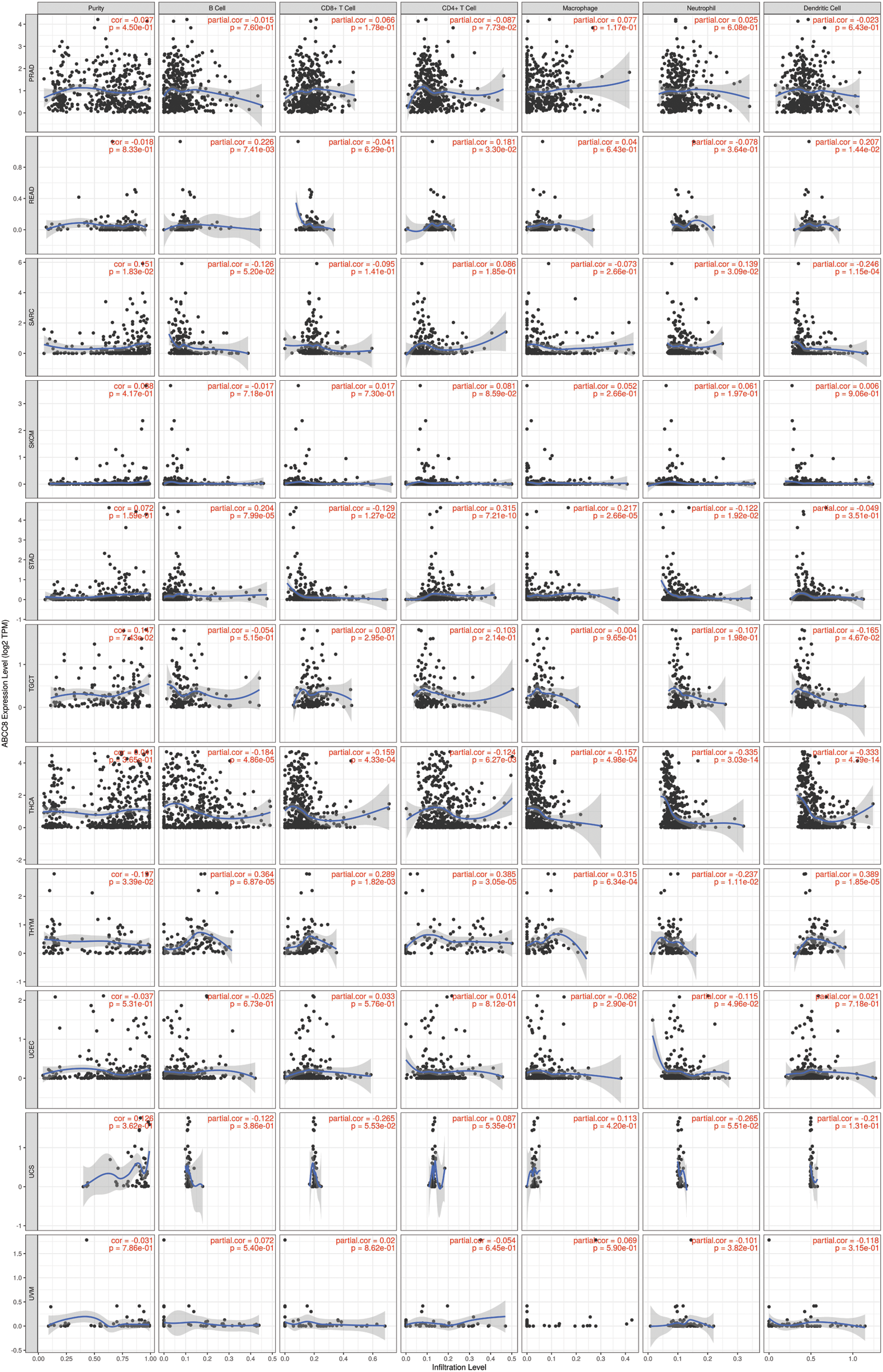
Figure S3: Association between ABCC8 expression levels and immune infiltration levels in PRAD, READ, SARC, SKCM, STAD, TGCT, THCA, THYM, UCEC, UCS and UVM.
Cite This Article
 Copyright © 2023 The Author(s). Published by Tech Science Press.
Copyright © 2023 The Author(s). Published by Tech Science Press.This work is licensed under a Creative Commons Attribution 4.0 International License , which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.


 Submit a Paper
Submit a Paper Propose a Special lssue
Propose a Special lssue View Full Text
View Full Text Download PDF
Download PDF Downloads
Downloads
 Citation Tools
Citation Tools