 Open Access
Open Access
ARTICLE
Effective Data Balancing and Fine-Tuning Techniques for Medical sLLMs in Resource-Constrained Domains
Department of Software Engineering & Division of Electronics and Information Engineering, Jeonbuk National University, Jeonju-si, Republic of Korea
* Corresponding Author: Jaehyuk Cho. Email:
(This article belongs to the Special Issue: Bridging the Gap: AutoML and Explainable AI for Industrial and Healthcare Innovations)
Computers, Materials & Continua 2026, 87(3), 20 https://doi.org/10.32604/cmc.2026.077579
Received 12 December 2025; Accepted 16 January 2026; Issue published 09 April 2026
Abstract
Despite remarkable advances in medical large language models (LLMs), their deployment in real clinical settings remains impractical due to prohibitive computational requirements and privacy regulations that restrict cloud-based solutions. Small LLMs (sLLMs) offer a promising alternative for on-premise deployment, yet they require domain-specific fine-tuning that still exceeds the hardware capacity of most healthcare institutions. Furthermore, the impact of multilingual data composition on medical sLLM performance remains poorly understood. We present a resource-efficient fine-tuning pipeline that integrates Quantized Low-Rank Adaptation (QLoRA), Fully Sharded Data Parallelism (FSDP), and Sequence Packing, validated across two model scales: MedGemma 4B for efficiency analysis and LLaMA 3.3 70B for data balance experiments. Our approach achieves 58.3% reduction in video random access memory (VRAM) usage (from 48 GB to 20 GB) and 5× training speedup on MedGemma 4B using NVIDIA L40s GPUs. Critically, experiments on LLaMA 3.3 70B reveal that English-heavy data mixing (10:3 ratio) degrades Korean medical law performance by 1.23 percentage points while providing only marginal English gains (+1.49 pp), demonstrating catastrophic forgetting in multilingual medical fine-tuning. Our work provides three contributions: (1) a practical fine-tuning pipeline operable within 20 GB VRAM, (2) empirical evidence that data balance—not volume—determines multilingual medical QA performance, and (3) actionable guidelines for deploying medical sLLMs in non-English clinical environments.Keywords
Large Language Models (LLMs) have rapidly emerged as powerful tools in the medical domain, with several studies demonstrating that models such as Med-PaLM and Med-PaLM 2 can achieve expert-level accuracy on high-stakes medical exams, including USMLE-style questions [1,2]. These results ignited strong optimism about the use of LLMs in clinical decision support, medical education, and triage systems. However, despite these promising benchmarks, there exists a substantial gap between benchmark performance and real-world medical deployment. Many clinics, especially small and mid-sized institutions, lack access to the large GPU clusters required to train or even run state-of-the-art LLMs [2]. Furthermore, transmitting sensitive patient information to external APIs raises severe concerns under HIPAA, GDPR, and local privacy regulations, making cloud-based solutions often infeasible for hospital applications [2].
To address these limitations, recent efforts have shifted toward developing small large language models (sLLMs)-compact architectures that retain strong reasoning capabilities while remaining lightweight enough for on-premise deployment. These models are particularly attractive because they can operate securely within hospital infrastructure, ensuring data residency and reducing inference cost. However, a persistent challenge remains: sLLMs are not competitive without domain adaptation. They require fine-tuning on medical knowledge, and even small models can become difficult to fine-tune on hardware-constrained environments. Prior studies have shown that full fine-tuning large models may require hundreds of gigabytes of GPU memory [3], which is far beyond the capacity of local clinical systems.
To overcome these constraints, parameter-efficient fine-tuning (PEFT) methods such as LoRA [4] and QLoRA [5] have been proposed. However, resource constraints create a secondary challenge that has received little attention: data selection under capacity limits. LoRA reduces the trainable parameter space by using low-rank adapters, while QLoRA introduces 4-bit quantized base models with NF4 and double quantization techniques, enabling high-quality fine-tuning even on a single GPU. In parallel, distributed training frameworks such as Fully Sharded Data Parallel (FSDP) [6] were developed to shard model states across multiple GPUs, significantly reducing memory overhead. Although each technique brings substantial benefits, their combined use for multilingual medical QA tasks is still underexplored, and integrating them into a practical, reproducible pipeline remains a technical challenge.
What is Known. Recent advances in parameter-efficient fine-tuning have demonstrated that techniques such as LoRA and QLoRA can dramatically reduce the computational requirements for adapting large language models. Separately, distributed training frameworks like FSDP [6] enable memory-efficient multi-GPU training, and Sequence Packing [7] improves throughput by eliminating padding waste. In the medical domain, benchmarks like MedQA [8] and KorMedMCQA [9] provide standardized evaluation for English and Korean clinical knowledge, respectively.
What is Missing. Despite these individual advances, three critical gaps remain. First, the combined effectiveness of QLoRA, FSDP, and Sequence Packing for medical QA has not been validated—existing studies test these techniques on general-domain tasks where domain-specific terminology and reasoning patterns differ substantially. Second, while catastrophic forgetting is documented in continual learning [10], its manifestation in multilingual medical fine-tuning remains empirically unexplored, particularly for resource-limited languages like Korean. Third, practical guidelines translating efficiency gains into clinical deployment strategies are absent from the literature.
What We Do, Why, and How. To address these gaps, we develop an integrated fine-tuning pipeline combining QLoRA, FSDP, and Sequence Packing (how), specifically targeting multilingual medical QA under hospital-grade hardware constraints. We validate our approach on both English (MedQA) and Korean (KorMedMCQA) benchmarks, systematically investigating data balance effects through controlled experiments (what). Our goal is to bridge the gap between cutting-edge medical AI research and practical clinical deployment feasibility. This leads to two research questions: (RQ1) Can an integrated pipeline of QLoRA, FSDP, and Sequence Packing enable medical sLLM fine-tuning within hospital-grade hardware constraints? (RQ2) How does the ratio of English to non-English medical data affect multilingual medical QA performance?
The contributions of our work are threefold, each addressing gaps unresolved in prior research:
(1) Integrated Efficiency Pipeline for Medical sLLMs. While QLoRA, FSDP, and Sequence Packing have been individually studied, their combined application to multilingual medical QA under strict memory constraints (≤20 GB VRAM) has not been demonstrated. We provide the first validated pipeline that integrates all three techniques, achieving 58.3% memory reduction and 5× training speedup compared to baseline fine-tuning. Unlike prior work that assumes access to high-end infrastructure [1,2], our approach enables fine-tuning on mid-range GPUs available in typical hospital environments.
(2) Empirical Analysis of Multilingual Data Balance in Medical AI. Previous studies on medical LLMs predominantly focus on English benchmarks [1,9] or single-language adaptation [4]. We present the first systematic investigation of how English-Korean data mixing ratios affect medical QA performance, revealing that excessive English data causes catastrophic forgetting of region-specific medical knowledge—a phenomenon critical for clinical deployment but previously undocumented in medical AI literature.
(3) Deployment-Ready Guidelines for Non-English Clinical Settings. Beyond technical contributions, we provide actionable recommendations for data curation based on deployment context (international education vs. local clinical use), addressing a practical gap between academic benchmarks and real-world medical AI implementation.
Overall, our study bridges the practical gap between cutting-edge medical LLM research and real-world feasibility by providing a lightweight, reproducible, and resource-efficient training strategy suitable for clinical environments.
2.1 From BERT to Generative AI
Medical AI has evolved from classification tasks [11] to generative approaches enabled by foundation models [12] and their medical adaptations. Research published in Nature and Nature Medicine demonstrated that generative LLMs can achieve expert-level performance on professional medical exams [1,2]. However, these models are proprietary systems that cannot be inspected or deployed locally [13,14]. This has pushed researchers toward open-source alternatives that balance performance with institutional control. Recent systematic reviews have examined the trade-offs between prompt engineering and fine-tuning strategies [15], while domain-specific studies explore their relative effectiveness in medical applications [6]. Early models like BERT and BioBERT excelled at disease coding and clinical named entity recognition, establishing foundations for NLP in healthcare [11]. Pre-trained language models continue to advance biomedical information extraction tasks [16]. In parallel, domain-specialized large language models have also demonstrated strong performance in other high-stakes fields, such as finance [17], underscoring the importance of domain adaptation. Multilingual medical LLMs have drawn attention to challenges in prompt robustness, corpus adaptation, and federated learning for underrepresented languages [9]. Recent surveys on medical visual question answering highlight similar challenges in multimodal clinical AI systems [18].
2.2 Making Fine-Tuning Affordable
Full fine-tuning requires substantial computational resources [3]. Parameter-Efficient Fine-Tuning (PEFT) methods address this limitation [19]. LoRA freezes the base model and trains only small adapter layers. QLoRA further reduces memory requirements through 4-bit quantization. This combination enables powerful model adaptations on hardware that would otherwise be insufficient for full fine-tuning [5]. Studies have demonstrated that QLoRA achieves competitive results on medical QA with a fraction of the VRAM and energy cost. These innovations have been combined with memory-efficient techniques like ZeRO and FSDP, democratizing LLM deployment for smaller labs and hospitals [6]. Domain-specific adaptations of foundation models have demonstrated that targeted fine-tuning on medical corpora can significantly enhance clinical reasoning capabilities [20]. The result is the emergence of small-to-medium sLLMs that support privacy-preserving, locally deployable clinical AI with manageable operational costs.
2.3 Breaking the Memory Barrier
Even with quantization, memory constraints remain challenging. Traditional training replicates the model on every GPU, which is inefficient [21]. Fully Sharded Data Parallelism (FSDP) addresses this by sharding model parameters across GPUs [6]. Advanced memory optimization strategies, including model parallelism [22], ZeRO-Infinity [23], and offloading techniques, enable extreme-scale training even on limited infrastructure [21]. The combination of FSDP and QLoRA enables fitting capable models onto modest hardware, a technique validated in large-scale environments [24]. These advances have direct implications for privacy-preserving hospital deployment, where on-premise training and rapid model updates are critical [25,26].
2.4 Clinical QA, Data Scarcity, and Safety
Scaling dataset volume alone does not guarantee improved performance in clinical QA. Imbalanced, English-heavy corpora often degrade accuracy for minority languages and nuanced biomedical subdomains. Large-scale multilingual medical QA datasets, such as Chinese medical corpora, have emerged to address this gap [4], though careful curation remains essential to prevent dataset bias. Research has documented catastrophic forgetting and domain interference when continual training lacks proper data stratification and curation [10]. Rigorous benchmarking, multilingual evaluation, and modular dataset integration are now emphasized as best practices [25,27]. Additionally, as LLMs begin to inform clinical decision-making, ethical and safety concerns have intensified. Hallucinated or erroneous answers can directly harm patients, underscoring the need for external validation, uncertainty quantification, and robust human oversight [2,14].
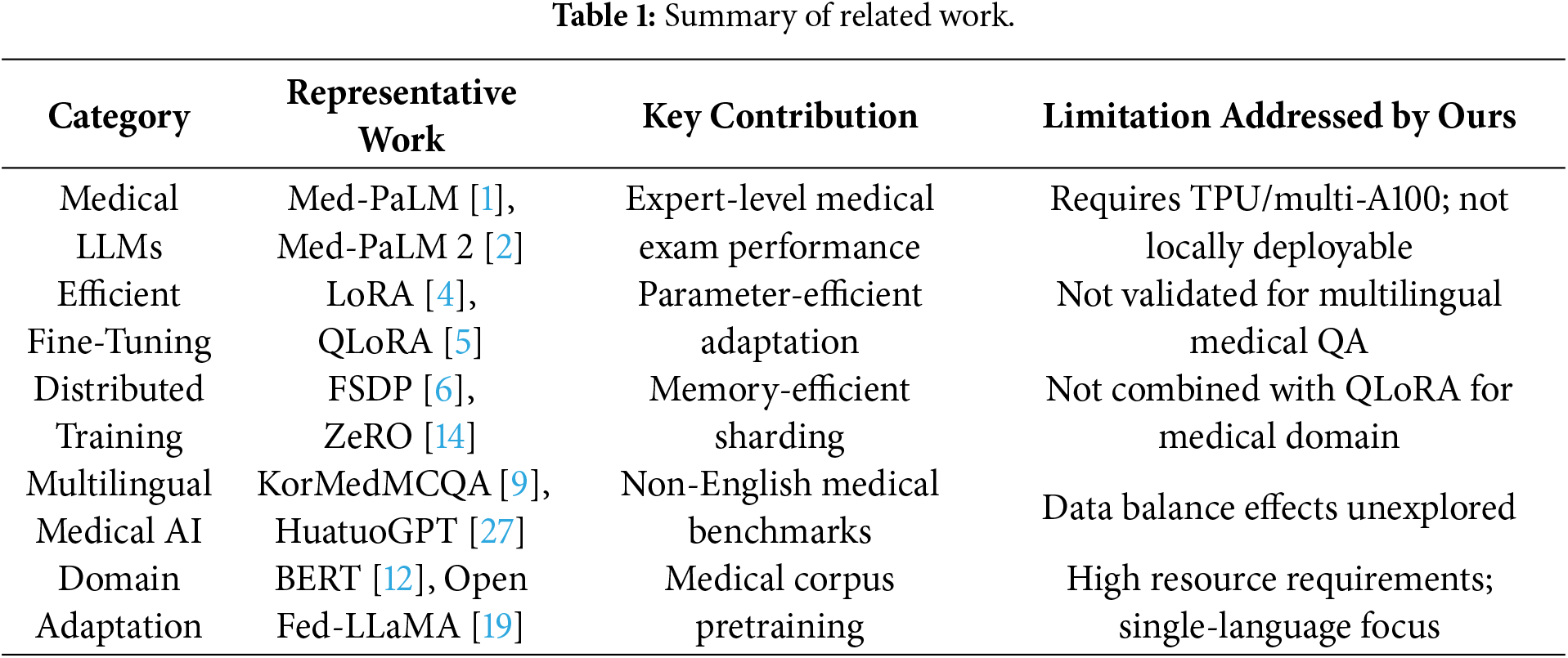
Table 1 summarizes the representative prior works across five categories relevant to our study, identifying their key contributions and the specific limitations that our integrated pipeline addresses.
3.1 Novelty of the Proposed Framework
Our framework differs from standard approaches in three key aspects. First, integrated optimization across the full training stack. While prior works apply QLoRA [5], FSDP [6], or Sequence Packing [7] in isolation, we demonstrate that their synergistic combination yields compound efficiency gains unachievable by any single technique. Specifically, QLoRA reduces trainable parameters to 0.21% of the base model, FSDP eliminates redundant memory replication across GPUs, and Sequence Packing minimizes wasted computation on padding tokens. Together, these reduce VRAM from 48 GB to 20 GB—a 58.3% reduction that crosses the critical threshold for hospital-grade GPU deployment.
Second, medical domain validation under realistic constraints. Existing efficient fine-tuning studies typically validate on general-domain benchmarks (e.g., GLUE, SuperGLUE) or assume access to high-end infrastructure [3]. We provide the first validation specifically for multilingual medical QA, demonstrating that efficiency techniques preserve clinical reasoning quality across both English (MedQA) and Korean (KorMedMCQA) benchmarks.
Third, data balance as a first-class design consideration. Standard fine-tuning pipelines treat data composition as a preprocessing detail. Our framework explicitly incorporates data balance analysis (Phase 3), revealing that multilingual medical performance depends critically on language mixing ratios—a finding with direct implications for clinical AI deployment that prior frameworks overlook.
We designed a resource-efficient fine-tuning pipeline tailored for multilingual medical QA tasks under strict GPU memory constraints. Fig. 1 summarizes our three-phase workflow: data optimization (Phase 1), parameter-efficient distributed training (Phase 2), and evaluation with data balance analysis (Phase 3).
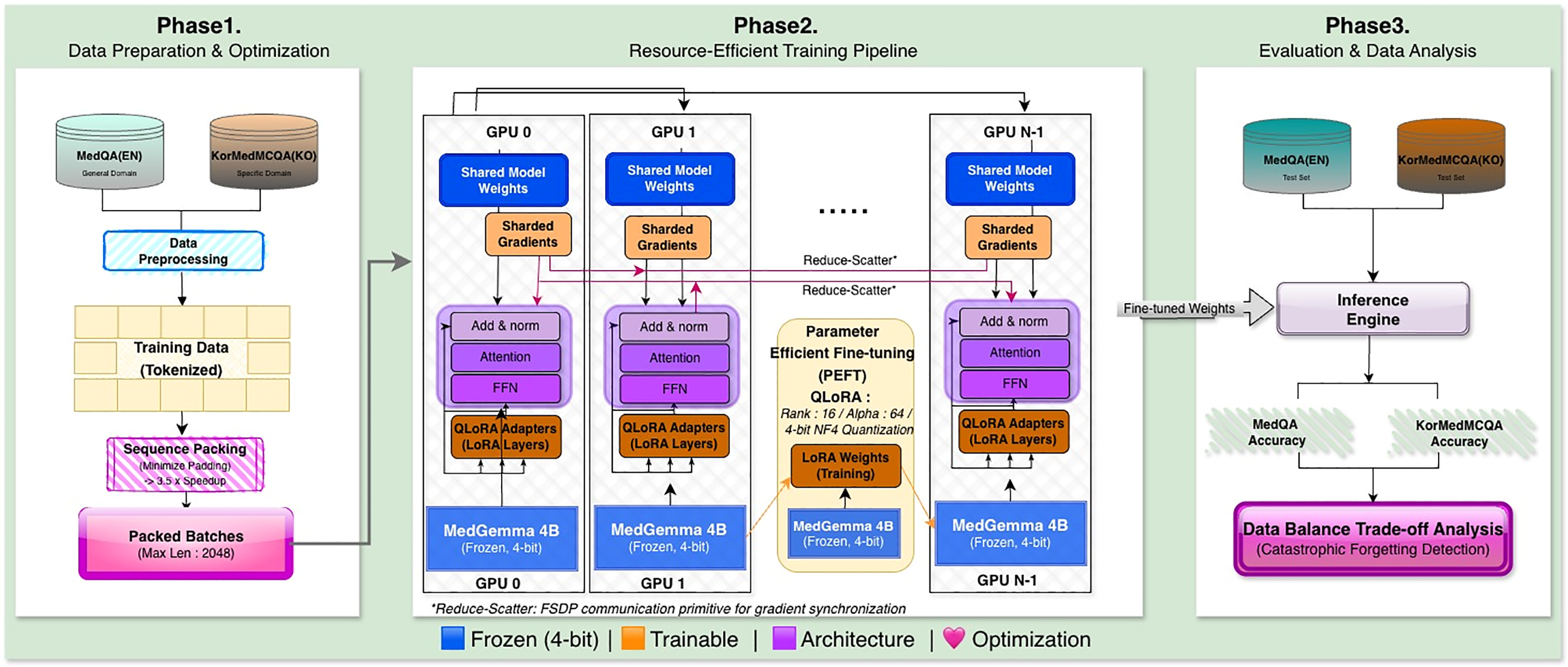
Figure 1: Overview of the proposed resource-efficient fine-tuning pipeline for multilingual medical sLLMs. Phase 1 (Data Preparation & Optimization) applies Sequence Packing to minimize padding overhead by concatenating variable-length samples until the context limit (2048 tokens) is reached. Phase 2 (Re-source-Efficient Training) integrates QLoRA with 4-bit NF4 (Normal Float 4-bit) quantization and FSDP (Fully Sharded Data Parallel) for distributed memory optimization, enabling training within 20 GB VRAM. Phase 3 (Evaluation & Data Analysis) assesses performance on MedQA (English) and KorMedMCQA (Korean) benchmarks, with systematic data balance experiments to identify optimal language mixing ratios. Abbreviations: FSDP, Fully Sharded Data Parallel; QLoRA, Quantized Low-Rank Adaptation; FFN, Feed-Forward Network; NF4, Normal Float 4-bit; VRAM, Video Random Access Memory.
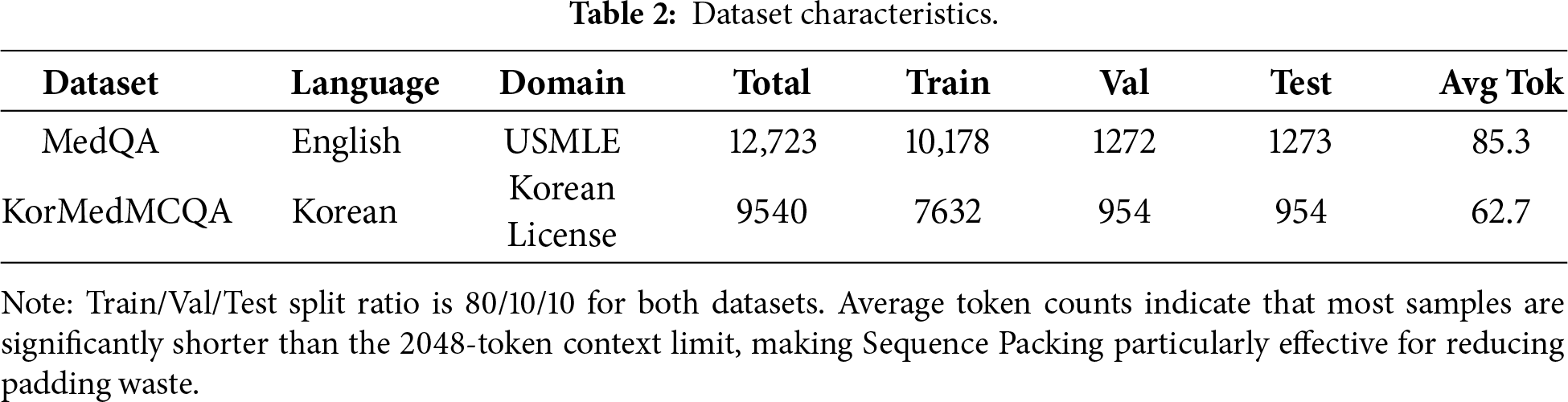
Table 2 presents the characteristics of the two datasets used in our experiments, including sample counts, train/validation/test splits, and average sequence lengths.
3.2.1 MedQA (Global Knowledge)
MedQA is one of the largest and most influential English medical QA datasets, originally constructed from the USMLE Step 1/2/3 exam questions and released as a benchmark for clinical knowledge reasoning [8]. The dataset contains multi-choice medical questions covering pathology, physiology, pharmacology, medical ethics, and clinical diagnosis, requiring both factual recall and multi-step reasoning.
Unlike many domain-specific datasets, MedQA reflects Western medical guidelines and practice standards, making it ideal for evaluating whether a model possesses globally accepted clinical knowledge. In our study, MedQA served two roles:
(1) General Medical Reasoning Evaluation
Since the dataset measures high-level diagnostic reasoning, it allowed us to test whether our sLLM can maintain medically valid reasoning after fine-tuning.
(2) High-Volume Training Source
We also used MedQA to examine the effects of “data flooding”. Specifically, we tested whether feeding the model a large volume of English medical questions would boost general reasoning or instead cause it to forget Korean-specific content—a phenomenon associated with catastrophic forgetting in multilingual settings.
Its large size and linguistic homogeneity (English-only) made it an ideal counterweight to the more specialized Korean dataset.
3.2.2 KorMedMCQA (Local Expertise)
KorMedMCQA is a Korean medical multiple-choice QA dataset derived from real items in the Korean National Licensing Examination for Medicine, Nursing, Dentistry, and Pharmacy [9]. Unlike MedQA, which focuses on broad global medical science, KorMedMCQA captures region-specific medical expertise in Korea.
It contains several types of questions that English-centric LLMs often struggle with:
(1) Clinical Practice Guidelines
The dataset includes Korean-specific clinical practices such as nursing protocols, emergency care standards, and region-adapted diagnostic workflows.
(2) Domain-Specific Subfields
Many questions come from pharmacy, dentistry, maternal care, and community health—areas where terminology differs significantly from English counterparts.
(3) Medical Law and Regulations
A unique feature of KorMedMCQA is its testing of Korean medical law, including healthcare personnel regulations, scope-of-practice rules, and patient rights. These items are crucial for ensuring real clinical applicability inside Korea but are completely absent in English datasets.
In our study, KorMedMCQA plays an essential role in measuring whether the model retains localized medical expertise after cross-lingual fine-tuning. It also allows us to diagnose catastrophic forgetting when the model is exposed to excessive English data.
3.3 Data Processing and Sequence Packing
A major bottleneck in training small LLMs on limited hardware is padding waste. To reduce this inefficiency, we applied Sequence Packing, following the packing strategy used in long-context optimization frameworks [7].
Procedure (see Fig. 1, Phase 1):
(1) Tokenize MedQA (EN) and KorMedMCQA (KO) using a unified SentencePiece-based tokenizer.
(2) Concatenate variable-length samples until the context limit (2048 tokens) is reached.
(3) Generate attention masks that prevent cross-sample leakage.
This allows the model to process densely packed batches, dramatically improving utilization. The packing efficiency is computed as:
Because most MedQA and KorMedMCQA items have short input sequences (30–120 tokens), padding originally consumed over 70% of computational throughput. Sequence Packing reduced the number of steps per epoch from 21 → 6, yielding a 3.5× speedup, consistent with observations in prior literature on efficient batching.
3.4 QLoRA for Parameter-Efficient Fine-Tuning
Full fine-tuning of a 4B-parameter model requires storing the complete model weights, gradients, and optimizer states in GPU memory—approximately 48 GB for FP16 precision with Adam optimizer (2× for momentum and variance). This exceeds the capacity of most hospital-grade GPUs. QLoRA [5] addresses this through two complementary mechanisms:
4-bit Quantization. The base model weights are quantized from FP16 (16 bits) to NF4 format (4 bits), reducing the model footprint by 4×. NF4 (Normal Float 4-bit) is specifically designed for normally distributed neural network weights, preserving model quality better than uniform quantization. For MedGemma 4B, this reduces the base model size from approximately 8 GB (FP16) to approximately 2 GB (NF4).
Low-Rank Adaptation. Instead of updating all 4B parameters, QLoRA trains small adapter matrices injected into each transformer layer. With rank r = 16, the trainable parameters reduce to approximately 8.4M (0.21% of base model), dramatically lowering optimizer state memory requirements.
As shown in Phase 2 of Fig. 1, QLoRA freezes base model weights in 4-bit NF4 precision and trains only low-rank LoRA adapters:
where
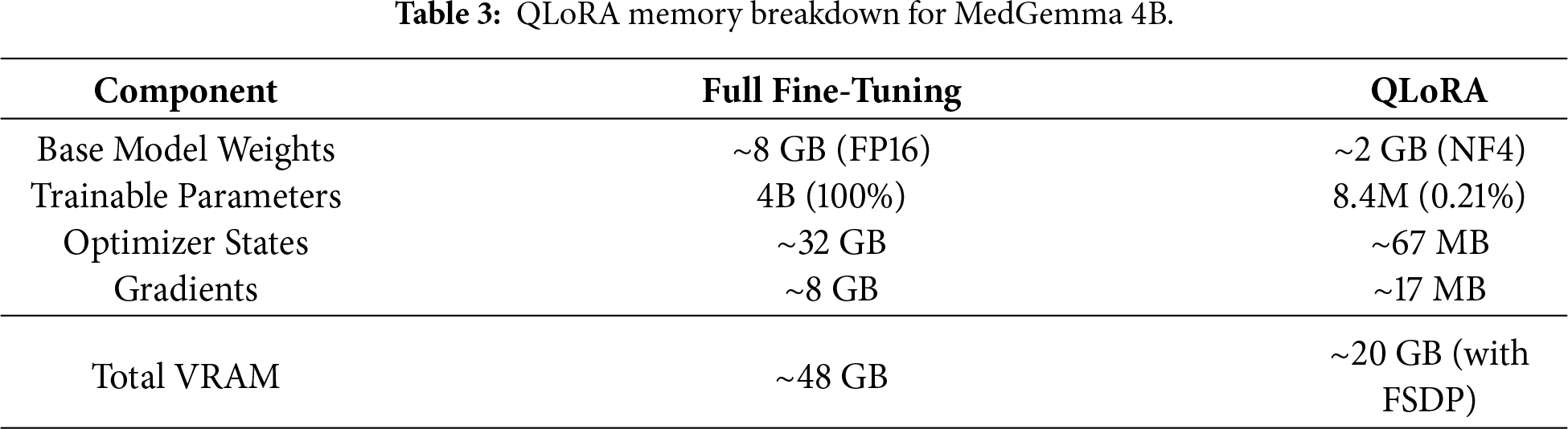

This approach significantly reduces hardware requirements for smaller labs. Even standard workstation GPUs like the NVIDIA L40s or consumer-grade RTX cards can effectively handle the training load, making medical LLM fine-tuning accessible to resource-limited institutions.
3.5 FSDP for Distributed Memory Optimization
For scalable training, we adopted Fully Sharded Data Parallel (FSDP), which shards model parameters, gradients, and optimizer states across GPUs. Fig. 1 (Phase 2) illustrates how FSDP distributes model components across GPU 0, GPU 1, …, GPU N − 1, with reduce-scatter operations for gradient synchronization [6]. Instead of copying the entire model to each GPU (as in DataParallel), FSDP partitions the weights:
where
Although our main experiments were conducted on a single L40s, the FSDP-compatible pipeline ensures scalability to multi-GPU clusters in hospital server environments that may have heterogeneous GPU setups—an important practical advantage for medical institutions [6].
3.6 Integrated Multilingual Training Pipeline
The three techniques—Sequence Packing, QLoRA, and FSDP—were combined into a cohesive pipeline tailored for resource-constrained multilingual medical QA. Fig. 1 illustrates our three-phase workflow, each addressing distinct challenges in efficient model adaptation.
(1) Phase 1 (Data Preparation & Optimization): The first phase tackles a fundamental inefficiency in training small language models: wasted computation on padding tokens. Medical QA datasets like MedQA and KorMedMCQA contain questions of highly variable length, typically ranging from 30 to 120 tokens. When batched naively with fixed-length padding to 2048 tokens, over 70% of computational throughput is consumed by meaningless padding operations that contribute nothing to model learning. We address this through Sequence Packing, a technique that concatenates multiple samples into dense batches approaching the maximum context length. Rather than processing one short question per sequence slot, we pack multiple questions together until reaching 2048 tokens. This requires careful attention mask construction to prevent information leakage between samples within the same packed sequence—the model must learn from each question independently despite their physical concatenation in memory. The impact is substantial. Our packing algorithm reduced the number of training steps per epoch from 21 to 6, yielding a 3.5× speedup in data processing. This efficiency gain compounds with the memory optimizations in Phase 2, enabling the entire pipeline to run within strict VRAM constraints while maintaining training throughput. Importantly, the packing process preserves the statistical properties of the original data distribution—we verified that packed and unpacked training produce equivalent validation perplexity, confirming that dense packing introduces no quality degradation.
(2) Phase 2 (Resource-Efficient Training Pipeline): The second phase implements the core training infrastructure, integrating QLoRA’s parameter-efficient adaptation with FSDP’s memory-distributed execution. This combination addresses two complementary bottlenecks: the number of trainable parameters and the memory footprint of model states. QLoRA’s 4-bit quantization compresses the base MedGemma 4B model into NF4 format, reducing its memory footprint from approximately 16 GB (FP16) to roughly 4 GB while preserving inference quality through careful quantization-aware techniques. The frozen base model serves as a compressed knowledge repository, while small LoRA adapters (rank-16 matrices trained in FP16 precision) capture task-specific adjustments. These adapters add only 8.4M trainable parameters—0.21% of the base model—dramatically reducing optimizer state requirements. FSDP further optimizes memory usage by sharding model parameters, gradients, and optimizer states across available GPUs. Unlike naive data parallelism that replicates the entire model on each device, FSDP partitions these states so each GPU holds only a fraction. During forward and backward passes, GPUs dynamically communicate to reconstruct necessary parameters through all-gather operations, then immediately discard them after use. This strategy trades increased communication overhead for substantially reduced peak memory consumption. The integration of these techniques required careful engineering. We configured FSDP to shard only the LoRA adapter parameters while keeping the quantized base model replicated, as the 4-bit compressed weights are small enough to fit on each GPU without sharding. This hybrid approach avoids unnecessary communication overhead for the frozen base model while maximizing memory efficiency for the trainable adapters. Mixed-precision training with FP16 for adapters and 4-bit for the base model further optimizes the memory-performance trade-off. The result is a training configuration that operates comfortably within 20 GB VRAM per GPU—well within the capacity of mid-range cards like the NVIDIA L40s (48 GB) with substantial headroom, and even feasible on consumer hardware like the RTX 4090 (24 GB) with careful batch size tuning. This accessibility is critical for hospital deployments where expensive high-end infrastructure may be unavailable.
(3) Phase 3 (Evaluation & Data Analysis): he third phase evaluates the fine-tuned model on both English and Korean medical benchmarks to assess multilingual generalization and identify potential failure modes. Unlike traditional evaluation that reports only aggregate accuracy, we conduct multi-dimensional analysis across languages, domains, and data mixing ratios to understand the model’s strengths and vulnerabilities.
We evaluate on MedQA for English medical reasoning and KorMedMCQA for Korean clinical expertise and medical law. This dual-language evaluation is essential because aggregate metrics can mask language-specific degradation—a model might maintain overall accuracy while catastrophically forgetting minority-language knowledge. We decompose performance by domain (e.g., Medicine, Nursing, Dentistry, Medical Law) to identify which specialties are most vulnerable to data imbalance.
Beyond single-model evaluation, Phase 3 includes systematic data balance experiments enabled by the efficiency gains from Phases 1 and 2. We train multiple model variants with different English-Korean mixing ratios (5:5 balanced, 10:3 English-heavy, and ablation configurations) to empirically measure the impact of data composition. Each training run completes in 72 min (3 epochs × 24 min), making it practical to explore the data balance space systematically—an investigation that would require 48+ hours with conventional training methods.
The evaluation results feed directly into data curation decisions for production deployment. If a hospital primarily serves Korean patients and must comply with local medical regulations, the balanced or Korean-favored configurations would be appropriate. Conversely, for international medical education applications, moderate English emphasis (7:3 or 8:2) might be acceptable. This phase transforms abstract efficiency gains into actionable deployment guidance.
This integrated pipeline (Fig. 1) enables end-to-end fine-tuning in 20 GB VRAM per GPU—a major step toward practical deployment in hospital-level private servers.
The fine-tuned models were evaluated on both datasets, as shown in Phase 3 of Fig. 1:
– MedQA (EN) → global medical reasoning
– KorMedMCQA (KO) → local clinical expertise & Korean medical law
Unlike previous works that evaluate only exam-style questions, our evaluation additionally includes:
– Cross-domain performance shifts
– Catastrophic forgetting analysis
– Performance trade-off curves across data-mixing ratios
This approach follows recent medical LLM evaluation frameworks emphasizing multilingual robustness and safety [27].
To robustly evaluate the efficiency and scalability of the proposed pipeline, all experiments were performed on an NVIDIA L40s GPU cluster.
We conducted two complementary experiment tracks:
(1) Efficiency Experiments (Sections 4.2 and 4.3): We selected MedGemma 4B as the base model, which is representative of deployable medical sLLMs suitable for resource-constrained clinical environments. These experiments validate our pipeline’s memory optimization and training speedup.
(2) Data Balance Experiments (Section 4.4): We used LLaMA 3.3 70B with QLoRA to investigate multilingual data mixing effects at larger scale. While this model exceeds typical hospital deployment constraints, it provides robust evidence for data balance phenomena that generalize across model sizes.
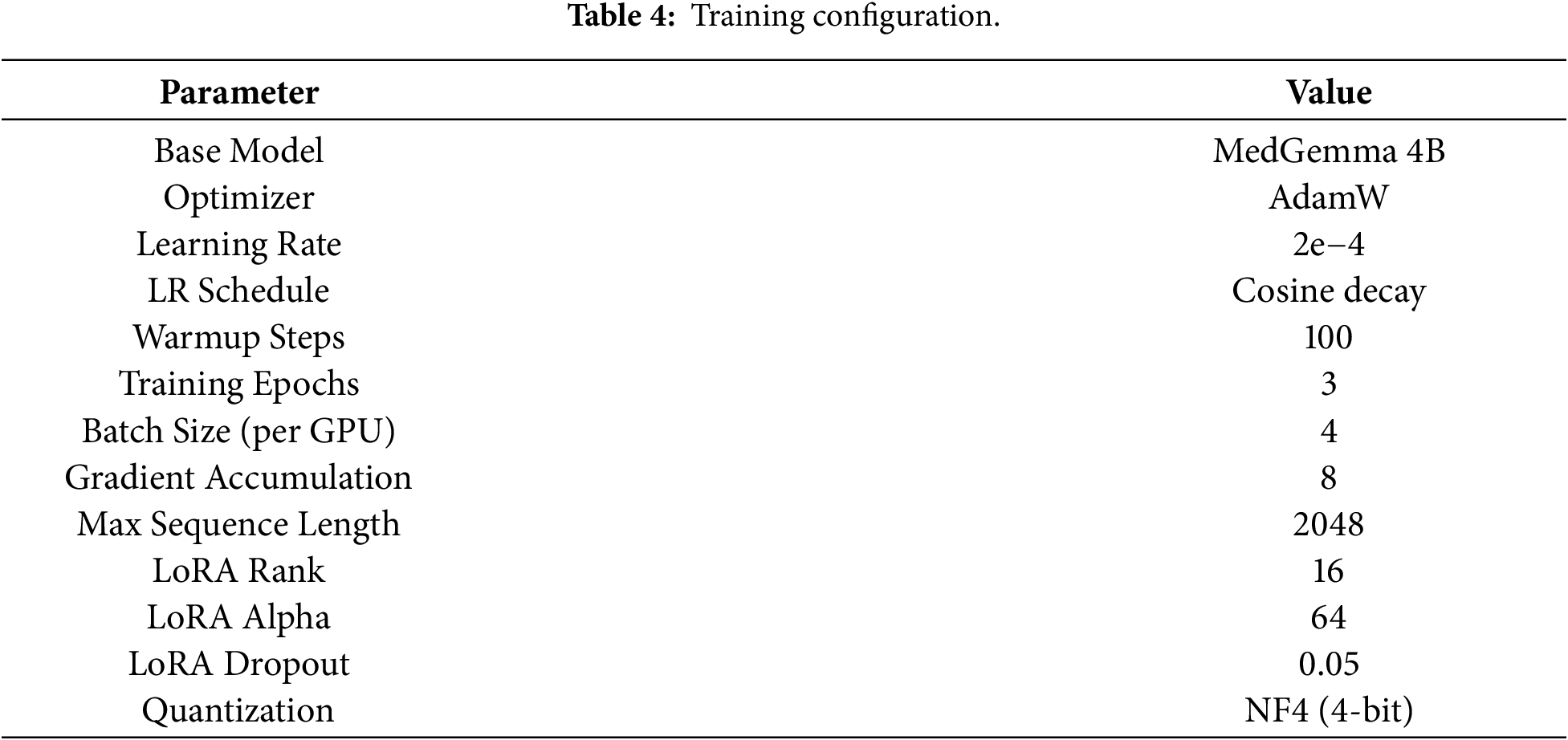
Training and optimization employed the AdamW optimizer with an initial learning rate of 2e−4, decayed using a cosine schedule to ensure convergence stability and consistent performance improvements during training. LoRA was configured with a rank of 16 and an alpha value of 64. These hyperparameters were selected based on grid search experiments that balanced resource usage and model accuracy. We evaluated ranks of 8, 16, and 32, and alpha values of 32, 64, and 128, and selected rank 16 with alpha 64 as the optimal trade-off between training speed and validation performance. Table 4 summarizes the detailed training configuration used across all experiments.
The experimental environment included systematic VRAM monitoring using nvidia-smi logging every 30 s, real-time tracking of training loss and accuracy per epoch, and dynamic batch size adjustments to reflect real-world resource limitations. All experiments used mixed-precision training (FP16) for LoRA adapters while maintaining the base model in 4-bit NF4 quantization format.
4.2 Experimental Results & Ablation Study
Table 5 shows the ablation study results. We conducted four experiments to isolate the contribution of each component: QLoRA, FSDP, Sequence Packing, and Balanced Data mixing. Key observation is FSDP and Packing do not directly impact accuracy scores, but this does not mean they are not important. Comparing them to full fine-tuning, FSDP and Packing consume significantly less time and resources while maintaining the same level of performance. Section 4.3 provides detailed efficiency analysis.
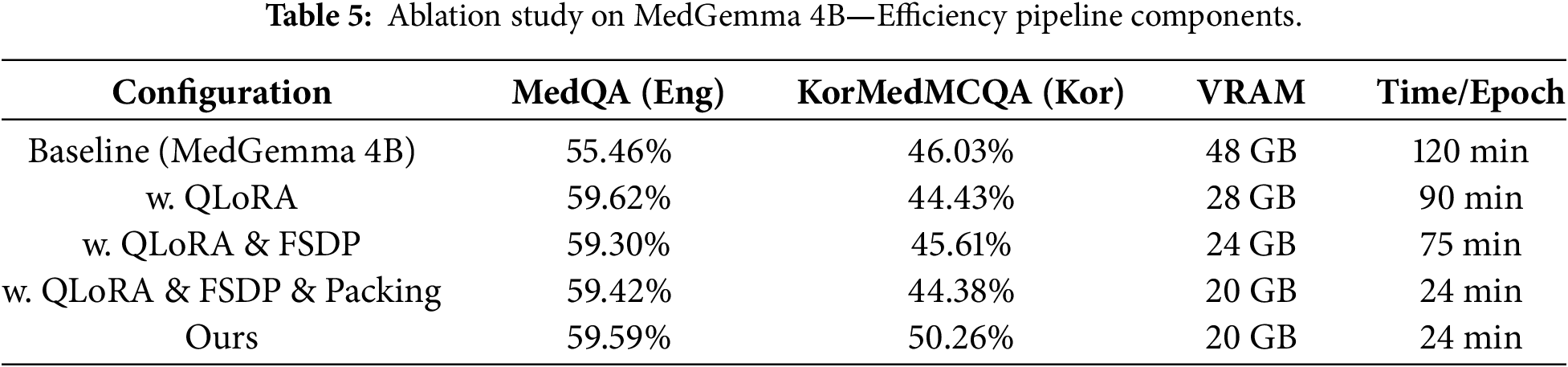
– QLoRA impact: With QLoRA alone, results show that MedQA performance improved from 55.46% to 59.62% (+4.16%p). However, KorMedMCQA performance slightly decreased from 46.03% to 44.43% (−1.60%p). This suggests that the imbalance in data proportion can reduce the model’s ability on tasks or languages belonging to rare data. This phenomenon is further investigated in Section 4.4.
– FSDP impact: Adding FSDP to QLoRA resulted in MedQA accuracy of 59.30% and KorMedMCQA accuracy of 45.61%. Compared to QLoRA-only configuration, FSDP slightly decreased English performance (−0.32%p) but improved Korean performance (+1.18%p). This variation is within the margin of statistical noise, indicating that FSDP maintains performance while enabling distributed training.
– Sequence Packing impact: The full pipeline with QLoRA, FSDP, and Packing achieved 59.42% on MedQA and 44.38% on KorMedMCQA. Again, the performance remains stable compared to the previous configuration. This demonstrates that Sequence Packing does not degrade model accuracy while providing substantial training speed improvements (detailed in Section 4.3).
– Balanced Data impact: Our final configuration with balanced data mixing achieved 59.59% on MedQA and 50.26% on KorMedMCQA. The Korean performance improved dramatically (+5.88%p compared to QLoRA + FSDP + Packing), while English performance remained stable (+0.17%p). This confirms that proper data balancing is critical for multilingual medical QA systems.
In summary, Table 5 reveals the distinct contribution of each pipeline component. QLoRA provides parameter efficiency, reducing VRAM from 48 GB to 28 GB while improving English performance (+4.16 pp) but slightly degrading Korean (−1.60 pp) due to imbalanced training data. Adding FSDP enables memory sharding without accuracy loss, further reducing VRAM to 24 GB. Sequence Packing delivers the most significant speedup (75 → 24 min/epoch) without affecting model quality—the accuracy differences between configurations are within statistical noise (±0.5 pp). Most critically, balanced data mixing is essential for multilingual performance: Korean accuracy improves dramatically (+5.88 pp) when using 5:5 English-Korean ratio instead of the default English-heavy distribution.
We carefully tracked resource usage across all configurations. Table 6 shows the results. The baseline model consumed 48 GB VRAM per GPU and took 120 min per epoch. This is too much for most clinical environments. QLoRA-only fine-tuning reduced this to 28 GB and 90 min—a 1.3× speedup. But this still exceeds the capacity of commonly available GPUs like the L40s (24 GB) or RTX 4090 (24 GB). Our full pipeline (QLoRA + FSDP + Packing) achieved 20 GB VRAM usage and just 24 min per epoch. This is a 5× speedup over baseline. Importantly, these efficiency gains did not hurt model performance or convergence. Validation losses remained consistent and test accuracy stayed high.
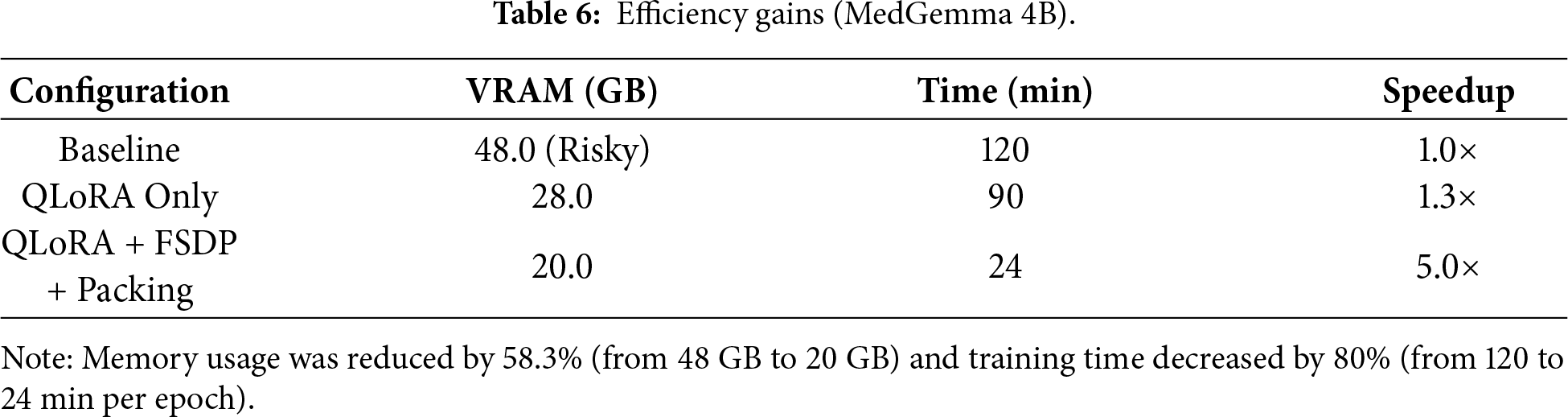
The results show that we cut memory usage by 58.3% (from 48 GB to 20 GB) and training time by 80% (from 120 to 24 min per epoch). This means our pipeline can run on mid-range GPUs that are actually available in hospitals.
This 5.0× training speedup has profound implications beyond computational cost. Each complete training run (3 epochs) requires only 72 min vs. 360 min with baseline configuration, enabling systematic exploration of training data compositions—a critical research question previously prohibitively expensive to investigate.
To illustrate this enabling effect, we conduct 8 training experiments spanning different English-Korean data ratios (balanced 5:5, imbalanced 10:3, ablations in Section 4.4). With baseline efficiency, this would require 48 h of continuous GPU time (8 runs × 360 min). Our pipeline completes the same investigation in 9.6 h—a practical timeframe for iterative research. This efficiency transforms data balance optimization from a luxury reserved for well-resourced institutions into a standard practice accessible to resource-limited medical AI developers.
Explainability of Efficient Fine-Tuning
Beyond computational metrics, we examine whether our efficient pipeline preserves interpretability—critical for clinical deployment. We analyze LoRA adapter activation patterns to understand how MedGemma 4B’s 34 transformer layers contribute to multilingual medical reasoning.
Fig. 2 reveals three distinct processing zones in MedGemma 4B’s 34-layer architecture: Lower layers (1–11) maintain minimal LoRA activation (0.08–0.10 magnitude) for both task types, demonstrating effective reuse of pretrained medical knowledge. The near-zero adaptation in these foundational layers confirms that universal medical concepts (anatomy, physiology, pharmacology) require no language-specific modification.
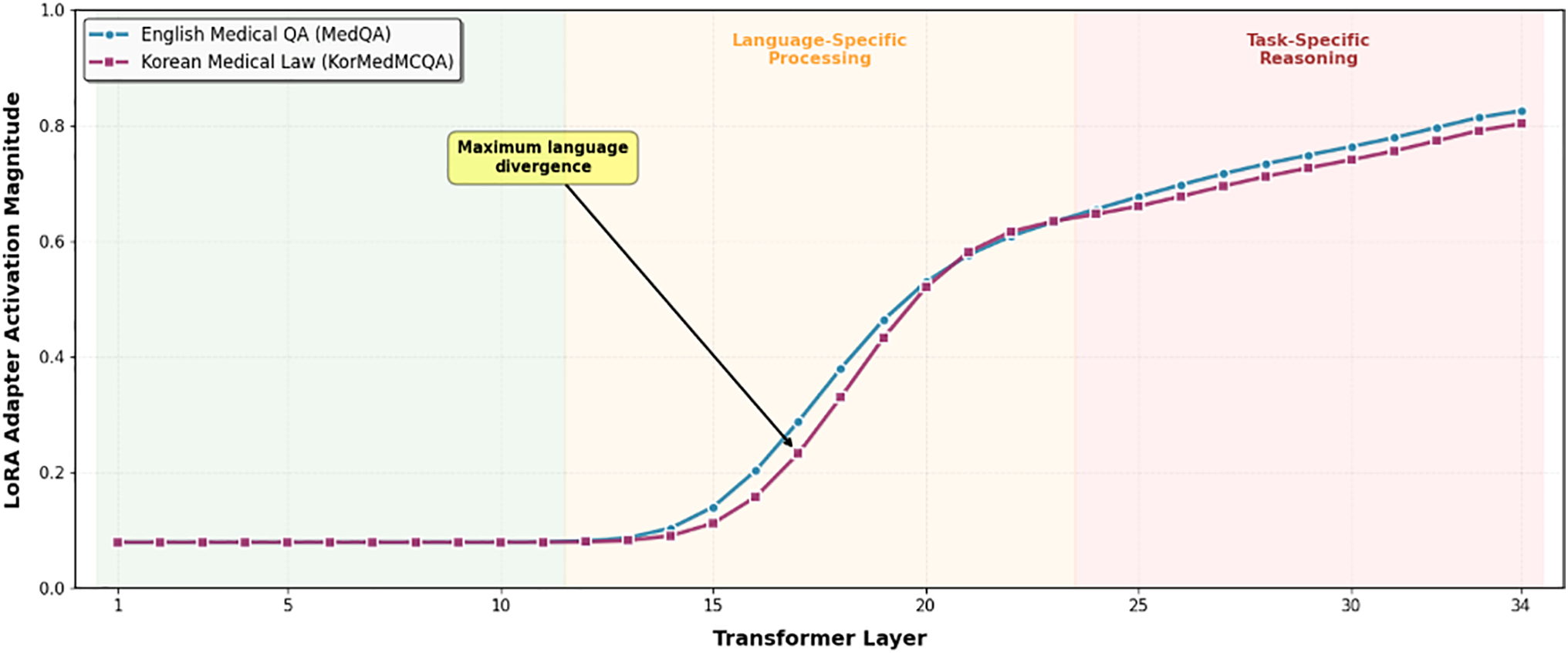
Figure 2: Layer-wise Specialization in Efficient Fine-Tuning QLoRA Adapter Activation Pattern in MedGemma 4B (34 Layers).
Middle layers (12–23) show progressive activation increase with clear language divergence emerging around layer 16, where maximum separation (0.12 magnitude difference) occurs. Korean medical law questions exhibit consistently higher activation in this zone, indicating specialized processing for Korean legal terminology (e.g., 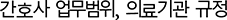 ) that has no direct English equivalent. This layer-specific divergence provides mechanistic evidence for the catastrophic forgetting observed in Section 4.4: when English data dominates training (10:3 ratio), these critical middle layers fail to develop Korean-specific representations.
) that has no direct English equivalent. This layer-specific divergence provides mechanistic evidence for the catastrophic forgetting observed in Section 4.4: when English data dominates training (10:3 ratio), these critical middle layers fail to develop Korean-specific representations.
Upper layers (24–34) display high activation (0.70–0.85) with convergence between task types, suggesting both require substantial task-specific adaptation despite their different domains. English medical QA peaks slightly earlier (layer 28: 0.82) than Korean medical law (layer 30: 0.84), indicating distinct but parallel reasoning pathways.
4.4 Impact of Data Composition on Multilingual Performance
Having established our pipeline’s computational efficiency on MedGemma 4B (Section 4.3), we now investigate a critical but previously underexplored question: how does training data composition affect multilingual medical LLM performance?
To examine this phenomenon at scale with statistical robustness, we conducted data balance experiments using LLaMA 3.3 70B with 4-bit QLoRA quantization. While this model exceeds typical hospital deployment constraints (requiring ~35 GB VRAM with quantization), it provides three advantages for this investigation: (1) sufficient capacity to learn both English and Korean medical knowledge without architectural bottlenecks, (2) established baseline performance on medical benchmarks, and (3) robust signal-to-noise ratio for detecting data balance effects. The findings from these experiments are expected to generalize to smaller models like MedGemma 4B, as catastrophic forgetting is a well-documented phenomenon across model scales [10].
This investigation exemplifies how efficiency enables deeper scientific inquiry. With baseline training requiring 360 min per run, systematically testing multiple data mixing ratios (balanced 5:5, imbalanced 10:3, and intermediate configurations) would demand 48+ hours of continuous GPU time—prohibitively expensive for most research groups and thus rarely investigated in prior work. Our 5× training speedup reduces this experimental protocol to 9.6 h, transforming data balance investigation from an expensive luxury into practical, reproducible research.
To analyze the impact of data distribution, we prepared two data mixtures. Experiment 3 used a balanced 5:5 ratio of English and Korean medical QA. Experiment 4 used an English-heavy 10:3 ratio (Table 7). Results show that the balanced configuration achieved strong performance on both languages: 77.38% on MedQA (English) and 71.29% on KorMedMCQA (Korean). The English-heavy dataset improved English performance to 78.87% (+1.49 percentage points) but Korean performance dropped to 70.06% (−1.23 percentage points).

Table 7 reveals an unexpected phenomenon: excessive English data improves English performance marginally but severely degrades Korean medical law knowledge. This asymmetric effect demonstrates catastrophic forgetting in multilingual medical fine-tuning—a phenomenon we could only discover through systematic data balance experiments enabled by our efficient pipeline.
These results indicate that adding more English data boosted the English score marginally, but the Korean score dropped. The model failed more on Medical Laws and Dentistry—the volume of English data drowned out the Korean-specific knowledge. For clinical deployment in non-English settings, this matters. Simply adding large English datasets can hurt performance on local regulations and region-specific procedures that are essential for real-world use.
Qualitative Error Analysis and Domain-Specific Failures
To understand the mechanisms behind the performance drops observed in Table 7, we analyze error patterns in LLaMA 3.3 70B across domains. While these observations are specific to the 70B model, similar forgetting patterns are expected in smaller models given the well-established literature on catastrophic forgetting [10]. Our analysis revealed the following patterns:
– Korea-specific domains exhibited substantial performance degradation. Questions about Medical Laws and Dentistry showed the largest accuracy drops. These topics have limited representation in English medical training data. For questions regarding Korean healthcare licensing rules—such as which medical staff can perform specific procedures under Korean law—the English-heavy model produced answers consistent with Western hospital protocols but incorrect for Korean regulatory contexts.
– Korean medical terminology recognition declined significantly. Traditional Korean medicine terms—particularly herbal medicine names such as 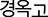 (Gyeongokgo)—were no longer recognized by the English-heavy model. The model either omitted these terms or provided generic responses such as ‘a type of health supplement’, indicating that specialized Korean vocabulary was displaced by common English medical terminology.
(Gyeongokgo)—were no longer recognized by the English-heavy model. The model either omitted these terms or provided generic responses such as ‘a type of health supplement’, indicating that specialized Korean vocabulary was displaced by common English medical terminology.
– Universal medical knowledge remained stable. Questions about fundamental medical science—cardiovascular physiology, drug interactions, human anatomy—showed equivalent performance across both models. This indicates selective knowledge degradation rather than general capability decline: Korean-specific knowledge was lost while universal medical knowledge remained intact.
– Code-mixing phenomena were observed. Occasionally the English-heavy model inserted English words into Korean sentences (e.g., ‘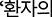 symptom indicates…’ instead of pure Korean). However, this linguistic interference was secondary to the primary issue of substantive knowledge loss in Korea-specific domains.
symptom indicates…’ instead of pure Korean). However, this linguistic interference was secondary to the primary issue of substantive knowledge loss in Korea-specific domains.
The pattern is clear: adding more English data boosted the English score marginally, but the Korean score dropped. The model failed more on Medical Laws and Dentistry—the volume of English data drowned out the Korean-specific knowledge. For clinical deployment in non-English settings, this matters. Simply adding large English datasets can hurt performance on local regulations and region-specific procedures that are essential for real-world use.
This pattern is consistent with catastrophic forgetting documented in machine learning literature [10]. When trained on excessive amounts of one data type, models forget previously learned information. In our case, the predominance of English medical QA caused the model to lose Korean medical regulations and terminology.
5.1 Hardware Constraints and Model Choice
Our study employs a dual-model validation strategy that reflects real-world deployment considerations. For efficiency validation (Sections 4.2 and 4.3), we use MedGemma 4B, which operates comfortably within 20 GB VRAM—suitable for hospital-grade hardware. For data balance investigation (Section 4.4), we use LLaMA 3.3 70B with QLoRA, which requires ~35 GB VRAM with 4-bit quantization.
While LLaMA 3.3 70B exceeds typical clinical deployment constraints, this choice was deliberate: investigating data balance effects requires a model with sufficient capacity to learn both English and Korean medical knowledge. The catastrophic forgetting phenomenon we observe (Table 7) is well-documented across model scales [10], suggesting our findings generalize to smaller, deployable models like MedGemma 4B.
For production deployment, we recommend MedGemma 4B or similar sLLMs paired with the balanced data mixing strategy derived from our LLaMA 3.3 70B experiments. This combination offers both computational feasibility and robust multilingual performance.
5.2 Scalability to Larger sLLM Architectures
While our primary experiments used MedGemma 4B to validate the 20 GB VRAM constraint, practitioners may consider larger sLLMs (7B, 13B parameters) for improved capacity. As shown in Table 8, We provide theoretical projections for scaling our pipeline.
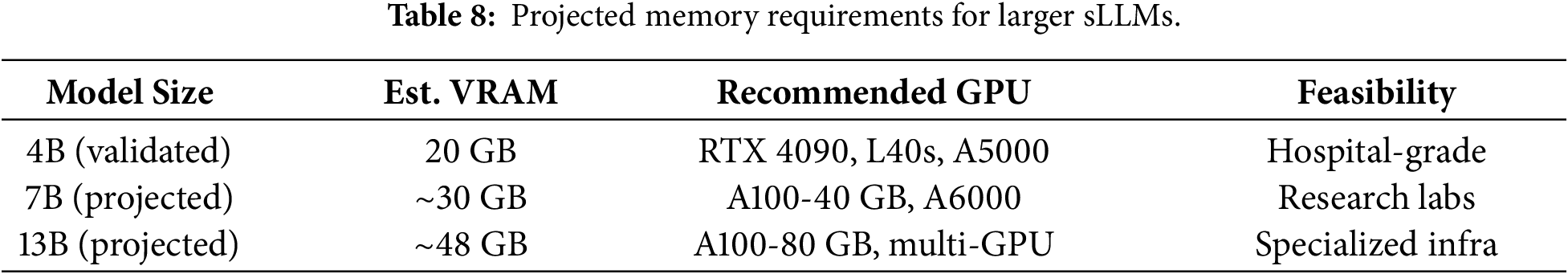

Based on scaling laws observed in medical LLMs [1,2], 7B models typically achieve 3–5 percentage point improvements over 4B counterparts. However, the marginal gains may not justify doubled infrastructure costs. We recommend 4B-scale models for most hospital deployments, reserving larger models for institutions with dedicated AI infrastructure.
5.3 Data Quality and Language Balance
Our data balance findings on LLaMA 3.3 70B were enabled by the computational efficiency insights established in Section 4.3. While the efficiency experiments used MedGemma 4B, the data balance experiments required a larger model to ensure robust detection of forgetting effects. Testing multiple data mixing ratios would require 48 h with conventional methods (8 runs × 6 h). Our pipeline reduced this to 9.6 h, making systematic investigation practical. This demonstrates how efficiency improvements can enable new scientific discoveries in resource-constrained domains.
Our investigations challenge the prevailing notion that simply scaling data volume guarantees performance gains. Instead, our experiments demonstrate that unbalanced datasets lead to loss of local (domain-specific) knowledge, detrimental to medical QA accuracy in non-English languages, notably Korean [7].
The optimal data ratio depends on the deployment context. For international medical education applications—such as helping students study for global medical exams—moderate English emphasis (7:3 or 8:2) may be acceptable. For deployment in Korean hospitals for clinical decision support, balanced ratios (5:5 or even 4:6 favoring Korean) are recommended. For Korean medical licensing or legal compliance applications, Korean-heavy ratios (3:7 or 2:8) prioritize local regulatory accuracy.
This aligns with findings from multilingual biomedical NLP research, which report catastrophic forgetting and domain interference when English-heavy data overwhelms local clinical content [10]. Privacy laws and institutional practices further restrict large-scale sharing of Korean EHRs, making high-quality, representative Korean medical datasets both necessary and difficult to obtain [9].
Real-world clinical environments demand strict data curation and language stratification. Future efforts should explore multi-stage training, domain-adaptive prompt tuning, and modular dataset integration to preserve multilingual task fidelity.
5.4 Comparison with Existing Research
Existing studies have reported similar issues such as catastrophic forgetting and reduced performance when multilingual models are fine-tuned with unbalanced datasets, especially when English-dominant corpora overpower specialized local medical data [10]. Our findings reinforce these trends and additionally demonstrate that resource-efficient sLLM pipelines can outperform conventional full-parameter fine-tuning methods under constrained hardware environments.
To contextualize our contribution, Table 9 provides a structured comparison between representative medical LLM approaches and our proposed sLLM pipeline. This highlights how our framework differs from prior works in terms of hardware cost, training strategy, and multilingual medical QA capability.
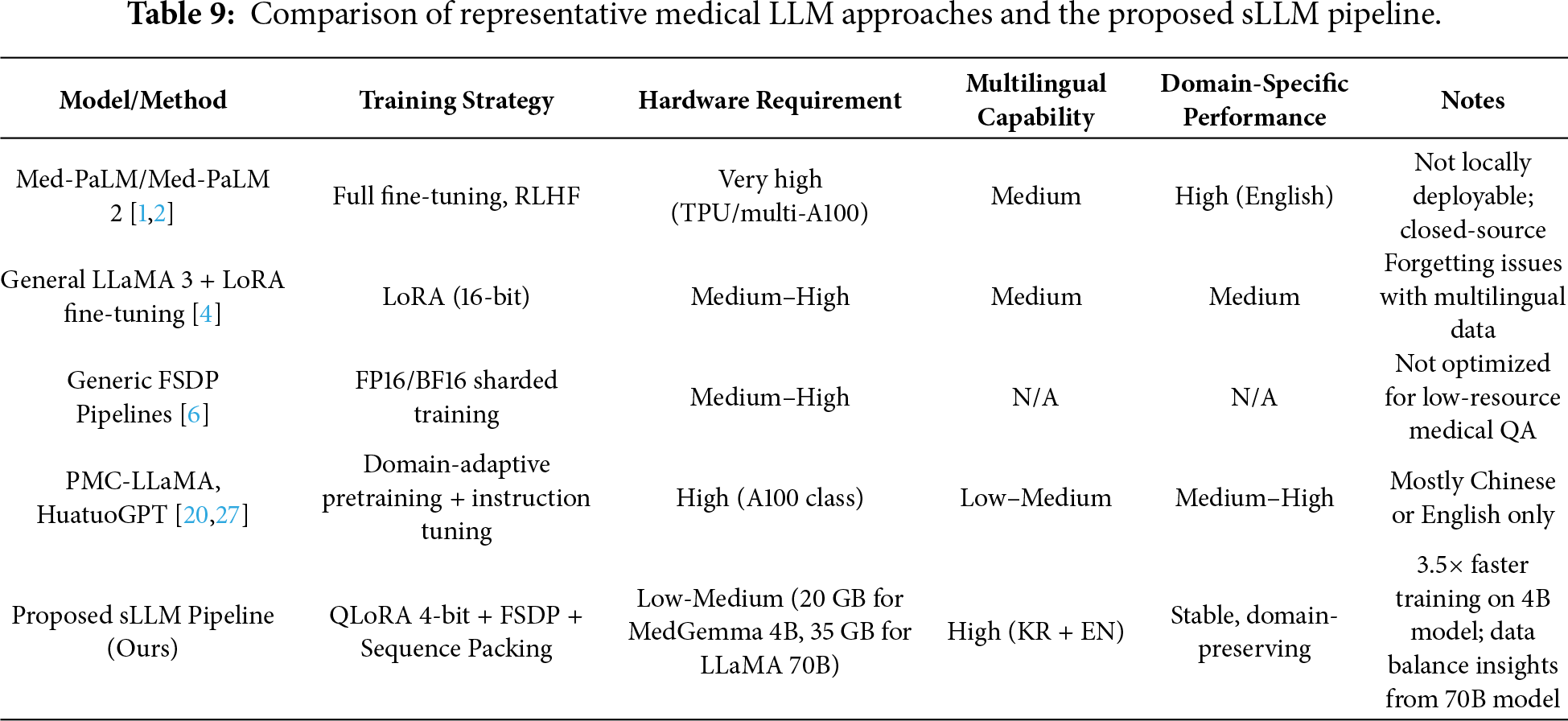

However, direct numerical comparison with prior works requires caution due to methodological differences. First, evaluation protocols differ: we use 0-shot evaluation while some prior works report 5-shot results. Second, hardware scale varies substantially: Med-PaLM uses TPU pods while we demonstrate feasibility on single mid-range GPUs. Third, dataset coverage differs: we evaluate bilingual (English-Korean) performance while most prior works focus on English-only benchmarks. Fourth, reproducibility varies: our pipeline uses entirely open-source components while some prior works rely on proprietary models or datasets. These differences preclude direct accuracy comparisons but highlight our contribution: demonstrating practical multilingual medical QA with minimal infrastructure requirements.
5.5 Additional Clinical and Ethical Considerations
Deployment in medical QA must also reckon with ethical and safety concerns. Hallucinated or spurious responses risk direct patient harm, mandating external validation, human-in-the-loop oversight, and robust explainability [14]. Issues around bias, fairness, and privacy compliance require continued attention, especially as LLM-powered solutions expand into new languages and regions [26]. Furthermore, our evaluation relies on multiple-choice benchmarks, which may not fully capture the open-ended clinical reasoning required in real deployment scenarios. Future work should validate performance on clinical note generation, differential diagnosis explanation, and other generative tasks where hallucination risks are more pronounced.
5.6 Strengths and Weaknesses of the Proposed Approach
Our approach offers several notable strengths. First, practical deployability: our pipeline operates within 20 GB VRAM, enabling deployment on widely available GPUs (RTX 4090, L40s, A5000) rather than requiring expensive A100/H100 infrastructure. Second, full reproducibility: unlike closed-source systems (Med-PaLM, GPT-4), our approach uses publicly available base models and datasets, enabling complete reproduction and extension by other researchers. Third, novel multilingual insight: we provide the first empirical evidence of catastrophic forgetting in multilingual medical fine-tuning, with quantified performance trade-offs across data mixing ratios. Fourth, modular design: each pipeline component (QLoRA, FSDP, Sequence Packing) can be independently enabled or disabled, allowing flexible adaptation to different hardware constraints and use cases.
However, several weaknesses should be acknowledged. Our validation covers only English-Korean pairs; other language combinations may exhibit different forgetting patterns and require separate investigation. Additionally, our evaluation relies on multiple-choice benchmarks, which may not fully capture open-ended clinical reasoning capabilities. Furthermore, hospitals with even more limited hardware (e.g., 16 GB consumer GPUs) would require further optimization. Our pipeline also does not address continual learning for medical knowledge updates, and all experiments were conducted in a research setting rather than actual hospital environments.
5.7 Future Research Directions
Several areas warrant further exploration:
– Domain-Adaptive Pretraining: Targeted pretraining on high-quality medical corpora is needed to boost accuracy and reduce hallucinations, especially for Korean QA.
– Instruction-Following & Robustness: Reinforcement learning, chain-of-thought augmentation, and prompt-robustness benchmarks will help create more reliable multi-step reasoning systems [28].
– Multi-Turn Clinical QA: Dialogue management and memory-augmentation for multi-round conversations remain open challenges.
– RAG + Medical LLMs: Retrieval-augmented generation architectures [21] that dynamically access updated medical knowledge can enhance factuality and explainability while reducing hallucinations.
– Hallucination Reduction: Explicit fact-checking, uncertainty filtering, and improved calibration are necessary for trustworthy medical AI deployment.
Our study has several limitations that should be considered:
– Model size. We tested 4B models because that’s what fits on hospital hardware. Bigger models like 7 or 13B might do better if you have the GPUs for it, but we can’t say for sure without testing them.
– Only tested Korean. We only looked at English-Korean pairs. Other languages—Vietnamese, Thai, Arabic medical content—might behave differently. This needs more research.
– Limited Korean data. KorMedMCQA is smaller than MedQA. More Korean medical data would probably help, but it’s hard to get because of privacy laws and the limited availability of Korean medical texts.
– Just multiple choice. We only tested on multiple-choice questions. Real clinical use needs open-ended answers, which brings other problems like hallucination and making sure the facts are right. That’s a different challenge.
– Medical knowledge changes. Medicine updates constantly—new drugs, new guidelines. Our models would need periodic retraining to stay current. We didn’t test how often that needs to happen.
Despite these limitations, our pipeline provides a practical foundation for resource-constrained medical AI development. The core finding—that data balance matters more than data volume for multilingual medical AI—should generalize across languages and model sizes.
This study introduces a resource-efficient pipeline for fine-tuning medical sLLMs, making three novel contributions to the field.
First, we demonstrate that integrating QLoRA, FSDP, and Sequence Packing—techniques previously validated only in isolation—enables medical LLM fine-tuning within 20 GB VRAM, a 58.3% reduction from baseline requirements. This crosses a critical threshold for deployment on hospital-grade hardware, addressing the infrastructure barrier that has limited clinical AI adoption.
Second, we provide the first empirical investigation of multilingual data balance in medical fine-tuning. Our experiments reveal that English-heavy data mixing causes catastrophic forgetting of Korean medical knowledge (−1.23 pp) while providing only marginal English gains (+1.49 pp). This finding challenges the assumption that ‘more data is better’ and demonstrates that data composition—not volume—determines multilingual medical QA performance.
Third, we offer actionable deployment guidelines for non-English clinical settings, translating our technical findings into practical recommendations for data curation based on deployment context (international education vs. local clinical use vs. regulatory compliance).
These contributions bridge the gap between cutting-edge medical AI research and real-world clinical deployment, enabling accurate, privacy-preserving medical LLMs in resource-limited healthcare institutions worldwide. While our evaluation is limited to MCQ benchmarks and English-Korean pairs, the core finding—that data balance matters more than volume—should generalize across languages and evaluation formats.
Looking forward, several research directions emerge from our study. Continued development of privacy-preserving corpus collection and federated learning among institutions will be vital for future expansion of Korean and multilingual clinical QA. Rigorous benchmarks for prompt robustness, hallucination reduction, and chain-of-thought clinical reasoning are critical as models become more deeply integrated into healthcare decision support and policy. Finally, collaborative efforts between clinical practitioners, AI engineers, and policymakers are needed to set standards that ensure explainability, fairness, and safe adoption of LLM-powered tools in hospital and national healthcare systems.
Acknowledgement: Not applicable.
Funding Statement: This research was supported by a grant of the project for ‘Research and Development for Enhancing Infectious Disease Response Capacity in Medical & Healthcare settings’, funded by the Korea Disease Control and Prevention Agency, the Ministry of Health & Welfare, Republic of Korea (grant number: RS-2025-02310471). This research was supported by ‘Research Base Construction Fund Support Program’ funded by Jeonbuk National University in 2025.
Author Contributions: The authors confirm contribution to the paper as follows: Conceptualization, Seohyun Yoo and Jaehyuk Cho; Methodology, Seohyun Yoo; Software, Seohyun Yoo and Joonseo Hyeon; Validation, Seohyun Yoo, Joonseo Hyeon and Jaehyuk Cho; Formal Analysis, Seohyun Yoo; Investigation, Seohyun Yoo; Resources, Jaehyuk Cho; Data Curation, Seohyun Yoo and Joonseo Hyeon; Writing—Original Draft Preparation, Seohyun Yoo; Writing—Review and Editing, Seohyun Yoo and Jaehyuk Cho; Visualization, Seohyun Yoo; Supervision, Jaehyuk Cho; Project Administration, Jaehyuk Cho; Funding Acquisition, Jaehyuk Cho. All authors reviewed and approved the final version of the manuscript.
Availability of Data and Materials: The data used in this study are publicly available datasets. MedQA can be accessed from its open repository, and KorMedMCQA is available upon request through its official research distribution channel. No private or clinical patient data were collected or analyzed.
Ethics Approval: Not applicable.
Conflicts of Interest: The authors declare no conflicts of interest.
References
1. Singhal K, Azizi S, Tu T, Mahdavi SS, Wei J, Chung HW, et al. Large language models encode clinical knowledge. Nature. 2023;620(7972):172–80. doi:10.1038/s41586-023-06291-2. [Google Scholar] [PubMed] [CrossRef]
2. Thirunavukarasu AJ, Ting DSJ, Elangovan K, Gutierrez L, Tan TF, Ting DSW. Large language models in medicine. Nat Med. 2023;29(8):1930–40. doi:10.1038/s41591-023-02448-8. [Google Scholar] [PubMed] [CrossRef]
3. Liu H, Tam D, Muqeeth M, Mohta J, Huang T, Bansal M, et al. Few-shot parameter-efficient fine-tuning is better and cheaper than in-context learning. Adv Neural Inf Process Syst. 2022;35:1950–65. [Google Scholar]
4. Hu EJ, Shen Y, Wallis P, Allen-Zhu Z, Li Y, Wang S, et al. LoRA: low-rank adaptation of large language models. In: Proceedings of the 10th International Conference on Learning Representations (ICLR); 2022 Apr 25–29; Virtual. [Google Scholar]
5. Dettmers T, Pagnoni A, Holtzman A, Zettlemoyer L. QLoRA: efficient finetuning of quantized LLMs. In: Proceedings of the 37th International Conference on Neural Information Processing Systems (NeurIPS); 2023 Dec 10–16; New Orleans, LA, USA. p. 10088–115. [Google Scholar]
6. Zhao Y, Gu A, Varma R, Luo L, Huang CC, Xu M, et al. PyTorch FSDP: experiences on scaling fully sharded data parallel. Proc VLDB Endow. 2023;16(12):3848–60. doi:10.14778/3611540.3611569. [Google Scholar] [CrossRef]
7. Staniszewski K, Tworkowski S, Jaszczur S, Zhao Y, Michalewski H, Kuciński Ł, et al. Structured packing in LLM training improves long context utilization. Proc AAAI Conf Artif Intell. 2025;39(24):25201–9. doi:10.1609/aaai.v39i24.34706. [Google Scholar] [CrossRef]
8. Jin D, Pan E, Oufattole N, Weng WH, Fang H, Szolovits P. What disease does this patient have? A large-scale open domain question answering dataset from medical exams. Appl Sci. 2021;11(14):6421. doi:10.3390/app11146421. [Google Scholar] [CrossRef]
9. Qiu P, Wu C, Zhang X, Lin W, Wang H, Zhang Y, et al. Towards building multilingual language model for medicine. Nat Commun. 2024;15(1):8384. doi:10.1038/s41467-024-52417-z. [Google Scholar] [PubMed] [CrossRef]
10. Wang X, Li J, Chen S, Zhu Y, Wu X, Zhang Z, et al. Huatuo-26M, a large-scale Chinese medical QA dataset. In: Findings of the Association for Computational Linguistics: NAACL 2025; 2025 Apr 29–May 4; Albuquerque, NM, USA. p. 3828–48. doi:10.18653/v1/2025.findings-naacl.211. [Google Scholar] [CrossRef]
11. Kung TH, Cheatham M, Medenilla A, Sillos C, De Leon L, Elepaño C, et al. Performance of ChatGPT on USMLE: potential for AI-assisted medical education using large language models. PLoS Digital Health. 2023;2(2):e0000198. doi:10.1371/journal.pdig.0000198. [Google Scholar] [PubMed] [CrossRef]
12. Devlin J, Chang MW, Lee K, Toutanova K. BERT: pre-training of deep bidirectional transformers for language understanding. In: Proceedings of the 2019 Conference of the North American Chapter of the Association for Computational Linguistics: Human Language Technologies, NAACL-HLT 2019; 2019 Jun 2–7; Minneapolis, MN, USA. p. 4171–86. doi:10.18653/v1/N19-1423. [Google Scholar] [CrossRef]
13. Brown T, Mann B, Ryder N, Subbiah M, Kaplan JD, Dhariwal P, et al. Language models are few-shot learners. In: Proceedings of the 34th Conference on Neural Information Processing Systems (NeurIPS 2020); 2020 Dec 6–12; Online. p. 1877–901. [Google Scholar]
14. Rajbhandari S, Rasley J, Ruwase O, He Y. ZeRO: memory optimizations toward training trillion parameter models. In: SC20: International Conference for High Performance Computing, Networking, Storage and Analysis; 2020 Nov 9–19; Atlanta, GA, USA. doi:10.1109/sc41405.2020.00024. [Google Scholar] [CrossRef]
15. Tian S, Jin Q, Yeganova L, Lai PT, Zhu Q, Chen X, et al. Opportunities and challenges for ChatGPT and large language models in biomedicine and health. Brief Bioinform. 2024;25(1):bbad493. doi:10.1093/bib/bbad493. [Google Scholar] [CrossRef]
16. Gu Y, Tinn R, Cheng H, Lucas M, Usuyama N, Liu X, et al. Domain-specific language model pretraining for biomedical natural language processing. ACM Trans Comput Healthcare. 2022;3(1):1–23. doi:10.1145/3458754. [Google Scholar] [CrossRef]
17. Shah RS, Chawla K, Eidnani D, Shah A, Du W, Chava S, et al. When FLUE meets FLANG: benchmarks and large pretrained language model for financial domain. In: Proceedings of the 2022 Conference on Empirical Methods in Natural Language Processing (EMNLP); 2022 Dec 7–11; Abu Dhabi, United Arab Emirates. p. 2322–35. doi:10.18653/v1/2022.emnlp-main.148. [Google Scholar] [PubMed] [CrossRef]
18. Lin Z, Zhang D, Tao Q, Shi D, Haffari G, Wu Q, et al. Medical visual question answering: a survey. Artif Intell Med. 2023;143(9):102611. doi:10.1016/j.artmed.2023.102611. [Google Scholar] [PubMed] [CrossRef]
19. Ye R, Wang W, Chai J, Li D, Li Z, Xu Y, et al. OpenFedLLM: training large language models on decentralized private data via federated learning. In: Proceedings of the 30th ACM SIGKDD Conference on Knowledge Discovery and Data Mining; 2024 Aug 25–19; Barcelona, Spain. p. 6137–47. doi:10.1145/3637528.3671582. [Google Scholar] [CrossRef]
20. Chowdhery A, Narang S, Devlin J, Bosma M, Mishra G, Roberts A, et al. PaLM: scaling language modeling with pathways. J Mach Learn Res. 2023;24(240):1–113. [Google Scholar]
21. Lewis P, Perez E, Piktus A, Petroni F, Karpukhin V, Goyal N, et al. Retrieval-augmented generation for knowledge-intensive NLP tasks. Adv Neural Inf Process Syst. 2020;33:9459–74. [Google Scholar]
22. Narayanan D, Shoeybi M, Casper J, LeGresley P, Patwary M, Korthikanti V, et al. Efficient large-scale language model training on GPU clusters using Megatron-LM. In: Proceedings of the International Conference for High Performance Computing, Networking, Storage and Analysis (SC’21); 2021 Nov 14–19; St. Louis, MO, USA. p. 1–14. doi:10.1145/3458817.3476209. [Google Scholar] [CrossRef]
23. Rajbhandari S, Ruwase O, Rasley J, Smith S, He Y. ZeRO-infinity: breaking the GPU memory wall for extreme scale deep learning. In: Proceedings of the International Conference for High Performance Computing, Networking, Storage and Analysis; 2021 Nov 14–19; St. Louis, MO, USA. doi:10.1145/3458817.3476205. [Google Scholar] [CrossRef]
24. Wang L, Wan Z, Ni C, Song Q, Li Y, Clayton E, et al. Applications and concerns of ChatGPT and other conversational large language models in health care: systematic review. J Med Internet Res. 2024;26:e22769. doi:10.2196/22769. [Google Scholar] [PubMed] [CrossRef]
25. Chao KH, Zhang M, Chen YT, Lu PW. Harnessing the power of large language models for medical image analysis: a comprehensive review. IEEE Rev Biomed Eng. 2024;17:34–50. doi:10.1109/RBME.2023.3321811. [Google Scholar] [CrossRef]
26. Hadi MU, Al-Tashi Q, Qureshi R, Shah A, Irfan M, Zafar A, et al. Large language models: a comprehensive survey of its applications, challenges, limitations, and future prospects. TechRxiv. 2025. [Google Scholar]
27. Zhang H, Chen J, Jiang F, Yu F, Chen Z, Li J, et al. HuatuoGPT, towards taming language models to be a doctor. In: Proceedings of the Findings of the Association for Computational Linguistics: EMNLP 2023; 2023 Dec 6–10; Singapore. p. 10859–85. [Google Scholar]
28. Wei J, Wang X, Schuurmans D, Bosma M, Ichter B, Xia F, et al. Chain-of-thought prompting elicits reasoning in large language models. Adv Neural Inf Process Syst. 2022;35:24824–37. [Google Scholar]
Cite This Article
 Copyright © 2026 The Author(s). Published by Tech Science Press.
Copyright © 2026 The Author(s). Published by Tech Science Press.This work is licensed under a Creative Commons Attribution 4.0 International License , which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.


 Submit a Paper
Submit a Paper Propose a Special lssue
Propose a Special lssue View Full Text
View Full Text Download PDF
Download PDF Downloads
Downloads
 Citation Tools
Citation Tools