 Open Access
Open Access
REVIEW
Next Generation DNA Damage Response Inhibitors: Harnessing Nanocarriers and Tumor Microenvironment for Precision Cancer Therapy
1 Division of Oral and Craniofacial Health Sciences, University of North Carolina at Chapel Hill, Chapel Hill, NC 27599, USA
2 Department of Molecular and Cellular Engineering, Jacob Institute of Biotechnology and Bioengineering, Prayagraj, 211007, India
3 Manipal Institute of Virology, Manipal Academy of Higher Education, Manipal, 576104, India
4 School of Biotechnology, Jawaharlal Nehru University, New Delhi, 110067, India
5 Department of Biochemistry and Biophysics, University of North Carolina, Chapel Hill, NC 27599, USA
6 Integrative Program for Biological and Genome Sciences (iBGS), UNC Chapel Hill, Chapel Hill, NC 27599, USA
7 Department of Chemical and Bio-Molecular Engineering, University of Nebraska-Lincoln, Lincoln, NE 68588, USA
8 ReNeuroGen LLC, Milwaukee, WI 53122, USA
9 Department of Surgery, Division of Pediatric Surgery, Medical College of Wisconsin, Milwaukee, WI 53226, USA
10 Children’s Research Institute, Children’s Wisconsin, Milwaukee, WI 53226, USA
* Corresponding Authors: Abhishikt David Solomon. Email: ; Tarun Pant. Email:
# Both authors contributed equally
Oncology Research 2026, 34(3), 5 https://doi.org/10.32604/or.2026.071632
Received 09 August 2025; Accepted 05 January 2026; Issue published 24 February 2026
Abstract
Tumor survival, genomic stability, and therapy resistance are dictated by the DNA damage response (DDR). Although poly (ADP-ribose) polymerase (PARP) inhibitors have established the DDR as a therapeutic target, many tumors evade first-generation drugs by rewiring their adaptive repair pathways and imposing microenvironmental constraints. This review synthesizes recent discoveries in key DDR pathways, such as PARP, ataxia telangiectasia and Rad3-related kinase (ATR), ataxia telangiectasia mutated kinase (ATM), checkpoint kinase 1 (CHK1), WEE1 G2 checkpoint kinase (WEE1), and DNA-dependent protein kinase (DNA-PK), and describes the next-generation inhibitors designed to increase selectivity and circumvent resistance. We also analyze the role of hypoxia, stromal remodeling, inflammatory cytokines, and immune-cell plasticity in the tumor microenvironment in determining DDR dependency and response. Special attention is paid to cGAS-STING, immunogenic signaling via damage-associated molecular patterns (DAMPs), and mechanisms that convert a cold tumor into a hot one. Lastly, we touch upon the new nanocarrier-based delivery approaches that enhance pharmacokinetics, target resistant tumor niches, and expand the possibilities for combinatorics with immunotherapy and radiotherapy. Collectively, these findings provide a guide to the implementation of next-generation DDR inhibitors and nanomedicines to deliver a more accurate, durable, and context-specific cancer therapy.Keywords
The DNA damage response (DDR) is a highly intricate network of cellular systems responsible for detecting, signaling, and repairing the DNA damage caused by endogenous (replication errors and oxidative stress) and exogenous factors (such as chemical, radiation, or ultraviolet (UV) light exposure) [1,2]. Disruption in DDR is a characteristic feature of cancer that contributes to genomic instability, excessive cellular proliferation, and resistance to treatment [3–5]. As a result, there has been significant research interest in developing new therapies to target weaknesses in the DDR of cancer cells.
The nucleus and mitochondria are the two primary organelles that contain DNA in mammalian cells [6]. The primary mechanisms for nucleolar DNA repair systems include direct reversal (repairing alkylation DNA, base excision repair (BER) targeting non-bulky damaged DNA bases and single strand breaks (SSBs), nucleotide excision repair (NER) for extensive helix-distorting DNA damages, insertion/deletion loop (IDL) repair and mismatch repair (MMR) for base-base mismatch repair, recombinational repair, which primarily works at DNA double strand breaks (DSBs) and is further subdivided into non-homologous end joining (NHEJ) and homologous recombination (HR) repair, alternative non-homologous end joining (alt-NHEJ), which is used to repair DSBs and translesion synthesis (TLS), which is generally a damage tolerance pathway [7].
Genomic instability is largely compromised by DSBs, which are generally the most deleterious type of DNA damage, according to mounting evidence [8]. Numerous essential DNA repair mechanisms have evolved in mammalian cells to counter various types of DNA damage throughout evolution. The NER, BER, and MMR pathways, for instance, have all been thoroughly studied [9–11]. However, aberrant DNA damage repair mechanisms and procedures have often been linked to the evolution of cancer cells. For example, Ataxia-Telangiectasia Mutated (ATM) kinases are frequently mutated in between 30% and nearly 50% of cancer cell lines, including breast, prostate, pancreatic, and lymphomas [12]. Chemotherapy resistance in cancer may be associated with these mutations [13]. Moreover, genes linked to cell cycle machinery are essential for causing cancer cells to evade the effects of chemotherapy and radiation therapy [14]. The majority of cancer cell death strategies involve either inducing abnormal HR in the G1 phase, causing mitotic arrest in cancer cells, or bypassing the cell cycle checkpoint.
The rapidly growing genomic instability of cancers also presents therapeutic opportunities to target DDR pathways, which can specifically kill cancer cells by inducing exogenous DNA damage, additional replication stress, and DDR suppression [1]. This is likely the primary reason why radiation and other DNA-damaging chemotherapeutics, such as alkylating drugs and topoisomerase inhibitors, were successful in their early stages of treatment [15]. These substances, however, tend to harm healthy tissues without discrimination, which can result in serious adverse effects [16].
Over the past few years, there has been a significant surge in publications addressing the role of genomic instability and mutations in cancer formation and pathogenesis. As a result, there have been significant preclinical and clinical investigations for developing new therapies to target weaknesses in the DDR of cancer cells. In addition, an immense body of research has focused on therapeutic opportunities to target DDR pathways, which can specifically kill cancer cells by inducing exogenous DNA damage, additional replication stress, and DDR suppression [1]. For instance, the ATM inhibitor (ATMi) KU-55933 has been widely used in vitro and in vivo investigations to advance the preclinical and clinical use of ATMs [17]. Also, ATR inhibitors (ATRis) like AZ20, VE-821, and VE-822 have demonstrated high selectivity over several other DDR kinases [18]. WEE1 inhibitor (WEE1i), adavosertib (MK1775), has also shown promising potential in combination therapies in clinical use [19,20]. However, their activity is limited to several resistance mechanisms that enable cancer cells to survive and proliferate, thereby activating compensatory signaling pathways. The progress and clinical development of ATMis KU-55933, CP466722, KU-60019, and KU-59403 demonstrated minimal cytotoxicity but exhibited off-target effects and poor pharmacokinetics when used alone. They faced obstacles due to their context-dependent efficacy, which is influenced by factors such as p53 status and mode of delivery [21]. Consequently, CHK1 inhibitors (CHK1i) such as UCN-01 and DNA-PK inhibitors (DNA-PKis) like wortmannin and KU-0060648 face specificity challenges and unfavorable pharmacokinetics (PK), poor stability, and low solubility [22,23].
This review examines the recent progress in targets and inhibitors within the DDR space. We discuss the development of specific inhibitors of DDR proteins by leveraging various modes of delivery, including nanotechnology and antibody-drug conjugates (ADCs), to enhance drug delivery and effectiveness [24,25]. Additionally, we discuss the challenges of conventional DDR inhibitors and the potential use of next-generation DDR inhibitors as a therapeutic strategy to combat genomic instability, which may prevent cancer growth and pathogenesis.
2 Overview of DNA Damage Response Initiation, Repair, and Replication Mechanisms
Several studies have substantiated that the genomic instability is comprised mainly of DSBs, which are generally the most deleterious type of DNA damage. Numerous essential DNA repair mechanisms have evolved in mammalian cells to counter various types of DNA damage throughout evolution. This section explicitly discusses the current understanding of the mechanisms associated with DNA damage. Fig. 1 illustrates the mechanisms of DNA damage from the preclinical and clinical studies.
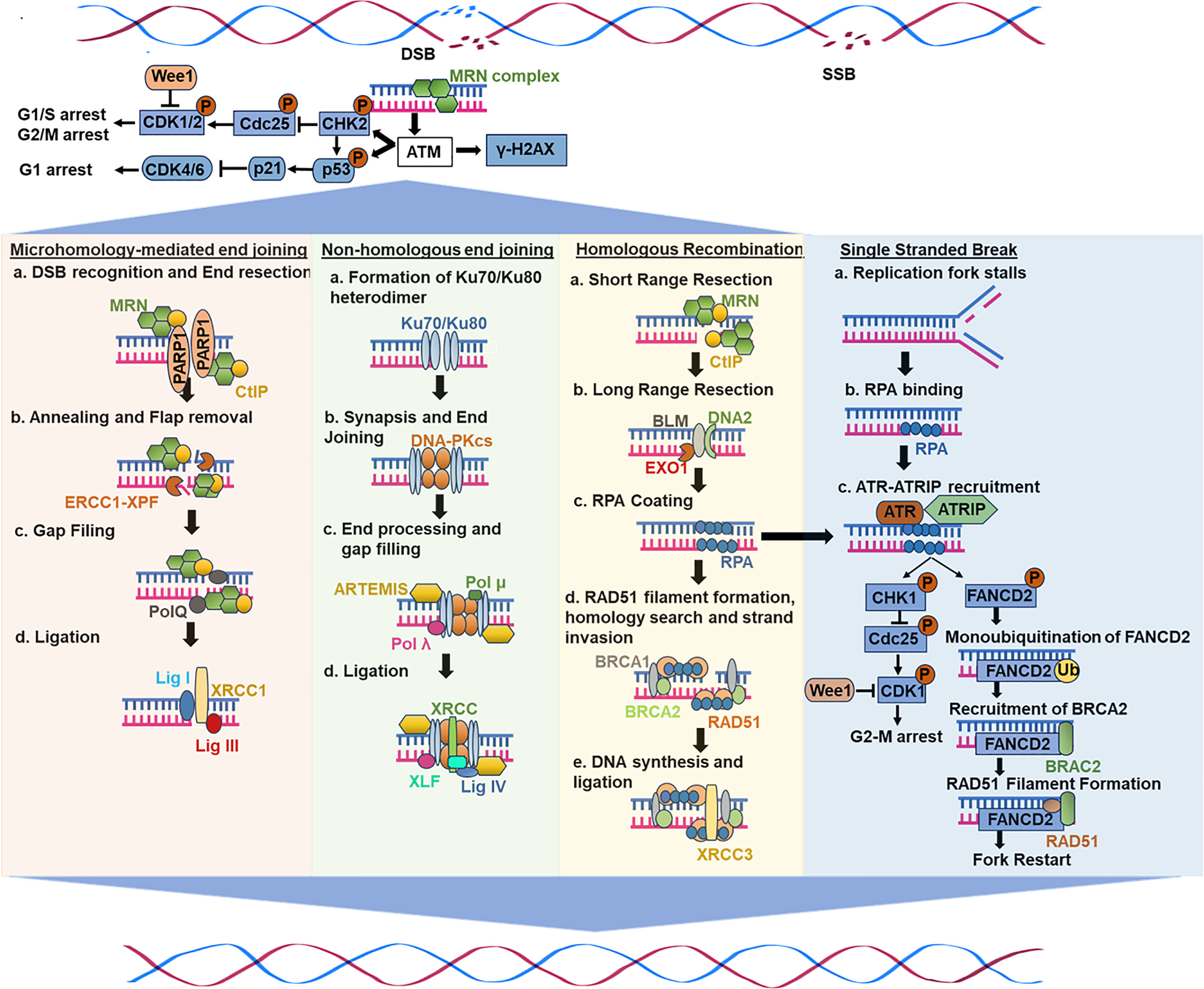
Figure 1: A general scheme of DNA damage response and repair mechanisms
Ataxia telangiectasia (A-T) is a rare inherited autosomal recessive condition characterized by immune system deficiency, hypersensitivity to radiation, increased cancer susceptibility, and progressive neurodegeneration, leading to impaired movement coordination (ataxia) [26–28]. A-T arises from ATM gene mutations that disrupt ATM kinase function, a pivotal regulator of the DDR [29]. Structurally akin to phosphoinositide 3-kinase (PI3K), ATM belongs to the phosphatidylinositol 3-kinase-related kinase (PIKK) family [28,30,31].
As a serine/threonine kinase, ATM acts as a master coordinator in DDR, governing DBS repair through HR, NHEJ, and cell cycle checkpoints [32,33]. Upon DNA damage, ATM undergoes self-activation (autophosphorylation) at serine 1981, relocating to damaged sites [34]. Thereafter, it phosphorylates crucial targets like checkpoint kinases (e.g., CHK2), the histone variant H2AX, and the mediator of DNA damage checkpoint 1 (MDC1) [34–36], triggering cell cycle arrest, activation of DNA repair pathways, or programmed cell death (apoptosis) [37]. ATM-mediated regulation of the G1/S, S, and G2/M checkpoints involves pathways such as the ATM-p53-p21, ATM-Chk2-CDc25, and ATM-BRCA1-Cyclin B1 pathways [38,39] (Fig. 1).
ATM influences both NHEJ and HR, but studies suggest a more prominent role in NHEJ, where it phosphorylates the DNA-dependent protein kinase catalytic subunit (DNA-PKcs), thereby impacting the accuracy and efficiency of DSB repair [40–42]. ATM also interacts intricately with A-T and Rad3-related (ATR), which predominantly responds to replication stress and SSBs [43,44]. Since ATR inhibition prompts ATM compensation, a dual-targeting approach involving ATM and ATR inhibitors (ATMis and ATRis) may offer a more promising therapeutic strategy for cancer treatment [36,45]. The WEE1 inhibitor (WEE1i) adavosertib (MK1775) has also shown promising potential in clinical use [19,20].
Early ATMs faced significant challenges, including feeble blood-brain barrier (BBB) penetration, which limited their use in brain tumors, such as glioblastomas (GBMs) [46]. Many compounds also lacked specificity, leading to off-target toxicity and a narrow therapeutic window. KU60019 exhibited potential to radiosensitize glioma-initiating cells (GICs), which provided the first evidence supporting ATM inhibition as a cancer treatment [47,48]. ATM’s restricted success highlighted its therapeutic potential, prompting the development of brain-penetrant inhibitors with improved sensitivity and safety profiles. The insights from these lessons have shaped the development of next-generation ATMs, which exhibit improved pharmacokinetics and demonstrate clinical potential.
2.2 ATR-CHK1 Axis in DNA Damage Response and Replication Repair
ATR, a key DDR kinase, responds to replication stress and SSBs by activating checkpoint kinase 1 (CHK1), leading to cell cycle arrest and DNA repair [49]. ATR inactivation increases replication stress, leading to DSB formation and cancer cell death [36]. ATRis are explored as monotherapies or in combination with immune checkpoint inhibitors (ICI), Poly(ADP-ribose) polymerase (PARP) inhibitors (PARPis), and DNA-damaging agents (DDAs) [50,51]. By disrupting the ATR/CHK1 pathway, ATRis promote genomic instability and mitotic catastrophe [52]. ATR inhibition also sensitizes cancer cells to replication stress-inducing agents by preventing ssDNA from causing cell cycle arrest [53].
CHK2, a serine/threonine kinase, regulates the cell cycle and DDR, particularly in response to replication stress caused by chemotherapy or repair deficiencies [54,55]. CHK1 inhibition sensitizes cancer cells to genotoxic treatments, particularly in those with G1-S checkpoint defects that depend on intra-S and G2/M checkpoints for survival [56]. Oncogene amplification on extrachromosomal DNA (ecDNA) is associated with a poor prognosis and resistance to therapy. These circular DNA elements exhibit high genomic plasticity but also induce elevated replication stress, making ecDNA-amplified tumors highly dependent on CHK1 [54]. Consequently, the application of CHK1 inhibitors is a promising strategy against such malignancies [57].
2.3 Non-Homologous End Joining (NHEJ)
NHEJ involves several mechanistically distinct stages in the repair of DSBs [28]. Upon DSB formation, the Ku70/80 heterodimer binds DNA ends, initiating the assembly of the NHEJ machinery [58,59]. Ku serves as a recruitment hub for downstream NHEJ components [28] (Fig. 1). Furthermore, DNA-PKcs, a major protein kinase, forms the DNA-PK holoenzyme upon binding to DNA-bound Ku [60–62]. DNA binding activates this process, which results in the phosphorylation of NHEJ components [63]. DNA-PKcs autophosphorylation represents an essential phase in NHEJ-mediated DSB repair processes, while its functional consequences await clarification. Moreover, for ligation to occur, DNA ends in the synaptic complex must be closely aligned, facilitated by NHEJ factors like X-ray repair cross-complementing protein 4 (XRCC4), XRCC4-like factor (XLF), and PAXX [64–66]. Finally, DNA ligase IV catalyzes ligation, forming a complex with XRCC4, and can tolerate terminal mismatches and damaged bases [67]. However, many DNA end structures cannot be directly ligated. As a result, many end-processing factors, such as nucleases and polymerases, are attracted to DSBs and work on the ends to prepare them for ligation. Deficits in this pathway have several detrimental effects on human health, as NHEJ is essential for repairing both spontaneous and planned DSBs [28]. Defective V(D)J recombination causes severe combined immunodeficiency (SCID) when mutations occur in the genes encoding XLF, LIG4, DNA-PKcs, and Artemis [28]. Human cells with abnormalities in essential NHEJ components are typically hypersensitive to infrared radiation, and a fraction of SCID patients also show radiosensitivity [28].
2.4 Homologous Recombination (HR)
Homologous recombination (HR) ensures genome stability by enabling error-free repair of DSBs, interstrand crosslinks (ICLs), and DNA gaps before and after DNA replication [68]. HR, which is mainly active in the S and G2 phases [69], heavily relies on homologous strands, typically the sister chromatid, as repair templates in somatic cells [70]. The process involves two key steps: DNA strand invasion and the homology search, which ensures precise repair. HR initiation requires extensive 5′ to 3′ end resection, generating 3′-OH single-stranded DNA (ssDNA) tails, facilitated by the MRN (MRE11-RAD50-NBS1) complex that also triggers DDR [71] (Fig. 1). C-terminal-binding protein-interacting protein (CtIP), an interacting partner of MRN, leads to its activation, shifting its role from DNA damage sensing to resection [72,73]. Exonuclease 1 (EXO1), endonuclease DNA2, and Bloom syndrome helicase further extend long-range resection [74–76]. Following resection, replication protein A (RPA) coats ssDNA, preventing secondary structures and nuclease degradation. RPA also inhibits RAD51 filament formation and nucleation, thereby preventing its interaction with ssDNA ends [77]. Thereafter, several mediators, including PALB2, RAD51 paralogs, and BRCA2, evict RPA through interactions with BRCA1 and BRCA1-associated RING domain protein 1 (BARD1), promoting RAD51 nucleofilament formation [77–79]. RAD51 recombinases search for a homologous sequence to facilitate strand invasion, forming a D-loop (displacement loop) intermediate. This structure stabilizes invading strands, allowing DNA polymerases-mediated synthesis using the homologous template. Following strand extension, the D-loop is resolved through one of the two pathways: synthesis-dependent strand annealing (SDSA), resulting in non-crossover-repair, or the double Holliday junction (dHJ) pathway that would lead to crossover or non-crossover outcomes [80,81]. Overall, these mechanisms collectively ensure high-fidelity DNA repair and error-free bypass, preventing mutations and maintaining genome stability [82].
3 Recent Advances and Challenges in Targeting DNA Damage Response Using Small Molecule Inhibitors
The above-described studies indicate that genomic instability is mainly comprised of DSBs, which are the most deleterious type of DNA damage. Conventional cancer therapies like radiation and chemotherapy exert their cytotoxic effects by directly damaging DNA [83,84]. However, cancer cells frequently harbor robust DNA repair mechanisms enabling them to survive and proliferate despite such damage [85,86]. This inherent resistance has fueled the development of small-molecule inhibitors (SMIs) targeting key DDR proteins [87]. Therefore, DNA damage inhibitors have been intensively investigated using animal models and cell culture-based studies (Fig. 2). Several classes of DDRis are currently under investigation, each with distinct mechanisms of action and therapeutic applications. The current section will examine preclinical and clinical drug models targeting DDR signaling using distinct DNA damage inhibitors like ATM, ATR, CHK1/2, Wee, and DNA-PK. Different preclinical and clinical studies investigating DDRi are summarized in (Table 1).
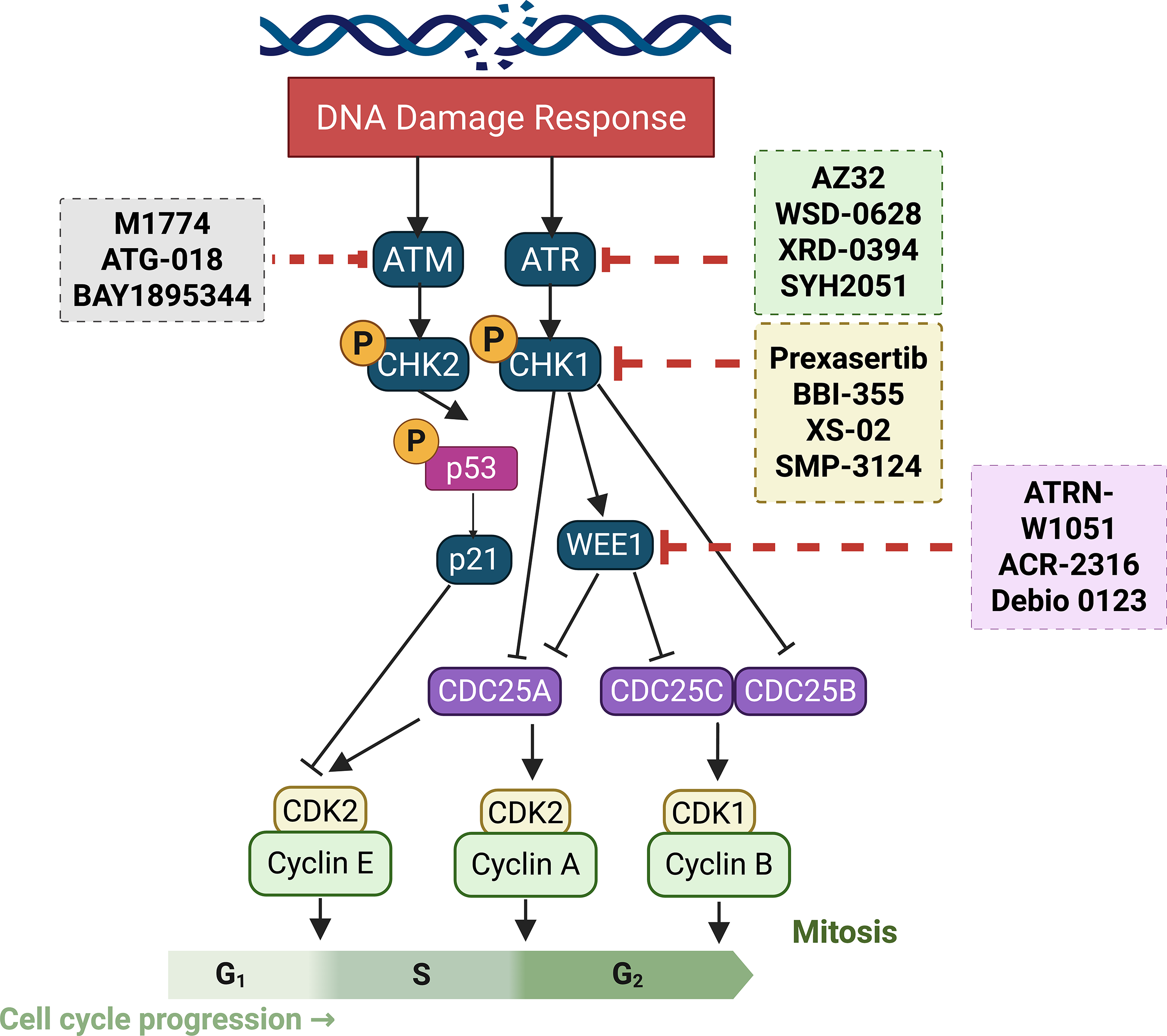
Figure 2: Illustration of current clinical and pre-clinical drug models targeting DNA damage response signaling

Several small-molecule ATMis are in development or undergoing clinical trials, exhibiting varying degrees of potency and selectivity [33,88]. Many feature bioisosteric variants or imidazoquinolinone cores to enhance efficacy and reduce off-target effects [89]. Combination strategies are being explored, including WSD-0628 with radiation for glioblastoma (GBM) and melanoma brain metastases [90] and ZN-B-2262 with topoisomerase inhibitor-based ADCs [91]. Since ATM repairs topoisomerase-induced DNA breaks, its inhibition may enhance cancer cell death. Additionally, AZD1390, a novel ATM kinase inhibitor, simultaneously targets HR and microhomology-mediated end joining (MMEJ), increasing TMZ cytotoxicity and radiosensitivity in resistant GBMs [92]. Further validation is needed to optimize ATM inhibition and overcome treatment resistance. Therefore, there is a need to validate several aspects of ATM inhibition, as well as multiple targets that may completely abrogate cancer cell proliferation and treatment resistance.
WSD-0628, designed by Wayshine Biopharmaceuticals and evaluated by the Mayo Clinic, is an effective ATMi reported to date, effectively inhibiting ATM activity in U251 glioblastoma cells at a concentration of 30 nM. In combination with IR, it showed highly significant survival rates (%) in GBM xenograft mice and demonstrated strong brain penetration [90,93]. Investigations by Rathi et al. reported a potent inhibition of ATM autophosphorylation and downstream targets CHK2 and KAP1 at 100 nM and reduced γH2AX foci in GBM43 cells. Additionally, preclinical studies using intracranial p53-mutant GBM43 xenograft models demonstrated rapid CNS distribution and prolonged retention at a dose of 10 mg/kg, compared to lower doses [94]. Oral dosing revealed non-linear pharmacokinetics, and only minor increases in brain exposure were observed in efflux transporter KO mice, suggesting limited BBB resistance [94]. It is potent, and its CNS accessibility supports its ongoing clinical evaluation as a powerful radiosensitizer for recurrent, advanced-grade gliomas (NCT05917145).
AZ32 is an orally bioavailable ATM inhibitor that blocks DDR and radiosensitizes gliomas. A study by Karlin et al. demonstrated the efficacy of AZ32 as a potential penetrant of the BBB [46]. It enhanced radiosensitivity in intracranial gliomas, outperforming AZ31 in syngeneic orthotopic glioma models. Its efficacy was particularly notable in glioma cell lines harboring mutant p53 or checkpoint defects, where combining AZ32 with low-dose radiation induced over a sixfold increase in tumor apoptosis compared to healthy brain tissue [46]. Furthermore, McCabe et al. revealed that ATM inhibition is synthetically lethal in PTEN-deficient tumors, which exhibit chronic ATM activation, elevated DNA damage, and heightened reactive oxygen species (ROS) levels. ATM blockade in these cells triggered apoptosis, irreversible DNA damage, and cell cycle arrest, whereas wild-type cells remained less affected [95]. These findings highlight AZ32 inhibitor as a promising therapeutic agent, particularly in tumors with p53 mutations or PTEN loss, warranting further clinical evaluation.
XRD-0394 functions as a sophisticated and potent dual inhibitor targeting ATM and DNA-PKcs, demonstrating increased tumor cell death when combined with IR, topoisomerase inhibitors, and PARPis, especially in BRCA1/2-deficient cells. This compound demonstrates independent-agent performance while exhibiting advantageous pharmacological properties. Gilmer et al. reported the completed Phase 1a trial and the identification and preliminary characterization of XRD-0394 [96]. The clinical potential of NCT05002140 emerges in combination therapies, warranting further investigation. XRD-0394 emerged as a solution to RT limitations by dual-inhibition of ATM and DNA-PK, thereby enhancing tumor sensitivity while countering resistance pathways. Preclinical data demonstrated that transient dual inhibition enhances tumor cell death while sparing non-cancerous cells in the absence of radiation. In a phase 1a trial, XRD-0394 was well tolerated with no dose-limiting toxicities when combined with palliative RT. At 160 mg, plasma levels exceeded preclinical efficacy thresholds for over 15 h, with confirmed ATM inhibition in tumor tissues [96]. Its pharmacokinetic profile supports co-administration with RT, providing a strong rationale for future trials combining XRD-0394 with RT, PARPis, immunotherapy, or topoisomerase-targeting ADCs.
Clinical investigations of the novel ATMi SYH2051 are underway. ZhongQi Pharmaceutical Technology Co. developed SYH2051 [21]. Following the successful completion of pre-clinical assessment, SYH2051 has entered clinical trials both as monotherapy and in combination therapy with IR [97]. The ongoing trial, NCT06011291, aims to characterize the pharmacokinetic profile and assess therapeutic efficacy in solid tumors, particularly in head and neck cancers. With the ability to enhance the effectiveness of current DNA-damaging treatments and circumvent resistance mechanisms, ATMis constitute an exciting intervention in cancer therapies. The role of ATMs in the treatment of cancer and other illnesses will likely be further defined by ongoing research and clinical trials, despite the obstacles that remain. Personalized cancer treatment and better patient outcomes are made possible by the ability to target the DDR precisely.
The ATR/CHK1 pathway is crucial for DDR and disrupted by ATRis, leading to the accumulation of DSBs, particularly in tumor cells with high replication [52,98]. This disruption induces mitotic catastrophe, genomic instability, and cell death. ATRis also prevent ssDNA from triggering cell cycle arrest, thereby enhancing cancer cell sensitivity to replication-stress-inducing agents. Several ATRis are currently in clinical trials, both as monotherapies and in combination with chemotherapeutics, aiming to identify predictive biomarkers for personalized treatment and evaluate their therapeutic efficacy [49,53].
M1774, an oral ATR inhibitor in clinical development, exhibits superior efficacy over ceralasertib and berzosertib in small cell lung cancer (SCLC), inhibiting cancer cell survival at nanomolar doses [52]. By blocking ATR/CK1 checkpoints, M1774 enhances TOP1 inhibitor-mediated tumor cell death by avoiding replication arrest and inducing DNA damage. Proteomic analysis revealed M1774, and SN-38 upregulate G2/M-related and replication proteins (PLK1 and CCNB1, TIPIN, CDC45, TIMELESS, and RPA1). It also synergizes with TOP1/2 inhibitors, cisplatin, and talazoparib, overcoming chemoresistance in SLFN11-deficient cells, positioning SLFN11 as a potential biomarker, and demonstrating therapeutic synergy with DNA-damaging agents in SCLC and colon cancer organoids, as well as in H82 SCLC xenografts. While promising, further evaluation of dosing and combination strategies is essential [52].
The antitumor effects of ATG-018 have also been demonstrated in vitro and in vivo [98]. In a study by Hui et al., normal peripheral blood mononuclear cells viability was unaffected by ATG-018; however, in 137 out of 143 cell lines, including solid tumors and hematologic malignancies, its IC50 ranged from 0.22 to 10 µM [98]. Numerous genetic alterations that can be used as predictive markers have been linked to ATG-018 sensitivity. Inhibiting downstream CHK1 phosphorylation in HT-29 cells required an IC50 of 1.4 nM, whereas ATR kinase activity required 16 nM [98]. Additionally, in CDX mouse models of OCI-LY-19 (lymphoma), OE21 (esophageal cancer), and LoVo (colorectal cancer), ATG-018 demonstrated dose-dependent anticancer effects [99].
BAY1895344 is another ATRi that has shown promise in preclinical and clinical studies. It was developed by optimizing a quinoline-based scaffold with initial weak ATR inhibition [100]. Chemical refinements enhanced its potency, selectivity, and oral bioavailability, establishing it as a strong clinical candidate. Preclinical studies also showed its efficacy both as monotherapy and in combination with DDAs, particularly in tumors with DNA repair deficiencies [100]. When combined with topoisomerase inhibitors (topotecan, irinotecan), it has been shown to exacerbate DNA damage and impair DNA repair, leading to enhanced tumor cell death [101]. Also, pairing with emcitabine or cisplatin showed greater cytotoxicity by blocking DNA repair of lesions induced by these agents [102]. Moreover, with PARPi talazoparib, it has been shown to disrupt DNA repair in HR-deficient tumors [103,104].
Another investigation by Tang et al. demonstrated that BAY1895344 represses ATR-CHK1 signaling, activates the Cyclin-dependent kinase-Speckle-type POZ (CDK1-SPOP) axis, and destabilizes programmed death-ligand 1 (PD-L1) protein in prostate cancer cells [105]. BAY1895344, combined with anti-PD-L1 therapy, robustly activated innate immunity while producing a synergistic T-cell-dependent therapeutic response in syngeneic mouse models [105]. The study outcomes justify the integration of ATR-targeted therapies with ICIs for treating castration-resistant prostate cancer patients. BAY1895344 is currently undergoing clinical evaluation in patients with advanced solid tumors and lymphomas (NCT03188965). This phase 1/2 trial is assessing safety, PK, and efficacy, both alone and in combination. Biomarker analysis within trials aims to refine patient selection and optimize therapeutic outcomes [106].
CHK1 inhibitors (CHK1i) function by blocking cell cycle checkpoints, specifically the intra-S phase and G2/M checkpoints [107]. By inhibiting CHK1, these inhibitors allow cells to bypass the checkpoint and proceed through the cell cycle despite damaged DNA, thereby preventing arrest in response to replication stress or DNA damage [108]. In the end, this results in cell death, genomic instability, and mitotic catastrophe [102]. CHK1 makes tumor cells more vulnerable to radiation and chemotherapy, among other DNA-damaging substances [109]. This is because cancer cells are more dependent on CHK1 for survival, as they frequently have defects in multiple DNA repair pathways [110]. Specific genetically altered cancer cells may be synthetically deadly when CHK1 is inhibited. Cancer cells that harbor ATM mutations, for instance, would be especially vulnerable to CHK1 inhibition [111]. Numerous clinical trials have assessed CHK1 as a standalone treatment and in conjunction with other cancer treatments [112,113]. Although the preclinical activity of specific CHK1 inhibitors has been encouraging, their clinical development has proven difficult [114].
Prexasertib, a dual CHK1/CHK2 inhibitor, has shown clinical efficacy in platinum-resistant, BRCA wild-type high-grade serous ovarian carcinoma [115]. However, its intravenous delivery poses toxicity and compliance challenges [116]. Resistance mechanisms include IGF-1/insulin pathway activation, immunosuppressive responses, a prolonged G2 delay via reduced CDK1/Cyclin B1 activity, and shielding cells from mitotic catastrophe [115]. Despite resistance, CHK1’s role in RAD51-mediated HR remains unchanged, allowing prexasertib to sensitize resistant cells to DNA-damaging agents like gemcitabine and hydroxyurea (HU). In triple-negative breast cancer (TNBC), epidermal growth factor receptor (EGFR) signaling contributes to resistance by phosphorylating BAD, inhibiting apoptosis. EGFR blockade restores BAD activity, enhancing prexasertib sensitivity [117].
BBI-355 exploits the vulnerability of cancers harboring ecDNA that encodes oncogenes such as MYC, EGFR, and fibroblast growth factor receptor 2 (FGFR2) [118,119]. ecDNA amplifications drive RS and resistance to targeted therapies through rapid, non-Mendelian oncogene distribution [54,57]. By inhibiting CHK1, BBI-355 disrupts the RS response and induces tumor cell death [120]. It has shown strong preclinical efficacy in vitro and in vivo. It selectively inhibits ecDNA-positive tumor cells and induces RS [54,118]. Oral administration in mice demonstrated dose-dependent target engagement, as evidenced by increased pCHK1-S345 levels, and robust tumor regression in ecDNA-positive models [118]. Combination therapy with FGFR or CDK4/6 inhibitors further enhances tumor suppression [118,119].
Furthermore, pCHK1-S345 has emerged as a reliable pharmacodynamic biomarker for CHK1 inhibition by BBI-355 [57]. It has been validated in vitro, in xenografts, and in patient skin biopsies from the ongoing POTENTIATE trial, confirming clinical target engagement and helping define optimal dosing. The phase 1/2 POTENTIATE trial evaluates BBI-355 alone or in combination with erlotinib (EGFR inhibitor) and futibatinib (FGFR inhibitor) in patients with solid tumors harboring oncogene amplifications. Initial results show good tolerability [57,118,119]. Initial results show good tolerability, manageable hematologic adverse events, dose-dependent exposure, and on-target pCHK1 modulation [119]. Since ecDNA-mediated resistance to targeted therapies poses a significant clinical challenge, BBI-355 prevents such resistance when combined with EGFR or FGFR inhibitors in preclinical gastric cancer models, yielding durable responses [118,119].
XS-02 is a new, orally bioavailable inhibitor of CHK1 with modest activity against CHK2. It exhibits strong anticancer activity across a variety of solid cancers, including those resistant to PARPis, with manageable side effects [116]. By inhibiting both CHK1 and CHK2 kinases in vitro, XS-02 reduces the phosphorylation of CHK1 entirely and CHK2 partly in OVCAR3 cancer cells. Dose-dependent suppression of tumor development is induced by oral administration of XS-02 alone [116]. In an animal model of acquired resistance to olaparib developed from patient-derived tumor xenografts, the synergy between XS-02 and olaparib, a PARPi, has been demonstrated to expedite tumor regression [116].
SMP-3124 is an innovative, liposome-encapsulated, and specific CHK1i [55]. Drug distribution is altered by this liposomal formulation, which results in extended drug retention in plasma and accumulation in tumors [55]. SMP-3124 demonstrates a strong anticancer property in several subcutaneously xenografted animals without any adverse effects [55]. In orthotopic and peritoneal dissemination models utilizing human ovarian cancer cells, it also increases overall survival [55]. Gemcitabine and SMP-3124 work together to synergistically reduce tumor development without causing any extra hematological effects [55].
Although several CHK1 inhibitors are currently under development, issues with toxicity, patient stratification, and resistance mechanisms remain. For CHK1i to realize its most significant therapeutic potential, future studies should focus on identifying predictive biomarkers, developing innovative combination strategies, and improving drug-delivery techniques. BBI-355, a new ecDNA-directed treatment, shows promise for targeting CHK1 in malignancies driven by extrachromosomal DNA oncogene amplification [118]. CHK1i’s safety and effectiveness in treating different types of cancer will be further clarified by ongoing clinical trials, such as the POTENTIATE study by Hansen et al. [118].
WEE1 regulates the G2/M transition by phosphorylating CDK1 at tyrosine 15 (Y15), preventing cell division in cells with damaged DNA [121]. When DNA damage occurs, WEE1 is activated to maintain the G2/M checkpoint, and its inhibition increases CDK1 activity, causing cells to bypass DNA repair, enter mitosis prematurely, and undergo mitotic catastrophe and cell death [122]. This makes WEE1 a key target in cancer treatment, especially in p53-deficient cancers, which are highly reliant on the G2/M checkpoint [123]. WEE1 inhibition induces apoptosis and mitotic catastrophe in these cells [124].
However; WEE1 inhibitors have previously been associated with severe hematological consequences; such as anemia; neutropenia; and thrombocytopenia; limiting their effectiveness [125]. Current strategies focus on developing more potent and selective inhibitors with improved pharmacokinetics to reduce off-target effects and extend the treatment window; thereby improving patient outcomes [122].
ATRN-W1051 is a selective WEE1i developed to reduce toxicity [125]. According to reports by Vacca et al., it has shown potential to inhibit WEE1 with an IC50 of 2.2 nM and suppress ovarian cancer cells at 100–200 nM concentrations [125]. Unlike AZD1775, it does not inhibit PLK-1-3, minimizing off-target effects. ATRN-W1051 also offers favorable pharmacokinetics, requiring 3–8 times lower doses than AZD1775 to achieve similar exposure in ZN-c3. It effectively reduces tumor growth in Cyclin E1-amplified high-grade serous ovarian cancer (CCNE1-HGSOC) xenograft models. It is well tolerated with daily oral dosing, highlighting its potential as a target therapeutic for CCNE1-overexpressing HGSOC.
ACR-2316 is a dual WEE1 and PKMYT1 inhibitor optimized via co-crystallography for improved single-agent activity and selectivity. In a study by Wigerup et al., ACR-2316 demonstrated potent inhibition of WEE1 at an IC50 of 2 nM and IC90 of 10 nM, following the moderate inhibition of PKYMT at IC20 of 44 nM, when compared to adavosertib and lunresertib, which resulted in superior activation of mitotic kinases CK1, CDK2, and PLK2 [126]. Selectivity profiling through AP3 (200+ kinases) and KINOMEscan (468 kinases) confirmed its improved specificity over existing WEE1/PKYMT1 inhibitors. Also, cell cycle analysis revealed greater S-G2/M accumulation than with adavosertib or lunresertib. Moreover, ACR-2316 showed the highest potential in a 19-cell line proliferation screening and 12 ovarian cancer patient-derived xenograft (PDX) models ex vivo. Consequently, orally administered ACR-2316 was well tolerated and produced robust, dose-dependent tumor growth inhibition in xenograft models [126]. These findings support its advancement toward clinical development as a monotherapy drug.
Debio 0123, a selective WEE1 Inhibitor, has demonstrated promising antitumor efficacy in preclinical GBM models and exhibits efficient brain penetration. According to Bellon et al., Debio 0123 reduced the IC50 of TMZ and enhanced radiation-induced cell death in primary GBM cell lines in vitro. In vivo studies showed favorable brain distribution of Debio 0123 in mice, rats, and monkeys, with brain-to-plasma ratios of 0.6, 1.52, and 4, respectively. In orthotopic tumor models, the tumor-to-plasma ratio was found to be 0.62. Also, in GBM xenografts (U87-MG), Debio 0123 suppressed tumor growth by up to 73.7% (brain) and 57.5% (subcutaneous). When combined with TMZ, it induced sustained complete regressions in 75% of mice for up to 100 days, with good tolerability over a 28-day treatment. Additionally, the oral bioavailability of Debio 0123 is pH-dependent, suggesting it should not be taken with proton pump inhibitors (PPIs) [127]. However, food intake had minimal impact on absorption, suggesting that administration with meals would improve patient compliance.
Inhibiting DNA-PK kinase activity is being investigated as a cancer treatment approach, particularly in conjunction with drugs that induce DNA-DSB140 [41,128]. DNA-PK inhibition is thought to disrupt NHEJ, thereby sensitizing cancer cells to radiation- or chemotherapy-induced DNA damage. There have been numerous SMIs targeting DNA-PK, some of which are currently undergoing clinical trials.
3.5.1 Targeting Ku70/80 Heterodimer for DNA-Pk Inhibition
Targeting KU70/80 heterodimer-DNA interaction is a novel strategy for DNA-Pk suppression, as Ku70/80 is essential for DNA-Pk activation by binding to DNA termini and recruiting DNA-Pk to damage sites [128,129]. Ku-DNA binding inhibitors (Ku-DBis) are designed to prevent this initial step, thereby blocking DNA-Pk catalytic activity, and disrupting the NHEJ pathway and increasing cancer cell sensitivity to DSB-inducing treatments [128,130].
Preclinical studies by Mendoza-Munoz et al. show that Ku-DBis effectively inhibits DNA-Pk and NHEJ with nanomolar activity, sensitizing non-small cell lung cancer (NSCLC) cells to ionizing radiation (IR), bleomycin, and etoposide [128,129]. Ku-DBis enhances DDR suppression by reducing DNA-Pkcs autophosphorylation, activating ATM-p53 signaling, leading to p53 phosphorylation [131]. To improve efficacy, researchers have optimized the structure-activity relationships (SARs) of Ku-DBis by developing oxindole derivatives within the X80 core that enhance cellular uptake, selectivity, and Ku inhibition. These modifications improve inhibition of DNA-DSB repair, reduce nonspecific protein binding, and enable serum-based cellular studies. In vivo studies in NSCLC xenograft models also confirmed the therapeutic potential of Ku-DBis, which inhibited IR-induced autophosphorylation of DNA-Pkcs and suppressed tumor cell proliferation and DDR signaling. These advancements mark a significant step toward the clinical application of Ku-DBis for cancer therapies.
SY-7021 is a highly potent and selective DNA-Pki that shows approximately 400-fold selectivity over ATM and ATR. According to investigations by Zhang et al., SY-7021 dose-dependently suppressed cellular NHEJ efficiency in a reporter assay and significantly inhibited cancer cell proliferation both alone and in combination with doxorubicin (DOX) [132]. The SY-7021/DOX combination enhanced phosphorylation of H2AX (Ser139), CHK2 (Tyr68) and p53 (Ser15) and induced G2/M arrest and cell death in MDA-MB-468 cells. In vivo, oral administration of SY-7021 twice daily led to dose-dependent tumor growth inhibition in NCI-H1703 xenografts, achieving 105.6% tumor growth inhibition (TGI) at 60 mg/kg without noticeable weight loss. SY-7021 also demonstrated favorable pharmacokinetics and safety profiles [132]. These results support its potential as a promising monotherapy or combinatorial treatment option across multiple cancer types.
Peposertib (M3814) is a potent, specific, orally available DNA-Pki that enhances the efficacy of DNA-damaging agents like topoisomerase II inhibitors and IR [133,134]. It effectively suppresses DNA-Pk catalytic activity, increasing cancer cell sensitivity to DSB-inducing treatments [134]. In preclinical studies, M3814, when combined with IR, induced complete tumor regression in xenograft models without toxicity, highlighting its potential in cancer RT. Combination treatments with peposertib and chemotherapeutics like DOX, epirubicin, and etoposide showed synergistic antiproliferative effects in TNBC cell lines [133]. Moreover, in xenograft models, combination treatments induced tumor regression and triggered pro-inflammatory responses. Additionally, peposertib in combination with IR significantly reduced tumor burden in cervical cancer models, supporting further clinical testing of peposertib with IR [135].
BAY-8400 originated from a screen aimed at identifying ATRis, during which a triazoloquinoxaline compound was found to inhibit ATR, ATM, and DNA-Pk [136]. This initial hit prompted lead optimization to enhance selectivity and potency toward DNA-Pk, ultimately yielding BAY-8400, a novel and selective DNA-PKi. This development underscores the value of cross-target screening in DDR pathways and highlights the potential of multi-kinase scaffolds [136]. BAY-8400 selectivity inhibits DNA-PK and prevents DSB repair, leading to DNA damage accumulation, genomic instability, and apoptosis, especially in cancer cells that rely heavily on NHEJ due to deficiencies in other repair pathways [137].
Preclinical studies have also demonstrated that BAY-8400 synergizes with DNA-damaging therapies, notably targeted alpha therapy (TAT), which utilizes alpha-emitting radionuclides for radiotherapeutics to induce DSBs. Furthermore, in combination with prostate-specific membrane antigen (PSMA)-targeted thorium-227 conjugates (BAY 2315497), BAY-8400 significantly enhanced antitumor efficacy in prostate cancer xenografts [136]. This effect stems from BAY-8400’s ability to inhibit DNA repair of alpha-particle-induced DSBs, making TAT more lethal to tumor cells [136,138]. BAY-8400 has also been optimized for high selectivity over PI3K and related kinases, minimizing off-target effects and improving its therapeutic window [139]. Related inhibitors, such as AZD7648, similarly showed minimal off-target activity across the kinome, underscoring the importance of DKA-Pk-specific inhibition in clinical development. Related inhibitors like AZD7648 similarly showed minimal off-target activity across the kinome, emphasizing the importance of DKA-Pk-specific inhibition in clinical development [139]. Thus, combining DDRis with DNA-damaging agents such as TAT represents a novel intervention in oncology, as blocking DSB repair sensitizes cancer cells to TAT-induced treatments, enhancing efficacy and potentially allowing dose reductions to mitigate toxicity [136,138].
WNC0901 is a highly potent and selective DNA-Pkcs inhibitor developed by Mladek and colleagues. Mladek et al. reported that WNC0901 exhibits potent kinase inhibition with an IC50 of 0.071 nM in a cell-free system, demonstrating over 30-fold greater selectivity than ATM, ATR, mTOR, and PI3K [140]. In U251 glioma and A549 lung cancer cells, 300 nM WNC0901 combined with 5 Gy radiation effectively suppressed DNA-Pkcs autophosphorylation. In HT29 cells, it showed an IC50 of 32.7 nM following 10 Gy exposure. Clonogenic assays revealed enhanced radio-sensitization, with survival dropping from 10% (radiation alone) to 1.5% at 100 and to 0.04% at 300 nM. Similarly, A549 cells showed reduced survival to 0.2% with combination therapy vs 19% with radiation alone [140]. These findings highlight the potential of WNC0901 as an effective radiosensitizer in GBM and lung cancer by targeting DNA-Pkcs.
In summary, based on data from numerous preclinical studies, DDR can be considered a class of anticancer drugs with the potential to inhibit cancer growth. However, clinical studies are warranted to assess the reproducibility of the preclinical findings to choose the most suitable DDR-based therapeutics to prevent cancer growth.
3.6 Challenges of DNA Damage Inhibitors
While the above-mentioned DNA damage inhibitors have demonstrated promising efficacy in targeting tumors with defective DNA repair pathways, numerous preclinical and clinical studies have indicated their potential to cause significant toxicity, especially when combined with chemotherapy. In this context, this section emphasizes the resistance mechanisms and drawbacks associated with DNA damage inhibitors.
Preclinical research suggests that ATMis can radiosensitize tumor cells, particularly those with deficiencies in the DDR pathway [141]. In colorectal cancer, ATMi has been shown to enhance the radiosensitizing effects of Bragg peak protons more than X-rays or entry protons, suggesting a potential strategy to improve RT [141]. Additionally, combining berzosertib with irinotecan showed promising activity in solid tumors with ATM mutations [142]. The clinical effectiveness of ATMi presents variability while researchers examine resistance mechanisms. A resistance mechanism activates ATM when ATR inhibitors are present, leading to G1 cell cycle arrest and reduced cytotoxicity.
The simultaneous inhibition of ATM and ATR proteins results in increased therapeutic effectiveness, according to Turchick et al. [36]. Furthermore, ATM-knockout (ATM-KO) prostate cancer cells can still repair IR-induced DNA damage through ATR and DNA-PKcs activation, highlighting the need for dual inhibition [143]. Overcoming ATM resistance involves combining ATM with ATR or DNA-PKis or targeting compensatory pathways [143–145].
Similarly, ATRi resistance arises through mechanisms such as enhanced replication fork progression, [146] cyclin C/CDK8 inhibition [147], or ATM pathway activation [36]. Combining ATRi with other DDRis such as ATMi and WEE1, along with biomarker-driven targeting of bypass pathways, offers a strategy to counteract resistance [52,146].
The use of CHK1i is justified because cancer cells, which often have impaired G1/S checkpoints, depend primarily on the CHK1-mediated G2/M checkpoint to survive. Clinical trials with CHK1i have demonstrated inconsistent efficacy despite encouraging preclinical results, and resistance mechanisms remain a significant obstacle. Numerous resistance mechanisms to CHK1i have been identified, including alterations in the thioredoxin system [148], PI3K/AKT signaling bypass [144], and increased WEE1 levels [149]. Studies also show a substantial correlation between the degree of acquired resistance and the messenger RNA (mRNA), protein levels, and DNA copy number of Wee1 [149]. Nevertheless, a combination of CHKi and PI3K/mTOR inhibitors has been shown to increase DNA damage, mitotic catastrophe, and attenuate cell viability [150].
WEE1i resistance has been shown to occur through the overexpression of MYT1 kinase, which blocks WEE1 by activating CDK1 [151]. This results in shortened premature mitotic entry and decreased mitotic duration, leading to enhanced survival rates (%) of adavosertib-treated cells [151]. This upregulation of MYT1 may also influence resistance to ATRi and CHK1i. Therefore, combining WEE1i with CHK1i ATRi, and Bcl-2 homology domain 3 (BH3) mimetics may be a useful strategy to inhibit treatment resistance [123]. Additionally, ZN-d5, as well as other BH3 mimetics, has been reported to cause caspase-induced DNA damage, along with Wee1 and ribonucleoside diphosphate reductase subunit M2 (RRM2) degradation, leading to synergistic efficacy when combined with ZN-c3 [123].
Despite the therapeutic potential of DNA-Pkis, resistance remains a significant hurdle. Intrinsic resistance can arise from tumor-specific genetic alterations or activation of alternative DNA repair pathways [96,152]. Acquired resistance may develop over time as cancer cells evolve in response to the selective stress. Notably, TP53-deficient tumors exhibit an increased reliance on MMEJ, conferring tolerance to DNA-PK inhibition [153]. In such cases, combining DNA-Pki with MMEJ (Polθ) inhibitors may induce synthetic lethality. Inhibition of DNA-PK has also been shown to enhance Polθ expression and end resection, promoting MMEJ activity and limiting the efficacy of monotherapy [153]. Additionally, normal tissues when exposed to DNA-Pki develop increased sensitivity to DNA-damaging agents like radiation and chemotherapy which prompts toxicity concerns [154]. The clinical translation of this approach demands the enhancement of the therapeutic index through increased tumor-selective cytotoxicity combined with reduced harm to normal tissues.
Across DDR-targeting agents, clinical outcomes remain heterogeneous and highly context-dependent. In pivotal early-phase studies, the ATR inhibitor BAY1895344 combined with topotecan achieved a median progression-free survival (PFS) of 9.2 months compared with 3.4 months with chemotherapy alone, while the WEE1 inhibitor adavosertib produced objective response rates (ORRs) of 26–38% in CCNE1-amplified tumors and platinum-resistant ovarian cancer [155,156]. Another phase I/II multi-tumor study using the ATR inhibitor ceralasertib (AZD6738) in ATM-deficient solid tumors and hematologic malignancies reported durable responses, with median overall survival (OS) approaching 12 months in select cohorts [157]. For PARP inhibitors, large-scale trials continue to demonstrate a survival benefit in BRCA-mutated ovarian and breast cancers, with median PFS improvements of 3–7 months compared with chemotherapy [158]. Although the therapeutic efficacy of DDR-targeting agents has shown great potential, class-related toxicities remain a significant limitation. For example, grade ≥ 3 hematologic adverse events (anemia, neutropenia, thrombocytopenia) have been reported in 25–45% of patients, alongside fatigue, nausea, and gastrointestinal intolerance [159]. More recently, Romesser et al. conducted a Phase Ib study of the DNA-PK inhibitor peposertib combined with neoadjuvant chemoradiation therapy (CRT) in patients with locally advanced rectal cancer (LARC). The patients were treated for 5 to 5.5 weeks with 50–250-mg peposertib administered once daily, capecitabine at 825 mg/m2 administered twice daily, and radiotherapy (RT) administered 5 days per week. In summary, while peposertib acts as a potent radiosensitizer, this Phase Ib study did not demonstrate clinical benefit in LARC patients. It was limited by significant gastrointestinal toxicity, highlighting challenges in combining DNA-PK inhibitors with full-dose chemoradiation [160]. Furthermore, intermittent or sequential dosing schedules, biomarker-guided patient selection (e.g., HRD or SLFN11 positivity), and nanoparticle-based formulations are now being explored to widen the therapeutic index and reduce overlapping myelosuppression in combination regimens [97].
4 Tumor Microenvironment-Induced Resistance to DDR Inhibitors and Immunologic Remodeling for Enhanced Effectiveness
The tumor microenvironment (TME) exerts decisive control over the efficacy of DDRis, functioning as both a refuge for resistant tumor cells and a dynamic regulator of treatment-induced immunity. Hypoxia, oxidative stress, and stromal remodeling activate compensatory repair pathways that sustain survival under genotoxic stress. At the same time, inflammatory cues and immune cell plasticity determine whether DDR perturbation triggers effective antitumor immunity or immune escape. Interactions among fibroblasts, endothelial cells, macrophages, and lymphocytes continuously reshape local cytokine gradients, vascular permeability, and redox balance factors that collectively dictate DDRi penetration and response. Understanding how these stromal, metabolic, and immune components intersect is essential for converting DDRi-induced stress into durable therapeutic benefit. The following subsections examine how hypoxia, stromal signaling, inflammation, and immune remodeling converge to create resistance, and how selective DDR targeting can reprogram the TME toward a more immunostimulatory state (Fig. 3).
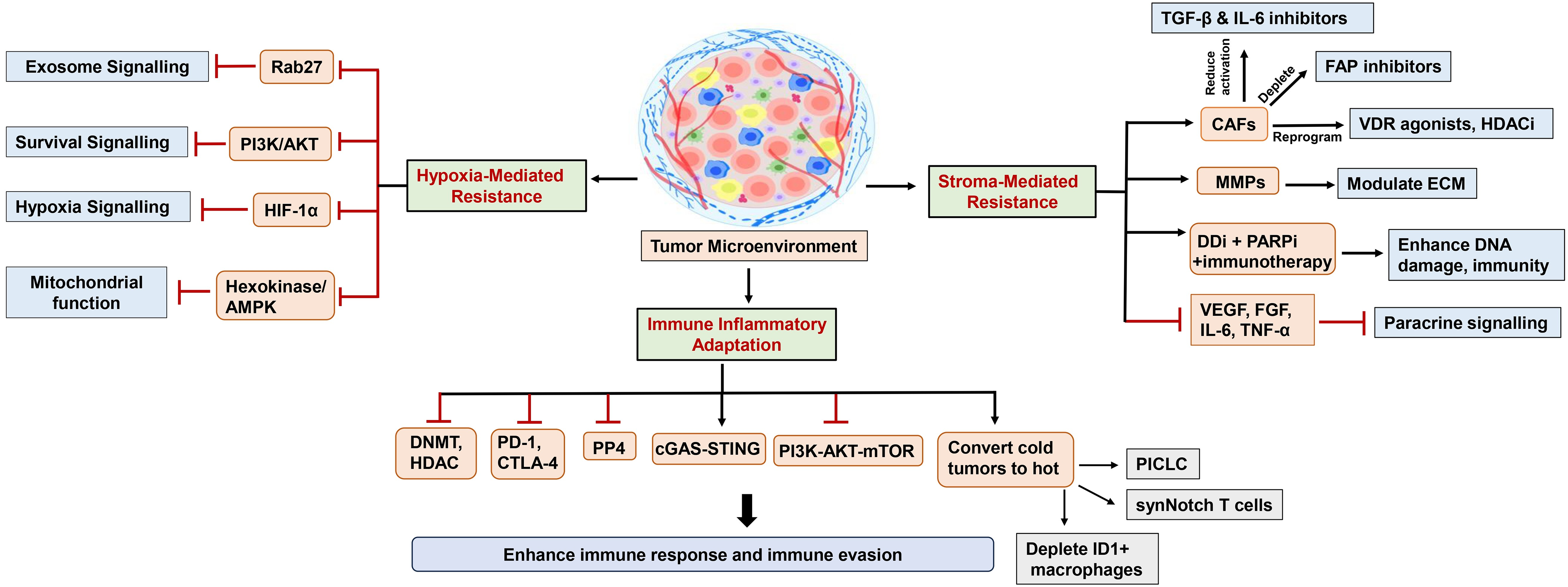
Figure 3: Schematic representation of mechanisms of TME-induced resistance to DNA damage response therapies
4.1 Hypoxia-Induced Resistance to DDR Inhibitors
Hypoxia, commonly found in the TME of solid tumors due to rapid growth and abnormal vasculature, drives therapy resistance by altering the metabolism and gene expression of cancer cells [161,162]. Under low-oxygen conditions, cancer cells activate hypoxia-inducible factors (HIFs), which drive adaptive mechanisms that support their survival, facilitate DNA repair, and enable them to evade treatment [163,164]. The activation of HIF-1α leads to the upregulation of DNA repair pathways, making tumor cells more resilient to genotoxic stress [165].
HIF-1 upregulates DNA repair genes, thereby enhancing repair capacity and reducing sensitivity to drugs such as PARP, ATM/ATR, and CHK1/CHK2 inhibitors [166,167]. It also disrupts cell cycle checkpoints and promotes epithelial-mesenchymal transition (EMT), thereby further enhancing resistance [167,168]. Additionally, hypoxia affects mitochondrial dynamics and promotes exosome-mediated signaling that spreads resistance traits [169,170]. In addition, hypoxia impairs drug delivery by disrupting vascular function and altering the tumor’s metabolic environment [171,172]. It also promotes the expansion of cancer stem-like cells and evasion of cell death by inducing autophagy and upregulating survival proteins [169,173,174].
A key resistance mechanism involves the hypoxia-induced upregulation of DDIT4 (also known as REDD1), which inhibits mTORC1 signaling, thereby aiding cell survival under stress and reducing sensitivity to radiotherapy and temozolomide. Whereas suppression of DDIT4 (REDD1) restores sensitivity [161]. Metabolic reprogramming in the TME plays a vital role in driving tumor progression by mediating interactions among cancer cells, fibroblasts, and immune cells [162]. Key regulators include mTOR, a downstream effector of PI3K/Akt signaling that controls cell growth, survival, and metabolism [175], and tAMPK, a central energy sensor that maintains metabolic balance [176].
The hypoxia-related gene COL5A1 is linked with poor prognosis while regulating immune pathways, which may affect chemotherapy response [162]. Moreover, hypoxic conditions promote genomic instability, which boosts mutation rates and develops resistance to DDRis [177]. Specific resistance mechanisms include enhanced homologous recombination under hypoxia, which undermines PARP inhibition, and HIF-1-driven bypass of ATM/ATR checkpoint signaling. CHK1/CHK2 inhibitor resistance arises through compensatory pathways and checkpoint regulation [167].
4.2 Stromal Interaction and DDR Inhibitors Resistance
Stromal cells, including fibroblasts, endothelial cells, and pericytes, are integral components of the TME, influencing resistance to DDRis through growth factor secretion, extracellular matrix (ECM) remodeling, immune modulation, and impaired drug delivery [178]. Among them, cancer-associated fibroblasts (CAFs) are particularly prominent. They secrete factors such as HGF and EGF that activate survival pathways, while their ECM remodeling creates a dense, drug-impermeable barrier [178].
CAFs also secrete cytokines such as transforming growth factor beta (TGF-β) and interleukin 6 (IL-6), which activate the PI3K/Akt, MAPK, and STAT3 pathways, thereby enhancing tumor cell survival and reducing the sensitivity of DDRis [179,180]. TGF-β promotes EMT, while IL-6 upregulates anti-apoptotic proteins, contributing to chemoresistance [181]. Additionally, CAF-derived ECM proteins, such as collagen, hinder drug penetration and sequester pro-survival cytokines [180,182]. Immunologically, CAFs recruit immunosuppressive cells, such as myeloid-derived suppressor cells (MDSCs) and regulatory T cells (Tregs), which suppress the activity of cytotoxic T cells and natural killer cells, thereby diminishing the immune-mediated response to DDRis [183–185].
Paracrine signaling from stromal cells further enhances resistance by activating pro-survival pathways in tumor cells. Growth factors like VEGF and FGF engage receptor tyrosine kinases (RTKs), promoting angiogenesis, ECM remodeling, and DNA repair while inhibiting apoptosis [179]. Cytokines, including IL-6 and TNF-α, activate inflammatory signaling pathways via STAT3 and NF-κB, which fosters genomic instability and drug resistance [186]. Stromal-derived exosomes also transfer resistance-promoting molecules such as microRNAs, proteins, and ECM components that bolster tumor survival and block drug penetration [187,188].
4.3 Inflammatory Signaling, Immune Cell Infiltration, and DDR Inhibitor Sensitivity
Inflammatory signaling in the TME significantly affects the response to DDRis [189]. Chronic inflammation supports tumor growth, angiogenesis, metastasis, and therapy resistance by creating a pro-survival environment. Pathways such as NF-κB and STAT3 upregulate anti-apoptotic proteins and DNA repair enzymes, thereby reducing the effectiveness of DDRis [189]. Since DNA damage itself activates NF-κB, DDR inhibition may amplify its signaling, promoting resistance to therapies that depend on inducing DNA damage [190]. Immune cell infiltration in the TME is essential for modulating sensitivity to DDRis [178,191]. DDRis increase tumor immunogenicity by elevating DNA damage, thereby enhancing the recruitment and activation of immune cells [189]. Nonetheless, immunosuppressive cells, such as myeloid-derived suppressor cells (MDSCs) and Tregs, can counteract this effect, promoting resistance [178].
Ultimately, the balance between immune activation and suppression in the TME determines the response to DDRis. Tumors with high cytotoxic immune infiltration and low immunosuppression respond better, while those dominated by suppressive cells are more resistant [178,189]. Combining DDRis with immunotherapies such as checkpoint blockade may improve therapeutic outcomes [191].
4.4 Immune Plasticity and DDR-Driven Remodeling of the Tumor Microenvironment
Inhibition of the DDR induces complex remodeling of the tumor immune landscape [192]. Myeloid and lymphoid populations adapt dynamically to DNA-damage-associated stress signals, balancing cytotoxic and suppressive functions [193]. For instance, tumor-associated macrophages (TAMs) exhibit remarkable plasticity, transitioning between pro-inflammatory (M1-like) and immunosuppressive (M2-like) states in response to signals from the TME, including cytokine cues (IL-12, IL-23, IL-4, IL-10, and IL-13) and ROS, which can be altered by DDR-targeting therapies [194]. Recent studies demonstrate that activation of the cGAS/STING pathway by accumulation of cytosolic DNA fragments following DDR inhibition in tumor or innate immune cells triggers type I interferon (such as IFN-β) release and chemokine production (CXCL8, CCL2), which contributes to both inflammation and recruitment of monocytes/macrophages into the TME, where additional cytokines direct their polarization [195,196]. However, accumulating evidence also suggests that chronic activation of the cGAS/STING pathway may exert an antitumor or pro-tumor effect, depending on numerous factors such as the rate of progression and the affected tissue [196,197]. Moreover, studies have reported an effect of DDR-targeted agents on immune cells such as Tregs and myeloid-derived suppressor cells (MDSCs) with immunosuppressive functions [194]. These context-dependent outcomes underscore that DDRi therapy can both ignite and dampen immunity, depending on exposure timing, tissue hypoxia, and stromal composition. Mechanistic dissection of this plasticity is therefore critical for identifying optimal sequencing with checkpoint blockade or cytokine modulation.
4.5 Mechanisms of “Cold-to-Hot” Tumor Conversion and Therapeutic Integration
In recent years, DDR-targeted therapies have emerged as potent activators of immunologic cell death (ICD) via enhancing tumor immunogenicity and activating tumor-specific immune sensing pathways (e.g., cGAS/STING-IFN), linking genotoxic stress to tumor-specific immune responses [198,199]. Inhibition of PARP, ATR, or WEE1 leads to accumulation of cytosolic DNA fragments and micronuclei, activating cGAS-STING-TBK1-IRF3 signaling and driving type I interferon and antigen-presentation pathways [200]. Cells have complex damage-associated molecular pathways that work harmoniously under stress or injury. As cancer cells die immunogenically, they release damage-associated molecular patterns (DAMPs) such as HMGB1, ATP and surface-exposed calreticulin [201–203]. These signals promote dendritic cell recruitment and the presentation of tumor-specific antigens to T cells, thereby inducing an immune response [203]. These processes together promote immunologically “cold” tumors into “hot,” inflamed lesions. However, immune escape via secondary up-regulation of programmed death-ligand 1 (PD-L1) and secretion of TGF-β or IL-6 can re-establish suppression, highlighting the importance of rational combinations [204,205]. For instance, studies using experimental preclinical models have demonstrated the favorable effects of DDR inhibitors when combined with immune checkpoint blockade, further enhancing the cytotoxic T-cell infiltration and tumor regression [206]. Additionally, targeting of stromal or angiogenic pathways (anti-TGF-β, anti-VEGF) can further stabilize immune activation. While preclinical studies have shown benefits, the overall success of the above-described strategies depends on several factors, including careful scheduling to exploit the early window of DAMP-driven stimulation before feedback inhibition predominates. Ultimately, integrating DDR modulation with immunotherapy offers a mechanistic route to overcoming TME-induced resistance and achieving durable “cold-to-hot” tumor reprogramming.
The impact of DDR modulation on the TME varies across tumor types, mutational landscapes, and baseline immune contexture [207]. Tumors with pre-existing interferon signaling or high neoantigen load, such as BRCA-mutated or mismatch-repair-deficient cancers, are especially susceptible to DDRi-driven immune activation. This susceptibility is linked to increased tumor mutation burden, neoantigen production, and heightened interferon pathway activity, collectively enhancing tumor immunogenicity and responsiveness to immunotherapy combined with DDR-targeted agents [208]. In contrast, hypoxic or desmoplastic tumors often remain refractory because of impaired drug penetration and persistent immunosuppressive signaling [209]. These tumors feature hypoxic regions that could induce genetic reprogramming and alter the DDR pathway, subsequently leading to resistance to DDR inhibitors and other therapies [210]. Also, persistent hypoxia in TME induces HIF-1 expression, which further promotes immunosuppression and reduces infiltration and function of immune effector cells [209,210].
While there have been advances in DDR-targeted therapies, these have not been without their limitations and challenges. In Section 5, we discuss advancements in nanocarrier-based delivery systems that may help overcome these barriers by improving intratumoral accumulation and reducing systemic toxicity, thereby enabling more effective immunotherapy combinations. Overall, DDRi-induced reprogramming of the TME reflects a dynamic balance between immunoregulatory mechanisms. Clarifying the temporal sequence of these responses, early DAMP-mediated activation followed by compensatory checkpoint induction, will be essential for optimizing dosing schedules and identifying biomarkers of durable “cold-to-hot” conversion. As this mechanistic insight deepens, DDR-targeted therapy is poised to emerge as a central modulator linking DNA repair dynamics to coordinated antitumor immunity.
4.6 Potential Strategies to Overcome TME-Induced Resistance
Overcoming TME-induced resistance to DDRis requires a comprehensive strategy targeting both cancer cells and their microenvironment [211,212]. Approaches include combining DDRis with chemotherapy, radiotherapy, immunotherapy, or agents like CHK1 or ATRis to enhance DNA damage and bypass resistance mechanisms [101,167,213]. In this section, we provide a detailed breakdown of current and emerging approaches to overcome TME-induced resistance.
4.6.1 Overcoming Hypoxia-Mediated DDR Inhibitor Resistance
Vasculature normalization using anti-angiogenic agents (e.g., bevacizumab) can reduce hypoxia and improve treatment delivery [161]. Targeting cancer metabolism, such as inhibiting pyrimidine synthesis, sensitizes cells to genotoxic agents [214]. Hypoxia-activated prodrugs (e.g., tirapazamine) selectively target low-oxygen environments [215], while HIF-1α pathway inhibition disrupts hypoxia-mediated signaling [165,216]. Improving oxygen delivery through vascular normalization, hyperbaric oxygen therapy, or the use of platinum-based nanoparticles can help reverse resistance [164,172,215,217]. Combining DDRis with chemotherapy, radiotherapy, or immunotherapy enhances efficacy in hypoxic tumors [172,216]. Importantly, combining DDRis with hypoxia-targeting agents (e.g., HIF-1 inhibitors or vascular normalization therapies) has shown promise [218,219]. Inhibiting the PI3K/AKT/HIF-1 axis or blocking exosome communication can also restore sensitivity [166,170]. Moreover, integrating DDRis with immunotherapy may overcome hypoxia-induced immunosuppression and improve outcomes [219]. Additionally, disrupting metabolic dependencies, such as glycolysis and mitochondrial function, also sensitizes these tumors [220,221].
Targeting the TME by inhibiting lysyl oxidase (LOX) or extracellular vesicle (EV) signaling improves drug delivery and reduces chemoresistance [220,222]. Hypoxia profiling and imaging technologies, such as photoacoustic imaging (PAI), can further guide precision treatment [223]. Clinical studies are evaluating hypoxia-targeted therapies to overcome resistance [224].
4.6.2 Overcoming Stromal-Mediated DDR Inhibitor Resistance
Effective treatment of stromal-mediated DDRi resistance demands simultaneous targeting of both tumor cells and stromal components. Practical approaches involve CAF targeting, ECM modulation, paracrine signaling inhibition, and the combined use of DDRis with additional treatments. Targeting the protective role of CAF enhances the efficacy of DDRi. This includes inhibiting CAF activation with TGF-β and IL-6 inhibitors (e.g., galunisertib, tocilizumab), depleting CAFs with FAP inhibitors such as talabostat, or reprogramming CAFs using VDR agonists or histone deacetylase inhibitors (HDACis) [182,185]. Also, modulating the ECM improves drug delivery and reduces resistance. Approaches include degrading collagen with collagenase [182], inhibiting LOX with BAPN [181], and disrupting ECM-integrin interactions using cilengitide [181].
Furthermore, inhibiting paracrine signaling can block protective signals, making tumor cells more susceptible to DDRis. Strategies include targeting growth factors (e.g., VEGF, FGF), cytokines (e.g., IL-6, TNF-α), and exosomes to enhance therapy sensitivity [179,187]. Combining DDRis with chemotherapy, radiation, or immunotherapy enhances treatment outcomes. PARPis (e.g., olaparib) and radiation therapy increase [225] DNA damage, while DDRis in immunotherapy, enhance the immune response by boosting antigen expression [211,226]. These strategies highlight the importance of combination therapies and TME targeting to overcome stromal-mediated resistance and improve cancer treatment outcomes [227].
4.6.3 Combination with Immune Checkpoint Inhibitors (ICIs)
DDRis can increase PD-L1 expression and tumor immunogenicity, making tumor cells more susceptible to ICIs. This synergy has shown promise in preclinical and clinical studies [228]. Raja et al. found that inhibiting protein phosphatase 4 (PP4) with carboplatin activates NF-κB and STAT1, thereby enhancing proinflammatory cytokine production and improving type I interferon responses, which suggests improved DDR inhibitor sensitivity through immune activation [189]. These pathways are interconnected, allowing NF-κB to trigger STAT3 through cytokine release, forming a feedback loop that maintains inflammation and promotes immune evasion by upregulating checkpoints, such as PD-L1 [229]. Therefore, targeting multiple inflammatory signals may be essential to enhance DDRi efficacy [189].
4.6.4 Enhancing Immune Infiltration into Cold Tumors
PP4 inhibition, achieved with fostriecin, combined with carboplatin, has been shown to enhance DNA damage and inflammatory signaling, resulting in increased infiltration of CD8+ T cells and NK cells and reduced tumor growth in ovarian cancer models [189]. This highlights PP4 as a target to enhance anti-tumor immunity, which targets interferon (IFN) responses and immune cell recruitment [191]. While DDRis can enhance this effect, chronic IFN signaling in some tumors may drive immunosuppression through the upregulation of checkpoint proteins, such as PD-L1 [229]. Approaches to convert cold tumors to hot tumors include 1) PICLC (polyinosinic-polycytidylic acid stabilized with poly-lysine and carboxymethylcellulose) to promote CD8+ T cell trafficking and activation, 2) localized delivery of IL-2 via engineered synNotch T cells to trigger localized immune activation, and 3) depleting ID1+ macrophages, with blocking CD8+ T cell infiltration, enhances antitumor immunity [230].
4.6.5 Activating the cGAS-STING Pathway
cGAS-STING agonists are being explored as adjuvants to overcome immune exclusion. For instance, the simultaneous use of DDRis and cGAS-STING activators enhances antitumor immune responses by augmenting type I interferon signaling, increasing cytoplasmic DNA levels, and improving T-cell infiltration, thereby overcoming resistance in BRCA1-deficient tumors [231]. Moreover, ATRi AZD6738, combined with radiotherapy and anti-PD-L1 treatment, has been shown to boost CD8+ T-cell activation while decreasing immunosuppressive populations and promoting long-term immune memory via cGAS-STING signaling pathways [232,233]. Additionally, STING agonists transform TAMs into anti-tumor cells, thereby restoring PARPi effectiveness through host STING-dependent type I IFN production and boosting CD8+ T-cell activity [234]. Furthermore, inhibition of PARP or CHK1 in SCLC has been shown to increase PD-L1 and chemokine levels through the cGAS-STING-TBK1-IRF3 pathway, making tumors more responsive to immune checkpoint blockade [235].
Similarly, the combination of PARPi with radiotherapy and anti-PD-L1 treatment stimulates cGAS-STING pathways while enhancing chemokine output and T-cell infiltration in SCLC, leading to improved antitumor results [236]. The combination of the CDC7 inhibitor XL413 and PARPi in ovarian cancer models has also been shown to enhance replication stress while activating cGAS–STING–mediated interferon responses, leading to significant tumor regression [237]. These findings collectively demonstrate the potential to improve cancer immunotherapy outcomes by activating innate immunity via DDR-induced cGAS-STING signaling. Therefore, the combination of cGAS-STING agonists and DDRis may provide new insights for treatment [235,238].
4.6.6 Inhibiting the PI3K-AKT-mTOR Pathway
Aberrant activation of the PI3K-AKT-mTOR Pathway contributes to immune evasion. Its inhibition can restore immune infiltration and sensitize tumors to ICIs, particularly those with PI3K pathway mutations [239].
Agents such as DNA methyltransferase inhibitors (DNMTis) and HDACis can enhance antigen presentation, increase T cell recruitment, and reactivate exhausted immune cells, making tumors more responsive to DDRis [228,240].
Overall, further research must focus on multi-pronged strategies that disrupt both the intrinsic survival mechanisms of tumor cells and the protective effects of the TME. Furthermore, future success will depend on rational trial designs, biomarker-driven patient selection, and continued mechanistic exploration.
5 Recent Advancements in Enhancing DDR Inhibitor Efficacy
The above-described studies indicate that while DNA damage inhibitors attenuate cancer growth, they have not been without their limitations. In this section, we explore the innovative use of nanoencapsulation techniques (Fig. 4) that enhance the delivery and efficacy of the DDR inhibitor.
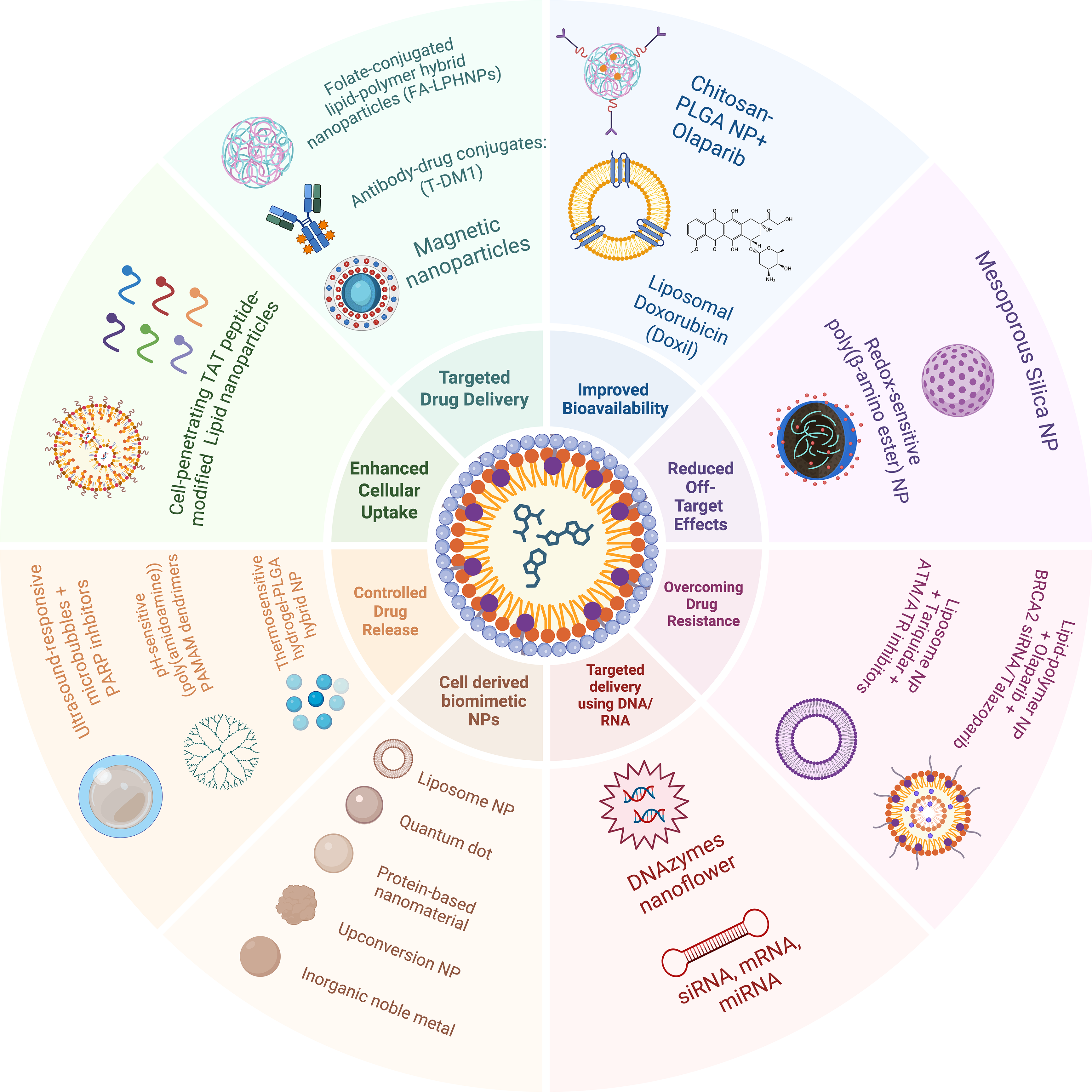
Figure 4: Potential application of nanoparticle-mediated drug delivery to enhance treatment responses to DNA damage response inhibitor therapies
5.1 Nanoparticle-Based Drug Delivery
Nanoparticle-based drug delivery systems enhance therapeutic precision through diverse platforms, such as liposomes, like Doxil®, which leverage the enhanced permeability and retention (EPR) effect for tumor accumulation [241]. For instance, folate-conjugated lipid-polymer hybrids (FA-LPHNPs) have been shown to target cancer cells via receptor-mediated endocytosis [242]. Also, stimuli-responsive systems, such as pH-sensitive mesoporous silica nanoparticles (MSNs), exhibit drug selectivity in the acidic TME [243–245]. Polymeric nanoparticles (e.g., PEG-PLA) have also been shown to improve the bioavailability of hydrophobic drugs, such as PARPis, by achieving sustained release [246–248]. Emerging innovations also include hybrid lipid-polymer nanoparticles (HLPNPs) for dual-drug delivery and biomimetic exosome-mimetic vesicles for enhanced glioblastoma targeting [249]. However, challenges persist in scalability, immune recognition, and long-term safety. Nevertheless, new interventions such as AI-driven drug design and CRISPR-loaded gold nanoparticles promise a breakthrough in personalized therapy [250–252]. Moreover, advances in sustainable nanomaterials and in organ-on-a-chip evaluation aim to address environmental and translational hurdles alongside DDRi interventions [253,254]. As discussed, the clinical success of DDRis targeting PARP, ATM, ATR, and CHK1 is often hindered by issues such as poor pharmacokinetics, limited tumor accumulation, synthetic lethality, and acquired drug resistance. Nanocarriers provide a versatile platform for overcoming these challenges by enhancing drug delivery, improving bioavailability, increasing specificity, and facilitating intracellular uptake. Below are representative studies on the strategies (Fig. 4) by which nanocarrier systems enhance the efficacy of DDRis.
Biocompatible nanocarriers reduce toxicity by utilizing degradable materials and employing surface modification techniques. FDA-approved PLGA-PEG NPs have demonstrated enhanced biocompatibility and biodegradability with several drugs in preclinical and clinical studies over the years, owing to their ability to degrade into CO2 and water. PEG minimizes immune recognition while biodegradable lipids like DSPC metabolize into non-toxic fatty acids. Clinical advancements include FDA-approved PLGA-PEG nanoparticles for leuprolide delivery, which degrade into CO2 and H2O [255], and albumin-bound paclitaxel (abraxane), which utilizes endogenous albumin to reduce immunogenicity in pancreatic cancer [256].
Targeted drug delivery uses tumor-specific ligands or antibodies to direct nanocarriers to receptors overexpressed on cancer cells, thereby minimizing off-target effects. A study by Gu et al. developed folate-conjugated, indocyanine green-loaded lipid-polymer hybrid nanoparticles (FA-LPHNPs) that bind folate receptors (FRs) on ovarian cancer cells, achieving 12-fold higher tumor accumulation in murine models compared to non-targeted systems [257,258]. These nanoparticles exploit receptor-mediated endocytosis, where FR binding triggers internalization, followed by liposomal acidification, which releases cisplatin directly into the cytoplasm, thereby bypassing efflux pumps. Over the years and recently, studies have demonstrated the clinical success of ADCs, such as trastuzumab-emtansine (T-DM1) (anti-HER2), in HER2+ breast cancer [259,260]. As well as the use of magnetic nanoparticles guided by external fields for tumor targeting, underscoring the versatility of ligand-mediated strategies [261,262].
Nanocarriers enhance the bioavailability of hydrophobic DDRis by shielding them from enzymatic degradation and rapid clearance. Anwer et al. demonstrated that Chitosan-coated PLGA nanoparticles encapsulating olaparib, which increased oral bioavailability by 4.75-fold in preclinical models [263]. PEGylation reduces opsonization by the mononuclear phagocyte system (MPS), while the PLA matrix enables sustained release, maintaining therapeutic plasma levels for 24 h. Studies have also highlighted similar improvements with liposomal doxorubicin (Doxil), where PEGylation extends the circulation half-life to 55 h [264], and a micellar SN-38 formulation that enhances solubility and reduces gastrointestinal toxicity [265,266].
5.5 Reduced Off-Target Effects
Stimuli-responsive nanocarrier minimizing off-target effects by releasing drugs selectively in the TME. The application of MSNs may offer an advanced strategy when encapsulated with DDRis, as the hydrazone bond grafted to the MSN’s surface hydrolyzes in acidic conditions, ensuring localized drug release [267]. Some recent advances include the formulation of doxorubicin-loaded redox-sensitive poly (β-amino ester) nanoparticles that degrade in high-glutathione environments to treat breast cancers [268], and MMP-2-expressed nano capsules for colorectal cancer, demonstrating the broad applicability of microenvironment-triggered systems [269].
Nanocarriers enhance the cellular uptake of DDRis via receptor-or peptide-mediated mechanisms. An early study by Futaki et al. highlights the role of lipid nanoparticles modified with cell-penetrating TAT peptides, which have been shown to achieve energy-independent enhanced internalization in breast cancer cells [270]. Consequently, a recent study shows that the PLGA-PEG shielded TAT peptide, when exposed only after selective binding to an angiotensin-converting enzyme (ACE2) inhibitor, is exposed and shows increased cellular uptake [271]. Moreover, siRNA-loaded cationic lipid carriers have also shown improved drug uptake in lung adenocarcinoma due to their ability to exploit electrostatic interactions for enhanced drug delivery to treat [272].
Controlled release systems maintain therapeutic drug levels while minimizing toxicity. For instance, thermosensitive hydrogel-PLGA hybrid (micro-TMZ@PLGA-PEG-PLGA) has been shown to be effective in the sustained release of TMZ in glioblastomas [273]. The hydrogel liquefies at body temperature, allowing PLGA to degrade via hydrolysis into lactic and glycolic acids, which govern zero-order release kinetics [274]. Innovations such as pH-sensitive (poly(amidoamine) PAMAM dendrimers for cisplatin delivery and ultrasound-responsive microbubbles for localized drug release further highlight the potential of programmable nanocarriers to sustain DDR inhibition [275,276].
5.8 Overcoming Drug Resistance
Co-delivery nanocarriers address resistance by simultaneously targeting DDR pathways and compensatory mechanisms. While previous investigations report the use of PARP siRNA-loaded lipidoid NPS in treating BRCA-resistant ovarian mice models [277]. Talazoparib-loaded solid lipid NPs (SLNPs) have been reported to directly overcome PARPi resistance in TNBCs that include BRCA-deficient as well as talazoparib-resistant tumor models [278]. The siRNA silences BRCA2 reversion mutations, turning off HR repair, while talazoparib blocks PARP-mediated BER, inducing synthetic lethality. Preclinical studies have also demonstrated similar success with liposome co-encapsulating paclitaxel and the P-gp inhibitor tariquidar, to circumvent efflux pump resistance, as well as dual ATM/ATR inhibitor NPs for TP53-mutant tumors [36,279].
5.9 Translational and Regulatory Challenges of Nanocarrier-Based DDR Inhibitor Delivery
While advances in the nanocarrier systems have transformed drug delivery by improving solubility, stability, and targeted accumulation, translating these constructs for DDRis remains complex. Accordingly, there are major obstacles in the clinical application of the nanocarrier systems, including variability in synthesis, limited tumor penetration, and evolving regulatory frameworks. A realistic appraisal of these challenges is essential to ensure that nanocarrier development advances in tandem with pharmacologic and biological understanding.
5.9.1 Manufacturing Scalability and Reproducibility
Recent studies have demonstrated major instabilities in nanoparticle formulations of DDRis, such as variability in size, surface charge (zeta potential), and drug payload loading, each of these factors impacts biodistribution and therapeutic index [280]. Additionally, studies have reported that alterations in size and charge affect tissue penetration, cellular uptake, and clearance rates, which subsequently impact the efficacy and toxicity profiles of DDRis [281,282]. Moreover, differences in solvent composition, mixing energy, or polymer molecular weight can shift key formulation attributes [283]. The above-mentioned discrepancies are often reported while transitioning from small laboratory scale-up process towards manufacturing. For instance, the scale-up process introduces new sources of variability in mixing, material handling, and process control, leading to batch-to-batch inconsistencies that directly impact nanoparticle characteristics critical for biodistribution and therapeutic efficacy [284,285]. Hence, adaptation of synthesis and stabilization protocols, well-optimized analytical methods, such as reverse-phase (RP) HPLC with dynamic light scattering, might enhance reproducibility and regulatory compliance could help to resolve a few concerns [286,287]. Moreover, for DDRi formulations, synchronizing drug release kinetics with transient DDR activation may also help.
5.9.2 Biodistribution, Tumor Delivery, and Biological Barriers
Another challenge in the nanocarrier system is the in vivo biological barriers [288]. For instance, hepatic and splenic macrophages mediated uptake often limits their tumor accumulation [289,290]. Additionally, the EPR effect observed in rodent tumor models is inconsistent in human tumors [291]. Therefore, surface modifications such as PEGylation or biomimetic coatings can prolong circulation and reduce immune recognition but may also impede cellular uptake or endosomal escape [292,293]. Quantitative imaging including advanced particle tracking with fluorescence/confocal microscopy, combined with mass spectrometric methods and mass spectrometric tracking, is increasingly used to assess actual nanoparticle delivery efficiency and refine carrier design for DDRi payloads [294,295].
5.9.3 Safety, Immunogenicity, and Off-Target Effects
Nanocarriers are not intrinsically inert. Chronic accumulation in the liver or spleen can induce inflammation or fibrosis, and surface chemistries such as PEGylation may trigger complement activation-related pseudoallergy (CARPA) or anti-PEG antibody formation, complicating safety profiles [296,297]. Because encapsulation does not eliminate the intrinsic cytotoxicity of DDRis, premature drug release may still damage normal tissues [296]. Preclinical evaluation using 3D microphysiological systems (MPS) and organ-on-chip (Ooc) platforms is therefore critical to define safe pharmacokinetic windows before human translation [253,298].
5.9.4 Regulatory Pathways and Clinical Translation
An overarching goal of research on nanocarrier systems is to improve unified global guidance and to emphasize early characterization of critical material attributes, particle stability, and immunological risk [299]. Additionally, establishing in vitro-in vivo correlation for release kinetics, defining bioequivalence criteria, and ensuring long-term toxicology data still impose significant challenges [300]. Moreover, the clinical success of approved nanomedicines liposomal doxorubicin, albumin-bound paclitaxel, and liposomal irinotecan, illustrates the relevance of manufacturing control and pharmacovigilance [301–304]. In the future, applying these lessons to DDRi nanocarriers will demand precise control of dual-drug ratios, robust stability under physiological conditions, and transparent reporting of biodistribution and immune responses.
Overall, advances in AI-assisted formulation design, scalable microfluidic synthesis, and improved analytical standards are poised to enhance reproducibility and accelerate regulatory review. It needs to be emphasized that, through early regulatory engagement, standardized safety testing, and comparative evaluation against free drugs, nanocarrier-based DDR inhibitors can evolve from experimental constructs into reproducible, patient-ready therapeutics.
6 Emerging Strategies and Future Directions
As we elucidate the therapeutic potential of DNA damage inhibitors in the modulation of cancer growth in preclinical and clinical studies, the horizon of DNA damage-targeting therapies continues to expand (Fig. 5). However, realizing the full therapeutic promise of DNA damage-targeting therapies requires a comprehensive and multidimensional future research and experimental models. Additionally, the following areas require focused attention to enhance the translational potential of DDR therapies:
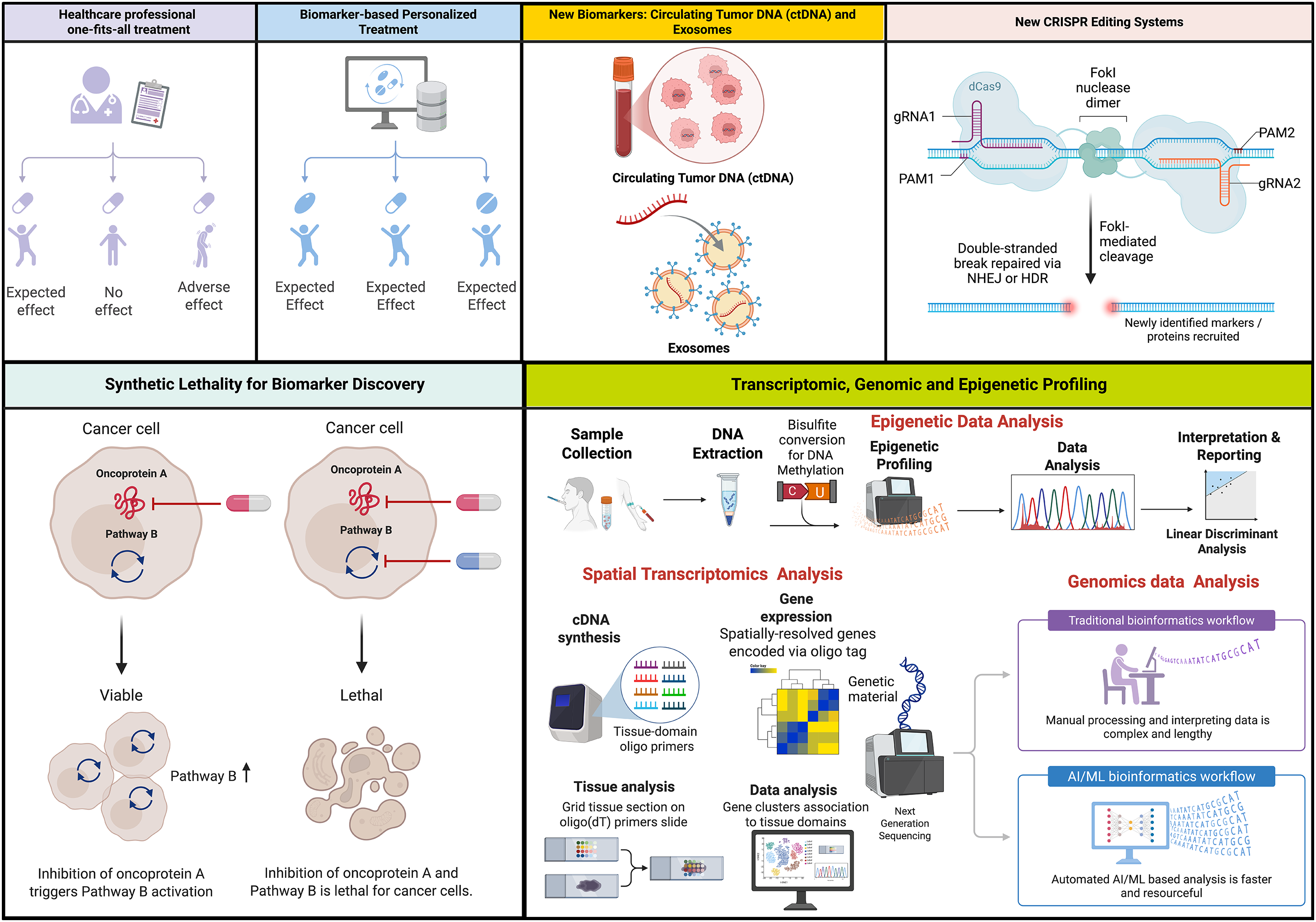
Figure 5: Schematic representation of emerging strategies offering enhanced disease detection and treatment outcomes to overcome long-term treatment resistance
6.1 Biomarker-Driven Patient Selection for DDR Inhibitor Therapy
DDR represents an essential cellular network responsible for maintaining genomic stability, whose dysregulation marks cancer presence [305]. Cancer cells often exhibit DDR gene deficiencies alongside replicative stress, making DDR pathways compelling therapeutic targets. Inhibiting DDR processes can induce excessive DNA damage, leading to cell death, particularly in cells with pre-existing DDR defects. PARPis have demonstrated the effectiveness of this approach, especially in tumors with BRCA mutations or homologous recombination deficiencies [305]. However, to expand the successful application of DDRis, refined strategies for patient selection are essential, with a focus on identifying predictive markers that go beyond simple BRCA mutations.
6.2 BRCA 1/2 Mutations and Beyond
Mutations in the BRCA1/2 genes are well-established predictors of sensitivity to PARP inhibitors. These mutations impair HR repair, rendering cancer cells more vulnerable to PARPis. However, a significant proportion of tumors without BRCA 1/2 mutations also exhibit DDR deficiencies and may respond to DDRis. Current research focuses on identifying non-BRCA-mutant tumors with potential responsiveness. Future clinical studies could assay key cancer-affected molecules to improve patient stratification for targeted therapy benefits [306].
6.3 Genomic Signatures: HRD Scores, RAD51 Foci and Related Markers
Genomic signatures, such as HR deficiency (HRD) scores, can refine patient selection for DDR inhibitor therapy. HRD scores assess the overall level of genomic instability resulting from defects in HR. These scores can identify tumors with BRCAness, a state of HRD that mimics the effects of BRCA 1/2 mutations [307]. Additionally, sequencing chromatin signatures of RAD51 and other assays, such as foci formation and immunoprecipitation, can be used to investigate RAD51-associated factors and assess the functional status of homology-directed repair (HDR). Tumors with impaired RAD51 foci formation are more likely to be sensitive to DDRis. Moreover, Genetic PD-L1 has also been recently shown to regulate HR-mediated end resection, where it regulates BRCAness and impacts synthetic lethality with PARPis, albeit some MAPK and mTORC1 signals are promoted PD-L1 surface receptors [308]. This indicates a new target for anticancer therapies: synthetic lethality approaches that mitigate alternative signaling pathways promoting metastasis.
6.4 Refined In Vitro Tools to Study New Biomarkers
Currently, in vitro assays utilizing Tus/Ter replication fork barriers to study short tract gene conversions (STGC) and long tract gene conversions (LTGC) in HDR [309], and I-SceI endonuclease reporters to study DSBs [310], provide invaluable information to identify several other regulatory markers other than repair factors that may recruit and stabilize DNA repair in cancer cells, thus defining futuristic models to treat severe malignancies and develop new clinical strategies. New CRISPR-based editing systems and enzymes are an interesting approach for identifying and mitigating new targets in cancer therapies [311–313].
6.5 Circulating Tumor DNA (ctDNA) and Exosome: The Emerging Biomarkers
Circulating tumor DNA (ctDNA) and exosomal markers are promising for real-time monitoring of DDRi efficacy [314,315]. CtDNA, or tumor-derived DNA, circulates in the bloodstream, and its levels can reflect the tumor burden and response to therapy [316]. Changes in ctDNA levels during DDRi treatment can provide early indications of drug efficacy or resistance. Also, exosome markers, such as DDR proteins or mutated DNA, can be used to trace DDRi response. For instance, exosomes have been implicated in mediating HR repair-directed end resection and recruiting RPA and RAD51 [317,318]. Therefore, ctDNA and exosomes represent novel therapeutic targets for enhancing DDRi treatment response.
Acquired resistance emerges through enhanced drug efflux pump expression, including ABC transporters, which actively expel DDRis from cells, thus lowering their intracellular levels and reducing therapeutic effectiveness [319]. The ATP-binding cassette (ABC) transporter superfamily comprises membrane proteins that utilize ATP hydrolysis to transport a wide range of substrates across cellular membranes actively. Overexpression of these transporters can significantly decrease the intracellular concentration of DDRis, preventing them from effectively targeting their intended molecular targets. For instance, gram-negative bacteria primarily resist antibiotics through drug efflux systems that work together with their outer membrane’s low permeability barrier and additional mutational and plasmid-borne resistance mechanisms [320]. These pumps exploit proton-motive force depletion across the IM to transport diverse substrates through the outer membrane (OM) while requiring accessory proteins for their operation which constitutes their major mechanistic advantage.
6.7 Altered DNA Repair Pathway Choice
Cancer cells can also develop resistance by altering their DNA repair pathway choice, shifting from HR to alternative, less precise pathways, such as NHEJ or polymerase theta-mediated repair [319]. These alternative pathways allow cancer cells to bypass the need for HR, which is the primary target of many DDRis, enabling them to repair DNA damage and survive despite DDR inhibition. Since the loss of the canonical DSB repair function (for example, due to defects in HR or altered checkpoint control, as in TP53-mutant tumors) forces the tumor to rely on alternative end-joining pathways, the principal path is microhomology-mediated end joining (MMEJ), which is driven by DNA polymerase theta (POLθ). This interesting target has been extensively studied more recently for its therapeutic potential [321]. Furthermore, POLθ is commonly overexpressed in HR-deficient and TP53-mutant cancers, where it is mechanistically linked to therapeutic resistance (e.g., escape from PARP inhibitors or genotoxic stress) by providing a mutagenic backup repair pathway that tolerates unresolved DSBs [322]. Therefore, several investigations show promising pharmacologic inhibition of POLθ by several chemotypes, including ART558 and novobiocin-like compounds and newer trapping inhibitors that suppress MMEJ and deplete PARP-inhibitor-resistant tumors from their acquired survival mechanism and are synthetically lethal with HR defects [323,324]. Similarly, combined targeting of POLθ and other DSB-repair nodes, such as DNA-PKi, NHEJ, HR inhibitors, and checkpoint kinase inhibitors, unleashes resection-dependent DNA damage in TP53-deficient backgrounds and provides new therapeutic outcomes by exploiting synthetic lethality against compensatory mechanisms.
6.8 Transcriptomic and Epigenetic Profiling
This method permits simultaneous identification of DDR weaknesses and counter-resistance strategies. Gene expression analysis identifies DDR deficiencies while DNA methylation together with histone modifications, represent epigenetic changes that alter chromatin accessibility and affect repair protein recruitment. These adaptations enable resistance through compensation for blocked DDR pathways or by activating alternative survival mechanisms. Through DNMT inhibition, scientists explore a re-sensitization approach, while HDAC inhibition boosts immune responses through enhancer reprogramming [319,325].
6.9 Lack of Standard Validation
A significant number of DDR-related biomarkers do not have standardized clinical validation which restricts their regular application. The identification of promising biomarkers demands extensive evaluation through future clinical trials before they can be applied in clinical settings. The process entails developing uniform assays alongside establishing optimal cutoff points while assessing biomarker predictive value across varied patient groups.
6.10 Need for Comprehensive Assays
Comprehensive genomic and functional assays are needed to guide therapy selection. Given the complexity of DDR pathways and the diverse mechanisms of DDR inhibitor resistance, single biomarkers maybe insufficient to predict treatment response accurately. Comprehensive assays that integrate genomics, spatial transcriptomics, epigenetics, and functional data are needed to provide a more complete picture of the DDR status of a tumor. These assays can help identify patients who are most likely to benefit from specific DDRis or combination therapies.
6.11 Exploiting Synthetic Lethality for Biomarker Discovery
The concept of synthetic lethality, where the synergistic efficacy of two or more defects leads to cell death, is central to the development of DDRis. Identifying new synthetic lethal interactions involving DDR pathways can uncover novel biomarkers for DDR response. For instance, the combination of ATR and PARPis has shown promise in IDH1/2-mutant cancers [326], suggesting that IDH1/2 mutations could serve as biomarkers for this combination therapy. Further, microtubule targeting agents (MTAs) have recently been implicated to target DNA DSB repair machinery and have shown significant efficacy to synergize with chemotherapy drugs and target tumor cells [327–329]. The essential function of microtubules, together with proteins such as γ-tubulin, kinesins, and intermediate filaments like vimentin, in sustaining cell structure and enabling inter-organelle communication has gained new significance through recent findings that connect these cytoskeletal elements to DSB repair, thereby creating novel opportunities in cancer research [328–330]. The role of these structural regulators as modulators of the DNA repair pathway highlights their therapeutic role in enhancing the efficacy of DNA damage treatment.
6.12 Cancer-Specific Considerations
In the case of pancreatic cancers, approximately 5%–10% of cases have DDR mutations, such as those in BRCA1 and BRCA2, which can be targeted with PARPis [249]. Defining the BRCAness phenotype in pancreatic cancer may be limited to patients harboring DDR genetic alterations [331]. For instance, olaparib demonstrated restricted antitumor effects in advanced platinum-sensitive pancreatic cancer patients with DDR genetic alterations, indicating potential treatment possibilities for specific patient groups [331]. Also, a considerable number of metastatic prostate cancer patients show germline mutations in DDR genes [332]. In a study, the presence of germline BRCA2 mutations negatively affected mCRPC patient outcomes, which showed dependence on the initial treatment approach used. The process of determining germline BRCA2 status can help select initial treatments for mCRPC [332].
6.13 Accelerating DDR-Targeted Precision Oncology
The advent of transformative technologies such as multi-omics, functional genomics, and computational modeling is redefining how the DDR is studied and therapeutically exploited. Representative studies demonstrate that these emerging platforms could reveal cellular heterogeneity, predict drug response, and reproduce patient-specific tumor complexity. Incorporating such technologies into DDR research is essential to overcome translational bottlenecks and to align DDRi development with the broader evolution of precision oncology. The merging role of spatial/single-cell omics, CRISPR screens, AI/ML predictive modeling, organoid/PDX systems, and an integrated translation framework is discussed in the present section.
6.13.1 Spatial and Single-Cell Omics Reveal DDR and TME Heterogeneity
Single-cell and spatial-omics approaches now expose the cellular diversity and microenvironmental context of DDR activation within tumors [333]. Platforms such as 10× Visium, MIBI, and CODEX imaging quantify phosphorylated markers (γH2AX, pCHK1) at subcellular resolution and map their distribution across hypoxic and immune niches [334,335]. There is a growing body of work combining transcriptomic, proteomic, and metabolic imaging to investigate “DDR-addicted” regions [336,337]. For example, combining transcriptomic, proteomic, and metabolic imaging has revealed that replication stress and oxygen gradients coincide with immune-exclusion areas that are most susceptible to DDR inhibition or nanoparticle-based delivery [338]. These data-driven atlases bridge molecular profiling and spatial pathology, allowing patient-specific mapping of DDR dependencies.
6.13.2 CRISPR Functional-Genomics Screens Define Synthetic-Lethal Networks
Genome-scale CRISPR/Cas9 and CRISPRi/a screens systematically identify vulnerabilities that condition DDRi sensitivity and resistance [339]. Recent studies have pinpointed POLQ as a synthetic-lethal partner of DNA-PK inhibition, MYT1 up-regulation as a cause of WEE1i resistance, and CDK8/Cyclin C loss as a determinant of ATRi escape [147,340,341]. Such discoveries clarify compensatory repair pathways and enable rational drug-combination design. When coupled with base-editing or enhancer-targeting platforms, CRISPR screens also map non-coding and regulatory elements that govern DDR gene expression, thereby enriching biomarker discovery for clinical translation [342].
6.13.3 Artificial Intelligence and Machine Learning for Predictive Modeling
As knowledge advances and artificial intelligence- and machine-learning-based mathematical models improve, such studies may significantly increase our understanding of DDRi response. Artificial-intelligence frameworks now integrate vast multi-omic and pharmacogenomic datasets to predict DDRi response [343,344]. For instance, models trained on DepMap and GDSC2 were found to correlate genomic instability signatures, homologous recombination deficiency scores, and transcriptomic features with therapeutic sensitivity and toxicity [345]. In drug design, deep-learning algorithms have also been reported to accelerate the optimization of kinase inhibitors, thereby enhancing selectivity and blood–brain barrier penetration [346,347]. At the same time, in-silico “digital-twin” simulations are rapidly advancing in tumor therapy via optimizing personalized treatment [348]. In the future, digital twin technology combined with genomic, radiomic, and pharmacokinetic derived inputs could ensure safety of DDRi regimens virtually before clinical exposure. Overall, these computational advances might improve and convert descriptive biology into predictive precision.
6.13.4 Patient-Derived Organoids and Xenografts Enable Functional Precision Testing
Patient-derived organoids (PDOs) and xenografts (PDXs) faithfully preserve tumor heterogeneity and microenvironmental cues, providing translational platforms to validate DDRi activity [349,350]. In the past few years, PDO biobanks from breast, ovarian, colorectal, and pancreatic cancers have demonstrated that replication stress levels and stromal interactions shape drug response [351]. Additionally, co-culture experiments with fibroblasts or immune cells have proven beneficial for recapitulating resistance mechanisms driven by cytokine or extracellular matrix signaling [352]. In the future, combining CRISPR perturbation with high-content imaging would enable researchers to assess synergy and toxicity in real time, effectively linking genomic prediction to pharmacologic validation.
In summary, an integrated use of spatial mapping, CRISPR functional genomics, AI analytics, and patient-derived modeling might allow a mechanistic resolution, predictive biomarkers, and experimentally testable therapeutic hypotheses. In the future, as integration expands to incorporate circulating tumor DNA dynamics and radiomic imaging, DDR-targeted therapy will transition from empirical evaluation to a data-driven paradigm in which each patient’s genomic instability landscape and microenvironmental architecture guide optimal inhibitor selection and combination strategies.
6.14 Effect of Gut Microbiome and Epigenetic Modulators on the Therapeutic Efficacy of DDR Inhibitor Response
In recent years, studies have reported that tumor-intrinsic alterations alone cannot fully explain the heterogeneity in therapeutic responses to DDRis. For example, the gut microbiome and the epigenetic landscape profoundly impact key signaling pathways related to the host immune system, metabolism, as well as chromatin architecture, thereby modulating DNA repair capacity, drug pharmacokinetics, and overall treatment efficacy. A better understanding of altered molecular networks is crucial for refining biomarker discovery and developing integrative precision oncology strategies.
6.14.1 Gut Microbiome as a Determinant of DDRi Efficacy and Toxicity
The intestinal microbiome regulates systemic inflammation, oxidative stress, and immunogenic signaling, all of which intersect with DDR and antitumor immunity [353]. Metabolites such as short-chain fatty acids (butyrate, propionate) and tryptophan-derived indoles regulate histone acetylation and type I interferon pathways, indirectly amplifying the cGAS–STING cascade triggered by DDR inhibition of commensal species, including Akkermansia muciniphila and Bifidobacterium spp [354]. An elevated immune activation has been associated with enhanced responses to PARP and ATR inhibitors [355,356]. At the same time, antibiotic-induced dysbiosis has been reported to dampen cytotoxic T-cell infiltration and increase gastrointestinal toxicity [357]. Additionally, microbial enzymes also modify NAD+ metabolism and drug conjugation, influencing the activity of NAD-dependent enzymes such as PARP [358]. These findings indicate that modulation of gut microbiota via dietary intervention, probiotics, and fecal transplantation could reduce systemic inflammation and enhance DDRi efficacy.
6.14.2 Epigenetic Regulation of DNA Repair and DDRi Sensitivity
Epigenetic control of chromatin structure determines accessibility of repair proteins and directly affects DDR pathway selection. Several studies show that promoter hypermethylation regulates genes involved in DNA repair pathways, such as BRCA1, MLH1, and RAD51C, potentially inducing a “BRCAness” phenotype that makes them sensitive to specific treatments, such as PARPi and ATRi, and enhances their sensitivity to these agents [359]. On the contrary, global hypomethylation promotes error-prone end joining and drug resistance. DNA-methyltransferase inhibitors (DNMTis) can reverse gene silencing and induce viral-mimicry responses by reactivating endogenous retroelements, thereby enhancing type I interferon signaling and immunogenic cell death when paired with DDR blockade [360,361]. Similarly, HDACis impair homologous recombination by disrupting RAD51 and 53BP1 assembly, thereby sensitizing tumors to PARP, WEE1, and ATR inhibitors [362,363]. Conversely, repressive marks such as H3K9me3 stabilize heterochromatin and confer radio resistance [364]. Non-coding RNAs, including LINC00460 and miR-182, add another layer of control by modulating molecular circuits related to immune response, checkpoint activity, and therapy response, thereby influencing tumor progression and synthetic-lethal responses [365,366].
6.14.3 The Interplay of Microbial and Epigenetic Modulation in Precision DDR Therapy
The interaction between microbial metabolites, immune tone, and chromatin remodeling establishes an epigenetic–metabolic axis that determines DDRi response [367]. Integrative profiling of fecal metabolites, circulating short chain fatty acids (SCFAs), and chromatin marks, such as LINE-1 methylation and histone H3 acetylation, may yield composite biomarkers of sensitivity and resistance [368]. Ongoing trials exploring combinations of DNMTis or HDACis with PARPi or ATRi aim to resensitize tumors that have become resistant [369]. At the same time, microbiome-directed interventions seek to maintain immune responsiveness during chronic DDR-targeted therapy [370,371]. Emerging areas of metagenomic sequencing, single-cell epigenomics, and AI-based modeling might pave the way for personalized DDRi regimens. Utilizing them in combination might fill a significant gap by addressing questions specific to each patient’s microbial and epigenetic landscape, providing a framework that links systemic biology with data-driven precision oncology.
Based on our insights, the DDR pathways represent a critical axis in maintaining genomic integrity and an increasingly valuable target in cancer therapy, with a focus on developing advanced inhibitors. Established strategies, particularly the use of PARPis in HR-deficient tumors, have demonstrated the therapeutic potential of exploiting synthetic lethality. Similarly, SMIs of key DDR kinases, such as ATM, ATR, and DNA-PK, have expanded the therapeutic landscape, although clinical efficacy remains hindered by both intrinsic and acquired resistance mechanisms. Therapeutic advancements in DDR-targeting now feature advanced and site-specific inhibitors designed to surpass existing treatment barriers while broadening synthetic lethality uses. The development of NP-based delivery systems is boosting DDRis’ precision and bioavailability while facing persistent toxicity challenges and concerns.
The response to DDRi treatments depends not only on tumor-intrinsic vulnerabilities but also on the complex interactions within the TME and immune landscape. A combination of immune infiltration, hypoxia, and stromal interactions drives resistance, which demands a transition to therapies that blend immunomodulatory agents with DDRis. The identification of promising biomarkers demands extensive evaluation through future clinical trials before they can be applied in medical settings. The process entails developing uniform assays alongside establishing optimal cutoff points while assessing biomarker predictive value across varied patient groups.
Future research should focus on understanding resistance through rational drug combinations, improving delivery platforms, and harnessing DDRis to enhance antitumor immunity using future technologies like AI-driven platforms. Advancing the clinical utility of DDR-targeting agents will require not only a deeper understanding of DDR biology but also integrated efforts to characterize the influence of the TME and immune system. Continued exploration of synthetic lethal interactions, biomarker discovery, and clinical trials involving combination regimens will be key to translating DDR inhibition into durable and widespread cancer therapies.
Acknowledgement: All the images in the present manuscript were created using BioRender.com.
Funding Statement: The authors received no specific funding for this study.
Author Contributions: Abhishikt David Solomon: Conceptualization, supervision, data curation, original draft, review and editing, figures and tables; Himanshu Kumar Vats: Data curation, original draft, review and editing; Shivam Chowdhary: Data curation, original draft, review and editing, figures and tables; Shivam Chowdhary: Data curation, original draft, review and editing, figures and tables; Supriya Nandlal Kanoujiya: Review and editing; Ajit Prakash: Review and editing; Hina Sultana: Review and editing; Sabyasachi Mohanty: Original draft, review and editing; Billy W. Day: Review and editing; Tarun Pant: Conceptualization, supervision, data curation, original draft, review and editing, figures and tables. All authors reviewed the results and approved the final version of the manuscript.
Availability of Data and Materials: Not applicable.
Ethics Approval: Not applicable.
Conflicts of Interest: Author Billy W. Day is employed by and is a co-owner of ReNeuroGen LLC. The remaining authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as potential conflicts of interest.
Abbreviations
| ATM | Ataxia-telangiectasia mutated |
| ATMi | ATM inhibitor |
| ATRi | ATR inhibitor |
| BER | Base excision repair |
| CHK | checkpoint kinases |
| DOX | Doxorubicin |
| DDAs | DNA-damaging agents |
| DDR | DNA damage response |
| DDRi | DNA damage response inhibitors |
| DNA-PK | DNA-dependent protein kinase |
| DNA-PKis | DNA-PK inhibitors |
| EGFR | Epidermal growth factor receptor |
| HDACis | Histone deacetylase inhibitors |
| HR | Homologous recombination |
| HIFs | Hypoxia-inducible factors |
| IDL | Insertion/deletion loop |
| IFN | Interferon |
| Ku-DBis | Ku-DNA binding inhibitors |
| NER | Nucleotide excision repair |
| NHEJ | Non-homologous end joining |
| PD-L1 | Programmed death-ligand 1 |
| PIKK | Phosphatidylinositol 3-kinase-related kinase |
| Tregs | Regulatory T cells |
| SMIs | Small-molecule inhibitors |
| SCLC | Small cell lung cancer |
| SSBs | Single strand breaks |
| ssDNA | Stranded DNA |
| TGF-β | Transforming growth factor beta |
| TLS | Translesion synthesis |
| WEE1 | WEE1 G2 checkpoint kinase |
References
1. Pilié PG, Tang C, Mills GB, Yap TA. State-of-the-art strategies for targeting the DNA damage response in cancer. Nat Rev Clin Oncol. 2019;16(2):81–104. doi:10.1038/s41571-018-0114-z. [Google Scholar] [PubMed] [CrossRef]
2. Tiwari V, Wilson DM. DNA damage and associated DNA repair defects in disease and premature aging. Am J Hum Genet. 2019;105(2):237–57. doi:10.1016/j.ajhg.2019.06.005. [Google Scholar] [PubMed] [CrossRef]
3. Curtin NJ. DNA repair dysregulation from cancer driver to therapeutic target. Nat Rev Cancer. 2012;12(12):801–17. doi:10.1038/nrc3399. [Google Scholar] [PubMed] [CrossRef]
4. Vadnais C, Möröy T. GFI1’s role in DNA repair suggests implications for tumour cell response to treatment. Cell Stress. 2018;2(8):213–5. doi:10.15698/cst2018.07.149. [Google Scholar] [PubMed] [CrossRef]
5. Kwok M, Agathanggelou A, Stankovic T. DNA damage response defects in hematologic malignancies: mechanistic insights and therapeutic strategies. Blood. 2024;143(21):2123–44. doi:10.1182/blood.2023019963. [Google Scholar] [PubMed] [CrossRef]
6. Li LY, Guan YD, Chen XS, Yang JM, Cheng Y. DNA repair pathways in cancer therapy and resistance. Front Pharmacol. 2021;11:629266. doi:10.3389/fphar.2020.629266. [Google Scholar] [PubMed] [CrossRef]
7. Li GM. Mechanisms and functions of DNA mismatch repair. Cell Res. 2008;18(1):85–98. doi:10.1038/cr.2007.115. [Google Scholar] [PubMed] [CrossRef]
8. Bröckelmann PJ, de Jong MRW, Jachimowicz RD. Targeting DNA repair, cell cycle, and tumor microenvironment in B cell lymphoma. Cells. 2020;9(10):2287. doi:10.3390/cells9102287. [Google Scholar] [PubMed] [CrossRef]
9. Krokan HE, Bjoras M. Base excision repair. Cold Spring Harb Perspect Biol. 2013;5(4):a012583. doi:10.1101/cshperspect.a012583. [Google Scholar] [PubMed] [CrossRef]
10. Kottemann MC, Smogorzewska A. Fanconi anaemia and the repair of Watson and Crick DNA crosslinks. Nature. 2013;493(7432):356–63. doi:10.1038/nature11863. [Google Scholar] [PubMed] [CrossRef]
11. Marteijn JA, Lans H, Vermeulen W, Hoeijmakers JHJ. Understanding nucleotide excision repair and its roles in cancer and ageing. Nat Rev Mol Cell Biol. 2014;15(7):465–81. doi:10.1038/nrm3822. [Google Scholar] [PubMed] [CrossRef]
12. Mareckova A, Malcikova J, Tom N, Pal K, Radova L, Salek D, et al. ATM and TP53 mutations show mutual exclusivity but distinct clinical impact in mantle cell lymphoma patients. Leuk Lymphoma. 2019;60(6):1420–8. doi:10.1080/10428194.2018.1542144. [Google Scholar] [PubMed] [CrossRef]
13. Choi M, Kipps T, Kurzrock R. ATM mutations in cancer: therapeutic implications. Mol Cancer Ther. 2016;15(8):1781–91. doi:10.1158/1535-7163.MCT-15-0945. [Google Scholar] [PubMed] [CrossRef]
14. Jiang H, Chang FC, Ross AE, Lee J, Nakayama K, Nakayama K, et al. Ubiquitylation of RAG-2 by Skp2-SCF links destruction of the V(D)J recombinase to the cell cycle. Mol Cell. 2005;18(6):699–709. doi:10.1016/j.molcel.2005.05.011. [Google Scholar] [PubMed] [CrossRef]
15. Jackson SP, Bartek J. The DNA-damage response in human biology and disease. Nature. 2009;461(7267):1071–8. doi:10.1038/nature08467. [Google Scholar] [CrossRef]
16. Groelly FJ, Fawkes M, Dagg RA, Blackford AN, Tarsounas M. Targeting DNA damage response pathways in cancer. Nat Rev Cancer. 2023;23(2):78–94. doi:10.1038/s41568-022-00535-5. [Google Scholar] [PubMed] [CrossRef]
17. Chwastek J, Jantas D, Lasoń W. The ATM kinase inhibitor KU-55933 provides neuroprotection against hydrogen peroxide-induced cell damage via a γH2AX/p-p53/caspase-3-independent mechanism: inhibition of calpain and cathepsin D. Int J Biochem Cell Biol. 2017;87:38–53. doi:10.1016/j.biocel.2017.03.015. [Google Scholar] [PubMed] [CrossRef]
18. Jossé R, Martin SE, Guha R, Ormanoglu P, Pfister TD, Reaper PM, et al. ATR inhibitors VE-821 and VX-970 sensitize cancer cells to topoisomerase I inhibitors by disabling DNA replication initiation and fork elongation responses. Cancer Res. 2014;74(23):6968–79. doi:10.1158/0008-5472.CAN-13-3369. [Google Scholar] [PubMed] [CrossRef]
19. Hirai H, Iwasawa Y, Okada M, Arai T, Nishibata T, Kobayashi M, et al. Small-molecule inhibition of Wee1 kinase by MK-1775 selectively sensitizes p53-deficient tumor cells to DNA-damaging agents. Mol Cancer Ther. 2009;8(11):2992–3000. doi:10.1158/1535-7163.MCT-09-0463. [Google Scholar] [PubMed] [CrossRef]
20. Kong A, Mehanna H. WEE1 inhibitor: clinical development. Curr Oncol Rep. 2021;23(9):107. doi:10.1007/s11912-021-01098-8. [Google Scholar] [PubMed] [CrossRef]
21. Ampolini EA, Jimenez-Sainz J, Long DT. The development of ATM inhibitors in cancer therapy. Target Oncol. 2025;20(2):281–97. doi:10.1007/s11523-025-01136-6. [Google Scholar] [PubMed] [CrossRef]
22. Munck JM, Batey MA, Zhao Y, Jenkins H, Richardson CJ, Cano C, et al. Chemosensitization of cancer cells by KU-0060648, a dual inhibitor of DNA-PK and PI-3K. Mol Cancer Ther. 2012;11(8):1789–98. doi:10.1158/1535-7163.MCT-11-0535. [Google Scholar] [PubMed] [CrossRef]
23. Karve S, Werner ME, Sukumar R, Cummings ND, Copp JA, Wang EC, et al. Revival of the abandoned therapeutic wortmannin by nanoparticle drug delivery. Proc Natl Acad Sci U S A. 2012;109(21):8230–5. doi:10.1073/pnas.1120508109. [Google Scholar] [PubMed] [CrossRef]
24. Chau CH, Steeg PS, Figg WD. Antibody-drug conjugates for cancer. Lancet. 2019;394(10200):793–804. doi:10.1016/S0140-6736(19)31774-X. [Google Scholar] [PubMed] [CrossRef]
25. Xiao Y, Pan T, Da W, Liu Y, Chen S, Chen D, et al. Aptamer-drug conjugates-loaded bacteria for pancreatic cancer synergistic therapy. Signal Transduct Target Ther. 2024;9(1):272. doi:10.1038/s41392-024-01973-3. [Google Scholar] [PubMed] [CrossRef]
26. Lavin MF. Ataxia-telangiectasia: from a rare disorder to a paradigm for cell signalling and cancer. Nat Rev Mol Cell Biol. 2008;9(10):759–69. doi:10.1038/nrm2514. [Google Scholar] [PubMed] [CrossRef]
27. Bunting SF, Nussenzweig A. End-joining, translocations and cancer. Nat Rev Cancer. 2013;13(7):443–54. doi:10.1038/nrc3537. [Google Scholar] [PubMed] [CrossRef]
28. Stinson BM, Loparo JJ. Repair of DNA double-strand breaks by the nonhomologous end joining pathway. Annu Rev Biochem. 2021;90(1):137–64. doi:10.1146/annurev-biochem-080320-110356. [Google Scholar] [PubMed] [CrossRef]
29. Karanam K, Kafri R, Loewer A, Lahav G. Quantitative live cell imaging reveals a gradual shift between DNA repair mechanisms and a maximal use of HR in mid S phase. Mol Cell. 2012;47(2):320–9. doi:10.1016/j.molcel.2012.05.052. [Google Scholar] [PubMed] [CrossRef]
30. Lovejoy CA, Cortez D. Common mechanisms of PIKK regulation. DNA Repair. 2009;8(9):1004–8. doi:10.1016/j.dnarep.2009.04.006. [Google Scholar] [PubMed] [CrossRef]
31. Huang X, You L, Nepovimova E, Psotka M, Malinak D, Valko M, et al. Inhibitors of phosphoinositide 3-kinase (PI3K) and phosphoinositide 3-kinase-related protein kinase family (PIKK). J Enzyme Inhib Med Chem. 2023;38(1):2237209. doi:10.1080/14756366.2023.2237209. [Google Scholar] [PubMed] [CrossRef]
32. Shiloh Y, Ziv Y. The ATM protein kinase: regulating the cellular response to genotoxic stress, and more. Nat Rev Mol Cell Biol. 2013;14(4):197–210. doi:10.1038/nrm3546. [Google Scholar] [PubMed] [CrossRef]
33. Zimmermann A, Zenke FT, Chiu LY, Dahmen H, Pehl U, Fuchss T, et al. A new class of selective ATM inhibitors as combination partners of DNA double-strand break inducing cancer therapies. Mol Cancer Ther. 2022;21(6):859–70. doi:10.1158/1535-7163.MCT-21-0934. [Google Scholar] [PubMed] [CrossRef]
34. Whiteaker JR, Wang T, Zhao L, Schoenherr RM, Kennedy JJ, Voytovich U, et al. Targeted mass spectrometry enables quantification of novel pharmacodynamic biomarkers of atm kinase inhibition. Cancers. 2021;13(15):3843. doi:10.3390/cancers13153843. [Google Scholar] [PubMed] [CrossRef]
35. Kciuk M, Gielecińska A, Kołat D, Kałuzińska Ż, Kontek R. Cancer-associated transcription factors in DNA damage response. Biochim Biophys Acta BBA Rev Cancer. 2022;1877(4):188757. doi:10.1016/j.bbcan.2022.188757. [Google Scholar] [PubMed] [CrossRef]
36. Turchick A, Zimmermann A, Chiu LY, Dahmen H, Elenbaas B, Zenke FT, et al. Selective inhibition of ATM-dependent double-strand break repair and checkpoint control synergistically enhances the efficacy of ATR inhibitors. Mol Cancer Ther. 2023;22(7):859–72. doi:10.1158/1535-7163.MCT-22-0685. [Google Scholar] [PubMed] [CrossRef]
37. Ueno S, Sudo T, Hirasawa A. ATM: functions of ATM kinase and its relevance to hereditary tumors. Int J Mol Sci. 2022;23(1):523. doi:10.3390/ijms23010523. [Google Scholar] [PubMed] [CrossRef]
38. Guleria A, Chandna S. ATM kinase: much more than a DNA damage responsive protein. DNA Repair. 2016;39:1–20. doi:10.1016/j.dnarep.2015.12.009. [Google Scholar] [PubMed] [CrossRef]
39. Lavin MF, Yeo AJ. Clinical potential of ATM inhibitors. Mutat Res. 2020;821:111695. doi:10.1016/j.mrfmmm.2020.111695. [Google Scholar] [PubMed] [CrossRef]
40. Lee JH, Paull TT. Cellular functions of the protein kinase ATM and their relevance to human disease. Nat Rev Mol Cell Biol. 2021;22(12):796–814. doi:10.1038/s41580-021-00394-2. [Google Scholar] [PubMed] [CrossRef]
41. Mendoza-Munoz PL, Gavande NS, VanderVere-Carozza PS, Pawelczak KS, Dynlacht JR, Garrett JE, et al. Abstract 2596: impact of novel Ku-DNA binding inhibitors on the DNA DSBs-induced DNA damage response. Cancer Res. 2022;82(12 Suppl):2596. doi:10.1158/1538-7445.am2022-2596. [Google Scholar] [CrossRef]
42. Sriramulu S, Thoidingjam S, Brown S, Siddiqui F, Movsas B, Green M, et al. Abstract A007: Ser/Thr kinase BUB1 stabilizes DNAPKcs in response to radiation induced DNA double strand break repair. Cancer Res. 2024;84(1_Suppl):A007. doi:10.1158/1538-7445.dnarepair24-a007. [Google Scholar] [CrossRef]
43. Maréchal A, Zou L. DNA damage sensing by the ATM and ATR kinases. Cold Spring Harb Perspect Biol. 2013;5(9):a012716. doi:10.1101/cshperspect.a012716. [Google Scholar] [PubMed] [CrossRef]
44. Marttila P, Bonagas N, Chalkiadaki C, Stigsdotter H, Schelzig K, Shen J, et al. The one-carbon metabolic enzyme MTHFD2 promotes resection and homologous recombination after ionizing radiation. Mol Oncol. 2024;18(9):2179–95. doi:10.1002/1878-0261.13645. [Google Scholar] [PubMed] [CrossRef]
45. Rinaldi C, Pizzul P, Casari E, Mangiagalli M, Tisi R, Longhese MP. The Ku complex promotes DNA end-bridging and this function is antagonized by Tel1/ATM kinase. Nucleic Acids Res. 2023;51(4):1783–802. doi:10.1093/nar/gkad062. [Google Scholar] [PubMed] [CrossRef]
46. Karlin J, Allen J, Ahmad SF, Hughes G, Sheridan V, Odedra R, et al. Orally bioavailable and blood-brain barrier-penetrating ATM inhibitor (AZ32) radiosensitizes intracranial gliomas in mice. Mol Cancer Ther. 2018;17(8):1637–47. doi:10.1158/1535-7163.mct-17-0975. [Google Scholar] [PubMed] [CrossRef]
47. Carruthers R, Ahmed SU, Strathdee K, Gomez-Roman N, Amoah-Buahin E, Watts C, et al. Abrogation of radioresistance in glioblastoma stem-like cells by inhibition of ATM kinase. Mol Oncol. 2015;9(1):192–203. doi:10.1016/j.molonc.2014.08.003. [Google Scholar] [PubMed] [CrossRef]
48. Vlatkovic T, Veldwijk MR, Giordano FA, Herskind C. Targeting cell cycle checkpoint kinases to overcome intrinsic radioresistance in brain tumor cells. Cancers. 2022;14(3):701. doi:10.3390/cancers14030701. [Google Scholar] [PubMed] [CrossRef]
49. Yano K, Shiotani B. Emerging strategies for cancer therapy by ATR inhibitors. Cancer Sci. 2023;114(7):2709–21. doi:10.1111/cas.15845. [Google Scholar] [PubMed] [CrossRef]
50. Roulston A, Zimmermann M, Papp R, Skeldon A, Pellerin C, Dumas-Bérube É, et al. RP-3500: a novel, potent, and selective ATR inhibitor that is effective in preclinical models as a monotherapy and in combination with PARP inhibitors. Mol Cancer Ther. 2022;21(2):245–56. doi:10.1158/1535-7163.mct-21-0615. [Google Scholar] [PubMed] [CrossRef]
51. Xie D, Jiang B, Wang S, Wang Q, Wu G. The mechanism and clinical application of DNA damage repair inhibitors combined with immune checkpoint inhibitors in the treatment of urologic cancer. Front Cell Dev Biol. 2023;11:1200466. doi:10.3389/fcell.2023.1200466. [Google Scholar] [PubMed] [CrossRef]
52. Jo U, Arakawa Y, Zimmermann A, Taniyama D, Mizunuma M, Jenkins LM, et al. Abstract LB301: molecular pharmacology and broad synergy of the novel ATR inhibitor M1774 with DNA damaging anticancer agents. Cancer Res. 2023;83(8_Suppl):LB301. doi:10.1158/1538-7445.am2023-lb301. [Google Scholar] [CrossRef]
53. Villette CC, Brightman F, Dupuy N, Zimmermann A, Lignet F, Zenke FT, et al. Abstract 5699: development and validation of a quantitative systems pharmacology model for prediction of preclinical efficacy of PARP inhibitors rucaparib and talazoparib combined with the ATR inhibitor gartisertib (M4344). Cancer Res. 2023;83(7_Suppl):5699. doi:10.1158/1538-7445.am2023-5699. [Google Scholar] [CrossRef]
54. Chowdhry S, Milutinovic S, Tse E, Garcia S, Perusse D, Ritland M, et al. Abstract 1626: tumors driven by oncogene amplified extrachromosomal DNA (ecDNA) demonstrate enhanced sensitivity to cell cycle checkpoint kinase 1 (CHK1) inhibition. Cancer Res. 2023;83(7_Suppl):1626. doi:10.1158/1538-7445.am2023-1626. [Google Scholar] [CrossRef]
55. Kumagai Y, Inagaki R, Shimizu K, Hirose W, Shimada N, Hayashi K, et al. Abstract 5724: preclinical evaluation of a novel CHK1 inhibitor encapsulated within the liposome, SMP-3124, for the treatment of solid cancer. Cancer Res. 2024;84(6_Suppl):5724. doi:10.1158/1538-7445.am2024-5724. [Google Scholar] [CrossRef]
56. van Bijsterveldt L, Durley SC, Maughan TS, Humphrey TC. The challenge of combining chemo- and radiotherapy with checkpoint kinase inhibitors. Clin Cancer Res. 2021;27(4):937–62. doi:10.1158/1078-0432.ccr-20-3358. [Google Scholar] [PubMed] [CrossRef]
57. Liao D, Hansen R, Chowdhry S, Holmes E, Wiese J, Steffy A, et al. Abstract 3631: preclinical and clinical pharmacodynamic characterization of BBI-355, a novel, orally bioavailable, and selective CHK1 inhibitor being evaluated in the first-in-human Phase 1/2 POTENTIATE clinical trial of patients with cancer harboring oncogene amplifications. Cancer Res. 2024;84(6_Suppl):3631. doi:10.1158/1538-7445.am2024-3631. [Google Scholar] [CrossRef]
58. Walker JR, Corpina RA, Goldberg J. Structure of the Ku heterodimer bound to DNA and its implications for double-strand break repair. Nature. 2001;412(6847):607–14. doi:10.1038/35088000. [Google Scholar] [PubMed] [CrossRef]
59. Zahid S, Seif El Dahan M, Iehl F, Fernandez-Varela P, Le Du MH, Ropars V, et al. The multifaceted roles of Ku70/80. Int J Mol Sci. 2021;22(8):4134. doi:10.3390/ijms22084134. [Google Scholar] [PubMed] [CrossRef]
60. Smith GCM, Jackson SP. The DNA-dependent protein kinase. Genes Dev. 1999;13(8):916–34. doi:10.1101/gad.13.8.916. [Google Scholar] [PubMed] [CrossRef]
61. Jiang W, Crowe JL, Liu X, Nakajima S, Wang Y, Li C, et al. Differential phosphorylation of DNA-PKcs regulates the interplay between end-processing and end-ligation during nonhomologous end-joining. Mol Cell. 2015;58(1):172–85. doi:10.1016/j.molcel.2015.02.024. [Google Scholar] [PubMed] [CrossRef]
62. Falck J, Coates J, Jackson SP. Conserved modes of recruitment of ATM, ATR and DNA-PKcs to sites of DNA damage. Nature. 2005;434(7033):605–11. doi:10.1038/nature03442. [Google Scholar] [PubMed] [CrossRef]
63. Jette N, Lees-Miller SP. The DNA-dependent protein kinase: a multifunctional protein kinase with roles in DNA double strand break repair and mitosis. Prog Biophys Mol Biol. 2015;117(2–3):194–205. doi:10.1016/j.pbiomolbio.2014.12.003. [Google Scholar] [PubMed] [CrossRef]
64. Li Z, Otevrel T, Gao Y, Cheng HL, Seed B, Stamato TD, et al. The XRCC4 gene encodes a novel protein involved in DNA double-strand break repair and V(D)J recombination. Cell. 1995;83(7):1079–89. doi:10.1016/0092-8674(95)90135-3. [Google Scholar] [PubMed] [CrossRef]
65. Ahnesorg P, Smith P, Jackson SP. XLF interacts with the XRCC4-DNA ligase IV complex to promote DNA nonhomologous end-joining. Cell. 2006;124(2):301–13. doi:10.1016/j.cell.2005.12.031. [Google Scholar] [PubMed] [CrossRef]
66. Ochi T, Blackford AN, Coates J, Jhujh S, Mehmood S, Tamura N, et al. PAXX, a paralog of XRCC4 and XLF, interacts with Ku to promote DNA double-strand break repair. Science. 2015;347(6218):185–8. doi:10.1126/science.1261971. [Google Scholar] [PubMed] [CrossRef]
67. Grawunder U, Wilm M, Wu X, Kulesza P, Wilson TE, Mann M, et al. Activity of DNA ligase IV stimulated by complex formation with XRCC4 protein in mammalian cells. Nature. 1997;388(6641):492–5. doi:10.1038/41358. [Google Scholar] [PubMed] [CrossRef]
68. Heyer WD, Ehmsen KT, Liu J. Regulation of homologous recombination in eukaryotes. Annu Rev Genet. 2010;44(1):113–39. doi:10.1146/annurev-genet-051710-150955. [Google Scholar] [PubMed] [CrossRef]
69. Spies J, Polasek-Sedlackova H, Lukas J, Somyajit K. Homologous recombination as a fundamental genome surveillance mechanism during DNA replication. Genes. 2021;12(12):1960. doi:10.3390/genes12121960. [Google Scholar] [PubMed] [CrossRef]
70. Choi EH, Yoon S, Koh YE, Seo YJ, Kim KP. Maintenance of genome integrity and active homologous recombination in embryonic stem cells. Exp Mol Med. 2020;52(8):1220–9. doi:10.1038/s12276-020-0481-2. [Google Scholar] [PubMed] [CrossRef]
71. Liu S, Kong D. End resection: a key step in homologous recombination and DNA double-strand break repair. Genome Instab Dis. 2021;2(1):39–50. doi:10.1007/s42764-020-00028-5. [Google Scholar] [CrossRef]
72. Sugiyama T, Zaitseva EM, Kowalczykowski SC. A single-stranded DNA-binding protein is needed for efficient presynaptic complex formation by the Saccharomyces cerevisiae Rad51 protein. J Biol Chem. 1997;272(12):7940–5. doi:10.1074/jbc.272.12.7940. [Google Scholar] [PubMed] [CrossRef]
73. Edlinger B, Schlögelhofer P. Have a break: determinants of meiotic DNA double strand break (DSB) formation and processing in plants. J Exp Bot. 2011;62(5):1545–63. doi:10.1093/jxb/erq421. [Google Scholar] [PubMed] [CrossRef]
74. Mazin AV, Alexeev AA, Kowalczykowski SC. A novel function of Rad54 protein. Stabilization of the Rad51 nucleoprotein filament. J Biol Chem. 2003;278(16):14029–36. doi:10.1074/jbc.M212779200. [Google Scholar] [PubMed] [CrossRef]
75. Wolner B, Peterson CL. ATP-dependent and ATP-independent roles for the Rad54 chromatin remodeling enzyme during recombinational repair of a DNA double strand break. J Biol Chem. 2005;280(11):10855–60. doi:10.1074/jbc.M414388200. [Google Scholar] [PubMed] [CrossRef]
76. Nimonkar AV, Genschel J, Kinoshita E, Polaczek P, Campbell JL, Wyman C, et al. BLM-DNA2-RPA-MRN and EXO1-BLM-RPA-MRN constitute two DNA end resection machineries for human DNA break repair. Genes Dev. 2011;25(4):350–62. doi:10.1101/gad.2003811. [Google Scholar] [PubMed] [CrossRef]
77. Wu Y, Sugiyama T, Kowalczykowski SC. DNA annealing mediated by Rad52 and Rad59 proteins. J Biol Chem. 2006;281(22):15441–9. doi:10.1074/jbc.M601827200. [Google Scholar] [PubMed] [CrossRef]
78. Baudat F, de Massy B. Regulating double-stranded DNA break repair towards crossover or non-crossover during mammalian meiosis. Chromosome Res. 2007;15(5):565–77. doi:10.1007/s10577-007-1140-3. [Google Scholar] [PubMed] [CrossRef]
79. Moynahan ME, Jasin M. Mitotic homologous recombination maintains genomic stability and suppresses tumorigenesis. Nat Rev Mol Cell Biol. 2010;11(3):196–207. doi:10.1038/nrm2851. [Google Scholar] [PubMed] [CrossRef]
80. Piazza A, Shah SS, Wright WD, Gore SK, Koszul R, Heyer WD. Dynamic processing of displacement loops during recombinational DNA repair. Mol Cell. 2019;73(6):1255–66.e4. doi:10.1016/j.molcel.2019.01.005. [Google Scholar] [PubMed] [CrossRef]
81. Yeh CD, Richardson CD, Corn JE. Advances in genome editing through control of DNA repair pathways. Nat Cell Biol. 2019;21(12):1468–78. doi:10.1038/s41556-019-0425-z. [Google Scholar] [PubMed] [CrossRef]
82. Bonilla B, Hengel SR, Grundy MK, Bernstein KA. RAD51 gene family structure and function. Annu Rev Genet. 2020;54(1):25–46. doi:10.1146/annurev-genet-021920-092410. [Google Scholar] [PubMed] [CrossRef]
83. Cidado J, Boiko S, Proia T, Ferguson D, Criscione SW, San Martin M, et al. AZD4573 is a highly selective CDK9 inhibitor that suppresses MCL-1 and Induces apoptosis in hematologic cancer cells. Clin Cancer Res. 2019;26(4):922–34. doi:10.1158/1078-0432.22473204.v1. [Google Scholar] [CrossRef]
84. Xu X, Xie K, Zhang XQ, Pridgen EM, Park GY, Cui DS, et al. Enhancing tumor cell response to chemotherapy through nanoparticle-mediated codelivery of siRNA and cisplatin prodrug. Proc Natl Acad Sci U S A. 2013;110(46):18638–43. doi:10.1073/pnas.1303958110. [Google Scholar] [PubMed] [CrossRef]
85. Gillespie MS, Ward CM, Davies CC. DNA repair and therapeutic strategies in cancer stem cells. Cancers. 2023;15(6):1897. doi:10.3390/cancers15061897. [Google Scholar] [PubMed] [CrossRef]
86. Ferri A, Stagni V, Barilà D. Targeting the DNA damage response to overcome cancer drug resistance in glioblastoma. Int J Mol Sci. 2020;21(14):4910. doi:10.3390/ijms21144910. [Google Scholar] [PubMed] [CrossRef]
87. Zhang J, Yang PL, Gray NS. Targeting cancer with small molecule kinase inhibitors. Nat Rev Cancer. 2009;9(1):28–39. doi:10.1038/nrc2559. [Google Scholar] [PubMed] [CrossRef]
88. Dimitrov T, Moschopoulou AA, Seidel L, Kronenberger T, Kudolo M, Poso A, et al. Design and optimization of novel benzimidazole- and imidazo[4,5-b]pyridine-based ATM kinase inhibitors with subnanomolar activities. J Med Chem. 2023;66(11):7304–30. doi:10.1021/acs.jmedchem.2c02104. [Google Scholar] [PubMed] [CrossRef]
89. Ivanenkov YA, Malyshev AS, Terentiev VA, Korzhenevskaya AA, Evteev SA, Vatsadze SZ, et al. Small molecule ATM inhibitors as potential cancer therapy: a patent review (2003–present). Expert Opin Ther Pat. 2025;35(2):111–36. doi:10.1080/13543776.2024.2446228. [Google Scholar] [PubMed] [CrossRef]
90. Tuma AM, Zhong W, Liu L, Burgenske DM, Carlson BL, Bakken KK, et al. Abstract 3305: WSD-0628, a novel brain penetrant ATM inhibitor, radiosensitizes GBM and melanoma patient derived xenografts. Cancer Res. 2022;82(12_Suppl):3305. doi:10.1158/1538-7445.am2022-3305. [Google Scholar] [CrossRef]
91. Zhou D, Wang Z, Liu YP, Fu T, Cheng ZZ. Abstract 7127: an ATM inhibitor: ZN-B-2262 in combination with radiation/ADCs containing topoisomerase inhibitors for the treatment of solid tumors. Cancer Res. 2024;84(6_Suppl):7127. doi:10.1158/1538-7445.am2024-7127. [Google Scholar] [CrossRef]
92. Laverty DJ, Gupta SK, Bradshaw GA, Hunter AS, Carlson BL, Calmo NM, et al. ATM inhibition exploits checkpoint defects and ATM-dependent double strand break repair in TP53-mutant glioblastoma. Nat Commun. 2024;15(1):5294. doi:10.1038/s41467-024-49316-8. [Google Scholar] [PubMed] [CrossRef]
93. Zhong W, Liu L, Sun C, Mu Z. Ddre-25. WSD0628: a brain penetrable atm inhibitor as a radiosensitizer for the treatment of gbm and metastatic cns tumor. Neuro Oncol. 2021;23(Suppl 6):vi79. doi:10.1093/neuonc/noab196.309. [Google Scholar] [CrossRef]
94. Rathi S, Oh JH, Zhang W, Mladek AC, Garcia DA, Xue Z, et al. Preclinical systemic pharmacokinetics, dose proportionality, and central nervous system distribution of the ATM inhibitor WSD0628, a novel radiosensitizer for the treatment of brain tumors. J Pharmacol Exp Ther. 2024;390(2):260–75. doi:10.1124/jpet.123.001971. [Google Scholar] [PubMed] [CrossRef]
95. McCabe N, Hanna C, Walker SM, Gonda D, Li J, Wikstrom K, et al. Mechanistic rationale to target PTEN-deficient tumor cells with inhibitors of the DNA damage response kinase ATM. Cancer Res. 2015;75(11):2159–65. doi:10.1158/0008-5472.can-14-3502. [Google Scholar] [PubMed] [CrossRef]
96. Gilmer TM, Lai CH, Guo K, Deland K, Ashcraft KA, Stewart AE, et al. Abstract LB181: first disclosure of XRD-0394, a novel dual ATM/DNA-PK inhibitor, that potently radiosensitizes and potentiates PARP and topoisomerase inhibitors. Cancer Res. 2024;84(7_Suppl):LB181. doi:10.1158/1538-7445.am2024-lb181. [Google Scholar] [CrossRef]
97. Qian J, Liao G, Chen M, Peng RW, Yan X, Du J, et al. Advancing cancer therapy: new frontiers in targeting DNA damage response. Front Pharmacol. 2024;15:1474337. doi:10.3389/fphar.2024.1474337. [Google Scholar] [PubMed] [CrossRef]
98. Hui Y, Kong Y, Jiang L, Deng M, Luo J, Jiang B, et al. Abstract 2604: the novel ATR inhibitor ATG-018 is efficacious in preclinical cancer models. Cancer Res. 2022;82(12 Suppl):2604. doi:10.1158/1538-7445.am2022-2604. [Google Scholar] [CrossRef]
99. Jo U, Senatorov IS, Zimmermann A, Saha LK, Murai Y, Kim SH, et al. Novel and highly potent ATR inhibitor M4344 kills cancer cells with replication stress, and enhances the chemotherapeutic activity of widely used DNA damaging agents. Mol Cancer Ther. 2021;20(8):1431–41. doi:10.1158/1535-7163.MCT-20-1026. [Google Scholar] [PubMed] [CrossRef]
100. Lücking U, Wortmann L, Wengner AM, Lefranc J, Lienau P, Briem H, et al. Damage incorporated: discovery of the potent, highly selective, orally available ATR inhibitor BAY 1895344 with favorable pharmacokinetic properties and promising efficacy in monotherapy and in combination treatments in preclinical tumor models. J Med Chem. 2020;63(13):7293–325. doi:10.1021/acs.jmedchem.0c00369. [Google Scholar] [PubMed] [CrossRef]
101. Wang C, Tang M, Chen Z, Nie L, Li S, Xiong Y, et al. Genetic vulnerabilities upon inhibition of DNA damage response. Nucleic Acids Res. 2021;49(14):8214–31. doi:10.1093/nar/gkab643. [Google Scholar] [PubMed] [CrossRef]
102. Shapiro GI, Wesolowski R, Devoe C, Lord S, Pollard J, Hendriks BS, et al. Phase 1 study of the ATR inhibitor berzosertib in combination with cisplatin in patients with advanced solid tumours. Br J Cancer. 2021;125(4):520–7. doi:10.1038/s41416-021-01406-w. [Google Scholar] [PubMed] [CrossRef]
103. Gralewska P, Gajek A, Marczak A, Mikuła M, Ostrowski J, Śliwińska A, et al. PARP inhibition increases the reliance on ATR/CHK1 checkpoint signaling leading to synthetic lethality-an alternative treatment strategy for epithelial ovarian cancer cells independent from HR effectiveness. Int J Mol Sci. 2020;21(24):9715. doi:10.3390/ijms21249715. [Google Scholar] [PubMed] [CrossRef]
104. Jette NR, Kumar M, Radhamani S, Arthur G, Goutam S, Yip S, et al. ATM-deficient cancers provide new opportunities for precision oncology. Cancers. 2020;12(3):687. doi:10.3390/cancers12030687. [Google Scholar] [PubMed] [CrossRef]
105. Tang Z, Pilié PG, Geng C, Manyam GC, Yang G, Park S, et al. ATR inhibition induces CDK1-SPOP signaling and enhances anti-PD-L1 cytotoxicity in prostate cancer. Clin Cancer Res. 2021;27(17):4898–909. doi:10.1158/1078-0432.CCR-21-1010. [Google Scholar] [PubMed] [CrossRef]
106. Yap TA, Krebs MG, Postel-Vinay S, El-Khouiery A, Soria JC, Lopez J, et al. Ceralasertib (AZD6738an oral ATR kinase inhibitor, in combination with carboplatin in patients with advanced solid tumors: a phase I study. Clin Cancer Res. 2021;27(19):5213–24. doi:10.1158/1078-0432.CCR-21-1032. [Google Scholar] [PubMed] [CrossRef]
107. Thompson R, Eastman A. The cancer therapeutic potential of Chk1 inhibitors: how mechanistic studies impact on clinical trial design. Br J Clin Pharmacol. 2013;76(3):358–69. doi:10.1111/bcp.12139. [Google Scholar] [PubMed] [CrossRef]
108. Chamoun K, Borthakur G. Investigational CHK1 inhibitors in early stage clinical trials for acute myeloid leukemia. Expert Opin Investig Drugs. 2018;27(8):661–6. doi:10.1080/13543784.2018.1508448. [Google Scholar] [PubMed] [CrossRef]
109. Ashwell S, Janetka JW, Zabludoff S. Keeping checkpoint kinases in line: new selective inhibitors in clinical trials. Expert Opin Investig Drugs. 2008;17(9):1331–40. doi:10.1517/13543784.17.9.1331. [Google Scholar] [PubMed] [CrossRef]
110. Rundle S, Bradbury A, Drew Y, Curtin NJ. Targeting the ATR-CHK1 axis in cancer therapy. Cancers. 2017;9(5):41. doi:10.3390/cancers9050041. [Google Scholar] [PubMed] [CrossRef]
111. Tozaki Y, Aoki H, Kato R, Toriuchi K, Arame S, Inoue Y, et al. The combination of ATM and Chk1 inhibitors induces synthetic lethality in colorectal cancer cells. Cancers. 2023;15(3):735. doi:10.3390/cancers15030735. [Google Scholar] [PubMed] [CrossRef]
112. Dent P. Investigational CHK1 inhibitors in early phase clinical trials for the treatment of cancer. Expert Opin Investig Drugs. 2019;28(12):1095–100. doi:10.1080/13543784.2019.1694661. [Google Scholar] [PubMed] [CrossRef]
113. Gorecki L, Andrs M, Korabecny J. Clinical candidates targeting the ATR-CHK1-WEE1 axis in cancer. Cancers. 2021;13(4):795. doi:10.3390/cancers13040795. [Google Scholar] [PubMed] [CrossRef]
114. Black EM, Joo YK, Kabeche L. Keeping RelApse in Chk: molecular mechanisms of Chk1 inhibitor resistance in lymphoma. Biochem J. 2022;479(22):2345–9. doi:10.1042/BCJ20220461. [Google Scholar] [PubMed] [CrossRef]
115. Giudice E, Zurcher G, Nair J, Huang T-T, Lee M-J, Trepel J, et al. Abstract 1562: correlative biomarker analysis of the phase II study of prexasertib, a cell cycle checkpoint kinase 1. Cancer Res. 2023;83(7 Suppl):1562. [Google Scholar]
116. Gao X, Liao Y, Li Q, Yang L, Zhao C, Teng Y, et al. Abstract 482: discovery of a novel and oral CHK1 inhibitor for the treatment of solid tumors. Cancer Res. 2023;83(7_Suppl):482. doi:10.1158/1538-7445.am2023-482. [Google Scholar] [CrossRef]
117. Lee KJ, Wright G, Bryant H, Wiggins LA, Schuler M, Gassman NR. EGFR signaling promotes resistance to CHK1 inhibitor prexasertib in triple negative breast cancer. Cancer Drug Resist. 2020;3(4):980–91. doi:10.20517/cdr.2020.73. [Google Scholar] [PubMed] [CrossRef]
118. Hansen RJ, Steffy A, Plum J, Ardeshiri A, Tse E, Garcia S, et al. Abstract 613: oral CHK1 inhibitor BBI-355 allows flexibility of dose and schedule with demonstration of monotherapy and combinational antitumor activity in extrachromosomal DNA (ecDNA) driven preclinical models. Cancer Res. 2024;84(6_Suppl):613. doi:10.1158/1538-7445.am2024-613. [Google Scholar] [CrossRef]
119. Maron SB, Sharma MR, Spira AI, Rodriguez Rivera II, Chawla SP, Philipovskiy A, et al. Preclinical and phase 1/2 data of the CHK1 inhibitor BBI-355 in development for esophageal and gastric cancers (EGC) withEGFRorFGFR2amplifications. J Clin Oncol. 2025;43(4 Suppl):tps517. doi:10.1200/jco.2025.43.4_suppl.tps517. [Google Scholar] [CrossRef]
120. Venkatachalam A, Peterson KL, Correia C, Flatten KS, Hou XN, Schneider PA, et al. CHK1 inhibitor prexasertib induces NOXA-dependent apoptosis in ovarian cancer. Cancer Res. 2022;82(12):1832. doi:10.1158/1538-7445.am2022-1832. [Google Scholar] [CrossRef]
121. Chao Y, Chen Y, Liu Y, Connelly JA, Wang H, Wang QJ. Abstract 5015: Wee1 inhibition and its synthetic lethal combination with Chk1 inhibition in mouse model of neuroendocrine prostate cancer. Cancer Res. 2023;83(7_Suppl):5015. doi:10.1158/1538-7445.am2023-5015. [Google Scholar] [CrossRef]
122. Yan J, Zhuang L, Wang Y, Jiang Y, Tu Z, Dong C, et al. Inhibitors of cell cycle checkpoint target Wee1 kinase—a patent review (2003–2022). Expert Opin Ther Pat. 2022;32(12):1217–44. doi:10.1080/13543776.2022.2166827. [Google Scholar] [PubMed] [CrossRef]
123. Izadi H, Ibrahim N, Hoang T, Ma J, de Jong PR, Pinchman J, et al. Abstract 2605: BH3 mimetics synergize with the Wee1 inhibitor ZN-c3 by activating caspases which induce DNA damage and degrade key proteins. Cancer Res. 2022;82(12_Suppl):2605. doi:10.1158/1538-7445.am2022-2605. [Google Scholar] [CrossRef]
124. Srivastava A, Heckman CA, Miettinen JJ, Harismendy O. Indicators of response to the Wee1 inhibitor adavosertib in acute myeloid leukemia. Blood. 2023;142(Suppl 1):5007. doi:10.1182/blood-2023-188608. [Google Scholar] [CrossRef]
125. Vacca J, Rocca S, Frye J, Schnell SJ, Hansbarger M, Korbel G, et al. Abstract 6177: ATRN-119 and ATRN-W1051: novel and potentially well tolerated ATR and WEE1 inhibitors for targeted cancer treatment. Cancer Res. 2023;83(7_Suppl):6177. doi:10.1158/1538-7445.am2023-6177. [Google Scholar] [CrossRef]
126. Wigerup C, Nilsson H, Shi L, Jung J, Baddour-Sousounis J, Cornelius R, et al. Abstract 1977: ACR-2316: a potentially first-in-class, potent, selective WEE1/PKMYT1 inhibitor rationally designed for superior single agent activity through synergistic disruption of cell cycle checkpoints. Cancer Res. 2024;84(6_Suppl):1977. doi:10.1158/1538-7445.am2024-1977. [Google Scholar] [CrossRef]
127. Bellon A, Saavedra O, Desar IM, Jalving M, Gietema JA, van Herpen C, et al. Abstract CT064: impact of food and high gastric pH on the bioavailability of the WEE1 inhibitor Debio 0123 assessed in a phase 1 dose escalation study. Cancer Res. 2024;84(7_Suppl):CT064. doi:10.1158/1538-7445.am2024-ct064. [Google Scholar] [CrossRef]
128. Mendoza-Munoz PL, Chauhan D, Gavande NS, Turchi JJ. Abstract B068: modulation of the DNA damage response by novel Ku-DNA binding inhibitors enhances cellular effects of DNA-double strand break inducing agents. Mol Cancer Ther. 2023;22(12_Suppl):B068. doi:10.1158/1535-7163.targ-23-b068. [Google Scholar] [CrossRef]
129. Mendoza-Munoz PL, Chauhan D, Gavande NS, Dynlacht JR, Garrett JE, Turchi JJ. Abstract 7128: novel Ku-DNA binding inhibitors impact on the cellular and in vivo DNA damage response to radio- and radiomimetic-therapy. Cancer Res. 2024;84(6_Suppl):7128. doi:10.1158/1538-7445.am2024-7128. [Google Scholar] [CrossRef]
130. Turchi JJ, Mendoza-Munoz PL, VanderVere-Carozza PS, Gavande NS, Dynlacht JR, Garrett JE, et al. Abstract A027: development of small molecule inhibitors of the Ku-DNA interaction: impacts on NHEJ, DDR signaling, optimizing genome editing technologies, and therapeutic intervention for the treatment of cancer. Cancer Res. 2024;84(1_Suppl):A027. doi:10.1158/1538-7445.dnarepair24-a027. [Google Scholar] [CrossRef]
131. Matsumoto Y. Development and evolution of DNA-dependent protein kinase inhibitors toward cancer therapy. Int J Mol Sci. 2022;23(8):4264. doi:10.3390/ijms23084264. [Google Scholar] [PubMed] [CrossRef]
132. Zhang K, Liu Z, Gao Y, Lu C, Cheng S, Liu X, et al. Abstract 4540: SY-7021, a novel DNA-PK inhibitor, exhibits significant anti-tumor activity in vitro and in vivo. Cancer Res. 2024;84(6_Suppl):4540. doi:10.1158/1538-7445.am2024-4540. [Google Scholar] [CrossRef]
133. Gordhandas SB, Manning-Geist B, Henson C, Iyer G, Gardner GJ, Sonoda Y, et al. Pre-clinical activity of the oral DNA-PK inhibitor, peposertib (M3814combined with radiation in xenograft models of cervical cancer. Sci Rep. 2022;12(1):974. doi:10.1038/s41598-021-04618-5. [Google Scholar] [PubMed] [CrossRef]
134. Revia S, Sirrenberg C, Schach A, Zimmermann A, Zenke FT, Albers J. Abstract 6215: peposertib, a DNA-PK inhibitor, enhances the antitumor efficacy of anthracyclines in triple-negative breast cancer models in vitro and in vivo. Cancer Res. 2023;83(7_Suppl):6215. doi:10.1158/1538-7445.am2023-6215. [Google Scholar] [CrossRef]
135. Hayrapetyan L, Roth SM, Hovhannisyan L, Medo M, Aebersold DM, Zimmer Y, et al. Abstract 498: p53-dependent treatment response to DNA-PK inhibition in combination with irradiation in head and neck squamous cell carcinoma (HNSCC) models. Cancer Res. 2023;83(7_Suppl):498. doi:10.1158/1538-7445.am2023-498. [Google Scholar] [CrossRef]
136. Berger M, Wortmann L, Buchgraber P, Lücking U, Zitzmann-Kolbe S, Wengner AM, et al. BAY-8400: a novel potent and selective DNA-PK inhibitor which shows synergistic efficacy in combination with targeted alpha therapies. J Med Chem. 2021;64(17):12723–37. doi:10.1021/acs.jmedchem.1c00762. [Google Scholar] [PubMed] [CrossRef]
137. Hu S, Hui Z, Lirussi F, Garrido C, Ye XY, Xie T. Small molecule DNA-PK inhibitors as potential cancer therapy: a patent review (2010–present). Expert Opin Ther Pat. 2021;31(5):435–52. doi:10.1080/13543776.2021.1866540. [Google Scholar] [PubMed] [CrossRef]
138. Wickstroem K, Hagemann UB, Cruciani V, Wengner AM, Kristian A, Ellingsen C, et al. Synergistic effect of a mesothelin-targeted (227)Th conjugate in combination with DNA damage response inhibitors in ovarian cancer xenograft models. J Nucl Med. 2019;60(9):1293–300. doi:10.2967/jnumed.118.223701. [Google Scholar] [PubMed] [CrossRef]
139. Goldberg FW, Finlay M, Ting A, Beattie D, Lamont GM, Fallan C, et al. The discovery of 7-methyl-2-[(7-methyl[1,2,4]triazolo[1,5-a]pyridin-6-yl)amino]-9-(tetrahydro-2h-pyran-4-yl)-7,9-dihydro-8h-purin-8-one (azd7648a potent and selective dna-dependent protein kinase (dna-pk) inhibitor. J Med Chem. 2019;63(7):3461–71. doi:10.1021/acs.jmedchem.9b01684. [Google Scholar] [PubMed] [CrossRef]
140. Mladek AC, Burgenske DL, Elmquist WF, Zhong W, Sarkaria JN. Abstract 701: preclinical data of a novel DNA-PK inhibitor in combination with radiation therapy shows promise in the treatment of established GBM and lung carcinoma cell lines. Cancer Res. 2024;84(6_Suppl):701. doi:10.1158/1538-7445.am2024-701. [Google Scholar] [CrossRef]
141. Callaghan CM, Dragojevic S. Bragg peak proton irradiation and ATM inhibition for rectal cancer. J Clin Oncol. 2025;43(4 Suppl):259. doi:10.1200/jco.2025.43.4_suppl.259. [Google Scholar] [CrossRef]
142. Villaruz LC, Kelly K, Waqar SN, Davis EJ, Shapiro G, LoRusso P, et al. NCI 9938: phase I clinical trial of ATR inhibitor berzosertib (M6620, VX-970) in combination with irinotecan in patients with advanced solid tumors. J Clin Oncol. 2022;40(16 Suppl):3012. doi:10.1200/jco.2022.40.16_suppl.3012. [Google Scholar] [CrossRef]
143. Hofstad ME, Yu L, Woods A, Sychev ZE, Mazzagatti A, Huo X, et al. Abstract A020: deciphering DNA damage repair in ATM mutant prostate cancers. Cancer Res. 2024;84(1_Suppl):A020. doi:10.1158/1538-7445.dnarepair24-a020. [Google Scholar] [CrossRef]
144. Hunter JE, Campbell AE, Kerridge S, Fraser C, Hannaway NL, Luli S, et al. Up-regulation of the PI3K/AKT and RHO/RAC/PAK signalling pathways in CHK1 inhibitor resistant Eμ-Myc lymphoma cells. Biochem J. 2022;479(19):2131–51. doi:10.1042/bcj20220103. [Google Scholar] [PubMed] [CrossRef]
145. Hofstad M, Yu L, Woods A, Sychev Z, Gilbreath C, Huo X, et al. Abstract 2405: delineating molecular vulnerabilities of ATM mutant prostate cancers. Cancer Res. 2023;83(7_Suppl):2405. doi:10.1158/1538-7445.am2023-2405. [Google Scholar] [CrossRef]
146. Schleicher EM, Dhoonmoon A, Jackson LM, Clements KE, Stump CL, Nicolae CM, et al. Dual genome-wide CRISPR knockout and CRISPR activation screens identify mechanisms that regulate the resistance to multiple ATR inhibitors. PLoS Genet. 2020;16(11):e1009176. doi:10.1371/journal.pgen.1009176. [Google Scholar] [PubMed] [CrossRef]
147. Lloyd RL, Urban V, Muñoz-Martínez F, Ayestaran I, Thomas JC, de Renty C, et al. Loss of Cyclin C or CDK8 provides ATR inhibitor resistance by suppressing transcription-associated replication stress. Nucleic Acids Res. 2021;49(15):8665–83. doi:10.1093/nar/gkab628. [Google Scholar] [PubMed] [CrossRef]
148. Prasad CB, Oo A, Qiu Z, Li N, Singh D, Xin X, et al. Abstract B024: the thioredoxin system determines CHK1 inhibitor sensitivity via redox-mediated regulation of ribonucleotide reductase activity. Cancer Res. 2024;84(1_Suppl):B024. doi:10.1158/1538-7445.dnarepair24-b024. [Google Scholar] [CrossRef]
149. Zhao X, Kim IK, Kallakury B, Chahine JJ, Iwama E, Pierobon M, et al. Acquired small cell lung cancer resistance to Chk1 inhibitors involves Wee1 up-regulation. Mol Oncol. 2021;15(4):1130–45. doi:10.1002/1878-0261.12882. [Google Scholar] [PubMed] [CrossRef]
150. Huang TT, Brill E, Nair JR, Zhang X, Wilson KM, Chen L, et al. Targeting the PI3K/mTOR pathway augments CHK1 inhibitor-induced replication stress and antitumor activity in high-grade serous ovarian cancer. Cancer Res. 2020;80(23):5380–92. doi:10.1158/0008-5472.CAN-20-1439. [Google Scholar] [PubMed] [CrossRef]
151. Sokhi S, Lewis CW, Bukhari AB, Hadfield J, Xiao EJ, Fung J, et al. Myt1 overexpression mediates resistance to cell cycle and DNA damage checkpoint kinase inhibitors. Front Cell Dev Biol. 2023;11:1270542. doi:10.3389/fcell.2023.1270542. [Google Scholar] [PubMed] [CrossRef]
152. Qian C, Li X, Zhang J, Wang Y. Small molecular inhibitors that target ATM for drug discovery: current research and potential prospective. J Med Chem. 2024;67(17):14742–67. doi:10.1021/acs.jmedchem.4c01064. [Google Scholar] [PubMed] [CrossRef]
153. Patterson-Fortin J, Bose A, Tsai WC, Grochala C, Nguyen H, Zhou J, et al. Abstract 796: dual inhibition of NHEJ and MMEJ induces synthetic lethality in TP53 mutant cancers. Cancer Res. 2022;82(12_Suppl):796. doi:10.1158/1538-7445.am2022-796. [Google Scholar] [CrossRef]
154. Baker JHE, Kyle AH, Liu NA, Wang T, Liu X, Teymori S, et al. Radiation and chemo-sensitizing effects of DNA-PK inhibitors are proportional in tumors and normal tissues. Mol Cancer Ther. 2024;23(9):1230–40. doi:10.1158/1535-7163.MCT-23-0681. [Google Scholar] [PubMed] [CrossRef]
155. Stockton S, Shyr C, Cecchini M, Aljumaily R, Halfdanarson TR, Sonbol MB, et al. A phase I study of ATR inhibitor BAY1895344 (elimusertib) plus topotecan (ETCTN 10402results of dose escalation. J Clin Oncol. 2024;42(16 Suppl):3076. doi:10.1200/jco.2024.42.16_suppl.3076. [Google Scholar] [CrossRef]
156. Heumann TR, Stockton S, Cecchini M, Aljumaily R, Shyr C, Whisenant J, et al. A phase I study of irinotecan combined with BAY1895344 (ATR inhibitor) in advanced solid tumors: results of ETCTN 10402 dose escalation. J Clin Oncol. 2024;42(16 Suppl):3077. doi:10.1200/jco.2024.42.16_suppl.3077. [Google Scholar] [CrossRef]
157. Kwon M, Kim G, Kim R, Kim KT, Kim ST, Smith S, et al. Phase II study of ceralasertib (AZD6738) in combination with durvalumab in patients with advanced gastric cancer. J Immunother Cancer. 2022;10(7):e005041. doi:10.1136/jitc-2022-005041. [Google Scholar] [PubMed] [CrossRef]
158. Shao F, Duan Y, Zhao Y, Li Y, Liu J, Zhang C, et al. PARP inhibitors in breast and ovarian cancer with BRCA mutations: a meta-analysis of survival. Aging. 2021;13(6):8975–88. doi:10.18632/aging.202724. [Google Scholar] [PubMed] [CrossRef]
159. Martorana F, Da Silva LA, Sessa C, Colombo I. Everything comes with a price: the toxicity profile of DNA-damage response targeting agents. Cancers. 2022;14(4):953. doi:10.3390/cancers14040953. [Google Scholar] [PubMed] [CrossRef]
160. Romesser PB, Capdevila J, Garcia-Carbonero R, Philip T, Fernandez Martos C, Tuli R, et al. A phase Ib study of the DNA-PK inhibitor peposertib combined with neoadjuvant chemoradiation in patients with locally advanced rectal cancer. Clin Cancer Res. 2024;30(4):695–702. doi:10.1158/1078-0432.CCR-23-1129. [Google Scholar] [PubMed] [CrossRef]
161. Foltyn M, Luger AL, Lorenz NI, Sauer B, Mittelbronn M, Harter PN, et al. The physiological mTOR complex 1 inhibitor DDIT4 mediates therapy resistance in glioblastoma. Br J Cancer. 2019;120(5):481–7. doi:10.1038/s41416-018-0368-3. [Google Scholar] [PubMed] [CrossRef]
162. Zhu H, Hu X, Feng S, Jian Z, Xu X, Gu L, et al. The hypoxia-related gene COL5A1 is a prognostic and immunological biomarker for multiple human tumors. Oxid Med Cell Longev. 2022;2022(1):6419695. doi:10.1155/2022/6419695. [Google Scholar] [PubMed] [CrossRef]
163. Hompland T, Fjeldbo CS, Lyng H. Tumor hypoxia as a barrier in cancer therapy: why levels matter. Cancers. 2021;13(3):499. doi:10.3390/cancers13030499. [Google Scholar] [PubMed] [CrossRef]
164. Zhuang Y, Liu K, He Q, Gu X, Jiang C, Wu J. Hypoxia signaling in cancer: implications for therapeutic interventions. MedComm. 2023;4(1):e203. doi:10.1002/mco2.203. [Google Scholar] [PubMed] [CrossRef]
165. Ullah A, Leong SW, Wang J, Wu Q, Ahmad Ghauri M, Sarwar A, et al. Cephalomannine inhibits hypoxia-induced cellular function via the suppression of APEX1/HIF-1α interaction in lung cancer. Cell Death Dis. 2021;12(5):490. doi:10.1038/s41419-021-03771-z. [Google Scholar] [PubMed] [CrossRef]
166. Sun S, Guo C, Gao T, Ma D, Su X, Pang Q, et al. Hypoxia enhances glioma resistance to sulfasalazine-induced ferroptosis by upregulating SLC7A11 via PI3K/AKT/HIF-1α axis. Oxid Med Cell Longev. 2022;2022(1):7862430. doi:10.1155/2022/7862430. [Google Scholar] [PubMed] [CrossRef]
167. Deng S, Vlatkovic T, Li M, Zhan T, Veldwijk MR, Herskind C. Targeting the DNA damage response and DNA repair pathways to enhance radiosensitivity in colorectal cancer. Cancers. 2022;14(19):4874. doi:10.3390/cancers14194874. [Google Scholar] [PubMed] [CrossRef]
168. Lu Y, Liu Y, Oeck S, Zhang GJ, Schramm A, Glazer PM. Hypoxia induces resistance to EGFR inhibitors in lung cancer cells via upregulation of FGFR1 and the MAPK pathway. Cancer Res. 2020;80(21):4655–67. doi:10.1158/0008-5472.CAN-20-1192. [Google Scholar] [PubMed] [CrossRef]
169. Shi R, Liao C, Zhang Q. Hypoxia-driven effects in cancer: characterization, mechanisms, and therapeutic implications. Cells. 2021;10(3):678. doi:10.3390/cells10030678. [Google Scholar] [PubMed] [CrossRef]
170. Zeng Y, Fu BM. Resistance mechanisms of anti-angiogenic therapy and exosomes-mediated revascularization in cancer. Front Cell Dev Biol. 2020;8:610661. doi:10.3389/fcell.2020.610661. [Google Scholar] [PubMed] [CrossRef]
171. Chen Z, Han F, Du Y, Shi H, Zhou W. Hypoxic microenvironment in cancer: molecular mechanisms and therapeutic interventions. Signal Transduct Target Ther. 2023;8(1):70. doi:10.1038/s41392-023-01332-8. [Google Scholar] [PubMed] [CrossRef]
172. Wrona A, Dziadziuszko R, Jassem J. Combining radiotherapy with targeted therapies in non-small cell lung cancer: focus on anti-EGFR, anti-ALK and anti-angiogenic agents. Transl Lung Cancer Res. 2021;10(4):2032–47. doi:10.21037/tlcr-20-552. [Google Scholar] [PubMed] [CrossRef]
173. Woźniak M, Makuch S, Winograd K, Wiśniewski J, Ziółkowski P, Agrawal S. 6-Shogaol enhances the anticancer effect of 5-fluorouracil, oxaliplatin, and irinotecan via increase of apoptosis and autophagy in colon cancer cells in hypoxic/aglycemic conditions. BMC Complement Med Ther. 2020;20(1):141. doi:10.1186/s12906-020-02913-8. [Google Scholar] [PubMed] [CrossRef]
174. Küper A, Baumann J, Göpelt K, Baumann M, Sänger C, Metzen E, et al. Overcoming hypoxia-induced resistance of pancreatic and lung tumor cells by disrupting the PERK-NRF2-HIF-axis. Cell Death Dis. 2021;12(1):82. doi:10.1038/s41419-020-03319-7. [Google Scholar] [PubMed] [CrossRef]
175. Mafi S, Mansoori B, Taeb S, Sadeghi H, Abbasi R, Cho WC, et al. mTOR-mediated regulation of immune responses in cancer and tumor microenvironment. Front Immunol. 2022;12:774103. doi:10.3389/fimmu.2021.774103. [Google Scholar] [PubMed] [CrossRef]
176. Keerthana CK, Rayginia TP, Shifana SC, Anto NP, Kalimuthu K, Isakov N, et al. The role of AMPK in cancer metabolism and its impact on the immunomodulation of the tumor microenvironment. Front Immunol. 2023;14:1114582. doi:10.3389/fimmu.2023.1114582. [Google Scholar] [PubMed] [CrossRef]
177. Moon J, Kitty I, Renata K, Qin S, Zhao F, Kim W. DNA damage and its role in cancer therapeutics. Int J Mol Sci. 2023;24(5):4741. doi:10.3390/ijms24054741. [Google Scholar] [PubMed] [CrossRef]
178. Cao J, Chow L, Dow S. Strategies to overcome myeloid cell induced immune suppression in the tumor microenvironment. Front Oncol. 2023;13:1116016. doi:10.3389/fonc.2023.1116016. [Google Scholar] [PubMed] [CrossRef]
179. Araujo A, Abaurrea A, Azcoaga P, Lpez-Velazco JI, Manzano S, Rodriguez J, et al. Stromal oncostatin m cytokine promotes breast cancer progression by reprogramming the tumor microenvironment. J Clin Investig. 2022;132(7):e148667. doi:10.1172/jci165107. [Google Scholar] [PubMed] [CrossRef]
180. Akkız H. Emerging role of cancer-associated fibroblasts in progression and treatment of hepatocellular carcinoma. Int J Mol Sci. 2023;24(4):3941. doi:10.3390/ijms24043941. [Google Scholar] [PubMed] [CrossRef]
181. Maneshi P, Mason J, Dongre M, Öhlund D. Targeting tumor-stromal interactions in pancreatic cancer: impact of collagens and mechanical traits. Front Cell Dev Biol. 2021;9:787485. doi:10.3389/fcell.2021.787485. [Google Scholar] [PubMed] [CrossRef]
182. Baldari S, Di Modugno F, Nisticò P, Toietta G. Strategies for efficient targeting of tumor collagen for cancer therapy. Cancers. 2022;14(19):4706. doi:10.3390/cancers14194706. [Google Scholar] [PubMed] [CrossRef]
183. Kapor S, Santibanez JF. Myeloid-derived suppressor cells and mesenchymal stem/stromal cells in myeloid malignancies. J Clin Med. 2021;10(13):2788. doi:10.3390/jcm10132788. [Google Scholar] [PubMed] [CrossRef]
184. Malla RR, Vasudevaraju P, Vempati RK, Rakshmitha M, Merchant N, Nagaraju GP. Regulatory T cells: their role in triple-negative breast cancer progression and metastasis. Cancer. 2022;128(6):1171–83. doi:10.1002/cncr.34084. [Google Scholar] [PubMed] [CrossRef]
185. Goenka A, Khan F, Verma B, Sinha P, Dmello CC, Jogalekar MP, et al. Tumor microenvironment signaling and therapeutics in cancer progression. Cancer Commun. 2023;43(5):525–61. doi:10.1002/cac2.12416. [Google Scholar] [PubMed] [CrossRef]
186. Dallavalasa S, Beeraka NM, Basavaraju CG, Tulimilli SV, Sadhu SP, Rajesh K, et al. The role of tumor associated macrophages (TAMs) in cancer progression, chemoresistance, angiogenesis and metastasis—current status. Curr Med Chem. 2021;28(39):8203–36. doi:10.2174/0929867328666210720143721. [Google Scholar] [PubMed] [CrossRef]
187. Li Y, Zhao W, Wang Y, Wang H, Liu S. Extracellular vesicle-mediated crosstalk between pancreatic cancer and stromal cells in the tumor microenvironment. J Nanobiotechnol. 2022;20(1):208. doi:10.1186/s12951-022-01382-0. [Google Scholar] [PubMed] [CrossRef]
188. Yang E, Wang X, Gong Z, Yu M, Wu H, Zhang D. Exosome-mediated metabolic reprogramming: the emerging role in tumor microenvironment remodeling and its influence on cancer progression. Signal Transduct Target Ther. 2020;5(1):242. doi:10.1038/s41392-020-00359-5. [Google Scholar] [PubMed] [CrossRef]
189. Raja R, Wu C, Bassoy EY, Rubino TE Jr, Utagawa EC, Magtibay PM, et al. PP4 inhibition sensitizes ovarian cancer to NK cell-mediated cytotoxicity via STAT1 activation and inflammatory signaling. J Immunother Cancer. 2022;10(12):e005026. doi:10.1136/jitc-2022-005026. [Google Scholar] [PubMed] [CrossRef]
190. Nakagawa Y, Sedukhina AS, Okamoto N, Nagasawa S, Suzuki N, Ohta T, et al. NF-κB signaling mediates acquired resistance after PARP inhibition. Oncotarget. 2015;6(6):3825–39. doi:10.18632/oncotarget.2868. [Google Scholar] [PubMed] [CrossRef]
191. Wilkins AC, Patin EC, Harrington KJ, Melcher AA. The immunological consequences of radiation-induced DNA damage. J Pathol. 2019;247(5):606–14. doi:10.1002/path.5232. [Google Scholar] [PubMed] [CrossRef]
192. Pilger D, Seymour LW, Jackson SP. Interfaces between cellular responses to DNA damage and cancer immunotherapy. Genes Dev. 2021;35(9–10):602–18. doi:10.1101/gad.348314.121. [Google Scholar] [PubMed] [CrossRef]
193. Law AMK, Valdes-Mora F, Gallego-Ortega D. Myeloid-derived suppressor cells as a therapeutic target for cancer. Cells. 2020;9(3):561. doi:10.3390/cells9030561. [Google Scholar] [PubMed] [CrossRef]
194. Lutfi N, Galindo-Campos MA, Yélamos J. Impact of DNA damage response-targeted therapies on the immune response to tumours. Cancers. 2021;13(23):6008. doi:10.3390/cancers13236008. [Google Scholar] [PubMed] [CrossRef]
195. Wu X, Kang X, Zhang X, Xie W, Su Y, Liu X, et al. WEE1 inhibitor and ataxia telangiectasia and RAD3-related inhibitor trigger stimulator of interferon gene-dependent immune response and enhance tumor treatment efficacy through programmed death-ligand 1 blockade. Cancer Sci. 2021;112(11):4444–56. doi:10.1111/cas.15108. [Google Scholar] [PubMed] [CrossRef]
196. Kwon J, Bakhoum SF. The cytosolic DNA-sensing cGAS-STING pathway in cancer. Cancer Discov. 2020;10(1):26–39. doi:10.1158/2159-8290.CD-19-0761. [Google Scholar] [PubMed] [CrossRef]
197. Bruand M, Barras D, Mina M, Ghisoni E, Morotti M, Lanitis E, et al. Cell-autonomous inflammation of BRCA1-deficient ovarian cancers drives both tumor-intrinsic immunoreactivity and immune resistance via STING. Cell Rep. 2021;36(3):109412. doi:10.1016/j.celrep.2021.109412. [Google Scholar] [PubMed] [CrossRef]
198. Tong J, Song J, Zhang W, Zhai J, Guan Q, Wang H, et al. When DNA-damage responses meet innate and adaptive immunity. Cell Mol Life Sci. 2024;81(1):185. doi:10.1007/s00018-024-05214-2. [Google Scholar] [PubMed] [CrossRef]
199. Chen G, Zhou S, Sun C, Han L. Editorial: the impact of the DNA damage response on anti-tumor immunity. Front Immunol. 2022;13:1026194. doi:10.3389/fimmu.2022.1026194. [Google Scholar] [PubMed] [CrossRef]
200. Carlsen L, El-Deiry WS. Anti-cancer immune responses to DNA damage response inhibitors: molecular mechanisms and progress toward clinical translation. Front Oncol. 2022;12:998388. doi:10.3389/fonc.2022.998388. [Google Scholar] [PubMed] [CrossRef]
201. Krysko DV, Garg AD, Kaczmarek A, Krysko O, Agostinis P, Vandenabeele P. Immunogenic cell death and DAMPs in cancer therapy. Nat Rev Cancer. 2012;12(12):860–75. doi:10.1038/nrc3380. [Google Scholar] [PubMed] [CrossRef]
202. Solari JIG, Filippi-Chiela E, Pilar ES, Nunes V, Gonzalez EA, Figueiró F, et al. Damage-associated molecular patterns (DAMPs) related to immunogenic cell death are differentially triggered by clinically relevant chemotherapeutics in lung adenocarcinoma cells. BMC Cancer. 2020;20(1):474. doi:10.1186/s12885-020-06964-5. [Google Scholar] [PubMed] [CrossRef]
203. Hernandez C, Huebener P, Schwabe RF. Damage-associated molecular patterns in cancer: a double-edged sword. Oncogene. 2016;35(46):5931–41. doi:10.1038/onc.2016.104. [Google Scholar] [PubMed] [CrossRef]
204. Kim JM, Chen DS. Immune escape to PD-L1/PD-1 blockade: seven steps to success (or failure). Ann Oncol. 2016;27(8):1492–504. doi:10.1093/annonc/mdw217. [Google Scholar] [PubMed] [CrossRef]
205. Briukhovetska D, Dörr J, Endres S, Libby P, Dinarello CA, Kobold S. Interleukins in cancer: from biology to therapy. Nat Rev Cancer. 2021;21(8):481–99. doi:10.1038/s41568-021-00363-z. [Google Scholar] [PubMed] [CrossRef]
206. Zhu L, Liu J, Chen J, Zhou Q. The developing landscape of combinatorial therapies of immune checkpoint blockade with DNA damage repair inhibitors for the treatment of breast and ovarian cancers. J Hematol Oncol. 2021;14(1):206. doi:10.1186/s13045-021-01218-8. [Google Scholar] [PubMed] [CrossRef]
207. Oliver AJ, Lau PKH, Unsworth AS, Loi S, Darcy PK, Kershaw MH, et al. Tissue-dependent tumor microenvironments and their impact on immunotherapy responses. Front Immunol. 2018;9:70. doi:10.3389/fimmu.2018.00070. [Google Scholar] [PubMed] [CrossRef]
208. Ye Z, Shi Y, Lees-Miller SP, Tainer JA. Function and molecular mechanism of the DNA damage response in immunity and cancer immunotherapy. Front Immunol. 2021;12:797880. doi:10.3389/fimmu.2021.797880. [Google Scholar] [PubMed] [CrossRef]
209. Bigos KJ, Quiles CG, Lunj S, Smith DJ, Krause M, Troost EG, et al. Tumour response to hypoxia: understanding the hypoxic tumour microenvironment to improve treatment outcome in solid tumours. Front Oncol. 2024;14:1331355. doi:10.3389/fonc.2024.1331355. [Google Scholar] [PubMed] [CrossRef]
210. Begg K, Tavassoli M. Inside the hypoxic tumour: reprogramming of the DDR and radioresistance. Cell Death Discov. 2020;6(1):77. doi:10.1038/s41420-020-00311-0. [Google Scholar] [PubMed] [CrossRef]
211. Jurkovicova D, Neophytou CM, Gašparović AČ, Gonçalves AC. DNA damage response in cancer therapy and resistance: challenges and opportunities. Int J Mol Sci. 2022;23(23):14672. doi:10.3390/ijms232314672. [Google Scholar] [PubMed] [CrossRef]
212. Minchom A, Aversa C, Lopez J. Dancing with the DNA damage response: next-generation anti-cancer therapeutic strategies. Ther Adv Med Oncol. 2018;10:1–18. doi:10.1177/1758835918786658. [Google Scholar] [PubMed] [CrossRef]
213. Ramkumar K, Stewart CA, Cargill KR, Della Corte CM, Wang Q, Shen L, et al. AXL inhibition induces DNA damage and replication stress in non-small cell lung cancer cells and promotes sensitivity to ATR inhibitors. Mol Cancer Res. 2020;19(3):485–97. doi:10.1158/1541-7786.MCR-20-0414. [Google Scholar] [PubMed] [CrossRef]
214. Brown KK, Spinelli JB, Asara JM, Toker A. Adaptive reprogramming of de novo pyrimidine synthesis is a metabolic vulnerability in triple-negative breast cancer. Cancer Discov. 2017;7(4):391–9. doi:10.1158/2159-8290.CD-16-0611. [Google Scholar] [PubMed] [CrossRef]
215. Chou YT, Lin CY, Wen JW, Hung LC, Chang YF, Yang CM, et al. Targeting triple-negative breast cancer with an aptamer-functionalized nanoformulation: a synergistic treatment that combines photodynamic and bioreductive therapies. J Nanobiotechnol. 2021;19(1):89. doi:10.1186/s12951-021-00786-8. [Google Scholar] [PubMed] [CrossRef]
216. Hunter FW, Wouters BG, Wilson WR. Hypoxia-activated prodrugs: paths forward in the era of personalised medicine. Br J Cancer. 2016;114(10):1071–7. doi:10.1038/bjc.2016.79. [Google Scholar] [PubMed] [CrossRef]
217. Liu R, Yang F, Yin JY, Liu YZ, Zhang W, Zhou HH. Influence of tumor immune infiltration on immune checkpoint inhibitor therapeutic efficacy: a computational retrospective study. Front Immunol. 2021;12:685370. doi:10.3389/fimmu.2021.685370. [Google Scholar] [PubMed] [CrossRef]
218. Salman S, Meyers DJ, Wicks EE, Lee SN, Datan E, Thomas AM, et al. HIF inhibitor 32-134D eradicates murine hepatocellular carcinoma in combination with anti-PD1 therapy. J Clin Invest. 2022;132(9):e156774. doi:10.1172/JCI156774. [Google Scholar] [PubMed] [CrossRef]
219. Abou Khouzam R, Brodaczewska K, Filipiak A, Zeinelabdin NA, Buart S, Szczylik C, et al. Tumor hypoxia regulates immune escape/invasion:influence on angiogenesis and potential impact of hypoxic biomarkers on cancer therapies. Front Immunol. 2021;11:613114. doi:10.3389/fimmu.2020.613114. [Google Scholar] [PubMed] [CrossRef]
220. Alharbi M, Lai A, Sharma S, Kalita-de Croft P, Godbole N, Campos A, et al. Extracellular vesicle transmission of chemoresistance to ovarian cancer cells is associated with hypoxia-induced expression of glycolytic pathway proteins, and prediction of epithelial ovarian cancer disease recurrence. Cancers. 2021;13(14):3388. doi:10.3390/cancers13143388. [Google Scholar] [PubMed] [CrossRef]
221. Owari T, Tanaka N, Nakai Y, Miyake M, Anai S, Kishi S, et al. 5-Aminolevulinic acid overcomes hypoxia-induced radiation resistance by enhancing mitochondrial reactive oxygen species production in prostate cancer cells. Br J Cancer. 2022;127(2):350–63. doi:10.1038/s41416-022-01789-4. [Google Scholar] [PubMed] [CrossRef]
222. Saatci O, Kaymak A, Raza U, Ersan PG, Akbulut O, Banister CE, et al. Targeting lysyl oxidase (LOX) overcomes chemotherapy resistance in triple negative breast cancer. Nat Commun. 2020;11(1):2416. doi:10.1038/s41467-020-16199-4. [Google Scholar] [PubMed] [CrossRef]
223. Nasri D, Manwar R, Kaushik A, Er EE, Avanaki K. Photoacoustic imaging for investigating tumor hypoxia: a strategic assessment. Theranostics. 2023;13(10):3346–67. doi:10.7150/thno.84253. [Google Scholar] [PubMed] [CrossRef]
224. Cisneros-Villanueva M, Hidalgo-Pérez L, Rios-Romero M, Cedro-Tanda A, Ruiz-Villavicencio CA, Page K, et al. Cell-free DNA analysis in current cancer clinical trials: a review. Br J Cancer. 2022;126(3):391–400. doi:10.1038/s41416-021-01696-0. [Google Scholar] [PubMed] [CrossRef]
225. Cha YJ, Koo JS. Role of tumor-associated myeloid cells in breast cancer. Cells. 2020;9(8):1785. doi:10.3390/cells9081785. [Google Scholar] [PubMed] [CrossRef]
226. Wu Y, Xu S, Cheng S, Yang J, Wang Y. Clinical application of PARP inhibitors in ovarian cancer: from molecular mechanisms to the current status. J Ovarian Res. 2023;16(1):6. doi:10.1186/s13048-023-01094-5. [Google Scholar] [PubMed] [CrossRef]
227. Baxter JS, Zatreanu D, Pettitt SJ, Lord CJ. Resistance to DNA repair inhibitors in cancer. Mol Oncol. 2022;16(21):3811–27. doi:10.1002/1878-0261.13224. [Google Scholar] [PubMed] [CrossRef]
228. Shi C, Qin K, Lin A, Jiang A, Cheng Q, Liu Z, et al. The role of DNA damage repair (DDR) system in response to immune checkpoint inhibitor (ICI) therapy. J Exp Clin Cancer Res. 2022;41(1):268. doi:10.1186/s13046-022-02469-0. [Google Scholar] [PubMed] [CrossRef]
229. Amodio V, Mauri G, Reilly NM, Sartore-Bianchi A, Siena S, Bardelli A, et al. Mechanisms of immune escape and resistance to checkpoint inhibitor therapies in mismatch repair deficient metastatic colorectal cancers. Cancers. 2021;13(11):2638. doi:10.3390/cancers13112638. [Google Scholar] [PubMed] [CrossRef]
230. Allen GM, Frankel NW, Reddy NR, Bhargava HK, Yoshida MA, Stark SR, et al. Synthetic cytokine circuits that drive T cells into immune-excluded tumors. Science. 2022;378(6625):eaba1624. doi:10.1126/science.aba1624. [Google Scholar] [PubMed] [CrossRef]
231. Concannon K, Morris BB, Gay CM, Byers LA. Combining targeted DNA repair inhibition and immune-oncology approaches for enhanced tumor control. Mol Cell. 2023;83(5):660–80. doi:10.1016/j.molcel.2022.12.031. [Google Scholar] [PubMed] [CrossRef]
232. Sheng H, Huang Y, Xiao Y, Zhu Z, Shen M, Zhou P, et al. ATR inhibitor AZD6738 enhances the antitumor activity of radiotherapy and immune checkpoint inhibitors by potentiating the tumor immune microenvironment in hepatocellular carcinoma. J Immunother Cancer. 2020;8(1):e000340. doi:10.1136/jitc-2019-000340. [Google Scholar] [PubMed] [CrossRef]
233. Vendetti FP, Karukonda P, Clump DA, Teo T, Lalonde R, Nugent K, et al. ATR kinase inhibitor AZD6738 potentiates CD8+ T cell-dependent antitumor activity following radiation. J Clin Invest. 2018;128(9):3926–40. doi:10.1172/JCI96519. [Google Scholar] [PubMed] [CrossRef]
234. Wang Q, Bergholz JS, Ding L, Lin Z, Kabraji SK, Hughes ME, et al. STING agonism reprograms tumor-associated macrophages and overcomes resistance to PARP inhibition in BRCA1-deficient models of breast cancer. Nat Commun. 2022;13(1):3022. doi:10.1038/s41467-022-30568-1. [Google Scholar] [PubMed] [CrossRef]
235. Sen T, Rodriguez BL, Chen L, Della Corte CM, Morikawa N, Fujimoto J, et al. Targeting DNA damage response promotes antitumor immunity through STING-mediated T-cell activation in small cell lung cancer. Cancer Discov. 2019;9(5):646–61. doi:10.1158/2159-8290.CD-18-1020. [Google Scholar] [PubMed] [CrossRef]
236. Ran X, Wu BX, Vidhyasagar V, Song L, Zhang X, Ladak RJ, et al. PARP inhibitor radiosensitization enhances anti-PD-L1 immunotherapy through stabilizing chemokine mRNA in small cell lung cancer. Nat Commun. 2025;16(1):2166. doi:10.1038/s41467-025-57257-z. [Google Scholar] [PubMed] [CrossRef]
237. Liu S, Deng P, Yu Z, Hong JH, Gao J, Huang Y, et al. CDC7 inhibition potentiates antitumor efficacy of PARP inhibitor in advanced ovarian cancer. Adv Sci. 2024;11(45):2403782. doi:10.1002/advs.202403782. [Google Scholar] [PubMed] [CrossRef]
238. Chen L, Lin J, Wen Y, Guo ZQ, Lan B, Xiong J, et al. DNA-PKcs dysfunction enhances the antitumor activity of radioimmunotherapy by activating the cGAS-STING pathway in HNSCC. J Inflamm Res. 2025;18:4177–93. doi:10.2147/JIR.S497295. [Google Scholar] [PubMed] [CrossRef]
239. Porta C, Paglino C, Mosca A. Targeting PI3K/Akt/mTOR signaling in cancer. Front Oncol. 2014;4:64. doi:10.3389/fonc.2014.00064. [Google Scholar] [PubMed] [CrossRef]
240. Bukowski K, Kciuk M, Kontek R. Mechanisms of multidrug resistance in cancer chemotherapy. Int J Mol Sci. 2020;21(9):3233. doi:10.3390/ijms21093233. [Google Scholar] [PubMed] [CrossRef]
241. Das KP, Chandra J. Nanoparticles and convergence of artificial intelligence for targeted drug delivery for cancer therapy: current progress and challenges. Front Med Technol. 2023;4:1067144. doi:10.3389/fmedt.2022.1067144. [Google Scholar] [PubMed] [CrossRef]
242. Zheng M, Gong P, Zheng C, Zhao P, Luo Z, Ma Y, et al. Lipid-polymer nanoparticles for folate-receptor targeting delivery of doxorubicin. J Nanosci Nanotechnol. 2015;15(7):4792–8. doi:10.1166/jnn.2015.9604. [Google Scholar] [PubMed] [CrossRef]
243. Wagner J, Gößl D, Ustyanovska N, Xiong M, Hauser D, Zhuzhgova O, et al. Mesoporous silica nanoparticles as pH-responsive carrier for the immune-activating drug resiquimod enhance the local immune response in mice. ACS Nano. 2021;15(3):4450–66. doi:10.1021/acsnano.0c08384. [Google Scholar] [PubMed] [CrossRef]
244. He Y, Fan X, Wu X, Hu T, Zhou F, Tan S, et al. pH-Responsive size-shrinkable mesoporous silica-based nanocarriers for improving tumor penetration and therapeutic efficacy. Nanoscale. 2022;14(4):1271–84. doi:10.1039/d1nr07513f. [Google Scholar] [PubMed] [CrossRef]
245. Gou K, Xin W, Lv J, Ma Z, Yang J, Zhao L, et al. A pH-responsive chiral mesoporous silica nanoparticles for delivery of doxorubicin in tumor-targeted therapy. Colloids Surf B Biointerfaces. 2023;221:113027. doi:10.1016/j.colsurfb.2022.113027. [Google Scholar] [PubMed] [CrossRef]
246. Baldwin P, Ohman A, Thong J, Tangutoori S, Dinulescu D, Sridhar S. Abstract A03: PARP inhibitor nanotherapy for ovarian cancer. Clin Cancer Res. 2016;22(2_Suppl):A03. doi:10.1158/1557-3265.ovca15-a03. [Google Scholar] [CrossRef]
247. Baldwin P, Ohman AW, Tangutoori S, Dinulescu DM, Sridhar S. Intraperitoneal delivery of NanoOlaparib for disseminated late-stage cancer treatment. Int J Nanomed. 2018;13:8063–74. doi:10.2147/IJN.S186881. [Google Scholar] [PubMed] [CrossRef]
248. Singh B, Yang S, Krishna A, Sridhar S. Nanoparticle formulations of poly (ADP-ribose) polymerase inhibitors for cancer therapy. Front Chem. 2020;8:594619. doi:10.3389/fchem.2020.594619. [Google Scholar] [PubMed] [CrossRef]
249. Gupta M, Iyer R, Fountzilas C. Poly(ADP-ribose) polymerase inhibitors in pancreatic cancer: a new treatment paradigms and future implications. Cancers. 2019;11(12):1980. doi:10.3390/cancers11121980. [Google Scholar] [PubMed] [CrossRef]
250. Andreana I, Repellin M, Carton F, Kryza D, Briançon S, Chazaud B, et al. Nanomedicine for gene delivery and drug repurposing in the treatment of muscular dystrophies. Pharmaceutics. 2021;13(2):278. doi:10.3390/pharmaceutics13020278. [Google Scholar] [PubMed] [CrossRef]
251. Lin Z, Chou WC, Cheng YH, He C, Monteiro-Riviere NA, Riviere JE. Predicting nanoparticle delivery to tumors using machine learning and artificial intelligence approaches. Int J Nanomed. 2022;17:1365–79. doi:10.2147/IJN.S344208. [Google Scholar] [PubMed] [CrossRef]
252. Sun L, Liu H, Ye Y, Lei Y, Islam R, Tan S, et al. Smart nanoparticles for cancer therapy. Signal Transduct Target Ther. 2023;8(1):418. doi:10.1038/s41392-023-01642-x. [Google Scholar] [PubMed] [CrossRef]
253. Ingber DE. Human organs-on-chips for disease modelling, drug development and personalized medicine. Nat Rev Genet. 2022;23(8):467–91. doi:10.1038/s41576-022-00466-9. [Google Scholar] [PubMed] [CrossRef]
254. Sean G, Banes AJ, Gangaraju R. Organoids and tissue/organ chips. Stem Cell Res Ther. 2024;15(1):241. doi:10.1186/s13287-024-03859-1. [Google Scholar] [PubMed] [CrossRef]
255. Dong WY, Körber M, López Esguerra V, Bodmeier R. Stability of poly(D,L-lactide-co-glycolide) and leuprolide acetate in in-situ forming drug delivery systems. J Control Release. 2006;115(2):158–67. doi:10.1016/j.jconrel.2006.07.013. [Google Scholar] [PubMed] [CrossRef]
256. Tomao S. Albumin-bound formulation of paclitaxel (Abraxane® ABI-007) in the treatment of breast cancer. Int J Nanomed. 2009;4:99–105. doi:10.2147/ijn.s3061. [Google Scholar] [PubMed] [CrossRef]
257. Gu L, Shi T, Sun Y, You C, Wang S, Wen G, et al. Folate-modified, indocyanine green-loaded lipid-polymer hybrid nanoparticles for targeted delivery of cisplatin. J Biomater Sci Polym Ed. 2017;28(7):690–702. doi:10.1080/09205063.2017.1296347. [Google Scholar] [PubMed] [CrossRef]
258. Werner ME, Karve S, Sukumar R, Cummings ND, Copp JA, Chen RC, et al. Folate-targeted nanoparticle delivery of chemo- and radiotherapeutics for the treatment of ovarian cancer peritoneal metastasis. Biomaterials. 2011;32(33):8548–54. doi:10.1016/j.biomaterials.2011.07.067. [Google Scholar] [PubMed] [CrossRef]
259. Verma S, Miles D, Gianni L, Krop IE, Welslau M, Baselga J, et al. Trastuzumab emtansine for HER2-positive advanced breast cancer. N Engl J Med. 2012;367(19):1783–91. doi:10.1056/NEJMoa1209124. [Google Scholar] [PubMed] [CrossRef]
260. Leone J, Recondo G, de la Vega M, Galanternik F, Díaz-Cantón E, Leone BA. Novel approaches to target HER2-positive breast cancer: trastuzumab emtansine. Cancer Manag Res. 2016;8:57. doi:10.2147/cmar.s104447. [Google Scholar] [PubMed] [CrossRef]
261. Arruebo M, Fernández-Pacheco R, Ibarra MR, Santamaría J. Magnetic nanoparticles for drug delivery. Nano Today. 2007;2(3):22–32. doi:10.1016/S1748-0132(07)70084-1. [Google Scholar] [CrossRef]
262. Yahya S, Pushpanathan S, Jan S, Chaudhary N, Parray R, Gandhi KA, et al. Development and characterization of pegylated Fe3O4-CAPE magnetic nanoparticles for targeted therapy and hyperthermia treatment of colorectal cancer. Sci Rep. 2025;15(1):26008. doi:10.1038/s41598-025-11927-6. [Google Scholar] [PubMed] [CrossRef]
263. Anwer MK, Ali EA, Iqbal M, Ahmed MM, Aldawsari MF, Al Saqr A, et al. Development of chitosan-coated PLGA-based nanoparticles for improved oral olaparib delivery: in vitro characterization, and in vivo pharmacokinetic studies. Processes. 2022;10(7):1329. doi:10.3390/pr10071329. [Google Scholar] [CrossRef]
264. Gabizon A, Shmeeda H, Barenholz Y. Pharmacokinetics of pegylated liposomal Doxorubicin: review of animal and human studies. Clin Pharmacokinet. 2003;42(5):419–36. doi:10.2165/00003088-200342050-00002. [Google Scholar] [PubMed] [CrossRef]
265. Matsumura Y, Kataoka K. Preclinical and clinical studies of anticancer agent-incorporating polymer micelles. Cancer Sci. 2009;100(4):572–9. doi:10.1111/j.1349-7006.2009.01103.x. [Google Scholar] [PubMed] [CrossRef]
266. Serras A, Faustino C, Pinheiro L. Functionalized polymeric micelles for targeted cancer therapy: steps from conceptualization to clinical trials. Pharmaceutics. 2024;16(8):1047. doi:10.3390/pharmaceutics16081047. [Google Scholar] [PubMed] [CrossRef]
267. Cheng SH, Liao WN, Chen LM, Lee CH. pH-controllable release using functionalized mesoporous silica nanoparticles as an oral drug delivery system. J Mater Chem. 2011;21(20):7130–7. doi:10.1039/c0jm04490c. [Google Scholar] [CrossRef]
268. Feng R, Chen Q, Zhou P, Wang Y, Yan H. Nanoparticles based on disulfide-containing poly(β-amino ester) and zwitterionic fluorocarbon surfactant as a redox-responsive drug carrier for brain tumor treatment. Nanotechnology. 2018;29(49):495101. doi:10.1088/1361-6528/aae122. [Google Scholar] [PubMed] [CrossRef]
269. Iaccarino G, Profeta M, Vecchione R, Netti PA. Matrix metalloproteinase-cleavable nanocapsules for tumor-activated drug release. Acta Biomater. 2019;89(4):265–78. doi:10.1016/j.actbio.2019.02.043. [Google Scholar] [PubMed] [CrossRef]
270. Futaki S, Suzuki T, Ohashi W, Yagami T, Tanaka S, Ueda K, et al. Arginine-rich peptides. An abundant source of membrane-permeable peptides having potential as carriers for intracellular protein delivery. J Biol Chem. 2001;276(8):5836–40. doi:10.1074/jbc.M007540200. [Google Scholar] [PubMed] [CrossRef]
271. Walter M, Bresinsky M, Zimmer O, Pockes S, Goepferich A. Conditional cell-penetrating peptide exposure as selective nanoparticle uptake signal. ACS Appl Mater Interfaces. 2024;16(29):37734–47. doi:10.1021/acsami.4c07821. [Google Scholar] [PubMed] [CrossRef]
272. Chen Y, Sen J, Bathula SR, Yang Q, Fittipaldi R, Huang L. Novel cationic lipid that delivers siRNA and enhances therapeutic effect in lung cancer cells. Mol Pharm. 2009;6(3):696–705. doi:10.1021/mp800136v. [Google Scholar] [PubMed] [CrossRef]
273. Gu W, Fan R, Quan J, Cheng Y, Wang S, Zhang H, et al. Intracranial in situ thermosensitive hydrogel delivery of temozolomide accomplished by PLGA-PEG–PLGA triblock copolymer blending for GBM treatment. Polymers. 2022;14(16):3368. doi:10.3390/polym14163368. [Google Scholar] [PubMed] [CrossRef]
274. Kamaly N, Yameen B, Wu J, Farokhzad OC. Degradable controlled-release polymers and polymeric nanoparticles: mechanisms of controlling drug release. Chem Rev. 2016;116(4):2602–63. doi:10.1021/acs.chemrev.5b00346. [Google Scholar] [PubMed] [CrossRef]
275. Lerchbammer-Kreith Y, Hejl M, Vician P, Jakupec MA, Berger W, Galanski MS, et al. Combination of drug delivery properties of PAMAM dendrimers and cytotoxicity of platinum(IV) complexes—a more selective anticancer treatment? Pharmaceutics. 2023;15(5):1515. doi:10.3390/pharmaceutics15051515. [Google Scholar] [PubMed] [CrossRef]
276. Rapoport NY, Kennedy AM, Shea JE, Scaife CL, Nam KH. Controlled and targeted tumor chemotherapy by ultrasound-activated nanoemulsions/microbubbles. J Control Release. 2009;138(3):268–76. doi:10.1016/j.jconrel.2009.05.026. [Google Scholar] [PubMed] [CrossRef]
277. Goldberg MS, Xing D, Ren Y, Orsulic S, Bhatia SN, Sharp PA. Nanoparticle-mediated delivery of siRNA targeting Parp1 extends survival of mice bearing tumors derived from Brca1-deficient ovarian cancer cells. Proc Natl Acad Sci U S A. 2011;108(2):745–50. doi:10.1073/pnas.1016538108. [Google Scholar] [PubMed] [CrossRef]
278. Guney Eskiler G, Cecener G, Egeli U, Tunca B. Synthetically lethal BMN 673 (talazoparib) loaded solid lipid nanoparticles for BRCA1 mutant triple negative breast cancer. Pharm Res. 2018;35(11):218. doi:10.1007/s11095-018-2502-6. [Google Scholar] [PubMed] [CrossRef]
279. Sanson C, Diou O, Thévenot J, Ibarboure E, Soum A, Brûlet A, et al. Doxorubicin loaded magnetic polymersomes: theranostic nanocarriers for MR imaging and magneto-chemotherapy. ACS Nano. 2011;5(2):1122–40. doi:10.1021/nn102762f. [Google Scholar] [PubMed] [CrossRef]
280. Rahman MA, Jalouli M, Yadab MK, Al-Zharani M. Progress in drug delivery systems based on nanoparticles for improved glioblastoma therapy: addressing challenges and investigating opportunities. Cancers. 2025;17(4):701. doi:10.3390/cancers17040701. [Google Scholar] [PubMed] [CrossRef]
281. Thomas A, Takahashi N, O’Connor O, Redon L, Mohindroo CE, Sciuto C, et al. Tumor-targeted top1 inhibitor delivery with optimized parp inhibition in advanced solid tumors: a phase I trial of gapped scheduling. Nat Commun. 2025;16(1):9457. doi:10.1038/s41467-025-64509-5. [Google Scholar] [PubMed] [CrossRef]
282. Saberian E, Jenčová J, Jenča A, Jenča A, Salehipoor F, Zare-Zardini H, et al. Bleomycin-loaded folic acid-conjugated nanoliposomes: a novel formulation for targeted treatment of oral cancer. Front Bioeng Biotechnol. 2025;13:1535793. doi:10.3389/fbioe.2025.1535793. [Google Scholar] [PubMed] [CrossRef]
283. Beck-Broichsitter M, Nicolas J, Couvreur P. Solvent selection causes remarkable shifts of the “Ouzo region” for poly(lactide-co-glycolide) nanoparticles prepared by nanoprecipitation. Nanoscale. 2015;7(20):9215–21. doi:10.1039/c5nr01695a. [Google Scholar] [PubMed] [CrossRef]
284. Paliwal R, Babu RJ, Palakurthi S. Nanomedicine scale-up technologies: feasibilities and challenges. AAPS PharmSciTech. 2014;15(6):1527–34. doi:10.1208/s12249-014-0177-9. [Google Scholar] [PubMed] [CrossRef]
285. Mehta M, Bui TA, Yang X, Aksoy Y, Goldys EM, Deng W. Lipid-based nanoparticles for drug/gene delivery: an overview of the production techniques and difficulties encountered in their industrial development. ACS Mater Au. 2023;3(6):600–19. doi:10.1021/acsmaterialsau.3c00032. [Google Scholar] [PubMed] [CrossRef]
286. Filippov SK, Khusnutdinov R, Murmiliuk A, Inam W, Zakharova LY, Zhang H, et al. Dynamic light scattering and transmission electron microscopy in drug delivery: a roadmap for correct characterization of nanoparticles and interpretation of results. Mater Horiz. 2023;10(12):5354–70. doi:10.1039/d3mh00717k. [Google Scholar] [PubMed] [CrossRef]
287. Bender V, Fuchs L, Süss R. RP-HPLC-CAD method for the rapid analysis of lipids used in lipid nanoparticles derived from dual centrifugation. Int J Pharm X. 2024;7:100255. doi:10.1016/j.ijpx.2024.100255. [Google Scholar] [PubMed] [CrossRef]
288. Blanco E, Shen H, Ferrari M. Principles of nanoparticle design for overcoming biological barriers to drug delivery. Nat Biotechnol. 2015;33(9):941–51. doi:10.1038/nbt.3330. [Google Scholar] [PubMed] [CrossRef]
289. Gustafson HH, Holt-Casper D, Grainger DW, Ghandehari H. Nanoparticle uptake: the phagocyte problem. Nano Today. 2015;10(4):487–510. doi:10.1016/j.nantod.2015.06.006. [Google Scholar] [PubMed] [CrossRef]
290. Zelepukin IV, Yaremenko AV, Yuryev MV, Mirkasymov AB, Sokolov IL, Deyev SM, et al. Fast processes of nanoparticle blood clearance: comprehensive study. J Control Release. 2020;326:181–91. doi:10.1016/j.jconrel.2020.07.014. [Google Scholar] [PubMed] [CrossRef]
291. Li X, Hu Y, Zhang X, Shi X, Parak WJ, Pich A. Transvascular transport of nanocarriers for tumor delivery. Nat Commun. 2024;15(1):8172. doi:10.1038/s41467-024-52416-0. [Google Scholar] [PubMed] [CrossRef]
292. Suk JS, Xu Q, Kim N, Hanes J, Ensign LM. PEGylation as a strategy for improving nanoparticle-based drug and gene delivery. Adv Drug Deliv Rev. 2016;99(Pt A):28–51. doi:10.1016/j.addr.2015.09.012. [Google Scholar] [PubMed] [CrossRef]
293. Becicka WM, Bielecki PA, Lorkowski ME, Moon TJ, Zhang Y, Atukorale PU, et al. The effect of PEGylation on the efficacy and uptake of an immunostimulatory nanoparticle in the tumor immune microenvironment. Nanoscale Adv. 2021;3(17):4961–72. doi:10.1039/d1na00308a. [Google Scholar] [PubMed] [CrossRef]
294. Ryu S, Ohuchi M, Yagishita S, Shimoi T, Yonemori K, Tamura K, et al. Visualization of the distribution of nanoparticle-formulated AZD2811 in mouse tumor model using matrix-assisted laser desorption ionization mass spectrometry imaging. Sci Rep. 2020;10(1):15535. doi:10.1038/s41598-020-72665-5. [Google Scholar] [PubMed] [CrossRef]
295. Xue J, Liu H, Chen S, Xiong C, Zhan L, Sun J, et al. Mass spectrometry imaging of the in situ drug release from nanocarriers. Sci Adv. 2018;4(10):eaat9039. doi:10.1126/sciadv.aat9039. [Google Scholar] [PubMed] [CrossRef]
296. Jakic K, Selc M, Razga F, Nemethova V, Mazancova P, Havel F, et al. Long-term accumulation, biological effects and toxicity of BSA-coated gold nanoparticles in the mouse liver, spleen, and kidneys. Int J Nanomed. 2024;19:4103–20. doi:10.2147/IJN.S443168. [Google Scholar] [PubMed] [CrossRef]
297. Verhoef JJF, Anchordoquy TJ. Questioning the use of PEGylation for drug delivery. Drug Deliv Transl Res. 2013;3(6):499–503. doi:10.1007/s13346-013-0176-5. [Google Scholar] [PubMed] [CrossRef]
298. Mansouri M, Lam J, Sung KE. Progress in developing microphysiological systems for biological product assessment. Lab Chip. 2024;24(5):1293–306. doi:10.1039/d3lc00876b. [Google Scholar] [PubMed] [CrossRef]
299. Liu Y, Liang Y, Jing Y, Xin P, Han JL, Du Y, et al. Advances in nanotechnology for enhancing the solubility and bioavailability of poorly soluble drugs. Drug Des Devel Ther. 2024;18:1469–95. doi:10.2147/DDDT.S447496. [Google Scholar] [PubMed] [CrossRef]
300. Lu Y, Kim S, Park K. In vitro-in vivo correlation: perspectives on model development. Int J Pharm. 2011;418(1):142–8. doi:10.1016/j.ijpharm.2011.01.010. [Google Scholar] [PubMed] [CrossRef]
301. Aloss K, Hamar P. Recent preclinical and clinical progress in liposomal doxorubicin. Pharmaceutics. 2023;15(3):893. doi:10.3390/pharmaceutics15030893. [Google Scholar] [PubMed] [CrossRef]
302. Solomon AD, Dabral S, Brajesh RG, Day BW, Juric M, Zielonka J, et al. Understanding the mechanisms of chemotherapy-related cardiotoxicity employing hiPSC-derived cardiomyocyte models for drug screening and the identification of genetic and epigenetic variants. Int J Mol Sci. 2025;26(9):3966. doi:10.3390/ijms26093966. [Google Scholar] [PubMed] [CrossRef]
303. Kundranda MN, Niu J. Albumin-bound paclitaxel in solid tumors: clinical development and future directions. Drug Des Dev Ther. 2015;9:3767–77. doi:10.2147/DDDT.S88023. [Google Scholar] [PubMed] [CrossRef]
304. Gupta A, De Jesus-Acosta A, Zheng L, Lee V, Kamel I, Le D, et al. Clinical outcomes of liposomal irinotecan in advanced pancreatic adenocarcinoma patients previously treated with conventional irinotecan-based chemotherapy: a real-world study. Front Oncol. 2023;13:1250136. doi:10.3389/fonc.2023.1250136. [Google Scholar] [PubMed] [CrossRef]
305. Li Q, Qian W, Zhang Y, Hu L, Chen S, Xia Y. A new wave of innovations within the DNA damage response. Signal Transduct Target Ther. 2023;8(1):338. doi:10.1038/s41392-023-01548-8. [Google Scholar] [PubMed] [CrossRef]
306. Criscuolo D, Morra F, Giannella R, Cerrato A, Celetti A. Identification of novel biomarkers of homologous recombination defect in DNA repair to predict sensitivity of prostate cancer cells to PARP-inhibitors. Int J Mol Sci. 2019;20(12):3100. doi:10.3390/ijms20123100. [Google Scholar] [PubMed] [CrossRef]
307. Lord CJ, Ashworth A. BRCAness revisited. Nat Rev Cancer. 2016;16(2):110–20. doi:10.1038/nrc.2015.21. [Google Scholar] [PubMed] [CrossRef]
308. Kornepati AVR, Boyd JT, Murray CE, Saifetiarova J, de la Peña Avalos B, Rogers CM, et al. Tumor intrinsic PD-L1 promotes DNA repair in distinct cancers and suppresses PARP inhibitor-induced synthetic lethality. Cancer Res. 2022;82(11):2156–70. doi:10.1158/0008-5472.CAN-21-2076. [Google Scholar] [PubMed] [CrossRef]
309. Willis NA, Chandramouly G, Huang B, Kwok A, Follonier C, Deng C, et al. BRCA1 controls homologous recombination at Tus/Ter-stalled mammalian replication forks. Nature. 2014;510(7506):556–9. doi:10.1038/nature13295. [Google Scholar] [PubMed] [CrossRef]
310. Gunn A, Stark JM. I-SceI-based assays to examine distinct repair outcomes of mammalian chromosomal double strand breaks. Methods Mol Biol. 2012;920:379–91. doi:10.1007/978-1-61779-998-3_27. [Google Scholar] [PubMed] [CrossRef]
311. Li SY, Cheng QX, Wang JM, Li XY, Zhang ZL, Gao S, et al. CRISPR-Cas12a-assisted nucleic acid detection. Cell Discov. 2018;4(1):20. doi:10.1038/s41421-018-0028-z. [Google Scholar] [PubMed] [CrossRef]
312. Stadtmauer EA, Fraietta JA, Davis MM, Cohen AD, Weber KL, Lancaster E, et al. CRISPR-engineered T cells in patients with refractory cancer. Science. 2020;367(6481):eaba7365. doi:10.1126/science.aba7365. [Google Scholar] [PubMed] [CrossRef]
313. Li H, Yang Y, Hong W, Huang M, Wu M, Zhao X. Applications of genome editing technology in the targeted therapy of human diseases: mechanisms, advances and prospects. Signal Transduct Target Ther. 2020;5(1):1. doi:10.1038/s41392-019-0089-y. [Google Scholar] [PubMed] [CrossRef]
314. Bettegowda C, Sausen M, Leary RJ, Kinde I, Wang Y, Agrawal N, et al. Detection of circulating tumor DNA in early- and late-stage human malignancies. Sci Transl Med. 2014;6(224):224ra24. doi:10.1126/scitranslmed.3007094. [Google Scholar] [PubMed] [CrossRef]
315. Alix-Panabières C, Pantel K. Clinical applications of circulating tumor cells and circulating tumor DNA as liquid biopsy. Cancer Discov. 2016;6(5):479–91. doi:10.1158/2159-8290.CD-15-1483. [Google Scholar] [PubMed] [CrossRef]
316. Tivey A, Church M, Rothwell D, Dive C, Cook N. Circulating tumour DNA—looking beyond the blood. Nat Rev Clin Oncol. 2022;19(9):600–12. doi:10.1038/s41571-022-00660-y. [Google Scholar] [PubMed] [CrossRef]
317. Domingo-Prim J, Endara-Coll M, Bonath F, Jimeno S, Prados-Carvajal R, Friedländer MR, et al. EXOSC10 is required for RPA assembly and controlled DNA end resection at DNA double-strand breaks. Nat Commun. 2019;10(1):2135. doi:10.1038/s41467-019-10153-9. [Google Scholar] [PubMed] [CrossRef]
318. Marin-Vicente C, Domingo-Prim J, Eberle AB, Visa N. RRP6/EXOSC10 is required for the repair of DNA double-strand breaks by homologous recombination. J Cell Sci. 2015;128(6):1097–107. doi:10.1242/jcs.158733. [Google Scholar] [PubMed] [CrossRef]
319. McMullen M, Karakasis K, Madariaga A, Oza AM. Overcoming platinum and PARP-inhibitor resistance in ovarian cancer. Cancers. 2020;12(6):1607. doi:10.3390/cancers12061607. [Google Scholar] [PubMed] [CrossRef]
320. Zgurskaya HI, Walker JK, Parks JM, Rybenkov VV. Multidrug efflux pumps and the two-faced Janus of substrates and inhibitors. Acc Chem Res. 2021;54(4):930–9. doi:10.1021/acs.accounts.0c00843. [Google Scholar] [PubMed] [CrossRef]
321. Patterson-Fortin J, D’Andrea AD. Targeting polymerase theta (POLθ) for cancer therapy. Cancer Treat Res. 2023;186(1):285–98. doi:10.1007/978-3-031-30065-3_15. [Google Scholar] [PubMed] [CrossRef]
322. Schrempf A, Slyskova J, Loizou JI. Targeting the DNA repair enzyme polymerase θ in cancer therapy. Trends Cancer. 2021;7(2):98–111. doi:10.1016/j.trecan.2020.09.007. [Google Scholar] [PubMed] [CrossRef]
323. Abeykoon JP, Asada S, Parmar K, Wu X, Witzig T, Shapiro G, et al. Abstract 6208: targeting DNA polymerase theta and ATM leads to synergistic killing of mantle cell lymphoma cells. Cancer Res. 2023;83(7_Suppl):6208. doi:10.1158/1538-7445.am2023-6208. [Google Scholar] [CrossRef]
324. Patterson-Fortin J, Jadhav H, Pantelidou C, Phan T, Grochala C, Mehta AK, et al. Polymerase theta inhibition activates the cGAS-STING pathway and cooperates with immune checkpoint blockade in BRCA-deficient cancers. Cancer Res. 2023;83(7_Suppl):6190. doi:10.1158/1538-7445.am2023-6190. [Google Scholar] [CrossRef]
325. Yang W, Feng Y, Zhou J, Cheung OK, Cao J, Wang J, et al. A selective HDAC8 inhibitor potentiates antitumor immunity and efficacy of immune checkpoint blockade in hepatocellular carcinoma. Sci Transl Med. 2021;13(588):eaaz6804. doi:10.1126/scitranslmed.aaz6804. [Google Scholar] [PubMed] [CrossRef]
326. Sule A, Van Doorn J, Sundaram RK, Ganesa S, Vasquez JC, Bindra RS. Targeting IDH1/2 mutant cancers with combinations of ATR and PARP inhibitors. NAR Cancer. 2021;3(2):zcab018. doi:10.1093/narcan/zcab018. [Google Scholar] [PubMed] [CrossRef]
327. Jordan MA, Wilson L. Microtubules as a target for anticancer drugs. Nat Rev Cancer. 2004;4(4):253–65. doi:10.1038/nrc1317. [Google Scholar] [PubMed] [CrossRef]
328. Solomon AD, Gouttia OG, Wang L, Zhu S, Wang F, Li Y, et al. γ-tubulin mediates DNA double-strand break repair. J Cell Sci. 2025;138(6):jcs262255. doi:10.1242/jcs.262255. [Google Scholar] [PubMed] [CrossRef]
329. Wang F, Rong M, Zhang L, Solomon AD, Gui W, Li J, et al. Vimentin intermediate filaments orchestrate DNA nonhomologous end joining repair and lipolysis after DNA damage. Oncogene. 2025;44(33):3025–36. doi:10.1038/s41388-025-03465-2. [Google Scholar] [PubMed] [CrossRef]
330. Zhu S, Paydar M, Wang F, Li Y, Wang L, Barrette B, et al. Kinesin Kif2C in regulation of DNA double strand break dynamics and repair. Elife. 2020;9:e53402. doi:10.7554/eLife.53402. [Google Scholar] [PubMed] [CrossRef]
331. Javle M, Shacham-Shmueli E, Xiao L, Varadhachary G, Halpern N, Fogelman D, et al. Olaparib monotherapy for previously treated pancreatic cancer with DNA damage repair genetic alterations other than germline BRCA variants: findings from 2 phase 2 nonrandomized clinical trials. JAMA Oncol. 2021;7(5):693–9. doi:10.1001/jamaoncol.2021.0006. [Google Scholar] [PubMed] [CrossRef]
332. Castro E, Romero-Laorden N, Del Pozo A, Lozano R, Medina A, Puente J, et al. PROREPAIR-B: a prospective cohort study of the impact of germline DNA repair mutations on the outcomes of patients with metastatic castration-resistant prostate cancer. J Clin Oncol. 2019;37(6):490–503. doi:10.1200/JCO.18.00358. [Google Scholar] [PubMed] [CrossRef]
333. Cheng X, Cao Y, Liu X, Li Y, Li Q, Gao D, et al. Single-cell and spatial omics unravel the spatiotemporal biology of tumour border invasion and haematogenous metastasis. Clin Transl Med. 2024;14(10):e70036. doi:10.1002/ctm2.70036. [Google Scholar] [PubMed] [CrossRef]
334. Phillips D, Schürch CM, Khodadoust MS, Kim YH, Nolan GP, Jiang S. Highly multiplexed phenotyping of immunoregulatory proteins in the tumor microenvironment by CODEX tissue imaging. Front Immunol. 2021;12:687673. doi:10.3389/fimmu.2021.687673. [Google Scholar] [PubMed] [CrossRef]
335. Keren L, Bosse M, Thompson S, Risom T, Vijayaragavan K, McCaffrey E, et al. MIBI-TOF: a multiplexed imaging platform relates cellular phenotypes and tissue structure. Sci Adv. 2019;5(10):eaax5851. doi:10.1126/sciadv.aax5851. [Google Scholar] [PubMed] [CrossRef]
336. Ortmayr K, Dubuis S, Zampieri M. Metabolic profiling of cancer cells reveals genome-wide crosstalk between transcriptional regulators and metabolism. Nat Commun. 2019;10(1):1841. doi:10.1038/s41467-019-09695-9. [Google Scholar] [PubMed] [CrossRef]
337. Kabatnik S, Zheng X, Pappas G, Steigerwald S, Padula MP, Mann M. Deep visual proteomics reveals DNA replication stress as a hallmark of signet ring cell carcinoma. npj Precis Oncol. 2025;9(1):37. doi:10.1038/s41698-025-00819-7. [Google Scholar] [PubMed] [CrossRef]
338. Vito A, El-Sayes N, Mossman K. Hypoxia-driven immune escape in the tumor microenvironment. Cells. 2020;9(4):992. doi:10.3390/cells9040992. [Google Scholar] [PubMed] [CrossRef]
339. Schleicher EM, Moldovan GL. CRISPR screens guide the way for PARP and ATR inhibitor biomarker discovery. FEBS J. 2022;289(24):7854–68. doi:10.1111/febs.16217. [Google Scholar] [PubMed] [CrossRef]
340. Ceccaldi R, Liu JC, Amunugama R, Hajdu I, Primack B, Petalcorin MIR, et al. Homologous-recombination-deficient tumours are dependent on Polθ-mediated repair. Nature. 2015;518(7538):258–62. doi:10.1038/nature14184. [Google Scholar] [PubMed] [CrossRef]
341. Bazan Russo TD, Mujacic C, Di Giovanni E, Vitale MC, Ferrante Bannera C, Randazzo U, et al. Polθ: emerging synthetic lethal partner in homologous recombination-deficient tumors. Cancer Gene Ther. 2024;31(11):1619–31. doi:10.1038/s41417-024-00815-2. [Google Scholar] [PubMed] [CrossRef]
342. Lin K, Chang YC, Billmann M, Ward HN, Le K, Hassan AZ, et al. A scalable platform for efficient CRISPR-Cas9 chemical-genetic screens of DNA damage-inducing compounds. Sci Rep. 2024;14(1):2508. doi:10.1038/s41598-024-51735-y. [Google Scholar] [PubMed] [CrossRef]
343. Wang L, Song Y, Wang H, Zhang X, Wang M, He J, et al. Advances of artificial intelligence in anti-cancer drug design: a review of the past decade. Pharmaceuticals. 2023;16(2):253. doi:10.3390/ph16020253. [Google Scholar] [PubMed] [CrossRef]
344. Papp O, Jordán V, Hetey S, Balázs R, Kaszás V, Bartha Á, et al. Network-driven cancer cell avatars for combination discovery and biomarker identification for DNA damage response inhibitors. npj Syst Biol Appl. 2024;10(1):68. doi:10.1038/s41540-024-00394-w. [Google Scholar] [PubMed] [CrossRef]
345. Yu G, Fan Q. Deep learning-driven drug response prediction and mechanistic insights in cancer genomics. Sci Rep. 2025;15(1):20824. doi:10.1038/s41598-025-91571-2. [Google Scholar] [PubMed] [CrossRef]
346. Plisson F, Piggott AM. Predicting blood−brain barrier permeability of marine-derived kinase inhibitors using ensemble classifiers reveals potential hits for neurodegenerative disorders. Mar Drugs. 2019;17(2):81. doi:10.3390/md17020081. [Google Scholar] [PubMed] [CrossRef]
347. Huang ETC, Yang JS, Liao KYK, Tseng WCW, Lee CK, Gill M, et al. Predicting blood-brain barrier permeability of molecules with a large language model and machine learning. Sci Rep. 2024;14(1):15844. doi:10.1038/s41598-024-66897-y. [Google Scholar] [PubMed] [CrossRef]
348. Shen S, Qi W, Liu X, Zeng J, Li S, Zhu X, et al. From virtual to reality: innovative practices of digital twins in tumor therapy. J Transl Med. 2025;23(1):348. doi:10.1186/s12967-025-06371-z. [Google Scholar] [PubMed] [CrossRef]
349. Wang E, Xiang K, Zhang Y, Wang XF. Patient-derived organoids (PDOs) and PDO-derived xenografts (PDOXsnew opportunities in establishing faithful pre-clinical cancer models. J Natl Cancer Cent. 2022;2(4):263–76. doi:10.1016/j.jncc.2022.10.001. [Google Scholar] [PubMed] [CrossRef]
350. Roh TT, Alex A, Chandramouleeswaran PM, Sorrells JE, Ho A, Iyer RR, et al. Predicting DNA damage response in non-small cell lung cancer organoids via simultaneous label-free autofluorescence multiharmonic microscopy. Redox Biol. 2024;75(10):103280. doi:10.1016/j.redox.2024.103280. [Google Scholar] [PubMed] [CrossRef]
351. Yang R, Wang S, Li Z, Yin C, Huang W, Huang W. Patient-derived organoid co-culture systems as next-generation models for bladder cancer stem cell research. Cancer Lett. 2025;625:217793. doi:10.1016/j.canlet.2025.217793. [Google Scholar] [PubMed] [CrossRef]
352. Strating E, Verhagen MP, Wensink E, Dünnebach E, Wijler L, Aranguren I, et al. Co-cultures of colon cancer cells and cancer-associated fibroblasts recapitulate the aggressive features of mesenchymal-like colon cancer. Front Immunol. 2023;14:1053920. doi:10.3389/fimmu.2023.1053920. [Google Scholar] [PubMed] [CrossRef]
353. Wu Z, Zhang S, Li L, Huang Z, Huang D, Hu Y. The gut microbiota modulates responses to anti-PD-1 and chemotherapy combination therapy and related adverse events in patients with advanced solid tumors. Front Oncol. 2022;12:887383. doi:10.3389/fonc.2022.887383. [Google Scholar] [PubMed] [CrossRef]
354. Xu Z, Wang T, Wang Y, Li Y, Sun Y, Qiu HJ. Short-chain fatty acids: key antiviral mediators of gut microbiota. Front Immunol. 2025;16:1614879. doi:10.3389/fimmu.2025.1614879. [Google Scholar] [PubMed] [CrossRef]
355. Wu Z, Cui P, Tao H, Zhang S, Ma J, Liu Z, et al. The synergistic effect of PARP inhibitors and immune checkpoint inhibitors. Clin Med Insights Oncol. 2021;15:1179554921996288. doi:10.1177/1179554921996288. [Google Scholar] [PubMed] [CrossRef]
356. Li A, Yi M, Qin S, Chu Q, Luo S, Wu K. Prospects for combining immune checkpoint blockade with PARP inhibition. J Hematol Oncol. 2019;12(1):98. doi:10.1186/s13045-019-0784-8. [Google Scholar] [PubMed] [CrossRef]
357. Wekking D, Silva CAC, Viscò R, Denaro N, Lambertini M, Maccioni A, et al. The interplay between gut microbiota, antibiotics, and immune checkpoint inhibitors in patients with cancer: a narrative review with biological and clinical aspects. Crit Rev Oncol Hematol. 2025;212(2):104767. doi:10.1016/j.critrevonc.2025.104767. [Google Scholar] [PubMed] [CrossRef]
358. Yong J, Cai S, Zeng Z. Targeting NAD+ metabolism: dual roles in cancer treatment. Front Immunol. 2023;14:1269896. doi:10.3389/fimmu.2023.1269896. [Google Scholar] [PubMed] [CrossRef]
359. Tao L, Zhou Y, Luo Y, Qiu J, Xiao Y, Zou J, et al. Epigenetic regulation in cancer therapy: from mechanisms to clinical advances. MedComm. 2024;3(1):e59. doi:10.1002/mog2.59. [Google Scholar] [CrossRef]
360. Fouad MA, Salem SE, Hussein MM, Zekri ARN, Hafez HF, El Desouky ED, et al. Impact of global DNA methylation in treatment outcome of colorectal cancer patients. Front Pharmacol. 2018;9:1173. doi:10.3389/fphar.2018.01173. [Google Scholar] [PubMed] [CrossRef]
361. Sato T, Issa JJ, Kropf P. DNA hypomethylating drugs in cancer therapy. Cold Spring Harb Perspect Med. 2017;7(5):a026948. doi:10.1101/cshperspect.a026948. [Google Scholar] [PubMed] [CrossRef]
362. Adimoolam S, Sirisawad M, Chen J, Thiemann P, Ford JM, Buggy JJ. HDAC inhibitor PCI-24781 decreases RAD51 expression and inhibits homologous recombination. Proc Natl Acad Sci U S A. 2007;104(49):19482–7. doi:10.1073/pnas.0707828104. [Google Scholar] [PubMed] [CrossRef]
363. Roos WP, Krumm A. The multifaceted influence of histone deacetylases on DNA damage signalling and DNA repair. Nucleic Acids Res. 2016;44(21):10017–30. doi:10.1093/nar/gkw922. [Google Scholar] [PubMed] [CrossRef]
364. Nicetto D, Zaret KS. Role of H3K9me3 heterochromatin in cell identity establishment and maintenance. Curr Opin Genet Dev. 2019;55:1–10. doi:10.1016/j.gde.2019.04.013. [Google Scholar] [PubMed] [CrossRef]
365. Cisneros-Villanueva M, Hidalgo-Pérez L, Cedro-Tanda A, Peña-Luna M, Mancera-Rodríguez MA, Hurtado-Cordova E, et al. LINC00460 is a dual biomarker that acts as a predictor for increased prognosis in basal-like breast cancer and potentially regulates immunogenic and differentiation-related genes. Front Oncol. 2021;11:628027. doi:10.3389/fonc.2021.628027. [Google Scholar] [PubMed] [CrossRef]
366. Cynthia Stafford MY, McKenna DJ. miR-182 is upregulated in prostate cancer and contributes to tumor progression by targeting MITF. Int J Mol Sci. 2023;24(3):1824. doi:10.3390/ijms24031824. [Google Scholar] [PubMed] [CrossRef]
367. Moretton A, Loizou JI. Interplay between cellular metabolism and the DNA damage response in cancer. Cancers. 2020;12(8):2051. doi:10.3390/cancers12082051. [Google Scholar] [PubMed] [CrossRef]
368. Chalova P, Tazky A, Skultety L, Minichova L, Chovanec M, Ciernikova S, et al. Determination of short-chain fatty acids as putative biomarkers of cancer diseases by modern analytical strategies and tools: a review. Front Oncol. 2023;13:1110235. doi:10.3389/fonc.2023.1110235. [Google Scholar] [PubMed] [CrossRef]
369. Luo L, Keyomarsi K. PARP inhibitors as single agents and in combination therapy: the most promising treatment strategies in clinical trials for BRCA-mutant ovarian and triple-negative breast cancers. Expert Opin Investig Drugs. 2022;31(6):607–31. doi:10.1080/13543784.2022.2067527. [Google Scholar] [PubMed] [CrossRef]
370. Sheikh A, Curran MA. The influence of the microbiome on radiotherapy and DNA damage responses. Front Oncol. 2025;15:1552750. doi:10.3389/fonc.2025.1552750. [Google Scholar] [PubMed] [CrossRef]
371. Huang L, Li Y, Zhang C, Jiang A, Zhu L, Mou W, et al. Microbiome meets immunotherapy: unlocking the hidden predictors of immune checkpoint inhibitors. npj Biofilms Microbiomes. 2025;11(1):180. doi:10.1038/s41522-025-00819-2. [Google Scholar] [PubMed] [CrossRef]
Cite This Article
 Copyright © 2026 The Author(s). Published by Tech Science Press.
Copyright © 2026 The Author(s). Published by Tech Science Press.This work is licensed under a Creative Commons Attribution 4.0 International License , which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.


 Submit a Paper
Submit a Paper Propose a Special lssue
Propose a Special lssue View Full Text
View Full Text Download PDF
Download PDF Downloads
Downloads
 Citation Tools
Citation Tools