 Open Access
Open Access
ARTICLE
Evaluation, Validation, and Application of Sex-Specific Molecular Marker in Kiwifruit (Actinidia spp.)
1 China-New Zealand the Belt and Road Joint Laboratory on Kiwifruit, Kiwifruit Breeding and Utilization Key Laboratory of Sichuan Province, Sichuan Academy of Natural Resource Sciences, Chengdu, China
2 Citrus Research Institute, Southwest University, Chongqing, China
3 Chendu Agricultural College, Chengdu, China
* Corresponding Author: Yue Xie. Email:
(This article belongs to the Special Issue: Advances in Plant Breeding and Genetic Improvement: Leveraging Molecular Markers and Novel Genetic Strategies)
Phyton-International Journal of Experimental Botany 2026, 95(2), 5 https://doi.org/10.32604/phyton.2026.074974
Received 22 October 2025; Accepted 08 January 2026; Issue published 28 February 2026
Abstract
The genus Actinidia is primarily functionally dioecious, and early sex identification plays a crucial role in improving breeding efficiency and reducing production costs. In this study, the accuracy of three sex-linked molecular markers (SyGI [Shy Girl], FrBy [Friendly Boy], and SmY1) in sex identification was evaluated in various Actinidia species. The selected marker products were subsequently cloned and sequenced in six wild Actinidia species. Ninety-six wild A. chinensis chinensis accessions and 74 A. chinensis deliciosa accessions, most of which were wild, with only one cultivated, were used for comprehensive primer validation. Thirty-three juvenile A. chinensis chinensis hybrid seedlings were used for practical application tests. The results showed that the marker SyGI accurately identified the sex of 20 samples from six Actinidia species and 96 A. chinensis chinensis accessions with 100% reliability. For Actinidia chinensis deliciosa, the identification accuracy reached 98.65%. Sequence analysis revealed that SyGI shared the highest similarity with the male-specific genomic region. Furthermore, SyGI achieved 100% accuracy in identifying the sex of 33 juvenile A. chinensis chinensis individuals. The findings confirm that the SyGI marker possesses high accuracy, strong specificity, and broad applicability, making it a valuable tool for kiwifruit breeding programs. The cloned sequences from wild Actinidia species also provide important references for future research on the mechanisms of sexual evolution and determination.Keywords
Supplementary Material
Supplementary Material FileKiwifruit (Actinidia Lindl.) is a perennial, dioecious, and economically important fruit crop native to China [1]. Often referred to as the “world’s precious fruit” and the “king of fruits”, kiwifruit is highly valued for its exceptional nutritional content, health benefits, and medicinal properties [2,3]. As a major global fruit crop, its market demand continues to rise steadily alongside the expansion of the kiwifruit industry.
Species within the Actinidia genus are primarily functionally dioecious, bearing Type I unisexual flowers. In these flowers, males possess vestigial pistils, while females have abortive stamens, resulting in the presence of only one functional reproductive organ per flower [4]. Based on the development of unisexual flowers, Mayer and Charlesworth (1991) classified dioecious plants with Type I unisexual flowers as showing cryptic dioecy [5].
Kiwifruit is a perennial, cryptically dioecious woody plant, and distinguishing between male and female plants at the seedling stage is not possible through morphology. Moreover, the juvenile phase lasts more than four years, with male individuals often comprising half or more of the progeny [6]. In breeding and improvement programs, growing seedlings to maturity involves considerable time and financial investment, especially when male plants must later be removed. Therefore, developing and applying accurate methods for early sex identification is essential for improving breeding efficiency and minimizing production costs in kiwifruit cultivation.
Extensive research has been dedicated to developing sex identification techniques in kiwifruit, employing approaches such as morphological characterization, physiological and biochemical analyses, and molecular biology methods [7,8,9]. Among these, DNA-based molecular markers have become the predominant tool due to their speed and reliable accuracy. Molecular markers, such as SSR, ISSR, RAPD, and sex-linked gene loci, have frequently been used for early sex identification in dioecious plants. For example, in persimmons, the male-linked gene OGI, located on the Y chromosome, plays a crucial role in sex determination and has been used for early sex identification of persimmon plants [10,11]. In kiwifruit, a variety of molecular markers have been developed and applied for sex identification, including RAPD markers (SmY, SmX), SCAR markers (SmY1, SmX), SSR markers (A001, A002, A003), etc. [12,13,14]. However, most of these identified molecular markers are polymorphic, and their linkage to sex remains inconclusive. In recent years, the practical performance of some molecular markers has proven inconsistent. For example, the male-linked RAPD marker S1032-850 has shown poor reliability during subsequent validation [15]. Among these markers, the SCAR marker SmY1 has demonstrated high accuracy for sex identification in A. chinensis chinensis [15,16]; however, its effectiveness in other Actinidia species remains uncertain. SmY1 is a marker of male sex in kiwifruit. It was originally developed as a RAPD marker by Harvey et al., and was later converted into a more stable SCAR marker by Gill et al. [12,13]. The present study investigated the use of SmY1 for identification in different Actinidia species to explore its universal applicability.
Understanding the mechanisms of sex determination is essential for guiding the development of reliable molecular markers in dioecious plants. Key areas of research include the study of sex chromosomes, sex-determining genes, sex-determination regions, and their associated regulatory elements. Research into plant sex determination is relatively recent and marked by considerable variability and evolutionary instability. In most plants, sex chromosomes remain at early evolutionary stages, retaining high levels of sequence homology [17]. The prevailing hypothesis posits that sex chromosomes originate from ancestral autosomes [18]. In the early stages of this transition, sex chromosomes are morphologically indistinguishable from autosomes. Over time, they undergo structural and functional differentiation, eventually forming heteromorphic sex chromosomes. This evolutionary process involves divergence between X (or Z) and Y (or W) chromosomes, as well as between sex chromosomes and autosomes [19].
In kiwifruit, studies have shown that the X and Y chromosomes in A. chinensis chinensis are similar in size and structure, aligning with Stage 2 of the six-phase model of sex chromosome evolution proposed by Ming et al. [20]. The subtelomeric region of the Y chromosome contains sex-determining loci that do not undergo recombination with the corresponding X chromosome—an established feature of sex-determining loci [21]. Additionally, a small male-specific region has been identified on the proto-Y chromosome, accompanied by the appearance of exclusively male or female plants. This pattern of sexual differentiation is comparable to that observed in asparagus (Asparagus officinalis), suggesting that Actinidia sex chromosomes are still in an early evolutionary phase.
Regarding genetic models of sex determination, Charlesworth B and Charlesworth D [22] proposed the two-gene mutation hypothesis, which suggests that at least two mutations are needed to develop a functional Y chromosome: a dominant mutation that suppresses female development (SuF), and a recessive mutation causing male sterility [23,24]. Kiwifruit employs an XY sex-determination system consistent with this two-gene sex-determination model [25].
In research on sex-determining regions (SDRs) and sex-determining genes, the sex-specific loci have been identified in several dioecious species, including white campion (Silene latifolia) [26], papaya (Carica papaya) [27], garden asparagus (Asparagus officinalis) [28], poplar (Populus spp.) [29,30], and hemp (Cannabis sativa) [31]. Advances in kiwifruit genomics have recently led to the identification of two key sex-determining genes: SyGI (Shy Girl), which suppresses female development, and FrBy (Friendly Boy), a masculinizing factor. These genes act together to regulate sexual dimorphism in Actinidia [25,32]. Recently, several reports have described the use of the SyGI and FrBy genes for sex identification in kiwifruit [33,34,35]. The design of the SyGI primers employed in these studies was based on the coding DNA sequence (CDS) of the SyGI gene. However, SyGI has a homolog, Achn384741, located on an autosome, with a sequence similarity of 99% [36], resulting in strong resemblance between the CDS regions of SyGI and Achn384741. Fang et al. [35] observed poor amplification efficiencies for primers designed for the exonic region of SyGI, suggesting the presence of additional homologs of the SyGI sequence within the kiwifruit genome. The present study selected a pair of specific primers from a recent study [36] that could avoid interference from homologous sequences during amplification and exclusively amplified the non-coding region sequences upstream of the CDS region of the SyGI gene.
Kiwifruit have significant genetic diversity, complex ploidy levels, and high heterozygosity [37]. Wild kiwifruit show marked genetic diversity and variability, thereby representing a crucial breeding resource [38,39,40]. While De Mori et al. have evaluated sex-specific molecular markers, including SyGI and FrBy, for several Actinidia species, there is limited information on the validation of the SyGI and FrBy markers for large-scale sex identification in wild kiwifruit [33]. Therefore, it is essential to investigate the applicability of these two sex-determining genes in various wild kiwifruit populations.
This study focused on the sex-linked SmY1 marker, as well as the recently reported SyGI and FrBy molecular markers. While SmY1 has been shown to be highly accurate in several studies, its universal applicability is unknown. These markers were investigated in different wild kiwifruit species to evaluate their suitability. Male and female plants from six Actinidia species were used to screen for highly conserved and accurate male-specific molecular marker. The identified marker products were cloned and analyzed through sequence alignment. Their reliability and broad applicability were further evaluated through large-scale validation in two important wild species, A. chinensis chinensis and A. chinensis deliciosa. The primary objective was to identify and validate robust sex-specific molecular marker with high universality and accuracy across Actinidia species. The selected marker aims to enable rapid and reliable sex identification in kiwifruit, thereby improving breeding efficiency and reducing associated time and costs. Additionally, comparative cloning in various wild Actinidia species will provide a molecular basis for further investigation into mechanisms of sex determination within the genus.
Female and male individuals from six Actinidia species (A. chinensis chinensis, A. chinensis deliciosa, A. eriantha, A. arguta, A. hemsleyana and A. callosa) were selected for initial primer screening (Table 1). For the molecular cloning of sex-specific marker products, leaf samples were collected from male individuals of six wild Actinidia species (A. chinensis chinensis, A. chinensis deliciosa, A. eriantha, A. arguta, A. hemsleyana, and A. callosa) (Table 2). For further validation and application of the molecular marker, leaf samples were collected from two groups: (1) 96 mature wild A. chinensis chinensis and 74 mature A. chinensis deliciosa plants, including both sexes, with geographical distributions shown in Table S1; (2) 33 two-year-old juvenile hybrid A. chinensis chinensis seedlings of unknown sex (Table S2). Young leaves from all specimens were immediately flash-frozen in liquid nitrogen and stored at −80°C for subsequent molecular analysis. Thirty-three of the seedlings shown in Table S2 were used for field verification when they were three years old. All plant materials were sourced from the kiwifruit breeding base of the Sichuan Provincial Academy of Natural Resource Sciences, located in Sichuan Province, China.
Table 1: Plant materials used for sex-specific molecular marker screening in Actinidia.
| Number | Sample Name | Species | Sex | Source of Materials |
|---|---|---|---|---|
| 1 | JXQS-EA-04238 | A. eriantha | female | Wild material; Jiangxi Province, China |
| 2 | JXQS-EA-04243 | A. eriantha | male | Wild material; Jiangxi Province, China |
| 3 | JXfujun-04212-OP01 | A. arguta | female | Wild material; Jiangxi Province, China |
| 4 | HNNX-AE-03098-OP02 | A. arguta | female | Wild material; Henan Province, China |
| 5 | HNNX-AE-03098-OP07 | A. arguta | male | Wild material; Henan Province, China |
| 6 | JXfujun-04212-OP04 | A. hemsleyana | male | Wild material; Jiangxi Province, China |
| 7 | JXfujun-04212-OP10 | A. hemsleyana | female | Wild material; Jiangxi Province, China |
| 8 | JXFX-CF-04146 | A. callosa | female | Wild material; Jiangxi Province, China |
| 9 | TJ-CF-10 | A. callosa | male | Wild material; Tongjiang County, Sichuan Province, China |
| 10 | FD-DA-2 | A. chinensis deliciosa | female | Wild material; Fengdu County, Chongqing City, China |
| 11 | WS-DA-20 | A. chinensis deliciosa | male | Wild material; Wushan mountains, Chongqing City, China |
| 12 | TJ-DD-19 | A. chinensis deliciosa | female | Wild material; Tongjiang County, Sichuan Province, China |
| 13 | Emei03103 | A. chinensis deliciosa | female | Wild material; Mount Emei, Sichuan Province, China |
| 14 | ChengMi | A. chinensis deliciosa | female | Cultivar |
| 15 | WS-DA-58 | A. chinensis deliciosa | female | Wild material; Wushan mountains, Chongqing City, China |
| 16 | WS-DA-17 | A. chinensis deliciosa | male | Wild material; Wushan mountains, Chongqing City, China |
| 17 | PZ-DA-7 | A. chinensis deliciosa | male | Wild material; Pengzhou City, Sichuan Province, China |
| 18 | TJ-DA-15♂ | A. chinensis deliciosa | female | Wild material; Tongjiang County, Sichuan Province, China |
| 19 | HNNX-CK-03095 | A. chinensis chinensis | male | Wild material; Henan Province, China |
| 20 | HNNX-CK-03086 | A. chinensis chinensis | female | Wild material; Henan Province, China |
Table 2: Plant materials used for molecular cloning of sex-specific marker products.
| Sample Name | Species | Sex | Source of Materials |
|---|---|---|---|
| 04266 | A. chinensis chinensis | male | Wild material; Jiangxi Province, China |
| WS-DA-54 | A. chinensis deliciosa | male | Wild material; Wushan mountains, Chongqing City, China |
| JXQS-EA-04243 | A. eriantha | male | Wild material; Jiangxi Province, China |
| HNNX-AE-03098-OP07 | A. arguta | male | Wild material; Henan Province, China |
| JXfujun-04212-OP04 | A. hemsleyana | male | Wild material; Jiangxi Province, China |
| TJ-CF-10 | A. callosa | male | Wild material; Tongjiang County, Sichuan Province, China |
Genomic DNA was extracted from kiwifruit leaf tissue using the Plant Genomic DNA Extraction Kit (Product No. DN32; BIOFIT BIOTECHNOLOGIES Co., Ltd., China), following the manufacturer’s protocol. DNA purity and concentration were measured using a microvolume spectrophotometer. Extracted DNA samples were stored at −80°C for later use.
2.2.2 Selection of Sex-Linked Molecular Markers
Sex-linked molecular markers previously reported in Actinidia spp. were selected based on published studies. These markers are listed in Table 3.
Table 3: Primers for sex-linked molecular markers used in kiwifruit.
| Primers | Forward Primer Sequences | Reverse Primer Sequences | Product Size (bp) | Source References |
|---|---|---|---|---|
| SyGI | CAAATACAACATCTGAATTTGTTTCTTA | TCTAAATGCCATGATCAATTCTCACTAATTATCAC | 308 | Akagi et al., 2018 [36] |
| SmY1 | TCGCAATTCGTTAGGGATGATGCG | CATAATCAACCATCCATAAAAACCAT | 770 | Gill et al., 1998 [13] |
| FRBY | ATGGCAAAGTGGTTCTCTCTCCAT | TTAACAAACCCAAACCCTAAAATAAAC | 723 | Akagi et al., 2019 [25] |
Polymerase chain reaction (PCR) was conducted in a 50 μL reaction volume to amplify target sequences (SmY1, SyGI, and FRBY). Each reaction contained 1 μL of template DNA, 25 μL of 2× EasyTaq® PCR SuperMix, 1 μL of each primer (10 μM), and 22 μL nuclease-free water. The thermal cycling protocol was as follows: initial denaturation at 94°C for 5 min; 35 cycles of denaturation at 94°C for 30 s, annealing at the primer-specific temperature for 30 s (SyGI: 58.5°C for 30 s; SmY1 and FRBY: 58°C for 30 s), and extension at 72°C for 20–60 s (SyGI: 72°C for 30 s; SmY1 and FRBY: 72°C for 45 s); followed by a final extension at 72°C for 10 min.
2.2.4 Molecular Cloning and Sequence Analysis of Sex-Specific Marker
PCR amplification was conducted using genomic DNA from male individuals of six Actinidia species (A. chinensis chinensis, A. chinensis deliciosa, A. eriantha, A. arguta, A. hemsleyana, and A. callosa) with primers specific to the SyGI marker. The reaction conditions and PCR system were identical to those described in Section 2.2.3. Amplification products were separated via electrophoresis on 1% agarose gels, and bands corresponding to the expected size were excised and purified.
Purified PCR fragments were ligated into the pClone007 Versatile Simple Vector using a commercial cloning kit and subsequently transformed into competent Escherichia coli cells. Positive colonies were selected from each transformation and verified by colony PCR. A total of 12 confirmed positive clones were submitted for sequencing to Tsingke Biotechnology Co., Ltd. (China).
The resulting sequences were aligned using DNAMAN software (DNAMAN Application 6.0.3.99) and analyzed using BLAST (Basic Local Alignment Search Tool) against the NCBI nucleotide database to assess sequence specificity and homology.
By aligning the cloned sequences with their corresponding reference sequences which have the highest similarity (GenBank: LC260493.1, LC260495.1, LC260498.1) [36], the cloned sequences were identified as the 308-bp (A. chinensis chinensis, A. chinensis deliciosa, A. arguta, and A. hemsleyana) or 307-bp (A. eriantha and A. callosa) upstream sequence of the start codon. As an extension of the core findings of this study, we conducted a comparative analysis of cis-acting elements within the candidate proximal regulatory sequence of the SyGI gene cloned from six wild Actinidia species. Additionally, we analyzed the impact of sequence variations on these cis-acting elements. This extension study aimed to preliminarily explore: (1) whether the sequence variations across different species lead to the generation or disruption of cis-acting elements; (2) how these variations influence the distribution and composition of conserved cis-acting elements. Through the above comparative analysis, specific candidate sites and hypotheses can be provided to understand the natural variation in SyGI expression regulation. The cis-acting elements in this region were identified and analyzed using the PlantCARE website (http://bioinformatics.psb.ugent.be/webtools/plantcare/html) [41].
3.1 Primer Evaluation and Selection
PCR amplification results using SyGI, SmY1, and FRBY primers across six Actinidia species (A. chinensis chinensis, A. chinensis deliciosa, A. eriantha, A. arguta, A. hemsleyana, and A. callosa) are shown in Fig. 1. Notably, the SyGI primers consistently amplified a single, clear band in all male samples, with no amplification observed in female plants. This marker accurately distinguished male and female individuals across all six species, achieving 100% identification accuracy.
In contrast, the SmY1 primers yielded target bands in only three male A. chinensis deliciosa and one male A. chinensis chinensis accessions, indicating limited applicability and lower accuracy. The FRBY primers produced multiple nonspecific bands in the female samples, undermining their specificity and reliability for sex identification. Based on these comparative results, SyGI was identified as the most specific and accurate molecular marker and was selected for use in subsequent analyses.
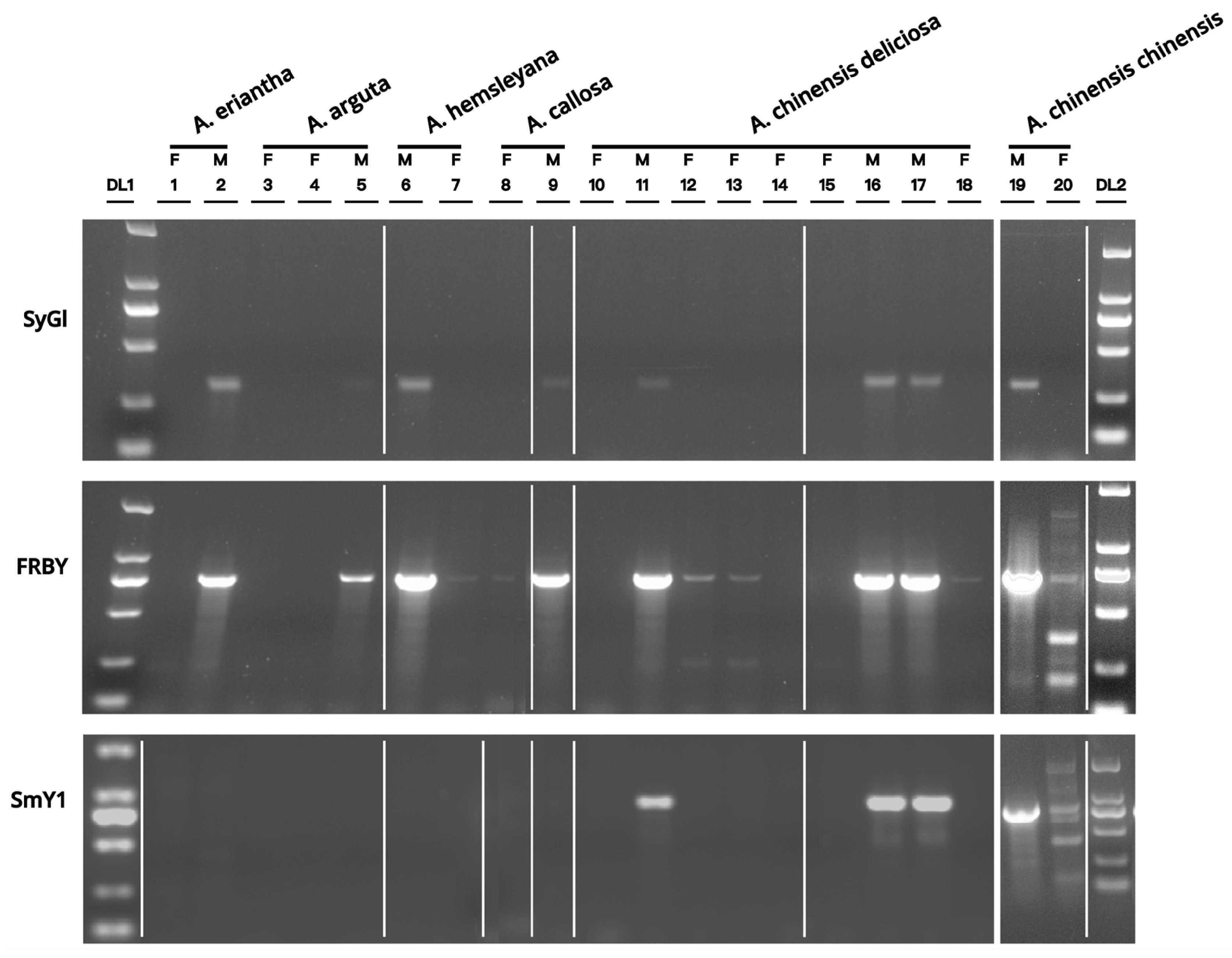
Figure 1: Electrophoresis of PCR products from sex-linked molecular markers. F: female sample. M: male sample. DL1-2: DL2000 DNA marker. 1–20: Plant materials used for the screening of sex-specific moleculars in Actinidia (details are shown in Table 1). The lanes were rearranged from the same gel for ease of comparison. White lines indicate the borders where lanes were rearranged.
3.2 Comparative Analysis of Cloned SyGI Sequences
Cloning and sequencing of the SyGI marker products revealed a 307-bp fragment in A. eriantha and A. callosa, while a 308-bp sequence was obtained from A. chinensis chinensis, A. chinensis deliciosa, A. arguta, and A. hemsleyana (Fig. 2). Sequence alignment demonstrated a high level of similarity (98.43%) among the SyGI sequences from male individuals across all six wild species, with few base differences.

Figure 2: Sequence comparison of cloned SyGI fragments from six Actinidia species.
BLAST analysis against the NCBI nucleotide database showed that the SyGI sequences from A. chinensis chinensis and A. chinensis deliciosa exhibited 100% identity with reference sequences LC260493.1, LC218546.1, and LC260494.1 (Fig. 3A). Sequences from A. callosa and A. eriantha also displayed high similarity to these references, with identity scores of 99.19% and 99.51%, respectively, with a few base differences between them (Fig. 3B,C).

Figure 3: BLAST alignment of SyGI sequences from six Actinidia species. (A) BLAST alignment of SyGI sequences from A. chinensis chinensis and A. chinensis deliciosa; (B) BLAST alignment of SyGI sequences from A. callosa; (C) BLAST alignment of SyGI sequences from A. eriantha; (D) BLAST alignment of SyGI sequences from A. arguta; (E) BLAST alignment of SyGI sequences from A. hemsleyana.
Notably, the reference sequences (LC260493.1, LC218546.1, and LC260494.1) are known male-specific Y-linked sequences from Actinidia [36].
Additionally, the wild A. arguta amplicon obtained in this study showed complete sequence identity (100%) with the A. arguta SyGI reference sequence (LC260498.1; Fig. 3D). The SyGI amplicons from A. hemsleyana displayed 99.68% similarity to the A. rufa SyGI reference sequence, with one base difference between them (LC260495.1; Fig. 3E).
3.3 Validation of Sex-Specific Molecular Marker in Kiwifruit
3.3.1 Validation of Mature Actinidia Plants
PCR amplification using SyGI-specific primers was carried out on 96 A. chinensis chinensis and 74 A. chinensis deliciosa accessions to assess marker performance in mature plants (most of the plants were wild, with only one cultivated; details are shown in Table S1). As shown in Fig. 4, the SyGI marker successfully identified the sex of all 96 A. chinensis chinensis samples with 100% accuracy. Specific target bands were amplified in all male samples, while no product was detected in female individuals (Fig. 4).
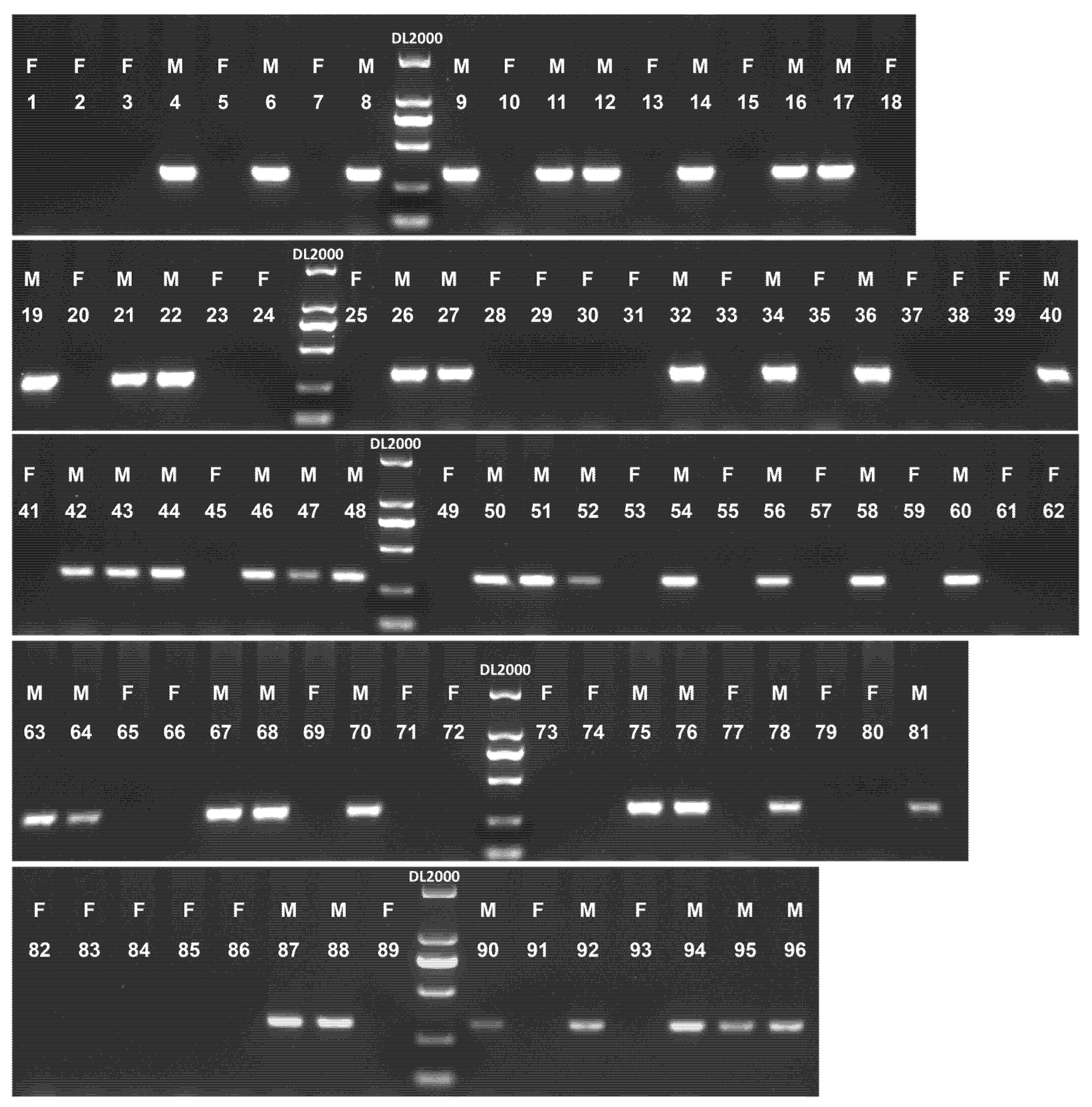

Figure 4: Sex identification of 96 mature A. chinensis chinensis accessions using the SyGI marker. F: female accession. M: male accession. DL2000: DL2000 DNA marker. 1–96: 96 A. chinensis chinensis samples used for validation (details are shown in Table S1).
Similarly, in A. chinensis deliciosa (n = 74, accessions #97–170), SyGI accurately determined the sex in 73 out of 74 samples. One accession (#130) failed to yield a detectable amplification product. The resulting sex identification accuracy was 98.65% (Fig. 5).
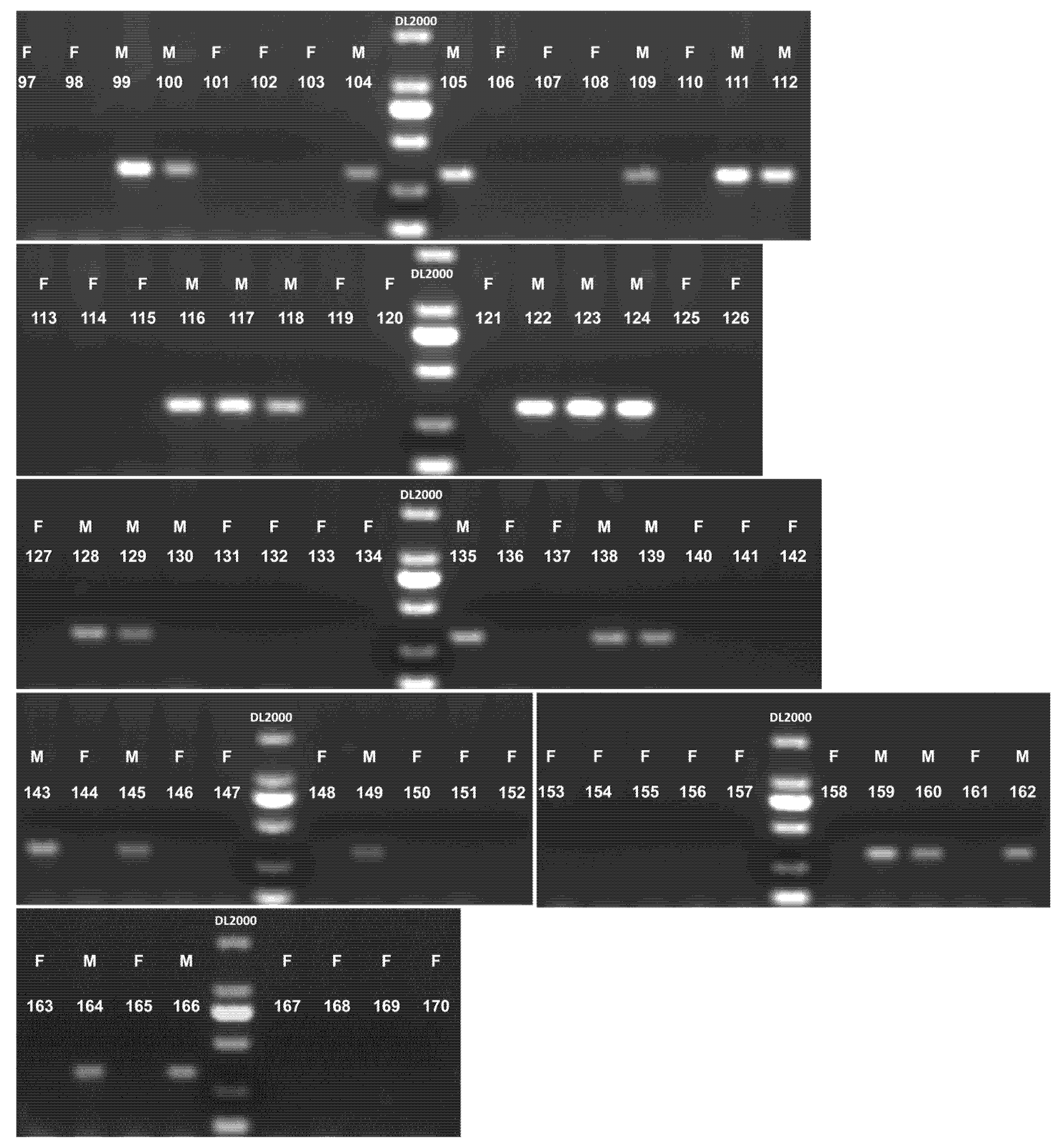
Figure 5: Sex identification of 74 mature A. chinensis deliciosa accessions using the SyGI marker. F: female accession. M: male accession. DL2000: DL2000 DNA marker. 97–170: 74 A. chinensis deliciosa samples used for validation (details are shown in Table S1).
3.3.2 Validation and Application of Juvenile Actinidia Plants
To assess the practical utility in early-stage breeding, the SyGI marker was applied to 33 juvenile A. chinensis chinensis seedlings (Table S2). The PCR results (Fig. 6) enabled the reliable identification of sex at the seedling stage. Subsequent field validation (Table 4) during the flowering phase confirmed complete agreement between molecular predictions and observed phenotypic sex expression, thereby validating the accuracy of the SyGI marker for early sex determination in A. chinensis chinensis.
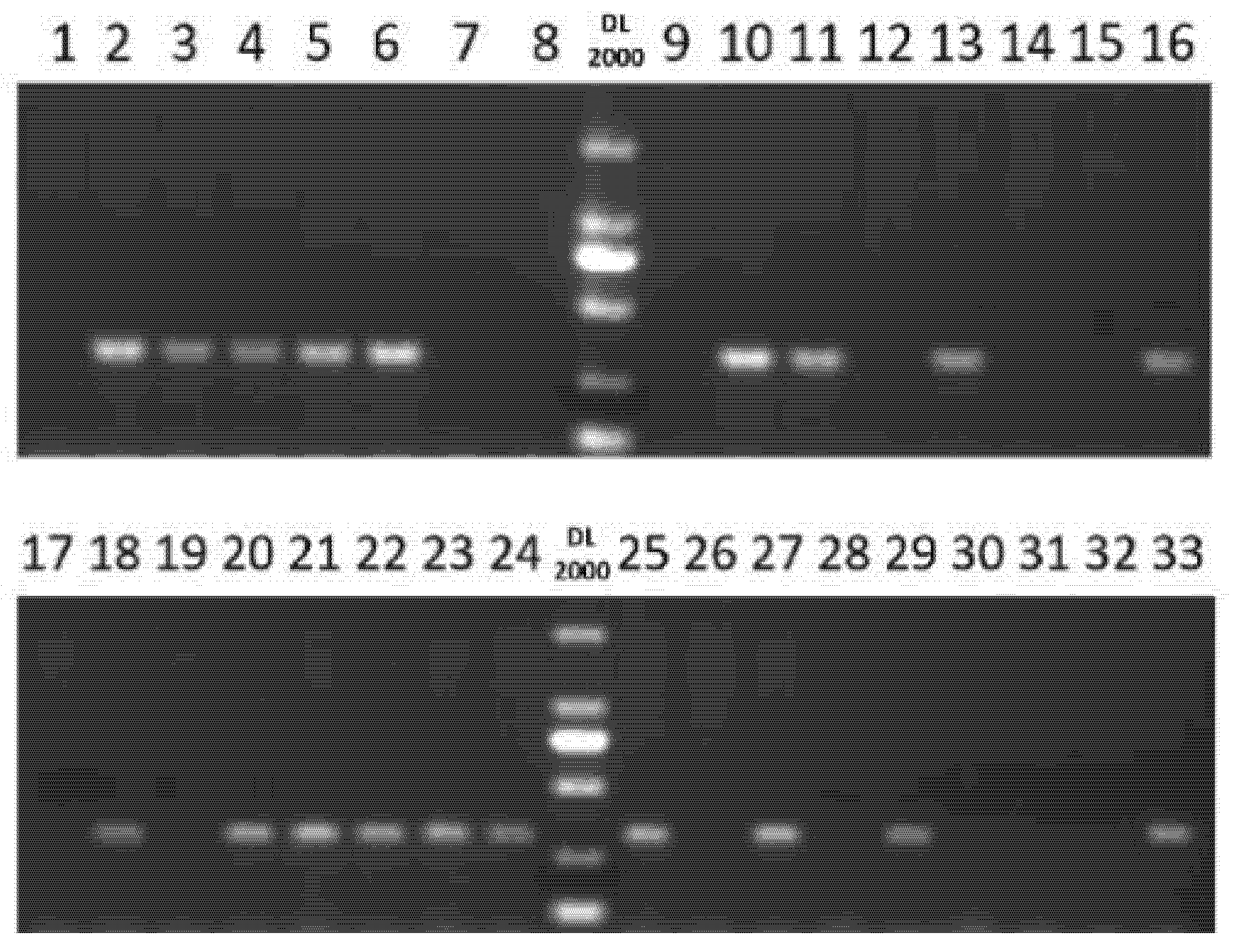
Figure 6: PCR-based sex identification of 33 juvenile A. chinensis chinensis seedlings using the SyGI marker. 1–33: 33 juvenile A. chinensis chinensis seedling materials (details are shown in Table S2). DL2000: DL2000 DNA marker.
Table 4: Early sex identification of juvenile A. chinensis chinensis seedlings.
| Sample Number | Sex Type (Laboratory Identification) | Sex Type (Field Identification) |
|---|---|---|
| 1 | female | female |
| 2 | male | male |
| 3 | male | male |
| 4 | male | male |
| 5 | male | male |
| 6 | male | male |
| 7 | female | female |
| 8 | female | female |
| 9 | female | female |
| 10 | male | male |
| 11 | male | male |
| 12 | female | female |
| 13 | male | male |
| 14 | female | female |
| 15 | female | female |
| 16 | male | male |
| 17 | female | female |
| 18 | male | male |
| 19 | female | female |
| 20 | male | male |
| 21 | male | male |
| 22 | male | male |
| 23 | male | male |
| 24 | male | male |
| 25 | male | male |
| 26 | female | female |
| 27 | male | male |
| 28 | female | female |
| 29 | male | male |
| 30 | female | female |
| 31 | female | female |
| 32 | female | female |
| 33 | male | male |
Actinidia spp. has become an economically significant fruit crop globally. However, its dioecious reproductive system and lengthy breeding cycle present challenges for breeding programs. Early-stage sex identification is therefore critical to optimize resource allocation and reduce labor and financial inputs. Traditional sex determination methods, such as morphological and physiological-biochemical assessments, are often limited by environmental influences and developmental timing. In contrast, advances in molecular biology have enabled the development of genetic approaches that offer greater reliability for sex determination.
In this study, the SmY1 marker showed limited accuracy and low universality in different wild species. SmY1 is a sex-linked molecular marker developed using the BSA method in diploid A. chinensis chinensis [12]. Therefore, it is possible that the location or sequence of sex-specific regions may vary among different kiwifruit species. The Y chromosome in kiwifruit is relatively primitive and has not fully evolved [20]. Akagi et al. investigated the evolution of dioecy in kiwifruit, showing that the sex-linked segments form a small 0.49-Mb male-specific region (MSY) on chromosome 25, flanked by zones of recombination [36]. This sex-determining region contains 61 genes. Based on the patterns of genetic diversity observed in these candidate genes, the authors concluded that dioecy likely emerged 20 million years ago in kiwifruit, possibly following an ancient polyploidization event [42]. A recent study identified the sex-determining region (SDR) of A. chinensis chinensis on chromosome 25 (Chr25), while the SDR of A. eriantha has been localized to chromosome 12 (Chr12), indicating a novel interchromosomal translocation between the sex chromosomes and autosomes in kiwifruit [43]. This study observed low chromosomal colinearity between the SDRs of A. chinensis chinensis and A. eriantha. Akagi et al. reported similar findings in A. rufa, A. arguta, and A. polygam [44].
Sex-determining genes are located in the sex-determining region of the sex chromosome and represent the molecular basis for the differentiation between the male and female sexes. The two-mutation model of sex determination has recently been reported in plants such as Actinidia spp. [25], garden asparagus (Asparagus officinalis) [28], and grapes (Vitis spp.) [45,46]. In kiwifruit, the SyGI and FrBy genes on the Y chromosome function independently as a respective suppressor of feminization (SuF) and for maintaining male (M) sex, thereby regulating sex differentiation. Previous studies have demonstrated conservation of both SyGI and FrBy in a variety of kiwifruit species [25]. However, the present study found that FrBy performed poorly, with some amplification of non-specific bands. The PCR amplification results revealed the presence of target bands corresponding to the size of the FrBy gene amplification product in female samples. De Mori et al. also observed amplification of FrBy in female genotypes of four different Actinidia species, but they ruled out the possibility of an intact and functional copy of the gene existing in these species [33]. Recently, Wen et al. also found that previously published FrBy primers [25] only amplified a specific band in one of the four male samples [34]. However, they subsequently designed a series of primers based on the FrBy sequence, leading to the successful identification of suitable primers in later studies. Collectively, these observations lead us to posit that: Sequence variations (e.g., SNPs) may exist in the primer-binding region of the FrBy gene in a population-specific manner, which could explain both the differences in amplification results of the same primer pair in male plants of different genetic backgrounds and the fact that redesigning primers can resolve amplification failures. On the X chromosome and/or autosomes, there may exist sequences highly homologous to the FrBy. The conservation of these homologous sequences likely varies across germplasms. In the wild materials tested in this study, these homologous sequences may remain well preserved, leading to the appearance of amplification bands in female samples. We will subsequently validate this hypothesis through sequencing and genomic localization of PCR products derived from wild materials in this study.
In this study, the use of the sex-specific primers of SyGI was validated in a large sample of wild kiwifruit. Wild kiwifruit is known to exhibit marked genetic diversity [40,47]. A degree of gene exchange occurs among wild genotypes [38]. Wild kiwifruit is widely distributed across China. The samples of wild kiwifruit investigated in this study were sourced from Jiangxi Province, Henan Province, Sichuan Province, Zhejiang Province, and the Wuling Mountains. The majority of these materials were concentrated in Jiangxi and Henan Provinces. Jiangxi Province boasts exceptionally abundant wild kiwifruit resources, including species such as A. chinensis chinensis and A. eriantha [38]. The application of SyGI-specific primers to these wild kiwifruit resources revealed that the marker was highly accurate and specific in the identification of sex in these wild materials. It also can serve as a valuable reference for sex identification and genomic collinearity analyses in other dioecious crops. SyGI has been shown to be derived from a lineage-specific gene duplication [43]. One male A. chinensis deliciosa accession did not yield a positive amplification, potentially attributed to sequence deletions or structural variation in the sex-determination region, possibly due to historical recombination or hybridization events. In future follow-up studies, sequencing the SyGI locus in this accession may provide important insights into the genetic mechanisms underlying sex determination and evolution in A. chinensis deliciosa.
To assess whether SyGI is located within the male-specific region of the Y chromosome (MSY), cloning and sequencing of SyGI-amplified fragments from male individuals of six wild Actinidia species were performed. The comparative analysis of cloned SyGI sequences from six wild Actinidia species showed that these sequences had the highest similarity to either Shy Girl or the MSY sequences, confirming the male specificity of this marker. Notably, we identified SNPs in candidate proximal regulatory sequences of SyGI across different Actinidia species. The specific mechanism by which SyGI modulates sex determination remains unknown [48]. To explore the potential functional impact of these SNPs on the expression regulation of the SyGI gene, we conducted an extended analysis in this study. This extended analysis revealed lineage-specific differentiation by comparing differences in cis-acting elements within candidate proximal regulatory regions of SyGI gene across six wild Actinidia species (Fig. S1). It also directly linked key SNPs to variations in cis-acting elements (Fig. S2). These findings provided sequence evidence and hypotheses for understanding the expression regulation diversity and adaptive evolution of SyGI, a key gene in kiwifruit sex determination. The analysis indicated that specific SNPs directly mediated variations in key regulatory elements. For instance, in A. callosa, the SNPs at key sites may not only lead to the loss of a conserved MYB element but could also create a key element—AT-TATA-box, thereby potentially affecting both its developmental signal response and basal transcription efficiency [49,50]. The unique SNPs in A. hemsleyana and A. arguta may create a low-temperature response element, suggesting that their SyGI expression might directly sense environmental temperature signals. In A. eriantha, a distinct SNP may form a novel W-box near the MYB element, which could couple developmental regulation with defense response pathways [49,51]. These SNP-driven variations in the “regulatory code” indicated that natural selection may fine-tune the upstream sequence of SyGI to adapt to different ecological and developmental contexts. It should be noted that this analysis has limitations. Firstly, the examined “candidate proximal regulatory region” was operationally defined, and key distal regulatory elements may reside outside this region. Secondly, the conclusions were based on bioinformatic predictions, and their functional impacts require further validation through experiments such as reporter gene assays or EMSA. Nonetheless, this study translated sequence variations into concrete regulatory hypotheses, providing clear and testable targets for subsequent functional studies.
The ultimate goal of determining molecular sex markers is their application in sex identification at the early/juvenile stages in kiwifruit development. Currently, most sex identification methods for kiwifruit were developed and validated using materials of known sex [14,15], with few studies on the application of molecular sex markers in juvenile seedlings. The successful application of the SyGI marker for sex identification in juvenile plants used in this study confirmed the practical value of this molecular marker technology. The breeding cycle of kiwifruit could potentially be shortened by 2 to 3 years. This provides an efficient and reliable tool for the identification of kiwifruit sex.
Although the sex determination model of kiwifruit has been elucidated, the specific regulatory mechanisms underlying the process are not known [48]. Scientists are attempting to identify these regulatory mechanisms using approaches such as gene editing [52]. Recently, researchers have investigated the regulatory networks of sex differentiation by integrating transcriptome and metabolome analysis [53]. Plant sex is fundamentally a quantitative trait, associated with complex genetic inheritance patterns and regulatory mechanisms [54,55]. In the future, techniques such as multiplex PCR and KASP genotyping could be integrated into high-throughput kiwifruit breeding workflows. These techniques can combine multiple loci or link the loci of multiple traits with the loci of functional markers of core economic traits, thereby maximizing automation, data-driven approaches, and efficiency in breeding operations.
In summary, the SyGI marker selected in this study exhibits high male specificity, strong cross-species applicability, and excellent accuracy, making it a promising tool for field-based sex identification in kiwifruit. Its use can significantly enhance breeding efficiency by enabling early selection, thereby reducing time and resource investments. Furthermore, the data generated here contribute valuable insights for future development of sex-linked molecular markers and further investigations into the genetic regulation of sex determination in Actinidia spp.
Acknowledgement:
Funding Statement: This research was funded by Sichuan Science and Technology Program, grant numbers 2021YFYZ0010, 2023YFH0006, 2025YFHZ0295; The Basic Research Program of Sichuan Provincial Research Institutes, grant numbers 2024JDKY0001 and 2023JDKY0001.
Author Contributions: Conceptualization: Yue Xie, Hui Zhang, Mingzhang Li. Methodology: Hui Zhang, Yingchun He, Yue Xie. Software: Yingchun He, Min Hong, Yang Wang. Formal analysis: Hui Zhang, Min Hong, Kui Du. Investigation: Yue Xie, Hui Zhang. Resources: Yue Xie, Mingzhang Li, Qiguo Zhuang. Data curation: Hui Zhang. Writing—original draft preparation: Hui Zhang. Writing—review and editing: Hui Zhang, Yue Xie. Visualization: Yingchun He, Kui Du, Yang Wang. Supervision: Yue Xie. Project administration: Qiguo Zhuang, Yue Xie. Funding acquisition: Yue Xie, Hui Zhang. All authors reviewed and approved the final version of the manuscript.
Availability of Data and Materials: The data supporting the findings of this study are available within the article/Supplementary Materials. Further inquiries can be directed to the corresponding authors.
Ethics Approval: Not applicable.
Conflicts of Interest: The authors declare no conflicts of interest.
Supplementary Materials: The supplementary material is available online at https://www.techscience.com/doi/10.32604/phyton.2026.074974/s1.
References
1. Wang Z , Hu G , Li Z , Zhong C , Yao X . Characterizing tetraploid populations of Actinidia chinensis for kiwifruit genetic improvement. Plants. 2022; 11( 9): 1154. doi:10.3390/plants11091154. [Google Scholar] [CrossRef]
2. Han X , Zhang Y , Zhang Q , Ma N , Liu X , Tao W , et al. Two haplotype-resolved, gap-free genome assemblies for Actinidia latifolia and Actinidia chinensis shed light on the regulatory mechanisms of vitamin C and sucrose metabolism in kiwifruit. Mol Plant. 2023; 16( 2): 452– 70. doi:10.1016/j.molp.2022.12.022. [Google Scholar] [CrossRef]
3. Wang S , Qiu Y , Zhu F . Kiwifruit (Actinidia spp.): A review of chemical diversity and biological activities. Food Chem. 2021; 350: 128469. doi:10.1016/j.foodchem.2020.128469. [Google Scholar] [CrossRef]
4. Luo L , Song W , Hua Q , Li D , Liang H , Zhang X . Research progress on plant sex-determination genes and their epigenetic regulation. Chin Bull Bot. 2024; 59( 02): 278– 90. (In Chinese). doi:10.11983/CBB23088. [Google Scholar] [CrossRef]
5. Mayer SS , Charlesworth D . Cryptic dioecy in flowering plants. Trends Ecol Evol. 1991; 6( 10): 320– 5. doi:10.1016/0169-5347(91)90039-Z. [Google Scholar] [CrossRef]
6. Khukhunaishvili RG , Dzhokhadze DI . Electrophoretic study of the proteins from Actinidia leaves and sex identification. Prikl Biokhim Mikrobiol. 2006; 42( 1): 117– 20. doi:10.1134/S0003683806010170. [Google Scholar] [CrossRef]
7. Zheng H , Li Y , Jin X , Zhao Y , Luo G . The variation of plant morphology and secondary metabolite of male and female Actinidia arguta during growth. Agric Sci J Yanbian Univ. 2018; 40( 04): 16– 21. (In Chinese). doi:10.13478/j.cnki.jasyu.2018.04.003. [Google Scholar] [CrossRef]
8. Biasi R , Falasca G , Speranza A , De Stradis A , Scoccianti V , Franceschetti M , et al. Biochemical and ultrastructural features related to male sterility in the dioecious species Actinidia deliciosa. Plant Physiol Biochem. 2001; 39( 5): 395– 406. doi:10.1016/S0981-9428(01)01255-4. [Google Scholar] [CrossRef]
9. Guo D , Wang R , Fang J , Zhong Y , Qi X . Development of sex-linked markers for gender identification of Actinidia arguta. Sci Rep. 2023; 13( 1): 12780 doi:10.1038/s41598-023-39561-0. [Google Scholar] [CrossRef]
10. Akagi T , Henry IM , Tao R , Comai L . A Y-chromosome–encoded small RNA acts as a sex determinant in persimmons. Science. 2014; 346( 6209): 646– 50. doi:10.1126/science.1257225. [Google Scholar] [CrossRef]
11. Zhang PX , Yang SC , Liu YF , Zhang QL , Xu LQ , Luo ZR . Validation of a male-linked gene locus (OGI) for sex identification in persimmon (Diospyros kaki Thunb.) and its application in F1 progeny. Plant Breed. 2016; 135( 6): 721– 7. doi:10.1111/pbr.12427. [Google Scholar] [CrossRef]
12. Harvey CF , Gill GP , Fraser LG , McNeilage MA . Sex determination in Actinidia. 1. Sex-linked markers and progeny sex ratio in diploid A. chinensis. Sex Plant Reprod. 1997; 10( 3): 149– 54. doi:10.1007/s004970050082. [Google Scholar] [CrossRef]
13. Gill GP , Harvey CF , Gardner RC , Fraser LG . Development of sex-linked PCR markers for gender identification in Actinidia. Theor Appl Genet. 1998; 97( 3): 439– 45. doi:10.1007/s001220050914. [Google Scholar] [CrossRef]
14. Zhang Q , Liu C , Liu Y , VanBuren R , Yao X , Zhong C , et al. High-density interspecific genetic maps of kiwifruit and the identification of sex-specific markers. DNA Res. 2015; 22( 5): 367– 75. doi:10.1093/dnares/dsv019. [Google Scholar] [CrossRef]
15. Zhang K , Jiang X , Wang Y , Tang W , Liu Y , Liang D , et al. Collection, evaluation and application of molecular markers for kiwifruit sex identification. J Sichuan Agric Univ. 2021; 39( 04): 541– 8. (In Chinese). doi:10.16036/j.issn.1000-2650.2021.04.016. [Google Scholar] [CrossRef]
16. Murakami S , Katai H , Yamada S , Taneishi M . The validity of marker-assisted selection using sex linked SCAR markers in kiwifruit (Actinidia chinensis cv ‘rainbow red’) seedlings. J Sci High Technol Agric. 2015; 27( 2): 68– 74. doi:10.2525/shita.27.68. [Google Scholar] [CrossRef]
17. Taravella AM , Sayres MAW . Fruitful analysis of sex chromosomes reveals X-treme genetic diversity. Genome Biol. 2016; 17( 1): 244. doi:10.1186/s13059-016-1115-9. [Google Scholar] [CrossRef]
18. Charlesworth D . Plant sex chromosome evolution. J Exp Bot. 2013; 64( 2): 405– 20. doi:10.1093/jxb/ers322. [Google Scholar] [CrossRef]
19. Zhao Y , Zhang T , Liu C , Huang X , Yuan Z . Progress on sex determinant mechanism in horticultural plants. Acta Hortic Sin. 2018; 45( 11): 2228– 42. (In Chinese). doi:10.16420/j.issn.0513-353x.2018-0487. [Google Scholar] [CrossRef]
20. Ming R , Bendahmane A , Renner SS . Sex chromosomes in land plants. Annu Rev Plant Biol. 2011; 62: 485– 514. doi:10.1146/annurev-arplant-042110-103914. [Google Scholar] [CrossRef]
21. He Z , Huang H , Zhong Y . Cytogenetic study of diploid Actinidia chinensis—karyotype, morphology of sex chromosomes at primary differentiation stage and evolutionary significance. Acta Hortic. 2003; 610: 379– 85. doi:10.17660/ActaHortic.2003.610.50. [Google Scholar] [CrossRef]
22. Charlesworth B , Charlesworth D . A model for the evolution of dioecy and gynodioecy. Am Nat. 1978; 112( 988): 975– 97. doi:10.1086/283342. [Google Scholar] [CrossRef]
23. Charlesworth D . Plant contributions to our understanding of sex chromosome evolution. New Phytol. 2015; 208( 1): 52– 65. doi:10.1111/nph.13497. [Google Scholar] [CrossRef]
24. Peng D , Wu Z . Progress on sex determination of dioecious plants. Biodivers Sci. 2022; 30( 3): 1– 12. doi:10.17520/biods.2021416. [Google Scholar] [CrossRef]
25. Akagi T , Pilkington SM , Varkonyi-Gasic E , Henry IM , Sugano SS , Sonoda M , et al. Two Y-chromosome-encoded genes determine sex in kiwifruit. Nat Plants. 2019; 5( 8): 801– 9. doi:10.1038/s41477-019-0489-6. [Google Scholar] [CrossRef]
26. Kazama Y , Ishii K , Aonuma W , Ikeda T , Kawamoto H , Koizumi A , et al. A new physical mapping approach refines the sex-determining gene positions on the Silene latifolia Y-chromosome. Sci Rep. 2016; 6: 18917. doi:10.1038/srep18917. [Google Scholar] [CrossRef]
27. Aryal R , Ming R . Sex determination in flowering plants: Papaya as a model system. Plant Sci. 2014; 217–218: 56– 62. doi:10.1016/j.plantsci.2013.10.018. [Google Scholar] [CrossRef]
28. Harkess A , Huang K , van der Hulst R , Tissen B , Caplan JL , Koppula A , et al. Sex determination by two Y-linked genes in garden Asparagus. Plant Cell. 2020; 32( 6): 1790– 6. doi:10.1105/tpc.19.00859. [Google Scholar] [CrossRef]
29. Müller NA , Kersten B , Leite Montalvão AP , Mähler N , Bernhardsson C , Bräutigam K , et al. A single gene underlies the dynamic evolution of poplar sex determination. Nat Plants. 2020; 6( 6): 630– 7. doi:10.1038/s41477-020-0672-9. [Google Scholar] [CrossRef]
30. Yang W , Wang D , Li Y , Zhang Z , Tong S , Li M , et al. A general model to explain repeated turnovers of sex determination in the Salicaceae. Mol Biol Evol. 2021; 38( 3): 968– 80. doi:10.1093/molbev/msaa261. [Google Scholar] [CrossRef]
31. Prentout D , Razumova O , Rhoné B , Badouin H , Henri H , Feng C , et al. An efficient RNA-seq-based segregation analysis identifies the sex chromosomes of Cannabis sativa. Genome Res. 2020; 30( 2): 164– 72. doi:10.1101/gr.251207.119. [Google Scholar] [CrossRef]
32. Varkonyi-Gasic E , Wang T , Cooney J , Jeon S , Voogd C , Douglas MJ , et al. Shy Girl, a kiwifruit suppressor of feminization, restricts gynoecium development via regulation of cytokinin metabolism and signalling. New Phytol. 2021; 230( 4): 1461– 75. doi:10.1111/nph.17234. [Google Scholar] [CrossRef]
33. De Mori G , Testolin R , Cipriani G . A molecular protocol for early sex discrimination (ESD) in Actinidia spp. J Berry Res. 2022; 12( 2): 249– 66. doi:10.3233/JBR-211530. [Google Scholar] [CrossRef]
34. Wen C , Wei J , Zhao Y , Zeng T , Li M , Zhang H , et al. Identification and development of molecular markers for sex in kiwifruit. J Sichuan Univ (Nat Sci Ed). 2025; 62( 02): 271– 8. (In Chinese). doi:10.19907/j.0490-6756.250025. [Google Scholar] [CrossRef]
35. Fang L , Zhang H , Chen Z , Wang F , Ji X . Development of specific markers for the characterization of kiwifruit sex trait based on fluorescence capillary electrophoresi. South China Fruits. 2024; 53( 03): 7– 15. (In Chinese). doi:10.13938/j.issn.1007-1431.20240127. [Google Scholar] [CrossRef]
36. Akagi T , Henry IM , Ohtani H , Morimoto T , Beppu K , Kataoka I , et al. A Y-encoded suppressor of feminization arose via lineage-specific duplication of a cytokinin response regulator in kiwifruit. Plant Cell. 2018; 30( 4): 780– 95. doi:10.1105/tpc.17.00787. [Google Scholar] [CrossRef]
37. Qi X , Wang R , Zhang M , Lin M , Li Y , Sun L . Research progress in kiwifruit breeding. J Fruit Sci. 2024; 41( 11): 2160– 72. (In Chinese). doi:10.13925/j.cnki.gsxb.20240601. [Google Scholar] [CrossRef]
38. He Y , Li Z , Liao G , Chen L , Zhong M , Huang C , et al. Variation in fruit quality within wild Actinidia eriantha germplasm. N Z J Crop Hortic Sci. 2020; 48( 3): 153– 63. doi:10.1080/01140671.2020.1761405. [Google Scholar] [CrossRef]
39. Yao X , Wang S , Wang Z , Li D , Jiang Q , Zhang Q , et al. The genome sequencing and comparative analysis of a wild kiwifruit Actinidia eriantha. Mol Hortic. 2022; 2( 1): 13. doi:10.1186/s43897-022-00034-z. [Google Scholar] [CrossRef]
40. Ma X , Han Z , Li T , Yang Y , Su W , Li W . Genetic diversity and population structure analysis of wild kiwifruit (Actinidia spp.) using SRAP markers. Biotechnol Biotechnol Equip. 2024; 38( 1): 2373845. doi:10.1080/13102818.2024.2373845. [Google Scholar] [CrossRef]
41. Lescot M , Déhais P , Thijs G , Marchal K , Moreau Y , Van de Peer Y , et al. PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res. 2002; 30( 1): 325– 7. doi:10.1093/nar/30.1.325. [Google Scholar] [CrossRef]
42. Caseys C . Shy girl gives kiwifruit male flowers. Plant Cell. 2018; 30( 4): 739– 40. doi:10.1105/tpc.18.00278. [Google Scholar] [CrossRef]
43. Yue J , Chen Q , Zhang S , Lin Y , Ren W , Li B , et al. Origin and evolution of the kiwifruit Y chromosome. Plant Biotechnol J. 2024; 22( 2): 287– 9. doi:10.1111/pbi.14213. [Google Scholar] [CrossRef]
44. Akagi T , Varkonyi-Gasic E , Shirasawa K , Catanach A , Henry IM , Mertten D , et al. Recurrent neo-sex chromosome evolution in kiwifruit. Nat Plants. 2023; 9( 3): 393– 402. doi:10.1038/s41477-023-01361-9. [Google Scholar] [CrossRef]
45. Massonnet M , Cochetel N , Minio A , Vondras AM , Lin J , Muyle A , et al. The genetic basis of sex determination in grapes. Nat Commun. 2020; 11: 2902. doi:10.1038/s41467-020-16700-z. [Google Scholar] [CrossRef]
46. Iocco-Corena P , Chaïb J , Torregrosa L , MacKenzie D , Thomas MR , Smith HM . VviPLATZ1 is a major factor that controls female flower morphology determination in grapevine. Nat Commun. 2021; 12( 1): 6995. doi:10.1038/s41467-021-27259-8. [Google Scholar] [CrossRef]
47. Guo R , Zhang YH , Zhang HJ , Landis JB , Zhang X , Wang HC , et al. Molecular phylogeography and species distribution modelling evidence of ‘oceanic’ adaptation for Actinidia eriantha with a refugium along the oceanic-continental gradient in a biodiversity hotspot. BMC Plant Biol. 2022; 22( 1): 89. doi:10.1186/s12870-022-03464-5. [Google Scholar] [CrossRef]
48. Ye LX , Luo MM , Wang Z , Bai FX , Luo X , Gao L , et al. Genome-wide analysis of MADS-box gene family in kiwifruit (Actinidia chinensis var. chinensis) and their potential role in floral sex differentiation. Front Genet. 2022; 13: 1043178. doi:10.3389/fgene.2022.1043178. [Google Scholar] [CrossRef]
49. Dubos C , Stracke R , Grotewold E , Weisshaar B , Martin C , Lepiniec L . MYB transcription factors in Arabidopsis. Trends Plant Sci. 2010; 15( 10): 573– 81. doi:10.1016/j.tplants.2010.06.005. [Google Scholar] [CrossRef]
50. Smale ST , Kadonaga JT . The RNA polymerase II core promoter. Annu Rev Biochem. 2003; 72: 449– 79. doi:10.1146/annurev.biochem.72.121801.161520. [Google Scholar] [CrossRef]
51. Rushton PJ , Reinstädler A , Lipka V , Lippok B , Somssich IE . Synthetic plant promoters containing defined regulatory elements provide novel insights into pathogen- and wound-induced signaling. Plant Cell. 2002; 14( 4): 749– 62. doi:10.1105/tpc.010412. [Google Scholar] [CrossRef]
52. De Mori G , Zaina G , Franco-Orozco B , Testolin R , De Paoli E , Cipriani G . Targeted mutagenesis of the female-suppressor SyGI gene in tetraploid kiwifruit by CRISPR/CAS9. Plants. 2020; 10( 1): 62. doi:10.3390/plants10010062. [Google Scholar] [CrossRef]
53. Wang B , Tang D , Zhou J , Zhong W , Liu Q , Zhang S , et al. The regulatory network of sex differentiation in the bisexual kiwifruit ‘Duie A10’ unveiled by integrated transcriptome and metabolome analysis. Sci Hortic. 2025; 354: 114516. doi:10.1016/j.scienta.2025.114516. [Google Scholar] [CrossRef]
54. Pannell JR . Plant sex determination. Curr Biol. 2017; 27( 5): R191– 7. doi:10.1016/j.cub.2017.01.052. [Google Scholar] [CrossRef]
55. Lu J , Shi Y , Yin X , Liu S , Liu C , Wen D , et al. The genetic mechanism of sex type, a complex quantitative trait, in Ricinus communis L. Ind Crops Prod. 2019; 128: 590– 8. doi:10.1016/j.indcrop.2018.11.023. [Google Scholar] [CrossRef]
Cite This Article
 Copyright © 2026 The Author(s). Published by Tech Science Press.
Copyright © 2026 The Author(s). Published by Tech Science Press.This work is licensed under a Creative Commons Attribution 4.0 International License , which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.


 Submit a Paper
Submit a Paper Propose a Special lssue
Propose a Special lssue View Full Text
View Full Text Download PDF
Download PDF Downloads
Downloads
 Citation Tools
Citation Tools