 Open Access
Open Access
ARTICLE
Evaluation of ASM for Ventricular Segmentation in Patients with Diverse Cardiac Abnormalities
1 Faculty of Electrical Engineering, Automatics, Computer Science and Biomedical Engineering, AGH University of Krakow, Kraków, Poland
2 Department of Biocybernetics and Biomedical Engineering, AGH University of Krakow, Kraków, Poland
3 Faculty of Geology, Geophysics and Environmental Protection, AGH University of Krakow, Kraków, Poland
4 Department of Algorithmics and Software, Silesian University of Technology, Gliwice, Poland
* Corresponding Author: Adam Piórkowski. Email:
(This article belongs to the Special Issue: Artificial Intelligence in Visual and Audio Signal Processing)
Computers, Materials & Continua 2026, 87(3), 86 https://doi.org/10.32604/cmc.2026.076062
Received 13 November 2025; Accepted 06 March 2026; Issue published 09 April 2026
Abstract
The efficacy of Active Shape Models (ASM) for automated ventricular segmentation was evaluated to address the computational demands of manual segmentation and the interpretability limitations of deep learning. A statistical shape model was constructed using a limited cohort of 19 Coronary Computed Tomography Angiography (CCTA) scans derived from patients with diverse cardiac abnormalities. Principal Component Analysis (PCA) was employed to encapsulate morphological variability, and strict point correspondence was enforced to maintain topological consistency. Validation was conducted via leave-one-out cross-validation, benchmarking automated segmentations against expert-delineated ground truths using the Dice Similarity Coefficient (DSC) and Hausdorff Distance (HD). It was found that mean Dice scores ranged from 0.50 to 0.64, with individual high-fidelity cases achieving scores up to 0.84. These results indicated that while quantitative performance reflected the complexity of pathological morphology, the methodology successfully accommodated high morphometric variance. It can be concluded that the ASM framework provides a resilient, interpretable foundation for managing complex clinical geometry where unconstrained models may fail.Keywords
Medical imaging data represents an invaluable asset for practitioners, facilitating the visualization and detailed analysis of complex internal physiological structures [1]. Currently, these modalities are extensively utilized within cardiology for the diagnosis and longitudinal monitoring of cardiovascular diseases [1]. Standard imaging techniques employed to evaluate cardiac morphology and function include echocardiography, magnetic resonance imaging (MRI), and computed tomography (CT). To enhance clinical utility, various post-processing methodologies are applied to these acquired datasets [2]. Specifically, the generation of patient-specific 3D models of anatomical structures enables clinicians to better comprehend complex pathologies and formulate informed therapeutic strategies [3]. Consequently, high-fidelity cardiac segmentation is a prerequisite for the quantitative assessment of cardiovascular health [2].
The heart’s right ventricle (RV) is responsible for propelling deoxygenated blood to the pulmonary circulation; thus, any deviation in its morphology serves as a critical indicator of underlying cardiac dysfunction [4]. Therefore, the precise segmentation of both the RV and the left ventricle (LV) from imaging data is essential for various clinical applications, ultimately leading to improved patient outcomes [5]. While automated segmentation has reached significant milestones, patients presenting with irregular cardiac geometry and structural abnormalities remain a formidable challenge for contemporary models, which are predominantly trained on “idealized” or healthy datasets [4].
To address these limitations, this study evaluates the efficacy of the Active Shape Model (ASM) algorithm for the automated segmentation of ventricles using Coronary Computed Tomography Angiography (CCTA) scans. The primary novelty of this research lies in the rigorous evaluation of the ASM framework within a cohort diagnosed with diverse and severe cardiac conditions. This methodology was specifically selected to overcome the inherent limitations of many Convolutional Neural Networks (CNNs). Although CNNs are powerful, they are frequently characterized as “black-box” systems; the intricate relationships between their internal neurons and weights are opaque, complicating the identification of segmentation logic or the root causes of error. Such a lack of interpretability is a significant concern in clinical environments, where transparency in diagnostic tools is paramount. In contrast, the ASM utilizes statistical shape modeling and Principal Component Analysis (PCA) to encapsulate variability in complex 3D structures while strictly enforcing anatomical constraints [6]. Given that the mathematical foundations of ASM are well-documented and transparent, this approach facilitates a more interpretable analysis of pathological cardiac images.
The Active Shape Model (ASM) algorithm has constituted a cornerstone of medical image analysis for nearly three decades, providing a robust framework for integrating statistical shape priors with the spatial appearance of target structures [6]. Researchers have extensively employed this model-based approach to mitigate the challenges of manual segmentation, specifically inter-observer variability and the prohibitive labor costs associated with expert annotation [7]. Notably, recent reviews by Khan et al. provide an overview of the ongoing use of three-dimensional Active Shape Models for left ventricle segmentation in cardiac computed tomography [8]. While linear models offered initial computational efficiency, recent hybrid ASM variants have introduced a more flexible representation of shape variations, which is crucial when delineating anatomical structures exhibiting complex pathological deformations [9].
A fundamental challenge in this domain arises from the significant morphological disparity between the ventricular chambers. The LV is characterized by a thick, symmetric myocardial wall that generates high-contrast intensity profiles [9]. Conversely, the right ventricle (RV) is distinguished by a significantly thinner wall and a complex, crescentic geometry that partially envelops the LV. These morphological characteristics, combined with the RV’s susceptibility to shading artifacts and extreme variability in diseased states, render it a substantially more challenging target for automated segmentation compared to the LV. To mitigate these difficulties, recent methodologies have sought to augment segmentation accuracy by leveraging clustering algorithms and diverse statistical priors [10].
In the contemporary landscape of medical imaging, Convolutional Neural Networks (CNNs) have emerged as the dominant paradigm for cardiac segmentation [11,12]. These architectures are highly effective at partitioning images into semantically distinct regions, facilitating the quantitative assessment of myocardial mass and ejection fraction [13]. In healthy cohorts, conventional architectures such as U-Net frequently achieve high Dice Similarity Coefficients (DSC), often exceeding 0.90 [14]. However, baseline CNNs may struggle to replicate expert-level performance when restricted to limited datasets. This limitation is intrinsic to the high-dimensional parameter space of deep learning architectures; without a vast repository of training examples to enforce feature invariance, these unconstrained models are prone to overfitting and frequently fail to preserve global topological constraints when generalized to pathological outliers or severe cardiac abnormalities. To enhance performance, recent studies have incorporated edge-attention mechanisms to prioritize boundary feature extraction [14]. Additionally, surface-mesh reconstruction techniques have been explored, demonstrating advantages over traditional voxel-based methods such as Voxel2Mesh [15].
Despite their efficacy, CNNs are frequently characterized as “black-box” algorithms due to the opacity of the intricate connections between neurons and weights [16,17]. This lack of interpretability constitutes a significant impediment to clinical adoption, where understanding the algorithmic rationale is critical for diagnosis. Furthermore, lacking the global anatomical priors inherent to ASM, CNNs may generate anatomically inconsistent outputs, such as disconnected voxel regions or irregular boundaries. Given that the mathematical formulation of ASM is both transparent and well-documented, it remains a vital tool for consistent, interpretable analysis in medical imaging [6,10].
The pursuit of high precision coupled with interpretability has driven the development of hybrid models. These strategies often integrate U-Net architectures for initial feature extraction with multiphase active contours for adaptive refinement [18]. Other hybrid approaches combine deep learning with multi-atlas mapping and geometric modeling to delineate substructures such as valves and coronary arteries [19]. Although these methods demonstrate high consistency for smaller structures, accurately segmenting patients with irregular geometries and structural abnormalities remains a persistent challenge. This study addresses this gap by establishing a versatile ASM framework capable of segmenting ventricles with diverse, abnormal morphologies using a training dataset derived directly from clinical pathology.
The proposed framework employs a 3D Active Shape Model (ASM) for the segmentation of both the left and right ventricles, constructed from manual annotations of a pilot training cohort (n = 19). Although the dataset is sparse, these models are explicitly engineered to encode complex morphological variability while enforcing global anatomical constraints. This ensures that the generated segmentations remain physiologically plausible, even in the presence of severe pathology. The ASM framework was selected for its mathematical transparency and its capacity to integrate expert anatomical knowledge through strict point correspondence. This provides a robust, interpretable alternative to data-intensive “black-box” architectures, which frequently exhibit poor generalization in clinical scenarios characterized by data scarcity.
The training dataset comprised 19 Coronary Computed Tomography Angiography (CCTA) scans acquired from patients diagnosed with diverse cardiac pathologies (n = 19). The dataset analyzed in this study is a pre-existing, fully anonymized dataset provided by a third party (MST). All data were de-identified by the provider prior to being accessed by the authors, ensuring that no individual participants can be identified. The data are not publicly available due to privacy. Each scan was associated with a segmentation mask delineating the boundaries of 13 anatomical structures. These masks were initially annotated by medical professionals and subsequently validated by independent clinical experts to ensure ground-truth fidelity. Fig. 1 illustrates representative data, displaying an acquired CCTA scan alongside its corresponding expert-verified segmentation. This integration of high-resolution imaging with precise annotations is intended to capture the significant morphological heterogeneity of ventricular anatomy within the patient cohort. Fig. 2 presents a three-dimensional reconstruction of the ventricles extracted from a single scan, highlighting the geometric complexity modeled in this study.
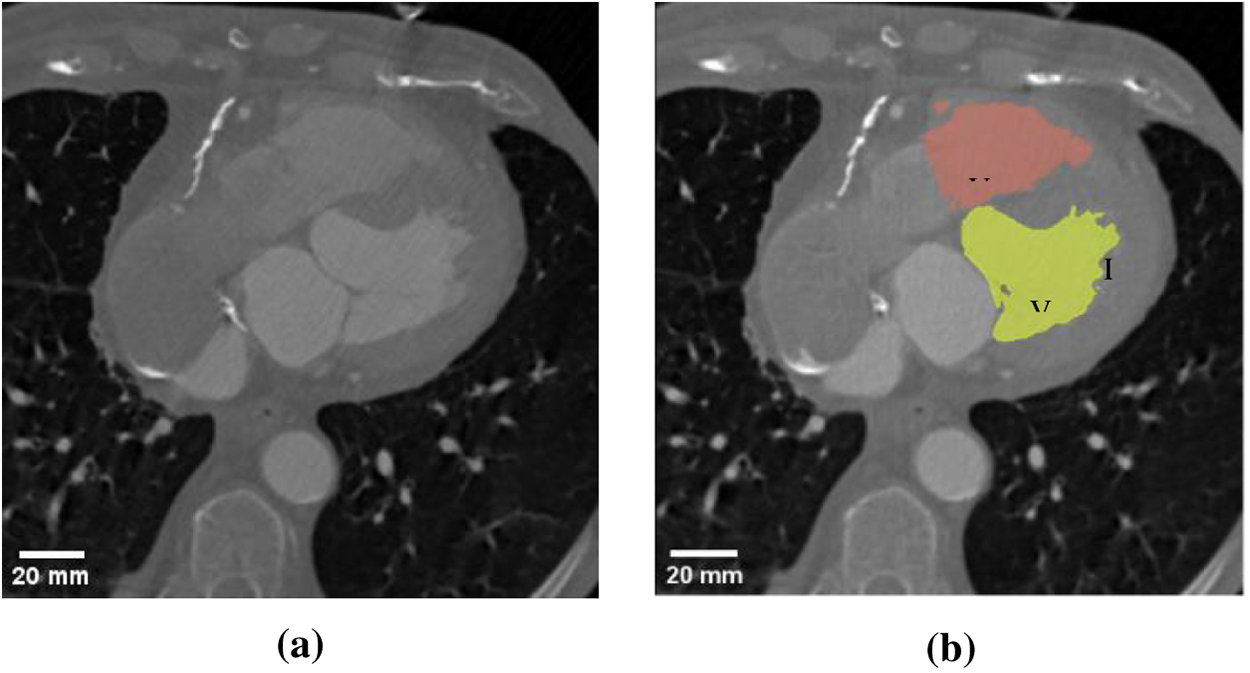
Figure 1: Representative sample of used data (a) Computed tomography scan; (b) Manual segmentation mask.
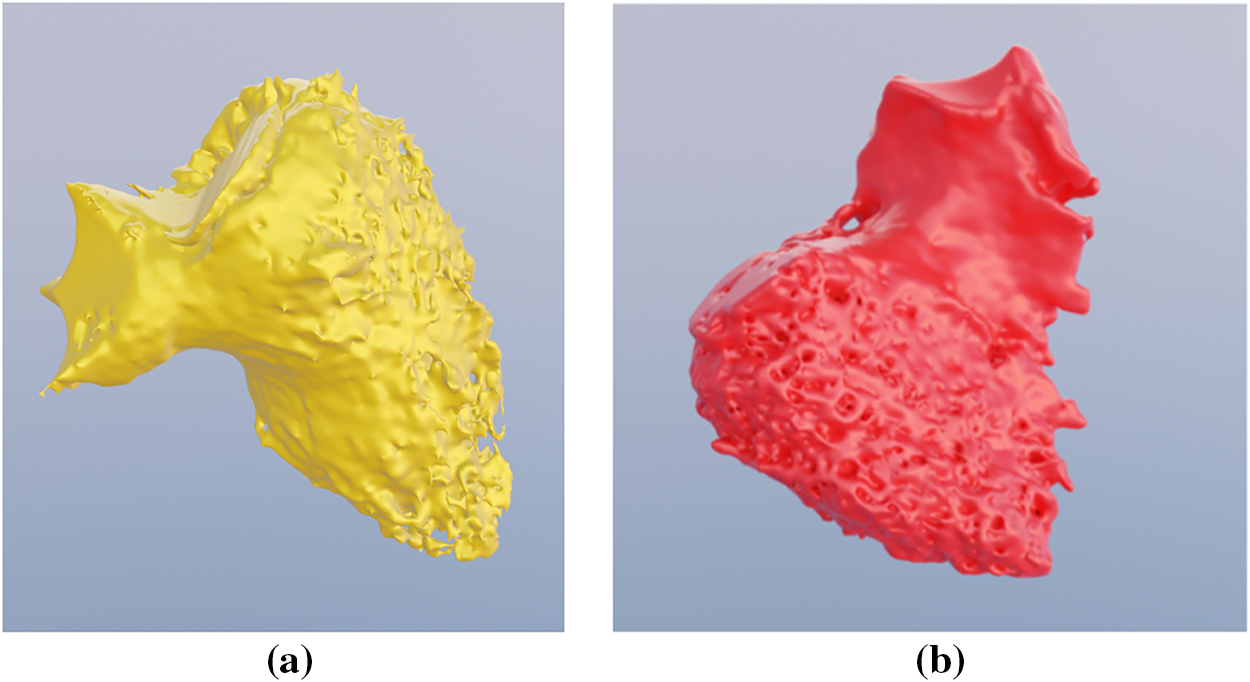

Figure 2: Three-dimensional representation (a) Left ventricle; (b) Right ventricle.
The source images were provided in DICOM (Digital Imaging and Communication in Medicine) format with a resolution of 512 × 512 pixels and an average voxel dimension of 0.4 mm × 0.4 mm × 0.4 mm. During the preprocessing phase, the original 16-bit depth was rescaled to 8-bit. This reduction was implemented primarily to optimize computational efficiency and accelerate model convergence. Two distinct normalization techniques were evaluated to assess their impact on the statistical appearance model: the Division Normalization (pixel intensities scaled by a factor of 1/16 and truncated to the nearest integer) and the MinMax Normalization (intensity values mapped linearly from their dynamic range to a standard 0–255 scale). The selection of a scaling strategy is pivotal, as the ASM algorithm relies on intensity profiles sampled perpendicular to contour points to update landmark positions. Since the magnitude of these profiles varies significantly between rescaling methods, both approaches were analyzed to determine their effect on gradient strength at anatomical boundaries. Additionally, to maintain coordinate consistency throughout the processing pipeline, the Hausdorff distance metrics utilized in subsequent validation steps are reported in pixels.
The Active Shape Model algorithm, originally established by Cootes and Taylor, provides a framework for segmenting intricate anatomical structures by synthesizing statistical shape priors with deformable model refinement [6]. This approach is particularly advantageous for cardiac modeling as it incorporates both global shape constraints and local pixel intensity profiles to navigate the complex morphology of the ventricles [20].
The foundation of the ASM is a mean shape model derived from the training set, capturing inherent variability in size and spatial orientation (Fig. 3). Each shape is defined by a vector of K landmark points
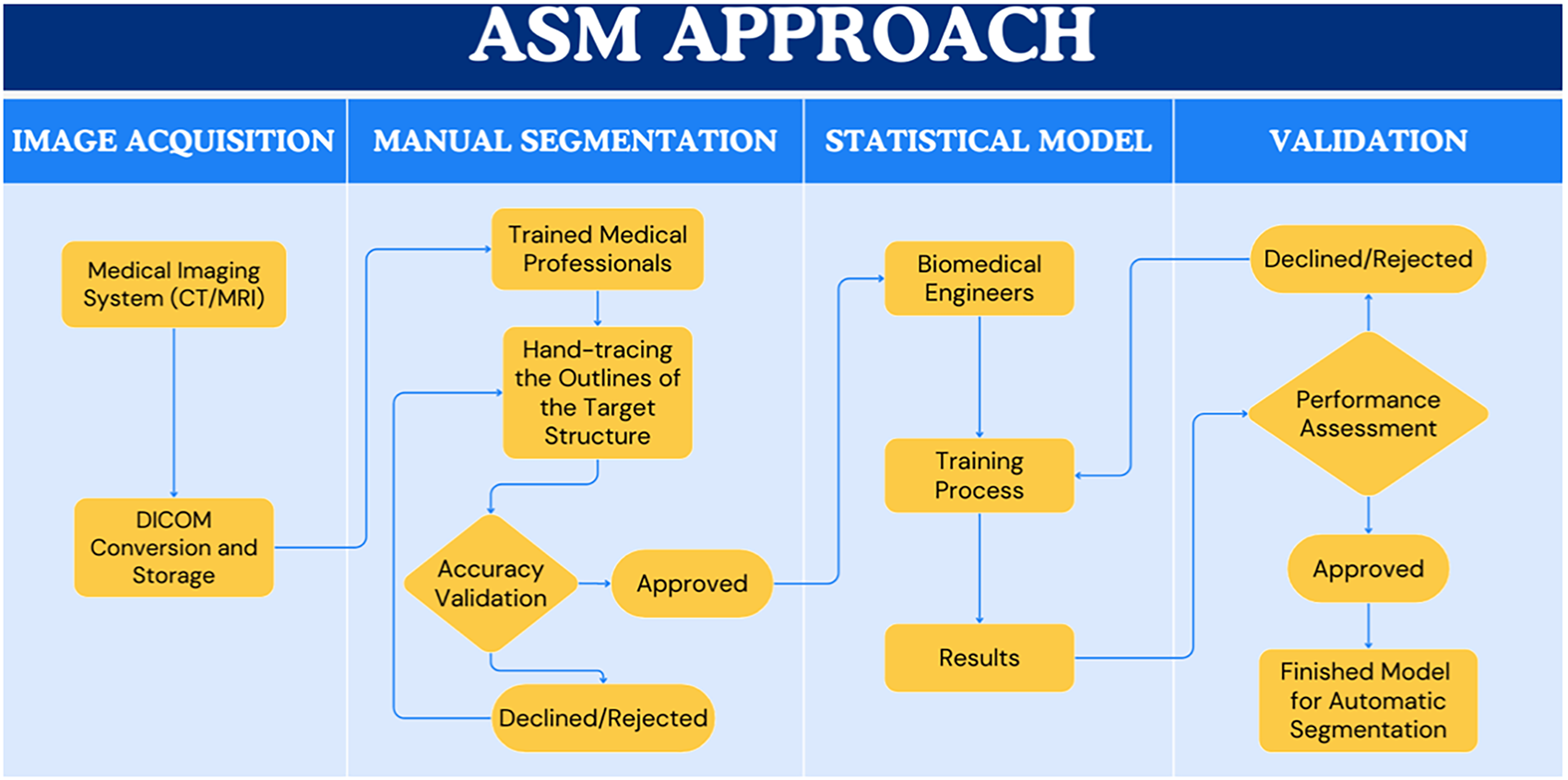
Figure 3: Core premise of the ASM approach.
To isolate intrinsic shape variability from pose differences, Procrustes analysis is utilized to align the shapes by eliminating translational, rotational, and scale variations. Rotational alignment is achieved by multiplying the shape vector by a rotation matrix R, as described in Eq. (1):
Shape dissimilarity is quantified using the Procrustes distance d, defined as the square root of the sum of squared differences between corresponding landmark coordinates (Eq. (2)).
where
The segmentation of novel scans is governed by an iterative energy minimization process. This refinement is driven by a total energy function E(x), which balances detected image features against the allowable statistical shape space (Eq. (3)):
here,
The ASM implementation was executed in MATLAB R2022b. All computational experiments were conducted on a workstation equipped with an Intel Core i7-9700KF processor (3.60 GHz), 64 GB of RAM, and an NVIDIA GeForce RTX 2070 SUPER graphics card. The algorithm utilized a high-fidelity template to encapsulate the morphological variability of the pathological cohort. For the statistical engine to accurately isolate shape variations via Principal Component Analysis (PCA), strict point correspondence is mathematically essential. This requires every model in the training set to possess an identical vertex count and topological connectivity.
The preprocessing workflow is outlined in Fig. 4. Initial ventricular meshes were extracted from segmentation masks using 3D Slicer; however, these meshes exhibited inconsistent vertex counts and connectivity. To establish a standardized reference, a uniform template of the left and right ventricles was manually sculpted in Blender 4.1 (Figs. 5 and 6). The resulting template, consisting of 4917 vertices and 9830 triangular faces, served as the baseline for all cases. This template was aligned to each patient-specific anatomy and fitted using elastic deformation and the shrinkwrap modifier, an iterative tool that non-rigidly projects the source mesh onto the target surface. However, due to the irregular ventricular geometry in the pathological dataset, the automated modifier frequently exhibited limitations in complex anatomical regions. Specifically, the tool often failed to accurately delineate the aortic and pulmonary valves and tended to overshoot the ventricular apex. Consequently, while the shrinkwrap modifier provided a preliminary approximation, manual expert deformation was required for every case to correct these misalignments. This intervention ensured that critical landmarks maintained correct spatial ordering, preventing the randomized point placement common in automated registration protocols.
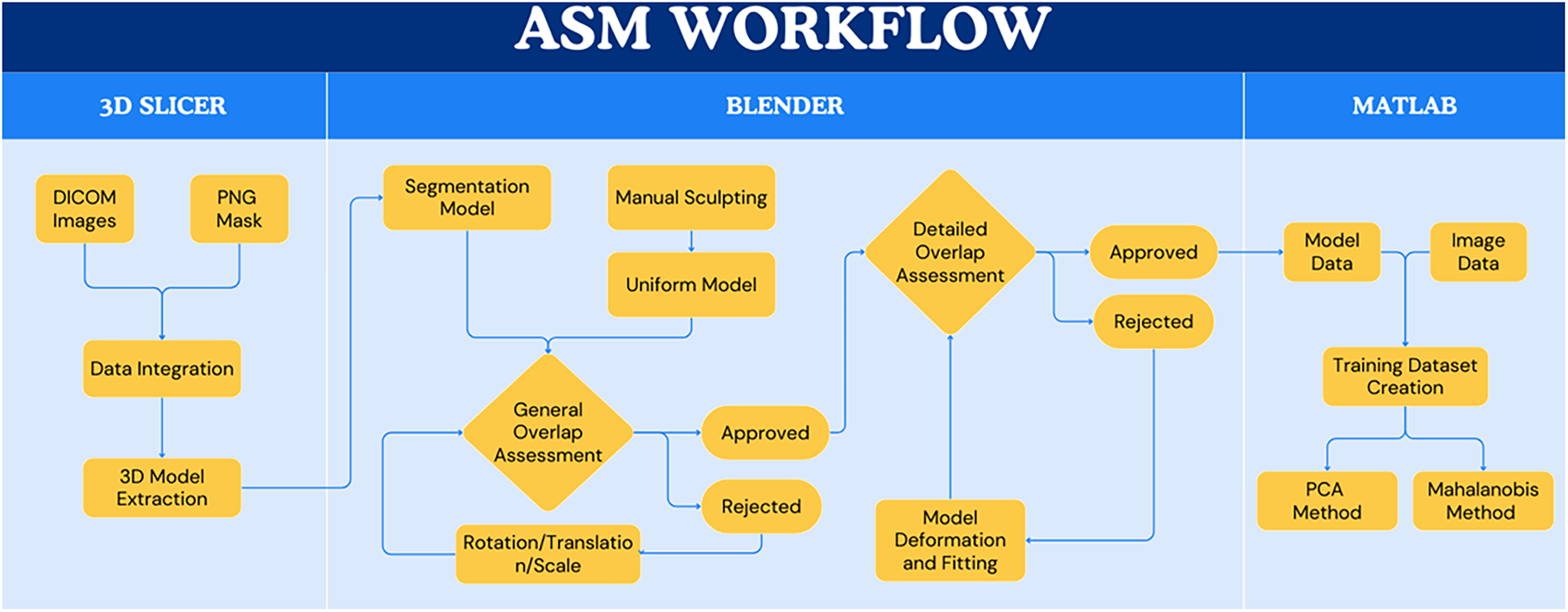
Figure 4: Workflow for ventricular segmentation using Active Shape Model.
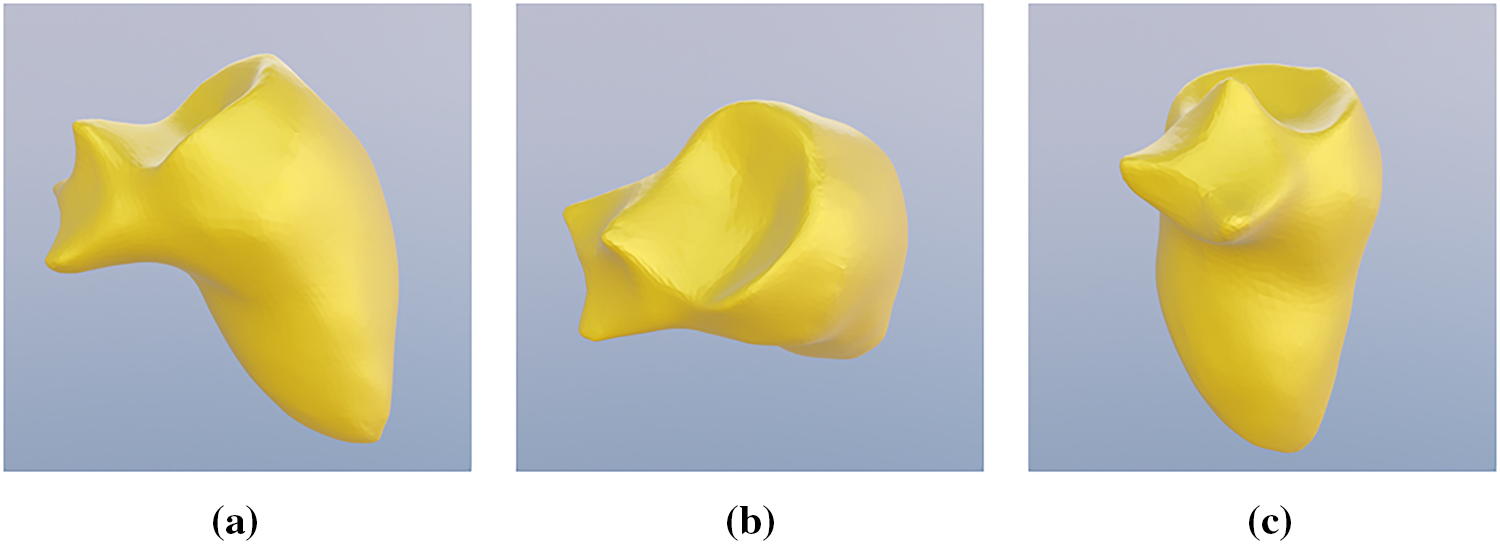

Figure 5: Manually sculpted template of a generic left ventricle, different perspective (a) side view; (b) top view; (c) front view.
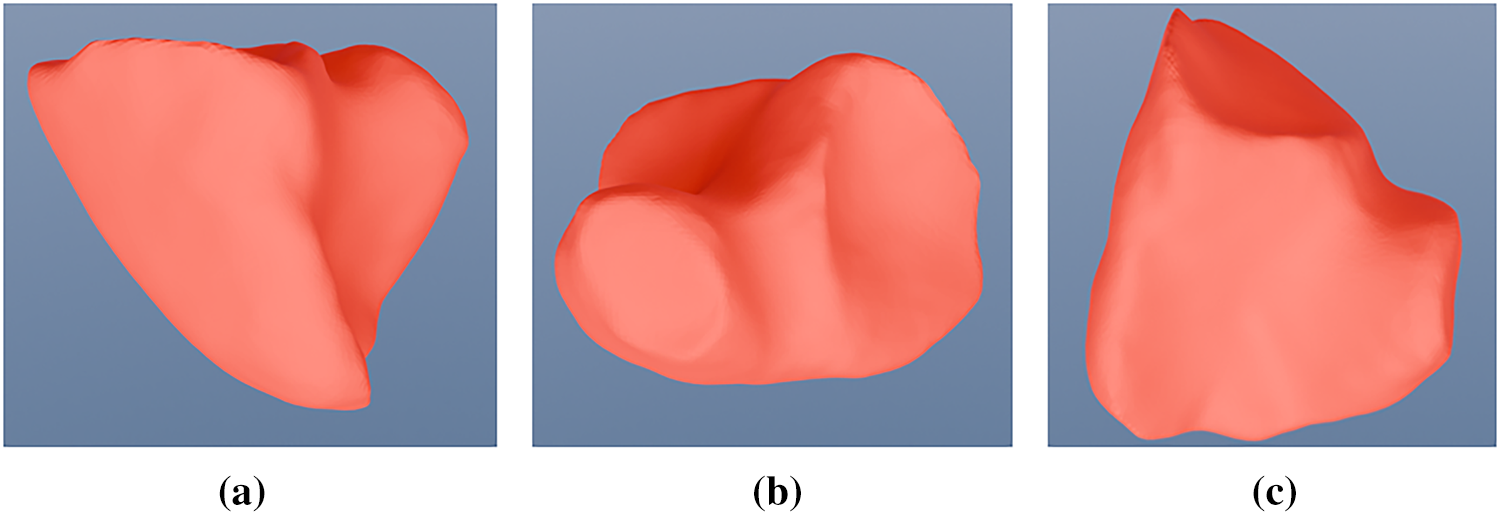
Figure 6: Manually sculpted template of a generic right ventricle, different perspective (a) side view; (b) top view; (c) back view.
The performance of the ASM algorithm was evaluated using leave-one-out cross-validation. In this framework, 19 distinct statistical models were generated; each was trained on 18 scans and subsequently validated against the remaining scan. This iterative process mitigated the risk of randomized outcomes and confirmed that the model learned meaningful, generalized shape statistics from the pathological data. Automated segmentations were benchmarked against manual segmentations created by expert clinicians, which served as the ground truth.
Accuracy was quantified using two primary metrics: the Dice Similarity Coefficient (DSC) and the Hausdorff Distance (HD). The DSC measures the volumetric overlap between the automated segmentation (X) and the manual segmentation (Y) (Eq. (4)).
A DSC value of 1.0 indicates perfect spatial alignment. In the context of complex cardiac structures with significant abnormalities, a coefficient exceeding 0.70 is typically regarded as acceptable for clinical research.
To assess local boundary precision and identify spatial discrepancies, the Hausdorff distance was also calculated. Unlike the DSC, which measures aggregate overlap, the HD quantifies the maximum distance between corresponding points in two sets (A and B), rendering it highly sensitive to outliers and localized segmentation errors (Eq. (5)).
where
To ensure consistency with the coordinate system of the 512 × 512 scan matrices, all Hausdorff distance metrics are reported in pixels. Following quantitative assessment, models were exported to Blender 4.1 for 3D visualization and secondary topological analysis to verify anatomical fidelity.
The evaluation of the Active Shape Model framework focused on three primary variables: the impact of image rescaling techniques, the efficacy of specific appearance modeling strategies, and the influence of vertex density on segmentation fidelity. Initial benchmarks confirmed that rescaling original 16-bit images to an 8-bit depth significantly attenuated computational overhead; average training durations decreased from 8 min 13 s to approximately 7 min 45 s for both the MinMax and Division methods.
4.1 Intensity Profile Analysis
To quantify the influence of intensity rescaling on the landmark update mechanism, mean intensity profiles were generated across the training cohort at multiple scaling factors (1.0, 0.5, and 0.25). These profiles, visualized in Figs. 7–9, illustrate the gradient characteristics utilized to guide the ASM deformation. In these plots, the ordinate represents mean pixel intensity, while the abscissa denotes the sampling position along the surface normal at each landmark.
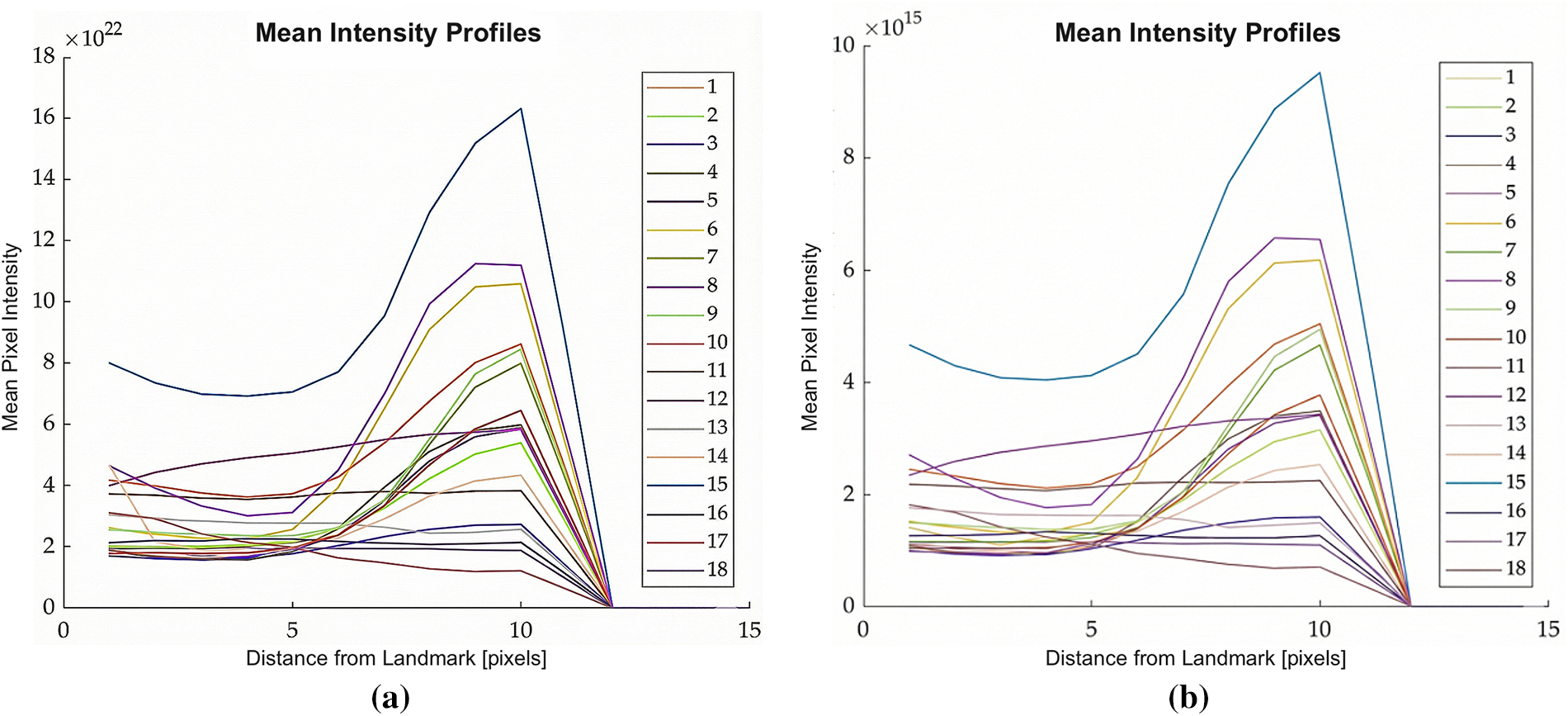
Figure 7: Plots showing average intensity values perpendicular to the contour points across all the training images used to build your ASM model, scaling factor: 1, (a) Division; (b) MinMax.
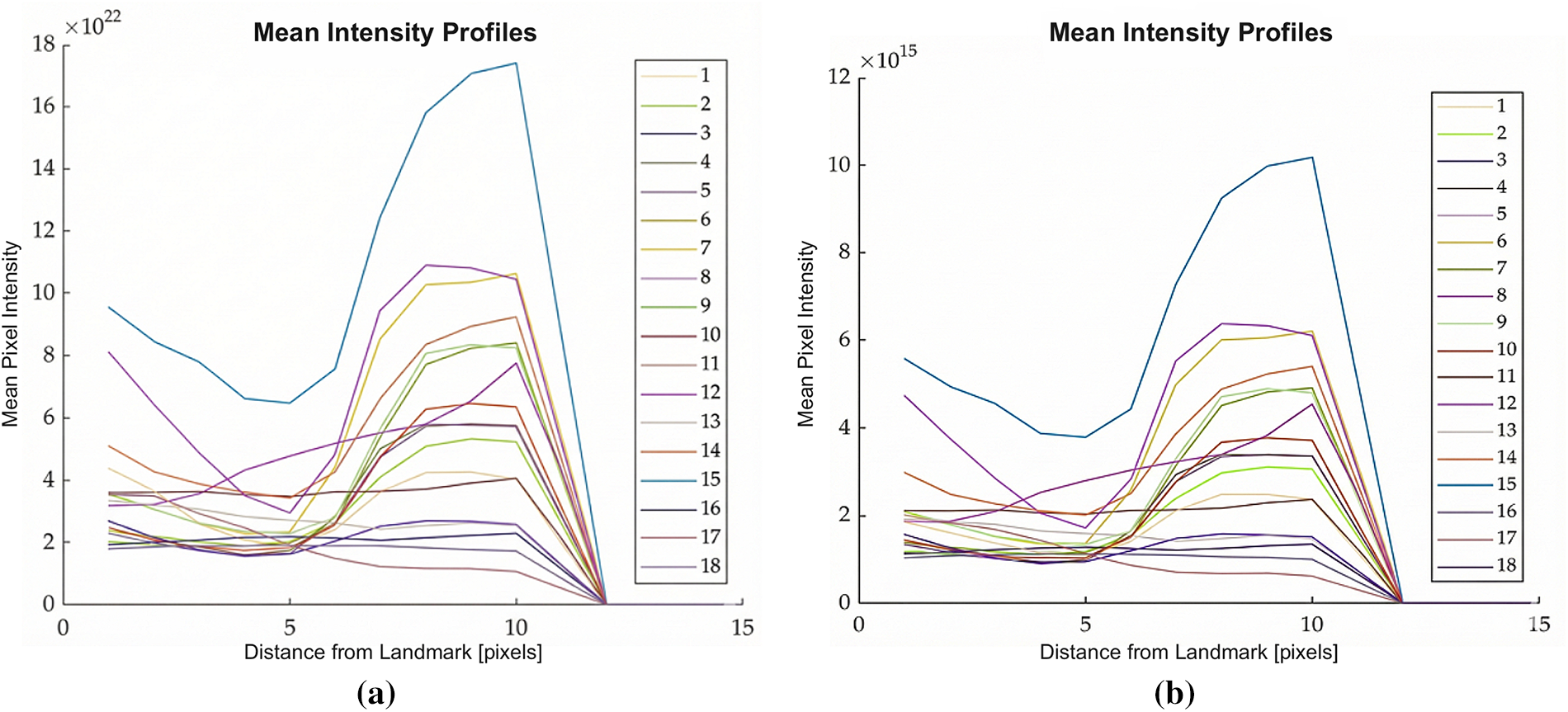
Figure 8: Plots showing average intensity values perpendicular to the contour points across all the training images used to build your ASM model, scaling factor: 0.5, (a) Division; (b) MinMax.
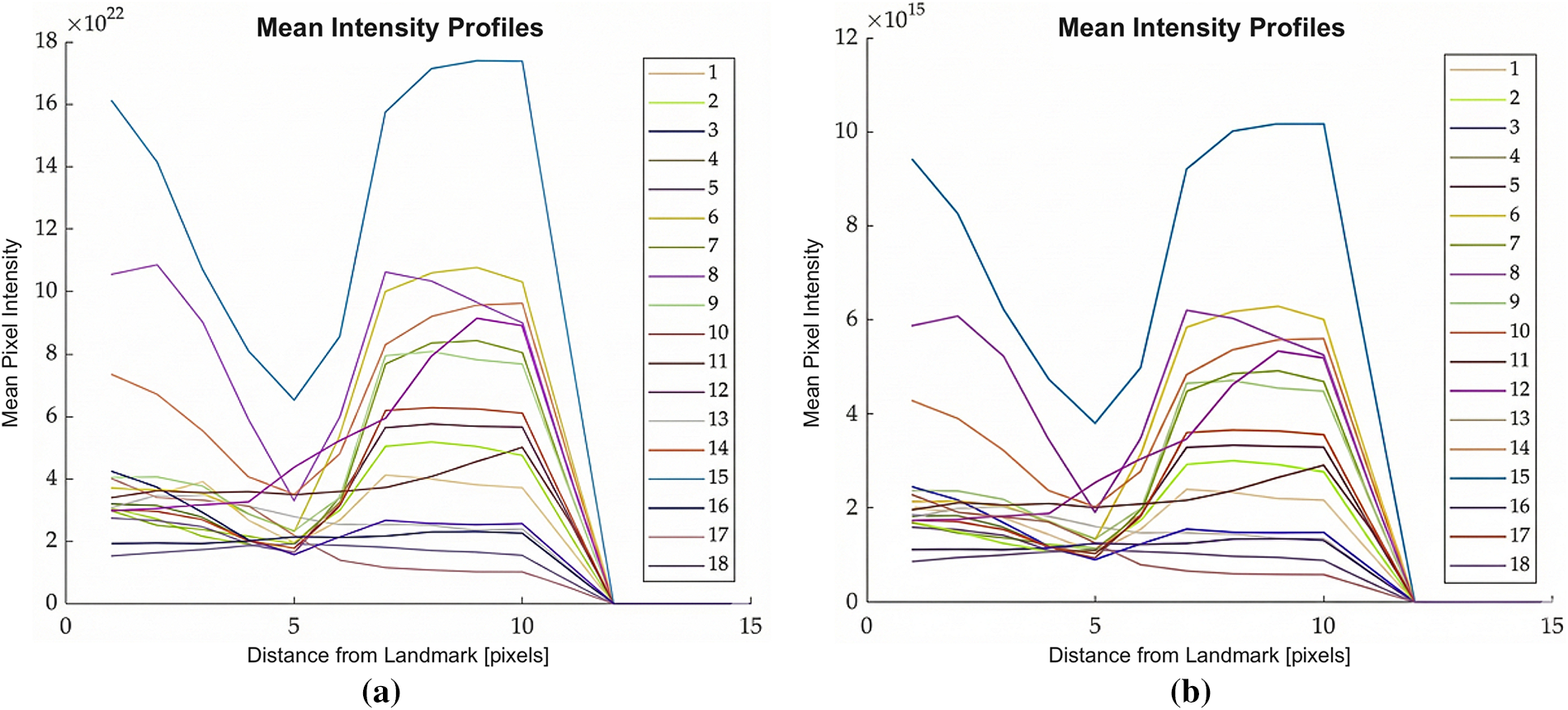
Figure 9: Plots showing average intensity values perpendicular to the contour points across all the training images used to build your ASM model, scaling factor: 0.25, (a) Division; (b) MinMax.
Comparative analysis revealed a substantial disparity in magnitude: intensity values produced by the MinMax method were seven orders of magnitude lower than those generated by the Division method. From an algorithmic perspective, this magnitude reduction is critical for the stability of the statistical appearance model. Lower-magnitude gradients are less sensitive to the high-frequency noise and localized shading artifacts prevalent in pathological CCTA scans, thereby enabling the landmark update process to maintain a smoother, more anatomically consistent trajectory during convergence.
To further investigate model sensitivity to these preprocessing parameters, an internal consistency analysis was performed. Fig. 10 depicts the Dice coefficient overlap between final fits produced specifically by the MinMax vs. the Division techniques. Results indicate that the right ventricle (RV) is significantly more resilient to spatial variability induced by rescaling, maintaining an average overlap of 0.958. This suggests that the global boundary of the RV remains stable regardless of the normalization technique employed. Conversely, the left ventricle (LV) exhibited a lower similarity coefficient of 0.746, indicating that its intricate myocardial and atrial anatomy is more sensitive to intensity normalization, necessitating precise parameter selection to ensure reproducibility.
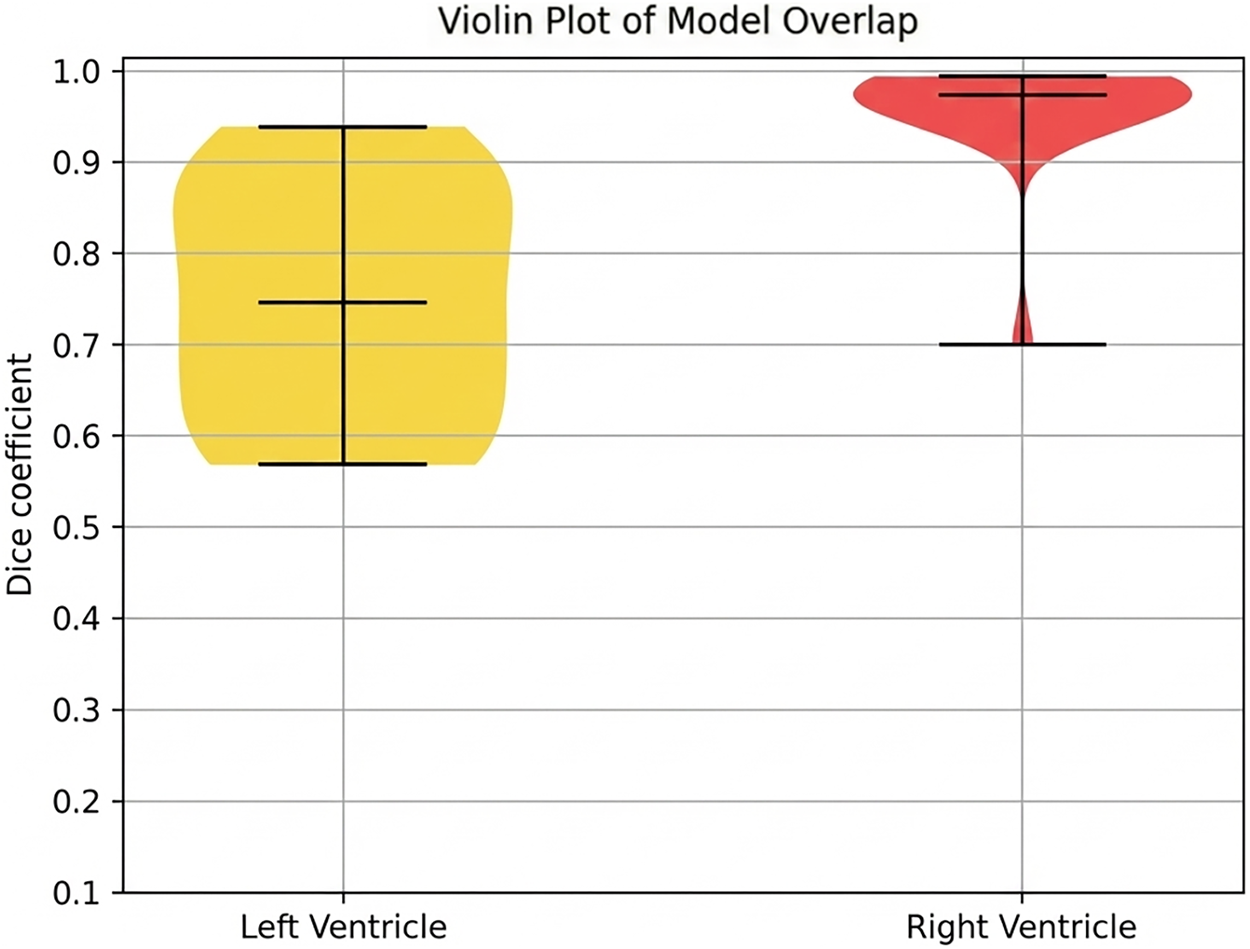
Figure 10: The overlap between final fits extracted from two different image rescaling techniques.
4.2 Comparative Performance and Validation
Following internal consistency verification, the ASM framework was validated against expert-delineated manual segmentations. The quantitative performance of the high-resolution models (4917 vertices) is detailed in Tables 1 and 2 and visualized in Figs. 11 and 12. The PCA MinMax approach demonstrated superior efficacy for LV segmentation, yielding the highest average Dice score of 0.6326 and the lowest average Hausdorff distance of 60.07 pixels.
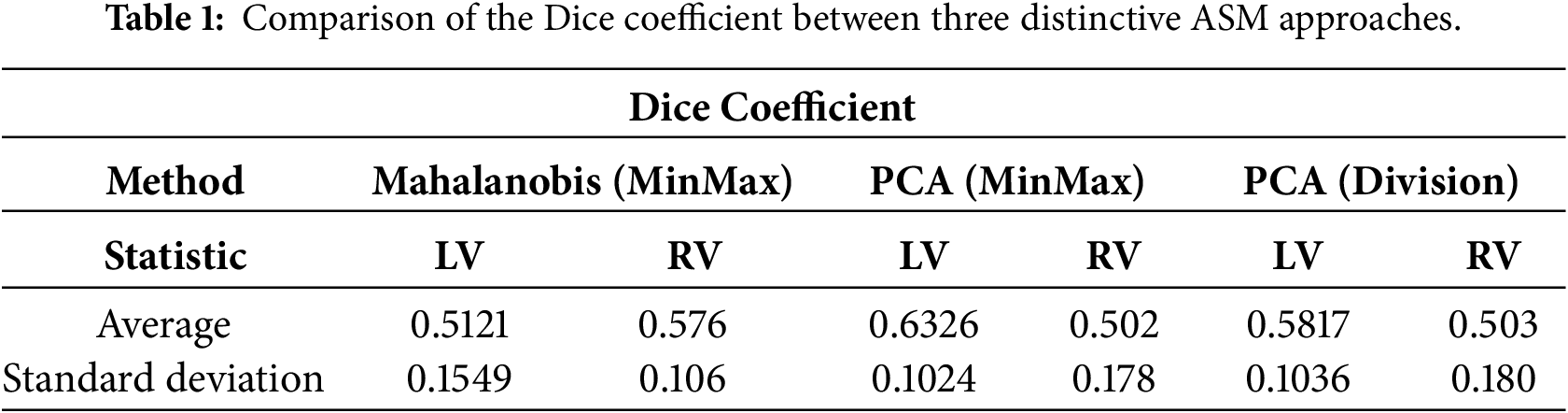
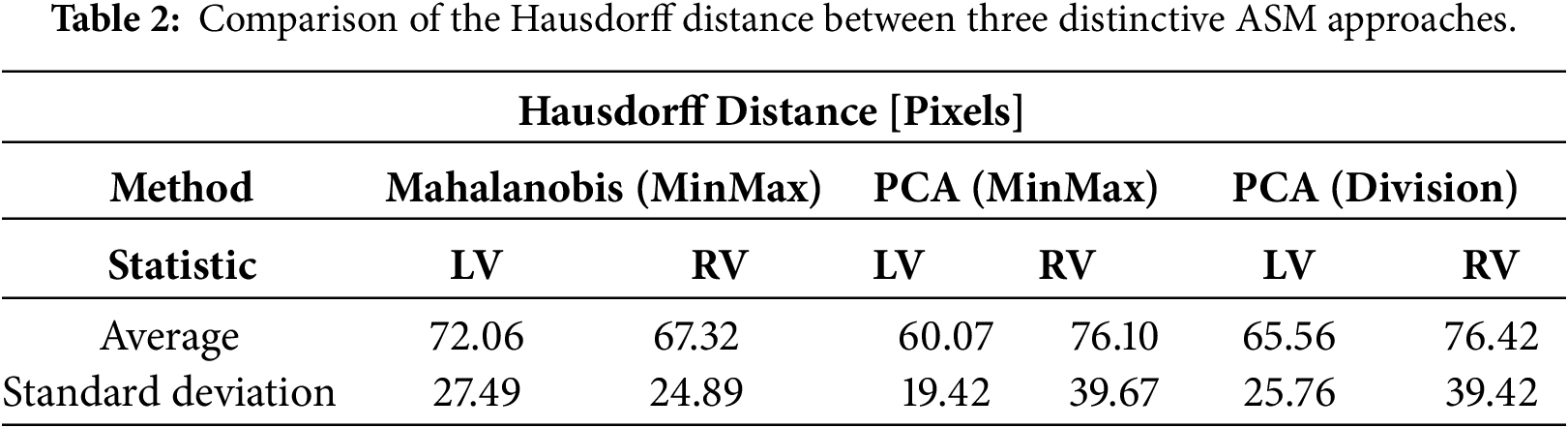
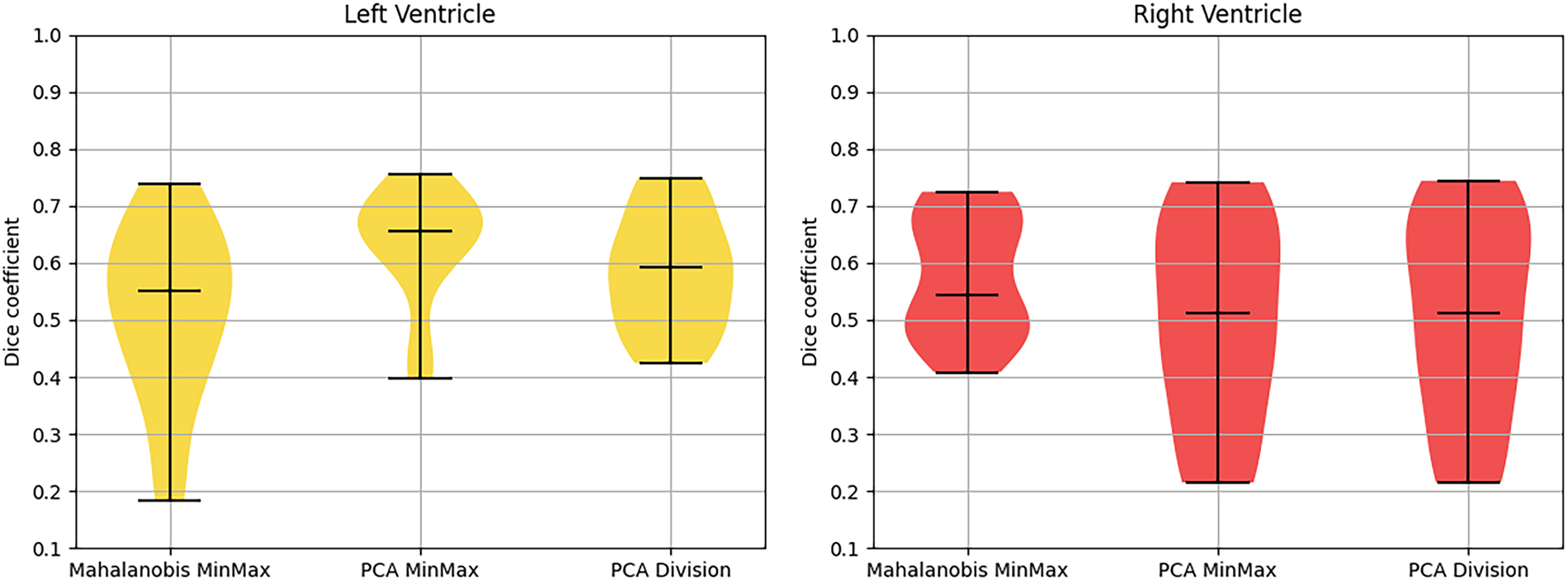
Figure 11: Violin plot of dice coefficient for different techniques.
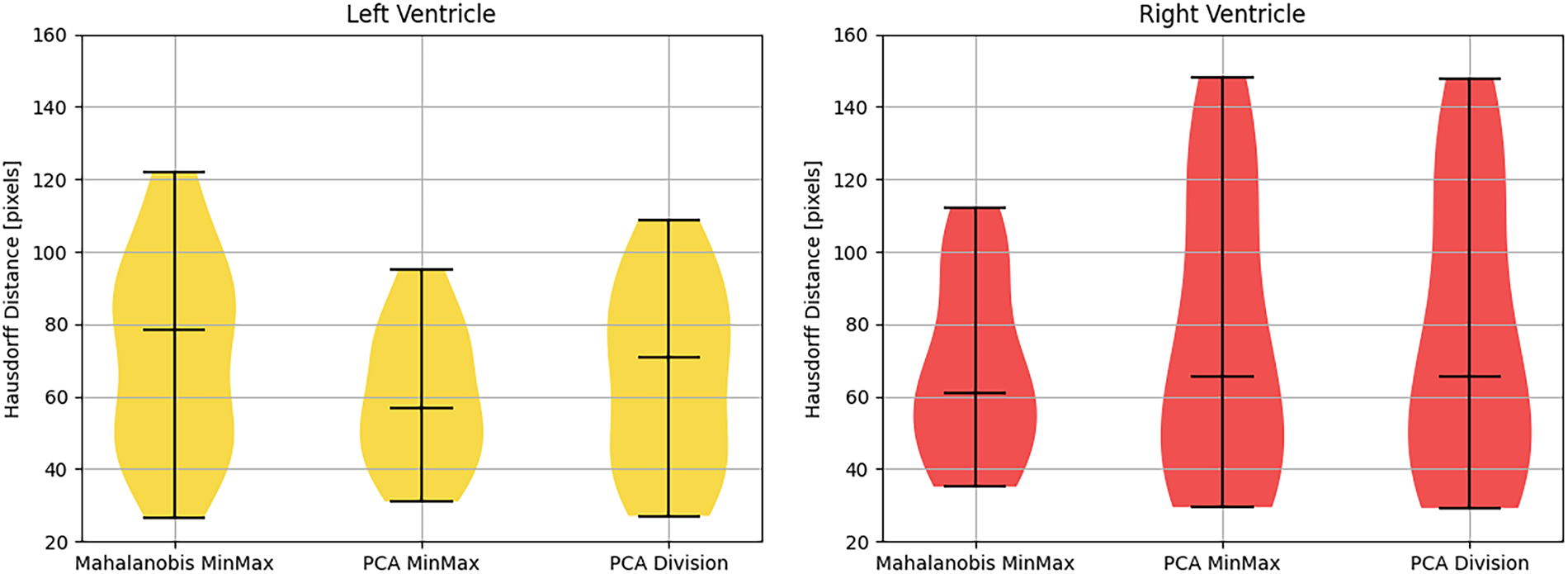
Figure 12: Violin plot of hausdorff distance for different techniques.
A notable discrepancy was observed in the right ventricle: while DSC scores were quantitatively lower (ranging between 0.502 and 0.576), visual inspection (Figs. 13 and 14) revealed high-quality spatial alignment with the ground truth. This discrepancy is fundamentally attributed to the RV’s unique crescentic geometry and its significantly thinner myocardial walls compared to the distinct, bullet-shaped LV. From a mathematical perspective, in thin-walled structures, spatial boundary shifts of only a few pixels lead to a disproportionate reduction in volumetric overlap (X ∩ Y). This phenomenon imposes a severe penalty on the DSC metric, even when the segmented morphology is anatomically faithful. In this context, the Hausdorff distance serves as a more reliable indicator of local boundary precision. The HD metric confirmed that the model’s borders remained closely aligned with physiological edges, despite the attenuation of volumetric scores.
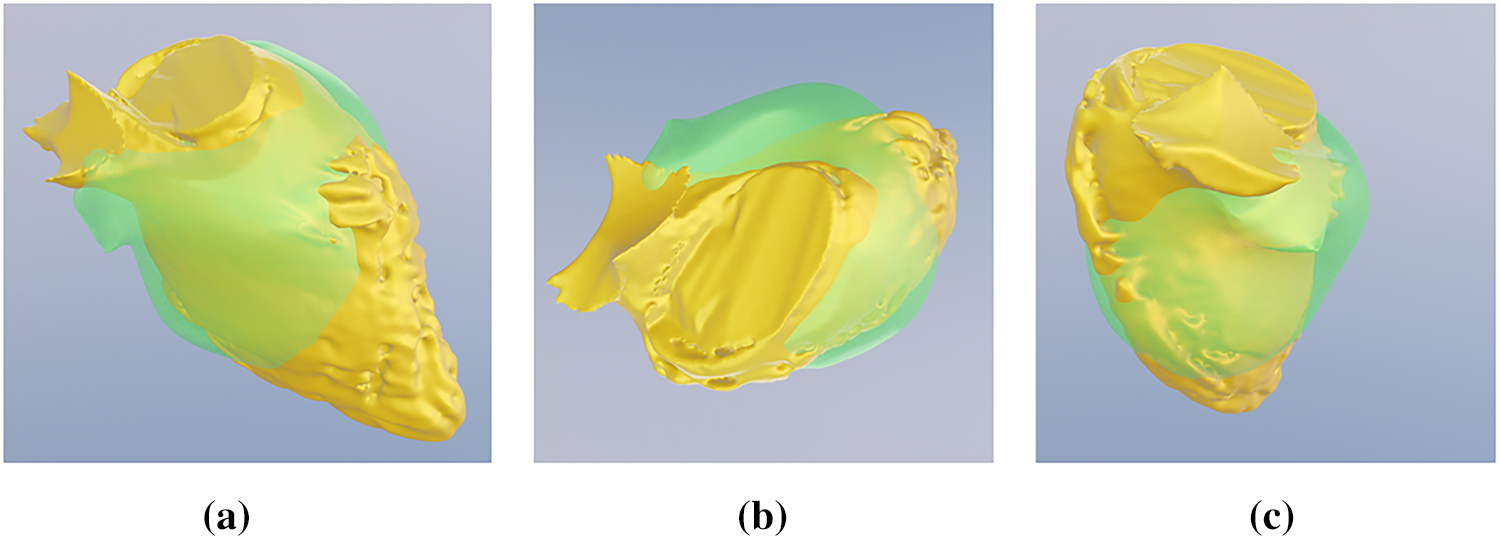
Figure 13: Visual overlap comparison of the best generated LV fit (Set Number: 10). The figure shows the ASM segmentation (green surface) overlapping the manual expert ground truth (yellow surface) from (a) side, (b) top, and (c) front perspectives.
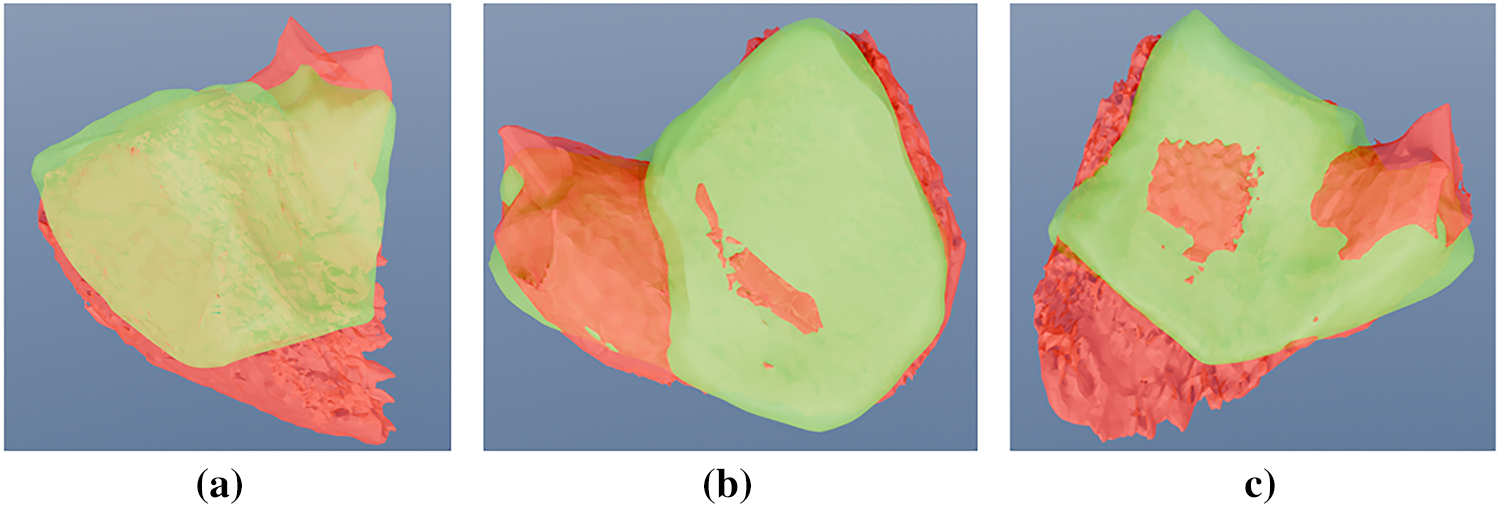

Figure 14: Visual overlap comparison of the best generated RV fit (Set Number: 7). The figure shows the ASM segmentation (green surface) overlapping the manual expert ground truth (red surface) from (a) side, (b) top, and (c) front perspectives.
From a clinical standpoint, these findings underscore the resilience of the ASM framework when processing pathological outliers. In diseased cardiac tissue, where intensity gradients are frequently attenuated or obscured by imaging artifacts, the capacity to preserve a continuous, anatomically plausible volume is often more diagnostically valuable than achieving maximal voxel-level congruency. While unconstrained deep learning architectures may yield superior Dice coefficients in healthy cohorts, they frequently produce topological anomalies, including fragmented regions (islands) or irregular boundaries, when applied to pathological data. By enforcing global anatomical priors, the ASM ensures structural continuity, providing the consistent orientation and geometric integrity indispensable for pre-procedural planning in complex clinical scenarios.
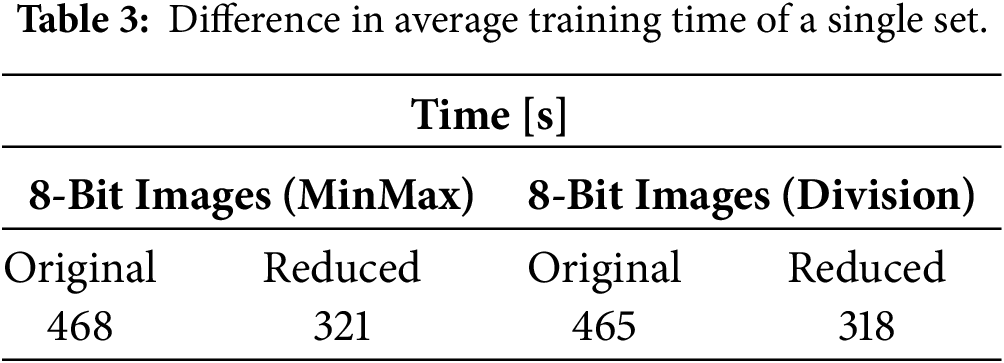
To investigate the trade-off between spatial resolution and algorithmic efficiency, a secondary evaluation was conducted using a simplified ventricular template. The vertex count was reduced by 50%, from the original 4917 vertices to a mesh of 2459 vertices. As summarized in Table 3, this reduction yielded a marked improvement in computational throughput, with average training times decreasing by approximately 31% (from 7 min 45 s to 5 min 18 s).
The impact on segmentation accuracy, detailed in Tables 4 and 5 and visualized in Figs. 15 and 16, revealed a counterintuitive finding. For the right ventricle, the PCA Division method demonstrated a significant performance enhancement, with the mean DSC increasing by 0.11 following vertex reduction. This suggests that for pathological hearts, high-resolution meshes may induce overfitting to local surface noise or imaging artifacts. By reducing vertex density, the statistical model effectively functions as a low-pass filter. This implicit smoothing forces the ASM to prioritize global anatomical shape over noisy local gradients, resulting in a more stable fit for the complex RV geometry.
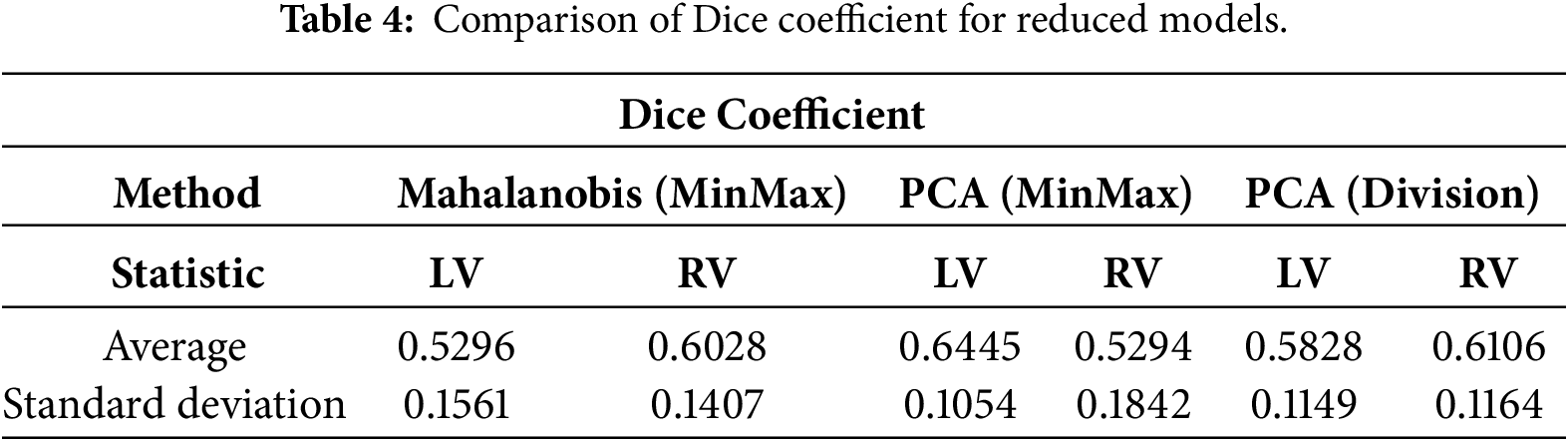
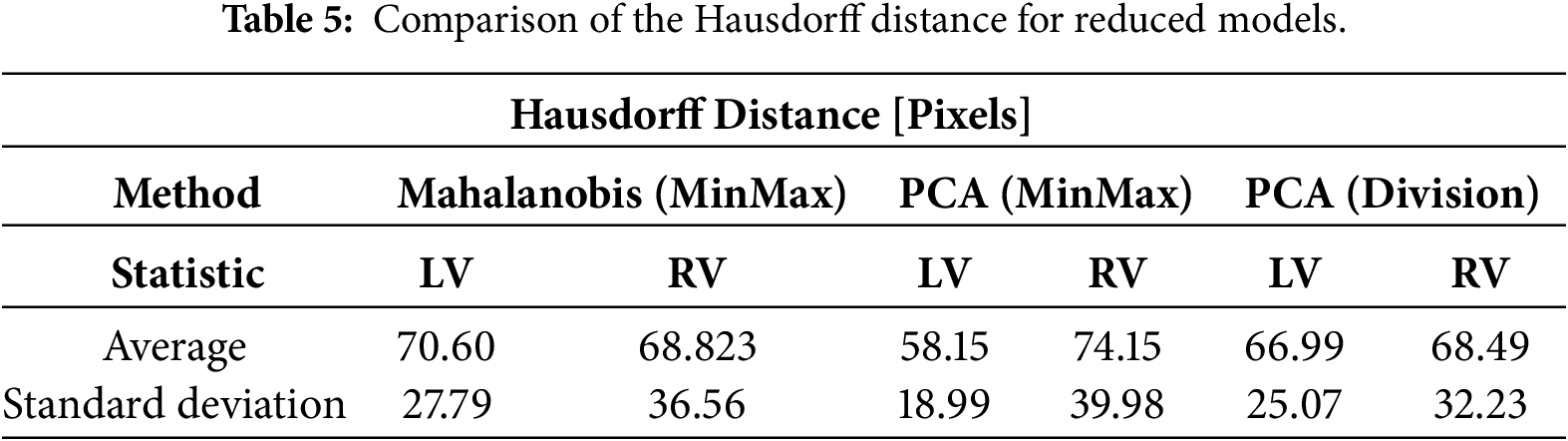
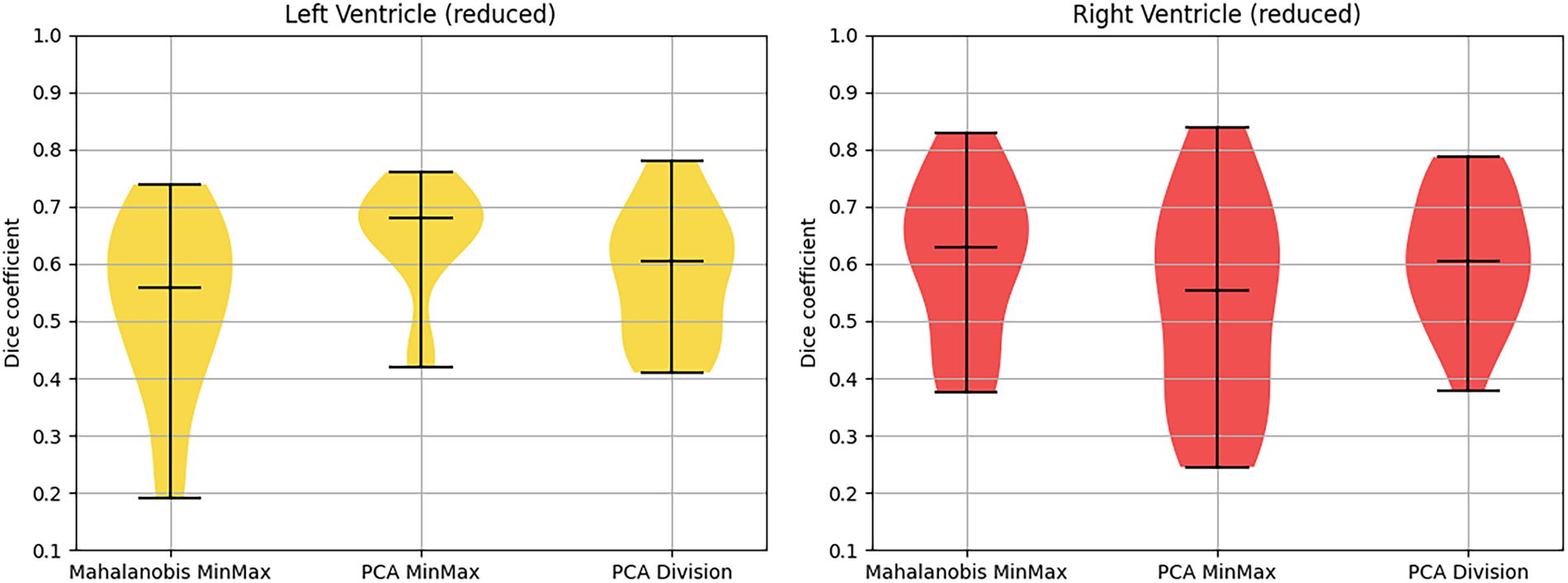
Figure 15: Violin plot serving as visual representation of Dice coefficient for reduced models.
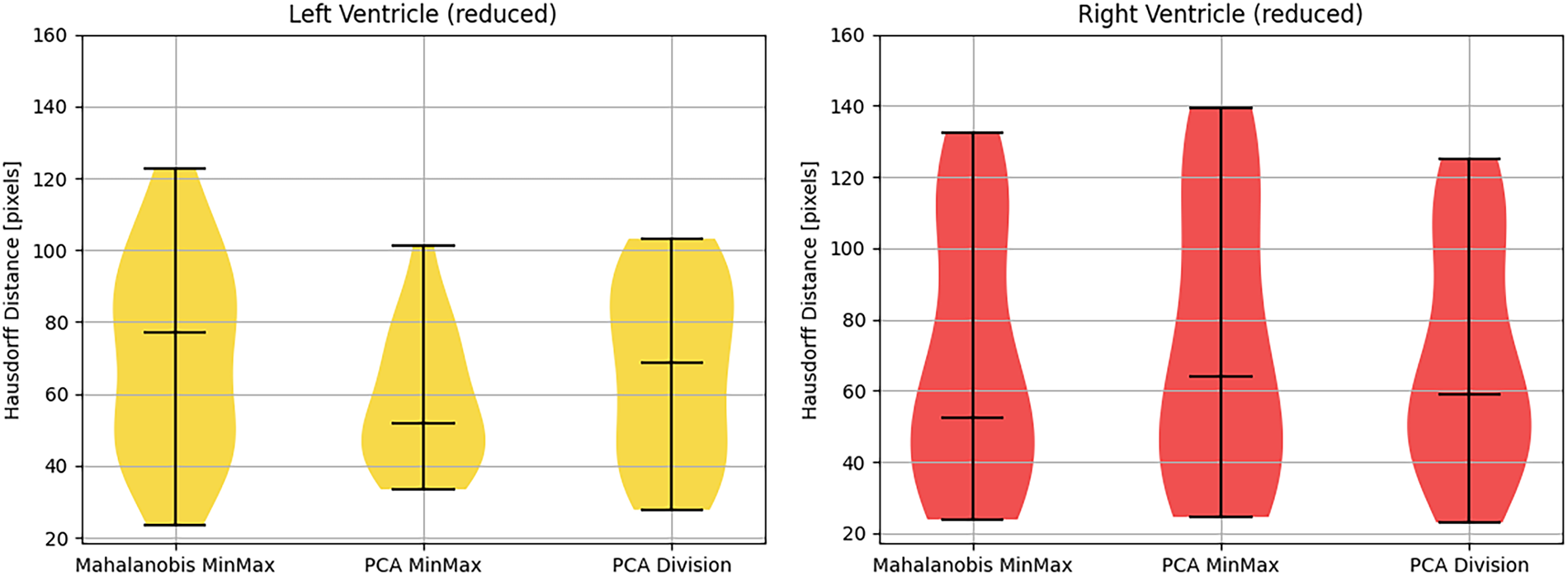
Figure 16: Violin plot serving as visual representation of the Hausdorff distance for reduced models.
Notwithstanding the improvements in the RV’s global volumetric overlap (DSC), the Hausdorff distance metrics exhibited negligible variance, indicating that the reduction in maximum boundary error was statistically insignificant. This dichotomy illuminates a fundamental trade-off: attenuated vertex counts facilitate robust global alignment for pathological orientations, whereas elevated vertex densities remain indispensable for resolving fine-grained anatomical substructures, such as the complex insertion points of the cardiac valves. Consequently, clinical deployment requires that model resolution be stratified according to the diagnostic objective. For rapid volumetric quantification and orientation of pathological ventricles, lower-resolution meshes offer superior stability and computational efficiency. Conversely, for applications demanding high-fidelity geometric precision, such as pre-surgical planning or myocardial wall thickness quantification, the high-resolution configuration with 4917 vertices constitutes the optimal standard, contingent upon sufficient image quality to support this level of geometric detail.
The results presented in this study demonstrate that the Active Shape Model (ASM) framework provides a resilient mechanism for segmenting pathological ventricular geometries, particularly in scenarios where data scarcity precludes the use of large-scale deep learning architectures. By constraining segmentation to a statistically learned shape space, the proposed method bridges the gap between the labour-intensive accuracy of manual delineation and the computational efficiency of automated systems.
5.1 Topological Reliability in Pathological Contexts
A primary advantage of the ASM approach observed in this study is the preservation of topological consistency. Unconstrained deep learning methods, such as U-Net or V-Net, operate on a voxel-wise classification basis. While efficient, these methods frequently produce topological artifacts—such as “islands” of false positives or “holes” within the myocardium—when trained on small or highly variable cohorts. In contrast, the ASM deformation is governed by global shape parameters (eigenvectors). This ensures that every segmented output remains a closed, continuous volume that creates a valid anatomical shape, regardless of image noise or artifacts. This intrinsic regularization is particularly vital for the “edge case” pathologies examined here, where standard intensity-based gradients are often ambiguous.
5.2 Statistical Adequacy of the Sample Size
A common challenge in modelling rare cardiac anomalies is the limited availability of high-resolution, validated datasets. While deep learning architectures require thousands of examples to approximate general features without overfitting, Statistical Shape Models operate as “low-shot” learners. The dimensionality of the search space in ASM is explicitly defined by the number of training shapes (n = 19). Widely cited literature in statistical shape modelling indicates that the primary modes of anatomical variation, which typically account for over 90% of the shape variance, converge rapidly, often requiring cohorts as small as n = 15−20 when dense point correspondence is strictly enforced [6]. Unlike a neural network estimating millions of weights, our PCA-based model estimates only n−1 modes. Consequently, the current cohort is statistically sufficient to construct the “mean shape” and the primary deformation fields required for a proof-of-concept in pathological segmentation.
5.3 Comparative Performance and Clinical Validity
To assess the clinical utility of the segmentation, quantitative metrics must be benchmarked against human performance rather than arbitrary numerical ideals. In the segmentation of pathological ventricles, inter-expert variability is significant; literature suggests that the Dice Similarity Coefficient (DSC) between two radiologists often falls between 0.75 and 0.85 due to the difficulty of defining boundaries in deformed tissues. Therefore, the mean DSC values achieved in this study (ranging from 0.50 to 0.84) should be interpreted in this context. Scores exceeding 0.70 indicate performance within the margin of human error for complex pathologies. Furthermore, for surgical planning applications, the Hausdorff Distance (HD) is often a more critical metric than volumetric overlap, as it penalizes maximum outliers. The ability of the ASM to maintain HD values generally below 20 mm implies that while the model may miss subtle volumetric nuances, it successfully avoids gross geometric errors that would endanger surgical intervention.
5.4 Limitations and Operational Constraints
Despite the demonstrated robustness, the current framework is subject to specific limitations. The reliance on manual landmarking for the initial training set ensures high-quality correspondence but introduces a significant bottleneck in scaling the system to larger populations. Additionally, the initialization of the mean shape requires a reasonable starting position; while the current search range is adequate for standard imaging protocols, significant translational misalignment could lead to local minima convergence. Future iterations of this work will likely require a hybrid approach, utilizing a lightweight neural network for pose initialization followed by ASM refinement to combine the best of both methodological paradigms.
This study successfully developed and evaluated a versatile Active Shape Model framework specifically tailored for ventricular segmentation in patients with heterogeneous cardiac pathologies. By utilizing a training dataset derived directly from clinical practice (n = 19), this research demonstrates that statistical shape priors can effectively navigate the complex morphological deformations and imaging artifacts that frequently characterize diseased hearts. The methodology presented herein offers a robust, interpretable alternative to data-intensive deep learning architectures, particularly in scenarios where high-fidelity pathological data is scarce.
Experimental results provided several key insights into the optimization of model-based segmentation for abnormal cardiac structures. Regarding preprocessing, the rescaling of 16-bit CCTA scans to 8-bit using the MinMax technique proved superior to the standard Division method. The significant reduction in intensity magnitude provided the stable gradients necessary for reliable ASM convergence, thereby reducing computational overhead without compromising accuracy. Quantitatively, the PCA MinMax configuration emerged as the most robust approach for left ventricle segmentation.
A significant contribution of this work is the technical clarification of the performance disparity between ventricular chambers. Despite lower volumetric scores in the thin-walled right ventricle, visual overlap analysis confirmed high anatomical fidelity. This underscores the primary advantage of the ASM: its ability to maintain a single, closed, and physiologically plausible volume even when local image gradients are weak or inconsistent. Furthermore, the investigation into vertex density revealed that reducing mesh resolution by 50% effectively acted as a statistical low-pass filter, improving RV segmentation stability by mitigating susceptibility to surface noise.
6.3 Summary of Contributions Limitations and Future Directions
While this patient cohort provided sufficient variability to establish a robust proof-of-concept, the results are presented as a pilot validation of the framework. The manual intervention required to establish strict point correspondence remains a technical necessity for ensuring ground-truth integrity in abnormal hearts, though it currently presents a bottleneck for large-scale clinical deployment. Future research will focus on expanding the training dataset to include a broader spectrum of congenital heart diseases and exploring hybrid architectures that combine the feature-extraction capabilities of convolutional neural networks with the anatomical safeguards of the ASM. Automating the point-correspondence workflow through non-rigid registration algorithms remains a priority for transitioning this framework toward real-time clinical applications.
Acknowledgement: Not applicable.
Funding Statement: This research utilized data collected under the Regional Operational Programme for the Małopolska Region 2014–2020 (Grant No. RPMP.01.02.01-12-0027/19). The data were used with the formal approval of the data administrator.
Author Contributions: Conceptualization, Oskar Kapuśniak, Adam Piórkowski, and Karolina Nurzyńska; methodology, Oskar Kapuśniak and Karolina Nurzyńska; software, Oskar Kapuśniak; validation, Oskar Kapuśniak; formal analysis, Oskar Kapuśniak and Karolina Nurzyńska; investigation, Oskar Kapuśniak; resources, Adam Piórkowski; data curation, Oskar Kapuśniak, and Julia Lasek; writing—original draft preparation, Oskar Kapuśniak; writing—review and editing, Oskar Kapuśniak, Julia Lasek, and Karolina Nurzyńska; visualization, Oskar Kapuśniak and Julia Lasek; supervision, Adam Piórkowski and Karolina Nurzyńska; project administration, Oskar Kapuśniak; funding acquisition, Adam Piórkowski. All authors reviewed and approved the final version of the manuscript.
Availability of Data and Materials: The data are not publicly available due to privacy. Data are available from the corresponding author upon reasonable request.
Ethics Approval: Ethics approval was not required for this study as it involved the analysis of a pre-existing, fully anonymized dataset provided by a third party (MST). All data were de-identified by the provider prior to being accessed by the authors, ensuring that no individual participants can be identified.
Conflicts of Interest: The authors declare no conflict of interests.
References
1. Zhou SK, Greenspan H, Davatzikos C, Duncan JS, Van Ginneken B, Madabhushi A, et al. A review of deep learning in medical imaging: imaging traits, technology trends, case studies with progress highlights, and future promises. Proc IEEE. 2021;109:820–38. doi:10.1109/JPROC.2021.3054390. [Google Scholar] [PubMed] [CrossRef]
2. Isola P, Zhu JY, Zhou T, Efros AA. Image-to-image translation with conditional adversarial networks. In: Proceedings of the 2017 IEEE Conference on Computer Vision and Pattern Recognition (CVPR); 2017 Jul 21–26; Honolulu, HI, USA. p. 1125–34. doi:10.1109/CVPR.2017.632. [Google Scholar] [CrossRef]
3. Alnasser T, Abdulaal L, Maiter A, Sharkey M, Dwivedi K, Salehi M, et al. Advancements in cardiac structures segmentation: a comprehensive systematic review of deep learning in CT imaging. Front Cardiovasc Med. 2024;11:1323461. doi:10.3389/fcvm.2024.1323461. [Google Scholar] [PubMed] [CrossRef]
4. Wang YR, Yang K, Wen Y, Wang P, Hu Y, Lai Y, et al. Screening and diagnosis of cardiovascular disease using artificial intelligence-enabled cardiac magnetic resonance imaging. Nat Med. 2024;30:1471–80. doi:10.1038/s41591-024-02971-2. [Google Scholar] [PubMed] [CrossRef]
5. Chanda PB, Sarkar SK. Cardiac MR images segmentation for identification of cardiac diseases using fuzzy based approach. In: Proceedings of the 3rd International Conference on Smart Systems and Inventive Technology (ICSSIT 2020); 2020 Aug 20–22; Tirunelveli, India. p. 1238–46. doi:10.1109/ICSSIT48917.2020.9214080. [Google Scholar] [CrossRef]
6. Cootes TF, Taylor CJ, Cooper DH, Graham J. Active shape models-their training and application. Comput Vis Image Underst. 1995;61(1):38–59. doi:10.1006/cviu.1995.1004. [Google Scholar] [CrossRef]
7. McGrath H, Li P, Dorent R, Bradford R, Saeed S, Bisdas S, et al. Manual segmentation versus semi-automated segmentation for quantifying vestibular schwannoma volume on MRI. Int J Comput Assist Radiol Surg. 2020;15:1445–55. doi:10.1007/s11548-020-02222-y. [Google Scholar] [PubMed] [CrossRef]
8. Khan K, Ullah F, Syed I, Ali H. Accurately assessing congenital heart disease using artificial intelligence. PeerJ Comput Sci. 2024;10:e2535. doi:10.7717/peerj-cs.2535. [Google Scholar] [PubMed] [CrossRef]
9. Hu H, Pan N, Liu H, Tu Z. Automatic segmentation of left and right ventricles in cardiac MRI using 3D-ASM and deep learning. Signal Process Image Commun. 2021;96:116303. doi:10.1016/j.image.2021.116303. [Google Scholar] [CrossRef]
10. Chai WY, Lin G, Wang CJ, Chiang HJ, Ng SH, Kuo YS, et al. A deep learning-based fully automated cardiac MRI segmentation approach for tetralogy of Fallot patients. J Magn Reson Imaging. 2026;63(1):264–76. doi:10.1002/jmri.70113. [Google Scholar] [PubMed] [CrossRef]
11. Jafari R, Verma R, Aggarwal V, Gupta RK, Singh A. Deep learning-based segmentation of left ventricular myocardium on dynamic contrast-enhanced MRI: a comprehensive evaluation across temporal frames. Int J CARS. 2024;19:2055–62. doi:10.1007/s11548-024-03221-z. [Google Scholar] [PubMed] [CrossRef]
12. Chen C, Qin C, Qiu H, Tarroni G, Duan J, Bai W, et al. Deep learning for cardiac image segmentation: a review. Front Cardiovasc Med. 2020;7:508599. doi:10.3389/FCVM.2020.00025. [Google Scholar] [PubMed] [CrossRef]
13. Aghapanah H, Rasti R, Kermani S, Tabesh F, Yousefi Banaem H, Pour Aliakbar H, et al. CardSegNet: an adaptive hybrid CNN-vision transformer model for heart region segmentation in cardiac MRI. Comput Med Imaging Graph. 2024;115(1):102382. doi:10.1016/j.compmedimag.2024.102382. [Google Scholar] [PubMed] [CrossRef]
14. Park S, Chung M. Cardiac segmentation on CT Images through shape-aware contour attentions. Comput Biol Med. 2022;147:105782. doi:10.1016/J.COMPBIOMED.2022.105782. [Google Scholar] [PubMed] [CrossRef]
15. Kong F, Wilson N, Shadden S. A deep-learning approach for direct whole-heart mesh reconstruction. Med Image Anal. 2021;74(1):102222. doi:10.1016/J.MEDIA.2021.102222. [Google Scholar] [PubMed] [CrossRef]
16. Gul F, Shah M, Ali M, Hussain L, Sadiq T, Abbasi AA, et al. A hybrid multi-panel image segmentation framework for improved medical image retrieval system. PLoS One. 2025;20:e0315823. doi:10.1371/journal.pone.0315823. [Google Scholar] [PubMed] [CrossRef]
17. Kinger S, Kulkarni V. A review of explainable AI in medical imaging: implications and applications. Int J Comput Appl. 2024;46:983–97. doi:10.1080/1206212X.2024.2404082. [Google Scholar] [CrossRef]
18. Carbajal-Degante E, Avendaño S, Ledesma L, Olveres J, Vallejo E, Escalante-Ramirez B. A multiphase texture-based model of active contours assisted by a convolutional neural network for automatic CT and MRI heart ventricle segmentation. Comput Methods Programs Biomed. 2021;211:106373. doi:10.1016/J.CMPB.2021.106373. [Google Scholar] [PubMed] [CrossRef]
19. Finnegan RN, Chin V, Chlap P, Haidar A, Otton J, Dowling J, et al. Open-source, fully-automated hybrid cardiac substructure segmentation: development and optimisation. Phys Eng Sci Med. 2023;46:377–93. doi:10.1007/S13246-023-01231-W. [Google Scholar] [PubMed] [CrossRef]
20. Freidoonimehr N, Arjomandi M, Sedaghatizadeh N, Chin R, Zander A. Transitional turbulent flow in a stenosed coronary artery with a physiological pulsatile flow. Int J Numer Method Biomed Eng. 2020;36:e3347. doi:10.1002/CNM.3347. [Google Scholar] [PubMed] [CrossRef]
Cite This Article
 Copyright © 2026 The Author(s). Published by Tech Science Press.
Copyright © 2026 The Author(s). Published by Tech Science Press.This work is licensed under a Creative Commons Attribution 4.0 International License , which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.


 Submit a Paper
Submit a Paper Propose a Special lssue
Propose a Special lssue View Full Text
View Full Text Download PDF
Download PDF Downloads
Downloads
 Citation Tools
Citation Tools