 Open Access
Open Access
REVIEW
Circular RNAs: Key Regulators of Tumor Metabolic Reprogramming and Clinical Translation
1 State Key Laboratory of Systems Medicine for Cancer, Shanghai Cancer Institute, Renji Hospital, Shanghai Jiao Tong University School of Medicine, Shanghai, China
2 Second Clinical Medical College, Guangdong Medical University, Zhanjiang, China
3 Department of Basic Medical Sciences, Kangda College of Nanjing Medical University, Lianyungang, China
4 College of Pharmacy, Inner Mongolia Medical University, Hohhot, China
5 Department of Pharmacy, The First Affiliated Hospital of Guangxi Medical University, Nanning, China
6 Shanghai Key Laboratory for Cancer Systems Regulation and Clinical Translation, Shanghai Jiading District Central Hospital, Shanghai, China
* Corresponding Authors: Qian Zhang. Email: ; Chen Li. Email:
; Mengyao Li. Email:
# These authors contributed equally to this work
(This article belongs to the Special Issue: Metabolic Heterogeneity in Cancer: Mechanisms, Biomarkers, and Therapeutic Implications)
Oncology Research 2026, 34(3), 1 https://doi.org/10.32604/or.2026.075012
Received 23 October 2025; Accepted 23 January 2026; Issue published 24 February 2026
Abstract
Tumor metabolic reprogramming is a core hallmark of cancer, characterized by pathways such as aerobic glycolysis, aberrant lipid metabolism, and glutaminolysis that support rapid proliferation and immunosuppressive microenvironments. Circular RNAs (circRNAs) are highly stable, evolutionarily conserved non-coding RNAs that have emerged as critical modulators of these metabolic shifts. This review aims to systematically elucidate the roles and mechanisms of circRNAs in reprogramming tumor metabolism, and to discuss their clinical potential as biomarkers and therapeutic targets. Through mechanisms including miRNA sponging, protein interactions, regulation of mitochondrial dynamics, and modulation of metabolic enzymes, circRNAs influence key metabolic pathways by targeting glycolytic enzymes, lipid synthesis regulators, and glutaminolysis-related molecules to either facilitate or inhibit their expression. This review systematically summarizes the unique contributions of circRNAs to tumor metabolic reprogramming, highlighting key mechanisms such as regulation of peptide-encoding protein translation, mitochondrial localization function, gene promoter-targeted transcriptional regulation, and cross-pathway metabolic mediation, which underscore their distinct biological advantages and regulatory roles in tumor metabolism. The stability and tissue specificity of circRNAs make them promising diagnostic biomarkers, while their role in drug resistance mediated by metabolic reprogramming highlights their potential as therapeutic targets. Strategies such as circRNA inhibitors, mimics, and nanoparticle-based delivery systems are being explored to modulate tumor metabolism. Despite challenges including complex regulatory networks and limited manipulation tools, advances in high-throughput technologies and clinical trials hold promise for translating circRNA research into novel cancer therapies.Keywords
Tumor metabolism is a defining hallmark of cancer, characterized by metabolic reprogramming that includes the Warburg effect (a preference for aerobic glycolysis in cancer cells), as well as aberrant lipid metabolism, glutaminolysis, and mitochondrial dynamics [1,2]. This reprogramming not only provides cancer cells with the energy and biosynthetic precursors necessary for rapid proliferation but also contributes to the formation of an immunosuppressive tumor microenvironment (TME) [3,4].
Circular RNAs are a class of predominantly non-coding RNAs characterized by their covalently closed circular structures. These molecules exhibit high stability, resistance to nuclease degradation, and strong evolutionary conservation. They regulate gene expression through various mechanisms, including acting as miRNA sponges, binding proteins, or translating functional polypeptides [5,6]. The formation of circular RNA involves back-splicing reactions of exons or introns in pre-mRNA, primarily relying on base-pairing sequences present in the intronic sequences flanking the circularization region [7,8]. This pairing facilitates the binding of the 5′ splice site with the 3′ splice site, leading to the formation of a circular structure in the RNA molecule [9]. Based on their generation pathways, circular RNAs can be categorized into two major types: one type comprises circular transcripts produced from intracellular non-coding sequences post-processing, while the other is generated through a spliceosome-dependent processing method, where precursor mRNA (pre-mRNA) in eukaryotes undergoes splicing to yield circular RNA [10,11].
The metabolic patterns of cancer cells are significantly different from those of normal cells. The survival environment of tumor cells is often characterized by hypoxia, acidity, oxidative stress, and nutritional deficiency. In order to survive in this harsh microenvironment, tumor cells must regulate their own metabolic pathways to meet the needs of survival, a process of adaptive changes known as “metabolic reprogramming” in oncology [12]. In recent years, the relationship between RNA and tumor metabolic reprogramming has become an important research direction in the field of tumors. There is a close relationship between N6-methyladenosine (m6A) modification of RNA and the Warburg effect. It can regulate glycolysis through various mechanisms: stabilizing the mRNA of key glycolytic molecules (such as HK2, GLUT1) to activate the pathway; promoting the translation of related mRNAs (such as PDK4, MYC) to enhance the activity of related enzymes; regulating the degradation of negative regulators’ mRNA of metabolic pathways (such as APC), so as to promote the aerobic glycolysis reprogramming of cancer cells [13,14]. Recent studies have found that circRNA plays an important role in regulating tumor metabolism. For example, circRNA can target key enzymes (such as HK2 and GLUT1) or signaling pathways (such as PI3K/AKT) to promote or inhibit the Warburg effect [2,4]. Strikingly, these regulatory effects of circRNAs frequently converge on central metabolic regulators, including HIF-1α, MYC, p53, AMPK, and mTOR—key mediators that orchestrate metabolic reprogramming by integrating signaling cues with metabolic demands [15]. CircRNAs modulate the activity, stability, or transcriptional output of these core regulators thereby indirectly controlling downstream metabolic pathways and strengthening a functional link between circRNA networks and tumor metabolic plasticity [16,17]. Through such interactions with central regulators, circRNAs also impact lipid metabolism—for example, circEPB41 modulate lipid synthesis to drive tumor growth in liver cancer [18]. Moreover, certain circRNAs contribute to chemoresistance through regulation of metabolic pathways, including glycolytic changes associated with 5-FU resistance [19].
CircRNAs display distinct regulatory properties during tumor metabolic reprogramming, setting them apart from other non-coding RNAs. Their unique attributes primarily arise from their covalently closed circular structure and specific molecular mechanisms. In comparison to lncRNAs, circRNAs offer enhanced stability, resisting degradation by exonucleases and boasting a half-life roughly ten times longer than that of linear RNA. Moreover, they exhibit notably elevated expression levels in tumor tissues and associated body fluids [20]. Remarkably, circRNAs can act as “super sponges,” binding multiple metabolism-associated miRNAs to collaboratively modulate crucial metabolic pathways, including glycolysis and lipid synthesis [8].
The same circRNA may have different effects in different tumor cells. The function of circHIPK3 is significantly heterogeneous in various cancers, and this contradictory phenotype provides a key entry point for understanding its mechanism of action. In most cancers, such as breast cancer, colorectal cancer, lung cancer, etc., circHIPK3 is highly expressed by binding to tumor suppressor miRNAs such as miR-124-3p and miR-637, activating classic oncogenic pathways such as PI3K/Akt and Wnt/β-catenin, thereby promoting the proliferation, metastasis, and chemotherapy resistance of tumor cells and exerting pro-cancer effects. However, in bladder cancer, multiple studies have confirmed that the expression of circHIPK3 is downregulated, which can regulate the miR-588/HPSE axis and inhibit the autophagy pathway, showing anti-cancer properties. The core reason for this functional contradiction may be related to specific differences in the tissue microenvironment: the bladder tissue and tumor microenvironment contain high concentrations of hydrogen peroxide (H2O2), and existing evidence shows that H2O2 can directly induce the downregulation of circHIPK3 expression [21,22]. This tissue microenvironment-dependent functional switch suggests the functional heterogeneity of circHIPK3, and its clinical application needs to be accurately evaluated according to the type of tumor and the characteristics of the microenvironment [22]. This characteristic also brings complexity to the selection and determination of therapeutic targets.
The regulatory mechanisms of circRNAs in tumor metabolism are of great value. Their stability and tissue specificity make them promising biomarkers, as demonstrated by the use of exosomal circRNAs in urinary system tumors [23,24]. Targeting metabolically relevant circRNAs such as circPVT1 in triple-negative breast cancer (TNBC) may reverse abnormal metabolism and treatment resistance [25–27]. However, there remain critical knowledge gaps and challenges: the precise molecular cascade through which circRNAs coordinate cross-pathway metabolic regulation is not fully defined; the upstream signals that control functional switching of circRNAs in a tissue-specific manner have not been fully elucidated; and translational efforts to apply circRNA-based strategies in clinical practice are hindered by limited understanding of their long-term regulatory effects and optimal delivery paradigms.
This review aims to address these unresolved issues by systematically summarizing the molecular mechanisms of circRNA-mediated tumor metabolic reprogramming, dissecting the drivers of their functional heterogeneity, exploring their clinical potential as therapeutic targets or biomarkers for metabolism, and outlining critical future research directions to bridge the gap between basic research and clinical translation. By integrating recent advances, this article provides a theoretical framework to guide the development of targeted circRNA-based strategies for tumor metabolic intervention and discusses key barriers to their clinical application.
2 Tumor Metabolism: A Core Hallmark of Cancer
2.1 Overview of Metabolic Reprogramming
Metabolic reprogramming refers to the adaptive rewiring of bioenergetic pathways that tumor cells exhibit in order to survive and proliferate under harsh conditions such as hypoxia and nutrient deprivation [28]. As a key hallmark distinguishing them from normal cells, this reprogramming involves reshaped lipid metabolism, glycolysis, glutaminolysis, and mitochondrial dynamics. cancer cells take up glucose and produce lactate (aerobic glycolysis) even in the presence of oxygen, rely on glutamine for anaplerosis and biosynthesis (glutaminolysis), increase lipogenesis, and alter fatty acid oxidation [29,30]. These metabolic shifts not only provide energy but also support unlimited proliferation by providing essential precursors for biomacromolecule synthesis, which enables survival in nutrient-poor environments (Fig. 1) [29,30].
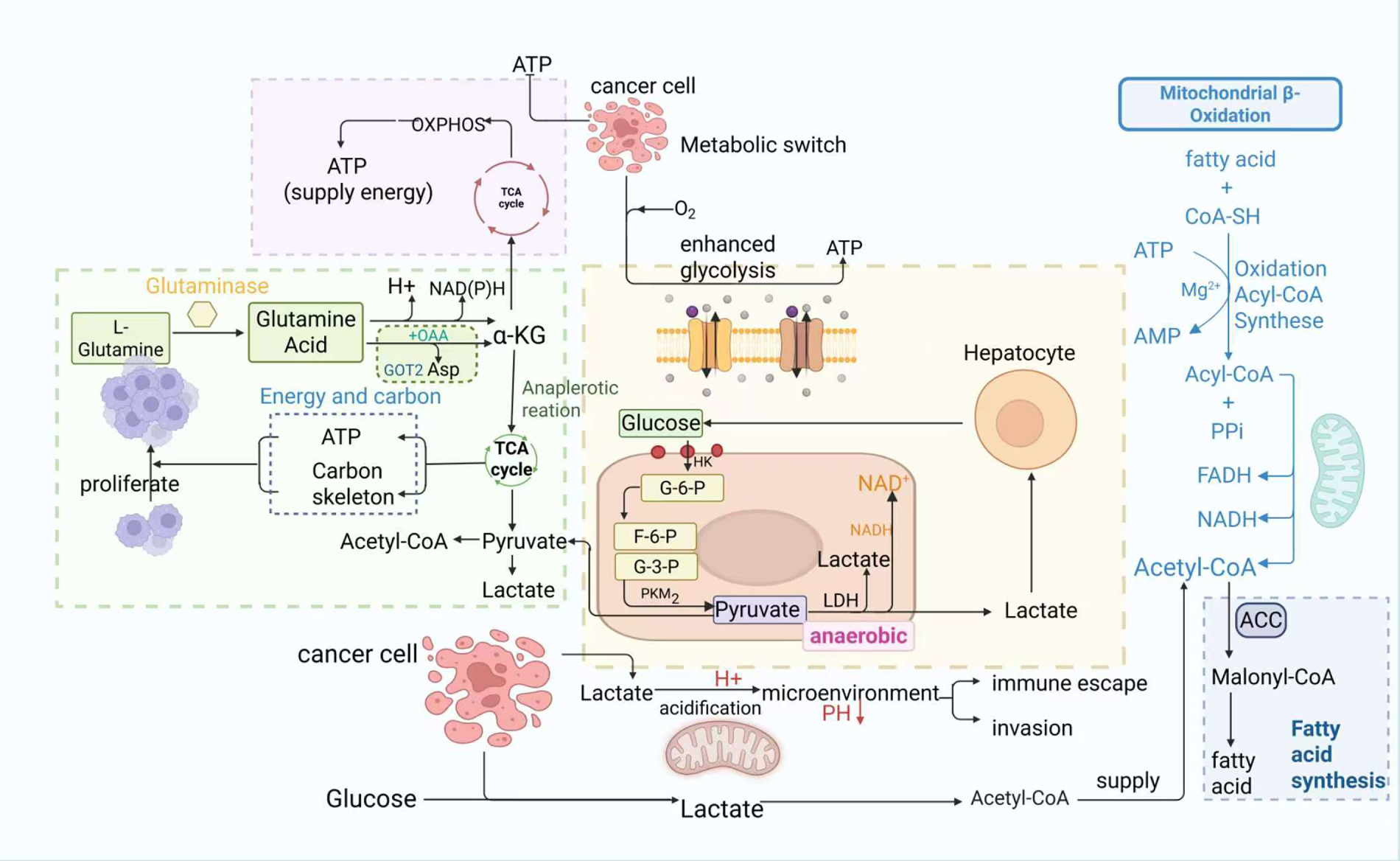
Figure 1: Schematic of metabolic reprogramming in cancer cells, highlighting key roles of glycolysis, glutaminolysis, and lipid metabolism, and lactate secretion’s impact on the tumor microenvironment. Glucose enters cancer cells and is converted to pyruvate via a series of enzymes, producing small amounts of ATP; unlike normal cells, this pyruvate is further converted to lactate by lactate dehydrogenase (LDH). L-glutamine is catalyzed by glutaminase to glutamate, which enters mitochondria as α-ketoglutarate (α-KG) to participate in the tricarboxylic acid (TCA) cycle, providing energy and carbon skeletons for tumor survival. Additionally, fatty acids and coenzyme A (CoA-SH) form acyl-CoA via acyl-CoA synthetase (consuming ATP), which enters mitochondria for oxidation, generating acetyl-CoA that feeds the TCA cycle and produces FADH2 and NADH—key for mitochondrial ATP production. Lactate secreted by cancer cells acidifies the tumor microenvironment, inhibiting immune cell function while promoting cancer cell proliferation and invasion, and also recycles acetyl-CoA to meet lipid precursor demands for membrane synthesis.
2.2 Key Metabolic Pathways in Reprogramming
2.2.1 Aerobic Glycolysis (Warburg Effect)
The Warburg effect describes the preference of cancer cells to generate ATP via glycolysis followed by lactate production, even in the presence of functional mitochondrial respiratory chains and intact oxidative phosphorylation capacity [31]. This ‘glycolytic-dominant’ metabolism upregulates expression of glycolytic enzymes to meet energy demands and provide critical metabolic intermediates for biosynthesis, thereby promoting tumor growth and invasion [32,33]. The elevated levels of lactate from this process lower TME pH, impairing macrophage reprogramming and T cell function, thus suppressing antitumor immunity [34]. Targeting oncogenic signaling pathways that regulate this effect—such as MAPK/ERK and MAPK/JNK, which are involved in tumorigenesis beyond their normal role in sensing extracellular stimuli—has emerged as a promising anticancer strategy [35].
2.2.2 Lipid Metabolic Reprogramming
Lipid metabolic reprogramming involves dysregulated (up- or down-regulation) processes of lipid uptake, desaturation, de novo synthesis, lipid droplet formation, and fatty acid oxidation in cancer cells [36]. Regulated by oncogenes, tumor suppressors, key transcription factors and growth factor signaling pathways [37], this reprogramming enhances lipid uptake, storage and synthesis to drive rapid tumor growth in various cancers [38]. Taking colorectal cancer cells as an example, tumor cells significantly increase their ability to uptake exogenous fatty acids by upregulating the expression of fatty acid transporters such as CD36 and FABPs. In CRC cells, the protein levels of key metabolic enzymes, including acetyl CoA carboxylase (ACC), fatty acid synthase (FASN) and stearoyl-CoA desaturase (SCD1) are significantly upregulated, thereby accelerating the de novo synthesis of endogenous fatty acids in tumor cells. The abnormally high expression of these metabolic enzymes not only provides sufficient precursors for lipid synthesis in tumor cells but also promotes tumor initiation and progression by activating multiple classic oncogenic signaling pathways such as Wnt/β-catenin and PI3K/AKT/mTOR [39,40]. Among them, the core metabolic enzyme FASN can enhance the proliferation and migration of colorectal cancer cells through AMPK/mTOR signaling pathway [41,42]. In addition to supporting tumor development via lipid acquisition and increased fatty acid oxidation, it also modulates immune cells to shape TME into a tumor-promoting niche [43]. For instance, enhanced lipid metabolism increases hyaluronan (HA) oligomers that facilitate cholesterol efflux in tumor-associated macrophages (TAMs) and drive their polarization toward M2 phenotype, thus accelerating tumor progression [44]. Such findings highlight the potential of therapeutically targeting TAMs to suppress their pro-tumor functions.
Mitochondria Dynamically undergo fusion and fission in response to environmental cues, a process known as mitochondrial dynamics [45] that is frequently deregulated in tumors and associated with drug resistance, cancer stemness, and metastatic potential [46–48]. Mitochondrial fission is mainly regulated by the cytoplasmic GTPase dynamin-related protein 1 (DRP1), which is recruited through a series of receptor proteins anchored on the outer mitochondrial membrane, including mitochondrial fission factor (MFF), mitofusin 49 and 51 kDa (MID49 and MID51) and fission protein 1 (FIS1). Studies have shown that different receptor proteins regulate distinct aspects of mitochondrial fission. Among them, MID49 and MID51 can bind acyl-CoA and act as sensors for β-oxidation metabolites, thus linking mitochondrial fission to changes in cellular metabolic status [49]. Fusion relies on mitofusins (MFN1/2) mediating outer membrane fusion and optic atrophy 1 (OPA1) mediating inner membrane fusion, whose core function is to maintain mitochondrial homogeneity and mtDNA integrity. When fission prevails, cells switch from oxidative phosphorylation (OXPHOS)-dependent metabolism to glycolysis; conversely, enhanced fusion favors OXPHOS [49,50]. The Increased activity of dynamin-related protein 1, a key regulator of fission, promotes mitochondrial fragmentation and increases metastatic capacity in breast, lung, and brain cancers [46]. Likewise, excessive fission drives hepatocellular carcinoma metastasis by regulating glucose metabolic reprogramming. Inhibitors of mitochondrial dynamics (mdivi-1) have demonstrated antitumor potential, underscoring the relevance of this pathway [51].
Glutaminolysis, the conversion of glutamine to glutamate and subsequently to α-ketoglutarate, is tightly coupled with mitochondrial dynamics [45]. Oncogene activation drives most cancer cells to rely heavily on the “glutamine anaplerosis” mechanism, shuttling glutamine-derived α-ketoglutarate into the tricarboxylic acid (TCA) cycle to fuel energy production [52,53]. As the most abundant circulating amino acid in blood and muscle, glutamine participates in multiple critical metabolic processes in cancer cells. Glutaminolysis also crosstalks with glycolysis, providing TCA cycle intermediates when glucose is limited [54]. Tumor cells, therefore, upregulate glutamine uptake and catabolism to support rapid proliferation [55], while glutamine also modulates intracellular signaling pathways to influence metabolic reprogramming [56]. The close interplay between the Warburg effect and glutaminolysis underscores their joint role in tumor progression, highlighting the therapeutic potential of targeting this crosstalk [53].
2.3 Drivers of Metabolic Reprogramming
Metabolic reprogramming is driven primarily by genetic mutations in oncogenes and tumor suppressors [29]. Activated oncogenes, for example, RAS and BRAF mutations, induce a glycolytic preference and increased glutamine dependence. Unlike normal cells that reduce energy consumption during nutrient scarcity, cancer cells with RAS or BRAF mutations bypass such regulation to continue glucose uptake, ensuring sufficient energy for proliferation and survival [57]. Genetic mutation remains the central driver from initiation through metastasis, although the roles of metabolism and epigenetics have only recently been recognized [58].
Epigenetic modifications, including DNA methylation, histone modification and chromatin remodelling, profoundly influence metabolic reprogramming [59]. Aberrant DNA methylation in tumor cells activates oncogenes and silences tumor suppressors and alters chromatin structure to rewire metabolic pathways. For example, hypermethylation of SLC27A6 downregulates its expression and promotes nasopharyngeal carcinoma (NPC) cell proliferation through altered lipid metabolism [60]. Histone modifications such as phosphorylation, acetylation, and methylation also have key roles. Histone methylation is mainly performed at lysine or arginine residues by histone methyltransferases (HMTs) using S-adenosylmethionine (SAM) as a cofactor [58]. Histone acetylation is regulated by histone acetyltransferases (HATs) and deacetylases (HDACs), which involves the HAT-mediated transfer of an acetyl group from acetyl-CoA to a histone lysine and is generally associated with transcriptional activation [61]. In addition to histones, acetylation directly modulates the function of proteins that are critical for oncogenesis; for instance, it can promote mitochondrial and upregulate glycolysis to drive metabolic reprogramming [62]. Furthermore, O-glcNAcylation of chromatin regulators changes chromatin morphology, further affecting gene expression. Together, epigenetic mechanisms regulate the expression of metabolic genes to modulate macromolecular synthesis and energy production, thereby orchestrating cancer cell reprogramming (Fig. 2) [63].
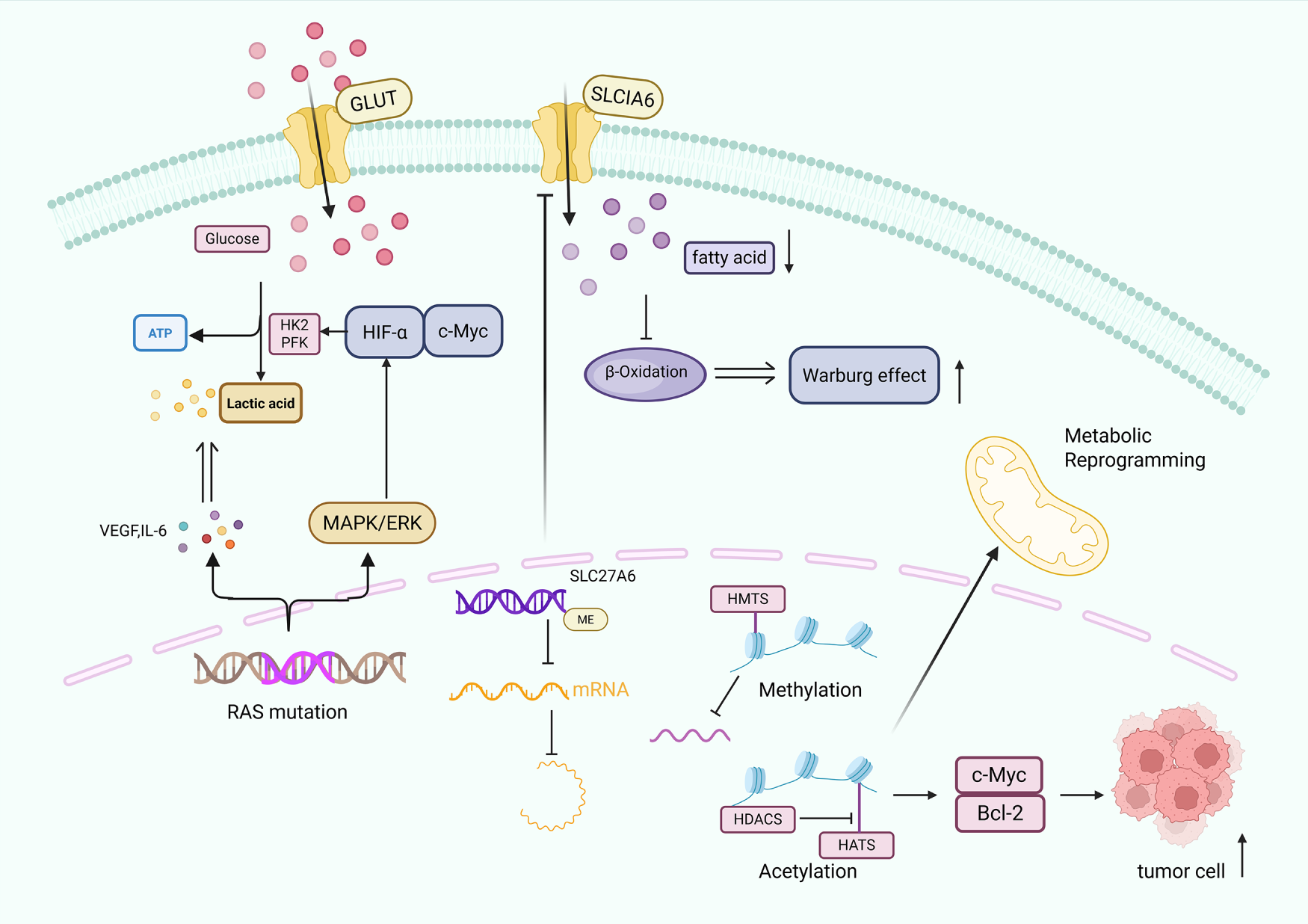
Figure 2: Molecular mechanisms driving tumor metabolic reprogramming: genetic mutations, DNA methylation, histone modifications, and their effects on glycolysis, lipid metabolism, and oncogene activation. Tumor metabolic reprogramming occurs via three main mechanisms: RAS mutations activate the mitogen-activated protein kinase/extracellular signal-regulated kinase (MAPK/ERK) pathway, upregulating glycolytic rate-limiting enzymes (hexokinase 2 (HK2), phosphofructokinase (PFK)) and prompting secretion of vascular endothelial growth factor (VEGF) and interleukin 6 (IL-6), thereby increasing glucose transporter (GLUT)-mediated glucose uptake and directing metabolism toward lactate production. DNA methylation of the solute carrier family 27 member 6 (SLC27A6) promoter downregulates its expression, reducing solute carrier family 1 member 6 (SLC1A6) (a related lipid transporter) to inhibit fatty acid β-oxidation and mitochondrial phosphorylation, shifting metabolism toward glycolysis. Histone methylation (regulated by histone methyltransferases (HMTs)) alters chromatin structure to repress tumor suppressors, while histone acetylation (regulated by histone deacetylases (HDACs) and histone acetyltransferases (HATs)) activates oncogenes like MYC proto-oncogene protein (c-Myc) and B-cell lymphoma 2 (Bcl-2), collectively driving metabolic reprogramming and abnormal proliferation. Additionally, hypoxia-inducible factor-α (HIF-α) is a key regulator in this process.
The TME, which is composed of fibroblasts, immune cells, and abundant stroma [64], shapes metabolic reprogramming by inducing scavenging mechanisms that force tumor cells to adapt to reduced nutrient availability [65,66]. For example, stromal cells secrete growth factors, cytokines and extracellular matrix (ECM) components that support tumor proliferation and invasion [67]; they also release amino acids via autophagy and pinocytosis to feed cancer cell metabolism [68]. Pancreatic cancer cells, for instance, use nucleosides and branched-chain keto acids produced by stromal cells under conditions of low glutamine [69,70]. Furthermore, metabolites derived from the TME modulate immune cell function, promoting an immunosuppressive niche that supports tumor progression (Fig. 3) [71–74].
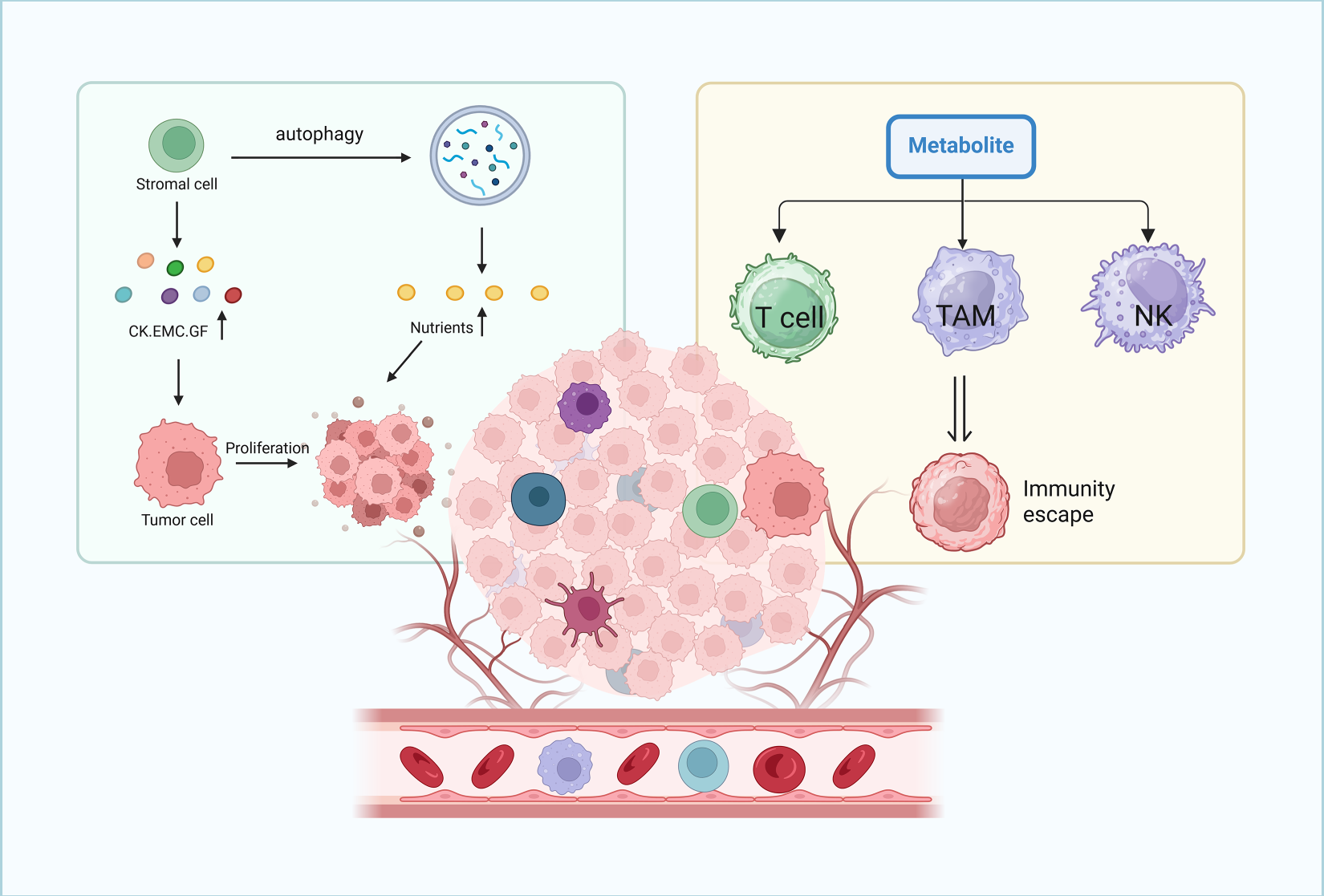
Figure 3: Role of the tumor microenvironment in supporting cancer cell proliferation and immune evasion through nutrient supply and immune cell modulation. The tumor microenvironment supports cancer cell proliferation and immune evasion: stromal cells release nutrients, cytokines (CK), extracellular matrix components (EMC), and growth factors (GF) via autophagy to promote tumor growth. Meanwhile, metabolites from cancer cell metabolism inhibit the function of T lymphocytes, macrophages, and natural killer cells, enabling tumor cells to evade immune recognition and elimination.
In summary, metabolic reprogramming in tumor cells is a well-recognized hallmark of cancer, driven by genetic mutations, epigenetic modifications, and TME cues. These changes collectively supply energy and biomolecules to sustain tumor initiation, growth, and metastasis.
3 Roles of CircRNAs in Metabolic Pathways
3.1 CircRNAs and the Warburg Effect
Recent studies have highlighted that circRNAs are key molecules in metabolic reprogramming, which are deeply involved in the regulation of aerobic glycolysis in tumor cells through various mechanisms. It is worth noting that circRNAs play dual roles in tumor metabolism: some act as oncogenic regulators by promoting abnormal metabolic reprogramming, while others function as tumor suppressors by inhibiting dysregulated metabolic pathways. This functional duality not only complicates therapeutic targeting but also reflects an unresolved controversy in the field: whether the context-dependent effects of circRNAs arise from their intrinsic molecular properties or are predominantly shaped by cues from the tumor microenvironment, a distinction that remains poorly defined and necessitates integrative analysis of circRNA-interacting networks across heterogeneous tumor settings [75,76]. These mechanisms mainly include direct modulation of glycolytic enzyme expression, translation of functional polypeptides and miRNA sponge by competing endogenous RNA (ceRNA) mechanism. Competitive endogenous RNA (ceRNA) is a special class of RNA molecules, mainly including protein coding messenger RNA (mRNA), pseudogene, long non-coding RNA (lncRNA) and circular RNA (circRNA) [77]. These RNA molecules competitively bind to microRNA (miRNA) to exert their biological functions, thereby relieving the inhibitory effect of miRNA on its target mRNA [78]. In the occurrence and development of tumors, they are involved in regulating the proliferation, invasion and migration ability of tumor cells, and are closely related to drug resistance of tumors, and have important influence on the overall progress of tumors [79].
Direct regulation of glycolytic enzymes by specific RNA binding domains (RBDs) in the enzyme, such as hexokinase 2 (HK2), phosphofructokinase (PFK), and lactate dehydrogenase A (LDHA), represents a major pathway. For HK2, circ_0000376 has been shown to significantly upregulate HK2 expression through miR-577-mediated regulation of HK2 and LDHA, thereby promoting glycolysis and tumor growth in osteosarcoma [80]. Overexpression of circVMP1 increases Hexokinase domain component 1(HKDC1) levels by competitively binding with miR-3167, enhancing glycolysis and colorectal cancer progression [81]. In glioblastoma (GBM), U3 snoRNA-derived circRNA directly binds to the promoter of HK2 via transcription factor ZBTB7A, which enhances HK2 expression through the U3/ZBTB7A/HK2 regulatory axis to drive tumor progression [82]. For PFK, some circRNAs inhibit PFK activity to reduce aerobic glycolytic flux and shift cellular metabolism toward mitochondrial oxidative phosphorylation [15], while circ_0031242 positively regulates PFK expression through MAD2L1/miR-944 pathways, exerting antagonistic effects on glycolysis [83]. For LDHA, circRARS modulates its activity by binding to LDHA and enhancing its enzymatic activity to promote glycolysis in non-small cell lung cancer (NSCLC) [84], and the transcription factor YY1 can directly bind to the LDHA promoter through circRNA-mediated mechanisms to enhance its expression [10]. These diverse signaling pathways collectively regulate glycolytic enzyme expression, providing the potential to develop targeted therapies for specific cancer types. ENO1 is a key enzyme in the glycolytic pathway and circUBE2G1-99aa can bind to the ENO1 enzyme, regulating the activity of the ENO1/PI3K/AKT signaling pathway, inhibiting glycolysis, altering tumor metabolic patterns, thereby contributing to the inhibition of gastric cancer cell proliferation [85]. In addition, circRNAs affect tumor growth and proliferation by regulating key transporters involved in glycolysis. For example, ITCH has been shown to potentially suppress cancer cell proliferation by downregulating GLUT1 expression, thereby reducing glucose uptake in melanoma cells [86].
CircRNAs sponge miRNAs to alleviate the repression of glycolytic genes through a ceRNA mechanism. For instance, circ_0000376 sequesters miR-577 to relieve its inhibition on HK2 and LDHA, thereby activating the Warburg effect and promoting tumor cell glycolysis [80]. Similarly, circ_MAPK9 binds to miR-642b-3p to reverse its suppression of STAT3 and LDHA, leading to increased ATP and lactate production that supports tumor cell energy metabolism [87]. In gastrointestinal cancers, circRNAs fine-tune miRNA networks to regulate the expression of glycolytic proteins such as GLUT1 and HK2, highlighting their intricate roles in glucose metabolic regulation [88,89]. The covalently closed circular structure of circRNAs confers high stability, making them ideal diagnostic biomarkers [90]. These findings position circRNAs as critical nodes in glycolytic regulation with promising potential as targets for metabolic intervention. Notably, many circRNAs that regulate glycolysis also crosstalk with lipogenesis and glutaminolysis pathways, integrating multiple metabolic processes to support tumor progression. This interconnected regulation ensures coordinated metabolic rewiring, whereby circRNA-mediated modulation of glycolytic enzymes or signaling cascades simultaneously influences lipid synthesis and glutamine catabolism.
3.2 CircRNAs and Lipid Metabolism
CircRNAs regulate lipid metabolism through multiple mechanisms, with the ceRNA mechanism being particularly prominent. Importantly, circRNAs involved in lipid metabolic reprogramming often exhibit crosstalk with glycolytic pathways, as metabolic networks are highly interconnected in tumor cells. CircRNAs may target key molecules of both lipid synthesis and glycolysis simultaneously to ensure that energy production and biosynthesis are coordinated to meet the needs of the rapid proliferation of tumor cells [16]. By competitively binding miRNAs such as miR-24-3p, circRNAs release the suppression on key lipid synthesis pathways (PPARγ signaling pathway and Igf2/PI3K-AKT-mTOR) and thus upregulate fatty acid synthase expression [91]. In the liver, spatially specific regulation by circRNAs promotes lipogenesis in mitochondria of the central zone (cental zone, referring to centrilobular regions) and enhances inhibition of fatty acid β-oxidation in mitochondria of the periportal zone (periportal, referring to periportal regions) [92], dysregulation of this balance results in abnormal lipid accumulation [93,94].
In intrahepatic cholangiocarcinoma (ICC), elevated circMBOAT2 expression binds to PTBP1, facilitating the export of Fatty Acid Synthase (FASN) mRNA and promoting lipid metabolism [95]. In liver fluke-associated cholangiocarcinoma, circRNAs upregulate FASN, leading to free fatty acid accumulation and creating an immunosuppressive microenvironment; inhibiting FASN enhances the efficacy of PD-1 inhibitors [96]. The aberrant upregulation of circ-SLC9A6 in non-alcoholic fatty liver disease (NAFLD) intensifies hepatocellular lipid metabolic disorders, as evidenced by comparative studies on human and mouse liver tissues [97]. Furthermore, circPRKAA1 plays a role in maintaining lipid homeostasis by enhancing fatty acid synthesis and lipid storage. Its biogenesis is negatively modulated by AMPK activity, marking it as a pivotal element in lipid metabolic reprogramming [91]. Within cancer cells, circRNAs influence lipogenesis by modulating key enzymes such as fatty acid synthase (FASN) and acetyl-CoA carboxylase (ACC), thereby affecting lipid synthesis pathways [98].
3.3 CircRNAs and Glutaminolysis
CircRNAs modulate glutaminolysis through various molecular mechanisms, among which the ceRNA mechanism is well-characterized [99]. For instance, circ_0000808/miR-1827/SLC1A5 regulatory cascade sponges miR-1827 to release solute carrier family 1 member 5 (SLC1A5) (a glutamate transporter) from repression, thereby promoting glutamine uptake [100]. CircPVT1 competitively binds with miR-33a-5p to liberate c-MYC, which activates transcription of glutaminase (GLS) and accelerates conversion of glutamine into glutamate [25]. These circRNA-driven metabolic reprogramming events affect α-ketoglutarate production and energy supply in tumor cells, thus regulating carcinogenesis [101,102]. It should be noted that such regulation of glutaminolysis by circRNAs does not exist in isolation but is intertwined with glycolysis and lipogenesis, as tumor metabolism relies on the coordination of multiple pathways. CircRNA-mediated alterations in glutamine catabolism can modify the availability of metabolites that support glucose utilization and lipid synthesis, creating a synergistic metabolic environment conducive to tumor growth.
Under conditions of nutrient deprivation, circRNAs facilitate tumor cell survival and proliferation by modulating glutamine uptake and its subsequent conversion to glutamate. Certain circRNAs enhance the adaptability of tumor cells through the regulation of enzyme expression involved in glutamine metabolism, thereby promoting mitochondrial metabolic reprogramming [103]. Additionally, these circRNAs can impact oxidative phosphorylation and the TCA cycle, thus governing the utilization of energy and metabolic intermediates stemming from glutaminolysis [104,105]. This presents novel avenues for personalized cancer therapy.
3.4 CircRNAs in Mitochondrial Dynamics and Oxidative Phosphorylation
CircRNAs are involved in mitochondrial dynamics and oxidative phosphorylation by regulating mitochondrial biogenesis, fusion/fission, and function of oxidative phosphorylation complexes, thereby affecting mitochondrial energy metabolism and cell proliferation [106,107]. For example, circMYLK4 inhibited glycolysis and promoted mitochondrial oxidative phosphorylation to regulate skeletal muscle energy metabolism via mechanisms involving voltage-gated channels and fatty acid β-oxidation in myofibers [108]. In cancer cells, circRNAs could promote mitochondrial metabolic reprogramming by modulating oxidative phosphorylation to support proliferation and survival [109,110]. Moreover, mitochondria-localized circRNAs have been implicated in the regulation of mitochondrial retrograde signaling that affects self-renewal of cancer stem cells. Key circRNAs involved in tumor metabolism regulation with their metabolic functions, targeted cancer types, and pathways are summarized in Table 1 and Fig. 4.
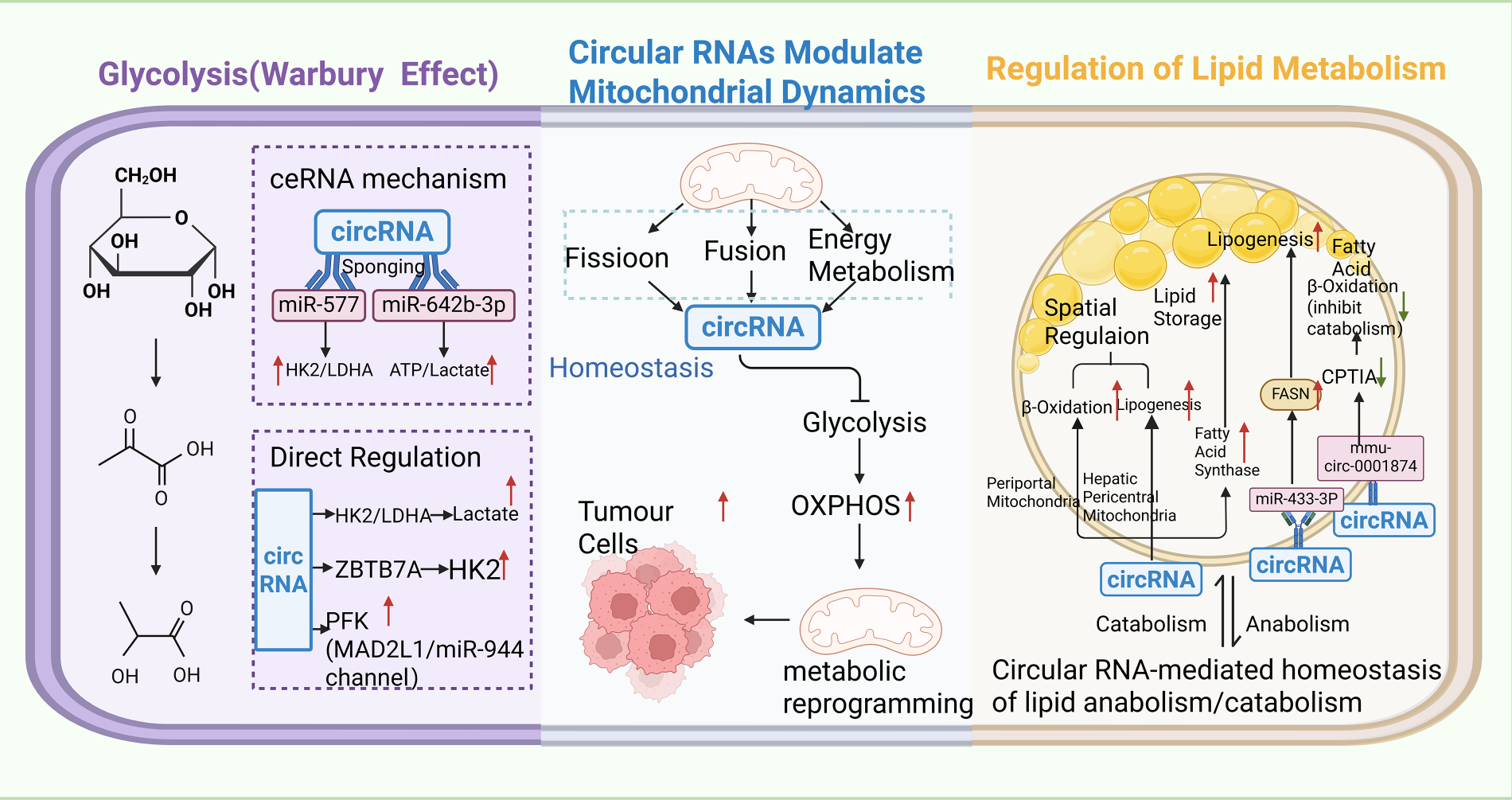
Figure 4: Mechanistic schematic of circRNA-mediated regulation of tumor metabolic reprogramming, encompassing glycolysis, lipid metabolism, and mitochondrial dynamics. CircRNAs regulate tumor metabolic reprogramming through three pathways: they act as competitive endogenous RNA (ceRNA) sponges for microRNA (miR) to affect hexokinase 2 (HK2)/lactate dehydrogenase A (LDHA), altering ATP and lactate levels, and directly bind enzymes to upregulate key glycolytic enzymes, promoting glucose conversion to pyruvate and then lactate to support glycolysis-dependent energy production. They also influence mitochondrial dynamics (fusion/fission) to regulate glycolysis and oxidative phosphorylation (OXPHOS), driving metabolic shifts. In lipid metabolism, circRNAs balance synthesis and degradation by spatially regulating mitochondrial lipid metabolism (pericentral vs. periportal regions), upregulating fatty acid synthase (FASN) to promote lipogenesis, and downregulating carnitine palmitoyltransferase 1A (CPT1A) to reduce fatty acid β-oxidation, meeting cancer cell growth demands. Additionally, genes such as zinc finger and BTB domain containing 7A (ZBTB7A) and mitotic arrest deficient 2 like 1 (MAD2L1) are involved in these regulatory networks.
4 Targeting CircRNAs for Tumor Metabolic Therapy
4.1 CircRNA-Based Therapeutic Strategies
Strategic approaches such as RNA interference (RNAi) technology, antisense oligonucleotides (ASOs), or the CRISPR-Cas system can silence oncogenic circular RNAs (circRNAs) through specific sequence recognition. RNAi tools like small interfering RNAs (siRNAs) and short hairpin RNAs (shRNAs) target circRNAs via complementary base pairing, incorporating the guide strand into the RNA-induced silencing complex (RISC) to facilitate the cleavage and downregulation of circRNAs [122]. To date, more than 20 siRNA-based therapies are in clinical trials for diseases including cancer [123]. In hepatocellular carcinoma, circRNA-SORE is upregulated in sorafenib-resistant cells; siRNAs targeting circRNA-SORE effectively silence this circRNA, significantly increasing cancer cell apoptosis and demonstrating therapeutic potential [124].
ASOs are another promising tool, consisting of unmodified or chemically modified single-stranded nucleic acid molecules (13–25 nt) that bind target RNAs through Watson–Crick base pairing to modulate their function [125]. In vivo studies have shown that ASO treatment inhibits tumor growth and alters the metabolism of tumor cells [126]. For example, ASO-circFOXK2 targets circFOXK2 in breast cancer cells and suppresses the growth of ER+ breast cancer in vivo. The antisense oligonucleotide (ASO-circFOXK2) downregulated circFOXK2 and reduced CCND1 expression at both mRNA and protein levels as well as p-RB and E2F target gene levels in BALB/C nude mice with MCF7 and drug-resistant MCF7 xenograft tumor models. This multitarget synergistic effect ultimately blocked the CCND1-CDK4/6 signaling pathway, thereby effectively inhibiting tumor cell proliferation and tumor growth [127].
While ASOs and siRNAs are widely used, they face challenges such as off-target effects. The CRISPR–Cas9 system provides a solution by efficiently targeting circRNA back-splice junctions and degrading partial or complete circRNA sequences without disrupting the function of linear coding mRNAs [128]. This revolutionary genome editing tool consists of an sgRNA and Cas9 nuclease that act as “molecular scissors”: the sgRNA–Cas9 complex recognizes PAM sequences near target DNA, guides Cas9 to induce double-strand breaks, and enables genome editing [129]. Innovations such as type II, III, and VI crRNP effector complexes have expanded CRISPR-based RNA targeting. Type VI Cas13–crRNA complexes cleaved target RNA in cis and non-target RNA in trans in vitro after binding to target RNA and undergoing conformational changes, with cleavage occurring outside the crRNA-binding region [130]. An Optimized sgRNA design for circRNA BSJ sites using CRISPR/Cas13d improved silencing specificity, which was validated through parallel screening of shRNA and CRISPR/Cas13d libraries [131]. Both long and short noncoding RNAs are now recognized as targets in precision oncology, offering robust approaches for cancer therapy (Fig. 5) [132].
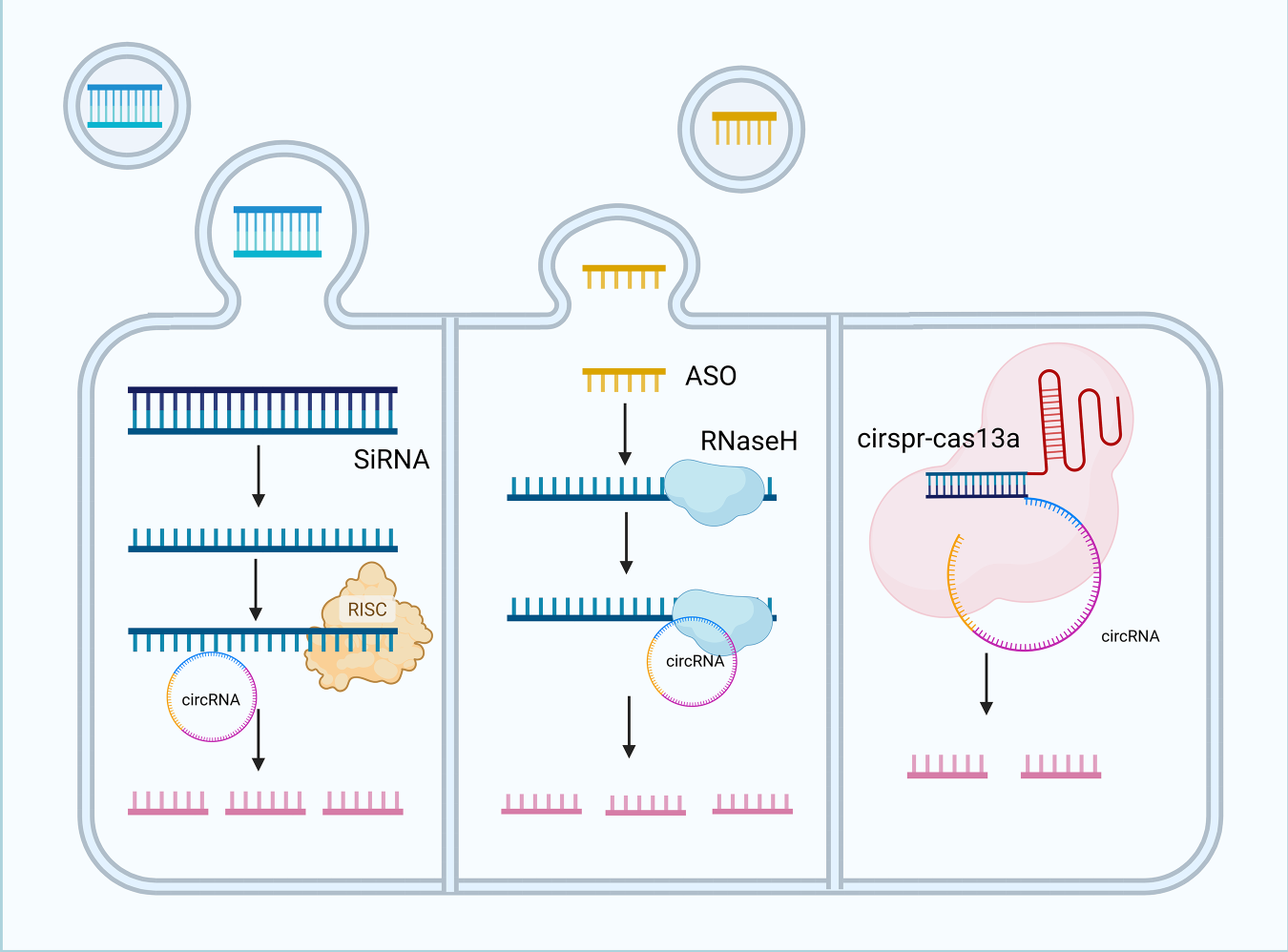
Figure 5: Strategies for targeting circular RNAs in tumor therapy. Small interfering RNA (siRNA)-mediated degradation, antisense oligonucleotide (ASO) mediated cleavage, and clustered regularly interspaced short palindromic repeats-CRISPR-associated protein 13a (CRISPR-Cas13a)-mediated disruption of circRNA functions. Three main strategies target circRNAs: siRNAs are delivered via lipid nanoparticles, bind endogenous RNA-induced silencing complex (RISC), and specifically degrade circRNAs to inhibit tumor-promoting activity. Antisense oligonucleotides form stable hybrids with circRNAs, recruiting ribonuclease H (RNaseH) to cleave circRNAs and block their functions. The CRISPR-Cas13a system uses guide RNAs to direct Cas13a to bind target circRNAs, activating its nuclease activity to degrade circRNAs and disrupt their regulatory roles in tumorigenesis and progression. siRNA, Small interfering RNA; RISC, RNA-induced silencing complex; ASO, Antisense oligonucleotide; RNaseH, Ribonuclease H; CRISPR-Cas13a, Clustered regularly interspaced short palindromic repeats-CRISPR-associated protein 13a.
Restoration of tumor suppressive circRNAs can remodel abnormal metabolism. Tumor suppressive circRNAs function as miRNA sponges through the circRNA-miRNA-mRNA regulatory network, regulate linear RNA transcription by binding to promoters or transcription factors, or interact with proteins to inhibit tumor progression [133]. For example, in bladder cancer, circFNDC3B binds miR-1178-3p to suppress tumors [134]; hsa_circ_0072309 overexpression in breast cancer inhibited miR-492 activity and circASS1 acted via the miR-4443/ASS1 axis to impede progression [135]; circPABPC1 promoted non-ubiquitin proteasomal degradation of ITGB1 to inhibit metastasis in hepatocellular carcinoma [136]; circRNA ciRS-7 bound p53 to disrupt the p53/MDM2 complex and suppress gliomagenesis [137,138]. These findings highlight circRNAs as a focal point for cancer investigations, providing new insights into tumorigenesis [139].
The development of small molecules designed to specifically restore tumor-suppressive circRNA function or regulate their expression is a promising strategy for precision oncology, although the large size of circRNAs hinders cellular uptake and necessitates the use of nanoparticle carriers. Advancing our understanding of circRNA-metabolism axes will enable the delivery of artificially synthesized or engineered circRNAs into the TME to mimic natural tumor-suppressive functions, thereby offering precision and programmability in treatment. A combined approach utilizing both circRNA inhibitors and mimics—which silences oncogenic circRNAs while replenishing tumor-suppressive ones—could facilitate comprehensive metabolic regulation and has the potential to revolutionize personalized cancer therapy.
4.2 Drug Resistance and CircRNA-Mediated Metabolism
CircRNAs contribute to drug resistance through metabolic pathways, and targeting circRNA-metabolism interactions can overcome such resistance. Tumor drug resistance—where cancer cells evade chemotherapy, thereby reducing drug efficacy—poses a major barrier to treatment [126]. CircRNAs mediate resistance through mechanisms including acting as miRNA sponges, regulating signaling pathways, modulating cancer stem cell properties, promoting epithelial-mesenchymal transition [140], and also contribute to radioresistance via metabolic reprogramming in nasopharyngeal carcinoma [141], with distinct circRNAs either enhancing or suppressing resistance by altering drug sensitivity [142].
Metabolic pathways are key mediators of this resistance. In renal cell carcinoma, bull RNA sequencing identified circME1 as significantly upregulated in resistant cells [143]. ME1 is a multifunctional enzyme involved in aerobic glycolysis, NADPH production and lipid metabolism that can promote tumor drug resistance and is cis-regulated by circME [144]. CircME1 can upregulate the expression of ME1 to enhance aerobic glycolysis in clear cell renal cell carcinoma (ccRCC) and contribute to the development of its malignant phenotype [143]. These findings indicate that circME1 facilitates ccRCC resistance to sunitinib-targeted therapy through an ME1-dependent mechanism involving metabolic reprogramming [145–147]. Similarly, circABCC4 enhances cancer-associated fibroblast (CAF)-induced oxaliplatin resistance in pancreatic cancer via glycolytic reprogramming [148]. In CRC, oxaliplatin-resistant cells transfer exosomal ciRS-122 to promote glycolysis and resistance; in lung cancer, hsa_circ_0002130 regulates glycolysis-related genes to enhance osimertinib resistance [149,150].
Targeting circRNAs can reverse drug resistance. For example, downregulated in non-small cell lung cancer (NSCLC) tissues and cell lines, circ_0002483 sequesters miR-182-5p to target GRB2, FOXO1, and FOXO3, inhibiting NSCLC proliferation and sensitizing it to paclitaxel (PTX) [151]. In PTX-resistant ovarian cancer, hsa_circ_0002782 (circSLC39A8) acts as an endogenous sponge for miR-185-5p to target BMF, restoring PTX sensitivity [152]. CircFOXO3 also sensitizes NSCLC cells to cisplatin by regulating the miR-543/Foxo3 axis and promoting glycolysis [153].
Modulation of circRNA expression—via small molecules, ASOs or CRISPR/Cas9—can enhance chemosensitivity [154]. In breast cancer, ASO-circFOXK2 inhibited ER+ tumor growth in mice and restored tamoxifen sensitivity in resistant cells [127]. Nek2-targeting siRNAs and ASOs synergized with paclitaxel and doxorubicin to promote apoptosis, suggesting that combined antisense–drug therapy is a viable strategy [155].
4.3 Nanoparticle Delivery Systems
Nanoparticle-based delivery systems facilitate the efficient and targeted administration of circRNA therapies, with notable advancements in lipid nanoparticles, stimulus-responsive nanoparticles, gold nanoparticles, and exosomes [156,157].
Exosomes, extracellular vesicles measuring 50–150 nm with an average diameter of 100 nm, are ideal carriers owing to their stability, small size, and low toxicity [158]. Their lipid bilayer protects the cargo from degradation and facilitates transport to target cells [159]. In vivo and in vitro studies have demonstrated that exosome-delivered si-ciRS-122 reverses oxaliplatin resistance and glycolysis, thereby inhibiting CRC proliferation; RGD-exo-circDIDO1 suppresses gastric cancer progression without detectable toxicity [160,161].
Lipid nanoparticles (LNPs), spherical vesicles with a phospholipid bilayer that mimic cell membranes, are capable of encapsulating drugs and releasing small interfering RNAs (siRNAs) into the cytoplasm following endocytosis. These nanoparticles safeguard RNA interference (RNAi) molecules from degradation, augment cellular uptake, and mitigate immune activation to ensure effective delivery [162]. Both in vitro cell experiments and in vivo animal model studies have substantiated that the LNP-si circPDHK1 complex, when modified with the AS1411 aptamer, can efficiently deliver therapeutic si circPDHK1 into clear cell renal cell carcinoma (ccRCC) tumor cells, significantly downregulating the expression level of the target circPDHK1 [163].
Stimuli-responsive nanoparticles release cargo in response to tumor-specific enzymes, pH, glutathione, temperature, hypoxia or reactive oxygen species (ROS) and enable targeted drug accumulation in tumors with reduced off-target effects by enhancing specificity [164,165]. Gold nanoparticles targeting circDNMT1 inhibit tumor growth and deliver tumor suppressive circRNAs such as circFoxo3 to suppress melanoma [166]. In breast cancer models, gold nanoparticles directly target circDNMT1 or disrupt its RNA-binding protein (RBP) interactions [167]; silencing of circDNMT1 via specific siRNAs or ASOs that block its p53 and Auf binding sites significantly reduces in vivo tumor growth as circDNMT1 regulates autophagy through these interactions [168]. Specific examples of circRNA-targeting strategies and their corresponding delivery systems for various cancer types are outlined in Table 2.
To fully harness the therapeutic potential of nanoparticle systems, it is imperative to address challenges related to biodistribution, pharmacokinetics, long-term safety, and material synthesis [169]. By Optimizing nanoparticle properties and investigating combined delivery strategies, we can expedite their clinical translation.
Notably, long-term safety profiles exhibit significant variation across nanoparticle platforms: lipid nanoparticles (LNPs) demonstrate potential hepatotoxicity in preclinical models due to lipid accumulation in hepatic tissues, whereas exosomes display reduced immunogenicity stemming from their endogenous origin but encounter manufacturing challenges in achieving scalable production with consistent safety parameters [170–172]. A critical unresolved challenge in nanoparticle-mediated circRNA delivery involves reconciling targeted delivery efficacy with off-target immune activation; emerging evidence indicates that carrier material immunogenicity represents not merely an adverse effect but a potentially tunable parameter that could synergize with circRNA-mediated metabolic modulation, although the mechanistic links between carrier properties and immune-metabolic crosstalk remain incompletely characterized [173,174]. While Gold nanoparticles exhibit favorable biocompatibility in short-term applications, concerns persist regarding their chronic accumulation in organs such as the spleen and kidneys, which may precipitate oxidative stress or inflammatory responses during prolonged exposure [175]. Regarding immunogenicity, a central debate centers on the equilibrium between nanoparticle-induced immune stimulation and off-target inflammatory consequences: certain investigations reveal that surface-engineered nanoparticles can attenuate complement system activation and cytokine secretion, yet conflicting reports indicate that repeated dosing may elicit adaptive immune responses targeting the carrier materials themselves, thereby compromising therapeutic effectiveness [176,177]. Mechanistically, the immunogenicity of nanoparticle carriers is closely related to their size, surface charge and chemical composition—cationic nanoparticles are more likely to interact with negatively charged cell membranes and immune receptors, thus inducing higher levels of pro-inflammatory cytokines compared to neutral or anionic counterparts [178]. An emerging hypothesis from recent studies proposes that co-delivery of circRNAs with small molecule immune modulators could synergistically suppress undesirable immune responses while maintaining targeted delivery efficiency. To address long-term safety concerns, biodegradable nanoparticle materials are being developed to minimize organ accumulation, and patient-specific dosing regimens based on individual immune profiles may further reduce risks [179]. These advances underscore the need for rigorous long-term toxicological studies and translational models that mimic clinical administration schedules to ensure clinical viability of nanoparticle-based circRNA therapies [180].
CircRNAs have the potential to indicate metabolic dysregulation in specific cancers and are promising as diagnostic biomarkers. They can be detected in plasma and saliva [76], and their stability (half-life of approximately 48 h compared with ~10 h for mRNA) makes them useful. The fact that they are often expressed in a disease-specific, tissue-specific and tumor stage-specific manner allows precise diagnosis and even prognosis. Their presence in extracellular vesicles further enables noninvasive detection by blood tests, which supports early tumor identification [186].
In gastrointestinal cancers, exosomal circRNAs related to glycolysis are upregulated or downregulated in plasma and can be used as liquid biopsy markers for early detection. For instance, circPDK1 is significantly upregulated in the plasma exosomes and tumor tissues of pancreatic cancer (PC) patients and promotes cell proliferation and glycolysis in vitro and in vivo, demonstrating its value in distinguishing PC patients from healthy controls by non-invasive means [187,188]. For example, exosomal circPDK1 has entered phase I clinical trials for the diagnosis of pancreatic cancer [189]. Similarly, hsa_circ_0000181 is markedly downregulated in gastric cancer tissues and plasma samples, indicating that it may serve as a stable diagnostic marker [190]. Many studies have shown that some circRNAs exhibit better stability and diagnostic value than their corresponding mRNAs while reflecting the tumorigenic stage, highlighting their great potential in cancer diagnosis [140]. To improve the practical application of circRNA biomarkers in different types of cancer, key selection criteria include robust stability in body fluids, distinct tissue-specific and cancer-specific expression patterns, reliable detectability through non-invasive approaches, and consistent correlation with disease progression or treatment response.
CircRNAs are associated with patient prognosis and tumor progression, making them valuable prognostic indicators. Their expression patterns can reflect cancer progression in clinical practice. For example, circ-LARP4 was downregulated in ovarian cancer (OC) tissues and significantly correlated with advanced FIGO stage, disease progression, and lymph node metastasis, positioning it as a potential prognostic marker for patients with OC [191]. Downregulation of circ_0009361 in CRC; its overexpression inhibited growth and metastasis, supporting its role as a prognostic biomarker [192]. In lung adenocarcinoma, upregulation of circPRKCI was associated with shorter overall survival in patients,thus identifying it as a poor prognostic factor [193]. A summary of the established diagnostic and prognostic values of circRNAs in various types of cancer is presented in Table 3.
Early detection and intervention are crucial for improving clinical outcomes. Advancing techniques to detect circRNA-associated metabolic dysregulation may improve the sensitivity and specificity of early diagnosis. Large-scale clinical studies investigating the associations between circRNA expression profiles and prognosis, coupled with the development of accurate prognostic models, will provide a solid basis for clinical decision-making and patient management.
The integration of circRNA-targeted therapies with existing cancer treatment regimens holds significant promise [210]. Synthetic circRNAs have shown potential in protein expression, vaccine development, and RNA editing [211]. Non-protein/peptide-encoding synthetic circRNAs have been tested as therapeutic agents based on their physiological functions.
One application is to use synthetic circRNAs as miRNA or protein sponge. For example, a synthetic circRNA acting as a miR-21 sponge inhibited gastric cancer cell proliferation and downstream protein activation [212]. In gene editing, synthetic circRNAs can function as small interfering RNA (siRNA). dumbbell-shaped circRNAs consisting of two loops and a stem are specifically recognized and processed by the endoribonuclease Dicer into double-stranded siRNAs that mediate target mRNA interference [213]. Another design, circular siRNAs composed of circular sense RNA and its linear complementary RNA, enhanced gene silencing efficiency and reduced off-target effects compared with conventional siRNAs [214].
ASOs can target circRNAs by complementary pairing. Although seldom used to target back-splice junctions for circRNA knockdown due to their length, they efficiently block protein interaction sites on circRNAs [215]. Furthermore, the CRISPR/Cas13 system enables more specific circRNA knockdown than RNAi due to its low mismatch tolerance, although its in vivo efficiency remains to be determined [216].
CircRNAs also exert therapeutic effects through protein interactions. For example, circGSK3β promotes migration and invasion in esophageal squamous cell carcinoma (ESCC) by reducing β-catenin phosphorylation via decreasing GSK3β and its subsequent ubiquitination [217]. CircHuR inhibits gastric cancer (GC) proliferation, invasion and metastasis by sequestering CCHC-type zinc finger nucleic acid binding protein (CNBP) from the promoter of human antigen R (HuR), leading to downregulation of HuR expression [218].
Targeting circRNAs or their associated pathways, which have been implicated in oncogenesis, offers therapeutic potential. For instance, circHIAT1 and circATP2B1 are key regulators of RCC development, rendering their signaling pathways as promising targets for metastatic RCC therapy [219].
Protein-encoding synthetic circRNAs hold promise for applications in metabolic cancer vaccines [220]. While mRNA vaccines provide high immunogenicity and ease of production, their inherent instability and limited duration of protein expression compromise efficacy. Single-cell omics technology, through the analysis of the cellular heterogeneity of tumors and the immune microenvironment, offers vital insights for the design of personalized vaccine strategies targeting specific patient subgroups or the overcoming of immunosuppressive microenvironments [221]. Cyclization of linear RNA into circRNA improves stability, and encapsulation of antigen-encoding circRNA within lipid nanoparticles (LNPs) elicits potent innate and adaptive immune responses, thereby establishing a novel circRNA vaccine platform [222].
A recent study reported a circRNA-based vaccine for melanoma, which was synthesized in vitro using a type I intron-mediated PIE system and incorporated the CVB3 IRES element to facilitate translation [223]. Formulated with LNPs, this vaccine demonstrated superior therapeutic efficacy against “immune-excluded” MC38 and “immune-desert” B16 orthotopic tumor models in mice at one week post injection. Remarkably, the OVA-encoding melaoma circRNA vaccine also showed preventive effects by protecting naive hosts [142,224]. Although circRNA vaccines can induce dendritic cell maturation and activate both innate and adaptive immunity, further mechanistic studies and optimization are required to improve the suboptimal antibody response induced by CART-circOVA before it can be advanced as cancer immunotherapy [225,226].
Traditional mRNA vaccines have shown promise in tumor immunotherapy, yet they are hindered by inherent limitations such as poor stability, dependence on nucleotide modifications for functionality, and complex regulation of immunogenicity. In contrast, the distinct covalently closed-loop structure of circRNA vaccines, which lacks 5′/3′ termini, confers resistance to RNase degradation, resulting in superior stability compared to linear mRNA and notable longevity [173,227]. Furthermore, circRNA vaccines exhibit enhanced safety profiles, with no observed cytokine storm during immune activation, while effectively stimulating dendritic cell activity, robust antigen-specific CD8 T cell responses in lymph nodes and tissues, and potent antitumor effects, positioning them as promising therapeutic cancer vaccines [225]. Consequently, circRNA vaccines outperform traditional linear mRNA vaccines in terms of stability, translational efficiency, immune activation potential, and antitumor efficacy, offering a safer and more effective platform for tumor immunotherapy and potentially accelerating the clinical translation of RNA vaccines. Nonetheless, challenges remain, including low in vitro cyclization efficiency, potential interference from residual linear fragments, and unresolved concerns regarding clinical research, long-term human safety, and standardization of large-scale production processes [228,229].
6 Challenges and Future Directions
Despite the growing recognition of circRNAs in recent investigations, their study is confronted with formidable challenges, including complex mechanisms and limitations in manipulation.
CircRNAs regulate glucose and lipid metabolism through diverse mechanisms, including miRNA sponging, protein interactions, and protein translation, forming highly complex regulatory networks. they are also involved in cancer cell metabolism, and understanding these processes is crucial for identifying innovative therapeutic approaches [1,15,98]. For instance, circDDX21 is upregulated by transcription factors such as HIF-1α under glucose deprivation, influencing metabolic reprogramming in hepatocytes and driving carcinogenesis [230]. Notably, the mechanisms of circRNAs vary significantly across different metabolic diseases, necessitating tissue-specific analyses to clarify their roles and guide personalized therapeutic strategies, which adds complexity to cancer translational research [25,98,231,232]. To address this complexity, actionable research directions include integrating multi-omics data to map comprehensive circRNA-mRNA-miRNA-protein interaction networks, developing specialized bioinformatics tools to decode context-dependent regulatory patterns, and constructing dynamic computational models to simulate how circRNA-mediated metabolic regulation responds to cellular or environmental changes [233].
Furthermore, tools for precise circRNA manipulation are still limited and the controversy of how to reconcile the specificity of circRNA targeting with the pleiotropy of metabolic pathways remains a critical gap in knowledge that hinders translational progress. The lack of tissue-specific, spatiotemporally controllable editing systems not only prevents accurate manipulation but also raises questions about whether it is even possible to achieve sustained metabolic reprogramming by targeting individual circRNAs without disrupting physiological metabolic homeostasis [234]. The CRISPR/Cas9 gene editing system faces two core technical challenges: first, it remains significantly difficult to accurately predict off-target effects; second, editing efficiency is affected by multiple factors such as sgRNA design quality and delivery system selection. More importantly, there is an unavoidable tradeoff between reducing off-target activity and maintaining sufficient on-target cleavage efficiency [235,236].
Overcoming drug resistance is another critical challenge; although ASO delivery shows promise in reversing resistance, targeting circRNAs offers a complementary strategy to counter treatment resistance, such approaches are still in the early stages of clinical trials [237]. Although circRNAs hold potential as diagnostic and prognostic biomarkers [238], substantial improvements are needed in bioinformatics analysis of circRNA sequencing data, analytical precision, and subsequent precise regulation—underscoring the long and challenging path ahead in circRNA investigations.
Research on circRNAs is rapidly evolving with emerging future directions. One focus is the identification of metabolism-related circRNAs using high-throughput technologies. The integration of single-cell sequencing, spatial transcriptomics and multi-omics approaches can systematically dissect dynamic circRNA expression profiles in metabolic diseases [239,240]. For example, 164 differentially expressed circRNAs associated with obesity have been identified by high-throughput sequencing. In addition, artificial intelligence could predict circRNA-metabolic target interaction networks to accelerate biomarker discovery and has great potential for improving therapeutic efficacy and prognosis [241].
Another direction is to conduct clinical trials of circRNA-based metabolic therapy. This includes developing targeted strategies, such as circRNA inhibitors (for example, siRNAs or antisense oligonucleotides) or mimics (synthetic circRNAs), to regulate key metabolic genes (such as GLUT1 and HK2) [2,15,83]. In addition, combining nanodelivery technologies to explore their roles in regulating drug metabolism, therapeutic efficacy and toxicity profiles will improve the targeting and safety of circRNA therapy [237,242].
For clinical translation, it is advisable to prioritize the validation of circRNA therapeutics in metabolic cancers (for example, pancreatic and liver cancers) given that there are ongoing clinical trials targeting lipid metabolism pathways (for example, ANGPTL3/8 antibodies) [243–246], further investigation into the application of circRNA in CAR-T cell therapy—such as enhancing metabolic reprogramming through electroporation-mediated delivery of circRNA—demonstrates the potential of circRNA-based technologies to advance cellular therapies [242,247] (Table 4).
Although this review systematically summarizes the critical roles of circRNA in tumor metabolic reprogramming and its clinical translational potential, several limitations remain. Firstly, circRNA exhibits significant functional heterogeneity across different tumor types and tissue microenvironments, with its upstream regulatory signals and downstream effector networks not yet fully elucidated, posing challenges for precise intervention targeting specific circRNAs. Secondly, most current studies remain at the preclinical stage, where the efficacy and safety of circRNA as therapeutic targets or diagnostic biomarkers have not been adequately validated through large-scale clinical trials. Finally, existing circRNA manipulation tools still face deficiencies in specificity, delivery efficiency, and in vivo stability, while the long-term biosafety, immunogenicity, and targeting efficiency of nanodelivery systems require further comprehensive evaluation. Future research should integrate multi-omics data, develop more precise regulatory tools, and advance prospective clinical studies to facilitate the translation of circRNA from basic research toward practical applications in tumor metabolism therapy.
CircRNAs contribute to tumor metabolic reprogramming by modulating key pathways including glycolysis, lipid metabolism, and glutaminolysis. Their inherent stability and tissue-specific expression profiles establish them as promising diagnostic biomarkers for applications in early detection and disease monitoring, while demonstrating significant correlations with clinical outcomes such as chemotherapy resistance and tumor progression. As therapeutic targets, circRNAs present opportunities for metabolic modulation through silencing oncogenic circRNAs using siRNA, ASOs, or CRISPR-based tools, or restoring tumor-suppressive circRNA expression. Advancements in nanoparticle delivery systems have further expanded their translational potential. Despite existing delivery challenges, future research should prioritize resolving the context-dependent functionality of circRNAs and optimizing LNP-circRNA complexes for phase III trials in hepatocellular carcinoma. This review proposes A novel hypothesis that circRNA-mediated metabolic regulation could function as a “metabolic checkpoint” to selectively reprogram tumor metabolism without affecting normal cells, exploiting the unique stability and tissue specificity of circRNAs to target the interconnectedness of glycolysis, lipid metabolism, and glutaminolysis—a mechanistic insight with transformative potential for developing circRNA-based therapeutic strategies.
Acknowledgement: Not applicable.
Funding Statement: This research was funded by National Natural Science Foundation of China (82360801).
Author Contributions: The authors confirm contribution to the paper as follows: conceptualization, Qian Zhang, Chen Li, Mengyao Li; investigation, Yimao Wu, Yitong Liu, Ruowei Sun; writing—original draft preparation, Yimao Wu, Yitong Liu, Ruowei Sun; writing—review and editing, Yimao Wu, Yitong Liu, Ruowei Sun, Yiyuan Zhang, Qian Zhang, Chen Li, Mengyao Li; visualization, Yimao Wu, Yitong Liu, Ruowei Sun; supervision, Qian Zhang, Chen Li, Mengyao Li; project administration, Qian Zhang, Chen Li, Mengyao Li. All authors reviewed and approved the final version of the manuscript.
Availability of Data and Materials: Not applicable.
Ethics Approval: Not applicable.
Conflicts of Interest: The authors declare no conflicts of interest.
Abbreviations
| 5-FU | 5-Fluorouracil |
| ACC | Acetyl-CoA carboxylase |
| ADARB1 | Adenosine deaminase RNA specific B1 |
| ANGPTL3/8 | Angiopoietin-like 3/8 |
| APC | Adenomatous polyposis coli |
| AUC | Area under the ROC curve |
| AS1411 | Aptamer AS1411 |
| ASOs | Antisense oligonucleotides |
| ATP | Adenosine triphosphate |
| B16 | Mouse melanoma cell line |
| BET | Bromodomain and extra-terminal domain |
| BMF | Bcl-2 modifying factor |
| BRAF | B-Raf proto-oncogene |
| BSJ | Back-splice junction |
| CAFs | Cancer-associated fibroblasts |
| CAR-T | Chimeric antigen receptor T-cell |
| CCND1 | Cyclin D1 |
| ccRCC | Clear cell renal cell carcinoma |
| CD36 | Cluster of differentiation 36 |
| CDK4/6 | Cyclin-dependent kinase 4/6 |
| CDKN2B-AS1 | CDKN2B antisense RNA 1 |
| CREB5 | cAMP-responsive element-binding protein 5 |
| ciRS-7 | Circular RNA sponge for miR-7 |
| CNBP | CCHC-type zinc finger nucleic acid binding protein |
| circPVT1 | Circular RNA plasmacytoma variant translocation 1 |
| CoA-SH | Coenzyme A (thiol form) |
| CRISPR-Cas | Clustered regularly interspaced short palindromic repeats-CRISPR-associated protein |
| CRISPR/Cas13d | Clustered regularly interspaced short palindromic repeats/CAS13d |
| crRNP | CRISPR ribonucleoprotein |
| CRC | Colorectal cancer |
| CSCs | Cancer Stem Cells |
| DCR | Disease control rate |
| DFS | Disease-free survival |
| DLL3 | Delta-like ligand 3 |
| DRP1 | Dynamin-related protein 1 |
| DNMT1 | DNA methyltransferase 1 |
| ECM | Extracellular matrix |
| ECE1 | Endothelin-converting enzyme 1 |
| ENO1 | Enolase 1 |
| ESCC | Esophageal squamous cell carcinoma |
| FABPs | Fatty acid-binding proteins |
| FASN | Fatty acid synthase |
| FIGO | International Federation of Gynecology and Obstetrics |
| FISH | Fluorescence in situ hybridization |
| FIS1 | Fission protein 1 |
| FOXO3 | Forkhead box O3 |
| FOXP1 | Forkhead box P1 |
| FADH2 | Flavin adenine dinucleotide hydrogen |
| GC | Gastric cancer |
| GBM | Glioblastoma |
| GEO | Gene Expression Omnibus |
| GLS | Glutaminase |
| GLUT1 | Glucose transporter 1 |
| GO | Gene Ontology |
| HA | Hyaluronan |
| HDACs | Histone deacetylases |
| HATs | Histone acetyltransferases |
| HIF-1α | Hypoxia-inducible factor 1 alpha |
| HIPK3 | Homeodomain-interacting protein kinase 3 |
| HK2 | Hexokinase 2 |
| HKDC1 | Hexokinase domain component 1 |
| HMGCS1 | 3-Hydroxy-3-methylglutaryl-CoA synthase 1 |
| HMTs | Histone methyltransferases |
| HuR | Human antigen R |
| IGF | Insulin-like growth factor |
| IGF2 | Insulin-like growth factor 2 |
| IGF2BP2 | Insulin-like growth factor 2 mRNA-binding protein 2 |
| IHC | Immunohistochemistry |
| IMP3 | Insulin-like growth factor 2 mRNA-binding protein 3 |
| ITCH | Itchy E3 ubiquitin protein ligase |
| KEGG | Kyoto Encyclopedia of Genes and Genomes |
| LDH | Lactate dehydrogenase |
| lncRNAs | Long non-coding RNAs |
| LMO7 | LIM domain only 7 |
| LNPs | Lipid nanoparticles |
| LXR | Liver X receptor |
| m6A | N6-methyladenosine |
| MAPK/ERK | Mitogen-activated protein kinase/Extracellular signal-regulated kinase |
| MAPK/JNK | Mitogen-activated protein kinase/c-Jun N-terminal kinase |
| MAD2L1 | Mitotic arrest deficient 2 like 1 |
| MC38 | Mouse colon adenocarcinoma cell line |
| MFF | Mitochondrial fission factor |
| MID49/51 | Mitofusin 49/51 kDa |
| MFN1/2 | Mitofusins 1/2 |
| mdivi-1 | Mitochondrial division inhibitor 1 |
| mtDNA | Mitochondrial DNA |
| mTOR | Mammalian target of rapamycin |
| mTORC1 | Mammalian target of rapamycin complex 1 |
| NADH | Nicotinamide adenine dinucleotide hydrogen |
| NAFLD | Non-alcoholic fatty liver disease |
| NPC | Nasopharyngeal carcinoma |
| NSCLC | Non-small cell lung cancer |
| OPA1 | Optic atrophy 1 |
| ORR | Objective response rate |
| OXPHOS | Oxidative phosphorylation |
| O-GlcNAcylation | O-linked β-N-acetylglucosamine modification |
| AMPK | Adenosine 5′-monophosphate-activated protein kinase |
| PI3K/AKT | Phosphatidylinositol 3-kinase/Protein kinase B |
| PD-1 | Programmed death 1 |
| PD-L1 | Programmed death-ligand 1 |
| PDK4 | Pyruvate dehydrogenase kinase 4 |
| PFK | Phosphofructokinase |
| PIE | Protein-intron-exon |
| PKLR | Pyruvate kinase, liver and RBC |
| PKM2 | Pyruvate kinase M2 |
| PPARs | Peroxisome proliferator-activated receptors |
| PPARγ | Peroxisome proliferator-activated receptor gamma |
| PPA2c | Pyruvate phosphate dikinase 2 cytoplasmic |
| PRMT5 | Protein arginine methyltransferase 5 |
| PTEN | Phosphatase and tensin homolog |
| PTBP1 | Polypyrimidine tract binding protein 1 |
| PTX | Paclitaxel |
| PFS | Progression-free survival |
| RAS | RAS oncogene |
| RHBDD1 | RH domain containing DEAD box protein 1 |
| RBP | RNA-binding protein |
| RNaseH | Ribonuclease H |
| RNAi | RNA interference |
| ROC | Receiver operating characteristic |
| RTKN2 | Rhotekin 2 |
| RISC | RNA-induced silencing complex |
| SAM | S-adenosylmethionine |
| SCD1 | Stearoyl-CoA desaturase 1 |
| SCNT | Single-walled carbon nanotube |
| sgRNA | Single-guide RNA |
| SLC1A5 | Solute carrier family 1 member 5 |
| SLIT2 | Slit guidance ligand 2 |
| SREBPs | Sterol regulatory element-binding proteins |
| SNRPA | Small nuclear ribonucleoprotein polypeptide A |
| siRNAs | Small interfering RNAs |
| shRNAs | Short hairpin RNAs |
| TAMs | Tumor-associated macrophages |
| TCA | Tricarboxylic acid cycle |
| TCGA | The Cancer Genome Atlas |
| TME | Tumor microenvironment |
| TNBC | Triple-negative breast cancer |
| TXNIP | Thioredoxin interacting protein |
| TXNDC5 | Thioredoxin domain-containing protein 5 |
| UBAP2 | Ubiquitin associated protein 2 |
| VAPA | Vesicle-associated membrane protein-associated protein A |
| Wnt/β-catenin | Wnt/β-catenin signaling pathway |
| YAP1 | Yes-associated protein 1 |
| YBX1 | Y-box binding protein 1 |
| ZBTB7A | Zinc finger and BTB domain containing 7A |
| ZFR | Zinc finger RNA binding protein |
| α-KG | α-Ketoglutarate |
References
1. Cai ZR, Hu Y, Liao K, Li H, Chen DL, Ju HQ. Circular RNAs: emerging regulators of glucose metabolism in cancer. Cancer Lett. 2023;552:215978. doi:10.1016/j.canlet.2022.215978. [Google Scholar] [PubMed] [CrossRef]
2. Wang Y, Chen X, Yang Y. CircRNA-regulated glucose metabolism in ovarian cancer: an emerging landscape for therapeutic intervention. Clin Transl Oncol. 2023;26(3):584–96. doi:10.1007/s12094-023-03285-4. [Google Scholar] [PubMed] [CrossRef]
3. Quail D, Joyce J. Microenvironmental regulation of tumor progression and metastasis. Nat Med. 2013;19(11):1423–37. doi:10.1038/nm.3394. [Google Scholar] [PubMed] [CrossRef]
4. Liu Y, Jiang C, Liu Q, Huang R, Wang M, Guo X. CircRNAs: emerging factors for regulating glucose metabolism in colorectal cancer. Clin Transl Oncol. 2023;25(8):2321–31. doi:10.1007/s12094-023-03131-7. [Google Scholar] [PubMed] [CrossRef]
5. Li Z, Li X, Xu D, Chen X, Li S, Zhang L, et al. An update on the roles of circular RNAs in osteosarcoma. Cell Prolif. 2021;54(1):e12936. doi:10.1111/cpr.12936. [Google Scholar] [PubMed] [CrossRef]
6. Geng Y, Wang M, Wu Z, Jia J, Yang T, Yu L. Research progress of circRNA in malignant tumour metabolic reprogramming. RNA Biol. 2023;20(1):641–51. doi:10.1080/15476286.2023.2247877. [Google Scholar] [PubMed] [CrossRef]
7. Wilusz JE. A 360° view of circular RNAs: from biogenesis to functions. WIREs RNA. 2018;9(4):e1478. doi:10.1002/wrna.1478. [Google Scholar] [PubMed] [CrossRef]
8. Shafaghat Z, Radmehr S, Saharkhiz S, Khosrozadeh A, Feiz K, Alkhathami AG, et al. Circular RNA, a molecule with potential chemistry and applications in RNA-based cancer therapeutics: an insight into recent advances. Top Curr Chem. 2025;383(2):21. doi:10.1007/s41061-025-00505-z. [Google Scholar] [PubMed] [CrossRef]
9. Chen LL, Yang L. Regulation of circRNA biogenesis. RNA Biol. 2015;12(4):381–8. doi:10.1080/15476286.2015.1020271. [Google Scholar] [PubMed] [CrossRef]
10. Alkhathami AG, Sahib AS, Al Fayi MS, Fadhil AA, Jawad MA, Shafik SA, et al. Glycolysis in human cancers: emphasis circRNA/glycolysis axis and nanoparticles in glycolysis regulation in cancer therapy. Environ Res. 2023;234(2):116007. doi:10.1016/j.envres.2023.116007. [Google Scholar] [PubMed] [CrossRef]
11. Wang J, Zhang Y, Li Z. Advancements in understanding the role of circular RNA in osteosarcoma. Mol Biotechnol. 2024;66(9):2157–67. doi:10.1007/s12033-023-00838-4. [Google Scholar] [PubMed] [CrossRef]
12. Jiao J, Zhao Y, Li Q, Jin S, Liu Z. LncRNAs in tumor metabolic reprogramming and tumor microenvironment remodeling. Front Immunol. 2024;15:1467151. doi:10.3389/fimmu.2024.1467151. [Google Scholar] [PubMed] [CrossRef]
13. Li Y, Huang H, Wu S, Zhou Y, Huang T, Jiang J. The role of RNA m6A modification in cancer glycolytic reprogramming. Curr Gene Ther. 2023;23(1):51–9. doi:10.2174/1566523222666220830150446. [Google Scholar] [PubMed] [CrossRef]
14. Huang H, Weng H, Chen J. m6A modification in coding and non-coding RNAs: roles and therapeutic implications in cancer. Cancer Cell. 2020;37(3):270–88. doi:10.1016/j.ccell.2020.02.004. [Google Scholar] [PubMed] [CrossRef]
15. Jiang M, Zhang K, Zhang Z, Zeng X, Huang Z, Qin P, et al. PI3K/AKT/mTOR axis in cancer: from pathogenesis to treatment. MedComm. 2025;6(8):e70295. doi:10.1002/mco2.70295. [Google Scholar] [PubMed] [CrossRef]
16. Li Q, Wang Y, Wu S, Zhou Z, Ding X, Shi R, et al. CircACC1 regulates assembly and activation of AMPK complex under metabolic stress. Cell Metab. 2019;30(1):157–73.e7. doi:10.1016/j.cmet.2019.05.009. [Google Scholar] [PubMed] [CrossRef]
17. Ma J, Xu J, Zhang X, Quan J. The emerging role and clinical significance of circRNAs in papillary thyroid cancer. Front Endocrinol. 2024;15:1351776. doi:10.3389/fendo.2024.1351776. [Google Scholar] [PubMed] [CrossRef]
18. Yang Y, Luo J, Wang Z, Liu K, Feng K, Wang F, et al. Energy stress–induced circEPB41(2) promotes lipogenesis in hepatocellular carcinoma. Cancer Res. 2025;85(4):723–38. doi:10.1158/0008-5472.CAN-24-1630. [Google Scholar] [PubMed] [CrossRef]
19. Mafi A, Hedayati N, Milasi YE, Kahkesh S, Daviran M, Farahani N, et al. The function and mechanism of circRNAs in 5-fluorouracil resistance in tumors: biological mechanisms and future potential. Pathol Res Pract. 2024;260(11):155457. doi:10.1016/j.prp.2024.155457. [Google Scholar] [PubMed] [CrossRef]
20. He J, Xie Q, Xu H, Li J, Li Y. Circular RNAs and cancer. Cancer Lett. 2017;396:138–44. doi:10.1016/j.canlet.2017.03.027. [Google Scholar] [PubMed] [CrossRef]
21. Wei Z, Shi Y, Xue C, Li M, Wei J, Li G, et al. Understanding the dual roles of CircHIPK3 in tumorigenesis and tumor progression. J Cancer. 2022;13(15):3674–86. doi:10.7150/jca.78090. [Google Scholar] [PubMed] [CrossRef]
22. Campelo MM, Reis-das-Mercês L, Vidal AF, Da Silva FRP, De Oliveira ACA, Monteiro JRDS, et al. The dual role of circHIPK3 in cancer and its implications for multiple drugs resistance: a systematic review and computational approach. Front Oncol. 2025;15:1547889. doi:10.3389/fonc.2025.1547889. [Google Scholar] [PubMed] [CrossRef]
23. Liu Q, Li S. Exosomal circRNAs: novel biomarkers and therapeutic targets for urinary tumors. Cancer Lett. 2024;588(4):216759. doi:10.1016/j.canlet.2024.216759. [Google Scholar] [PubMed] [CrossRef]
24. Gao QX, Wu QQ, Dong ZY, Qin XY, Zhai LY, Xie BP, et al. In situ detection of piRNA-651 in exosomes and cells for cancer diagnosis by a new gold nanoparticle nucleic acid probe. Interdiscip Med. 2025;3(1):e20240058. doi:10.1002/INMD.20240058. [Google Scholar] [CrossRef]
25. Palcau AC, Pulito C, De Pascale V, Casadei L, Valerio M, Sacconi A, et al. CircPVT1 weakens miR-33a-5p unleashing the c-MYC/GLS1 metabolic axis in breast cancer. J Exp Clin Cancer Res. 2025;44(1):100. doi:10.1186/s13046-025-03355-1. [Google Scholar] [PubMed] [CrossRef]
26. Yang L, Yi Y, Mei Z, Huang D, Tang S, Hu L, et al. Circular RNAs in cancer stem cells: insights into their roles and mechanisms. Int J Mol Med. 2025;55(3):50. doi:10.3892/ijmm.2025.5491. [Google Scholar] [PubMed] [CrossRef]
27. Liu W, Sun Y, Huo Y, Zhang L, Zhang N, Yang M. Circular RNAs in lung cancer: implications for preventing therapeutic resistance. EBioMedicine. 2024;107:105309. doi:10.1016/j.ebiom.2024.105309. [Google Scholar] [PubMed] [CrossRef]
28. Yoshida GJ. Metabolic reprogramming: the emerging concept and associated therapeutic strategies. J Exp Clin Cancer Res. 2015;34(1):111. doi:10.1186/s13046-015-0221-y. [Google Scholar] [PubMed] [CrossRef]
29. Tufail M, Jiang CH, Li N. Altered metabolism in cancer: insights into energy pathways and therapeutic targets. Mol Cancer. 2024;23(1):203. doi:10.1186/s12943-024-02119-3. [Google Scholar] [PubMed] [CrossRef]
30. Fukushi A, Kim HD, Chang YC, Kim CH. Revisited metabolic control and reprogramming cancers by means of the Warburg effect in tumor cells. Int J Mol Sci. 2022;23(17):10037. doi:10.3390/ijms231710037. [Google Scholar] [PubMed] [CrossRef]
31. Zhou Z, Ibekwe E, Chornenkyy Y. Metabolic alterations in cancer cells and the emerging role of oncometabolites as drivers of neoplastic change. Antioxidants. 2018;7(1):16. doi:10.3390/antiox7010016. [Google Scholar] [PubMed] [CrossRef]
32. Abdel-Wahab AF, Mahmoud W, Al-Harizy RM. Targeting glucose metabolism to suppress cancer progression: prospective of anti-glycolytic cancer therapy. Pharmacol Res. 2019;150(18):104511. doi:10.1016/j.phrs.2019.104511. [Google Scholar] [PubMed] [CrossRef]
33. Kooshan Z, Cárdenas-Piedra L, Clements J, Batra J. Glycolysis, the sweet appetite of the tumor microenvironment. Cancer Lett. 2024;600(4):217156. doi:10.1016/j.canlet.2024.217156. [Google Scholar] [PubMed] [CrossRef]
34. Barba I, Carrillo-Bosch L, Seoane J. Targeting the Warburg effect in cancer: where do we stand? Int J Mol Sci. 2024;25(6):3142. doi:10.3390/ijms25063142. [Google Scholar] [PubMed] [CrossRef]
35. Papa S, Choy PM, Bubici C. The ERK and JNK pathways in the regulation of metabolic reprogramming. Oncogene. 2019;38(13):2223–40. doi:10.1038/s41388-018-0582-8. [Google Scholar] [PubMed] [CrossRef]
36. Fu Y, Zou T, Shen X, Nelson PJ, Li J, Wu C, et al. Lipid metabolism in cancer progression and therapeutic strategies. MedComm. 2021;2(1):27–59. doi:10.1002/mco2.27. [Google Scholar] [PubMed] [CrossRef]
37. Butler LM, Perone Y, Dehairs J, Lupien LE, De Laat V, Talebi A, et al. Lipids and cancer: emerging roles in pathogenesis, diagnosis and therapeutic intervention. Adv Drug Deliv Rev. 2020;159(10):245–93. doi:10.1016/j.addr.2020.07.013. [Google Scholar] [PubMed] [CrossRef]
38. Cheng C, Geng F, Cheng X, Guo D. Lipid metabolism reprogramming and its potential targets in cancer. Cancer Commun. 2018;38(1):1–14. doi:10.1186/s40880-018-0301-4. [Google Scholar] [PubMed] [CrossRef]
39. Li B, Mi J, Yuan Q. Fatty acid metabolism-related enzymes in colorectal cancer metastasis: from biological function to molecular mechanism. Cell Death Discov. 2024;10(1):350. doi:10.1038/s41420-024-02126-9. [Google Scholar] [PubMed] [CrossRef]
40. Liu X, Lu J, Ni X, He Y, Wang J, Deng Z, et al. FASN Promotes lipid metabolism and progression in colorectal cancer via the SP1/PLA2G4B axis. Cell Death Discov. 2025;11(1):122. doi:10.1038/s41420-025-02409-9. [Google Scholar] [PubMed] [CrossRef]
41. Liu W, Dong S, Hao F, Gao Y, Wei Q. Lipid metabolic reprogramming in colorectal cancer: mechanisms and therapeutic strategies. Front Immunol. 2025;16:1603032. doi:10.3389/fimmu.2025.1603032. [Google Scholar] [PubMed] [CrossRef]
42. Zhuang Z, Chen Y, Yao Y, Zhu X. Metabolic reprogramming in colorectal cancer: the impact of fatty acid metabolism. Hum Mutat. 2025;2025(1):9567214. doi:10.1155/humu/9567214. [Google Scholar] [PubMed] [CrossRef]
43. Yang K, Wang X, Song C, He Z, Wang R, Xu Y, et al. The role of lipid metabolic reprogramming in tumor microenvironment. Theranostics. 2023;13(6):1774–808. doi:10.7150/thno.82920. [Google Scholar] [PubMed] [CrossRef]
44. Wu Y, Chen Z, Lee J, Zengin G, Li MY. Circular RNAs: epigenetic puppeteers pulling the strings of cancer pathogenesis. Med Bull. 2026;2(1):124–36. doi:10.1002/mdb2.70022. [Google Scholar] [CrossRef]
45. Frattaruolo L, Brindisi M, Curcio R, Marra F, Dolce V, Cappello AR. Targeting the mitochondrial metabolic network: a promising strategy in cancer treatment. Int J Mol Sci. 2020;21(17):6014. doi:10.3390/ijms21176014. [Google Scholar] [PubMed] [CrossRef]
46. Carles-Fontana R, Heaton N, Palma E, Khorsandi S. Extracellular vesicle-mediated mitochondrial reprogramming in cancer. Cancers. 2022;14(8):1865. doi:10.3390/cancers14081865. [Google Scholar] [PubMed] [CrossRef]
47. Brahimi-Horn MC, Giuliano S, Saland E, Lacas-Gervais S, Sheiko T, Pelletier J, et al. Knockout of Vdac1 activates hypoxia-inducible factor through reactive oxygen species generation and induces tumor growth by promoting metabolic reprogramming and inflammation. Cancer Metab. 2015;3(1):8. doi:10.1186/s40170-015-0133-5. [Google Scholar] [PubMed] [CrossRef]
48. Wu Z, Xiao C, Long J, Huang W, You F, Li X. Mitochondrial dynamics and colorectal cancer biology: mechanisms and potential targets. Cell Commun Signal. 2024;22(1):91. doi:10.1186/s12964-024-01490-4. [Google Scholar] [PubMed] [CrossRef]
49. Wu Z, Xiao C, Li F, Huang W, You F, Li X. Mitochondrial fusion–fission dynamics and its involvement in colorectal cancer. Mol Oncol. 2024;18(5):1058–75. doi:10.1002/1878-0261.13578. [Google Scholar] [PubMed] [CrossRef]
50. Iwata W, Haggerty N, Sesaki H, Iijima M. Targeting mitochondrial structure and dynamics for therapeutic intervention in cancer. PLoS Biol. 2025;23(10):e3003453. doi:10.1371/journal.pbio.3003453. [Google Scholar] [PubMed] [CrossRef]
51. Dai W, Wang G, Chwa J, Oh ME, Abeywardana T, Yang Y, et al. Mitochondrial division inhibitor (Mdivi-1) decreases oxidative metabolism in cancer. Br J Cancer. 2020;122(9):1288–97. doi:10.1038/s41416-020-0778-x. [Google Scholar] [PubMed] [CrossRef]
52. Li X, Peng X, Li Y, Wei S, He G, Liu J, et al. Glutamine addiction in tumor cell: oncogene regulation and clinical treatment. Cell Commun Signal. 2024;22(1):12. doi:10.1186/s12964-023-01449-x. [Google Scholar] [PubMed] [CrossRef]
53. Smith B, Schafer XL, Ambeskovic A, Spencer CM, Land H, Munger J. Addiction to coupling of the Warburg effect with glutamine catabolism in cancer cells. Cell Rep. 2016;17(3):821–36. doi:10.1016/j.celrep.2016.09.045. [Google Scholar] [PubMed] [CrossRef]
54. Yang L, Venneti S, Nagrath D. Glutaminolysis: a hallmark of cancer metabolism. Annu Rev Biomed Eng. 2017;19(1):163–94. doi:10.1146/annurev-bioeng-071516-044546. [Google Scholar] [PubMed] [CrossRef]
55. Li Z, Zhang H. Reprogramming of glucose, fatty acid and amino acid metabolism for cancer progression. Cell Mol Life Sci. 2016;73(2):377–92. doi:10.1007/s00018-015-2070-4. [Google Scholar] [PubMed] [CrossRef]
56. Andryszkiewicz W, Gąsiorowska J, Kübler M, Kublińska K, Pałkiewicz A, Wiatkowski A, et al. Glucose metabolism and tumor microenvironment: mechanistic insights and therapeutic implications. Int J Mol Sci. 2025;26(5):1879. doi:10.3390/ijms26051879. [Google Scholar] [PubMed] [CrossRef]
57. Morrison AJ. Cancer cell metabolism connects epigenetic modifications to transcriptional regulation. FEBS J. 2022;289(5):1302–14. doi:10.1111/febs.16032. [Google Scholar] [PubMed] [CrossRef]
58. Thakur C, Chen F. Connections between metabolism and epigenetics in cancers. Semin Cancer Biol. 2019;57(5):52–8. doi:10.1016/j.semcancer.2019.06.006. [Google Scholar] [PubMed] [CrossRef]
59. Ge T, Gu X, Jia R, Ge S, Chai P, Zhuang A, et al. Crosstalk between metabolic reprogramming and epigenetics in cancer: updates on mechanisms and therapeutic opportunities. Cancer Commun. 2022;42(11):1049–82. doi:10.1002/cac2.12374. [Google Scholar] [PubMed] [CrossRef]
60. Sun L, Zhang H, Gao P. Metabolic reprogramming and epigenetic modifications on the path to cancer. Protein Cell. 2022;13(12):877–919. doi:10.1007/s13238-021-00846-7. [Google Scholar] [PubMed] [CrossRef]
61. Johnson C, Warmoes MO, Shen X, Locasale JW. Epigenetics and cancer metabolism. Cancer Lett. 2015;356(2 Pt A):309–14. doi:10.1016/j.canlet.2013.09.043. [Google Scholar] [PubMed] [CrossRef]
62. Kinnaird A, Zhao S, Wellen KE, Michelakis ED. Metabolic control of epigenetics in cancer. Nat Rev Cancer. 2016;16(11):694–707. doi:10.1038/nrc.2016.82. [Google Scholar] [PubMed] [CrossRef]
63. Singh JP, Zhang K, Wu J, Yang X. O-GlcNAc signaling in cancer metabolism and epigenetics. Cancer Lett. 2015;356(2 Pt A):244–50. doi:10.1016/j.canlet.2014.04.014. [Google Scholar] [PubMed] [CrossRef]
64. Liu S, Zhang X, Wang W, Li X, Sun X, Zhao Y, et al. Metabolic reprogramming and therapeutic resistance in primary and metastatic breast cancer. Mol Cancer. 2024;23(1):261. doi:10.1186/s12943-024-02165-x. [Google Scholar] [PubMed] [CrossRef]
65. Wu Y, Sun R, Ren S, Zengin G, Li MY. Neuronal reshaping of the tumor microenvironment in tumorigenesis and metastasis: bench to clinic. Med Adv. 2025;3(4):364–71. doi:10.1002/med4.70044. [Google Scholar] [CrossRef]
66. Che X, Wang H, Wang H, Jiang YZ. Intratumoral nerve phased development: a promising therapeutic target. Med Bull. 2025;1(2):154–70. doi:10.1002/mdb2.70009. [Google Scholar] [CrossRef]
67. Martínez-Reyes I, Chandel NS. Cancer metabolism: looking forward. Nat Rev Cancer. 2021;21(10):669–80. doi:10.1038/s41568-021-00378-6. [Google Scholar] [PubMed] [CrossRef]
68. Liu QQ, Dong ZK, Wang YF, Jin WL. Reprogramming neural-tumor crosstalk: emerging therapeutic dimensions and targeting strategies. Mil Med Res. 2025;12(1):73. doi:10.1186/s40779-025-00661-9. [Google Scholar] [PubMed] [CrossRef]
69. Niu Y, Feng J, Ma J, Xiao T, Yuan W. Targeting the tumor microbiota in cancer therapy basing on nanomaterials. Exploration. 2025;5(4):e20210185. doi:10.1002/EXP.20210185. [Google Scholar] [PubMed] [CrossRef]
70. Anastasiou D. Tumour microenvironment factors shaping the cancer metabolism landscape. Br J Cancer. 2017;116(3):277–86. doi:10.1038/bjc.2016.412. [Google Scholar] [PubMed] [CrossRef]
71. Gu A, Li J, Qiu S, Hao S, Yue ZY, Zhai S, et al. Pancreatic cancer environment: from patient-derived models to single-cell omics. Mol Omics. 2024;20(4):220–33. doi:10.1039/D3MO00250K. [Google Scholar] [PubMed] [CrossRef]
72. Xie J, Liu M, Deng X, Tang Y, Zheng S, Ou X, et al. Gut microbiota reshapes cancer immunotherapy efficacy: mechanisms and therapeutic strategies. iMeta. 2024;3(1):e156. doi:10.1002/imt2.156. [Google Scholar] [PubMed] [CrossRef]
73. Qin L, Liang T, Zhu X, Hu W, Li B, Wei M, et al. Senescent macrophages and the lung cancer microenvironment: a new perspective on tumor immune evasion. Aging Dis. 2025;16(6):3453–65. doi:10.14336/AD.2024.1404. [Google Scholar] [PubMed] [CrossRef]
74. Zhang H, Li S, Wang D, Liu S, Xiao T, Gu W, et al. Metabolic reprogramming and immune evasion: the interplay in the tumor microenvironment. Biomark Res. 2024;12(1):96. doi:10.1186/s40364-024-00646-1. [Google Scholar] [PubMed] [CrossRef]
75. Long F, Lin Z, Li L, Ma M, Lu Z, Jing L, et al. Comprehensive landscape and future perspectives of circular RNAs in colorectal cancer. Mol Cancer. 2021;20(1):26. doi:10.1186/s12943-021-01318-6. [Google Scholar] [PubMed] [CrossRef]
76. Cui X, Wang J, Guo Z, Li M, Li M, Liu S, et al. Emerging function and potential diagnostic value of circular RNAs in cancer. Mol Cancer. 2018;17(1):123. doi:10.1186/s12943-018-0877-y. [Google Scholar] [PubMed] [CrossRef]
77. Saranya I, Dharshini VS, Akshaya RL, Subhashini PS, Selvamurugan N. Regulatory and therapeutic implications of competing endogenous RNA network in breast cancer progression and metastasis: a review. Int J Biol Macromol. 2024;266(Pt 1):131075. doi:10.1016/j.ijbiomac.2024.131075. [Google Scholar] [PubMed] [CrossRef]
78. Salmena L, Poliseno L, Tay Y, Kats L, Pandolfi PP. A ceRNA hypothesis: the Rosetta Stone of a hidden RNA language? Cell. 2011;146(3):353–8. doi:10.1016/j.cell.2011.07.014. [Google Scholar] [PubMed] [CrossRef]
79. Mao M, Zhang J, Xiang Y, Gong M, Deng Y, Ye D. Role of exosomal competitive endogenous RNA (ceRNA) in diagnosis and treatment of malignant tumors. Bioengineered. 2022;13(5):12156–68. doi:10.1080/21655979.2022.2073130. [Google Scholar] [PubMed] [CrossRef]
80. Song J, Zheng J, Liu X, Dong W, Yang C, Wang D, et al. A novel protein encoded by ZCRB1-induced circHEATR5B suppresses aerobic glycolysis of GBM through phosphorylation of JMJD5. J Exp Clin Cancer Res. 2022;41(1):171. doi:10.1186/s13046-022-02374-6. [Google Scholar] [PubMed] [CrossRef]
81. Chen H, Li X, Yang L, Ye C, Chen Z, Wang Z. CircVMP1 promotes glycolysis and disease progression by upregulating HKDC1 in colorectal cancer. Environ Toxicol. 2024;39(3):1617–30. doi:10.1002/tox.24061. [Google Scholar] [PubMed] [CrossRef]
82. Dong W, Liu Y, Wang P, Ruan X, Liu L, Xue Y, et al. U3 snoRNA-mediated degradation of ZBTB7A regulates aerobic glycolysis in isocitrate dehydrogenase 1 wild-type glioblastoma cells. CNS Neurosci Ther. 2023;29(10):2811–25. doi:10.1111/cns.14218. [Google Scholar] [PubMed] [CrossRef]
83. Dai Q, Liu Y, Ding F, Guo R, Cheng G, Wang H. CircRNAs: a promising target for intervention regarding glycolysis in gastric cancer. Heliyon. 2024;10(14):e34658. doi:10.1016/j.heliyon.2024.e34658. [Google Scholar] [PubMed] [CrossRef]
84. Lei Q, Hu X, Han H, Wang J, Liu W, Zhou Y, et al. Integrative analysis of circRNA, miRNA, and mRNA profiles to reveal ceRNA regulation in chicken muscle development from the embryonic to post-hatching periods. BMC Genomics. 2022;23(1):342. doi:10.1186/s12864-022-08525-5. [Google Scholar] [PubMed] [CrossRef]
85. Lu L, Guo G, Guo J, Li H, Chen K, Chen Y, et al. A novel protein encoded by circUBE2G1 suppresses glycolysis in gastric cancer through binding to ENO1. Cell Death Discov. 2025;11(1):350. doi:10.1038/s41420-025-02644-0. [Google Scholar] [PubMed] [CrossRef]
86. Lin Q, Jiang H, Lin D. Circular RNA ITCH downregulates GLUT1 and suppresses glucose uptake in melanoma to inhibit cancer cell proliferation. J Dermatolog Treat. 2021;32(2):231–5. doi:10.1080/09546634.2019.1654069. [Google Scholar] [PubMed] [CrossRef]
87. Xu L, Shen Y, Zhang C, Shi T, Sheng X. Exploring the link between noncoding RNAs and glycolysis in colorectal cancer. J Cell Mol Med. 2025;29(4):e70443. doi:10.1111/jcmm.70443. [Google Scholar] [PubMed] [CrossRef]
88. Fan N, Fu H, Feng X, Chen Y, Wang J, Wu Y, et al. Long non-coding RNAs play an important regulatory role in tumorigenesis and tumor progression through aerobic glycolysis. Front Mol Biosci. 2022;9:941653. doi:10.3389/fmolb.2022.941653. [Google Scholar] [PubMed] [CrossRef]
89. Wang X, Guan X, Zhu X, Zhang L, Ma C, He S, et al. CircNAP1L4 regulates pulmonary artery smooth muscle cell proliferation via the NAP1L4-mediated super-enhancer-driven glycolysis gene Hexokinase II (HK II) in pulmonary hypertension. FASEB J. 2024;38(15):e23868. doi:10.1096/fj.202400585RRR. [Google Scholar] [PubMed] [CrossRef]
90. Wen L, Tan C, Ma S, Li X. lncRNAs: key regulators of signaling pathways in tumor glycolysis. Dis Markers. 2022;2022(13):2267963–9. doi:10.1155/2022/2267963. [Google Scholar] [PubMed] [CrossRef]
91. Li Q, Yao H, Wang Y, Wu Y, Thorne RF, Zhu Y, et al. circPRKAA1 activates a Ku80/Ku70/SREBP-1 axis driving de novo fatty acid synthesis in cancer cells. Cell Rep. 2022;41(8):111707. doi:10.1016/j.celrep.2022.111707. [Google Scholar] [PubMed] [CrossRef]
92. Kang SWS, Cunningham RP, Miller CB, Brown LA, Cultraro CM, Harned A, et al. A spatial map of hepatic mitochondria uncovers functional heterogeneity shaped by nutrient-sensing signaling. Nat Commun. 2024;15(1):1799. doi:10.1038/s41467-024-45751-9. [Google Scholar] [PubMed] [CrossRef]
93. Zou P, Wang L. Dietary pattern and hepatic lipid metabolism. Liver Res. 2023;7(4):275–84. doi:10.1016/j.livres.2023.11.006. [Google Scholar] [PubMed] [CrossRef]
94. Meng X, Li W, Yu T, Lu F, Wang C, Yuan H, et al. Hsa_circ_0086414/Transducer of ERBB2 (TOB2) axis-driven lipid elimination and tumor suppression in clear cell renal cell cancer via perilipin 3. Int J Biol Macromol. 2024;261(Pt 1):129636. doi:10.1016/j.ijbiomac.2024.129636. [Google Scholar] [PubMed] [CrossRef]
95. Yu X, Tong H, Chen J, Tang C, Wang S, Si Y, et al. CircRNA MBOAT2 promotes intrahepatic cholangiocarcinoma progression and lipid metabolism reprogramming by stabilizing PTBP1 to facilitate FASN mRNA cytoplasmic export. Cell Death Dis. 2023;14(1):20. doi:10.1038/s41419-022-05540-y. [Google Scholar] [PubMed] [CrossRef]
96. Xu L, Zhang Y, Lin Z, Deng X, Ren X, Huang M, et al. FASN-mediated fatty acid biosynthesis remodels immune environment in Clonorchis sinensis infection-related intrahepatic cholangiocarcinoma. J Hepatol. 2024;81(2):265–77. doi:10.1016/j.jhep.2024.03.016. [Google Scholar] [PubMed] [CrossRef]
97. Wang Y, Tian X, Wang Z, Liu D, Zhao X, Sun X, et al. A novel peptide encoded by circ-SLC9A6 promotes lipid dyshomeostasis through the regulation of H4K16ac-mediated CD36 transcription in NAFLD. Clin Transl Med. 2024;14(8):e1801. doi:10.1002/ctm2.1801. [Google Scholar] [PubMed] [CrossRef]
98. Chen S, Song P, Wang Y, Wang Z, Xue J, Jiang Y, et al. CircMAPK9 promotes adipogenesis through modulating Hsa-miR-1322/FTO axis in obesity. iScience. 2023;26(10):107756. doi:10.1016/j.isci.2023.107756. [Google Scholar] [PubMed] [CrossRef]
99. Liu X, Feng S, Zhang XD, Li J, Zhang K, Wu M, et al. Non-coding RNAs, metabolic stress and adaptive mechanisms in cancer. Cancer Lett. 2020;491:60–9. doi:10.1016/j.canlet.2020.06.024. [Google Scholar] [PubMed] [CrossRef]
100. Cai Y, Dong Z, Wang J. Circ_0000808 promotes the development of non-small cell lung cancer by regulating glutamine metabolism via the miR-1827/SLC1A5 axis. World J Surg Oncol. 2022;20(1):329. doi:10.1186/s12957-022-02777-x. [Google Scholar] [PubMed] [CrossRef]
101. Yang L, Wang L, Wu J, Wang Y. Circ_0000069 contributes to the growth, metastasis and glutamine metabolism in renal cell carcinoma (RCC) via regulating miR-125a-5p-dependent SLC1A5 expression. Transpl Immunol. 2023;77(1–6):101764. doi:10.1016/j.trim.2022.101764. [Google Scholar] [PubMed] [CrossRef]
102. Wu R, Tang S, Wang Q, Kong P, Liu F. Hsa_circ_0003602 contributes to the progression of colorectal cancer by mediating the miR-149-5p/SLC38A1 axis. Gut Liver. 2023;17(2):267–79. doi:10.5009/gnl210542. [Google Scholar] [PubMed] [CrossRef]
103. Eltayeb K, Alfieri R, Fumarola C, Bonelli M, Galetti M, Cavazzoni A, et al. Targeting metabolic adaptive responses induced by glucose starvation inhibits cell proliferation and enhances cell death in osimertinib-resistant non-small cell lung cancer (NSCLC) cell lines. Biochem Pharmacol. 2024;228:116161. doi:10.1016/j.bcp.2024.116161. [Google Scholar] [PubMed] [CrossRef]
104. Musicco C, Signorile A, Pesce V, Loguercio Polosa P, Cormio A. Mitochondria deregulations in cancer offer several potential targets of therapeutic interventions. Int J Mol Sci. 2023;24(13):10420. doi:10.3390/ijms241310420. [Google Scholar] [PubMed] [CrossRef]
105. Liu J, Bai X, Zhang M, Wu S, Xiao J, Zeng X, et al. Energy metabolism: a new target for gastric cancer treatment. Clin Transl Oncol. 2023;26(2):338–51. doi:10.1007/s12094-023-03278-3. [Google Scholar] [PubMed] [CrossRef]
106. Yan MQ, Zhu BH, Liu XH, Yang YM, Duan XY, Wang Y, et al. Mitoguardin 1 and 2 promote granulosa cell proliferation by activating AKT and regulating the Hippo-YAP1 signaling pathway. Cell Death Dis. 2023;14(11):779. doi:10.1038/s41419-023-06312-y. [Google Scholar] [PubMed] [CrossRef]
107. Huang C, Deng K, Wu M. Mitochondrial cristae in health and disease. Int J Biol Macromol. 2023;235(1):123755. doi:10.1016/j.ijbiomac.2023.123755. [Google Scholar] [PubMed] [CrossRef]
108. Cao H, Li C, Sun X, Yang J, Li X, Yang G, et al. Circular RNA circMYLK4 shifts energy metabolism from glycolysis to OXPHOS by binding to the calcium channel auxiliary subunit CACNA2D2. J Biol Chem. 2024;300(7):107426. doi:10.1016/j.jbc.2024.107426. [Google Scholar] [PubMed] [CrossRef]
109. De Rasmo D, Cormio A, Cormio G, Signorile A. Ovarian cancer: a landscape of mitochondria with emphasis on mitochondrial dynamics. Int J Mol Sci. 2023;24(2):1224. doi:10.3390/ijms24021224. [Google Scholar] [PubMed] [CrossRef]
110. Ma Z, Chen H, Xia Z, You J, Han C, Wang S, et al. Energy stress-induced circZFR enhances oxidative phosphorylation in lung adenocarcinoma via regulating alternative splicing. J Exp Clin Cancer Res. 2023;42(1):169. doi:10.1186/s13046-023-02723-z. [Google Scholar] [PubMed] [CrossRef]
111. Li J, Wang X, Shi L, Liu B, Sheng Z, Chang S, et al. A mammalian conserved circular RNA circLARP1B regulates hepatocellular carcinoma metastasis and lipid metabolism. Adv Sci. 2024;11(2):2305902. doi:10.1002/advs.202305902. [Google Scholar] [PubMed] [CrossRef]
112. Wu Z, Gu D, Wang R, Zuo X, Zhu H, Wang L, et al. CircRIC8B regulates the lipid metabolism of chronic lymphocytic leukemia through miR199b-5p/LPL axis. Exp Hematol Oncol. 2022;11(1):51. doi:10.1186/s40164-022-00302-0. [Google Scholar] [PubMed] [CrossRef]
113. Xing Z, Wang X, Liu J, Zhang M, Feng K, Wang X. Hsa_circ_0069094 accelerates cell malignancy and glycolysis through regulating the miR-591/HK2 axis in breast cancer. Cell Signal. 2021;79(862):109878. doi:10.1016/j.cellsig.2020.109878. [Google Scholar] [PubMed] [CrossRef]
114. Liu A, Xu J. Circ_03955 promotes pancreatic cancer tumorigenesis and Warburg effect by targeting the miR-3662/HIF-1α axis. Clin Transl Oncol. 2021;23(9):1905–14. doi:10.1007/s12094-021-02599-5. [Google Scholar] [PubMed] [CrossRef]
115. Li G, She FF, Liao CY, Wang ZW, Wang YT, Wu YD, et al. cNEK6 induces gemcitabine resistance by promoting glycolysis in pancreatic ductal adenocarcinoma via the SNRPA/PPA2c/mTORC1 axis. Cell Death Dis. 2024;15(10):742. doi:10.1038/s41419-024-07138-y. [Google Scholar] [PubMed] [CrossRef]
116. Huang M, He YR, Liang LC, Huang Q, Zhu ZQ. Circular RNA hsa_circ_0000745 may serve as a diagnostic marker for gastric cancer. World J Gastroenterol. 2017;23(34):6330–8. doi:10.3748/wjg.v23.i34.6330. [Google Scholar] [PubMed] [CrossRef]
117. Zheng D, Chen W, Peng J, Huang X, Zhang S, Zhuang Y. Hsa_circ_0007590/PTBP1 complex reprograms glucose metabolism by reducing the stability of m6A-modified PTEN mRNA in pancreatic ductal adenocarcinoma. Cancer Gene Ther. 2024;31(7):1090–102. doi:10.1038/s41417-024-00786-4. [Google Scholar] [PubMed] [CrossRef]
118. Shen S, Yao T, Xu Y, Zhang D, Fan S, Ma J. CircECE1 activates energy metabolism in osteosarcoma by stabilizing c-Myc. Mol Cancer. 2020;19(1):151. doi:10.1186/s12943-020-01269-4. [Google Scholar] [PubMed] [CrossRef]
119. Zhang Y, Zhao L, Yang S, Cen Y, Zhu T, Wang L, et al. CircCDKN2B-AS1 interacts with IMP3 to stabilize hexokinase 2 mRNA and facilitate cervical squamous cell carcinoma aerobic glycolysis progression. J Exp Clin Cancer Res. 2020;39(1):281. doi:10.1186/s13046-020-01793-7. [Google Scholar] [PubMed] [CrossRef]
120. Zhou J, Zhang S, Chen Z, He Z, Xu Y, Li Z. CircRNA-ENO1 promoted glycolysis and tumor progression in lung adenocarcinoma through upregulating its host gene ENO1. Cell Death Dis. 2019;10(12):885. doi:10.1038/s41419-019-2127-7. [Google Scholar] [PubMed] [CrossRef]
121. Cao J, Zhang X, Xu P, Wang H, Wang S, Zhang L, et al. Circular RNA circLMO7 acts as a microRNA-30a-3p sponge to promote gastric cancer progression via the WNT2/β-catenin pathway. J Exp Clin Cancer Res. 2021;40(1):6. doi:10.1186/s13046-020-01791-9. [Google Scholar] [PubMed] [CrossRef]
122. Lee SJ, Kim MJ, Kwon IC, Roberts TM. Delivery strategies and potential targets for siRNA in major cancer types. Adv Drug Deliv Rev. 2016;104(Suppl. 1):2–15. doi:10.1016/j.addr.2016.05.010. [Google Scholar] [PubMed] [CrossRef]
123. Dong Y, Siegwart DJ, Anderson DG. Strategies, design, and chemistry in siRNA delivery systems. Adv Drug Deliv Rev. 2019;144(60):133–47. doi:10.1016/j.addr.2019.05.004. [Google Scholar] [PubMed] [CrossRef]
124. Xu J, Ji L, Liang Y, Wan Z, Zheng W, Song X, et al. CircRNA-SORE mediates sorafenib resistance in hepatocellular carcinoma by stabilizing YBX1. Signal Transduct Target Ther. 2020;5(1):298. doi:10.1038/s41392-020-00375-5. [Google Scholar] [PubMed] [CrossRef]
125. Tamm I, Dörken B, Hartmann G. Antisense therapy in oncology: new hope for an old idea? Lancet. 2001;358(9280):489–97. doi:10.1016/S0140-6736(01)05629-X. [Google Scholar] [PubMed] [CrossRef]
126. Zhu G, Chang X, Kang Y, Zhao X, Tang X, Ma C, et al. CircRNA: a novel potential strategy to treat thyroid cancer. Int J Mol Med. 2021;48(5):201. doi:10.3892/ijmm.2021.5034. [Google Scholar] [PubMed] [CrossRef]
127. Yi J, Du J, Chen X, Nie R, Hu G, Wang L, et al. A circRNA–mRNA pairing mechanism regulates tumor growth and endocrine therapy resistance in ER-positive breast cancer. Proc Natl Acad Sci U S A. 2025;122(8):e2420383122. doi:10.1073/pnas.2420383122. [Google Scholar] [PubMed] [CrossRef]
128. Aquino-Jarquin G. CircRNA knockdown based on antisense strategies. Drug Discov Today. 2024;29(8):104066. doi:10.1016/j.drudis.2024.104066. [Google Scholar] [PubMed] [CrossRef]
129. Karimian A, Azizian K, Parsian H, Rafieian S, Shafiei-Irannejad V, Kheyrollah M, et al. CRISPR/Cas9 technology as a potent molecular tool for gene therapy. J Cell Physiol. 2019;234(8):12267–77. doi:10.1002/jcp.27972. [Google Scholar] [PubMed] [CrossRef]
130. Terns MP. CRISPR-based technologies: impact of RNA-targeting systems. Mol Cell. 2018;72(3):404–12. doi:10.1016/j.molcel.2018.09.018. [Google Scholar] [PubMed] [CrossRef]
131. Zhang Y, Nguyen TM, Zhang XO, Wang L, Phan T, Clohessy JG, et al. Optimized RNA-targeting CRISPR/Cas13d technology outperforms shRNA in identifying functional circRNAs. Genome Biol. 2021;22(1):41. doi:10.1186/s13059-021-02263-9. [Google Scholar] [PubMed] [CrossRef]
132. Yang J, Meng X, Pan J, Jiang N, Zhou C, Wu Z, et al. CRISPR/Cas9-mediated noncoding RNA editing in human cancers. RNA Biol. 2018;15(1):35–43. doi:10.1080/15476286.2017.1391443. [Google Scholar] [PubMed] [CrossRef]
133. Jia S, Yu L, Wang L, Peng L. The functional significance of circRNA/miRNA/mRNA interactions as a regulatory network in lung cancer biology. Int J Biochem Cell Biol. 2024;169(4):106548. doi:10.1016/j.biocel.2024.106548. [Google Scholar] [PubMed] [CrossRef]
134. Liang Z, Guo W, Fang S, Zhang Y, Lu L, Xu W, et al. CircRNAs: emerging bladder cancer biomarkers and targets. Front Oncol. 2021;10:606485. doi:10.3389/fonc.2020.606485. [Google Scholar] [PubMed] [CrossRef]
135. Yan L, Zheng M, Wang H. Circular RNA hsa_circ_0072309 inhibits proliferation and invasion of breast cancer cells via targeting miR-492. Cancer Manag Res. 2019;11:1033–41. doi:10.2147/CMAR.S186857. [Google Scholar] [PubMed] [CrossRef]
136. Shi L, Liu B, Shen DD, Yan PJ, Zhang Y, Tian Y, et al. A tumor-suppressive circular RNA mediates uncanonical integrin degradation by the proteasome in liver cancer. Sci Adv. 2021;7(13):eabe5043. doi:10.1126/sciadv.abe5043. [Google Scholar] [PubMed] [CrossRef]
137. Li Z, Ruan Y, Zhang H, Shen Y, Li T, Xiao B. Tumor-suppressive circular RNAs: mechanisms underlying their suppression of tumor occurrence and use as therapeutic targets. Cancer Sci. 2019;110(12):3630–8. doi:10.1111/cas.14211. [Google Scholar] [PubMed] [CrossRef]
138. Li C, Yang C, Yang Q. The role of miRNAs in the extracellular vesicle-mediated interplay between breast tumor cells and cancer-associated fibroblasts. J Cancer Metastasis Treat. 2024;10:27. doi:10.20517/2394-4722.2024.76. [Google Scholar] [CrossRef]
139. Shang Q, Yang Z, Jia R, Ge S. The novel roles of circRNAs in human cancer. Mol Cancer. 2019;18(1):6. doi:10.1186/s12943-018-0934-6. [Google Scholar] [PubMed] [CrossRef]
140. Zhang Y, Liang W, Zhang P, Chen J, Qian H, Zhang X, et al. Circular RNAs: emerging cancer biomarkers and targets. J Exp Clin Cancer Res. 2017;36(1):152. doi:10.1186/s13046-017-0624-z. [Google Scholar] [PubMed] [CrossRef]
141. Wang D, Tang L, Chen M, Gong Z, Fan C, Qu H, et al. Nanocarriers targeting circular RNA ADARB1 boost radiosensitivity of nasopharyngeal carcinoma through synergically promoting ferroptosis. ACS Nano. 2024;18(45):31055–75. doi:10.1021/acsnano.4c07676. [Google Scholar] [PubMed] [CrossRef]
142. Niu D, Wu Y, Lian J. Circular RNA vaccine in disease prevention and treatment. Signal Transduct Target Ther. 2023;8(1):341. doi:10.1038/s41392-023-01561-x. [Google Scholar] [PubMed] [CrossRef]
143. Zhang MX, Wang JL, Mo CQ, Mao XP, Feng ZH, Li JY, et al. CircME1 promotes aerobic glycolysis and sunitinib resistance of clear cell renal cell carcinoma through cis-regulation of ME1. Oncogene. 2022;41(33):3979–90. doi:10.1038/s41388-022-02386-8. [Google Scholar] [PubMed] [CrossRef]
144. Nakashima C, Yamamoto K, Fujiwara-Tani R, Luo Y, Matsushima S, Fujii K, et al. Expression of cytosolic malic enzyme (ME1) is associated with disease progression in human oral squamous cell carcinoma. Cancer Sci. 2018;109(6):2036–45. doi:10.1111/cas.13594. [Google Scholar] [PubMed] [CrossRef]
145. Adams VR, Leggas M. Sunitinib malate for the treatment of metastatic renal cell carcinoma and gastrointestinal stromal tumors. Clin Ther. 2007;29(7):1338–53. doi:10.1016/j.clinthera.2007.07.022. [Google Scholar] [PubMed] [CrossRef]
146. Li J, Gu A, Tang N, Zengin G, Li M, Liu Y. Patient-derived xenograft models in pan-cancer: from bench to clinic. Interdiscip Med. 2025;3(5):e20250016. doi:10.1002/INMD.20250016. [Google Scholar] [CrossRef]
147. Zhou T, Yan H, Deng Y, Zhu Y, Xia X, Wu W, et al. The role of long non-coding RNA Maternally Expressed Gene 3 in cancer-associated fibroblasts at single cell pan-cancer level. Interdiscip Med. 2024;2(4):e20240018. doi:10.1002/INMD.20240018. [Google Scholar] [CrossRef]
148. He R, Hu C, Yuan Y, Li T, Tian Q, Huang T, et al. Glycolysis reprogramming in CAFs promotes oxaliplatin resistance in pancreatic cancer through circABCC4 mediated PKM2 nuclear translocation. Cell Death Dis. 2025;16(1):126. doi:10.1038/s41419-025-07431-4. [Google Scholar] [PubMed] [CrossRef]
149. Guo X, Gao C, Yang DH, Li S. Exosomal circular RNAs: a chief culprit in cancer chemotherapy resistance. Drug Resist Updat. 2023;67(1):100937. doi:10.1016/j.drup.2023.100937. [Google Scholar] [PubMed] [CrossRef]
150. Li J, Gu A, Tang N, Li MY, Liu Y. Efficacy and mechanisms of Ci-Gu-Tang against pancreatic cancer: a study using patient-derived models. J Future Foods. 2025;18:493. In press. doi:10.1016/j.jfutfo.2024.10.009. [Google Scholar] [CrossRef]
151. Li Q, Zhang Y, Jin P, Chen Y, Zhang C, Geng X, et al. New insights into the potential of exosomal circular RNAs in mediating cancer chemotherapy resistance and their clinical applications. Biomed Pharmacother. 2024;177:117027. doi:10.1016/j.biopha.2024.117027. [Google Scholar] [PubMed] [CrossRef]
152. Liu Y, Shen Z, Wei X, Gu L, Zheng M, Zhang Y, et al. CircSLC39A8 attenuates paclitaxel resistance in ovarian cancer by regulating the miR-185-5p/BMF axis. Transl Oncol. 2023;36(1):101746. doi:10.1016/j.tranon.2023.101746. [Google Scholar] [PubMed] [CrossRef]
153. Zhang Y, Ge P, Zhou D, Xing R, Bai L. Circular RNA FOXO3 accelerates glycolysis and improves cisplatin sensitivity in lung cancer cells via the miR-543/Foxo3 axis. Oncol Lett. 2021;22(5):839. doi:10.3892/ol.2021.13100. [Google Scholar] [PubMed] [CrossRef]
154. Gu J, Dai Z, Shen T, Chen X, Yang Z, Sun S, et al. CDK4/6 and BET inhibitors synergistically suppress pancreatic tumor growth and epithelial-to-mesenchymal transition by regulating the GSK3β-mediated Wnt/β-catenin pathway. Cancer Drug Resist. 2025;8:52. doi:10.20517/cdr.2025.38. [Google Scholar] [PubMed] [CrossRef]
155. Lee J, Gollahon L. Nek2-Targeted ASO or siRNA pretreatment enhances anticancer drug sensitivity in triple-negative breast cancer cells. Int J Oncol. 2013;42(3):839–47. doi:10.3892/ijo.2013.1788. [Google Scholar] [PubMed] [CrossRef]
156. Guo C, Zhang J, Cai X, Dou R, Tang J, Huang Z, et al. Emerging nanomedicine strategies for hepatocellular carcinoma therapy. iMetaOmics. 2024;1(1):e12. doi:10.1002/imo2.12. [Google Scholar] [PubMed] [CrossRef]
157. Sahu RC, Arora S, Kumar D, Agrawal AK. Machine learning for predictive modeling in nanomedicine-based cancer drug delivery. Med Res. 2025. doi:10.1002/mdr2.70043. [Google Scholar] [CrossRef]
158. Dhar R, Subramaniyan V. Exosomes: a next-generation cancer theranostics. Nano TransMed. 2025;75(8):100096. In press. doi:10.1016/j.ntm.2025.100096. [Google Scholar] [CrossRef]
159. Wang S, Dong Y, Gong A, Kong H, Gao J, Hao X, et al. Exosomal circRNAs as novel cancer biomarkers: challenges and opportunities. Int J Biol Sci. 2021;17(2):562–73. doi:10.7150/ijbs.48782. [Google Scholar] [PubMed] [CrossRef]
160. Zhang F, Jiang J, Qian H, Yan Y, Xu W. Exosomal circRNA: emerging Insights into Cancer Progression and Clinical Application Potential. J Hematol Oncol. 2023;16(1):67. doi:10.1186/s13045-023-01452-2. [Google Scholar] [PubMed] [CrossRef]
161. Song S, Liu Z, Wang Y, Gong B. Human organoids and their application in tumor models, disease modeling, and tissue engineering. Med Bull. 2025;1(1):17–36. doi:10.1002/mdb2.70001. [Google Scholar] [CrossRef]
162. Wang SW. Current applications and future perspective of CRISPR/Cas9 gene editing in cancer. Mol Cancer. 2022;21(1):57. doi:10.1186/s12943-022-01518-8. [Google Scholar] [PubMed] [CrossRef]
163. Ning H, Jiang Y, Li B, Ren J, Ran A, Li W, et al. Targeted delivery of circPDHK1 siRNA via aptamer functionalized lipid nanoparticles inhibits ccRCC growth and migration. Int J Pharm. 2025;677(5):125666. doi:10.1016/j.ijpharm.2025.125666. [Google Scholar] [PubMed] [CrossRef]
164. Chen B, Dragomir MP, Yang C, Li Q, Horst D, Calin GA. Targeting non-coding RNAs to overcome cancer therapy resistance. Signal Transduct Target Ther. 2022;7(1):121. doi:10.1038/s41392-022-00975-3. [Google Scholar] [PubMed] [CrossRef]
165. Luo Y, He X, Du Q, Xu L, Xu J, Wang J, et al. Metal-based smart nanosystems in cancer immunotherapy. exploration. 2024;4(6):20230134. doi:10.1002/EXP.20230134. [Google Scholar] [PubMed] [CrossRef]
166. Yu HJ, Liu JH, Liu W, Niu R, Zhang B, Xiong Y, et al. Metal-based nanomedicines for cancer theranostics. Mil Med Res. 2025;12(1):41. doi:10.1186/s40779-025-00627-x. [Google Scholar] [PubMed] [CrossRef]
167. Sun L, Liu Y, Yang N, Ye X, Liu Z, Wu J, et al. Gold nanoparticles inhibit tumor growth via targeting the Warburg effect in a c-Myc-dependent way. Acta Biomater. 2023;158:583–98. doi:10.1016/j.actbio.2022.12.054. [Google Scholar] [PubMed] [CrossRef]
168. Racca L, Liuzzi E, Comparato S, Giordano G, Pignochino Y. Nanoparticles-delivered circular RNA strategy as a novel antitumor approach. Int J Mol Sci. 2024;25(16):8934. doi:10.3390/ijms25168934. [Google Scholar] [PubMed] [CrossRef]
169. He S, Gou X, Zhang S, Zhang X, Huang H, Wang W, et al. Nanodelivery systems as a novel strategy to overcome treatment failure of cancer. Small Methods. 2024;8(1):2301127. doi:10.1002/smtd.202301127. [Google Scholar] [PubMed] [CrossRef]
170. Lv K, Yu Z, Wang J, Li N, Wang A, Xue T, et al. Discovery of ketal-ester ionizable lipid nanoparticle with reduced hepatotoxicity, enhanced spleen tropism for mRNA vaccine delivery. Adv Sci. 2024;11(45):2404684. doi:10.1002/advs.202404684. [Google Scholar] [PubMed] [CrossRef]
171. Hosseini-Kharat M, Bremmell KE, Prestidge CA. Why do lipid nanoparticles target the liver? Understanding of biodistribution and liver-specific tropism. Mol Ther Methods Clin Dev. 2025;33(1):101436. doi:10.1016/j.omtm.2025.101436. [Google Scholar] [PubMed] [CrossRef]
172. Kim SB. Function and therapeutic development of exosomes for cancer therapy. Arch Pharm Res. 2022;45(5):295–308. doi:10.1007/s12272-022-01387-1. [Google Scholar] [PubMed] [CrossRef]
173. Gong Z, Hu W, Zhou C, Guo J, Yang L, Wang B. Recent advances and perspectives on the development of circular RNA cancer vaccines. NPJ Vaccin. 2025;10(1):41. doi:10.1038/s41541-025-01097-x. [Google Scholar] [PubMed] [CrossRef]
174. Jeong M, Lee Y, Park J, Jung H, Lee H. Lipid nanoparticles (LNPs) for in vivo RNA delivery and their breakthrough technology for future applications. Adv Drug Deliv Rev. 2023;200(2018):114990. doi:10.1016/j.addr.2023.114990. [Google Scholar] [PubMed] [CrossRef]
175. Lim ZZJ, Li JEJ, Ng CT, Yung LYL, Bay BH. Gold nanoparticles in cancer therapy. Acta Pharmacol Sin. 2011;32(8):983–90. doi:10.1038/aps.2011.82. [Google Scholar] [PubMed] [CrossRef]
176. Wang J, Ding Y, Chong K, Cui M, Cao Z, Tang C, et al. Recent advances in lipid nanoparticles and their safety concerns for mRNA delivery. Vaccines. 2024;12(10):1148. doi:10.3390/vaccines12101148. [Google Scholar] [PubMed] [CrossRef]
177. Hald Albertsen C, Kulkarni JA, Witzigmann D, Lind M, Petersson K, Simonsen JB. The role of lipid components in lipid nanoparticles for vaccines and gene therapy. Adv Drug Deliv Rev. 2022;188:114416. doi:10.1016/j.addr.2022.114416. [Google Scholar] [PubMed] [CrossRef]
178. Eygeris Y, Gupta M, Kim J, Sahay G. Chemistry of lipid nanoparticles for RNA delivery. Acc Chem Res. 2022;55(1):2–12. doi:10.1021/acs.accounts.1c00544. [Google Scholar] [PubMed] [CrossRef]
179. Zhang Z, Yao S, Hu Y, Zhao X, Lee RJ. Application of lipid-based nanoparticles in cancer immunotherapy. Front Immunol. 2022;13:967505. doi:10.3389/fimmu.2022.967505. [Google Scholar] [PubMed] [CrossRef]
180. Li MY, Zhang H, Li J, Chen YS, Ren S, Sun WL, et al. Reshaping the future of cancer therapy: taming toxicity and side effects. Biomed Eng Commun. 2026;5(2):11. doi:10.53388/BMEC2026011. [Google Scholar] [CrossRef]
181. Chen Z, Kang F, Xie C, Liao C, Li G, Wu Y, et al. A novel Trojan horse nanotherapy strategy targeting the cPKM-STMN1/TGFB1 axis for effective treatment of intrahepatic cholangiocarcinoma. Adv Sci. 2023;10(32):2303814. doi:10.1002/advs.202303814. [Google Scholar] [PubMed] [CrossRef]
182. You S, Luo Z, Cheng N, Wu M, Lai Y, Wang F, et al. Magnetically responsive nanoplatform targeting circRNA Circ_0058051 inhibits hepatocellular carcinoma progression. Drug Deliv Transl Res. 2023;13(3):782–94. doi:10.1007/s13346-022-01237-z. [Google Scholar] [PubMed] [CrossRef]
183. Yu C, Zhang Y, Wang N, Wei W, Cao K, Zhang Q, et al. Treatment of bladder cancer by geoinspired synthetic chrysotile nanocarrier-delivered circPRMT5 siRNA. Biomater Res. 2022;26(1):6. doi:10.1186/s40824-022-00251-z. [Google Scholar] [PubMed] [CrossRef]
184. Du WW, Yang W, Li X, Awan FM, Yang Z, Fang L, et al. A circular RNA circ-DNMT1 enhances breast cancer progression by activating autophagy. Oncogene. 2018;37(44):5829–42. doi:10.1038/s41388-018-0369-y. [Google Scholar] [PubMed] [CrossRef]
185. Li Y, Wang Z, Gao P, Cao D, Dong R, Zhu M, et al. CircRHBDD1 promotes immune escape via IGF2BP2/PD-L1 signaling and acts as a nanotherapeutic target in gastric cancer. J Transl Med. 2024;22(1):704. doi:10.1186/s12967-024-05498-9. [Google Scholar] [PubMed] [CrossRef]
186. Harper K, Mcdonnell E, Whitehouse A. CircRNAs: from anonymity to novel regulators of gene expression in cancer. Int J Oncol. 2019;55(6):1183–93. doi:10.3892/ijo.2019.4904. [Google Scholar] [PubMed] [CrossRef]
187. Luo X, Peng Y, Fan X, Xie X, Jin Z, Zhang X. The crosstalk and clinical implications of CircRNAs and glucose metabolism in gastrointestinal cancers. Cancers. 2023;15(8):2229. doi:10.3390/cancers15082229. [Google Scholar] [PubMed] [CrossRef]
188. Gu A, Li J, Li M, Liu Y. Patient-derived xenograft model in cancer: establishment and applications. Med Comm. 2025;6(2):e70059. doi:10.1002/mco2.70059. [Google Scholar] [PubMed] [CrossRef]
189. Lin J, Wang X, Zhai S, Shi M, Peng C, Deng X, et al. Hypoxia-induced exosomal circPDK1 promotes pancreatic cancer glycolysis via c-Myc activation by modulating miR-628-3p/BPTF axis and degrading BIN1. J Hematol Oncol. 2022;15(1):128. doi:10.1186/s13045-022-01348-7. [Google Scholar] [PubMed] [CrossRef]
190. Shan C, Zhang Y, Hao X, Gao J, Chen X, Wang K. Biogenesis, functions and clinical significance of circRNAs in gastric cancer. Mol Cancer. 2019;18(1):136. doi:10.1186/s12943-019-1069-0. [Google Scholar] [PubMed] [CrossRef]
191. Ma Y, Zheng L, Gao Y, Zhang W, Zhang Q, Xu Y. A comprehensive overview of circRNAs: emerging biomarkers and potential therapeutics in gynecological cancers. Front Cell Dev Biol. 2021;9:709512. doi:10.3389/fcell.2021.709512. [Google Scholar] [PubMed] [CrossRef]
192. Zhang Y, Luo J, Yang W, Ye WC. CircRNAs in colorectal cancer: potential biomarkers and therapeutic targets. Cell Death Dis. 2023;14(6):353. doi:10.1038/s41419-023-05881-2. [Google Scholar] [PubMed] [CrossRef]
193. Li J, Sun D, Pu W, Wang J, Peng Y. Circular RNAs in cancer: biogenesis, function, and clinical significance. Trends Cancer. 2020;6(4):319–36. doi:10.1016/j.trecan.2020.01.012. [Google Scholar] [PubMed] [CrossRef]
194. Zhu X, Wang X, Wei S, Chen Y, Chen Y, Fan X, et al. Hsa_circ_0013958: a circular RNA and potential novel biomarker for lung adenocarcinoma. FEBS J. 2017;284(14):2170–82. doi:10.1111/febs.14132. [Google Scholar] [PubMed] [CrossRef]
195. Siedlecki E, Remiszewski P, Stec R. The role of circHIPK3 in tumorigenesis and its potential as a biomarker in lung cancer. Cells. 2024;13(17):1483. doi:10.3390/cells13171483. [Google Scholar] [PubMed] [CrossRef]
196. Chen J, Wang H, Xu J, Chen E, Meng Q, Wang J, et al. CircZFR promotes colorectal cancer progression via stabilizing BCLAF1 and regulating the miR-3127-5p/RTKN2 axis. Sci China Life Sci. 2024;67(9):1881–98. doi:10.1007/s11427-023-2514-y. [Google Scholar] [PubMed] [CrossRef]
197. Li K, Liao X, Li S, Wei H, Liang Z, Ge L, et al. Identification and diagnostic potential of Hsa_circ_101303 in colorectal cancer: unraveling a regulatory network. BMC Cancer. 2024;24(1):671. doi:10.1186/s12885-024-12458-5. [Google Scholar] [PubMed] [CrossRef]
198. Wang H, Xiao Y, Wu L, Ma D. Comprehensive circular RNA profiling reveals the regulatory role of the circRNA-000911/miR-449a pathway in breast carcinogenesis. Int J Oncol. 2018;52(3):743–54. doi:10.3892/ijo.2018.4265. [Google Scholar] [PubMed] [CrossRef]
199. Zhu J, Ye J, Zhang L, Xia L, Hu H, Jiang H, et al. Differential expression of circular RNAs in glioblastoma multiforme and its correlation with prognosis. Transl Oncol. 2017;10(2):271–9. doi:10.1016/j.tranon.2016.12.006. [Google Scholar] [PubMed] [CrossRef]
200. Wang S, Zhang Y, Cai Q, Ma M, Jin LY, Weng M, et al. Circular RNA FOXP1 promotes tumor progression and Warburg effect in gallbladder cancer by regulating PKLR expression. Mol Cancer. 2019;18(1):145. doi:10.1186/s12943-019-1078-z. [Google Scholar] [PubMed] [CrossRef]
201. Yang C, Yuan W, Yang X, Li P, Wang J, Han J, et al. Circular RNA circ-ITCH inhibits bladder cancer progression by sponging miR-17/miR-224 and regulating p21, PTEN expression. Mol Cancer. 2018;17(1):19. doi:10.1186/s12943-018-0771-7. [Google Scholar] [PubMed] [CrossRef]
202. Zhou H, Cui X, Zhu L, Xu Z, Wang Z, Shao J. circPVT1 Inhibits the proliferation and aids in prediction of the prognosis of bladder cancer. Pharmgenomics Pers Med. 2024;17:1–11. doi:10.2147/PGPM.S427147. [Google Scholar] [PubMed] [CrossRef]
203. Yu J, Yang M, Zhou B, Luo J, Zhang Z, Zhang W, et al. CircRNA-104718 acts as competing endogenous RNA and promotes hepatocellular carcinoma progression through microRNA-218-5p/TXNDC5 signaling pathway. Clin Sci. 2019;133(13):1487–503. doi:10.1042/CS20190394. [Google Scholar] [PubMed] [CrossRef]
204. Zhang H, Wang G, Ding C, Liu P, Wang R, Ding W, et al. Increased circular RNA UBAP2 acts as a sponge of miR-143 to promote osteosarcoma progression. Oncotarget. 2017;8(37):61687–97. doi:10.18632/oncotarget.18671. [Google Scholar] [PubMed] [CrossRef]
205. Zhang X, Xu Y, Yamaguchi K, Hu J, Zhang L, Wang J, et al. Circular RNA circVAPA knockdown suppresses colorectal cancer cell growth process by regulating miR-125a/CREB5 axis. Cancer Cell Int. 2020;20(1):103. doi:10.1186/s12935-020-01178-y. [Google Scholar] [PubMed] [CrossRef]
206. Weng W, Wei Q, Toden S, Yoshida K, Nagasaka T, Fujiwara T, et al. Circular RNA ciRS-7—a promising prognostic biomarker and a potential therapeutic target in colorectal cancer. Clin Cancer Res. 2017;23(14):3918–28. doi:10.1158/1078-0432.CCR-16-2541. [Google Scholar] [PubMed] [CrossRef]
207. Zhang H, Ma X, Wang L, Li X, Feng D, Liu M, et al. Circular RNA hsa_circ_0007367 promotes the progression of pancreatic ductal adenocarcinoma by sponging miR-6820-3p and upregulating YAP1 expression. Cell Death Dis. 2022;13(8):736. doi:10.1038/s41419-022-05188-8. [Google Scholar] [PubMed] [CrossRef]
208. Weng Q, Chen M, Li M, Zheng YF, Shao G, Fan W, et al. Global microarray profiling identified hsa_circ_0064428 as a potential immune-associated prognosis biomarker for hepatocellular carcinoma. J Med Genet. 2019;56(1):32–8. doi:10.1136/jmedgenet-2018-105440. [Google Scholar] [PubMed] [CrossRef]
209. Wang L, Xiao S, Zheng Y, Gao Z. CircRNA circSLIT2 is a novel diagnostic and prognostic biomarker for gastric cancer. Wien Klin Wochenschr. 2023;135(17–18):472–7. doi:10.1007/s00508-023-02155-x. [Google Scholar] [PubMed] [CrossRef]
210. Li MY, Gu A, Li J, Tang N, Matin M, Yang Y, et al. Exploring food and medicine homology: potential implications for cancer treatment innovations. Acta Mater Med. 2025;4(2):200–6. doi:10.15212/AMM-2025-0003. [Google Scholar] [CrossRef]
211. Zhang C, Zhang B. RNA therapeutics: updates and future potential. Sci China Life Sci. 2023;66(1):12–30. doi:10.1007/s11427-022-2171-2. [Google Scholar] [PubMed] [CrossRef]
212. Li J, Luo P, Li Z, Wang Q, Huang X, Wang K, et al. Extrachromosomal circular DNA in cancer: mechanisms and clinical applications. Cell Prolif. 2025;58(6):e70040. doi:10.1111/cpr.70040. [Google Scholar] [PubMed] [CrossRef]
213. Liu X, Zhang Y, Zhou S, Dain L, Mei L, Zhu G. Circular RNA: an emerging frontier in RNA therapeutic targets, RNA therapeutics, and mRNA vaccines. J Control Release. 2022;348(3):84–94. doi:10.1016/j.jconrel.2022.05.043. [Google Scholar] [PubMed] [CrossRef]
214. Liang Y, Yang Q. The role of circular RNAs in glucose metabolic reprogramming: impact on cancer progression and therapeutic implications in breast cancer. J Cancer Metastasis Treat. 2025;11:3. doi:10.20517/2394-4722.2024.93. [Google Scholar] [CrossRef]
215. Dhuri K, Bechtold C, Quijano E, Pham H, Gupta A, Vikram A, et al. Antisense oligonucleotides: an emerging area in drug discovery and development. J Clin Med. 2004 2020;9(6):2004. doi:10.3390/jcm9062004. [Google Scholar] [PubMed] [CrossRef]
216. Li S, Li X, Xue W, Zhang L, Yang LZ, Cao SM, et al. Screening for functional circular RNAs using the CRISPR-Cas13 system. Nat Methods. 2021;18(1):51–9. doi:10.1038/s41592-020-01011-4. [Google Scholar] [PubMed] [CrossRef]
217. Hu X, Wu D, He X, Zhao H, He Z, Lin J, et al. circGSK3β promotes metastasis in esophageal squamous cell carcinoma by augmenting β-catenin signaling. Mol Cancer. 2019;18(1):160. doi:10.1186/s12943-019-1095-y. [Google Scholar] [PubMed] [CrossRef]
218. Zhou WY, Cai ZR, Liu J, Wang DS, Ju HQ, Xu RH. Circular RNA: metabolism, functions and interactions with proteins. Mol Cancer. 2020;19(1):172. doi:10.1186/s12943-020-01286-3. [Google Scholar] [PubMed] [CrossRef]
219. Wang Y, Zhang Y, Wang P, Fu X, Lin W. Circular RNAs in renal cell carcinoma: implications for tumorigenesis, diagnosis, and therapy. Mol Cancer. 2020;19(1):149. doi:10.1186/s12943-020-01266-7. [Google Scholar] [PubMed] [CrossRef]
220. Zhang J, Luo Z, Zheng Y, Duan M, Qiu Z, Huang C. CircRNA as an achilles heel of cancer: characterization, biomarker and therapeutic modalities. J Transl Med. 2024;22(1):752. doi:10.1186/s12967-024-05562-4. [Google Scholar] [PubMed] [CrossRef]
221. Tang N, Li J, Gu A, Li M, Liu Y. Single-cell multi-omics in biliary tract cancers: decoding heterogeneity, microenvironment, and treatment strategies. Mol Biomed. 2025;6(1):82. doi:10.1186/s43556-025-00330-2. [Google Scholar] [PubMed] [CrossRef]
222. Li H, Peng K, Yang K, Ma W, Qi S, Yu X, et al. Circular RNA cancer vaccines drive immunity in hard-to-treat malignancies. Theranostics. 2022;12(14):6422–36. doi:10.7150/thno.77350. [Google Scholar] [PubMed] [CrossRef]
223. Yu J, Kong X, Feng Y. Tumor microenvironment-driven resistance to immunotherapy in non-small cell lung cancer: strategies for cold-to-hot tumor transformation. Cancer Drug Resist. 2025;8:21. doi:10.20517/cdr.2025.14. [Google Scholar] [PubMed] [CrossRef]
224. Yang M, Zhou J, Lu L, Deng D, Huang J, Tang Z, et al. Tumor cell membrane-based vaccines: a potential boost for cancer immunotherapy. Exploration. 2024;4(6):20230171. doi:10.1002/EXP.20230171. [Google Scholar] [PubMed] [CrossRef]
225. Amaya L, Grigoryan L, Li Z, Lee A, Wender PA, Pulendran B, et al. Circular RNA vaccine induces potent T cell responses. Proc Natl Acad Sci U S A. 2023;120(20):e2302191120. doi:10.1073/pnas.2302191120. [Google Scholar] [PubMed] [CrossRef]
226. Wu J, Xue X, Qu H. Tumor microenvironment-responsive nanoplatforms for enhanced cancer immunotherapy: advances and synergistic strategies. Nano TransMed. 2025;4:100092. doi:10.1016/j.ntm.2025.100092. [Google Scholar] [CrossRef]
227. Zhao X, Zhong Y, Wang X, Shen J, An W. Advances in circular RNA and its applications. Int J Med Sci. 2022;19(6):975–85. doi:10.7150/ijms.71840. [Google Scholar] [PubMed] [CrossRef]
228. Yu H, Wen Y, Yu W, Lu L, Yang Y, Liu C, et al. Optimized circular RNA vaccines for superior cancer immunotherapy. Theranostics. 2025;15(4):1420–38. doi:10.7150/thno.104698. [Google Scholar] [PubMed] [CrossRef]
229. Sharma R, Malviya R. Modifying the electrical, optical, and magnetic properties of cancer cells: a comprehensive approach for cancer management. Med Adv. 2024;2(1):3–19. doi:10.1002/med4.51. [Google Scholar] [CrossRef]
230. Luo J, Yang Y, Zhang G, Fang D, Liu K, Mei Y, et al. Energy stress-induced circDDX21 promotes glycolysis and facilitates hepatocellular carcinogenesis. Cell Death Dis. 2024;15(5):354. doi:10.1038/s41419-024-06743-1. [Google Scholar] [PubMed] [CrossRef]
231. Yepmo M, Potier JB, Pinget M, Grabarz A, Bouzakri K, Dumond Bourie A. Discussing the role of circular RNA in the pathogenesis of non-alcoholic fatty liver disease and its complications. Front Endocrinol. 2022;13:1035159. doi:10.3389/fendo.2022.1035159. [Google Scholar] [PubMed] [CrossRef]
232. Ye M, Liu Y, Li H. Pan-cancer analysis and mechanistic insights of tuft cell-like tumors. Genes Dis. 2025;18(1):101939. In press. doi:10.1016/j.gendis.2025.101939. [Google Scholar] [CrossRef]
233. Yang C, Yang L, Feng Y, Song X, Bai S, Zhang S, et al. Modeling methods of different tumor organoids and their application in tumor drug resistance research. Cancer Drug Resist. 2025;8:32. doi:10.20517/cdr.2025.34. [Google Scholar] [PubMed] [CrossRef]
234. Yi Z, Qu L, Tang H, Liu Z, Liu Y, Tian F, et al. Engineered circular ADAR-recruiting RNAs increase the efficiency and fidelity of RNA editing in vitro and in vivo. Nat Biotechnol. 2022;40(6):946–55. doi:10.1038/s41587-021-01180-3. [Google Scholar] [CrossRef]
235. Bartoszewski R, Sikorski AF. Editorial focus: understanding off-target effects as the key to successful RNAi therapy. Cell Mol Biol Lett. 2019;24(1):69. doi:10.1186/s11658-019-0196-3. [Google Scholar] [PubMed] [CrossRef]
236. Li T, Li S, Kang Y, Zhou J, Yi M. Harnessing the evolving CRISPR/Cas9 for precision Oncology. J Transl Med. 2024;22(1):749. doi:10.1186/s12967-024-05570-4. [Google Scholar] [PubMed] [CrossRef]
237. Alqahtani S, Alqahtani T, Venkatesan K, Sivadasan D, Ahmed R, Elfadil H, et al. Unveiling pharmacogenomics insights into circular RNAs: toward precision medicine in cancer therapy. Biomolecules. 2025;15(4):535. doi:10.3390/biom15040535. [Google Scholar] [PubMed] [CrossRef]
238. Mei S, Ma X, Zhou L, Wuyun Q, Cai Z, Yan J, et al. CircSMAD3 represses VSMC phenotype switching and neointima formation via promoting hnRNPA1 ubiquitination degradation. Cell Prolif. 2025;58(1):e13742. doi:10.1111/cpr.13742. [Google Scholar] [PubMed] [CrossRef]
239. Lu X, Ying Y, Chen J, Chen Z, Wu Y, Prasanna P, et al. From digitized whole-slide histology images to biomarker discovery: a protocol for handcrafted feature analysis in brain cancer pathology. Brain-X. 2025;3(2):e70030. doi:10.1002/brx2.70030. [Google Scholar] [CrossRef]
240. Li MY, Zhang Q, Zengin G, Guo Q, Sun W. Spatial omics: deciphering heterogeneity in the tumor immune microenvironment and resistance to immunotherapy. Curr Proteomics. 2025;22(5):100055. doi:10.1016/j.curpro.2025.100055. [Google Scholar] [CrossRef]
241. Zhang Q, Liu J, Li J, Li MY, Jin CH. Precision application of food and medicine with a multi-omics network and biomimetic delivery system. Food Med Homol. 2026;3(1):9420129. doi:10.26599/FMH.2026.9420129. [Google Scholar] [CrossRef]
242. Malhotra D, Gabrani R. Metabolic shifts in glioblastoma: unraveling altered pathways and exploring novel therapeutic avenues. Mol Biol Rep. 2025;52(1):146. doi:10.1007/s11033-025-10242-7. [Google Scholar] [PubMed] [CrossRef]
243. Li MY, Zhang Q, Li J, Zengin G. Food and medicine homology in cancer treatment: traditional thoughts collide with scientific evidence. Food Med Homol. 2025;2(3):9420120. doi:10.26599/FMH.2025.9420120. [Google Scholar] [CrossRef]
244. Yin X, Xu R, Song J, Ruze R, Chen Y, Wang C, et al. Lipid metabolism in pancreatic cancer: emerging roles and potential targets. Cancer Commun. 2022;42(12):1234–56. doi:10.1002/cac2.12360. [Google Scholar] [PubMed] [CrossRef]
245. Chan DC, Watts GF. Inhibition of the ANGPTL3/8 complex for the prevention and treatment of atherosclerotic cardiovascular disease. Curr Atheroscler Rep. 2024;27(1):6. doi:10.1007/s11883-024-01254-y. [Google Scholar] [PubMed] [CrossRef]
246. Bremova-Ertl T, Hofmann J, Stucki J, Vossenkaul A, Gautschi M. Inborn errors of metabolism with ataxia: current and future treatment options. Cells. 2023;12(18):2314. doi:10.3390/cells12182314. [Google Scholar] [PubMed] [CrossRef]
247. Cai J, Liu Z, Chen S, Zhang J, Li H, Wang X, et al. Engineered circular RNA-based DLL3-targeted CAR-T therapy for small cell lung cancer. Exp Hematol Oncol. 2025;14(1):35. doi:10.1186/s40164-025-00625-8. [Google Scholar] [PubMed] [CrossRef]
248. Judge A, MacLachlan I. Overcoming the innate immune response to small interfering RNA. Hum Gene Ther. 2008;19(2):111–24. doi:10.1089/hum.2007.179. [Google Scholar] [PubMed] [CrossRef]
249. Robbins M, Judge A, MacLachlan I. siRNA and innate immunity. Oligonucleotides. 2009;19(2):89–102. doi:10.1089/oli.2009.0180. [Google Scholar] [PubMed] [CrossRef]
250. Draz MS, Fang BA, Zhang P, Hu Z, Gu S, Weng KC, et al. Nanoparticle-mediated systemic delivery of siRNA for treatment of cancers and viral infections. Theranostics. 2014;4(9):872–92. doi:10.7150/thno.9404. [Google Scholar] [PubMed] [CrossRef]
251. Hama Faraj GS, Hussen BM, Abdullah SR, Fatih Rasul M, Hajiesmaeili Y, Baniahmad A, et al. Advanced approaches of the use of circRNAs as a replacement for cancer therapy. Noncoding RNA Res. 2024;9(3):811–30. doi:10.1016/j.ncrna.2024.03.012. [Google Scholar] [PubMed] [CrossRef]
252. Chen Z, Krishnamachary B, Pacheco-Torres J, Penet MF, Bhujwalla ZM. Theranostic small interfering RNA nanoparticles in cancer precision nanomedicine. Interdiscip Rev Nanomed Nanobiotechnol. 2020;12(2):e1595. doi:10.1002/wnan.1595. [Google Scholar] [PubMed] [CrossRef]
253. Dancy JG, Wadajkar AS, Connolly NP, Galisteo R, Ames HM, Peng S, et al. Decreased nonspecific adhesivity, receptor-targeted therapeutic nanoparticles for primary and metastatic breast cancer. Sci Adv. 2020;6(3):eaax3931. doi:10.1126/sciadv.aax3931. [Google Scholar] [PubMed] [CrossRef]
254. Kara G, Calin GA, Ozpolat B. RNAi-based therapeutics and tumor targeted delivery in cancer. Adv Drug Deliv Rev. 2022;182:114113. doi:10.1016/j.addr.2022.114113. [Google Scholar] [PubMed] [CrossRef]
Cite This Article
 Copyright © 2026 The Author(s). Published by Tech Science Press.
Copyright © 2026 The Author(s). Published by Tech Science Press.This work is licensed under a Creative Commons Attribution 4.0 International License , which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.


 Submit a Paper
Submit a Paper Propose a Special lssue
Propose a Special lssue View Full Text
View Full Text Download PDF
Download PDF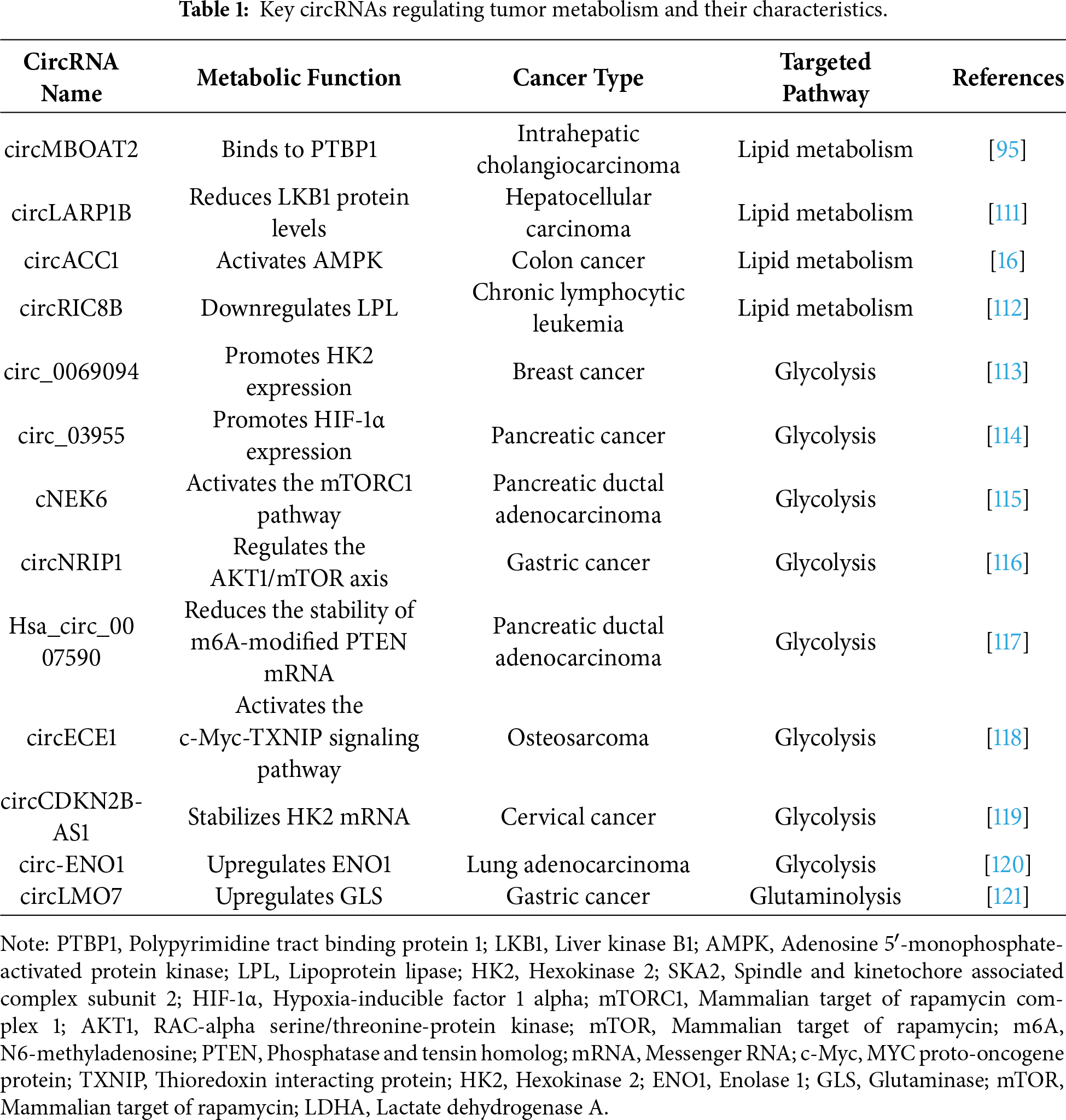
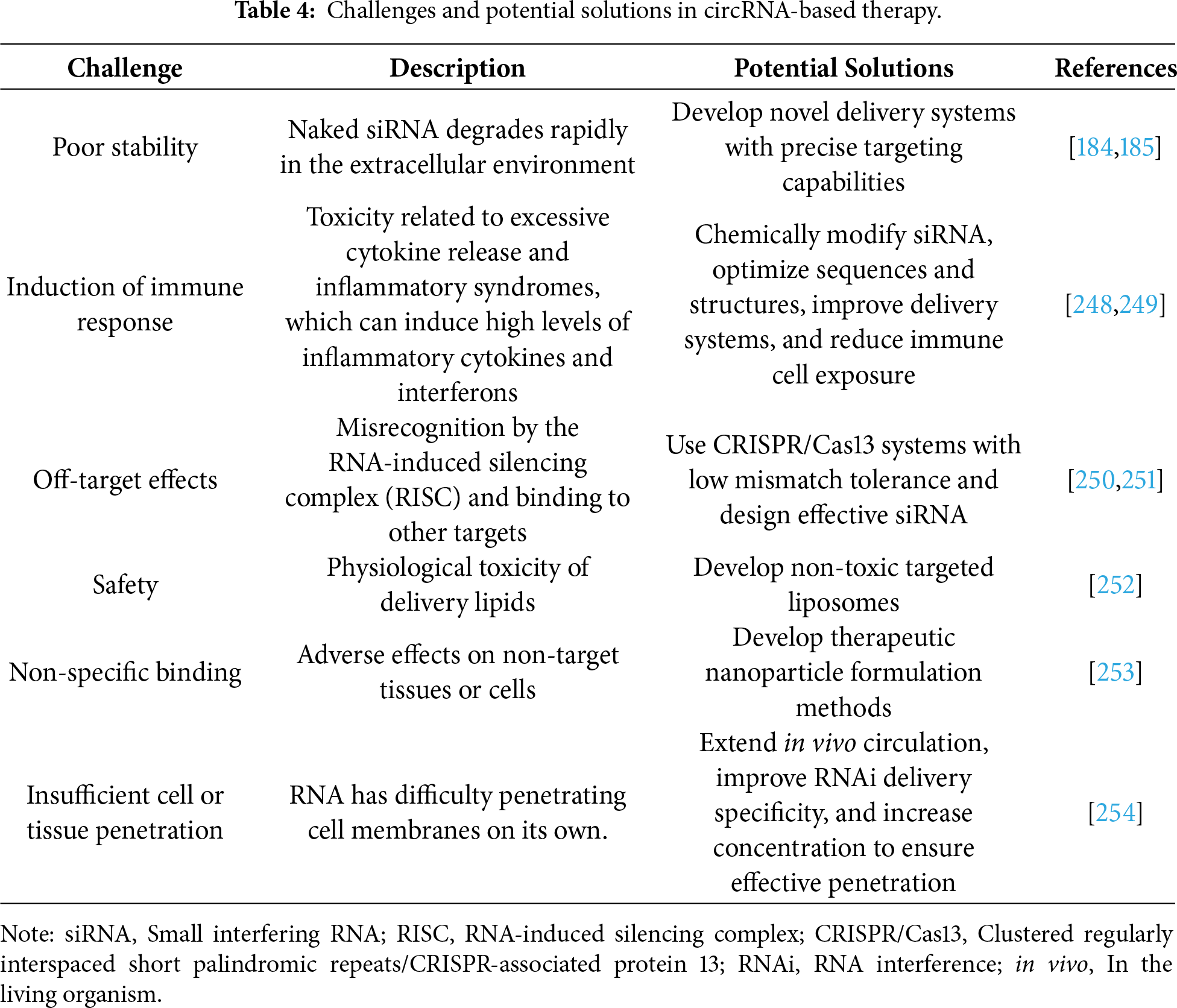
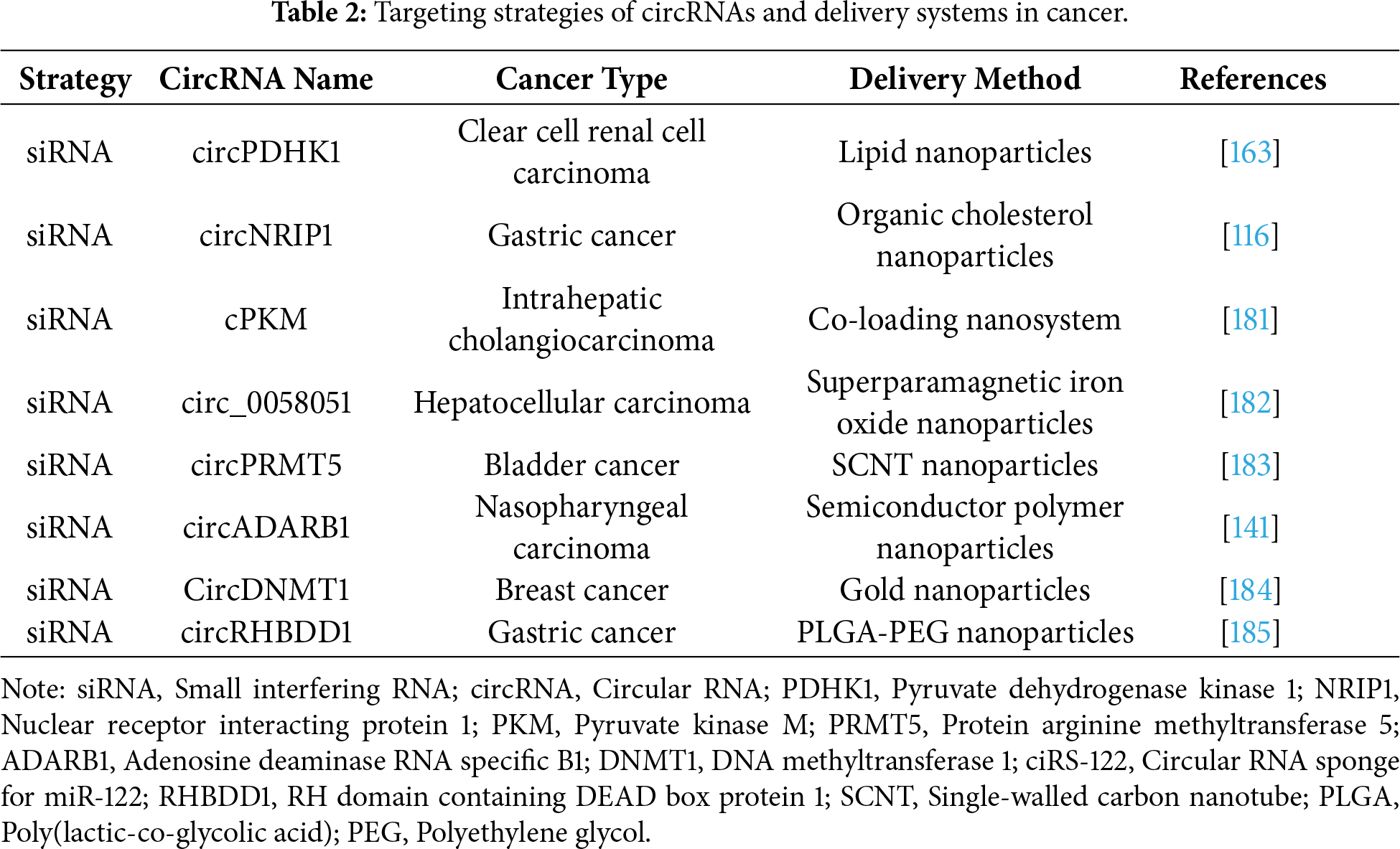
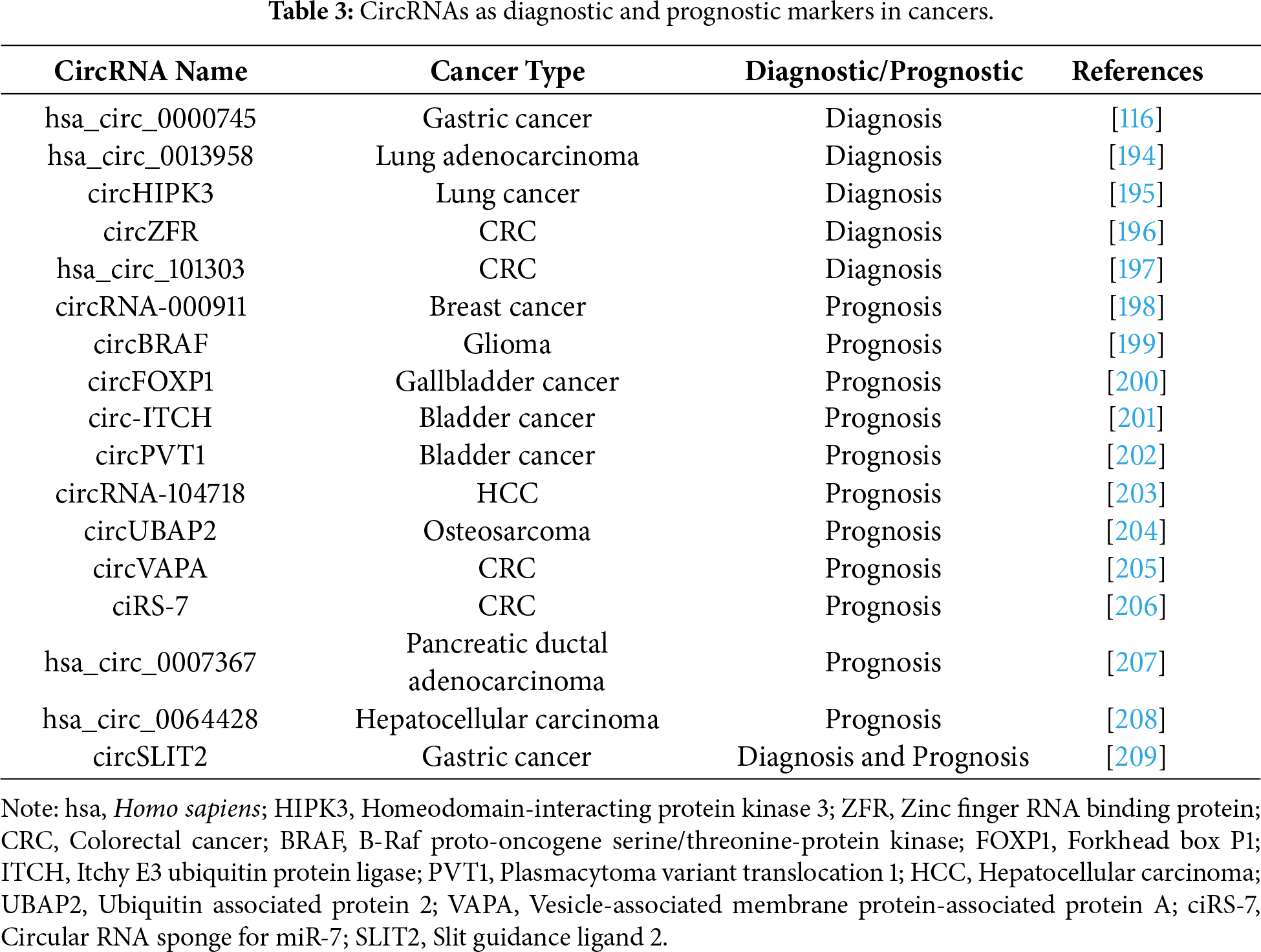

 Downloads
Downloads
 Citation Tools
Citation Tools