 Open Access
Open Access
ARTICLE
Automated Severity Classification of Knee Osteoarthritis from Radiographs Using Transfer Learning Based Deep Neural Networks
National Institute of Electronics and Information Technology (NIELIT), Srinagar, India
* Corresponding Author: Syed Nisar Hussain Bukhari. Email:
Journal on Artificial Intelligence 2026, 8, 137-152. https://doi.org/10.32604/jai.2026.077943
Received 19 December 2025; Accepted 28 January 2026; Issue published 11 March 2026
Abstract
Knee osteoarthritis is a progressive degenerative joint disorder that leads to pain, stiffness, and reduced mobility, significantly affecting quality of life. Early and reliable diagnosis is essential for effective disease management, yet conventional radiographic assessment remains time-consuming and subject to inter-observer variability. This study presents a comparative deep learning (DL) based approach for automated severity classification of knee osteoarthritis using plain radiographic images. Multiple pretrained convolutional neural network architectures, including EfficientNetB3, InceptionNet, VGG19, ResNet, and EfficientNetV2S, were evaluated within a transfer learning paradigm. All models were trained and assessed on a publicly available dataset to classify knee osteoarthritis severity into clinically relevant categories. Among the evaluated architectures, EfficientNetB3 demonstrated the most consistent performance, achieving an accuracy of 0.97. Statistical significance analysis further confirmed that the performance differences between EfficientNetB3 and the other models were significant. The results indicate that modern DL architectures can provide reliable and consistent severity assessment, supporting their potential use as clinical decision support tools for knee osteoarthritis diagnosis.Keywords
Knee osteoarthritis (KO) is one of the most common chronic musculoskeletal disorders and represents a major cause of disability among the adult population worldwide [1]. The disease primarily affects older individuals, with prevalence increasing significantly with age due to progressive degeneration of joint tissues. Epidemiological studies indicate that a large proportion of individuals above the age of fifty exhibit fradiographic signs of osteoarthritic changes, even in the absence of pronounced clinical symptoms. The condition is characterized by gradual degradation of articular cartilage, leading to joint space narrowing, altered biomechanics, and increased friction between articulating surfaces. As the disease progresses, patients experience persistent pain, stiffness, reduced range of motion, swelling, and difficulty in performing weight-bearing activities, all of which adversely affect functional independence and quality of life [2]. In advanced stages, when conservative management strategies such as physical therapy, pharmacological intervention, or lifestyle modification fail to provide adequate relief, total knee arthroplasty often becomes the final treatment option. While effective, this surgical procedure is invasive, costly, and associated with prolonged recovery periods. Consequently, early diagnosis and accurate grading of disease severity are essential to guide appropriate treatment decisions, delay structural deterioration, and improve long-term clinical outcomes [3].
Clinical diagnosis of KO continues to rely heavily on the interpretation of plain radiographic images. Radiologists assess structural indicators such as tibiofemoral joint space width, osteophyte formation, subchondral sclerosis, and surface deformities to estimate disease severity [4]. These features form the basis of the Kellgren Lawrence grading system, which categorizes osteoarthritis into discrete stages ranging from normal to severe [5]. While widely used in clinical practice, this grading process requires substantial expertise and remains susceptible to inter-observer and intra-observer variability, particularly in early and moderate stages where radiographic differences are subtle [6]. In addition, the increasing volume of medical imaging data places a growing burden on radiology workflows, further motivating the need for automated and reliable diagnostic support systems. Fig. 1 illustrates how joint space narrowing is used as a central indicator of cartilage loss. Healthy knees show a clear and preserved joint space, while moderate and severe conditions exhibit progressive narrowing that reflects structural degeneration and loss of load-bearing capacity. Fig. 2 provides an enlarged view of a healthy and an osteoarthritic knee, highlighting pathological changes such as cartilage erosion, the formation of osteophytes, and the thickening of subchondral bone. These observable characteristics help clinicians identify the stage of knee osteoarthritis, but their interpretation requires considerable expertise.
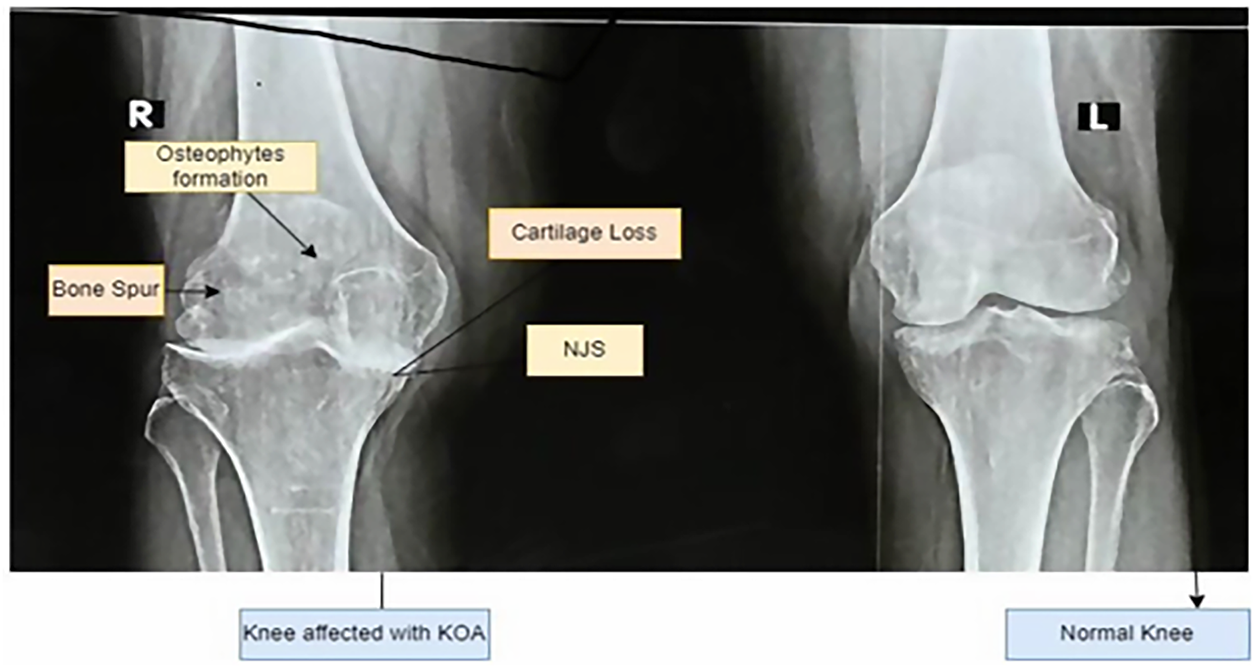
Figure 1: Knee with Osteoarthritis and without Osteoarthritis.
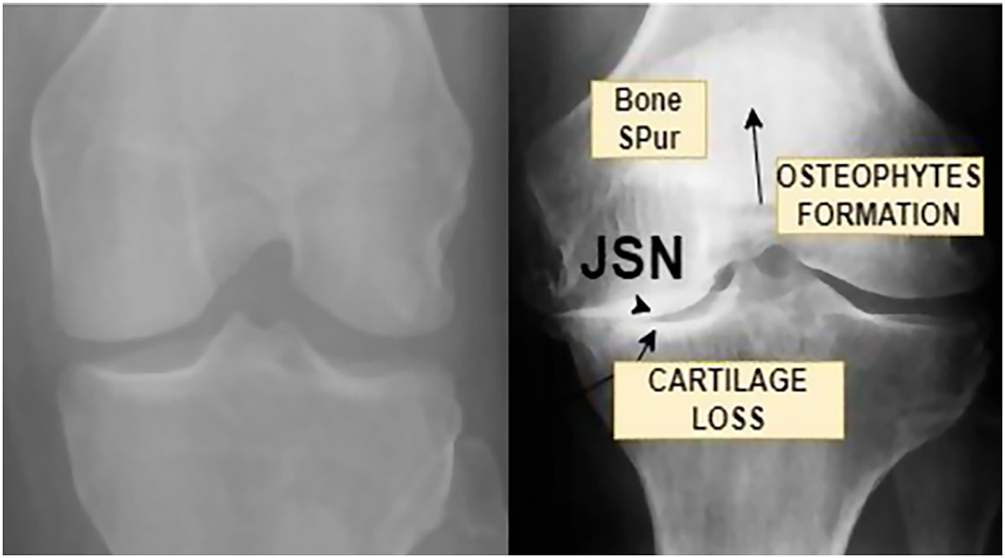
Figure 2: Enlarged view of a normal knee and Knee osteoarthritis affected.
Traditional ML approaches have attempted to address these limitations by extracting handcrafted features related to texture, shape, or intensity patterns. However, such methods depend heavily on domain-specific feature engineering and often fail to capture the complex and heterogeneous patterns present in knee radiographs. These shortcomings have driven increasing interest in deep learning techniques, which can learn discriminative features directly from imaging data without manual intervention [7].
Recent advances in deep learning (DL) have transformed medical image analysis by enabling models to automatically identify both global structural changes and fine-grained textural variations within radiographs. Convolutional neural networks (CNNs), in particular, have demonstrated strong performance in a range of diagnostic imaging tasks due to their ability to model hierarchical feature representations [6]. When combined with transfer learning (TL) strategies, these models can be effectively adapted to medical datasets, even when labeled data are limited. Automated systems built on such architectures have the potential to reduce subjectivity, improve reproducibility, and provide consistent decision support in clinical environments.
In this study, multiple pre-trained CNN architectures are investigated within a TL framework for automated classification of KO severity from radiographic images. The evaluated models include EfficientNetB3, InceptionNet, VGG19, ResNet, and EfficientNetV2S, each representing distinct architectural design principles and feature extraction capabilities. The dataset is organized into three clinically relevant categories: healthy, moderate, and severe. Careful preprocessing and balancing strategies are applied to ensure robust learning across all severity levels.
The primary objective of this research is to develop and validate an automated DL based system capable of classifying KO severity with high reliability using plain radiographs. By providing consistent and objective assessments, the proposed framework aims to support clinicians in routine diagnostic workflows, reduce variability in disease grading, and contribute to improved management of knee osteoarthritis in clinical practice.
The application of artificial intelligence (AI) to musculoskeletal imaging has gained substantial attention in recent years, with knee osteoarthritis emerging as a prominent research focus due to its high prevalence and diagnostic complexity [6]. Conventional diagnostic workflows typically involve physical examination combined with radiological assessment using X-ray or magnetic resonance imaging [5]. Although widely adopted, these approaches are inherently subjective and dependent on clinician expertise, which can result in variability across institutions and observers. One of the most significant challenges identified in the literature is the limited ability of conventional imaging assessment to reliably detect early-stage osteoarthritis, where morphological changes are minimal and difficult to quantify [7]. The anatomical complexity of the knee joint, overlapping bone structures, and gradual progression of cartilage loss further complicate visual interpretation. These challenges have motivated researchers to explore computational methods that can provide objective and reproducible assessments.
Early studies employed traditional ML techniques that relied on handcrafted features extracted from radiographs, including texture descriptors, edge information, and shape metrics [8]. While these methods demonstrated initial promise, their performance was often constrained by limited feature expressiveness and poor generalization across datasets. With the advancement of deep learning, convolutional neural networks have increasingly replaced traditional approaches due to their ability to automatically learn relevant features from raw image data [9].
Several studies have reported the successful application of CNN-based models for knee osteoarthritis classification. Researchers have explored region-specific analysis, such as focusing on the patellar or tibiofemoral regions, to improve feature discrimination [10]. Others have developed models to directly predict Kellgren Lawrence grades, demonstrating improved consistency compared to manual assessment [11]. More recent work has incorporated visualization techniques to improve interpretability by highlighting image regions that contribute most to model predictions, addressing an important requirement for clinical acceptance [12].
Advanced architectures such as ResNet and EfficientNet have further improved performance by enabling deeper networks and more efficient parameter utilization [13]. Despite these advancements, the literature reveals considerable variation in reported results, often due to differences in dataset composition, preprocessing pipelines, and handling of class imbalance [14]. Many studies also lack detailed methodological descriptions, limiting reproducibility and comparability across experiments.
These observations highlight the need for methodologically rigorous frameworks that combine modern deep learning architectures with transparent preprocessing, balanced training strategies, and comprehensive evaluation protocols. The present study addresses these gaps by systematically evaluating multiple state-of-the-art architectures within a unified experimental framework and emphasizing reproducibility and clinical relevance.
Contributions
The main contributions of this work are summarized as follows:
1. A comprehensive DL framework is developed for automated classification of KO severity from plain radiographic images.
2. Multiple state-of-the-art CNN architectures are systematically evaluated under a unified experimental setup to identify the most reliable model for KOA severity assessment.
3. A robust data preprocessing and class balancing strategy is implemented to mitigate bias and improve generalization across healthy, moderate, and severe categories.
4. The effectiveness of TL for KO classification is demonstrated, highlighting its suitability for medical imaging tasks with limited labeled data.
5. Extensive performance evaluation using clinically meaningful metrics is conducted to ensure reliability and applicability of the proposed system in real-world diagnostic settings.
The proposed methodology follows a carefully structured DL framework aimed at achieving reliable and clinically meaningful multi-class classification of knee osteoarthritis severity. The overall pipeline integrates standardized image preprocessing, explicit handling of class imbalance, advanced data augmentation strategies, and the use of multiple high-capacity pretrained convolutional neural networks within a transfer learning paradigm. This design ensures robustness, reproducibility, and fair evaluation across all disease severity categories. The complete workflow of the proposed approach is illustrated in Fig. 3.
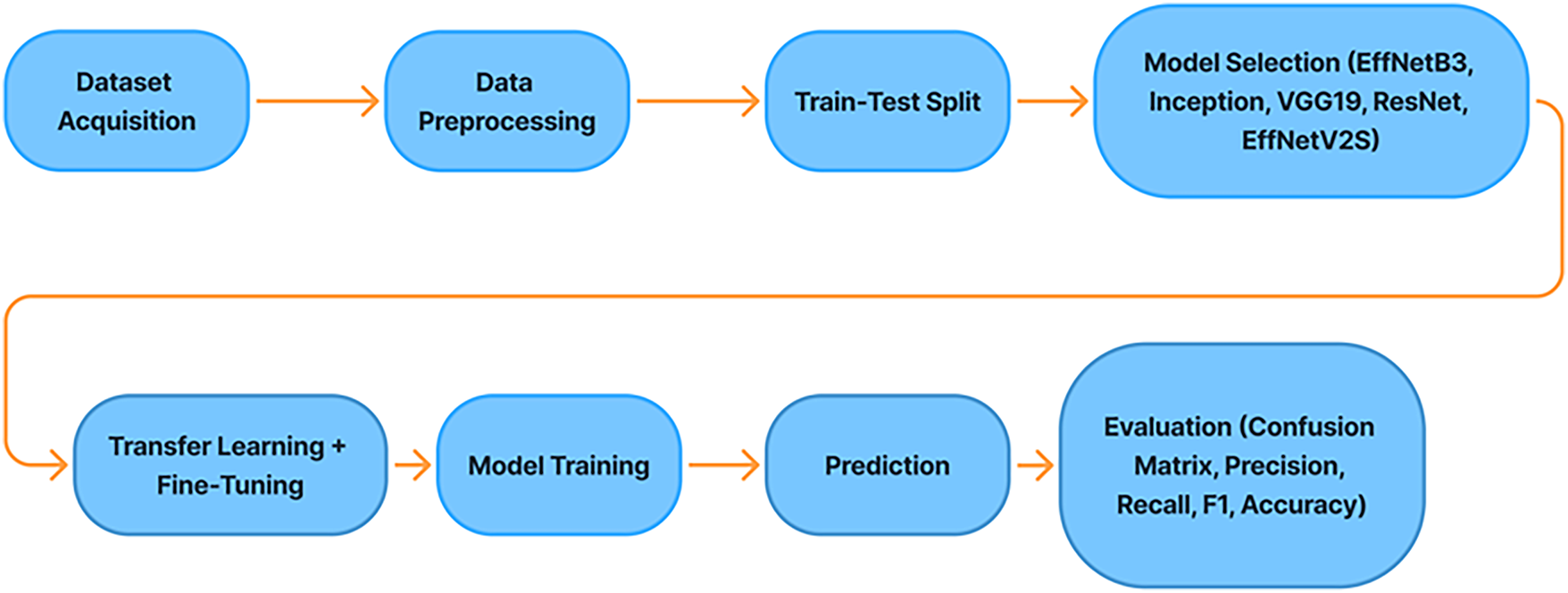
Figure 3: Proposed methodology.
During data augmentation, controlled geometric transformations were applied to the training images to introduce realistic variability while preserving anatomical plausibility. These included random rotations within ±15°, horizontal flipping, zooming within a range of 0.9–1.1, and small width and height translations of up to 10% of the image dimensions. All augmentation parameters were kept consistent across models and applied exclusively to the training set to prevent information leakage.
3.1 Data Acquisition and Pre-Processing
A publicly available dataset of knee radiographs was obtained from an open-access repository, consisting of frontal-view X-ray images annotated according to the Kellgren Lawrence grading system [15]. Each image was categorized into one of three clinically relevant severity classes: healthy, moderate, and severe. The dataset was organized into separate training, validation, and test subsets to prevent information leakage and to ensure unbiased performance assessment as shown in Table 1.
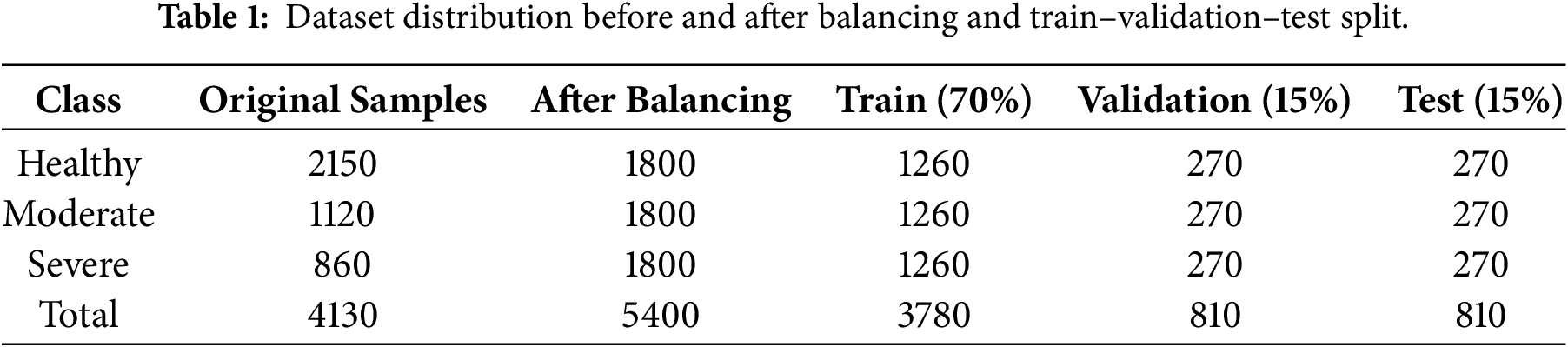
Table 1 presents the class-wise distribution of knee radiographs before and after dataset balancing, along with the allocation into training, validation, and test subsets. The dataset was split using a fixed ratio of 70% for training, 15% for validation, and 15% for testing. To avoid information leakage and ensure unbiased evaluation, class balancing and augmentation were applied exclusively to the training subset, while validation and test sets preserved the original data distribution. The dataset used in this study is publicly available and anonymized; therefore, detailed patient-level demographic information such as age, gender, or body mass index was not consistently accessible. As a result, the present work focuses exclusively on image-based severity classification, in line with prior studies using the same dataset. The training-validation-test split ratios were selected to balance robust model learning with unbiased evaluation, while preserving a sufficiently large independent test set for reliable performance assessment.
All images were standardized prior to model training. Pixel intensity normalization was applied to reduce variations arising from different imaging conditions and acquisition devices. Subsequently, all radiographs were resized to a uniform spatial resolution of 224 × 224 pixels, which is compatible with the input requirements of modern deep convolutional architectures. Initial exploratory analysis revealed a pronounced imbalance among the severity classes, a well-known issue in medical imaging that can significantly bias model learning if left unaddressed. To mitigate this problem and ensure equitable learning across classes, a dedicated two-stage data balancing strategy was incorporated, as described in the following subsection [16].
3.2 Data Balancing and Augmentation
The importance of data balancing in knee osteoarthritis classification has been consistently highlighted in prior studies, where imbalance has been shown to bias predictions toward majority classes and reduce sensitivity to moderate disease stages. Class imbalance was addressed using a combination of controlled down sampling and targeted augmentation applied exclusively to the training data. In the first stage, over-represented classes were reduced using random uniform sampling to limit their dominance during optimization [17]. This trimming step ensured that the learning process did not become biased toward majority classes, which is particularly important in multi-class clinical classification problems.
In the second stage, under-represented classes were augmented until each severity category contained an equal number of samples. Synthetic variations were generated through spatial transformations such as rotation, horizontal flipping, zooming, and small translations. These transformations were carefully selected to preserve anatomical plausibility while introducing realistic variability that improves generalization. The resulting balanced dataset allowed the models to learn class-specific patterns more uniformly and reduced sensitivity to noise and acquisition-related variations [18].
3.3 Deep Learning Architectures
To ensure a comprehensive evaluation, five pretrained CNN architectures were employed, each reflecting different design philosophies and representational strengths.
EfficientNetB3 was selected due to its compound scaling strategy, which jointly optimizes network depth, width, and input resolution using a principled scaling coefficient [17]. This design enables EfficientNet models to achieve high accuracy with fewer parameters, making them particularly effective for detecting subtle textural and structural variations in medical images.
InceptionNet employs parallel convolutional paths with multiple receptive field sizes, allowing simultaneous extraction of local and global features. This multi-scale representation is advantageous for radiographic analysis, where disease indicators may appear at different spatial resolutions.
VGG19 represents a deeper yet structurally simple architecture based on sequential stacking of small convolutional filters. Although computationally intensive, its depth enables hierarchical feature learning, making it a strong baseline for medical image classification tasks.
ResNet introduces residual connections that facilitate gradient propagation in very deep networks [19]. These skip connections allow the network to learn fine-grained anatomical details while mitigating vanishing gradient issues, which is beneficial for distinguishing closely related disease stages.
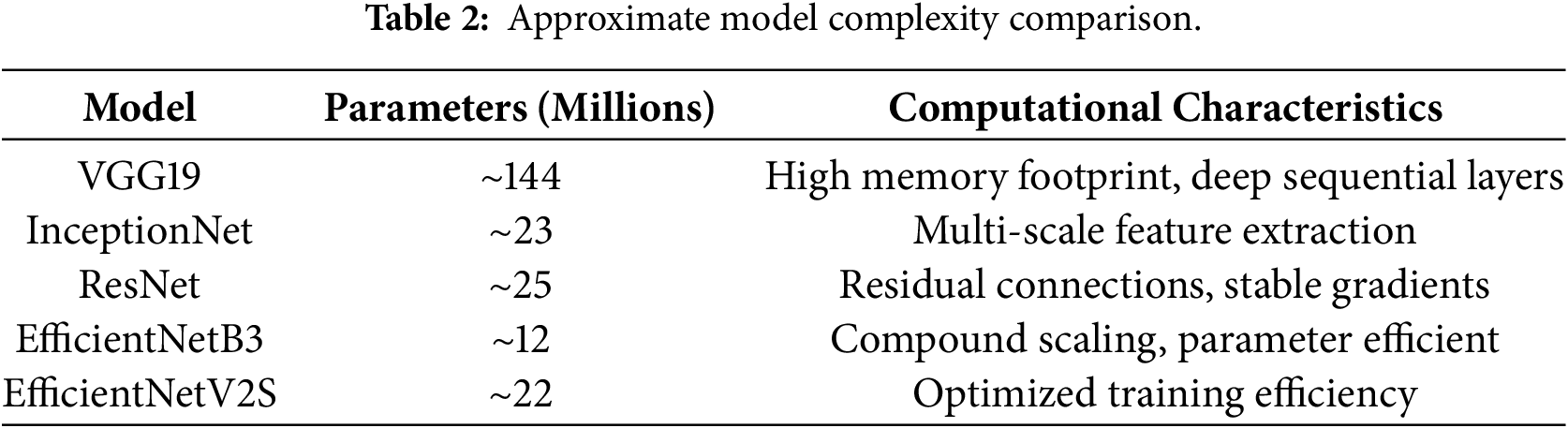
EfficientNetV2S is a more recent evolution of the EfficientNet family, optimized for faster training and improved parameter efficiency through fused convolutional blocks and refined scaling strategies. Its design is particularly well suited for medium-sized medical datasets, balancing performance and computational cost. The inclusion of these diverse architectures enabled a robust comparative analysis and ensured that the final conclusions were not dependent on a single network design.
As depicted in Table 2, EfficientNetB3 achieves a favorable balance between representational capacity and parameter efficiency, which contributes to its superior generalization performance.
3.4 Transfer Learning Strategy
Transfer learning (TL) was employed to leverage knowledge learned from large-scale natural image datasets, most notably ImageNet, where these models were originally trained. Since labeled medical datasets are often limited in size, TL provides a practical means of improving convergence speed and generalization performance [19]. During model training, the pretrained convolutional layers were retained as fixed feature extractors, while the newly added classification head was trained on the knee radiograph dataset. The custom classification head consisted of global average pooling to reduce spatial dimensionality, followed by batch normalization to stabilize learning. A fully connected layer with rectified linear activation was used to learn high-level representations relevant to KOA, and dropout regularization was applied to reduce overfitting. The final softmax layer produced probability estimates for the three severity classes [20].
All models were trained using the Adamax optimizer, chosen for its numerical stability when handling sparse or noisy gradients as shown in Algorithm 1. The categorical cross-entropy loss function was employed to reflect the multi-class nature of the classification task. Model training involved continuous monitoring of validation loss to detect overfitting, along with early stopping and checkpointing mechanisms to preserve the best-performing weights. Final evaluation was conducted exclusively on the unseen test set to provide an unbiased estimate of real-world performance [21]. The batch size was set to 32 and an initial learning rate to 1 × 10−4 during model training. Training was conducted for a maximum of 50 epochs, with early stopping applied based on validation loss using a patience of 7 epochs. Dropout regularization with a rate of 0.5 was employed in the fully connected layer of the classification head to mitigate overfitting. These hyperparameters were kept consistent across all evaluated architectures to ensure a fair and controlled comparative analysis.

A fixed train-validation-test split strategy was adopted instead of cross-validation to preserve a fully independent test set and ensure unbiased evaluation, which is particularly important in clinical imaging studies. This approach is consistent with prior deep learning-based osteoarthritis studies using comparable datasets. Model stability under controlled training conditions was further supported by consistent performance metrics and statistical significance analysis.
All experiments were conducted on a workstation equipped with an NVIDIA GPU with 12 GB memory, an Intel multi-core CPU, and 32 GB system RAM. Model training was performed using TensorFlow with GPU acceleration. On average, a single training epoch required approximately 20–30 s depending on the architecture, and full model convergence was typically achieved within 30–40 epochs due to early stopping. The total training time for individual models ranged from approximately 15 to 25 min. Inference time per image was on the order of milliseconds, supporting the feasibility of the proposed framework for practical clinical deployment.
Model performance was assessed using a comprehensive set of evaluation metrics commonly adopted in medical image analysis. Confusion matrix analysis was used to examine class-specific prediction behavior, providing insights into misclassification patterns and potential confusion between adjacent severity grades [22]. This analysis is particularly important for KOA, where moderate cases often share visual similarities with neighboring stages.
Precision, recall, and F1-score were computed separately for each class to capture different aspects of diagnostic reliability [23]. Precision reflects the model’s ability to avoid false positive predictions, while recall measures sensitivity to true disease cases [24]. The F1-score provides a balanced assessment by combining both measures, especially under conditions of class imbalance.
Overall accuracy was reported to summarize global correctness, while class-wise accuracy ensured that performance for each severity category was evaluated independently [25]. In addition, macro-averaged metrics treated all classes equally, providing a fairness-oriented evaluation, whereas weighted averages incorporated class frequencies to reflect real-world data distributions [26]. Together, these metrics offer a statistically robust and clinically meaningful assessment of model performance [27]
To assess the stability of the proposed framework, experiments were conducted using fixed data splits and controlled training configurations. Minor variability across repeated training sessions was observed to be negligible, and the reported results reflect stable model behavior under consistent experimental settings. Statistical significance analysis was further performed using McNemar’s test to validate performance differences among the evaluated models.
The performance of five DL architectures, namely EfficientNetB3, EfficientNetV2S, InceptionNet, VGG19, and ResNet, was evaluated on an independent test set to assess their effectiveness in classifying KO severity into Healthy, Moderate, and Severe categories. Model performance was quantified using precision, recall, F1-score, and overall accuracy, providing a comprehensive assessment of classification reliability [28,29]. The comparative results are summarized in Table 3, while the Confusion matrix of EfficientNetB3 is depicted in Fig. 4 and training and validation behavior of the best-performing model is illustrated through accuracy and loss curves in Fig. 5.
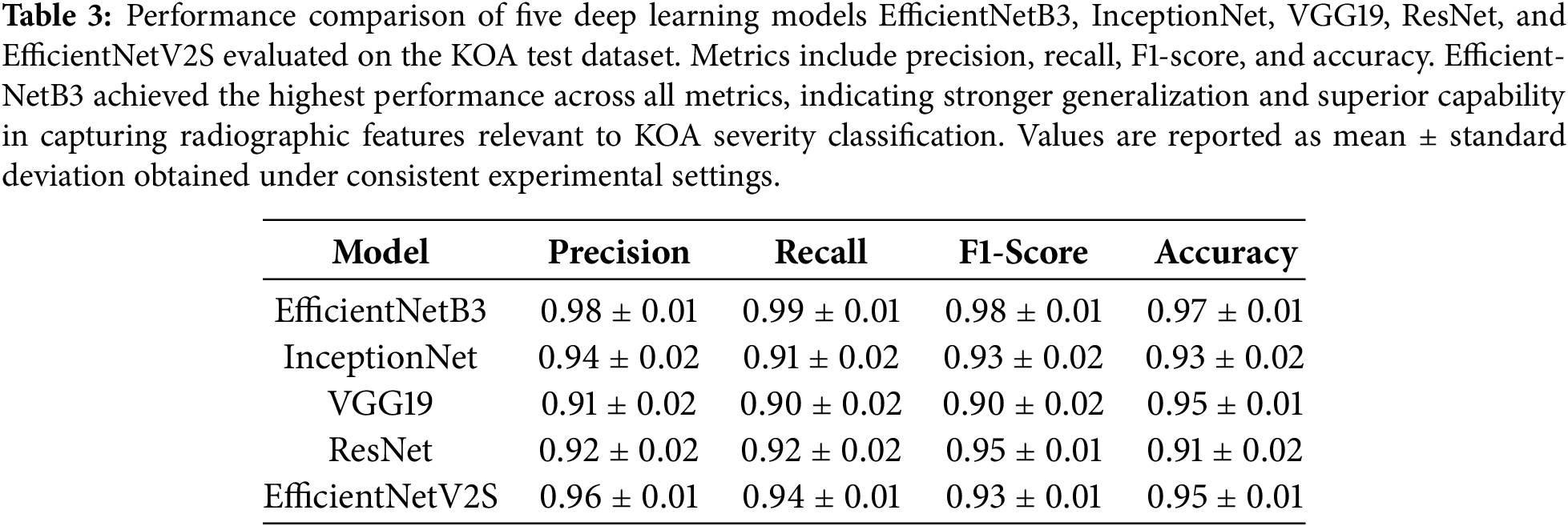
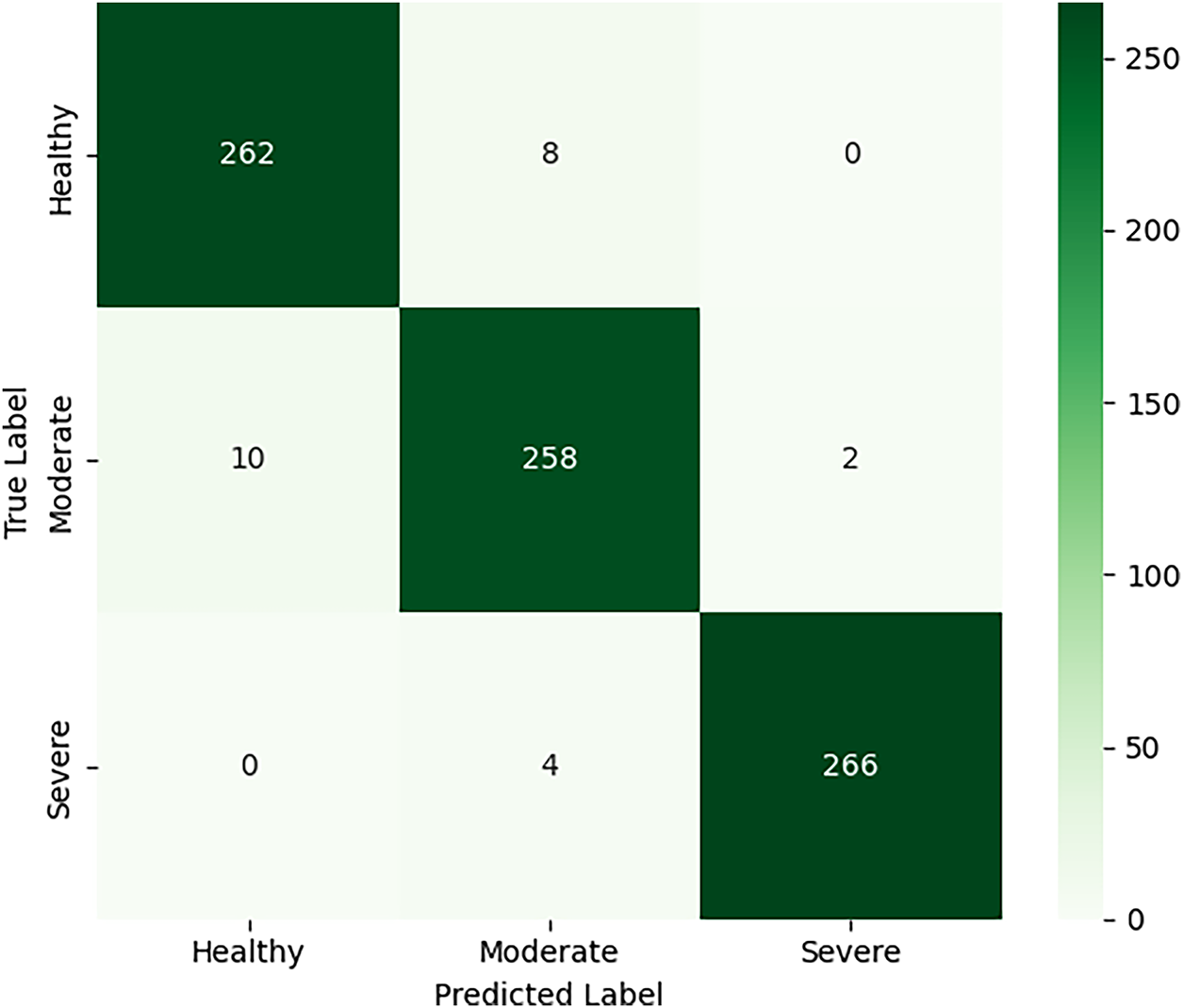
Figure 4: Confusion matrix of the EfficientNetB3 model on the independent test set for knee osteoarthritis severity classification. The matrix illustrates class-wise prediction performance across Healthy, Moderate, and Severe categories.
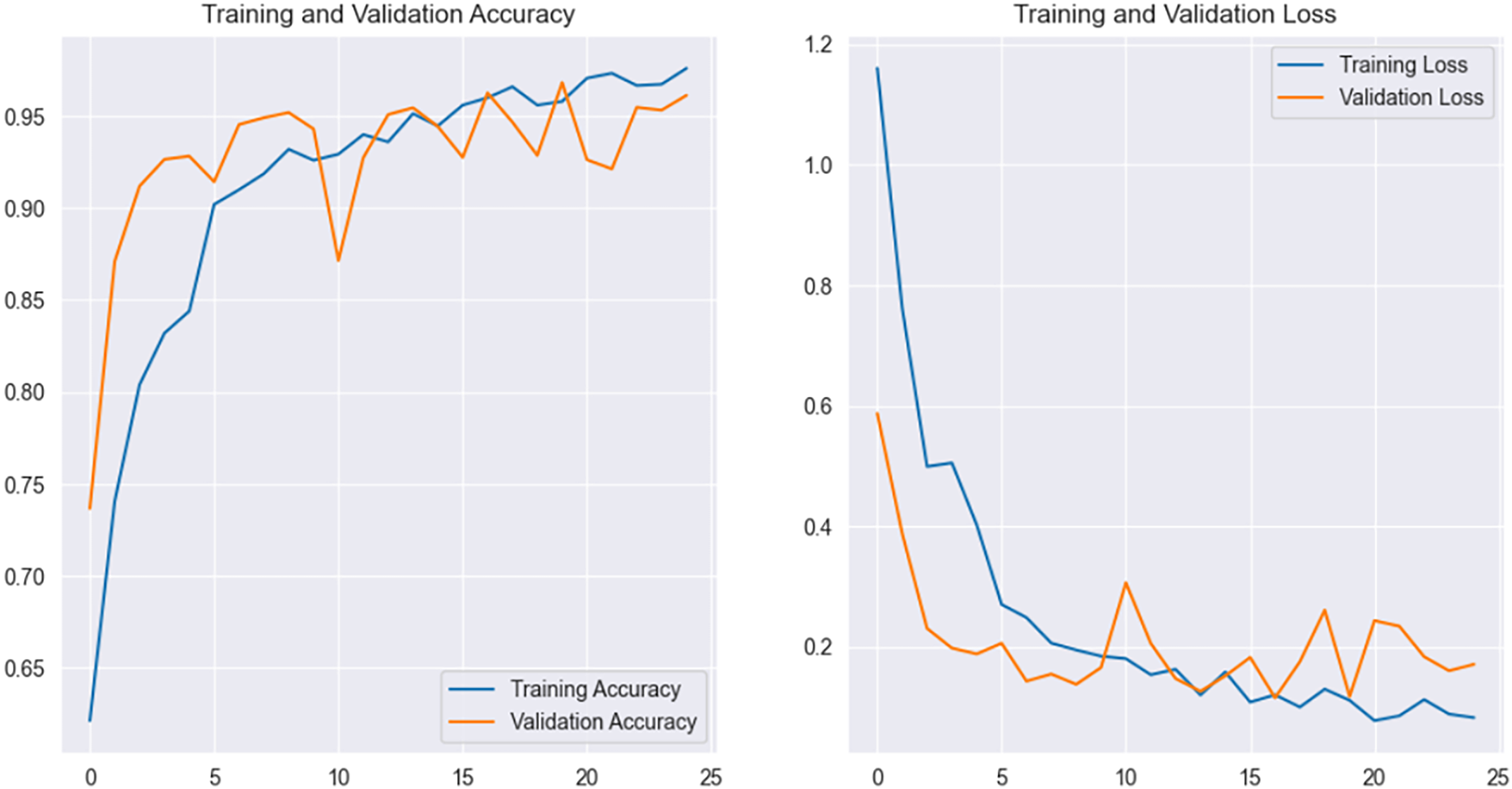
Figure 5: Accuracy and loss curve of EfficientNetB3.
The confusion matrix analysis of the EfficientNetB3 model indicates that the majority of correct predictions are concentrated along the main diagonal, consistent with the achieved overall accuracy of 0.97. Most misclassifications occur between adjacent severity levels, particularly between Healthy and Moderate cases. This behavior is expected, as early osteoarthritic changes often present subtle radiographic differences that are difficult to distinguish visually. In contrast, Severe cases are classified with high consistency, reflecting the presence of more pronounced structural abnormalities such as significant joint space narrowing and osteophyte formation. Overall, the observed misclassification patterns align with known clinical challenges in knee osteoarthritis severity assessment.
Among all evaluated architectures, EfficientNetB3 demonstrated the strongest overall performance, achieving a precision of 0.98, recall of 0.99, F1-score of 0.98, and an accuracy of 0.97. The high recall value indicates exceptional sensitivity, suggesting that the model was able to correctly identify nearly all osteoarthritic cases, which is particularly critical in clinical screening scenarios where false negatives can delay treatment. The smooth convergence observed in the training and validation curves further indicates stable optimization and effective generalization. EfficientNetV2S achieved the second-best performance with an accuracy of 0.95, reflecting the benefit of recent architectural refinements designed to improve efficiency and learning stability on medium-sized datasets.
In contrast, InceptionNet and VGG19 exhibited comparatively lower performance. InceptionNet achieved moderate precision and recall, indicating difficulty in consistently distinguishing between adjacent severity levels, particularly Moderate cases where radiographic changes are subtle. VGG19, while achieving relatively high accuracy, showed lower precision and recall, suggesting that its predictions were biased toward dominant classes and lacked sensitivity to nuanced structural variations. ResNet produced a high F1-score but a lower overall accuracy, indicating balanced class-wise predictions but inconsistency across the dataset as a whole. These observations highlight that classical CNN architectures, although effective to a degree, are less capable of capturing the fine-grained structural and textural patterns necessary for robust knee osteoarthritis severity classification. Overall, the results confirm that architectures incorporating compound scaling and optimized parameter utilization provide a clear advantage in medically relevant image analysis tasks.
Statistical Significance Analysis
To examine whether the observed performance differences among the evaluated deep learning models were statistically significant, McNemar’s test was employed. This non-parametric test is well suited for paired nominal data and is commonly used to compare the classification performance of two models evaluated on the same test set. The test assesses whether the disagreement patterns between two classifiers occur by chance.
In this study, McNemar’s test was applied by taking EfficientNetB3 as the reference model, as it achieved the highest overall performance, and comparing its predictions against those produced by InceptionNet, VGG19, ResNet, and EfficientNetV2S. The null hypothesis assumes that there is no statistically significant difference in classification performance between the two models being compared. A p-value below 0.05 indicates sufficient evidence to reject this hypothesis. The results of McNemar’s test, summarized in Table 4, show that all pairwise comparisons yielded p-values less than 0.05. This indicates that the performance improvements observed with EfficientNetB3 over the other architectures are statistically significant and not attributable to random variation. These findings provide additional validation that the superior performance of EfficientNetB3 reflects genuine differences in classification capability, reinforcing its suitability for reliable knee osteoarthritis severity assessment.

In addition to statistical significance, the observed McNemar’s test outcomes indicate a consistent and non-trivial disagreement pattern favoring EfficientNetB3 over the compared models. The low p-values reflect systematic differences in misclassification behavior rather than marginal performance fluctuations, suggesting a meaningful practical effect. In particular, EfficientNetB3 demonstrated fewer false negatives in moderate and severe cases, which is clinically relevant given the importance of sensitivity in osteoarthritis screening and severity assessment.
The results obtained in this study are consistent with and extend findings reported in recent knee osteoarthritis diagnostic research. Prior studies have shown that conventional deep learning architectures such as VGG and early Inception variants often achieve acceptable accuracy but struggle with inter-class variability, particularly when differentiating moderate disease stages from healthy or advanced cases. The performance trends observed for VGG19 and InceptionNet in this work reflect these limitations, as both models exhibited reduced sensitivity to subtle radiographic cues such as early joint space narrowing and minor osteophyte formation. The superior performance of EfficientNetB3 can be attributed primarily to its compound scaling strategy, which jointly optimizes network depth, width, and input resolution. Unlike conventional architectures such as VGG19 or InceptionNet, which scale individual dimensions independently, EfficientNetB3 maintains a balanced representational capacity that is particularly effective for capturing subtle structural and textural variations in radiographic images. In addition, its relatively efficient parameter utilization allows improved generalization on medium-sized medical datasets, reducing overfitting while preserving discriminative power. These architectural characteristics collectively explain its consistent advantage over other evaluated models under identical training conditions. EfficientNetV2S also benefited from these architectural principles, although its performance was marginally lower, suggesting that model capacity and optimization dynamics must be carefully matched to dataset characteristics. Another important observation relates to the role of data balancing and augmentation. Several studies have emphasized that class imbalance significantly affects the reliability of osteoarthritis classification models, often leading to biased predictions toward advanced disease stages. Many prior works focus on binary classification or direct prediction of Kellgren–Lawrence grades using proprietary or institution-specific datasets, limiting reproducibility. Nevertheless, the performance achieved in this study is comparable to or exceeds reported results in recent works employing similar deep learning architectures for knee osteoarthritis analysis, indicating that the proposed framework represents a meaningful advancement within a reproducible and standardized experimental setting. The consistent performance across metrics in this study indicates that the adopted balancing strategy enabled more uniform learning across severity levels. This reinforces the view that architectural advancements alone are insufficient without carefully designed preprocessing and training pipelines. Collectively, the findings demonstrate that combining modern deep learning architectures with rigorous data handling practices yields models that are both statistically robust and clinically relevant.
From a practical deployment perspective, the evaluated models are compatible with standard GPU-enabled clinical systems, and inference can be performed in near real-time once training is completed. Among the tested architectures, as mentioned above, EfficientNetB3 offers a favorable balance between accuracy and computational efficiency due to its relatively low parameter count and optimized scaling strategy. This makes it suitable for integration into clinical decision support pipelines where rapid inference and limited computational resources are important considerations. While explicit benchmarking of inference latency was not the focus of the current study, the architectural efficiency of EfficientNet-based models suggests their feasibility for routine clinical use.
From a clinical perspective, the proposed framework is intended to function as a decision support tool rather than a standalone diagnostic system. Its primary role would be to assist radiologists and clinicians by providing consistent severity assessments that complement routine visual interpretation of knee radiographs. Integration into existing clinical workflows could be achieved through picture archiving and communication systems (PACS), where automated predictions are presented alongside radiographic images. However, translation into real-world clinical practice requires careful consideration of regulatory and validation requirements. Extensive external validation on multi-center datasets, robustness testing across diverse imaging protocols, and compliance with medical device regulations would be necessary prior to deployment. In addition, real-world implementation may be influenced by factors such as variability in image acquisition quality, population heterogeneity, and the need for transparent model behavior to support clinical trust. These considerations highlight that, while the proposed approach demonstrates strong potential, further validation and standardization are required before routine clinical adoption.
Limitations
Despite the promising results, several limitations of the present study should be acknowledged. First, the proposed framework relies exclusively on plain radiographic images, which primarily capture bone-level structural changes and may not fully reflect early cartilage degeneration detectable through magnetic resonance imaging or clinical assessment. Second, the dataset was obtained from a single publicly available source, which may introduce dataset-specific biases related to imaging protocols or population characteristics. Consequently, although the models demonstrated strong performance on the test set, further validation using multi-center datasets is necessary to confirm generalizability. In addition, the study focused on image-based features alone and did not incorporate patient-specific clinical variables such as age, body mass index, or pain severity, which may provide complementary diagnostic information. These limitations define important directions for future research. Another limitation of the present study is the absence of explicit visualization-based interpretability analysis to highlight image regions influencing model predictions. While the primary focus of this work was on comparative performance evaluation and statistical robustness, incorporating visual explanations would further enhance clinical interpretability and user trust. This aspect is identified as an important direction for future research.
This study presented a comprehensive DL approach for automated classification of KO severity using plain radiographic images. By systematically evaluating five widely used pretrained CNN architectures under a unified experimental setup, the work demonstrated that model selection has a substantial impact on diagnostic reliability. Among the evaluated models, EfficientNetB3 consistently outperformed other architectures across all evaluation metrics, highlighting the effectiveness of compound-scaled networks in capturing fine-grained structural variations associated with osteoarthritis progression. The results confirm that deep learning-based systems can provide consistent and objective severity assessment, offering valuable support for clinicians in routine diagnostic workflows. While the proposed framework achieved strong performance, several avenues remain for future exploration. The current study focused exclusively on X-ray images, which primarily capture bone-level changes. Building on the findings of this study, future work will focus on extending the proposed framework by integrating multimodal data, including magnetic resonance imaging and relevant clinical variables, to improve sensitivity in early and intermediate disease stages. Multi-center validation using heterogeneous radiographic datasets will be explored to enhance generalizability and reduce dataset-specific bias. Additionally, incorporating model interpretability techniques such as activation-based visualization methods could further improve clinical trust by highlighting radiographic regions contributing to severity predictions. These extensions aim to move toward clinically deployable, transparent, and robust AI-assisted diagnostic tools for knee osteoarthritis.
Acknowledgement: Not applicable.
Funding Statement: The authors received no specific funding for this study.
Author Contributions: The authors confirm contribution to the paper as follows: conceptualization, Syed Nisar Hussain Bukhari and Sehar Altaf; methodology, Sehar Altaf and Syed Nisar Hussain Bukhari; software, Sehar Altaf and Syed Nisar Hussain Bukhari; validation, Syed Nisar Hussain Bukhari and Sehar Altaf; formal analysis, Syed Nisar Hussain Bukhari and Sehar Altaf; investigation, Syed Nisar Hussain Bukhari; resources, Syed Nisar Hussain Bukhari and Sehar Altaf; data curation, Syed Nisar Hussain Bukhari and Sehar Altaf; writing—original draft preparation, Syed Nisar Hussain Bukhari and Sehar Altaf; writing—review and editing, Syed Nisar Hussain Bukhari and Sehar Altaf; visualization, Syed Nisar Hussain Bukhari and Sehar Altaf; supervision, Syed Nisar Hussain Bukhari; project administration, Syed Nisar Hussain Bukhari. All authors reviewed and approved the final version of the manuscript.
Availability of Data and Materials: The data that support the findings of this study are openly available in Kaggle at https://www.kaggle.com/datasets/shashwatwork/knee-osteoarthritis-dataset-with-severity?resource=download&select=auto_test.
Ethics Approval: Not applicable.
Conflicts of Interest: The authors declare no conflicts of interest.
References
1. Bin Tufail A, Ma YK, Kaabar MKA, Rehman AU, Khan R, Cheikhrouhou O. Classification of initial stages of Alzheimer’s disease through pet neuroimaging modality and deep learning: quantifying the impact of image filtering approaches. Mathematics. 2021;9(23):3101. doi:10.3390/math9233101. [Google Scholar] [CrossRef]
2. Yeoh PSQ, Lai KW, Goh SL, Hasikin K, Wu X, Li P. Transfer learning-assisted 3D deep learning models for knee osteoarthritis detection: data from the osteoarthritis initiative. Front Bioeng Biotechnol. 2023;11:1164655. doi:10.3389/fbioe.2023.1164655. [Google Scholar] [PubMed] [CrossRef]
3. Joseph GB, McCulloch CE, Sohn JH, Pedoia V, Majumdar S, Link TM. AI MSK clinical applications: cartilage and osteoarthritis. Skeletal Radiol. 2022;51(2):331–43. doi:10.1007/s00256-021-03909-2. [Google Scholar] [PubMed] [CrossRef]
4. Vos T, Flaxman AD, Naghavi M, Lozano R, Michaud C, Ezzati M, et al. Years lived with disability (YLDs) for 1160 sequelae of 289 diseases and injuries 1990–2010: a systematic analysis for the Global Burden of Disease Study 2010. Lancet. 2012;380(9859):2163–96. doi:10.1016/s0140-6736(12)61729-2. [Google Scholar] [PubMed] [CrossRef]
5. Rani S, Memoria M, Choudhury T, Sar A. A comprehensive review of machine learning’s role within koa. EAI Endorsed Trans Internet Things. 2024;10:5329. doi:10.4108/eetiot.5329. [Google Scholar] [CrossRef]
6. Almansour SHS, Singh R, Alyami SMH, Sharma N, Al Reshan MS, Gupta S, et al. A convolution neural network design for knee osteoarthritis diagnosis using X-ray images. Int J Onl Eng. 2023;19(7):125–41. doi:10.3991/ijoe.v19i07.40161. [Google Scholar] [CrossRef]
7. Antony J, McGuinness K, O’Connor NE, Moran K. Quantifying radiographic knee osteoarthritis severity using deep convolutional neural networks. In: Proceedings of the 2016 23rd International Conference on Pattern Recognition (ICPR); 2016 Dec 4–8; Cancun, Mexico. doi:10.1109/icpr.2016.7899799. [Google Scholar] [CrossRef]
8. Singh R, Sharma N, Chauhan R, Rawat D, Gupta R. Knee osteoarthritis classification using EfficientNet B3 transfer learning model. In: Proceedings of the 2023 2nd International Conference on Futuristic Technologies (INCOFT); 2023 Nov 24–2; Belagavi, India. doi:10.1109/incoft60753.2023.10425390. [Google Scholar] [CrossRef]
9. Brahim A, Jennane R, Riad R, Janvier T, Khedher L, Toumi H, et al. A decision support tool for early detection of knee OsteoArthritis using X-ray imaging and machine learning: data from the OsteoArthritis Initiative. Comput Med Imag Graph. 2019;73(7):11–8. doi:10.1016/j.compmedimag.2019.01.007. [Google Scholar] [PubMed] [CrossRef]
10. Tiwari A, Poduval M, Bagaria V. Evaluation of artificial intelligence models for osteoarthritis of the knee using deep learning algorithms for orthopedic radiographs. World J Orthop. 2022;13(6):603–14. doi:10.5312/wjo.v13.i6.603. [Google Scholar] [PubMed] [CrossRef]
11. Harish H, Patrot A, Bhavan S, Gousiya S, Livitha A. Knee osteoarthritis prediction using deep learning. In: Proceedings of the 2023 International Conference on Recent Advances in Information Technology for Sustainable Development (ICRAIS); 2023 Nov 6–7; Manipal, India. p. 14–9. doi:10.1109/icrais59684.2023.10367065. [Google Scholar] [CrossRef]
12. Kinger S. Deep learning for automatic knee osteoarthritis severity grading and classification. Indian J Orthop. 2024;58(10):1458–73. doi:10.1007/s43465-024-01259-4. [Google Scholar] [PubMed] [CrossRef]
13. Tiulpin A, Thevenot J, Rahtu E, Lehenkari P, Saarakkala S. Automatic knee osteoarthritis diagnosis from plain radiographs: a deep learning-based approach. Sci Rep. 2018;8(1):1727. doi:10.1038/s41598-018-20132-7. [Google Scholar] [PubMed] [CrossRef]
14. Abimannan S, El-Alfy EM, Chang YS, Hussain S, Shukla S, Satheesh D. Ensemble multifeatured deep learning models and applications: a survey. IEEE Access. 2023;11:107194–217. doi:10.1109/access.2023.3320042. [Google Scholar] [CrossRef]
15. Tiwari S. Knee osteoarthritis dataset with severity grading. Kaggle. [cited 2025 Aug 18]. Available from: https://www.kaggle.com/datasets/shashwatwork/knee-osteoarthritis-dataset-with-severity?resource=download&select=auto_test. [Google Scholar]
16. Sabah Afroze AA, Tamilselvi R, Parisa Beham MG. Machine learning based osteoarthritis detection methods in different imaging modalities: a review. Curr Med Imaging. 2023;19(14):e300123213263. doi:10.2174/1573405619666230130143020. [Google Scholar] [PubMed] [CrossRef]
17. Schiratti JB, Dubois R, Herent P, Cahané D, Dachary J, Clozel T, et al. A deep learning method for predicting knee osteoarthritis radiographic progression from MRI. Arthritis Res Ther. 2021;23(1):262. doi:10.1186/s13075-021-02634-4. [Google Scholar] [PubMed] [CrossRef]
18. Halilaj E, Le Y, Hicks JL, Hastie TJ, Delp SL. Modeling and predicting osteoarthritis progression: data from the osteoarthritis initiative. Osteoarthr Cartil. 2018;26(12):1643–50. doi:10.1016/j.joca.2018.08.003. [Google Scholar] [PubMed] [CrossRef]
19. Abdullah SS, Rajasekaran MP. Automatic detection and classification of knee osteoarthritis using deep learning approach. Radiol Med. 2022;127:398–406. doi:10.1007/s11547-022-01476-7. [Google Scholar] [PubMed] [CrossRef]
20. Zhang W, McWilliams DF, Ingham SL, Doherty SA, Muthuri S, Muir KR, et al. Nottingham knee osteoarthritis risk prediction models. Ann Rheum Dis. 2011;70(9):1599–604. doi:10.1136/ard.2011.149807. [Google Scholar] [PubMed] [CrossRef]
21. Mahum R, Irtaza A, El-Meligy MA, Sharaf M, Tlili I, Butt S, et al. A robust framework for severity detection of knee osteoarthritis using an efficient deep learning model. Int J Patt Recogn Artif Intell. 2023;37(7):2352010. doi:10.1142/S0218001423520109. [Google Scholar] [CrossRef]
22. Hayashi D, Roemer FW, Guermazi A. Magnetic resonance imaging assessment of knee osteoarthritis: current and developing new concepts and techniques. Clin Exp Rheumatol. 2019;37(Suppl 1):88–95. [Google Scholar] [PubMed]
23. Kerkhof HJM, Bierma-Zeinstra SMA, Arden NK, Metrustry S, Castano-Betancourt M, Hart DJ, et al. Prediction model for knee osteoarthritis incidence, including clinical, genetic and biochemical risk factors. Ann Rheum Dis. 2014;73(12):2116–21. doi:10.1136/annrheumdis-2013-203620. [Google Scholar] [PubMed] [CrossRef]
24. Ganesh Kumar M, Das Goswami A. Automatic classification of the severity of knee osteoarthritis using enhanced image sharpening and CNN. Appl Sci. 2023;13(3):1658. doi:10.3390/app13031658. [Google Scholar] [CrossRef]
25. Jamshidi A, Pelletier JP, Martel-Pelletier J. Machine-learning-based patient-specific prediction models for knee osteoarthritis. Nat Rev Rheumatol. 2019;15(1):49–60. doi:10.1038/s41584-018-0130-5. [Google Scholar] [PubMed] [CrossRef]
26. Ningrum DNA, Kung WM, Tzeng IS, Yuan SP, Wu CC, Huang CY, et al. A deep learning model to predict knee osteoarthritis based on nonimage longitudinal medical record. J Multidiscip Healthc. 2021;14:2477–85. doi:10.2147/jmdh.s325179. [Google Scholar] [PubMed] [CrossRef]
27. Jahan M, Hasan MZ, Jahan Samia I, Fatema K, Rony MAH, Shamsul Arefin M, et al. KOA-CCTNet: an enhanced knee osteoarthritis grade assessment framework using modified compact convolutional transformer model. IEEE Access. 2024;12(1):107719–41. doi:10.1109/access.2024.3435572. [Google Scholar] [CrossRef]
28. Vijaya Kishore V, Kalpana V, Kumar GH. Evaluating the efficacy of deep learning models for knee osteoarthritis prediction based on Kellgren-Lawrence grading system. e-Prime—Adv Electr Eng Electron Energy. 2023;5:100266. doi:10.1016/j.prime.2023.100266. [Google Scholar] [CrossRef]
29. Castagno S, Birch M, van der Schaar M, McCaskie A. Prediction of the rapid progression of knee osteoarthritis using automated machine learning: a novel precision health approach for chronic degenerative diseases. SSRN. 2024. doi:10.2139/ssrn.4561796. [Google Scholar] [CrossRef]
Cite This Article
 Copyright © 2026 The Author(s). Published by Tech Science Press.
Copyright © 2026 The Author(s). Published by Tech Science Press.This work is licensed under a Creative Commons Attribution 4.0 International License , which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.


 Submit a Paper
Submit a Paper Propose a Special lssue
Propose a Special lssue View Full Text
View Full Text Download PDF
Download PDF Downloads
Downloads
 Citation Tools
Citation Tools