 Open Access
Open Access
REVIEW
Exploring Novel E3 Ligases and Neosubstrates for Molecular Glue Degraders and Therapeutic Applications in Cancer
Department of Anatomy, College of Medicine, Yeungnam University, Daegu, Republic of Korea
* Corresponding Author: Tae-Jin Lee. Email:
(This article belongs to the Special Issue: Advances in Cancer Therapeutics)
Oncology Research 2026, 34(6), 7 https://doi.org/10.32604/or.2026.073660
Received 23 September 2025; Accepted 30 December 2025; Issue published 21 May 2026
Abstract
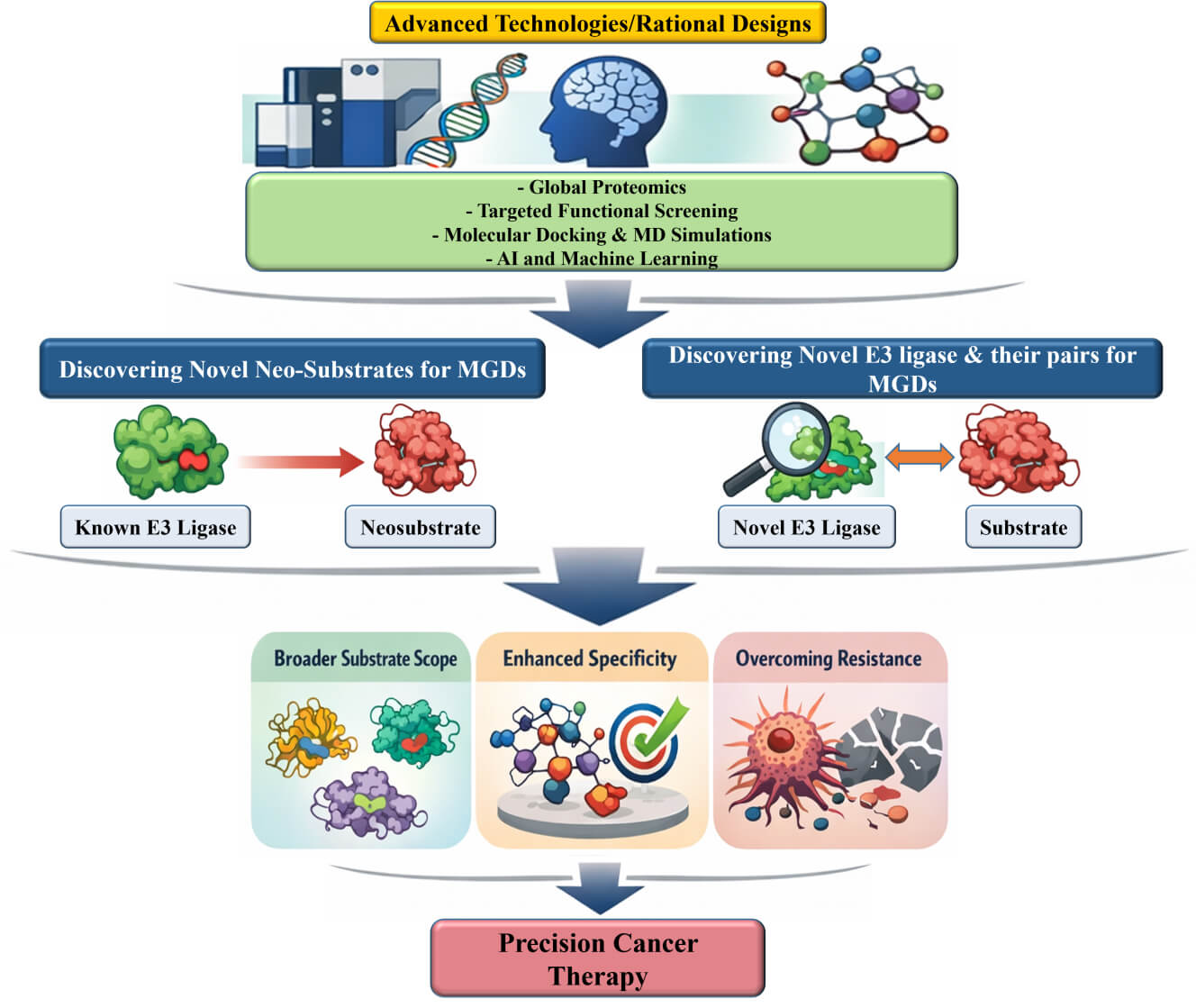
Molecular glue degraders (MGDs) are an emerging class of small molecules that promote selective protein degradation by inducing neomorphic interactions between E3 ubiquitin ligases and non-native substrates, referred to as neosubstrates. Clinically validated examples include thalidomide analogs that recruit cereblon (CRBN) to degrade IKAROS family zinc finger 1/3 in multiple myeloma, and arylsulfonamide-based MGDs that promote the degradation of RNA-binding protein 39 in acute myeloid leukemia and solid tumors. These molecules highlight the therapeutic potential of this modality in oncology. These findings underscore the promise of MGDs for eliminating oncogenic proteins previously considered undruggable and overcoming resistance to conventional inhibitors. Despite these successes, the current MGD landscape relies heavily on a limited set of E3 ligases-mainly CRBN which constrains substrate diversity, tissue selectivity, and durability of clinical response. Expanding the therapeutic utility of MGDs requires the systematic identification of novel ligases and their neosubstrates, accompanied by a deeper understanding of the mechanistic basis of ligase-substrate recognition. Recent technological advances, including chemoproteomics, ubiquitin-remnant profiling, degron mapping, clustered regularly interspaced short palindromic repeats-based functional genomics, and artificial intelligence-driven structural modeling, are advancing the discovery of new ligase-substrate pairs and enabling the rational design of degraders. Parallel progress in next-generation CRBN E3 ligase modulators, noncanonical MGDs, and structure-guided engineering further illustrates the expanding therapeutic versatility of this approach. By integrating multidisciplinary discovery strategies with translational oncology, the field is moving toward the development of next-generation MGDs with enhanced specificity, broader substrate scope, and improved resistance profiles. This study aims to elucidate how these innovations expand the degradable proteome and establish MGDs as a cornerstone of precision cancer therapy, thereby redefining the boundaries of drug discovery and providing customizable degraders tailored to diverse cancer contexts.Graphic Abstract
Keywords
Cite This Article
 Copyright © 2026 The Author(s). Published by Tech Science Press.
Copyright © 2026 The Author(s). Published by Tech Science Press.This work is licensed under a Creative Commons Attribution 4.0 International License , which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.


 Submit a Paper
Submit a Paper Propose a Special lssue
Propose a Special lssue View Full Text
View Full Text Download PDF
Download PDF Downloads
Downloads
 Citation Tools
Citation Tools