 Open Access
Open Access
ARTICLE
From Limited Samples to Mechanistic Insights: Exploring Ferroptosis-Related Genes in Hypoplastic Left Heart Syndrome
1 Cardiology Department, The Second Affiliated Hospital of Guangxi Medical University, Nanning, 530007, China
2 School of Basic Medical Sciences, Guangxi Medical University, Nanning, 530022, China
3 Second School of Clinical Medicine, Guangxi Medical University, Nanning, 530007, China
4 Department of Science and Education, The Reproductive Hospital of Guangxi Zhuang Autonomous Region, Nanning, 530022, China
* Corresponding Authors: Chunxia Wang. Email: ; Ge Xu. Email:
# These authors contributed equally to this work
Structural and Congenital Heart Disease 2026, 21(1), 8 https://doi.org/10.32604/schd.2026.072858
Received 05 September 2025; Accepted 05 December 2025; Issue published 31 March 2026
Abstract
Background: Hypoplastic left heart syndrome (HLHS) is a congenital heart disease (CHD), and accumulating evidence has implicated ferroptosis in the pathogenesis of HLHS. Therefore, exploring ferroptosis-related genes (FRGs) in HLHS is of clinical significance. Materials and Methods: Gene Expression Omnibus (GEO) was accessed to obtain analytical data. WGCNA was employed to screen relevant module genes, and the limma package was used to identify differentially expressed genes (DEGs). The rfe function in the R package caret and the glmnet package were utilized to conduct SVM-RFE and LASSO regression analyses, and the intersection of these two analyses was taken as the HLHS-related genes. The reliability of the HLHS-related genes was verified by receiver operating characteristic (ROC) curve. Single-sample GSEA (ssGSEA) was used to calculate the scores of each pathway, and the R package clusterProfiler was employed to perform enrichment analysis on the genes. The NetworkAnalyst was used to predict transcription factors (TFs). Results: WGCNA analysis identified that the module genes were chiefly enriched in inflammation and immunity pathways, especially neutrophil-related module genes. Two HLHS-related genes, Cystathionine β-synthase (CBS) and Heparan-α-glucosaminide N-acetyltransferase (HGSNAT), were obtained. A high area under the curve (AUC) values confirmed the reliability of these two genes as the HLHS-related genes. CHD signaling pathway analysis revealed higher scores in most pathways in control group than HLHS group. Additionally, the score of CBS showed a positive association with hypoxia pathway and VEGF signaling pathway, whereas HGSNAT exhibited a negative association. Ultimately, CBS and HGSNAT shared three TFs, namely JUN, EGR1, and TFAP2A. Conclusion: Collectively, this study identified CBS and HGSNAT as ferroptosis-related candidate biomarkers for HLHS. This study offered new directions for HLHS, contributing to the effective treatment of the disease.Keywords
Supplementary Material
Supplementary Material FileHypoplastic left heart syndrome (HLHS) [1] is characterized by aortic valve and mitral valve atresia or stenosis, left ventricular (LV) hypoplasia, and ascending aortic stenosis [2]. According to relevant reports, incidence of HLHS accounts for 1%–3.8% of all congenital heart malformations [3,4]. Cardiac mortality within the first week after birth can reach as high as 23% [5]. Currently, surgical intervention remains the primary treatment for HLHS, with transformation of the right ventricle (RV) into a systemic circulation pumping chamber as the primary treatment [6]. Nonetheless, the 5-year survival rate following surgery ranges from 50% and 70% [7]. At present, it is imperative to explore the molecular mechanism of HLHS and identify HLHS-related genes [8,9].
Ferroptosis, as an iron-dependent, non-apoptotic form of cell death, is characterized by progressive accumulation of lipid reactive oxygen species (ROS) [10,11,12], which is closely linked to an imbalance in intracellular redox status [13,14]. Ferroptotic cells often display abnormal mitochondrial morphology, including volume condensation or swelling, increased membrane density, reduction or disappearance of cristae, and outer membrane rupture [15]. In recent years, accumulating evidence has revealed the potential physiological significance of ferroptosis in tumor suppression and immunomodulation [16]. Current research findings indicated that iron metabolism disorders and oxidative stress responses are the key mechanisms of myocardial injury related to congenital heart disease (CHD) [17]. Ferroptosis can potentially exacerbate the damage to myocardial cells and cardiac structural cells by influencing lipid peroxidation process and glutathione metabolism pathway [18]. It has been confirmed that ferroptosis could serve as a crucial target for the prevention of cardiomyopathy [19]. Moreover, ferroptosis interacts with several key developmental signaling pathways, including WNT, TGF-β, and NOTCH signaling pathways [20,21]. Ferroptosis-induced lipid peroxidation disrupts mitochondrial structure. As the core of cellular energy metabolism, mitochondrial dysfunction leads to impaired energy production in cardiomyocytes, further exacerbating the pathological progression of heart failure and myocardial infarction [22]. Given these findings, it is particularly necessary to explore the intrinsic relationship between ferroptosis and HLHS.
This study employed computational methods such as differential analysis, weighted gene correlation network analysis (WGCNA), and machine learning to identify HLHS-related ferroptotic genes. Moreover, this study further investigated the inherent relationship between the HLHS-related genes and the scores of CHD-related pathways, and screened transcription factors (TFs) closely linked to these HLHS-related genes. The current findings were expected to contribute to effective detection and treatment of HLHS.
The gene expression profile data GSE23959 was downloaded from the Gene Expression Omnibus (GEO) database based on the GPL5188 (Affymetrix Human Exon 1.0 ST Array) platform. The samples comprised myocardial tissues from the right ventricles of 6 neonates with HLHS, alongside left and right ventricular samples from 10 age-matched controls. Ferroptosis-related genes (FRGs) were obtained from a published literature [23].
2.2 Identification of HLHS-Related Genes by WGCNA
Gene modules related to HLHS in GSE23959 were screened by WGCNA [24,25]. First, the pickSoftThreshold() function was used to calculate the scale-free topology fit index (R2) and mean connectivity under different soft-thresholding powers (β), and the relationship curve between R2 and β was plotted. A scale-free network was established when the R2 reached 0.85 and remained stable, at which point the optimal β soft-thresholding power was selected. Following β weighting, the expression matrix was converted into an adjacency matrix. Average linkage hierarchical clustering was performed based on topological overlap matrix (TOM), with initial modules identified via the dynamic tree cut algorithm. Modules were merged according to the correlation between module eigengenes (MEs), using the parameters: height = 0.35, deepSplit = 2, and minModuleSize = 50. The flowchart of the WGCNA was shown in Fig. S1.
The limma package was employed to conduct differential analysis between HLHS and control samples in GSE23959. Differential analysis was performed based on a linear modeling framework combined with empirical Bayesian methods to improve statistical robustness under small-sample conditions. Resulting p-values were adjusted with the Benjamini–Hochberg method for multiple hypothesis testing to control the false discovery rate (FDR). Genes meeting the criteria of adjusted p-value < 0.05 and |Fold Change| > 1.2 were designated as DEGs. Candidate genes were subsequently obtained by intersecting the DEGs, genes identified by WGCNA, and FRGs [26].
2.4 Identifying HLHS-Related Genes by Machine Learning
The rfe function in the R package caret [27] was utilized. Least Absolute Shrinkage and Selection Operator (LASSO) feature selection for candidate genes was performed using the R package glmnet, with 10-fold cross-validation (nfolds = 10) to determine the optimal value of the penalty parameter λ. The model was configured with family = “binomial” to adapt to the binary classification framework (HLHS vs. control group). During the cross-validation process, we selected the λ value that minimized the mean squared error (lambda.min) as the optimal penalty parameter for the final model.
Recursive feature elimination (RFE) was implemented using the rfe() function in the R package caret, and a support vector machine (SVM) model was developed with the svmLinear method. The model performance was also evaluated via 10-fold cross-validation (method = “cv”, number = 10). The process started with the full gene set, iteratively removing the least important features while retraining and assessing the model via 10-fold cross-validation. The final feature set was determined as the smallest subset yielding the highest classification accuracy. The receiver operating characteristic (ROC) curve was used to validate the reliability of the HLHS-related genes obtained.
2.5 Pathway Correlation and Enrichment Analysis
Single-sample GSEA (ssGSEA) [28] was used to compute the pathway scores for each sample. Differences in pathway scores between the disease group and the control group were subsequently compared. The R package clusterProfiler [29] was used to perform enrichment analysis on the HLHS-related genes [30].
2.6 Analysis of TF Regulatory Network
To elucidate the upstream regulatory mechanisms of cystathionine β-synthase (CBS) and heparan-α-glucosaminide N-acetyltransferase (HGSNAT), we utilized the “TF-miRNA coregulatory interactions” module in the NetworkAnalyst platform (https://www.networkanalyst.ca/, accessed on 01 January 2025) to construct a regulatory network. The regulatory interaction information of this module was derived from the RegNetwork database, which integrates large-scale literature curation and experimentally validated TF/miRNA regulatory relationships.
All statistical data were analyzed using the R language (version 3.6.0). The t-test was employed to calculate differences between two groups of continuous variables. The spearman method was used to calculate the correlations between HLHS-related genes and pathways. And p < 0.05 was considered to indicate a statistically significant difference.
3.1 Screening of HLHS-Related Genes by WGCNA
The scale-free topology fit index reached stability when the soft threshold was set at 16 (Fig. 1A). Based on this soft threshold, hierarchical clustering was performed on the genes (Fig. 1B). A total of 12 modules were obtained, and gene number in each module was presented in Fig. 1C. The heatmap of the associations between the modules and phenotypes revealed significant associations between specific modules and HLHS (Fig. 1D). Among them, the skyblue module exhibited the greatest positive correlation (Fig. 1D,E). Enrichment analysis demonstrated that the genes in the skyblue module were notably enriched in inflammation- and immunity-related biological processes such as neutrophil-mediated immunity, neutrophil activation, neutrophil degranulation, neutrophil activation involved in immune, leukocyte migration (Fig. 1F).
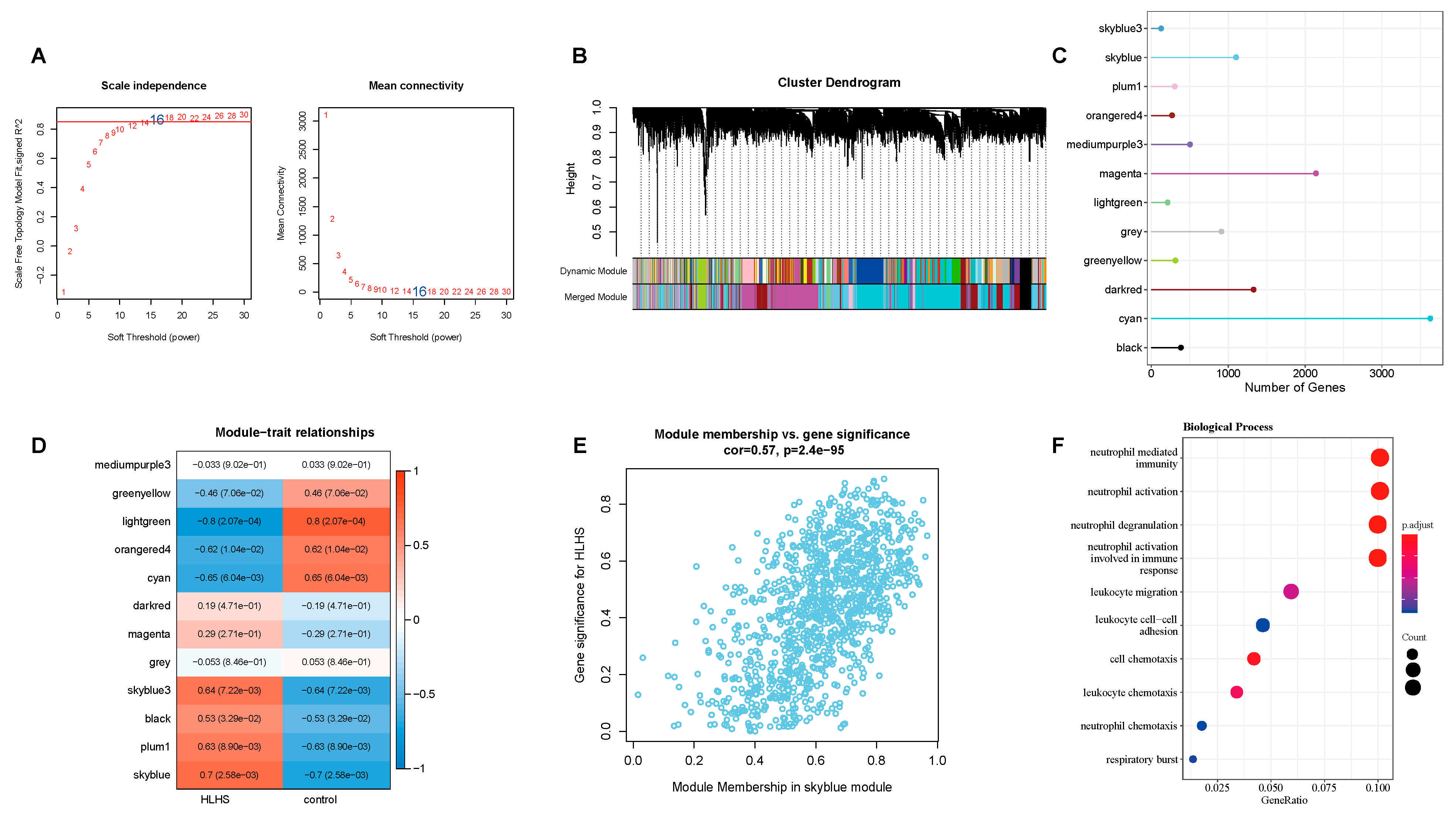
Figure 1: Results of WGCNA analysis on HLHS-related genes. (A) The soft threshold selection was used to determine the scale-free property of the network. (B) The gene clustering dendrogram shows the allocation of different modules. (C) The number of genes in each module. (D) The heatmap of correlations between different gene modules and HLHS as well as the control. (E) Correlation analysis between gene member values and gene significance in the skyblue module, demonstrating a significant correlation between this module and HLHS. (F) The enrichment analysis of biological processes of genes in the skyblue module.
For the GSE23959 dataset, differential gene expression was compared between HLHS samples and control samples. Fig. 2A was the volcano plot of the DEGs, while Fig. 2B was the expression heatmap of the top 50 DEGs. Finally, five candidate genes were obtained by intersecting the DEGs, genes in the skyblue module of WGCNA, and FRGs (Fig. 2C).
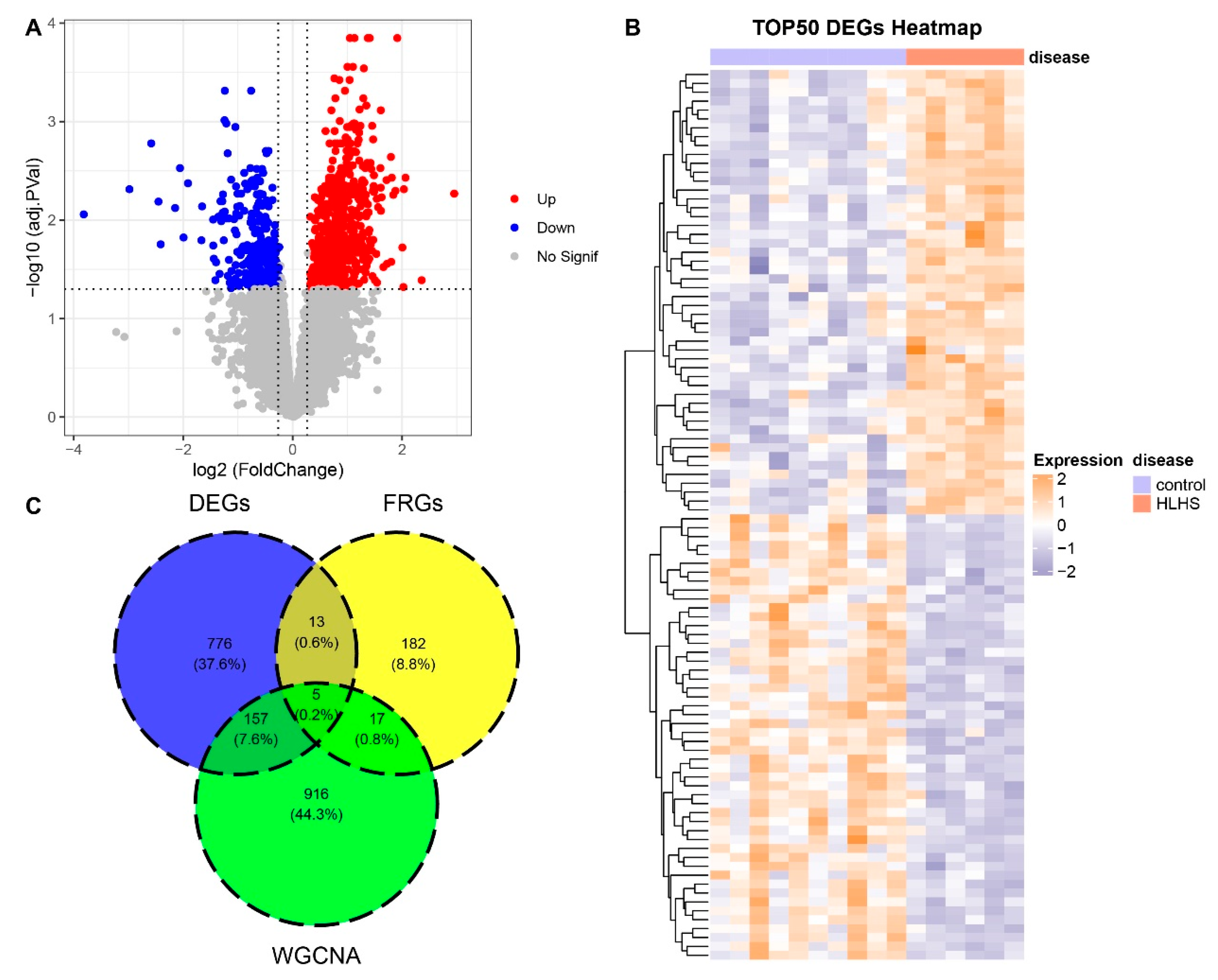
Figure 2: Analysis of DEGs. (A) Volcano plot of the results of differential gene analysis. (B) Expression heatmap of the top 50 DEGs. (C) The intersection of DEGs, skyblue module genes and FRGs in Venn diagram.
3.3 Machine Learning for Screening HLHS-Related Genes
LASSO regression analysis was used to screen the candidate genes. Fig. 3A showed the variation of regression coefficients of different gene features with the penalty parameter (λ) in the LASSO regression. Fig. 3B presented the deviance when determining the optimal λ value through 10-fold CV. The dotted line indicated the selected optimal λ value, which achieved a balance between maintaining model performance and minimizing the number of features. Fig. 3C presented the trend in cross-validation (CV) accuracy of the SVM model as the number of features varies under the RFE method. Through intersecting the results yielded by the two machine learning methods, 2 HLHS-related genes, CBS and HGSNAT, were finally obtained (Fig. 3D).
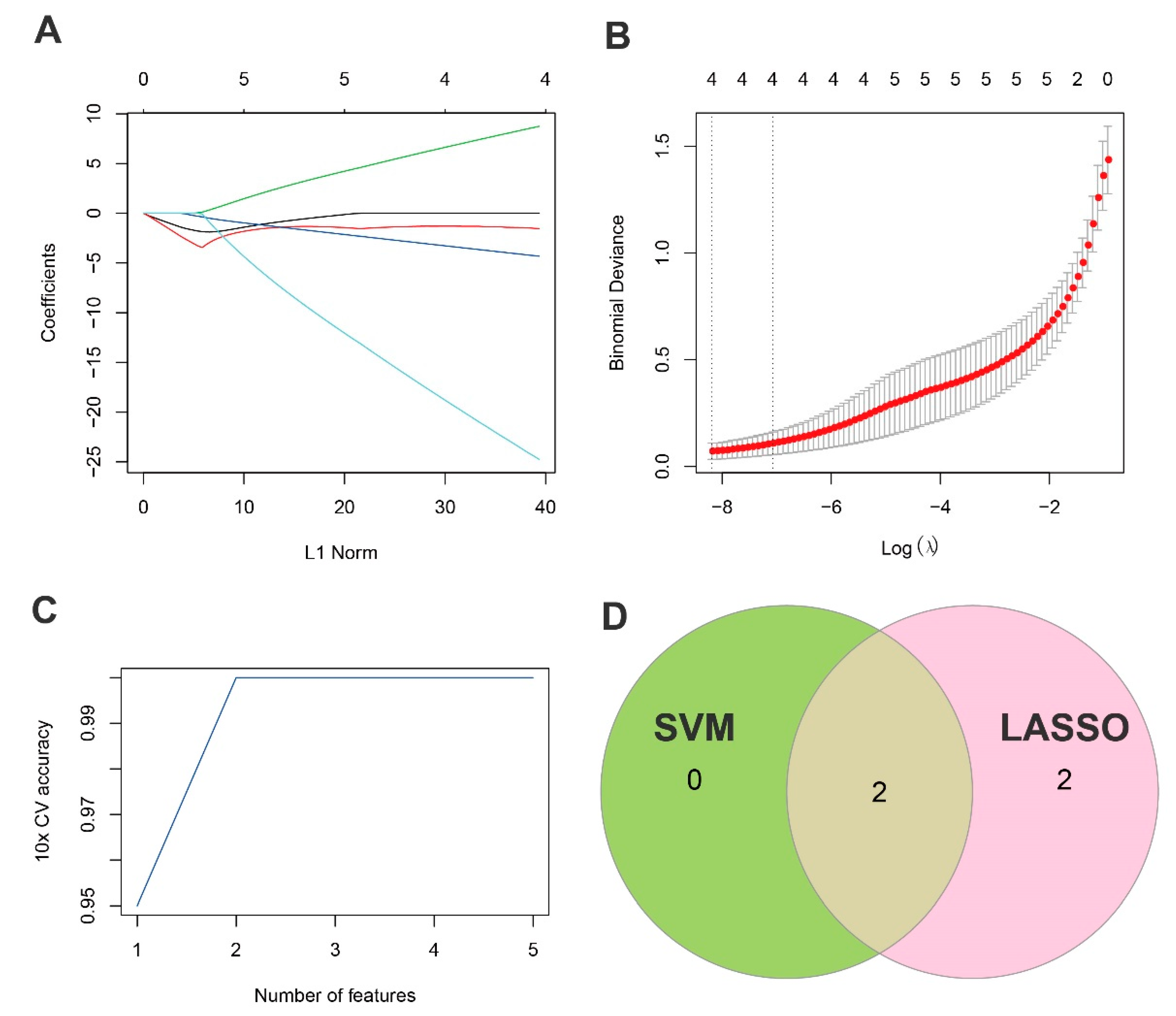
Figure 3: Screening of HLHS-related genes by machine learning. (A,B) The regression coefficient changes of gene features in the LASSO regression model and the optimal penalty parameter (λ) determined by CV. (C) The curve of the CV accuracy of feature screening by the RFE method of the SVM model with the change of the number of features. (D) The Venn diagram of the intersection of HLHS-related genes screened by the SVM-RFE and LASSO methods. By intersecting the results of two machine learning methods, 2 HLHS-related genes were ultimately identified.
3.4 Validation of HLHS-Related Genes
Both CBS and HGSNAT demonstrated strong high diagnostic performance for HLHS, as indicated by their relatively high individual area under the curve (AUC) values (Fig. 4A,B). The AUC value of their combined predictive model reached 1.000, further confirming the excellent diagnostic performance of the two genes as a diagnostic model (Fig. 4C). Comparison of the gene expression profiles of CBS and HGSNAT revealed that CBS was significantly downregulated in the HLHS group than the control group, whereas HGSNAT exhibited a marked upregulation in the HLHS group than the control group (Fig. 4D, p < 0.001).
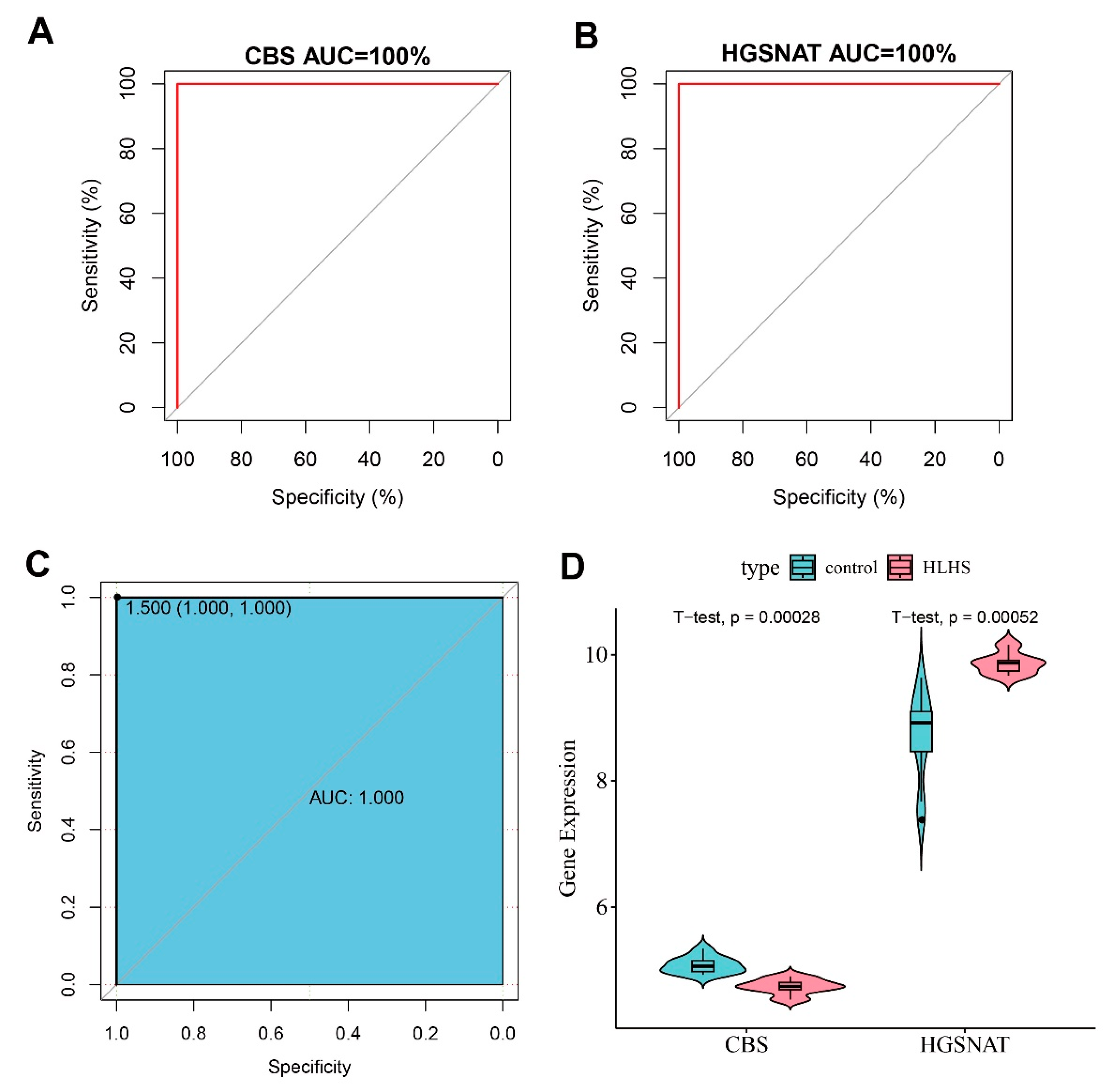
Figure 4: Verification of CBS and HGSNAT as HLHS-related genes. (A) ROC curve of CBS. (B) ROC curve of HGSNAT. (C) ROC curve of the combination of CBS and HGSNAT. (D) Violin plot of the expressions of CBS and HGSNAT in the control group and HLHS samples.
3.5 Pathway Scores Related to CHD and Their Correlations with HLHS-Related Genes
We performed pathway activity scoring across multiple CHD-related signaling pathways in each sample of the two groups. WNT, NOTCH, VEGF, and hypoxia showed notably higher scores in the control group (Fig. 5A, p < 0.05), while Hedgehog pathway exhibited distinct regulatory patterns in HLHS but its activity did not reduce significantly. This may be related to its persistent requirement during the development of specific cardiac structures. Furthermore, analysis on the association between CBS and HGSNAT and the pathway scores demonstrated that CBS was strongly positively linked to the scores of hypoxia pathway and VEGF signaling pathway (Fig. 5B), while the HGSNAT gene exhibited an opposite trend (Fig. 5C).
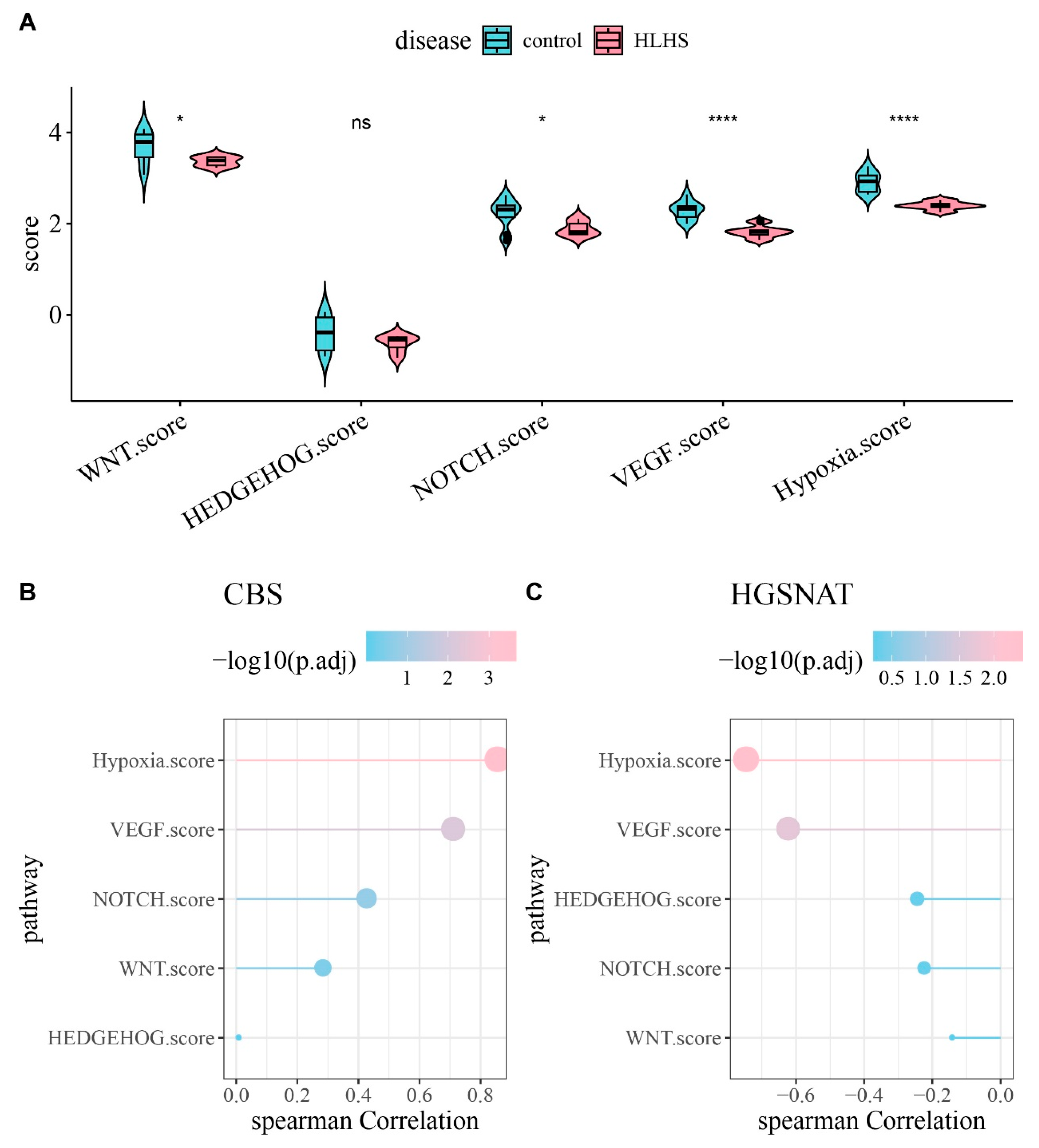
Figure 5: Analysis of pathway score differences between HLHS and control. (A) Differences in pathway scores between HLHS and the control group, with * indicating p < 0.05, **** indicating p < 0.0001 and ns indicating p > 0.05. (B) Correlations of CBS with pathway scores. (C) Correlations of HGSNAT with pathway scores.
3.6 TF-TARGET Regulatory Network
Taking CBS and HGSNAT as the input genes, we constructed a three-layer regulatory network comprising TFs, miRNAs, and target genes. The results of the analysis of the TF-TARGET regulatory network showed that there were a total of 15 TFs for CBS and 11 TFs for HGSNAT in the cardiac muscle tissues (Fig. 6). Moreover, CBS and HGSNAT shared three common TFs, namely JUN, EGR1, and TFAP2A (Fig. 6).
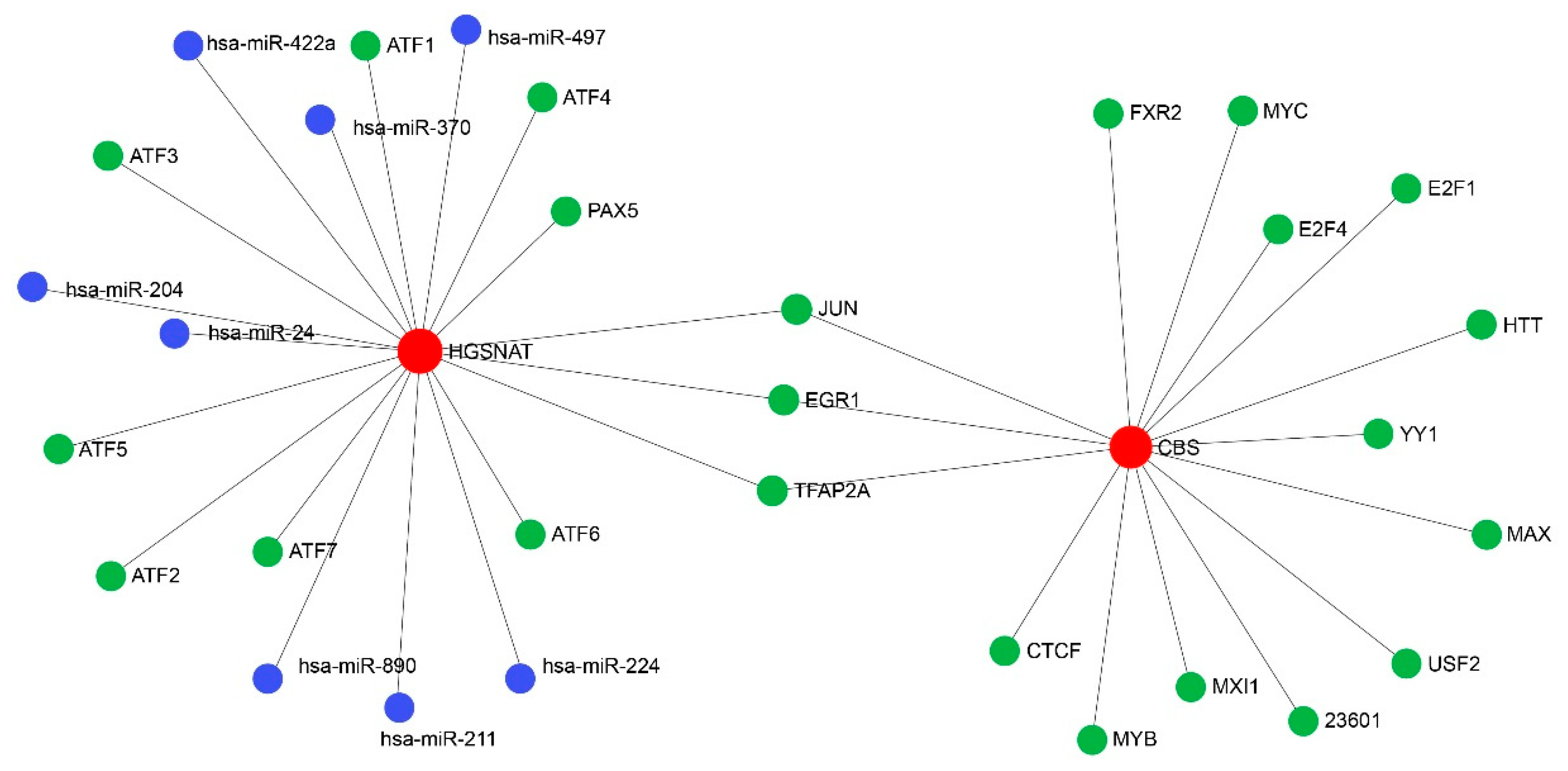
Figure 6: The TF regulatory network diagram, in which red represents HLHS-related genes, green represents TFs, and blue represents miRNAs.
HLHS is characterized by underdevelopment of the left ventricle and associated structures, which are consequently unable to support systemic circulation. Cell death is commonly accompanied by a large amount of iron accumulation and lipid peroxidation [31]. Study reported that ferroptosis is closely associated with the pathogenesis of cardiovascular diseases [32,33]. Previous transcriptomic studies of HLHS addressed limitedly regulated cell death mechanisms. For instance, WGCNA and DEG analyses in ventricular tissues identified hub genes including Fbn1, Itga8, Itga11, Itgb5, and Thbs2, and revealed pathways involved in cardiac development, apoptosis, and BMP signaling during HLHS progression [34]. In addition, recent scRNA-seq analysis on peripheral blood mononuclear cells (PBMCs) from single ventricle (SV)/HLHS patients revealed widespread immune dysregulation, with significant alterations in Th1/17 cells, TFH cells, NK cells, and Th2 cells, alongside altered MYC and IFN-γ activity [35]. However, ferroptosis-related mechanisms remained unexplored. The present research investigated the association between ferroptosis and HLHS, and found that HLHS was mainly enriched in inflammation and immune-correlated biological processes. Additionally, two HLHS-related genes related to ferroptosis, namely CBS and HGSNAT, were successfully identified in the limited samples, and three TFs (JUN, EGR1, TFAP2A) shared by the two HLHS-related genes were also found. However, this discovery provided a hypothesis for future functional studies rather than an established biomarkers.
We discovered two HLHS-related genes, namely HGSNAT and CBS. HGSNAT is the only known lysosomal enzyme that is not a hydrolase, and it catalyzes the N-acetylation of terminal α-D-glucosamine using acetyl coenzyme A [36]. HGSNAT has been previously confirmed as a characteristic gene related to heart diseases [37]. CBS can catalyze the condensation reaction between homocysteine and serine to generate cystathionine [38]. The H2S derived from CBS is associated with the inhibition of mitochondrial function within cells [39], and CBS domain can regulate cell functions in response to energy changes. Ferroptosis is also closely related to cell respiration [40]. The generation of ROS could directly induce ferroptosis in cardiomyocytes by catalyzing oxidation of phospholipids in cell membrane [41]. According to relevant reports, the expression of CBS in the nervous and cardiac systems is significantly high, indicating that this gene may influence cardiac development. These findings suggested that CBS may play a role in the occurrence of HLHS through ferroptosis pathway.
Our results demonstrated that CBS was positively correlated with the hypoxia/VEGF pathway score, suggesting that CBS may regulate ferroptosis in cardiomyocytes and thereby affect cellular respiration. Notably, HGSNAT exhibited a negative correlation with hypoxia/VEGF pathway, contrasting with the positive correlation observed for CBS. Specifically, CBS may be involved in cardiac development by enhancing the activity of the hypoxia/VEGF pathway through promoting angiogenesis and alleviating myocardial hypoxia. CBS has been confirmed to be able to inhibit cell death induced by ferroptosis. In contrast, HGSNAT may prevent abnormal angiogenesis or cardiomyocyte damage by suppressing excessive activation of this pathway. Additionally, mutations in the HGSNAT gene have been shown to impair mitochondrial energy metabolism in mice. This complementary “promotion-inhibition” regulatory pattern may serve as a crucial mechanism for maintaining the homeostasis of the hypoxia/VEGF pathway. In conclusion, HGSNAT and CBS were likely to influence cellular energy by modulating mitochondrial function. Mitochondrial dysfunction leads to cellular energy metabolic disorders, thereby driving the development of cardiac diseases including myocardial hypertrophy and cardiomyopathy [42], which may also be relevant to the occurrence of HLHS. Previous studies also found that hypoxia [43], inflammation, and other pathways can similarly regulate the expression of FRGs [44]. Nonetheless, the roles of these potential candidate genes still require further experimental validation.
By constructing a TF regulatory network, this study discovered three TFs shared by CBS and HGSNAT, namely JUN, TFAP2A, and EGR1. As a proto-oncogene, JUN can accelerate cell proliferation and is essential for cell cycle progression from G1 phase to subsequent phases. It was confirmed that JUN is involved in both embryonic heart development and the occurrence of heart failure. Mechanistically, JUN can bind to AP-1 motifs within promoter regions of target genes and directly regulate transcriptional programs that control oxidative stress responses and lipid metabolism, thereby linking its activity to ferroptosis-associated pathways. TFAP2A has been previously recognized as a key regulatory mediator in embryonic development, especially in the formation of neural crest and epidermis [45]. In terms of cardiac morphogenesis, TFAP2A also functions critically and is closely related to the viability of cardiomyocytes. Studies have shown the essential role of TFAP2A in ferroptosis through specific signal transduction pathways [46,47]. This may be achieved by modulating the expression of antioxidant genes and iron-handling proteins, suggesting that TFAP2A could exert transcriptional control over ferroptosis sensitivity by balancing redox status in cardiac cells. EGR1 encodes a protein belonging to the early growth response (EGR) family, which is mainly involved in tissue damage, immune response, and fibrosis processes [48]. Our enrichment analysis of HLHS-related module genes demonstrated that the related genes were mainly enriched in inflammation and immune-related biological pathways, indicating that the occurrence of HLHS might be closely related to the inflammation and immune pathways correlated with neutrophils. EGR1 is a direct regulator of multiple tumor suppressors. EGR1 in the plasma of patients with coronary heart disease is significantly reduced, and the regulation of EGR1 by miRNAs is related to cardiovascular diseases [49], which affects cardiomyocyte autophagy and leads to cardiomyocyte damage in turn [50]. EGR1 plays a crucial role in tumor cell proliferation, angiogenesis, invasion and immune response [51]. It is also involved in cell proliferation, growth, apoptosis and ferroptosis processes via signal transduction pathways [52]. Study reported that EGR1 is involved in the regulation of ferroptosis in acute myocardial infarction [53]. Importantly, EGR1 binds to GC-rich consensus sequences in promoter regions of stress-responsive genes, thereby directly activating or repressing gene programs associated with iron metabolism and lipid peroxidation. This specific transcriptional regulation provides a mechanistic bridge between EGR1 activity, inflammatory signaling, and ferroptosis execution in cardiomyocytes. In summary, the three TFs, namely JUN, TFAP2A, and EGR1, are all closely associated with embryonic cardiac development. They may participate in the pathogenesis of HLHS by regulating ferroptosis to influence cardiac development or by mediating redox-related reactions. However, this speculation still required further in-depth research and verification.
This study had several limitations. First, the core training set only contained a small number of HLHS and control samples, resulting in insufficient statistical power, unstable results, and susceptibility to outliers. Accordingly, the HLHS-related genes identified merely provided a hypothesis for future functional research rather than established HLHS-related genes. Therefore, we plan to prospectively collect samples according to standardized inclusion and exclusion criteria. Secondly, limited independent cohorts available for HLHS and effect attenuation affected by different tissues might hinder the generalizability of our discoveries. Therefore, our future study will prioritize validation using cohorts from the same tissue, same platform, or calibratable platforms. Third, the entire study was based solely on in silico predictive analysis. The putative functions of CBS, HGSNAT, JUN, EGR1, and TFAP2A proposed were purely hypothetical, which required confirmation through direct experimental validation, such as in vitro or in vivo models of HLHS or cardiac development. Future studies are encouraged to introduce embryonic/neonatal myocardial models to conduct immunohistochemical and functional assays of CBS/HGSNAT and GPX4/ACSL4, followed by performing rescue experiments and genetic manipulations to validate the causal chain between genes, ferroptosis, and phenotype.
Collectively, our integrative analyses identified CBS and HGSNAT as HLHS- and ferroptosis-related candidate genes. These findings provided mechanistic clues that linked ferroptosis-relevant pathways and inflammation/immune processes to HLHS pathobiology. We also predicted upstream TFs (JUN, EGR1, TFAP2A) as potential regulators. Nonetheless, future work incorporating developmentally matched cohorts, protein-level assays, and genetic/pharmacologic perturbations of the CBS/HGSNAT–TF axes are still required before clinical application of the two genes.
Acknowledgement:
Funding Statement: This research was supported by Guangxi Natural Science Foundation under Grant No. 2024JJA141354, Guangxi Medical and Health Appropriate Technology and Promotion Application Project: No. S2024053 and Guangxi Zhuang Autonomous Region Health Commission Self-Funded Scientific Research Project: No. Z-A20230697.
Author Contributions: All authors contributed to this present work: Yiheng Pang, Naixia Chao, Chunxia Wang and Ge Xu designed the research; Hongji Li interpreted the results; Mingxi Xie analyzed the results; Yiheng Pang, Naixia Chao, Hongji Li drafted the manuscript; Mingxi Xie, Chunxia Wang and Ge Xu revised the manuscript and gave the final approval of the version to be published. All authors read and approved the final manuscript.
Availability of Data and Materials: The datasets generated and/or analyzed during the current study are available in the [GSE23959] repository, [https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE23959, accessed on 01 January 2025].
Ethics Approval: Not applicable.
Informed Consent: Not applicable.
Conflicts of Interest: The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Supplementary Materials: The supplementary material is available online at https://www.techscience.com/doi/10.32604/schd.2026.072858/s1.
Abbreviation
| HLHS | hypoplastic left heart syndrome |
| CHD | congenital heart disease |
| LV | left ventricular |
| RV | right ventricle |
| ROS | reactive oxygen species |
| WGCNA | weighted gene co-expression network analysis |
| TF | transcription factor |
| GEO | Gene Expression Omnibus |
| TOM | topological overlap matrix |
| FDR | false discovery rate |
| FRG | ferroptosis-related genes |
| DEG | differentially expressed gene |
| RFE | Recursive Feature Elimination |
| CV | cross-validation |
| SVM | Support Vector Machine |
| LASSO | Least Absolute Shrinkage and Selection Operator |
| ROC | Receiver Operating Characteristic |
| ssGSEA | single sample gene set enrichment analysis |
| CBS | cystathionine β-synthase |
| HGSNAT | heparan-α-glucosaminide N-acetyltransferase |
| AUC | Area Under the Curve |
| miRNA | microRNA |
References
1. Miao Y , Tian L , Martin M , Paige SL , Galdos FX , Li J , et al. Intrinsic endocardial defects contribute to hypoplastic left heart syndrome. Cell Stem Cell. 2020; 27( 4): 574– 89.e8. doi:10.1016/j.stem.2020.07.015. [Google Scholar] [CrossRef]
2. Theis JL , Hu JJ , Sundsbak RS , Evans JM , Bamlet WR , Qureshi MY , et al. Genetic Association between hypoplastic left heart syndrome and cardiomyopathies. Circ Genom Precis Med. 2021; 14( 1): e003126. doi:10.1161/CIRCGEN.120.003126. [Google Scholar] [CrossRef]
3. Kačar P , Tamborrino PP , Iannaccone G , Butera G , Brida M , Prokšelj K , et al. Hypoplastic left heart syndrome (HLHS) becomes of age: Assessing the young adult with HLHS including the neoaorta/aortic arch. Int J Cardiol Congenit Heart Dis. 2025; 19: 100555. doi:10.1016/j.ijcchd.2024.100555. [Google Scholar] [CrossRef]
4. Del Nido PJ . Improving the Outcomes for hypoplastic left heart syndrome: a time for reflection or a call to action? J Am Coll Cardiol. 2025; 85( 24): 2399– 400. doi:10.1016/j.jacc.2025.04.063. [Google Scholar] [CrossRef]
5. Saraf A , Book WM , Nelson TJ , Xu C . Hypoplastic left heart syndrome: from bedside to bench and back. J Mol Cell Cardiol. 2019; 135: 109– 18. doi:10.1016/j.yjmcc.2019.08.005. [Google Scholar] [CrossRef]
6. Alonzo M , Contreras J , Bering J , Zhao MT . In vivo and in vitro approaches to modeling hypoplastic left heart syndrome. Curr Cardiol Rep. 2024; 26( 11): 1221– 9. doi:10.1007/s11886-024-02122-6. [Google Scholar] [CrossRef]
7. Ramonfaur D , Zhang X , Garza AP , García-Pons JF , Britton-Robles SC . Hypoplastic left heart syndrome: a review. Cardiol Rev. 2023; 31( 3): 149– 54. doi:10.1097/CRD.0000000000000435. [Google Scholar] [CrossRef]
8. Datta S , Cao W , Skillman M , Wu M . Hypoplastic left heart syndrome: signaling & molecular perspectives, and the road ahead. Int J Mol Sci. 2023; 24( 20): 15249. doi:10.3390/ijms242015249. [Google Scholar] [CrossRef]
9. Xu X , Huang Y , Han X . Single-nucleus RNA sequencing reveals cardiac macrophage landscape in hypoplastic left heart syndrome. Congenit Heart Dis. 2024; 19( 2): 233– 46. doi:10.32604/chd.2024.050231. [Google Scholar] [CrossRef]
10. Zhang X , Yao L , Yan Y , Fu M . Evolution of natural polymer nerve conduit technology in peripheral nerve repair: a narrative review. Adv Technol Neurosci. 2024; 1( 2): 229– 43. doi:10.4103/ATN.ATN-D-24-00011. [Google Scholar] [CrossRef]
11. Huang L , Wang J , Xu J , Bian M , Wang J , Lu S . Ferroptosis in osteogenic differentiation: a narrative review of bone regeneration metabolism. Regen Med Rep. 2025; 2( 3): 100– 7. doi:10.4103/REGENMED.REGENMED-D-25-00007. [Google Scholar] [CrossRef]
12. Tang H , Zhang Q , Zhang P , Li Y , Guo Y , Gao J , et al. Chaigui Bitong decoction inhibits rheumatoid arthritis fibroblast-like synoviocyte injury through regulating the ferroptosis signaling pathway via Nrf2/HO-1/GPX4 axis. Curr Pharm Anal. 2025; 21( 3): 179– 91. doi:10.1016/j.cpan.2025.03.003. [Google Scholar] [CrossRef]
13. Zefrei F-J , Shormij M , Dastranj L , Alvandi M , Shaghaghi Z , Farzipour S , et al. Ferroptosis inducers as promising radiosensitizer agents in cancer radiotherapy. Curr Radiopharm. 2024; 17( 1): 14– 29. doi:10.2174/0118744710262369231110065230. [Google Scholar] [CrossRef]
14. Li Y , Zhang L , Dong R . Research progress in regulation of ferroptosis by Epigallocatechin-3-gallate in tumor cells. Lett Drug Des Discov. 2023; 20( 12): 1877– 83. doi:10.2174/1570180819666220819150351. [Google Scholar] [CrossRef]
15. Dixon SJ , Olzmann JA . The cell biology of ferroptosis. Nat Rev Mol Cell Biol. 2024; 25( 6): 424– 42. doi:10.1038/s41580-024-00703-5. [Google Scholar] [CrossRef]
16. Fischer C , Volani C , Komlódi T , Seifert M , Demetz E , Valente de Souza L , et al. Dietary iron overload and Hfe−/− related hemochromatosis alter hepatic mitochondrial function. Antioxidants. 2021; 10( 11): 1818. doi:10.3390/antiox10111818. [Google Scholar] [CrossRef]
17. Fan X , Li A , Yan Z , Geng X , Lian L , Lv H , et al. From iron metabolism to ferroptosis: pathologic changes in coronary heart disease. Oxidative Med Cell Longev. 2022; 2022( 1): 6291889. doi:10.1155/2022/6291889. [Google Scholar] [CrossRef]
18. Bajic VP , Van Neste C , Obradovic M , Zafirovic S , Radak D , Bajic VB , et al. Glutathione “redox homeostasis” and its relation to cardiovascular disease. Oxidative Med Cell Longev. 2019; 2019( 1): 5028181. doi:10.1155/2019/5028181. [Google Scholar] [CrossRef]
19. Zhang B , Li S , Liu H , Wang D , Gao A , Wang Y , et al. Immune infiltration in atherosclerosis is mediated by cuproptosis-associated ferroptosis genes. Cardiovasc Innov Appl. 2022; 7( 1): 978. doi:10.15212/CVIA.2023.0003. [Google Scholar] [CrossRef]
20. Zhao J , Ma W , Wang S , Zhang K , Xiong Q , Li Y , et al. Differentiation of intestinal stem cells toward goblet cells under systemic iron overload stress are associated with inhibition of Notch signaling pathway and ferroptosis. Redox Biol. 2024; 72: 103160. doi:10.1016/j.redox.2024.103160. [Google Scholar] [CrossRef]
21. Liu T , Xu P , Ke S , Dong H , Zhan M , Hu Q , et al. Histone methyltransferase SETDB1 inhibits TGF-β-induced epithelial–mesenchymal transition in pulmonary fibrosis by regulating SNAI1 expression and the ferroptosis signaling pathway. Arch Biochem Biophys. 2022; 715: 109087. doi:10.1016/j.abb.2021.109087. [Google Scholar] [CrossRef]
22. Zhang Y , Xin L , Xiang M , Shang C , Wang Y , Wang Y , et al. The molecular mechanisms of ferroptosis and its role in cardiovascular disease. Biomed Pharmacother. 2022; 145: 112423. doi:10.1016/j.biopha.2021.112423. [Google Scholar] [CrossRef]
23. Zou Y , Xie J , Zheng S , Liu W , Tang Y , Tian W , et al. Leveraging diverse cell-death patterns to predict the prognosis and drug sensitivity of triple-negative breast cancer patients after surgery. Int J Surg. 2022; 107: 106936. doi:10.1016/j.ijsu.2022.106936. [Google Scholar] [CrossRef]
24. Yan S , Han Z , Wang T , Wang A , Liu F , Yu S , et al. Exploring the immune-related molecular mechanisms underlying the comorbidity of temporal lobe epilepsy and major depressive disorder through integrated data set analysis. Curr Mol Pharmacol. 2025; 17: E18761429380394. doi:10.2174/0118761429380394250217093030. [Google Scholar] [CrossRef]
25. Rajalingam A , Sekar K , Ganjiwale A . Identification of potential genes and critical pathways in postoperative recurrence of crohn’s disease by machine learning and WGCNA network analysis. Curr Genom. 2023; 24( 2): 84– 99. doi:10.2174/1389202924666230601122334. [Google Scholar] [CrossRef]
26. Zhu X , Zhang Y , Pan P , Liu X , Zhang J , Du X , et al. Identification and validation of amino acid metabolism-related biomarkers and investigation of their potential mechanisms in lung adenocarcinoma. Curr Gene Ther. 2025; 25: 1– 19. doi:10.2174/0115665232364091250203090710. [Google Scholar] [CrossRef]
27. Kuhn M . Building predictive models in R using the caret package. J Stat Softw. 2008; 28( 5): 1– 26. doi:10.18637/jss.v028.i05. [Google Scholar] [CrossRef]
28. Hänzelmann S , Castelo R , Guinney J . GSVA: gene set variation analysis for microarray and RNA-Seq data. BMC Bioinform. 2013; 14( 1): 7. doi:10.1186/1471-2105-14-7. [Google Scholar] [CrossRef]
29. Yu G , Wang L-G , Han Y , He Q-Y . clusterProfiler: an R package for comparing biological themes among gene clusters. Omics J Integr Biol. 2012; 16( 5): 284– 7. doi:10.1089/omi.2011.0118. [Google Scholar] [CrossRef]
30. Peng Z , Fang C , Tong Z , Rao Q , Ren Z , Hu K . Crosstalk between cancer-associated fibroblasts and myeloid cells shapes the heterogeneous microenvironment of gastric cancer. Curr Genom. 2024; 25( 5): 390– 411. doi:10.2174/0113892029300608240531111743. [Google Scholar] [CrossRef]
31. Li J , Cao F , Yin H-L , Huang Z-J , Lin Z-T , Mao N , et al. Ferroptosis: past, present and future. Cell Death Dis. 2020; 11( 2): 88. doi:10.1038/s41419-020-2298-2. [Google Scholar] [CrossRef]
32. Song Y , Wang B , Zhu X , Hu J , Sun J , Xuan J , et al. Human umbilical cord blood–derived MSCs exosome attenuate myocardial injury by inhibiting ferroptosis in acute myocardial infarction mice. Cell Biol Toxicol. 2021; 37( 1): 51– 64. doi:10.1007/s10565-020-09530-8. [Google Scholar] [CrossRef]
33. Lu X , Hu W , Wang X , Wang Z , Yang P , Wang W . Protective role of methane in traumatic nervous system diseases. Med Gas Res. 2024; 14( 3): 159– 62. doi:10.4103/mgr.mgr_23_23. [Google Scholar] [CrossRef]
34. Liu X , Shang H , Li B , Zhao L , Hua Y , Wu K , et al. Exploration and validation of hub genes and pathways in the progression of hypoplastic left heart syndrome via weighted gene co-expression network analysis. BMC Cardiovasc Disord. 2021; 21( 1): 300. doi:10.1186/s12872-021-02108-0. [Google Scholar] [CrossRef]
35. Qu HQ , Ostberg K , Slater DJ , Wang F , Snyder J , Hou C , et al. Single-cell RNA sequencing of peripheral blood mononuclear cells in patients with single ventricle/hypoplastic left heart syndrome. J Gene Med. 2025; 27( 8): e70030. doi:10.1002/jgm.70030. [Google Scholar] [CrossRef]
36. Huizing M , Gahl WA . Inherited disorders of lysosomal membrane transporters. Biochim Biophys Acta Biomembr. 2020; 1862( 12): 183336. doi:10.1016/j.bbamem.2020.183336. [Google Scholar] [CrossRef]
37. Volodko O , Volinsky N , Yarkoni M , Margalit N , Kusniec F , Sudarsky D , et al. Characterization of systemic and culprit-coronary artery miR-483-5p expression in chronic CAD and acute myocardial infarction male patients. Int J Mol Sci. 2023; 24( 10): 8551. doi:10.3390/ijms24108551. [Google Scholar] [CrossRef]
38. Liu N , Lin X , Huang C . Activation of the reverse transsulfuration pathway through NRF2/CBS confers erastin-induced ferroptosis resistance. Br J Cancer. 2020; 122( 2): 279– 92. doi:10.1038/s41416-019-0660-x. [Google Scholar] [CrossRef]
39. Wen X , Ye Y , Yu Z , Shen H , Cui G , Chen G . The role of nitric oxide and hydrogen sulfide in spinal cord injury: an updated review. Med Gas Res. 2024; 14( 3): 96– 101. doi:10.4103/2045-9912.385946. [Google Scholar] [CrossRef]
40. Li C , Zhang Y , Liu J , Kang R , Klionsky DJ , Tang D . Mitochondrial DNA stress triggers autophagy-dependent ferroptotic death. Autophagy. 2021; 17( 4): 948– 60. doi:10.1080/15548627.2020.1739447. [Google Scholar] [CrossRef]
41. Fang X , Ardehali H , Min J , Wang F . The molecular and metabolic landscape of iron and ferroptosis in cardiovascular disease. Nat Rev Cardiol. 2023; 20( 1): 7– 23. doi:10.1038/s41569-022-00735-4. [Google Scholar] [CrossRef]
42. Fatima S , Zhou H , Chen Y , Liu Q . Role of ferroptosis in the pathogenesis of heart disease. Front Physiol. 2024; 15: 1450656. doi:10.3389/fphys.2024.1450656. [Google Scholar] [CrossRef]
43. Feng L , Yin X , Hua Q , Ren T , Ke J . Advancements in understanding the role of ferroptosis in hypoxia-associated brain injury: a narrative review. Transl Pediatr. 2024; 13( 6): 963– 75. doi:10.21037/tp-24-47. [Google Scholar] [CrossRef]
44. Li R , Wei R , Liu C , Zhang K , He S , Liu Z , et al. Heme oxygenase 1-mediated ferroptosis in Kupffer cells initiates liver injury during heat stroke. Acta Pharm Sin B. 2024; 14( 9): 3983– 4000. doi:10.1016/j.apsb.2024.05.007. [Google Scholar] [CrossRef]
45. Xiong Y , Feng Y , Zhao J , Lei J , Qiao T , Zhou Y , et al. TFAP2A potentiates lung adenocarcinoma metastasis by a novel miR-16 family/TFAP2A/PSG9/TGF-β signaling pathway. Cell Death Dis. 2021; 12( 4): 352. doi:10.1038/s41419-021-03606-x. [Google Scholar] [CrossRef]
46. Huang H-X , Yang G , Yang Y , Yan J , Tang X-Y , Pan Q . TFAP2A is a novel regulator that modulates ferroptosis in gallbladder carcinoma cells via the Nrf2 signalling axis. Eur Rev Med Pharmacol Sci. 2020; 24( 9): 4745– 55. [Google Scholar]
47. Jin C , Luo Y , Liang Z , Li X , Kołat D , Zhao L , et al. Crucial role of the transcription factors family activator protein 2 in cancer: current clue and views. J Transl Med. 2023; 21( 1): 371. doi:10.1186/s12967-023-04189-1. [Google Scholar] [CrossRef]
48. Saha SK , Islam SMR , Saha T , Nishat A , Biswas PK , Gil M , et al. Prognostic role of EGR1 in breast cancer: a systematic review. BMB Rep. 2021; 54( 10): 497– 504. doi:10.5483/BMBRep.2021.54.10.087. [Google Scholar] [CrossRef]
49. Kristensen LS , Andersen MS , Stagsted LVW , Ebbesen KK , Hansen TB , Kjems J . The biogenesis, biology and characterization of circular RNAs. Nat Rev Genet. 2019; 20( 11): 675– 91. doi:10.1038/s41576-019-0158-7. [Google Scholar] [CrossRef]
50. Xie Y , Li Y , Chen J , Ding H , Zhang X . Early growth response-1: Key mediators of cell death and novel targets for cardiovascular disease therapy. Front Cardiovasc Med. 2023; 10: 1162662. doi:10.3389/fcvm.2023.1162662. [Google Scholar] [CrossRef]
51. Chen J , Zhan Y , Xu J , Wang Y , Gao Q . EGR1 Overexpression inhibits the occurrence of preeclampsia by binding to MicroRNA-574 promoter and upregulating GAB1. Reprod Sci. 2021; 28( 4): 1112– 21. doi:10.1007/s43032-020-00392-3. [Google Scholar] [CrossRef]
52. Lin Z , Liu Z , Pan Z , Zhang Y , Yang X , Feng Y , et al. EGR1 promotes erastin-induced ferroptosis through activating Nrf2-HMOX1 signaling pathway in breast cancer cells. J Cancer. 2024; 15( 14): 4577– 90. doi:10.7150/jca.95328. [Google Scholar] [CrossRef]
53. Fan K , Huang W , Qi H , Song C , He C , Liu Y , et al. The Egr-1/miR-15a-5p/GPX4 axis regulates ferroptosis in acute myocardial infarction. Eur J Pharmacol. 2021; 909: 174403. doi:10.1016/j.ejphar.2021.174403. [Google Scholar] [CrossRef]
Cite This Article
 Copyright © 2026 The Author(s). Published by Tech Science Press.
Copyright © 2026 The Author(s). Published by Tech Science Press.This work is licensed under a Creative Commons Attribution 4.0 International License , which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.


 Submit a Paper
Submit a Paper Propose a Special lssue
Propose a Special lssue View Full Text
View Full Text Download PDF
Download PDF Downloads
Downloads
 Citation Tools
Citation Tools