 Open Access
Open Access
REVIEW
Artificial intelligence in urological malignancy diagnosis and prognosis: current status and future prospects
1 Department of Urology, Hangzhou TCM Hospital of Zhejiang Chinese Medical University (Hangzhou Hospital of Traditional Chinese Medicine), Hangzhou, China
2 Department of Urology, The Second Xiangya Hospital of Central South University, Changsha, China
3 Department of Urology, The First Affiliated Hospital of Guangzhou Medical University, Guangzhou, China
4 Department of Oncology, Jiangsu Cancer Hospital, Nanjing, China
5 Department of Urology, Shenshan Medical Center, Sun Yat-sen Memorial Hospital, Sun Yat-sen University, Shanwei, China
6 Department of Urology, Shanghai Tenth People’s Hospital, School of Medicine, Tongji University, Shanghai, China
7 Department of Pharmacy, Shanghai Pudong Hospital, Fudan University Pudong Medical Center, Shanghai, China
8 Molecular and Experimental Surgery, Clinic for Visceral, General, Vascular and Transplantation Surgery, University Medicine Magdeburg, Otto-von Guericke University Magdeburg, Magdeburg, Germany
9 Proteomics and Cancer Cell Signaling Group, German Cancer Research Center, Heidelberg, Germany
10 Department of Radiation Oncology, Lueneburg Hospital, Lueneburg, Germany
11 Department of Urology, Suzhou Municipal Hospital, Suzhou, China
12 Department of Oncology, The First Affiliated Hospital of Nanjing Medical University, Nanjing, China
13 Department of Urology, Hangzhou Integrative Medicine Hospital Affiliated to Zhejiang Chinese Medical University (Hangzhou Red Cross Hospital), Hangzhou, China
* Corresponding Authors: Weizhuo Wang. Email: ; Run Shi. Email:
; Jingyu Zhu. Email:
# These authors contributed equally to this work
Canadian Journal of Urology 2026, 33(1), 35-49. https://doi.org/10.32604/cju.2026.076084
Received 13 November 2025; Accepted 13 January 2026; Issue published 28 February 2026
Abstract
Artificial intelligence (AI) is transforming the diagnostic landscape of malignant tumors in the urinary system, including prostate cancer, bladder cancer, and renal cell carcinoma (RCC). By integrating imaging, pathology, and molecular data, AI enhances the precision and reproducibility of tumor detection, grading, and risk stratification. In prostate cancer, AI-assisted multiparametric Magnetic resonance imaging (MRI) and digital pathology systems improve lesion localization and Gleason scoring. For bladder cancer, deep learning-based cystoscopy and radiomics models from Computed tomography/magnetic resonance imaging (CT/MRI) enable real-time lesion segmentation and non-invasive biomarker prediction, such as Programmed Cell Death-Ligand 1 (PD-L1) expression. In RCC, AI, combined with CT/MRI and multi-omics data, aids in subtype classification and prognostic prediction, supporting personalized therapy. However, despite these promising advances, challenges such as data standardization, model generalizability, interpretability, and regulatory compliance hinder AI’s clinical translation. This review outlines the current state of AI in urological cancer diagnosis and prognosis, its technological innovations, and the clinical challenges and opportunities that lie ahead.Keywords
Urologic malignancies—principally prostate cancer, bladder cancer, and renal cell carcinoma (RCC)—remain a significant public health challenge, accounting for a substantial proportion of cancer incidence and mortality worldwide.1 Prostate cancer is the most common cancer in men, accounting for roughly one quarter of all male cancers.1 Bladder cancer ranks ninth in global cancer incidence, with ~550,000 new cases and ~200,000 deaths annually.2 As the primary malignant tumor of the kidney, RCC causes about 140,000 new cases and 60,000 deaths every year.3 While these diseases seriously threaten the life of patients, they also have a profound impact on their quality of life. There are still many unmet clinical needs for early diagnosis, precise classification, and individualized treatment of these diseases.
Imaging, endoscopy, and histopathological evaluation together constitute the cornerstone of the diagnosis of urologic cancers.4–6 These traditional methods have their own limitations: imaging examinations have limited sensitivity for detecting early or microscopic lesions; cystoscopy and tissue biopsy are invasive procedures that not only cause discomfort to patients but also carry risks such as bleeding and infection. Although pathological diagnosis is regarded as the “gold standard”, its results still depend on pathologists’ subjective judgment, and inter- and intra-observer variability can lead to inconsistent diagnoses or delayed treatment.5–8 In recent years, the rapid development of artificial intelligence (AI) technology has provided a new opportunity to solve the above problems. In particular, machine learning and deep learning algorithms are increasingly used in the diagnosis of urologic cancers, with great potential to improve diagnostic accuracy, reduce invasive procedures, and enable early detection.9,10 AI can analyse complex data such as medical imaging, digital pathology, and genomic profiles, capture subtle features that are difficult for the human eye to detect, and provide more objective and accurate decision support for clinicians. By improving diagnostic efficiency, supporting clinical decision-making, and reducing human error, AI is gradually driving innovation in the diagnosis and treatment of urologic cancers.11–13 This review provides new insight into the application of multi-modal AI for the diagnosis and prognosis of urologic cancers, including prostate cancer, bladder cancer, and RCC. It systematically reviews emerging applications and research developments in AI, with particular emphasis on integrating multi-modal data, including medical imaging, clinical data, genomic data, and pathology. In contrast to previous surveys that mainly focus on specific AI techniques or single-modal data, this article investigates the potential of a more holistic, multi-modal approach to AI-powered diagnostic tools. These advanced systems enable real-time processing of high-dimensional, multi-source data to build more patient-specific and widely applicable diagnostic models, significantly promoting clinical standardization, reducing misdiagnosis, and improving patient prognosis. In addition, this review discusses the revolutionary implications of AI applications in clinical settings, the limited prospects in the current literature, and issues of data standardization, model generalizability, and regulatory compliance. In doing so, it hopes to facilitate the development of increasingly potent clinical applications of AI for the diagnosis of urological cancers.
Applications of AI in the Diagnosis of Urologic Cancers
AI has demonstrated significant application value and broad prospects in multi-modal imaging, including multi-parameter MRI (mpMRI), ultrasound, and PET-CT for prostate cancer.14,15 Of these, deep-learning approaches have been most comprehensively combined with mpMRI to support early lesion detection, risk assessment using the Prostate Imaging-Reporting and Data System, and clinical staging, facilitating more precise decisions.14,16,17 Applications in ultrasound and the rapidly developing field of PSMA PET imaging are also accelerating. The clinical trial has also shown that AI models can improve tumor detection and significantly reduce the risk of false negatives, which is particularly useful for early screening and diagnosis of prostate cancer.18–20 In addition, deep learning models have become increasingly adept at detecting subtle image features invisible to the human eye. These developments have demonstrated excellent potential for predicting Gleason scores and tumor aggressiveness, respectively, providing tumor-critical information unavailable to traditional methods and thereby increasing the accuracy of patient prognosis.21,22
The convergence of digital pathology and AI marks a new era in diagnostic practice and grading. Among the most representative recent advances, the operationalization of the Gleason system on whole-slide images using AI is achieved through algorithms that automatically delineate cancerous areas while simultaneously providing Gleason patterns independent of human operators, thereby reducing inter- and intra-observer variability.12,23,24 In terms of histological classification and outcome prediction, the AI model can identify subtle tumor structural changes by analysing histological slice images and provide accurate prognostic information for the patient.25,26 AI can also help pathologists make more accurate diagnoses in complex histological classifications. The application of AI in the early stage of prostate cancer improves the accuracy of early detection and diagnosis of cancer.
Using AI models to combine imaging, clinical, and molecular data for diagnosis and prognosis evaluation of prostate cancer has become a research hotspot in prostate cancer precision medicine. Multi-modal data integration can combine different types of data (such as imaging, genomic, and clinical) to build a more accurate predictive model.27,28 The application of AI in individualized medical care and decision-making support has also gradually advanced, helping doctors formulate personalized treatment plans tailored to each patient’s specific situation.28,29 There is an increasing embedding of AI into individualized care pathways, allowing clinicians to tailor therapy to patient-specific risk and biology. Joint modeling of imaging and genomics may predict recurrence risk and inform adjuvant or surveillance strategies.30,31 Similarly, several AI systems are under development for decision support in treatment: these synthesize multi-omics data to surface evidence-based options and optimize therapeutic outcomes.32–34
The rapid development of cross-sectional imaging, endoscopy, and digital pathology has catalyzed the application of AI in bladder cancer diagnosis. Deep-learning models of CT, MRI, ultrasound, and cystoscopic video are now contributing to tumor detection and staging; multiple studies in muscle-invasive disease report generally strong performance—AUC typically ~0.85–0.92.35
In mpMRI, a CRDL model integrating radiomic, deep-learning, and clinical features accurately predicted 5-year recurrence risk in NMIBC, achieving an AUC of 0.909 in an independent test cohort and underscoring the value of quantitative imaging for prognosis.36 In the key endoscopic technology of cystoscopy, the application of AI is moving towards real-time and precision. The BTS-Net, a Transformer-based model, has been developed to segment early bladder cancer and its satellite foci in real time, significantly improving the detection rate of tiny lesions.37,38 These developments together show that AI can not only enhance the reading efficiency of traditional images but also directly embed the endoscope operation process, thereby improving real-time diagnostic ability.
In pathology, the integration of AI into whole-slide digital workflows is changing the face of diagnostic classification and grading. In a multi-institution study, deep-learning models were trained and validated on >12,500 bladder WSI. They achieved high-performance differentiation among normal urothelium, non-invasive urothelial tumors, and invasive bladder cancer, with a best-model AUC of 0.983.39 These tools provide strong decision support and enhance objectivity and consistency among readers. Additional work further underlines how AI might help overcome subjectivity and observer-to-observer variability typical of conventional assessment by providing novel, reproducible biomarkers for prognostication.40
Multi-modal modeling that integrates imaging, clinical data, and molecular/pathologic features, such as PD-L1 expression and genomic signatures, is emerging as a key strategy to refine diagnosis and risk stratification. In a dual-center study, an interpretable deep-learning model trained on triphasic contrast-enhanced computed tomography scans predicted PD-L1 status non-invasively and demonstrated intense discrimination in external validation (AUC 0.857), illustrating the feasibility of imaging-based biomarker prediction for immunotherapy planning.41 To our knowledge, truly integrated multi-modal approaches for the comprehensive analysis of bladder cancer remain relatively underdeveloped, and many efforts are still at an early exploratory stage.42
AI technology has made remarkable progress in diagnosing RCC. AI models based on CT, MRI, and ultrasound images have been widely used for the detection, classification, and grading of kidney cancer. In CT image analysis, AI models, especially convolutional neural networks (CNNs), showed high accuracy in the automatic detection and subtype differentiation of kidney tumors.43 A study used a CNN to automatically detect RCC, achieving about 85% accuracy in benign and malignant identification on CT images, and RCC subtype classification accuracy of 0.68-0.77, demonstrating feasibility for clinical applications.44 On MRI, AI exploits multiparametric sequences, including contrast-enhanced imaging and diffusion-weighted imaging, to assist in staging and benign–malignant differentiation.45 The coupling of AI with radiomics allows for automated segmentation and extraction of deep quantitative features, a particular advantage in cases demonstrating indistinct tumor outlines or atypical signal characteristics.46,47 In ultrasound, AI-assisted models enhance the recognition of solid renal masses, with significantly improved sensitivity and specificity.48 Imaging based on deep learning is increasingly important for predicting the aggressiveness and metastatic risk of kidney cancer.49
In terms of pathology, AI has become an essential tool for analyzing RCC tissue. Research confirms that AI can accurately identify the main subtypes of RCC by analysing digital tissue slice images and can further predict patient progression and prognosis.50 The combination of digital pathology and AI has shown high efficiency in realising automatic classification of kidney cancer and reducing differences in human diagnosis, providing a more objective and reproducible basis for pathological diagnosis.51 Not limited to pathology, the application of AI has extended to the molecular level. By integrating genomic data, AI models can deeply explore the complex relationships between gene states, such as the Von Hippel-Lindau (VHL) gene, and treatment responses. Some studies use AI to predict kidney cancer patients’ responses to immune checkpoint inhibitor treatment, providing a new strategy for the formulation of individualized treatment plans.52 These applications jointly promote, more accurate diagnosis and treatment of kidney cancer.
Combining multi-modal AI models of imaging, clinical, and molecular markers is a key direction for improving the accuracy of RCC diagnosis and treatment. The multi-modal AI model builds a more powerful early diagnosis and risk prediction system for RCC by integrating CT/MRI image features, clinical indicators, and molecular biomarkers (such as genetic mutations and PD-L1 expression).53 Such fusion models build much more powerful systems for early detection and risk prediction compared with single-source approaches.54–56 There is also growing evidence that they provide superior prognostic information, especially for predicting recurrence and estimating the efficacy of targeted and immunotherapies.57 On this foundation, AI is increasingly integrated within clinical decision-support tools that synthesize patient-specific data to guide therapy selection and optimize outcomes.58 As shown in Figure 1, the multi-modal AI model has significantly improved the diagnostic ability of urologic cancers by integrating medical images, pathological data, and genomic information, and has shown excellent advantages in early detection and risk stratification.
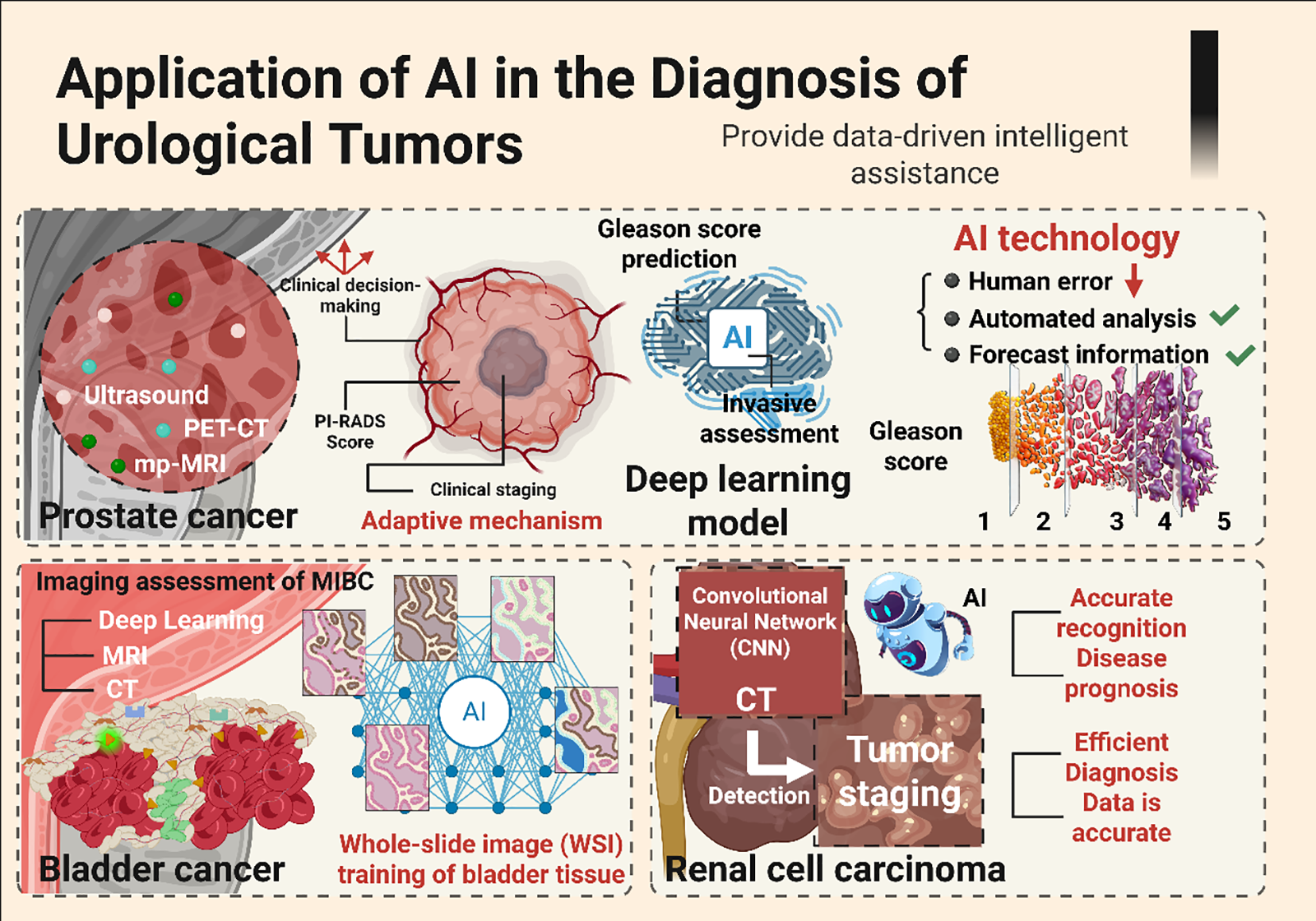
FIGURE 1. The multi-modal AI model integrates medical imaging, pathology, and genomic data to improve tumor diagnosis accuracy and support individualized treatment in the urinary system. AI: Artificial intelligence; PET-CT: Positron emission tomography-computed tomography; MRI: Magnetic resonance imaging; MIBC: Muscle-invasive bladder cancer
Multi-Modal Evidence in AI Diagnosis
Definition and type of multi-modal data
Multi-modal data usually refers to information collected from multiple sources and structures to address specific clinical problems. In the field of urologic cancers, this mainly includes medical imaging (such as CT, MRI, ultrasound, and PET/PSMA-PET), digital pathology (such as full-slice images and their characteristics), structured clinical variables (such as age, PSA level, TNM staging, and medication history), and genomic, transcriptomic, proteomic, and metabolomic data.59 Compared with single-modal learning, multi-modal learning integrates information across different strategies, such as data, feature, and decision-making layers, yielding more robust data characterisation and stronger model generalization in complex clinical tasks.60–62 In recent years, several systematic reviews have summarised multi-modal fusion strategies, including early/mid-term/late fusion, attention mechanisms, Transformers, and graph neural networks, and their medical applications.63–66 In urinary system oncology, these integrated strategies provide a feasible technical path for the unified interpretation of imaging, pathology, and molecular evidence, and vigorously promote the implementation of “integrated diagnosis” in clinical practice.64
The core advantages of multi-modal AI are mainly reflected in two key aspects: first, the integration of multi-source information from imaging, pathology, clinical, and molecular data can significantly improve the tumor detection rate, grading accuracy, and clinical decision-making efficiency; Second, through cross-modal knowledge transfer and uncertainty modelling, multi-modal AI can enhance model generalizability, decision stability, and interpretability, thereby providing a stronger foundation for truly individualized treatment.27,62,64
Multi-modal AI advantages in prostate cancer
In the diagnosis of prostate cancer, multi-modal AI shows significant value by integrating imaging characteristics and clinical variables. Research confirms that combining the depth characteristics of MRI images with key clinical information, such as PSA level and prostate volume, can significantly improve the identification ability of clinically significant prostate cancer.65 Compared with a single model that relies solely on imaging features, multi-modal models that integrate clinical variables achieve better performance on key indicators such as AUC and sensitivity, providing a more reliable basis for early, accurate diagnosis and individualized treatment planning.65–67 When higher-dimensional image data is introduced, the performance of multi-modal AI improves. For example, the multi-modal network built by integrating multi-parameter MRI and PSMA-PET/CT images at the same time is not only more accurate in the location of the lesion, but also optimizes risk stratification and clinical staging evaluation, and provides strong imaging support for the guidance of prostate targeted biopsy and the formulation of follow-up treatment strategies.68–70 This integrated model is beneficial for identifying small foci that are easily overlooked in traditional images and for improving diagnostic sensitivity and the accuracy of clinical decision-making.
Advantages of multi-modal AI in bladder cancer
In the field of bladder cancer, the advantages of multi-modal AI are mainly reflected in image-molecular linkage and real-time auxiliary diagnosis. The explainable deep learning model based on CT image construction can predict PD-L1 expression status non-invasively, providing an essential tool for screening the beneficiary population for immunotherapy and highlighting the potential of “image-molecule” integration in individualized treatment design.1,71 By combining imaging features with key biomarkers, such models not only improve disease detection but also provide a new perspective for predicting treatment response. At the same time, the combination of AI and cystoscopy technology has dramatically improved the real-time accuracy of the diagnosis process. Traditional cystoscopy is highly dependent on the operator’s experience and can easily miss early or small lesions. The real-time enhanced cystoscopy system based on deep learning can dynamically identify suspicious lesions in the video stream and automatically segment focal lesions and outline their boundaries, significantly improving lesion detection rate and operational efficiency, and providing technical support for intraoperative decision-making and follow-up treatment planning.72,73
Advantages of multi-modal AI in RCC
In RCC diagnosis and treatment, multi-modal AI plays a vital role in prognosis prediction and treatment decision-making by integrating images and clinical pathology data. Research shows that combining CT image characteristics with clinical variables such as patient age, tumor stage, kidney function index, etc., can effectively construct a predictive model, accurately assess total survival and the risk of recurrence, and provide a quantitative basis for individualized follow-up and treatment strategies.58,74,75 After the introduction of radiogenomics methods, imaging phenotypes are associated with genomic data (such as characteristic gene mutations), which can provide deeper insights into tumor behaviour and predict patients’ possible responses to targeted or immunotherapy before surgery.76 This cross-modal integration not only improves the objectivity of diagnostic staging, but also helps to establish a consistent biological interpretation framework between different data types of “the same patient”, reduces the judgement bias that may be caused by single-modal analysis, and lays the foundation for the realisation of actual precision medical treatment.64,76
Challenges of multi-modal data integration
Despite the significant advantages of multi-modal AI, its clinical transformation road still faces a series of complex challenges that run through the levels of data, algorithms, biology, and supervision.
Data heterogeneity and standardisation bottleneck
The core challenge of multi-modal data lies in its inherent heterogeneity. The CT and MRI equipment at different medical centres use different acquisition protocols and reconstruction algorithms, and the scanning process for pathological slices lacks unified standards. These factors introduce systematic deviations, resulting in a significant lack of generalization ability of the model between different scanning equipment and medical institutions.77–79 To address this problem, the field of imaging has adopted the IBSI standard to standardise feature extraction and uses ComBat, CovBat, and other methods for data harmonisation.30,80 However, the effectiveness of these methods depends heavily on algorithm selection and parameter settings, and their universality still needs to be further verified.
Questioning the rigour of methodology and the consistency of labelling
The design of multi-modal research is complex, more prone to selection bias and data leakage, and most studies lack sufficient external verification. A fundamental obstacle is that the data labelling standards are not uniform.35 Especially in digital pathology and cystoscopy video analysis, subjective differences among annotators will directly “contaminate” the quality of the training set, leading the model to learn the wrong correlation. To this end, the latest guidelines, such as TRIPOD+AI, STARD-AI, and PROBAST-AI, set stricter specifications across multiple dimensions, including prediction model construction, diagnostic accuracy evaluation, and bias risk, and emphasize clear data division, external verification, and reports on fairness and explainability.81–84
Complexity of tumor biology and model explainability dilemma
Beyond the technical level, the inherent heterogeneity of tumors and the atypical imaging characteristics of small lesions pose fundamental challenges to building biologically meaningful models. The model must be able to recognise robust patterns in complex biological background noise, and the lack of interpretability in the multi-modal model’s decision-making process remains a core bottleneck.85–88 When the model integrates high-dimensional imaging and genomic data, its decision-making logic often becomes a “black box”, which not only hinders clinicians’ understanding and trust but also makes it difficult to trace back to the original AI discovery and to deepen understanding of the disease’s biological mechanisms.
Compliance, Clinical Deployment, and Governance Barriers
Data privacy protection, cross-institutional data sharing barriers, and continuous model learning and performance drift monitoring all lack mature solutions. Seamlessly integrating AI tools into existing clinical workflows, such as PACS, LIS, EMR, etc., poses significant engineering challenges.64 In addition, the European Union’s AI Act classifies medical AI systems as “high-risk” categories, and its “superimposed” compliance model with the medical device regulations of various countries poses a severe test for the cross-border deployment of products.89
Case studies of multi-modal AI models
In the specific application of tumors in the urinary system, successful cases of multi-modal AI are accumulating rapidly. With the increasing abundance of multi-modal datasets, AI technology can integrate imaging, pathology, clinical data, and genomic information to provide strong support for the accurate diagnosis, staging, and treatment of urologic cancers.64 By integrating data across different modes, AI has shown great potential for focal detection, clinical decision-making, and personalized treatment support.
AI application of prostate cancer
At present, the research focuses on the integrated analysis of multi-modal images, including MRI and PSMA-PET/CT. Evidence shows that the AI model that integrates MRI depth characteristics with clinical variables such as PSA level and prostate volume can significantly improve the identification ability of clinically significant prostate cancer.90 Such models still maintain stable performance in external validation, demonstrating good generalization and support for clinical decision-making.27,69 The integration of mpMRI and PSMA-PET/CT into a multi-modal network shows significant advantages in the precise localisation and staging of the lesion, providing a reliable basis for biopsy guidance and treatment planning.70 With the progress of deep learning technology, AI has made a significant breakthrough in the extraction of subtle features of images. For example, through automated analysis, AI can identify early lesions that are difficult to detect by traditional methods, thereby effectively improving the sensitivity and specificity of early diagnosis.11 At the pathological level, AI has realised the automation and accuracy of Gleason scoring, significantly reducing subjective differences and providing stable and repeatable auxiliary tools for pathological grading.39 The AI system that integrates images, molecular markers, and clinical information is gradually promoting the development of prostate cancer diagnosis and treatment in the direction of individualisation, showing a certain transformation potential in risk stratification and treatment strategy optimization.91
AI application of bladder cancer
The diagnosis and staging of bladder cancer are highly dependent on integrating multi-modal data, and the application of AI in this field is gradually reshaping its diagnosis and treatment. In terms of imaging, AI can not only non-invasively predict PD-L1 expression status based on CT and other imaging features, but also provide a key reference for stratifying patients for immunotherapy.92 Furthermore, the real-time enhanced cystoscopy system based on deep learning can automatically identify and divide the lesion during the examination process, significantly improving the detection rate and boundary definition accuracy of early lesions, and providing real-time and visual support for perioperative decision-making.93–95 In the field of pathological analysis, AI has effectively improved the consistency and objectivity of diagnosis by integrating digital pathological images. For example, a deep learning model trained on full-slice images (WSI) can accurately distinguish between normal tissues, non-invasive urinary tract thelial tumors, and invasive bladder cancer, providing stable and reliable auxiliary tools for pathological diagnosis and significantly reducing inter-observer variability.40
In terms of multi-modal integration, AI builds an auxiliary system with more comprehensive judgment capabilities by integrating images, pathology, and molecular data (e.g., PD-L1 expression, gene mutations). Research shows that AI models based on image-molecular coupling perform well in predicting immunotherapy responses and provide a powerful tool for the formulation of individualized treatment strategies for bladder cancer.83 These advances jointly promote the diagnosis and treatment of bladder cancer in the direction of precision and minimally invasive procedures.
By integrating CT images, clinical information, and even pathology and transcriptomic characteristics using multi-modal deep learning, the model shows strong performance in predicting overall survival and recurrence risk in patients. Several models have undergone external validation and shown better discrimination and calibration than traditional scoring systems, including those based on multi-omics or multi-group information.58,74,75 A recent review also emphasised that incorporating genetic mutations, expression profiles, and imaging phenotypes into a unified analytical framework can substantially improve our understanding of disease mechanisms and better inform treatment decisions.74,76 Compared with the single-modal model, multi-modal AI has improved both statistical and clinical performance in most studies. To provide a more systematic overview of its core components and technical bottlenecks, Table 1 summarises the main data types, integration strategies, typical technologies, advantages, and key challenges involved in multi-modal AI.
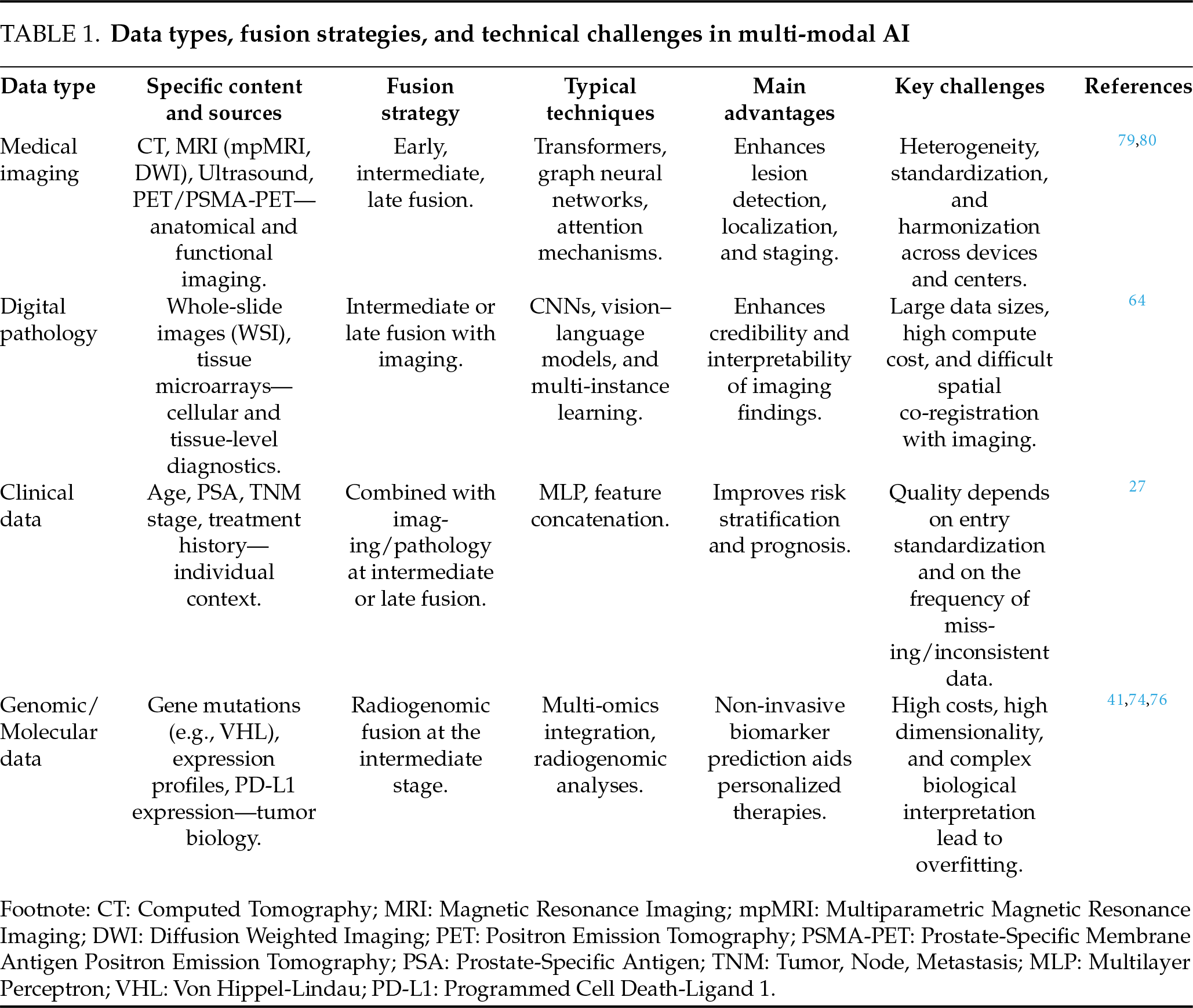
In summary, multi-modal AI is creating a continuous chain of evidence for urologic cancers, linking detection, stratification, and treatment decision-making. Upstream of diagnosis, AI models combine imaging or endoscopy with pathology to enhance tumor detection and grading. In the midstream, image–molecular fusion enables non-invasive prediction of key biomarkers. Downstream, AI integrates clinical variables with multi-omics information to provide more accurate risk assessment and treatment response prediction.79,80,96
Clinical Translation of AI in Urologic Cancer Diagnosis
Barriers to clinical translation
The path from “research-grade algorithms” to “clinically deployable tools” is bridged by three domains: regulation, interpretability, and workflow integration. On the regulatory front, a governance framework for “adaptive” or continuously learning algorithms has taken form. In the United States, the FDA’s final guidance on PCCP for AI/ML-enabled devices allows sponsors to pre-specify, in the initial marketing submission, the scope and boundaries of post-market model updates, while also requiring real-world performance monitoring, risk management, and change-impact analyses—requirements directly linked to the safety and effectiveness of diagnostic AI in hospital settings.97,98
Medical AI systems have been classified as “high-risk” since the entry into force of the European Union’s AI Act in August 2024, which imposes statutory obligations for risk management, data governance, transparency, and human oversight, with a 24–36-month transition period.99 Together with the MDR, IVDR, and GDPR, the AI Act results in a multi-layered regulatory framework that substantially increases the compliance burden for deploying urologic-oncology AI products across borders.89,99 Real-world performance surveillance remains, however, a weak link: recent work argues that metric monitoring alone cannot reliably detect data drift. Postdeployment monitoring and recalibration mechanisms are needed, while calls are for greater requirements—under STARD-AI, TRIPOD+AI, and PROBAST-AI—for external validation, bias assessment, and transparent reporting.100
From a regulatory perspective, current frameworks acknowledge the promise of AI in urologic cancer diagnosis while imposing essential constraints on its widespread use. Most approved tools are still narrow, task-specific applications (e.g., prostate pathology or mpMRI reading), with limited multi-centre and real-world evidence. In addition, stringent requirements for external validation, lifecycle performance monitoring, and bias assessment make it difficult for many research-grade, single-centre models to progress towards authorization, meaning that more complex multi-modal AI systems, such as those discussed in this review, are not yet ready for routine deployment in everyday practice.
A small but growing number of AI systems have moved beyond the research stage and into routine or near-routine clinical use in urologic oncology, providing concrete examples of how these technologies are being embedded into real workflows.
Digital pathology. Paige Prostate was the first prostate pathology AI tool to receive FDA De Novo authorization, marking a significant milestone for computer-assisted diagnosis on whole-slide images. Paige Prostate functions as a second reader: it highlights suspicious foci on digitized biopsy slides and prompts pathologists to examine these regions more carefully. In simulated and retrospective diagnostic studies, researchers have reported that Paige Prostate increases cancer detection rates and reduces missed small tumor foci, while maintaining or even improving specificity.101–103 Several independent groups have also evaluated the system and found that it performs robustly across different slide scanners and institutions, which supports its safe integration into high-volume diagnostic laboratories to improve consistency and efficiency.101–103
Prostate MRI. In radiology, several commercial AI solutions for prostate MRI have obtained 510(k) clearance and are now incorporated into standard imaging pipelines. Products such as Quantib® Prostate and Siemens AI-Rad Companion Prostate MR automatically segment the prostate gland, assist with PI-RADS scoring by pre-marking candidate lesions, and facilitate targeted fusion biopsy planning. In daily practice, these systems are typically launched directly from the PACS, generate structured reports or overlays, and are used by radiologists for decision support rather than as stand-alone readers.104,105 By reducing manual contouring and standardising lesion annotation, they aim to shorten reading time and decrease inter-reader variability, particularly among less experienced readers.104,105
Large-scale clinical evidence. The international, paired, non-inferiority PI-CAI study provides a good illustration of how MRI-based AI can perform under conditions close to real practice. In this trial, multi-centre-trained AI models for clinically significant prostate cancer detection were prospectively compared with radiologists’ interpretations under a preregistered protocol. The AI system achieved non-inferior—and in some analyses slightly superior—discrimination compared with human readers, while maintaining robust performance across centres and scanner types.91 Taken together, these examples outline a typical pathway for AI deployment in urologic oncology: initial regulatory authorization (De Novo or 510(k)), followed by integration with existing digital pathology or imaging infrastructure, and finally prospective or large-scale validation to demonstrate added clinical value beyond technical performance metrics.
Looking ahead, several priorities will determine whether AI can move from proof-of-concept tools to reliable, practice-changing technologies in urologic cancer diagnosis.11,79 In the short term, the most realistic advances are likely to come from narrowly focused, task-specific models that address clearly defined clinical pain points, such as triaging prostate biopsies, standardising PI-RADS or VI-RADS scoring, or flagging high-risk lesions on CT and MRI. These systems should be prospectively embedded into real-world workflows and evaluated not only by traditional metrics such as AUC or accuracy, but also by their impact on biopsy rates, time to diagnosis, inter-reader variability, and downstream treatment decisions.21,91
Over the medium to long term, future work will need to move beyond single-modality models towards truly multi-modal AI systems that integrate imaging, digital pathology, clinical variables, and, where available, genomic and molecular data.60,61 Such models may enable more precise risk stratification, dynamic prediction of treatment response, and identification of patients who are most likely to benefit from intensified surveillance or early intervention. To achieve this, large multi-centre consortia, federated learning frameworks, and harmonised data standards will be essential to overcome data silos, reduce batch effects, and improve generalizability across scanners, institutions, and populations.30,77
Finally, the sustainable deployment of AI in urologic oncology will depend on strengthening interpretability, governance, and regulatory alignment.68,79,106,107 Future research should prioritise models that can provide case-level explanations or visual evidence, thereby enhancing clinician trust and facilitating shared decision-making with patients.22,106 At the same time, adherence to emerging reporting and evaluation standards, together with continuous postdeployment monitoring for performance drift, bias, and safety events, will be crucial to satisfy regulatory expectations. Suppose these technical, clinical, and regulatory pillars can be aligned. In that case, AI has the potential not only to refine existing diagnostic pathways but also to reshape how urologic cancers are screened, detected, and managed in routine practice. Figure 2 shows the AI-based flowchart for urinary system tumor diagnosis, which seamlessly integrates imaging, pathology, and clinical data to help doctors more accurately evaluate the patient’s condition.
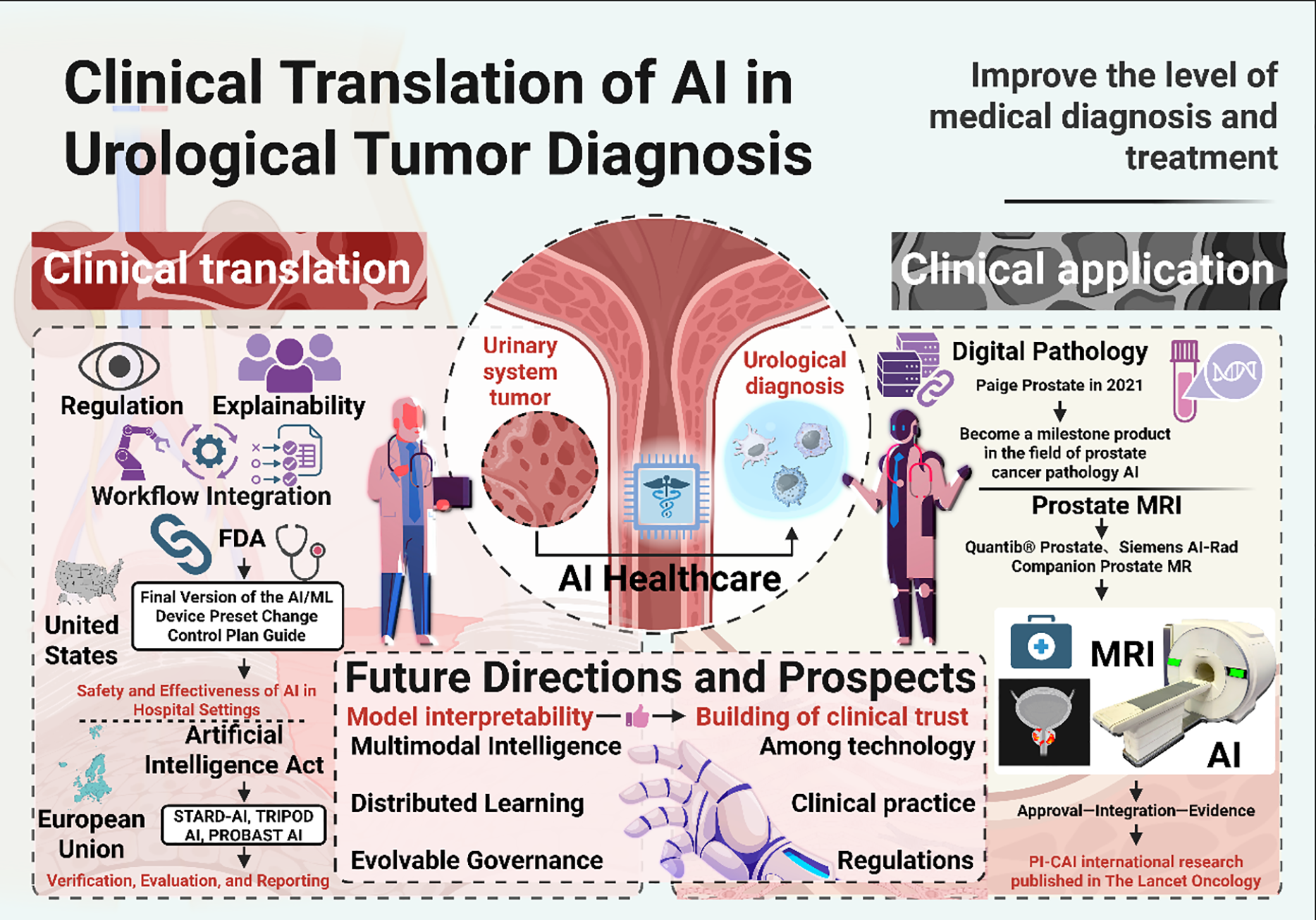
FIGURE 2. The flowchart of AI-based urinary system tumor diagnosis shows the multi-modal fusion of imaging, pathology, and clinical data. AI: Artificial intelligence; FDA: Food and drug administration; MRI: Magnetic resonance imaging; STARD-AI: Standards for reporting of diagnostic accuracy-artificial intelligence; TRIPOD: Transparent reporting of a multivariable prediction model for individual prognosis or diagnosis; PROBAST: Prediction model risk of bias assessment tool
Although multi-modal AI has shown great potential in urologic oncology, several limitations must be considered. First, many studies rely on small, single-center datasets that may not fully reflect the diversity of patient populations in real-world clinical settings. This limits the generalizability of AI models across different institutions and regions. Second, data quality and consistency are crucial. Inconsistent labeling and variations in imaging protocols can affect model accuracy.
Additionally, most studies to date have been retrospective, which may introduce biases and limit the reliability of the results. Finally, while AI tools have been promising, their clinical adoption remains slow. Clinicians need proper training and trust in these systems before widespread use. Moreover, regulatory hurdles and ethical concerns, such as patient privacy and transparency in decision-making, must also be addressed.
Reviewing the available evidence, AI has shown promising potential across the entire diagnostic process for urologic cancers. In radiology, AI significantly increases the sensitivity of lesion detection, grading, and staging in different modalities, and in pathology, whole-slide image analytics yields more objective and reproducible evaluations. At the same time, at the molecular level, integrative modeling improves risk stratification and prediction of treatment response. Multi-modal integration not only enhances the model’s performance but also lays the foundation for an explainable chain of clinical evidence. However, AI clinical transformation still faces key challenges, including insufficient model generalization, imperfect interpretability mechanisms, limited data standardization, insufficient privacy protection, and a lack of continuous governance. To this end, it is necessary to conduct multi-centre forward-looking verification and to follow international standards to promote technological transparency and standardization. In terms of clinical impact, the application of AI will show gradual development: as an auxiliary tool to improve diagnostic efficiency in the short term, support individualized treatment decision-making in the medium term, and promote the comprehensive implementation of precision medicine through the deep integration of continuous learning systems and medical information systems in the long term. In a word, AI is promoting urinary oncology towards precision and efficiency. The key to achieving this goal is to coordinate technological development, clinical verification, and standardized governance, and steadily complete the final leap from algorithm innovation to clinical benefit.
Acknowledgement: The authors would like to thank all colleagues from the participating institutions for their constructive comments and support during the preparation of this manuscript.
Funding Statement: This work was supported by grants from the Hangzhou Key Project for Agricultural and Social Development under Grant No. 20231203A12 (JZ) and the General Program of the Scientific Research Special Project for Post-Marketing Clinical Research of Innovative Drugs, Development Center for Medical Science & Technology, National Health Commission of the People’s Republic of China under Grant No. WKZX2024CX104202 (JZ).
Author Contributions: The authors confirm contribution to the paper as follows: Mingwei Zhan, Zhaokai Zhou, and Jianpeng Zhang contributed equally to this work. Conceptualization, Mingwei Zhan and Jingyu Zhu; methodology, Mingwei Zhan, Zhaokai Zhou, Jianpeng Zhang and Jingyu Zhu; software, Mingwei Zhan, Zhaokai Zhou and Jianpeng Zhang; validation, Mingwei Zhan, Zhaokai Zhou and Jianpeng Zhang; formal analysis, Mingwei Zhan, Zhaokai Zhou and Jianpeng Zhang; investigation, Zhaokai Zhou, Jianpeng Zhang, Xin Wang, Canxuan Li and Bochen Pan; resources, Jingyu Zhu, Weizhuo Wang and Run Shi; data curation, Zhaokai Zhou, Jianpeng Zhang, Xin Wang, Canxuan Li and Bochen Pan; writing—original draft preparation, Mingwei Zhan, Zhaokai Zhou and Jianpeng Zhang; writing—review and editing, Xin Wang, Canxuan Li, Bochen Pan, Zhanyang Luo, Wenjie Shi, Yongjie Wang, Minglun Li and Jingyu Zhu; visualization, Mingwei Zhan, Jianpeng Zhang, Zhanyang Luo and Wenjie Shi; supervision, Weizhuo Wang, Run Shi and Jingyu Zhu; project administration, Weizhuo Wang, Run Shi and Jingyu Zhu; funding acquisition, Jingyu Zhu. All authors reviewed and approved the final version of the manuscript.
Availability of Data and Materials: No new data were created or analyzed in this study. Data sharing is not applicable.
Ethics Approval: Not applicable. This review does not involve studies with human participants or animals performed by the authors.
Informed Consent: Not applicable.
Conflicts of Interest: The authors declare no conflicts of interest.
References
1. Siegel RL, Kratzer TB, Giaquinto AN, Sung H, Jemal A. Cancer statistics, 2025. CA Cancer J Clin 2025 2025 Jan–Feb;75(1):10–45. doi:10.3322/caac.21871. [Google Scholar] [PubMed] [CrossRef]
2. IARC. Bladder cancer–IARC. International Agency for Research on Cancer [Internet]. [cited 2025 Nov 11]. Available from: https://www.iarc.who.int/cancer-type/bladder-cancer/. [Google Scholar]
3. Bukavina L, Bensalah K, Bray F et al. Epidemiology of renal cell carcinoma: 2022 update. Eur Urol 2022 Nov;82(5):529–542. doi:10.1016/j.eururo.2022.08.019. [Google Scholar] [PubMed] [CrossRef]
4. Hafeez S, Huddart R. Advances in bladder cancer imaging. BMC Med 2013 Apr 10;11(1):104. doi:10.1186/1741-7015-11-104. [Google Scholar] [PubMed] [CrossRef]
5. Ozkan TA, Eruyar AT, Cebeci OO et al. Interobserver variability in Gleason histological grading of prostate cancer. Scand J Urol 2016 Dec;50(6):420–424. doi:10.1080/21681805.2016.1206619. [Google Scholar] [PubMed] [CrossRef]
6. Williamson SR, Rao P, Hes O et al. Challenges in pathologic staging of renal cell carcinoma: a study of interobserver variability among urologic pathologists. Am J Surg Pathol 2018 Sep;42(9):1253–1261. doi:10.1097/pas.0000000000001087. [Google Scholar] [PubMed] [CrossRef]
7. Zhu CZ, Ting HN, Ng KH, Ong TA. A review on the accuracy of bladder cancer detection methods. J Cancer 2019 Jul 8;10(17):4038–4044. doi:10.7150/jca.28989. [Google Scholar] [PubMed] [CrossRef]
8. Abouelkheir RT, Abdelhamid A, Abou El-Ghar M, El-Diasty T. Imaging of bladder cancer: standard applications and future trends. Medicina 2021 Mar 1;57(3):220. doi:10.3390/medicina57030220. [Google Scholar] [PubMed] [CrossRef]
9. Liu K, Zhang M, Li H et al. Research status and prospect of artificial intelligence technology in the diagnosis of urinary system tumors. Sheng Wu Yi Xue Gong Cheng Xue Za Zhi 2021 Dec 25;38(6):1219–1228. [Google Scholar]
10. Li M, Jiang Z, Shen W, Liu H. Deep learning in bladder cancer imaging: a review. Front Oncol 2022 Oct 20;12:930917. doi:10.3389/fonc.2022.930917. [Google Scholar] [PubMed] [CrossRef]
11. Liu X, Shi J, Li Z et al. The present and future of artificial intelligence in urological cancer. J Clin Med 2023 Jul 29;12(15):4995. doi:10.3390/jcm12154995. [Google Scholar] [PubMed] [CrossRef]
12. Zhu L, Pan J, Mou W et al. Harnessing artificial intelligence for prostate cancer management. Cell Rep Med 2024 Apr 16;5:101506. doi:10.1016/j.xcrm.2024.101506. [Google Scholar] [PubMed] [CrossRef]
13. Esteva A, Kuprel B, Novoa RA et al. Dermatologist-level classification of skin cancer with deep neural networks. Nature 2017 Jun 28;542(7639):115–118. doi:10.1038/nature21056. [Google Scholar] [PubMed] [CrossRef]
14. Alqahtani S. Systematic review of AI-assisted MRI in prostate cancer diagnosis: enhancing accuracy through second opinion tools. Diagnostics 2024 Nov 15;14(22):2576. doi:10.3390/diagnostics14222576. [Google Scholar] [PubMed] [CrossRef]
15. Bhattacharya I, Khandwala YS, Vesal S et al. A review of artificial intelligence in prostate cancer detection on imaging. Ther Adv Urol 2022 Oct 10;14:17562872221128791. doi:10.1177/17562872221128791. [Google Scholar] [PubMed] [CrossRef]
16. Parikh RB, Ferrell WJ, Girard A et al. The impact of machine learning mortality risk prediction on clinician prognostic accuracy and decision support: a randomized vignette study. Med Decis Making 2025 Aug;45(6):690–702. doi:10.1177/0272989x251349489. [Google Scholar] [PubMed] [CrossRef]
17. Zhao Y, Zhang L, Zhang S et al. Machine learning-based MRI imaging for prostate cancer diagnosis: systematic review and meta-analysis. Prostate Cancer Prostatic Dis 2025 Jul;28:1–8. doi:10.1038/s41391-025-00997-2. [Google Scholar] [PubMed] [CrossRef]
18. Egevad L, Camilloni A, Delahunt B et al. The role of artificial intelligence in the evaluation of prostate pathology. Pathol Int 2025 May;75(5):213–220. doi:10.1111/pin.70015. [Google Scholar] [PubMed] [CrossRef]
19. Trägårdh E, Ulén J, Enqvist O et al. A fully automated AI-based method for tumor detection and quantification on [18F]PSMA-1007 PET-CT images in prostate cancer. EJNMMI Phys 2025 Aug 20;12(1):78. doi:10.1186/s40658-025-00786-9. [Google Scholar] [PubMed] [CrossRef]
20. Zhou SR, Zhang L, Choi MH et al. ProMUS-NET: artificial intelligence detects more prostate cancer than urologists on micro-ultrasonography. BJU Int 2025 Dec;136(6):1071–1079. doi:10.1111/bju.16892. [Google Scholar] [PubMed] [CrossRef]
21. Bulten W, Balkenhol M, Belinga JA et al. Artificial intelligence assistance significantly improves Gleason grading of prostate biopsies by pathologists. Mod Pathol 2021 Mar;34(3):660–671. doi:10.1038/s41379-020-0640-y. [Google Scholar] [PubMed] [CrossRef]
22. Mittmann G, Laiouar-Pedari S, Mehrtens HA et al. Pathologist-like explainable AI for interpretable Gleason grading in prostate cancer. Nat Commun 2025 Oct 8;16(1):8959. doi:10.1038/s41467-025-64712-4. [Google Scholar] [PubMed] [CrossRef]
23. Frewing A, Gibson AB, Robertson R, Urie PM, Corte DD. Don’t fear the artificial intelligence: a systematic review of machine learning for prostate cancer detection in pathology. Arch Pathol Lab Med 2024 May 1;148(5):603–612. doi:10.5858/arpa.2022-0460-ra. [Google Scholar] [PubMed] [CrossRef]
24. Huo X, Ong KH, Lau KW et al. A comprehensive AI model development framework for consistent Gleason grading. Commun Med 2024 May 9;4(1):84. doi:10.1038/s43856-024-00502-1. [Google Scholar] [PubMed] [CrossRef]
25. Zhu M, Sali R, Baba F et al. Artificial intelligence in pathologic diagnosis, prognosis and prediction of prostate cancer. Am J Clin Exp Urol 2024 Aug 25;12(4):200–215. doi:10.62347/jsae9732. [Google Scholar] [PubMed] [CrossRef]
26. He M, Cao Y, Chi C et al. Research progress on deep learning in magnetic resonance imaging-based diagnosis and treatment of prostate cancer: a review on the current status and perspectives. Front Oncol 2023 Jun 13;13:1189370. doi:10.3389/fonc.2023.1189370. [Google Scholar] [PubMed] [CrossRef]
27. Roest C, Yakar D, Rener Sitar DI et al. Multi-modal AI combining clinical and imaging inputs improves prostate cancer detection. Invest Radiol 2024 Dec 1;59(12):854–860. doi:10.1097/rli.0000000000001102. [Google Scholar] [PubMed] [CrossRef]
28. Bozgo V, Roest C, van Oort I et al. Prostate MRI and artificial intelligence during active surveillance: should we jump on the bandwagon? Eur Radiol 2024 Dec;34(12):7698–7704. doi:10.1007/s00330-024-10869-3. [Google Scholar] [PubMed] [CrossRef]
29. Martinez-Marroquin E, Chau M, Turner M, Haxhimolla H, Paterson C. Use of artificial intelligence in discerning the need for prostate biopsy and readiness for clinical practice: a systematic review protocol. Syst Rev 2023 Jul 17;12(1):126. doi:10.1186/s13643-023-02282-6. [Google Scholar] [PubMed] [CrossRef]
30. Giganti F, Moreira da Silva N, Yeung M et al. AI-powered prostate cancer detection: a multi-centre, multi-scanner validation study. Eur Radiol 2025 Aug;35(8):4915–4924. doi:10.1007/s00330-024-11323-0. [Google Scholar] [PubMed] [CrossRef]
31. Jambor I, Falagario U, Ratnani P et al. Prediction of biochemical recurrence in prostate cancer patients who underwent prostatectomy using routine clinical prostate multiparametric MRI and decipher genomic score. J Magn Reson Imaging 2020 Apr;51(4):1075–1085. doi:10.1002/jmri.26928. [Google Scholar] [PubMed] [CrossRef]
32. Kim H, Kang SW, Kim JH et al. The role of AI in prostate MRI quality and interpretation: opportunities and challenges. Eur J Radiol 2023 Aug;165:110887. doi:10.1016/j.ejrad.2024.111585. [Google Scholar] [PubMed] [CrossRef]
33. Ursprung S, Agrotis G, van Houdt PJ et al. Prostate MRI using deep learning reconstruction in response to cancer screening demands—a systematic review and meta-analysis. J Pers Med 2025 Jul 2;15(7):284. doi:10.3390/jpm15070284. [Google Scholar] [PubMed] [CrossRef]
34. Fay M, Liao RS, Lone ZM et al. Artificial intelligence–based digital histologic classifier for prostate cancer risk stratification: independent blinded validation in patients treated with radical prostatectomy. JCO Clin Cancer Inform 2025 Jun;9:e2400292. doi:10.1200/CCI-24-00292. [Google Scholar] [PubMed] [CrossRef]
35. Ma X, Zhang Q, He L et al. Artificial intelligence application in the diagnosis and treatment of bladder cancer: advance, challenges, and opportunities. Front Oncol 2024 Nov 7;14:1487676. doi:10.3389/fonc.2024.1487676. [Google Scholar] [PubMed] [CrossRef]
36. Huang H, Huang Y, Kaggie JD et al. Multiparametric MRI-based deep learning radiomics model for assessing 5-year recurrence risk in non-muscle invasive bladder cancer. J Magn Reson Imaging 2025 Mar;61(3):1442–1456. doi:10.1002/jmri.29574. [Google Scholar] [PubMed] [CrossRef]
37. He C, Xu H, Yuan E et al. The accuracy and quality of image-based artificial intelligence for muscle-invasive bladder cancer prediction. Insights Imaging 2024 Aug 1;15(1):185. doi:10.1186/s13244-024-01780-y. [Google Scholar] [PubMed] [CrossRef]
38. Li L, Jiang L, Yang K, Luo B, Wang X. A novel artificial intelligence segmentation model for early diagnosis of bladder tumors. Abdom Radiol 2025 Jul;50(7):3092–3099. doi:10.1007/s00261-024-04715-9. [Google Scholar] [PubMed] [CrossRef]
39. Park JY, Kim J, Kim YJ et al. Multi-institutional validation of AI models for classifying urothelial neoplasms in digital pathology. Sci Rep 2025 Oct 24;15(1):37215. doi:10.1038/s41598-025-21096-1. [Google Scholar] [PubMed] [CrossRef]
40. Khoraminia F, Fuster S, Kanwal N et al. Artificial intelligence in digital pathology for bladder cancer: hype or hope? a systematic review. Cancers 2023 Sep 12;15(18):4518. doi:10.3390/cancers15184518. [Google Scholar] [PubMed] [CrossRef]
41. Han X, Guan J, Guo L et al. A CT-based interpretable deep learning signature for predicting PD-L1 expression in bladder cancer: a two-center study. Cancer Imaging 2025 Mar 10;25(1):27. doi:10.1186/s40644-025-00849-1. [Google Scholar] [PubMed] [CrossRef]
42. O’Sullivan NJ, Temperley HC, Corr A et al. Current role of radiomics and radiogenomics in predicting oncological outcomes in bladder cancer. Curr Urol 2025 Jan;19(1):43–48. doi:10.1097/cu9.0000000000000235. [Google Scholar] [PubMed] [CrossRef]
43. Wang Z, Zhang X, Wang X et al. Deep learning techniques for imaging diagnosis of renal cell carcinoma: current and emerging trends. Front Oncol 2023 Sep 1;13:1152622. doi:10.3389/fonc.2023.1152622. [Google Scholar] [PubMed] [CrossRef]
44. Han JH, Kim BW, Kim TM et al. Fully automated segmentation and classification of renal tumors on CT scans via machine learning. BMC Cancer 2025 Jan 29;25(1):173. [Google Scholar]
45. Xi IL, Zhao Y, Wang R et al. Deep learning to distinguish benign from malignant renal lesions based on routine MR imaging. Clin Cancer Res 2020 Apr 15;26(8):1944–1952. doi:10.1158/1078-0432.ccr-19-0374. [Google Scholar] [PubMed] [CrossRef]
46. Dai C, Xiong Y, Zhu P et al. Deep learning assessment of small renal masses at contrast-enhanced multiphase CT. Radiology 2024 May;311(2):e232178. doi:10.1148/radiol.232178. [Google Scholar] [PubMed] [CrossRef]
47. Bleker J, Kwee TC, Rouw D et al. A deep learning masked segmentation alternative to manual segmentation in biparametric MRI prostate cancer radiomics. Eur Radiol 2022 Sep;32(9):6526–6535. doi:10.1007/s00330-022-08712-8. [Google Scholar] [PubMed] [CrossRef]
48. Klontzas ME, Kalarakis G, Koltsakis E et al. Convolutional neural networks for the differentiation between benign and malignant renal tumors with a multicenter international computed tomography dataset. Insights Imaging 2024 Jan 25;15(1):26. doi:10.1186/s13244-023-01601-8. [Google Scholar] [PubMed] [CrossRef]
49. Xiong Y, Yao L, Lin J et al. Artificial intelligence links CT images to pathologic features and survival outcomes of renal masses. Nat Commun 2025 Feb 7;16(1):1425. doi:10.1038/s41467-025-56784-z. [Google Scholar] [PubMed] [CrossRef]
50. Zhu M, Ren B, Richards R et al. Development and evaluation of a deep neural network for histologic classification of renal cell carcinoma on biopsy and surgical resection slides. Sci Rep 2021 Mar 29;11(1):7080. doi:10.1038/s41598-021-86540-4. [Google Scholar] [PubMed] [CrossRef]
51. Chanchal AK, Lal S, Suresh S. Development and evaluation of deep neural networks for the classification of subtypes of renal cell carcinoma from kidney histopathology images. Sci Rep 2025 Aug 5;15(1):28585. doi:10.1038/s41598-025-10712-9. [Google Scholar] [PubMed] [CrossRef]
52. Jiang W, Qi S, Chen C, Wang W, Chen X. Diagnosis of clear cell renal cell carcinoma via a deep learning model with whole-slide images. Ther Adv Urol 2025 May 3;17:17562872251333865. doi:10.1177/17562872251333865. [Google Scholar] [PubMed] [CrossRef]
53. Schulz S, Woerl AC, Jungmann F et al. Multimodal deep learning for prognosis prediction in renal cancer. Front Oncol 2021 Nov 24;11:788740. doi:10.3389/fonc.2021.788740. [Google Scholar] [PubMed] [CrossRef]
54. Tang H, Zhao H, Yu S et al. Automatic segmentation of clear cell renal cell carcinoma based on deep learning and a preliminary exploration of the tumor microenvironment. Transl Androl Urol 2025 Jul 30;14(7):2059–2074. doi:10.21037/tau-2025-400. [Google Scholar] [PubMed] [CrossRef]
55. Bai Y, An ZC, Li F et al. Deep learning using contrast-enhanced ultrasound images to predict the nuclear grade of clear cell renal cell carcinoma. World J Urol 2024 Mar 21;42(1):184. doi:10.1007/s00345-024-04889-3. [Google Scholar] [PubMed] [CrossRef]
56. Chen YF, Fu F, Zhuang JJ et al. Ultrasound-based radiomics for predicting the WHO/ISUP grading of clear-cell renal cell carcinoma. Ultrasound Med Biol 2024 Nov;50(11):1619–1627. doi:10.1016/j.ultrasmedbio.2024.06.004. [Google Scholar] [PubMed] [CrossRef]
57. Luo Y, Liu X, Jia Y, Zhao Q. Ultrasound contrast-enhanced radiomics model for preoperative prediction of the tumor grade of clear cell renal cell carcinoma: an exploratory study. BMC Med Imaging 2024 Jun 6;24(1):135. doi:10.1186/s12880-024-01317-1. [Google Scholar] [PubMed] [CrossRef]
58. Mahootiha M, Qadir HA, Bergsland J, Balasingham I. Multi-modal deep learning for personalized renal cell carcinoma prognosis: Integrating CT imaging and clinical data. Comput Methods Programs Biomed 2024 Feb;244:107978. doi:10.1016/j.cmpb.2023.107978. [Google Scholar] [PubMed] [CrossRef]
59. Wu P, Wu K, Li Z et al. Multi-modal investigation of bladder cancer data based on computed tomography, whole slide imaging, and transcriptomics. Quant Imaging Med Surg 2023 Feb 1;13(2):1023–1035. doi:10.21037/qims-22-679. [Google Scholar] [PubMed] [CrossRef]
60. Stahlschmidt SR, Ulfenborg B, Synnergren J. Multi-modal deep learning for biomedical data fusion: a review. Brief Bioinform 2022 Mar 10;23:bbab569. doi:10.1093/bib/bbab569. [Google Scholar] [PubMed] [CrossRef]
61. Yang H, Yang M, Chen J et al. Multi-modal deep learning approaches for precision oncology: a comprehensive review. Brief Bioinform 2024 Nov 22;26(1):bbae699. doi:10.1093/bib/bbae699. [Google Scholar] [PubMed] [CrossRef]
62. Taguelmimt K, Andrade-Miranda G, Harb H et al. Towards more reliable prostate cancer detection: Incorporating clinical data and uncertainty in MRI deep learning. Comput Biol Med 2025 Aug;194:110440. doi:10.1016/j.compbiomed.2025.110440. [Google Scholar] [PubMed] [CrossRef]
63. Li Y, El Habib Daho M, Conze PH et al. A review of deep learning-based information fusion techniques for multi-modal medical image classification. Comput Biol Med 2024 Jul;177:108635. doi:10.1016/j.compbiomed.2024.108635. [Google Scholar] [PubMed] [CrossRef]
64. Simon BD, Moise D, Patel S, Harmon SA, Türkbey B. The future of multi-modal artificial intelligence models for integrating imaging and clinical metadata: a narrative review. Diagn Interv Radiol 2025 Jul 8;31(4):303–312. doi:10.4274/dir.2024.242631. [Google Scholar] [PubMed] [CrossRef]
65. Bacchetti E, De Nardin A, Giannarini G et al. A deep learning model integrating clinical and MRI features improves risk stratification and reduces unnecessary biopsies in men with suspected prostate cancer. Cancers 2025 Jul 7;17(13):2257. doi:10.3390/cancers17132257. [Google Scholar] [PubMed] [CrossRef]
66. Woźnicki P, Westhoff N, Huber T et al. Multiparametric MRI for Prostate cancer characterization: combined use of radiomics model with PI-RADS and clinical parameters. Cancer 2020 Jul 2;12(7):1767. doi:10.3390/cancers12071767. [Google Scholar] [PubMed] [CrossRef]
67. Hiremath A, Shiradkar R, Fu P et al. An integrated nomogram combining deep learning, Prostate Imaging-Reporting and Data System (PI-RADS) scoring, and clinical variables for identification of clinically significant prostate cancer on biparametric MRI: a retrospective multi-centre study. Lancet Digit Health 2021 Jul;3(7):e445–e454. doi:10.1016/s2589-7500(21)00082-0. [Google Scholar] [PubMed] [CrossRef]
68. Lambert B, Forbes F, Doyle S, Dehaene H, Dojat M. Trustworthy clinical AI solutions: a unified review of uncertainty quantification in Deep Learning models for medical image analysis. Artif Intell Med 2024 Apr;150:102830. doi:10.1016/j.artmed.2024.102830. [Google Scholar] [PubMed] [CrossRef]
69. Takeda H, Akatsuka J, Kiriyama T et al. Clinically significant prostate cancer prediction using multi-modal deep learning with prostate-specific antigen restriction. Curr Oncol 2024 Nov 15;31(11):7180–7189. doi:10.3390/curroncol31110530. [Google Scholar] [PubMed] [CrossRef]
70. Yao F, Lin H, Xue YN et al. Multi-modal imaging deep learning model for predicting extraprostatic extension in prostate cancer using MpMRI and 18 F-PSMA-PET/CT. Cancer Imaging 2025 Aug 19;25(1):103. doi:10.1186/s40644-025-00927-4. [Google Scholar] [PubMed] [CrossRef]
71. Cao Y, Zhu H, Li Z, Liu C, Ye J. CT image-based radiomic analysis for detecting PD-L1 expression status in bladder cancer patients. Acad Radiol 2024 Sep;31(9):3678–3687. doi:10.1016/j.acra.2024.02.047. [Google Scholar] [PubMed] [CrossRef]
72. Kim J, Ham WS, Koo KC et al. Evaluation of the diagnostic efficacy of an AI-based software in detecting suspicious areas of bladder cancer using cystoscopy images. J Clin Med 2024 Nov 24;13(23):7110. doi:10.3390/jcm13237110. [Google Scholar] [PubMed] [CrossRef]
73. Ye Z, Li D, Wu J et al. Leveraging deep learning in real-time intelligent cystoscopy for bladder lesion detection. Ann Surg Oncol 2025 May;32(5):3220–3226. doi:10.1245/s10434-025-17015-3. [Google Scholar] [PubMed] [CrossRef]
74. Ferro M, Musi G, Marchioni M et al. Radiogenomics in renal cancer management-current evidence and future prospects. Int J Mol Sci 2023 Feb 27;24(5):4615. doi:10.3390/ijms24054615. [Google Scholar] [PubMed] [CrossRef]
75. Wang S, Zhu C, Jin Y et al. A multi-model based on radiogenomics and deep learning techniques associated with histological grade and survival in clear cell renal cell carcinoma. Insights Imaging 2023 Nov 27;14(1):207. doi:10.1186/s13244-023-01557-9. [Google Scholar] [PubMed] [CrossRef]
76. Guo Y, Li T, Gong B et al. From images to genes: radiogenomics based on artificial intelligence to achieve non-invasive precision medicine in cancer patients. Adv Sci 2025 Jan;12(2):e2408069. doi:10.1002/advs.202408069. [Google Scholar] [PubMed] [CrossRef]
77. Zhao B. Understanding sources of variation to improve the reproducibility of radiomics. Front Oncol 2021 Mar 29;11:633176. doi:10.3389/fonc.2021.633176. [Google Scholar] [PubMed] [CrossRef]
78. Harder C, Pryalukhin A, Quaas A et al. Enhancing prostate cancer diagnosis: artificial intelligence-driven virtual biopsy for optimal magnetic resonance imaging-targeted biopsy approach and gleason grading strategy. Mod Pathol 2024 Oct;37(10):100564. doi:10.1016/j.modpat.2024.100564. [Google Scholar] [PubMed] [CrossRef]
79. Pak S, Park SG, Park J et al. Applications of artificial intelligence in urologic oncology. Investig Clin Urol 2024 May;65(3):202–216. doi:10.4111/icu.20230435. [Google Scholar] [PubMed] [CrossRef]
80. Gelikman DG, Rais-Bahrami S, Pinto PA, Turkbey B. AI-powered radiomics: revolutionizing detection of urologic malignancies. Curr Opin Urol 2024 Jan 1;34(1):1–7. doi:10.1097/mou.0000000000001144. [Google Scholar] [PubMed] [CrossRef]
81. Collins GS, Moons KGM, Dhiman P et al. TRIPOD+AI statement: updated guidance for reporting clinical prediction models that use regression or machine learning methods. BMJ 2024 Apr 16;385:e078378. doi:10.1136/bmj.q902. [Google Scholar] [PubMed] [CrossRef]
82. Sounderajah V, Guni A, Liu X et al. The STARD-AI reporting guideline for diagnostic accuracy studies using artificial intelligence. Nat Med 2025 Oct;31(10):3283–3289. doi:10.1038/s41591-025-03953-8. [Google Scholar] [PubMed] [CrossRef]
83. Moons KGM, Damen JAA, Kaul T et al. PROBAST+AI: an updated quality, risk of bias, and applicability assessment tool for prediction models using regression or artificial intelligence methods. BMJ 2025 Mar 24;388:e082505. doi:10.1136/bmj-2024-082505. [Google Scholar] [PubMed] [CrossRef]
84. Park SH, Suh CH. Reporting guidelines for artificial intelligence studies in healthcare (for Both Conventional and Large Language Modelswhat’s new in 2024. Korean J Radiol 2024 Aug;25(8):687–690. doi:10.3348/kjr.2024.0598. [Google Scholar] [PubMed] [CrossRef]
85. Wu Y, Shieh A, Cen S et al. Prediction of PD-L1 and CD68 in clear cell renal cell carcinoma with green learning. J Imaging 2025 Jun 10;11(6):191. doi:10.3390/jimaging11060191. [Google Scholar] [PubMed] [CrossRef]
86. Wu J, Li J, Huang B et al. Radiomics predicts the prognosis of patients with clear cell renal cell carcinoma by reflecting the tumor heterogeneity and microenvironment. Cancer Imaging 2024 Sep 16;24(1):124. doi:10.1186/s40644-024-00768-7. [Google Scholar] [PubMed] [CrossRef]
87. Gouravani M, Shahrabi Farahani M, Salehi MA et al. Diagnostic performance of artificial intelligence in detection of renal cell carcinoma: a systematic review and meta-analysis. BMC Cancer 2025 Jan 27;25(1):155. doi:10.1186/s12885-025-13547-9. [Google Scholar] [PubMed] [CrossRef]
88. Sheikhy A, Dehghani Firouzabadi F, Lay N et al. State of the art review of AI in renal imaging. Abdom Radiol 2025 Nov;50(11):5305–5323. doi:10.1007/s00261-025-04963-3. [Google Scholar] [PubMed] [CrossRef]
89. European Commission. Artificial intelligence in healthcare [Internet]. [cited 2025 Nov 11]. Available from: https://health.ec.europa.eu/ehealth-digital-health-and-care/artificial-intelligence-healthcare_en. [Google Scholar]
90. Tong L, Song K, Wang Y. Zinc oxide nanoparticles dissolution and toxicity enhancement by polystyrene microplastics under sunlight irradiation. Chemosphere 2022 Jul;299(15):134421. doi:10.1016/j.chemosphere.2022.134421. [Google Scholar] [PubMed] [CrossRef]
91. Sun Z, Wang K, Kong Z et al. A multicenter study of artificial intelligence-aided software for detecting visible clinically significant prostate cancer on mpMRI. Insights Imaging 2023 Apr 30;14(1):72. doi:10.1186/s13244-023-01421-w. [Google Scholar] [PubMed] [CrossRef]
92. Lee CM, Hong SB, Lee NK et al. Preliminary data on computed tomography-based radiomics for predicting programmed death ligand 1 expression in urothelial carcinoma. Kosin Med J 2024 Jul 18;39(3):186–194. doi:10.7180/kmj.24.103. [Google Scholar] [CrossRef]
93. Chang TC, Shkolyar E, Del Giudice F et al. Real-time detection of bladder cancer using augmented cystoscopy with deep learning: a pilot study. J Endourol 2023 Jul 11;36(8):e1. doi:10.1089/end.2023.0056. [Google Scholar] [PubMed] [CrossRef]
94. Shkolyar E, Jia X, Chang TC et al. Augmented bladder tumor detection using deep learning. Eur Urol 2019 Dec;76(6):714–718. doi:10.1016/j.eururo.2019.11.020. [Google Scholar] [PubMed] [CrossRef]
95. Chan TH, Haworth A, Wang A et al. Detecting localised prostate cancer using radiomic features in PSMA PET and multiparametric MRI for biologically targeted radiation therapy. EJNMMI Res 2023 Apr 26;13(1):34. doi:10.1186/s13550-023-00984-5. [Google Scholar] [PubMed] [CrossRef]
96. Yan K, Fong S, Li T, Song Q. Multi-modal machine learning for prognosis and survival prediction in renal cell carcinoma patients: a two-stage framework with model fusion and interpretability analysis. Appl Sci 2024;14(13):5686. doi:10.1109/iceeie66203.2025.11252428. [Google Scholar] [CrossRef]
97. Marketing submission recommendations for a predetermined change control plan for artificial intelligence-enabled device software functions [Internet]. [cited 2025 Nov 11]. Available from: https://www.fda.gov/regulatory-information/search-fda-guidance-documents/marketing-submission-recommendations-predetermined-change-control-plan-artificial-intelligence. [Google Scholar]
98. Request for public comment on measuring and evaluating AI-enabled medical device performance in the real-world [Internet]. [cited 2025 Nov 11]. Available from: https://www.fda.gov/medical-devices/digital-health-center-excellence/request-public-comment-measuring-and-evaluating-artificial-intelligence-enabled-medical-device. [Google Scholar]
99. Aboy M, Minssen T, Vayena E. Navigating the EU AI Act: implications for regulated digital medical products. npj Digit Med 2024 Sep 6;7(1):237. doi:10.1038/s41746-024-01232-3. [Google Scholar] [PubMed] [CrossRef]
100. Andersen ES, Birk-Korch JB, Hansen RS et al. Monitoring performance of clinical artificial intelligence in health care: a scoping review. JBI Evid Synth 2024 Dec 1;22(12):2423–2446. doi:10.11124/jbies-24-00042. [Google Scholar] [PubMed] [CrossRef]
101. Device Classification Under Section 513(f)(2) (De NovoPaige Prostate (DEN200080) [Internet]. [cited 2025 Nov 11]. Available from: https://www.accessdata.fda.gov/scripts/cdrh/cfdocs/cfpmn/denovo.cfm?id=DEN200080. [Google Scholar]
102. Perincheri S, Levi AW, Celli R et al. An independent assessment of an artificial intelligence system for prostate cancer detection shows strong diagnostic accuracy. Mod Pathol 2021 Aug;34(8):1588–1595. doi:10.1038/s41379-021-00794-x. [Google Scholar] [PubMed] [CrossRef]
103. Raciti P, Sue J, Retamero JA et al. Clinical validation of artificial intelligence-augmented pathology diagnosis demonstrates significant gains in diagnostic accuracy in prostate cancer detection. Arch Pathol Lab Med 2023 Oct 1;147(10):1178–1185. doi:10.5858/arpa.2022-0066-oa. [Google Scholar] [PubMed] [CrossRef]
104. K221106: Quantib Prostate—510(k) Premarket Notification [Internet]. [cited 2025 Nov 11]. Available from: https://www.accessdata.fda.gov/scripts/cdrh/cfdocs/cfpmn/pmn.cfm?ID=K221106. [Google Scholar]
105. K193283: AI-Rad Companion Prostate MR—510(k) Premarket Notification [Internet]. [cited 2025 Nov 11]. Available from: https://www.accessdata.fda.gov/scripts/cdrh/cfdocs/cfpmn/pmn.cfm?ID=K193283. [Google Scholar]
106. Kore A, Abbasi Bavil E, Subasri V et al. Empirical data drift detection experiments on real-world medical imaging data. Nat Commun 2024 Feb 29;15(1):1887. doi:10.1038/s41467-024-46142-w. [Google Scholar] [PubMed] [CrossRef]
107. Marey A, Arjmand P, Alerab ADS et al. Explainability, transparency and black box challenges of AI in radiology: impact on patient care in cardiovascular radiology. Egypt J Radiol Nucl Med 2024 Sep 13;55(1):183. doi:10.1186/s43055-024-01356-2. [Google Scholar] [CrossRef]
Cite This Article
 Copyright © 2026 The Author(s). Published by Tech Science Press.
Copyright © 2026 The Author(s). Published by Tech Science Press.This work is licensed under a Creative Commons Attribution 4.0 International License , which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.


 Submit a Paper
Submit a Paper Propose a Special lssue
Propose a Special lssue View Full Text
View Full Text Download PDF
Download PDF Downloads
Downloads
 Citation Tools
Citation Tools