 Open Access
Open Access
REVIEW
A Survey of Generative Adversarial Networks for Medical Images
1 Department of Biomedical Engineering, KMCT College of Engineering for Women, Kozhikode, 673601, Kerala, India
2 Department of Electronics and Communication Engineering, National Institute of Technology Calicut, Kozhikode, 673601, Kerala, India
3 Electrical Engineering Department, College of Engineering, King Khalid University, Abha, 61413, Saudi Arabia
4 Departamento de Ciencias de la Construcción, Facultad de Ciencias de la Construcción Ordenamiento Territorial, Universidad Tecnológica Metropolitana, Santiago, 7800002, Chile
* Corresponding Author: Sameera V. Mohd Sagheer. Email:
# These authors contributed equally to this work
(This article belongs to the Special Issue: Advances in AI-Driven Computational Modeling for Image Processing)
Computer Modeling in Engineering & Sciences 2026, 146(2), 4 https://doi.org/10.32604/cmes.2025.067108
Received 25 April 2025; Accepted 08 August 2025; Issue published 26 February 2026
Abstract
Over the years, Generative Adversarial Networks (Keywords
Supplementary Material
Supplementary Material FileMedical imaging is a fundamental component of modern healthcare, offering non-invasive methods to visualize the internal structures of the human body. It supports diagnosing, planning treatment, and monitoring a range of medical conditions, utilizing common imaging techniques such as X-rays, Magnetic Resonance
These imaging methods, such as MR, CT, PET, and ultrasound, serve various diagnostic purposes, offering detailed insights into the body’s internal structures. Each modality is suited for specific clinical applications, with MR excelling in soft tissue imaging, CT providing high-resolution bone images, PET detecting metabolic activity, and ultrasound enabling real-time visualization of soft tissues. An outline of the different medical imaging techniques is provided below, outlining their specific uses and advantages in healthcare.
1. Ultrasound (US) Images: US imaging is widely utilized in diagnostic fields like cardiology, obstetrics, and gynecology due to its ability to generate high-resolution images without subjecting patients to ionizing radiation [1]. The technique works by emitting high-frequency sound waves (typically between
2. Magnetic Resonance (MR) Images: MR imaging uses magnetic fields and radio waves to generate detailed images of internal body structures that are difficult to capture with other imaging modalities. The human body is composed of billions of hydrogen atoms, which align with the magnetic field when exposed to it. This alignment causes the hydrogen atoms, which are positively charged, to orient uniformly. A pulse of radio frequency energy is applied to disrupt this alignment, causing the protons to shift. The protons emit energy when they return to the initial position. The intensity of this released energy is measured and displayed on a gray scale, forming cross-sectional images of the body. MR images are created using complex values that correspond to the Fourier transform of the magnetization distribution [7–9]. Fig. 2 illustrates the formation of MR image.
3. Computed Tomography (CT) Images: A CT scan employs computer algorithms to process multiple X-ray images procured from various angles. The combination of these images generate cross-sectional (tomographic) images of a given region within the scanned object. This technique is particularly useful in detecting hemorrhages and other conditions that may resemble a stroke, such as tumors or subdural/extradural hematomas [10]. However, CT imaging relies on ionizing radiation, and the exposure from this radiation accumulates over time. To minimize the impact of ionizing radiation, Low Dose Computed Tomography (LDCT) images are generated as an alternative [11–14].
4. Positron Emission Tomography (PET) Images: PET is a molecular imaging method that has quickly become a vital tool for functional imaging. PET works by generating images of the body based on the radiation emitted by radioactive substances introduced into the body. These substances, often tagged with short-lived radioactive isotopes like Carbon-
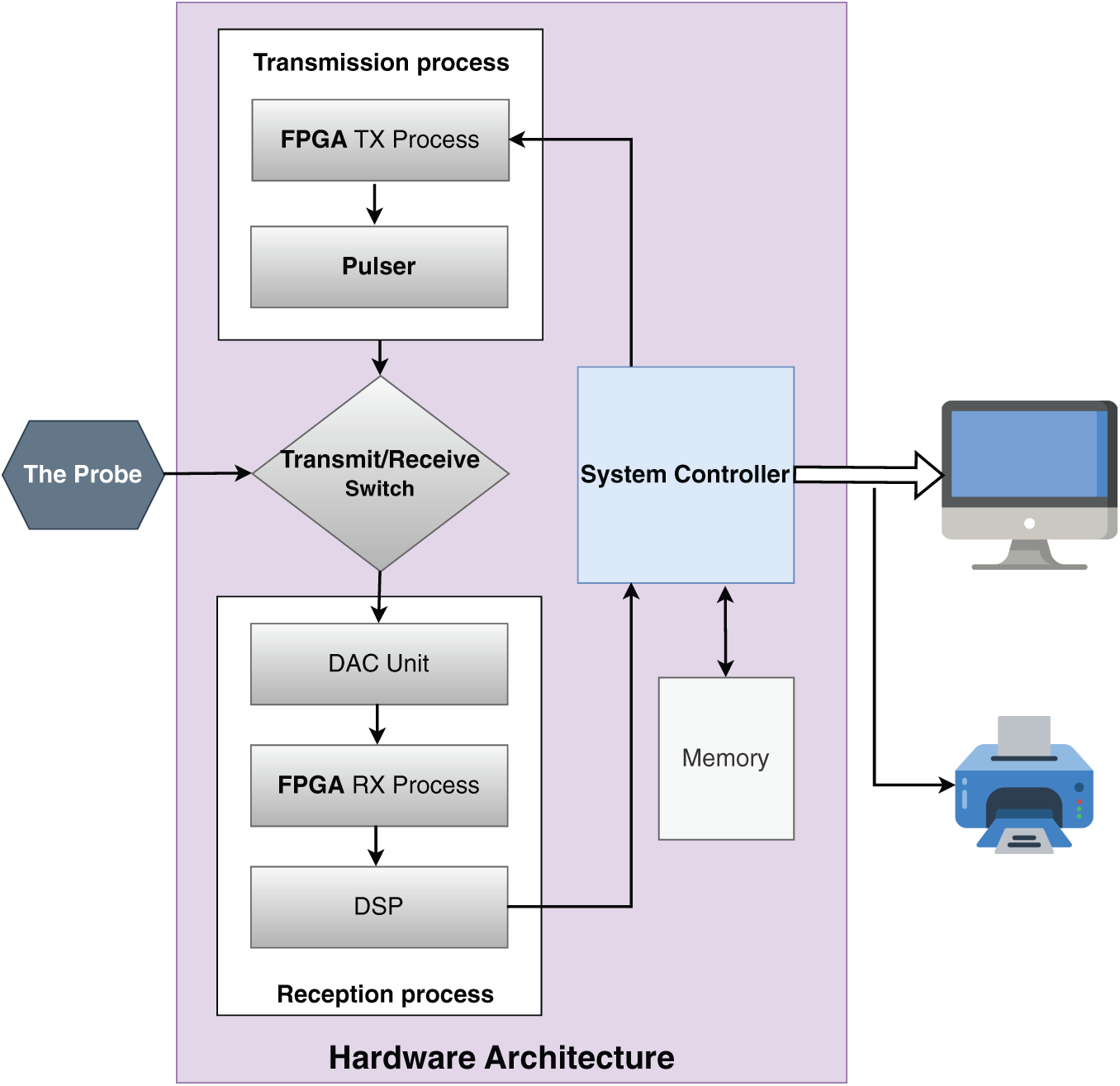
Figure 1: Formation of US image [2]
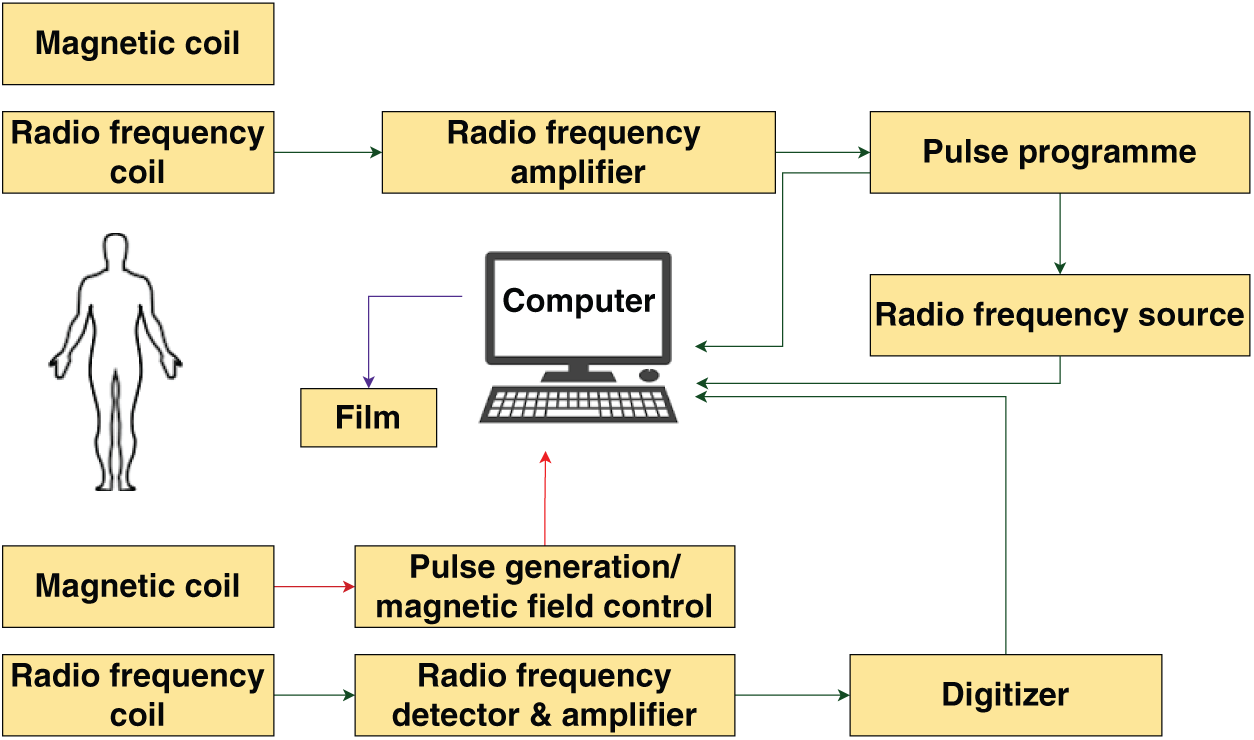
Figure 2: Formation of MR image [8]
Each imaging modality provides unique understanding, helping clinicians in making better decisions. The complexity and quantity of medical imaging data emphasizes for advanced computational tools to support analysis and interpretation. Generative Adversarial Networks
Moreover,
1.1 Overview of Generative Adverserial Network (GAN)
Recent advancements in computing power and big data analysis have significantly boosted the development of Artificial Intelligence (AI)—systems that mimic human cognitive abilities, such as learning, problem-solving, and decision-making [18–22]. AI can process and analyze large datasets efficiently. Machine Learning (ML), a subset of AI, learns from data by identifying patterns and features [23]. Two fundamental types of machine learning are supervised learning and unsupervised learning. While supervised learning [24,25] requires labeled data for training, unsupervised learning discovers patterns in unlabeled data, making it more applicable in scenarios where labeling is infeasible. Among these, supervised learning is the most widely utilized and successful approach. In supervised learning, algorithms are provided with a data set comprising pairs of input and output examples. The algorithm learns to map each input with its corresponding output, effectively associating input examples to output examples. A widely used form of supervised learning is classification. Once trained, supervised learning algorithms can achieve accuracy levels that exceed human performance, making them essential in various products and services. Despite these advancements, the learning process has limitations compared to human abilities. Current supervised learning approaches typically require millions of training examples [26]. To address these challenges, researchers are increasingly focusing on unsupervised learning, to reduce dependence on extensive human supervision and decrease the number of training examples needed. In general, the purpose of unsupervised learning is to extract meaningful information from a data set containing unlabeled input examples. Unlike supervised learning, unsupervised learning seeks to uncover useful patterns from unlabeled data [27,28]. Two well-known applications of unsupervised learning are clustering and dimensionality reduction.
A significant approach in unsupervised learning is generative modeling. Generative modeling aims to approximate the true data distribution
Formally, a GAN consists of two models: a generator
This adversarial process drives the generator to improve its outputs such that they are indistinguishable from real samples, while the discriminator becomes better at detection.
In general, the two neural networks: a generator and a discriminator, are designed to compete with each other [35]. Fig. 3 illustrates the structure of a GAN. The generator and discriminator architectures generally consist of multi-layer convolutional or fully connected layers. The generator learns the statistical properties of the training data and generates new images, while the discriminator evaluates and distinguishes between real and synthetic images [30]. Both networks serve as mappings between data domains [36]. The generator, without direct access to the real dataset, aims to create convincing synthetic images to deceive the discriminator. If the discriminator makes an incorrect classification, an error signal is generated to refine the generator’s output, progressively enhancing the quality of generated images. The generator transforms a latent space into the data space, while the discriminator maps image data to a probability score, indicating whether an image is real or synthetic. If the discriminator identifies an image as real, it outputs a value close to
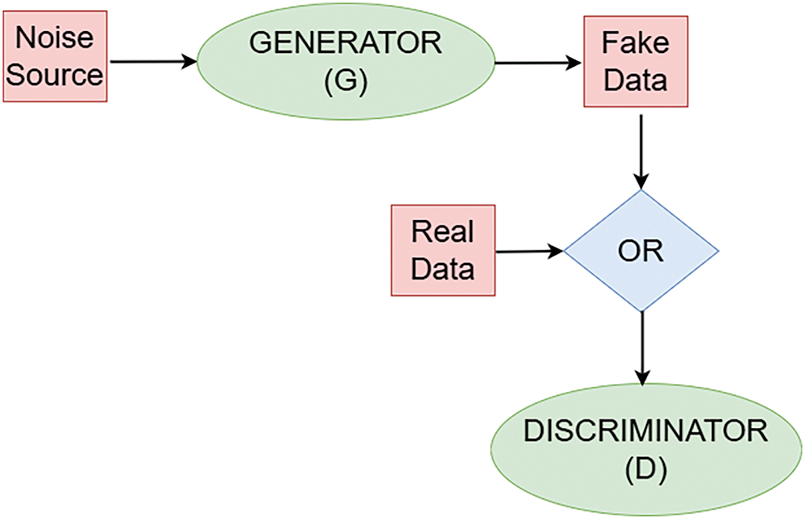
Figure 3: Block diagram of GAN
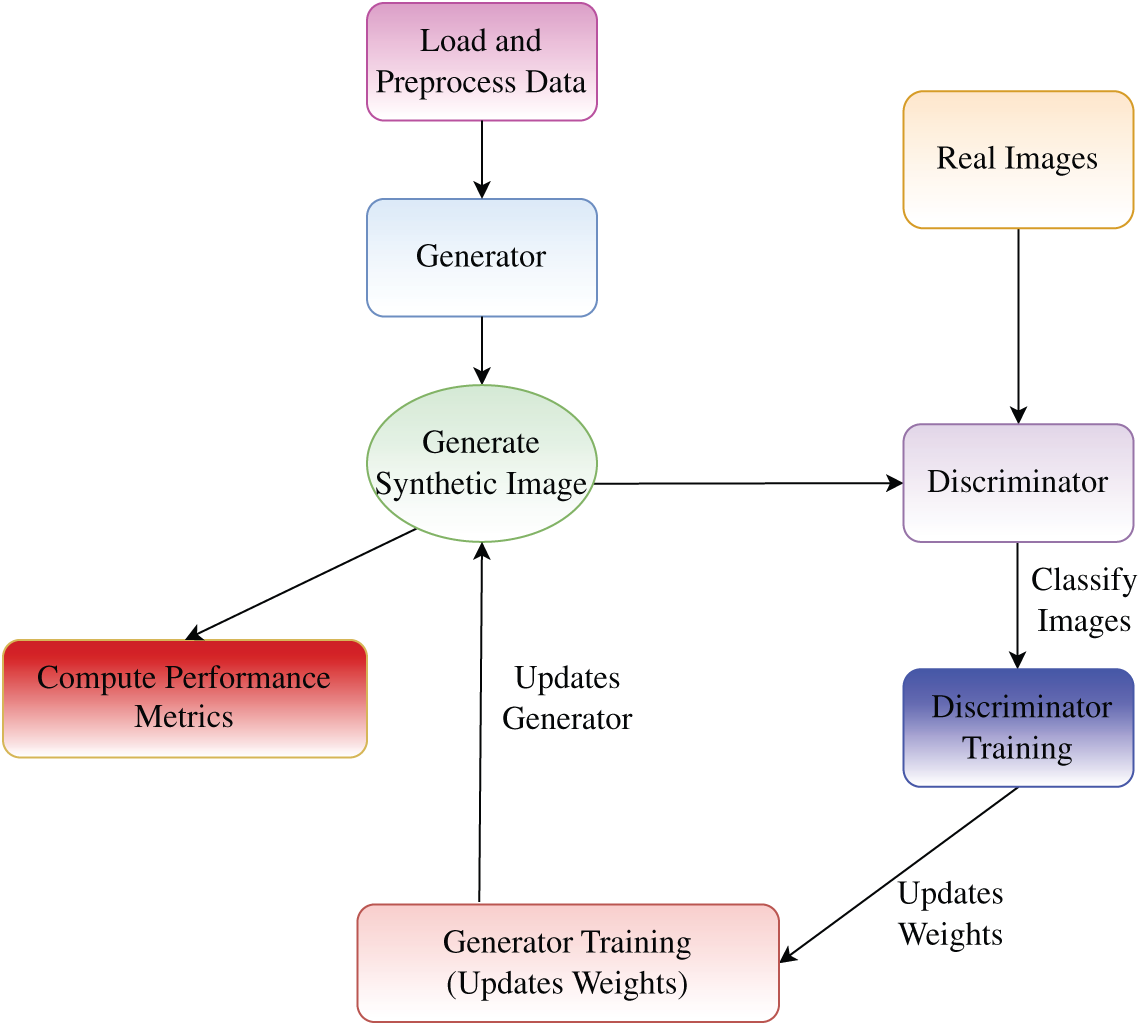
Figure 4: Flowchart of GAN training process
Over the years,
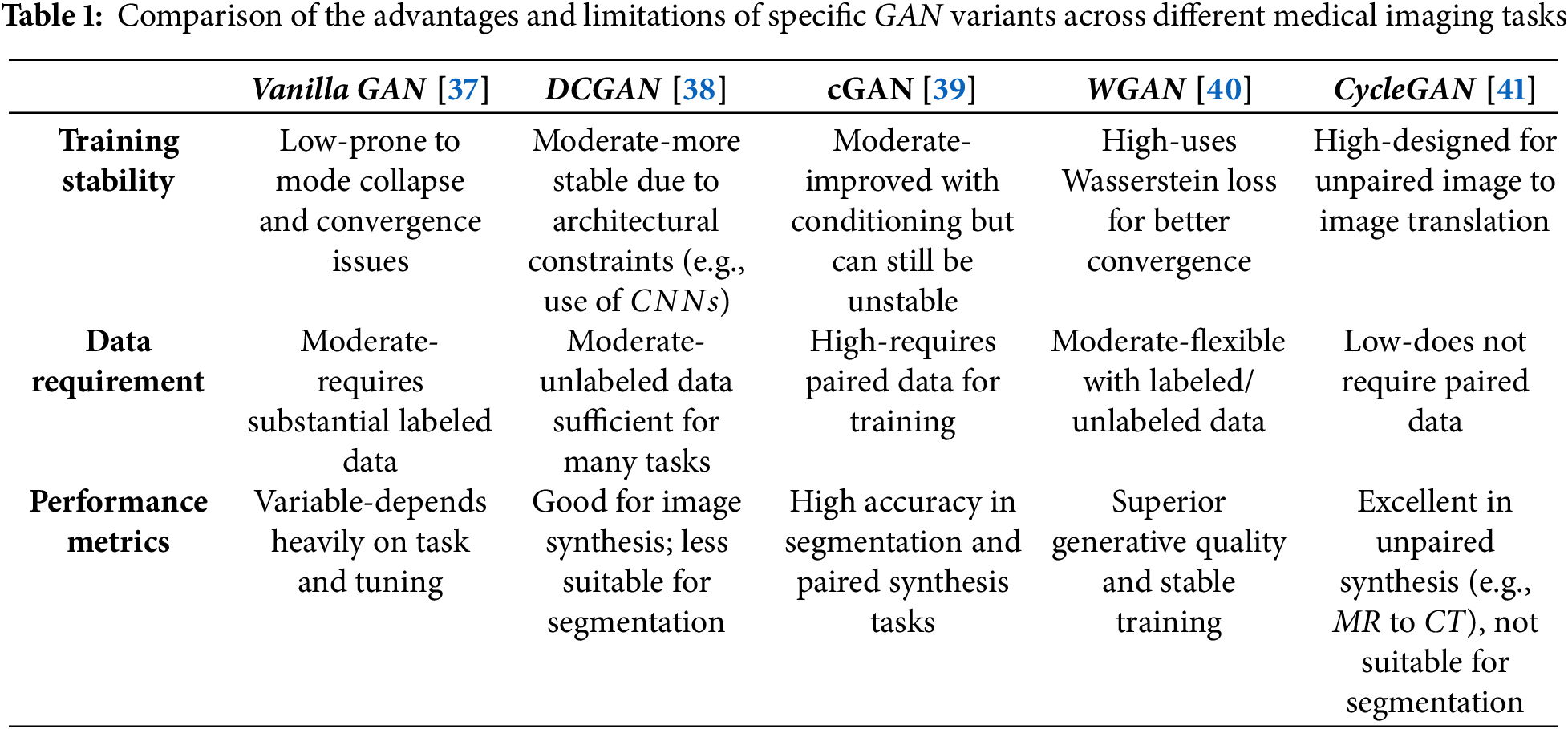
With the variety of GAN models available for different applications, the base model or its specific variants can be selected based on the task. For instance, DCGAN is suitable for image generation, SRGAN for super-resolution, U-Net-based
• Image Denoising: Medical images, such as low-dose CT scans or accelerated MRI, often have lower quality due to noise or reduced resolution.
• Image Segmentation: Segmentation is a crucial task in medical imaging, as it helps in determining and extracting areas of interest from medical images which has lately been well achieved using
• Image Super Resolution: Super-Resolution enhances the resolution thereby improving the clarity and detail of anatomical structures.
• Image Translation: Translation involves converting images from one modality to another.
• Image Reconstruction:
• Data Augmentation:
• Anomaly Detection: By learning the distribution of normal anatomical structures,
• Domain Adaptation: Variations in medical images resulting from differences in imaging devices, acquisition protocols, or healthcare institutions can affect model performance.
The use of
Ongoing research and technological progress continue to expand the scope and impact of
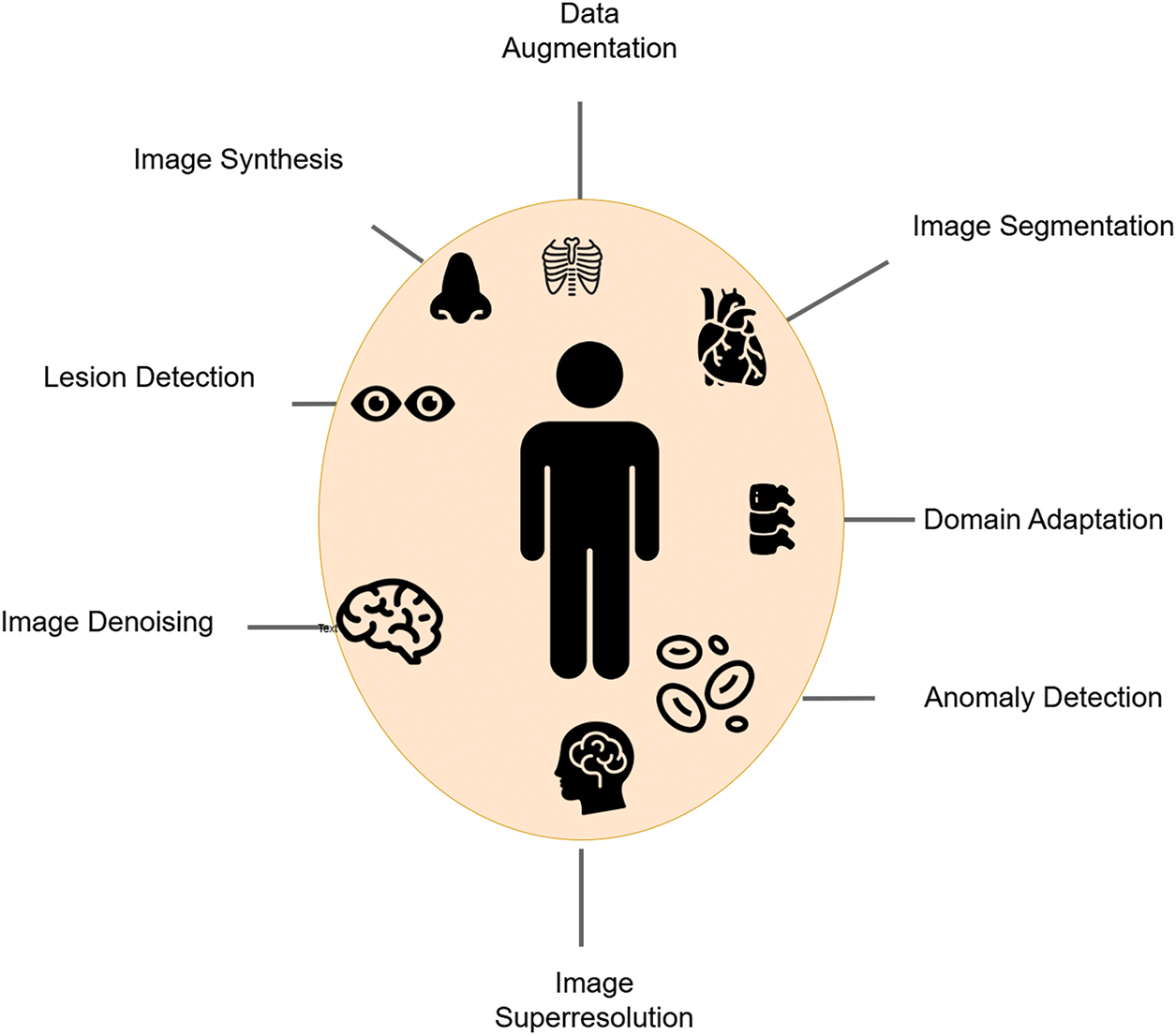
Figure 5: Applications of GAN in medical imaging
Despite their potential,
This research adopts a Systematic Literature Review
1. Databases Searched: The review targets scholarly, peer-reviewed articles sourced from well-established databases, including ScienceDirect, SpringerLink, the ACM Digital Library, IEEE Xplore, PubMed, Scopus and Web of Science. At the outset of this study, a total of
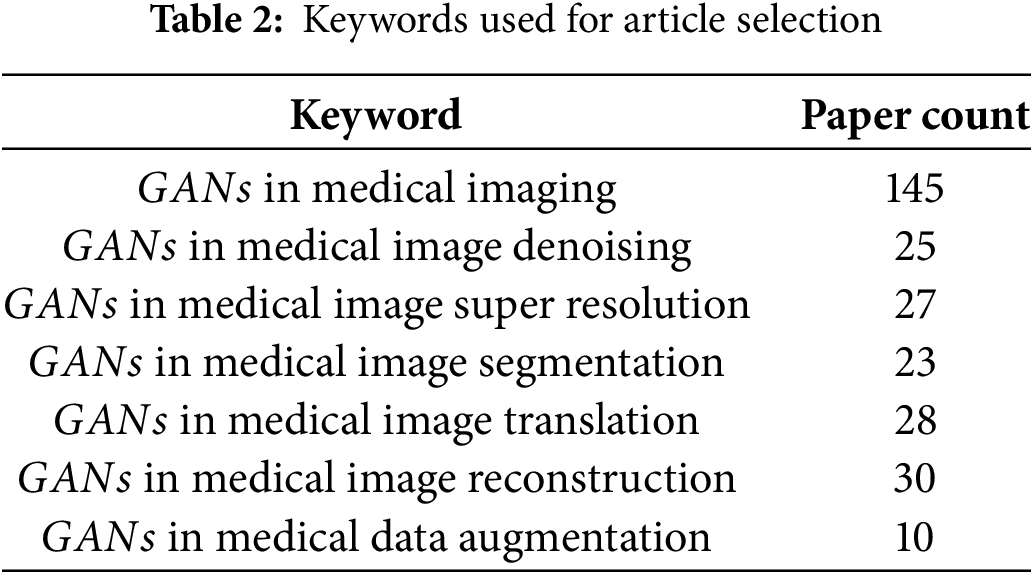
2. Search Terms and Keyword Strategy: To capture the full range of applications of
3. Boolean Operators and Search String Construction: Boolean operators were applied to systematically combine the two groups of keywords. The operator OR was used within each group to capture synonyms and variations, while the operator AND was used to combine the technology-related terms with the application-related terms. The following is an example of the search string used:
(“Generative Adversarial Networks” or “GAN” or “
This logic ensures that the search retrieves studies that discuss
4. Search Execution and Documentation: The search was performed using both keyword and
5. Screening Process: Following retrieval, titles and abstracts were screened independently by two reviewers. Full texts were then assessed for eligibility based on predefined inclusion and exclusion criteria. Table 3 indicates the inclusion and exclusion criteria used for selecting the articles.
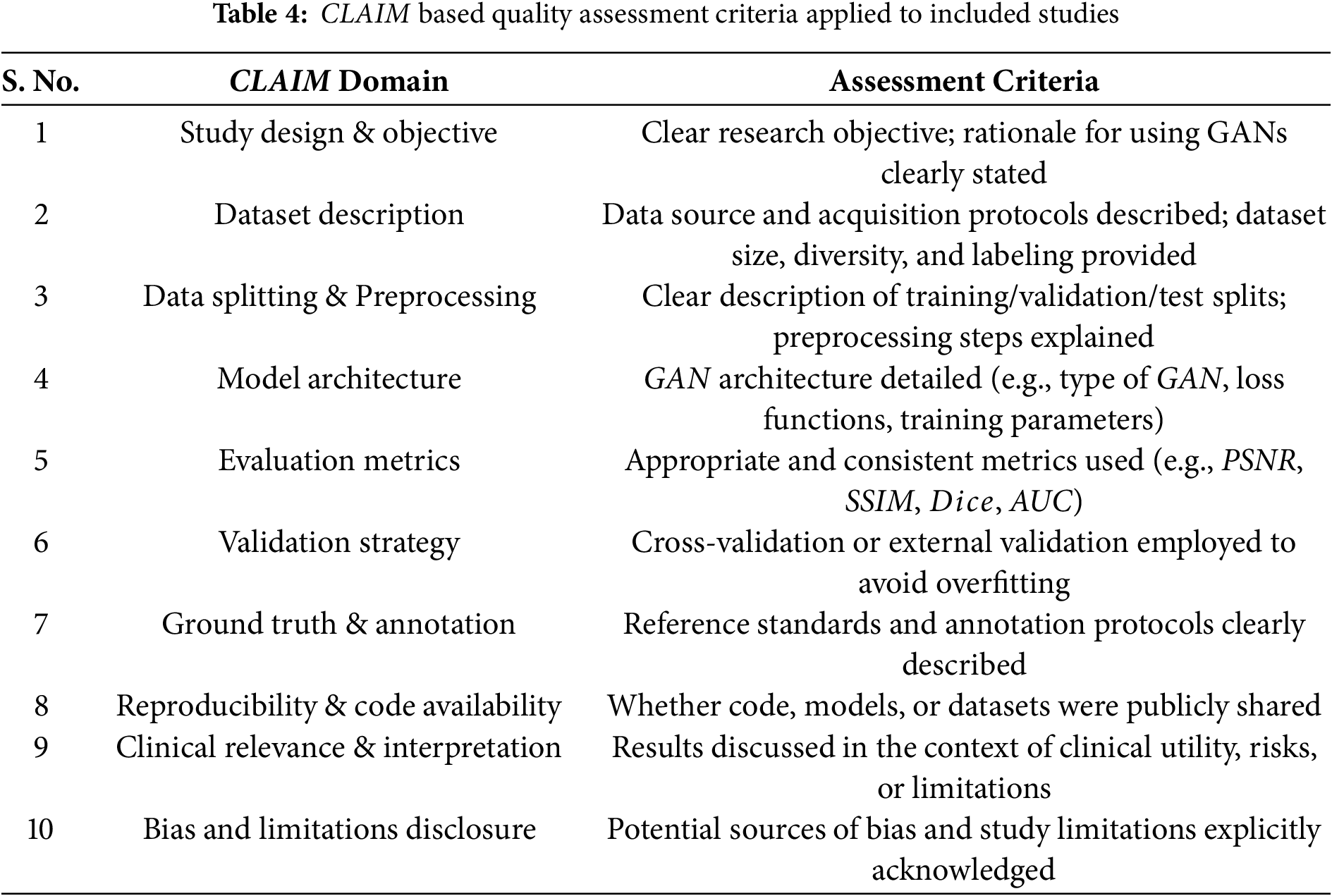
6. Quality Assessment Method: To assess the methodological quality and reporting transparency of included studies, the CLAIM (Checklist for Artificial Intelligence in Medical Imaging) guideline was used. This checklist evaluates critical domains relevant to AI studies, including dataset characteristics, model evaluation procedures, validation methodology, and reproducibility. Table 4 adapted from the CLAIM, indicates the criterion used to evaluate the methodological quality of included studies. Only studies meeting key CLAIM criteria were retained for final synthesis to ensure reliability of the review findings. Each study was assessed independently by two reviewers. Disagreements were resolved through discussion or by consulting a third reviewer.



This approach ensured a comprehensive, reproducible, and methodologically sound search process in alignment with PRISMA standards. The PRISMA flow diagram of the article selection procedure is shown in Fig. 6. PRISMA checklists can be found in the supplementary files.
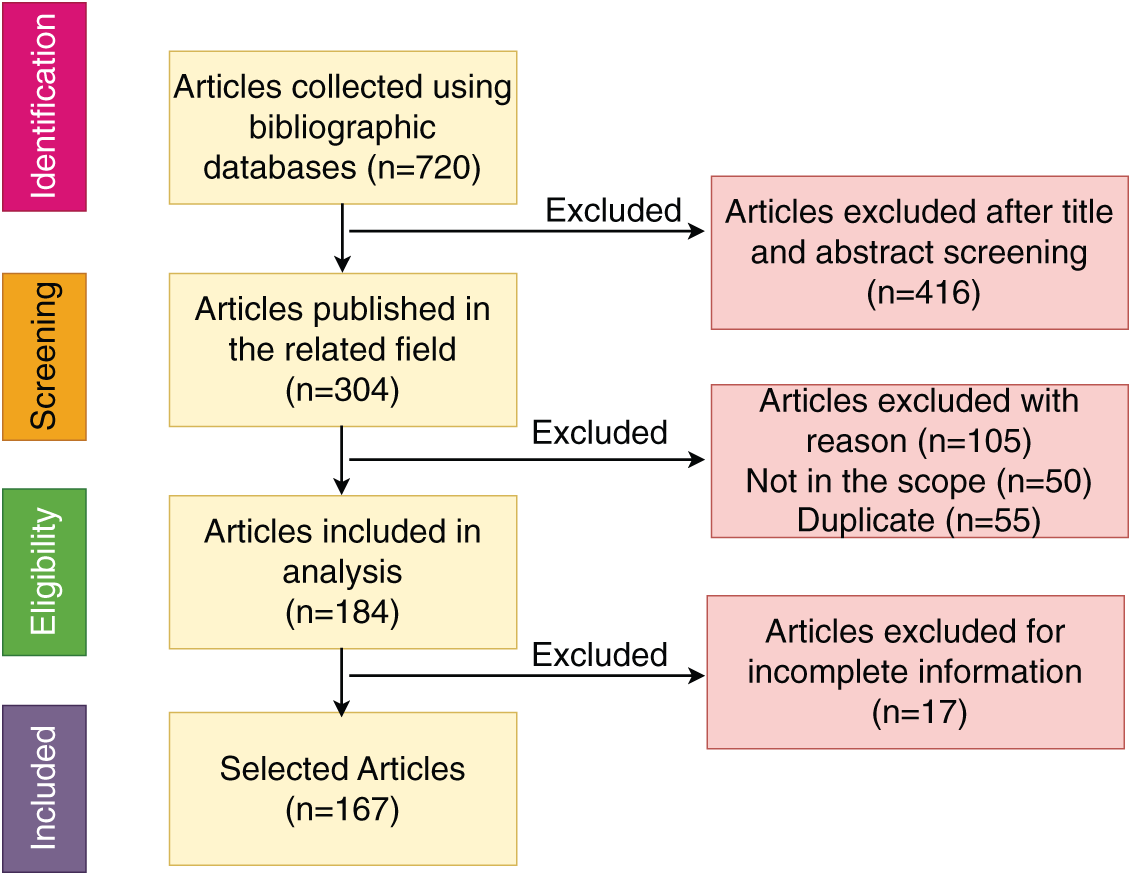
Figure 6: PRISMA flow diagram of the article selection procedure
The remaining sections of this paper explore in detail how
Medical imaging application uses GAN in two separate ways; one as generator which examines the underlying data distribution and generates new (synthetic) images. The discriminator section can classify normal and abnormal images. An overview of usage of
Image denoising is a vital preprocessing act in analysis, as all types of medical images are susceptible to noise [100–104]. The sources of noise in medical imaging can be categorized into sensor-related, acquisition-related, and radiation-related factors. Computed Tomography (CT) is a widely utilized technique for disease diagnosis, but it carries the potential risk of radiation exposure [105,106]. Reducing radiation levels can lead to increased noise in CT images. Reconstruction of Low-Dose CT (LDCT) images offers an effective approach to address this issue. Fig. 7 shows a GAN based framework for medical image denoising. A GAN utilizing Wasserstein distance (WGAN) and perceptual similarity was applied to denoise CT images in [94]. The perceptual loss minimized noise by aligning output and ground truth features in a defined space, while the GAN shifted the noise distribution from strong to weak. Wasserstein distance was used to compare the distributions of normal-dose CT (NDCT) and LDCT data. Feature extraction was performed using Convolutional Neural Networks (CNN) based on the Visual Geometry Group (VGG). Performance metrics such as PSNR and SSIM were employed to evaluate the outputs of different networks, with WGAN−VGG achieving superior performance. Huang et al. in [107] presented a denoising method (DU−GAN) which utilized U-Net-based discriminators to assess the global and local variations in the denoised and normal images. A denoising method based on Conditional GAN was popularized by Li et al. in [53] in which the image context relationship and structural information was preserved. The method was tested on LIDC dataset and was seen to outperform the state of the art works. Zhu et al. had introduced a denoising method based on GAN in [108]. Molecular activity in human tissues was captured using Single-Photon Emission Computed Tomography (SPECT), which relies on gamma rays for image acquisition in [91]. High-noise SPECT images are input into the generator, while the discriminator assesses the generated images against real samples, specifically low-noise SPECT images. The loss, which quantifies the difference between generated and real images, is used to update both the generator and discriminator simultaneously. Both components were optimized using the Adam optimizer, with a learning rate of
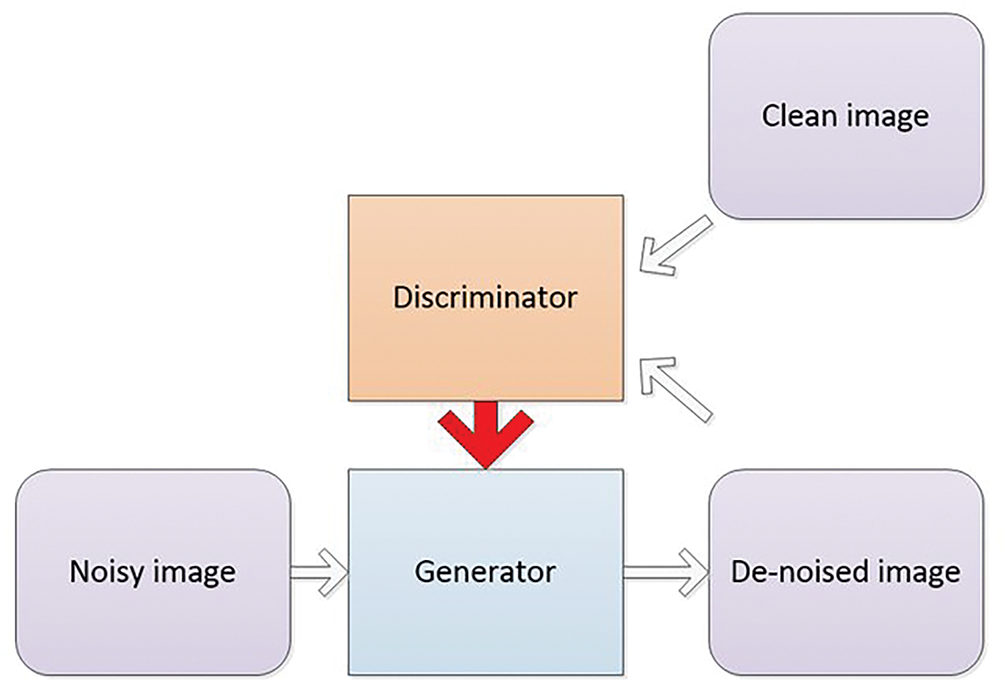
Figure 7: GAN-based framework for medical image denoising: enhancing image quality with adversarial training
Segmentation plays a crucial role in medical image analysis. Automating the segmentation process is highly challenging due to variations in anatomical structures across different patients [55,109–111]. Skin cancer, common among individuals with fair skin, is classified into melanoma (pigmented lesions) and non-melanoma (non-pigmented lesions). Early detection of melanoma is critical. Dermoscopic images captured via smartphones can be analyzed to detect pigmented lesions. A GAN was trained using
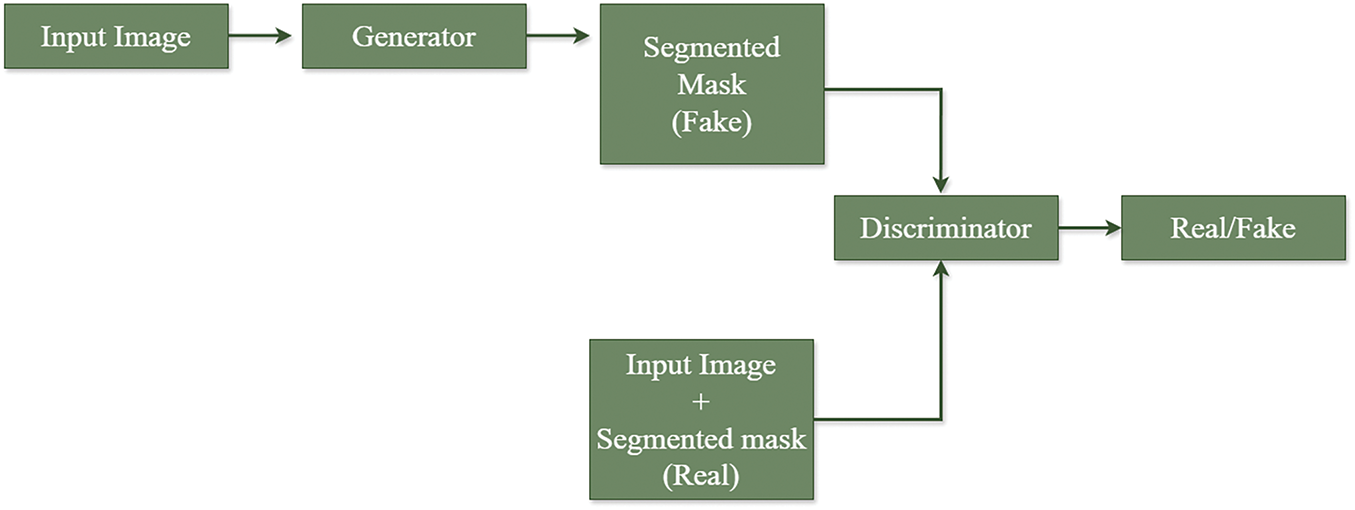
Figure 8: Framework for medical image segmentation using GAN for lesion, infection detection, etc.
Fig. 8 shows a framework for medical image segmentation using GAN. A healthy immune system defends against foreign bodies in the body. In autoimmune disorders, the immune system attacks healthy tissues. Human epithelial type
The neural network architecture proposed in [113] is composed of two main components: a Segmentor and a Critic. To enhance semantic feature extraction, a novel model named
High-resolution images play a crucial role in accurate disease diagnosis. Despite advancements in medical imaging techniques, factors such as imaging conditions, equipment limitations, and environmental obstructions can result in low-resolution images [115–119]. Resolution can be improved by various methods such as enhancing the spatial resolution, interpolation, multi image super resolution methods. MR image resolution can be improved using GAN technique by taking advantage of the volumetric information in the
Histopathology is the study of presence of disease in biopsy or surgical samples using microscopes [121]. Histopathological images provides a comprehensive view of cell tissues. Different segments in a tissue are visualized by pigmenting it using different dyes. Histopathological images analysis involves segmentation, detection, feature extraction, classification, etc. A unified GAN framework with a newly designed loss function was introduced for the same in [96]. The loss function is derived from both WGAN with gradient penalty (WGAN−GP) and Information Maximizing Generative Adversarial Networks (
Diabetic retinopathy, glaucoma are the diseases which are diagnosed with the help of retinal fundus images. Retinal fundus image resolution are not good enough to detect microaneurysms, hemorrhages as they cover small image areas. Progressive GAN (P−GAN) generated a high resolution image from a low resolution input image. The proposed P−GAN architecture incorporated two stages-stage
X-Ray Computed Tomography (X-Ray CT) uses X-Rays for screening, diagnosis, image guided surgery, etc. The quality of X-Ray CT images can be improved in two ways-either by using a good quality hardware or by computationally enhancing the images. The former is not economical and also are of high radiations [123]. High radiations cause gene damages and can cause cancer. Low Dose CT (LDCT) uses lower radiation, which results in low quality CT images. Computational techniques can be employed to obtain High Resolution CT (HRCT) from LDCT. A novel residual CNN-based network in the
• cycle consistency to ensures strong across domain consistency between LRCT and HRCT,
• exclusion of Nash equilibrium [30] problem for training GAN,
• omission of over fitting problem by optimizing the network,
• Inclusion of multiple cascaded layers to extract hierarchical features,
• to enhance deblurring
The main challenges in recovering HRCT images from noisy LRCT images are-complex spatial variations in the images, presence of unique noise patterns, sampling and degradation makes the image blur. To handle these limitations, non linear SR functional blocks with residual module is included in the proposed framework, which have the ability to learn high frequency details. Adversarial learning in a cyclic manner is followed, which ensures superior quality CT images. The proposed network has two generators—G & F, with feature extraction network and reconstruction network. Reconstruction network has a Parallelized CNN with multi layer perceptron (MLP) to carry out nonlinear projection in the spatial domain. Parallelized CNN are network within network which can perform dimensionality reduction with faster computation and less information loss, able to learn the complex mapping at finer levels with better accuracy. The framework has been trained and tested using two datasets-twenty five images from Tibia dataset and
A deep learning approach named
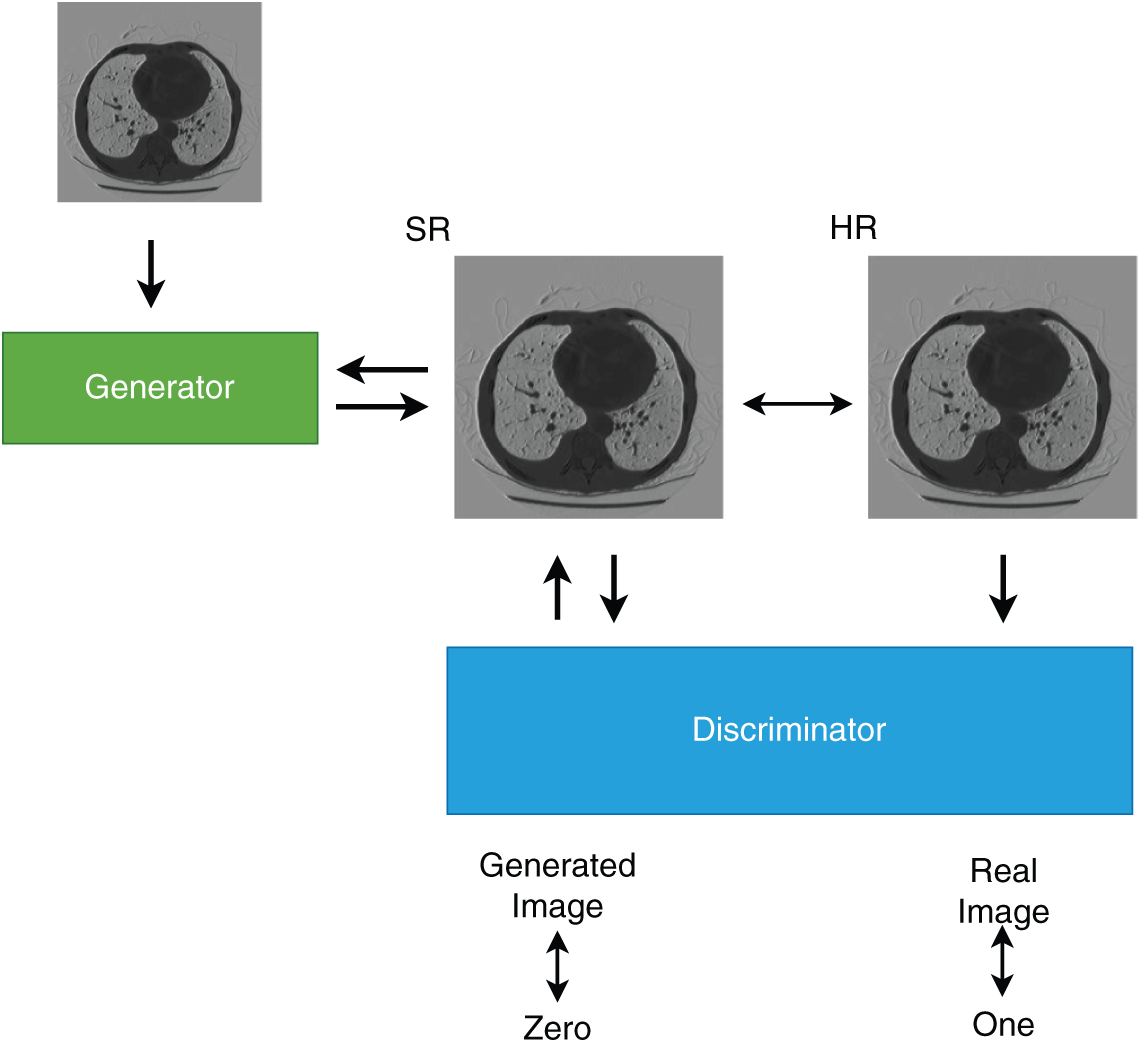
Figure 9: Architecture of
Degraded images are the major drawback of portable imaging devices. Spatial resolution, contrast and noise are the three issues related with ultrasound images. Portable devices produce low resolution, low contrast and high noise images, which makes the disease diagnosis inaccurate. GAN based method provides promising results for resolution enhancement with the following advantages-non linear multi level mapping between LR and HR images, adaptive feature extraction without human intervention, image quality enhancement using discriminator D, direct and efficient one step feed forward reconstruction procedure and easier implementation on hardware like
Disease diagnosis and treatment with single image modality may not be adequate in most of the cases. The images acquired cannot outline the complete anatomical details or fails to acquire the details with the desired imaging modality [85,90,130–132]. Image translation is an optimum solution, where required image is synthesized from a different image modality, without inducing much cost or risk. The challenge involved in translating one image modality to another is the presence of unrealistic data in the output image [133]. Fully supervised learning methods are among the most widely used deep learning approaches for this task [134]. However, these methods require paired low- and high-quality images for training, which is especially challenging in medical imaging, where obtaining such aligned image pairs is difficult in real-world scenarios. To address this limitation, several unsupervised learning frameworks have been developed [132,135]. Despite their potential, these approaches often face issues such as instability, noise amplification, and the occurrence of halo artifacts. A well-known solution for unpaired image-to-image translation is the Cycle-consistent Generative Adversarial Network
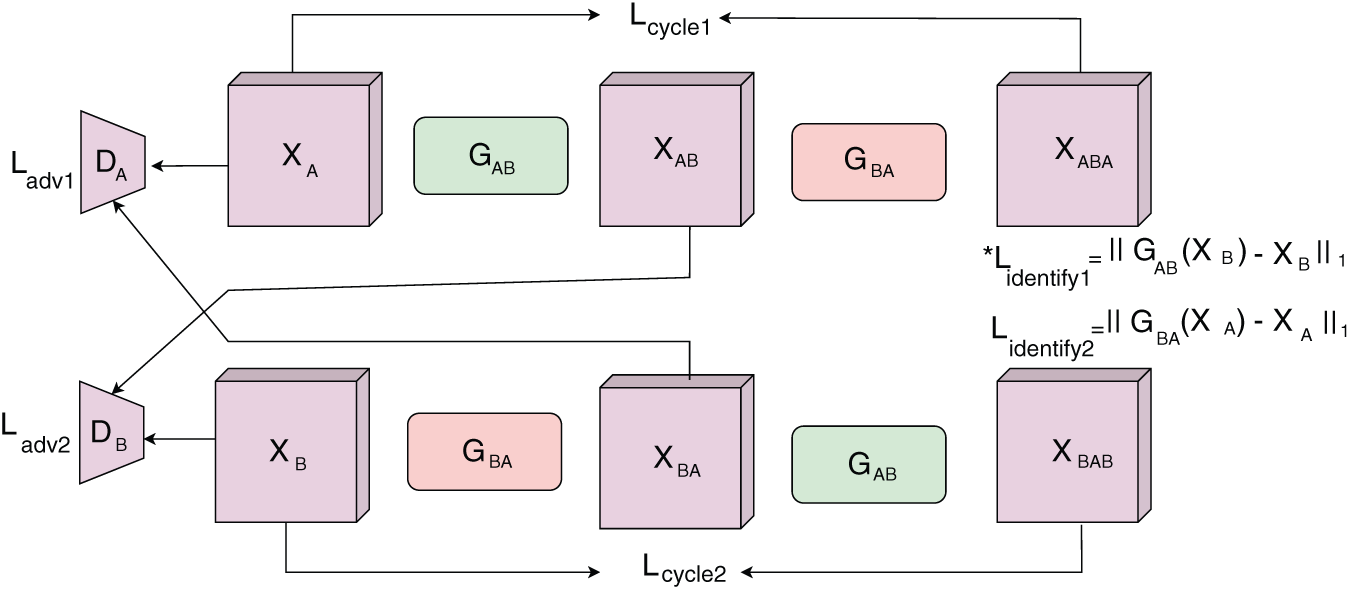
Figure 10: The
A skin lesion synthesizer based on GAN was proposed in [98], having a coarse to fine generator, multi scale discriminator and a robust objective function for learning. This method synthesized images from a semantic label map and an instance map [60]. Images of resolution
Yang et al. in [9] introduced a method to perform Image Modality Translation
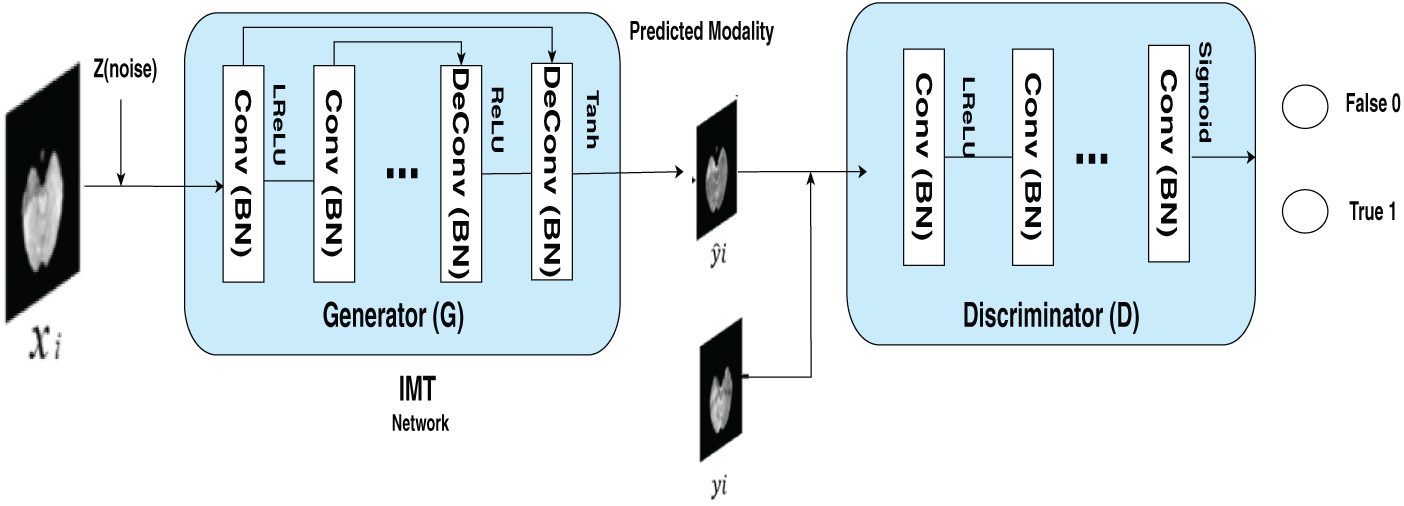
Figure 11: The figure outlines the structure of an end-to-end IMT network designed for cross-modality image generation. The training dataset is defined as
A novel GAN architecture named
Chen et al. in [139] introduced MI−GAN, a novel multi-domain medical image translation algorithm that incorporates a key transfer branch. By analyzing the imbalance present in medical imaging datasets, the approach identified critical target domain images and constructed a specialized transfer branch. Utilizing a single generator, the method facilitates multi-domain image translation in the medical context. This structure enhanced both the model’s attention mechanism and the quality of the generated images. Additionally, a lung image classification model was presented, leveraging synthetic image data for augmentation. The training dataset combines both synthetic lung CT scans and original real-world images to evaluate the effectiveness of the model in diagnosing normal individuals, as well as patients with mild and severe cases of
Medical image reconstruction is a crucial process for generating high-quality images needed for accurate analysis. However, the quality of these images is often affected by noise and artifacts [141–143]. To address these limitations, there has been a paradigm shift from traditional analytical and iterative reconstruction methods to data-driven machine learning approaches [144,145].
A review of the literature reveals that frameworks like
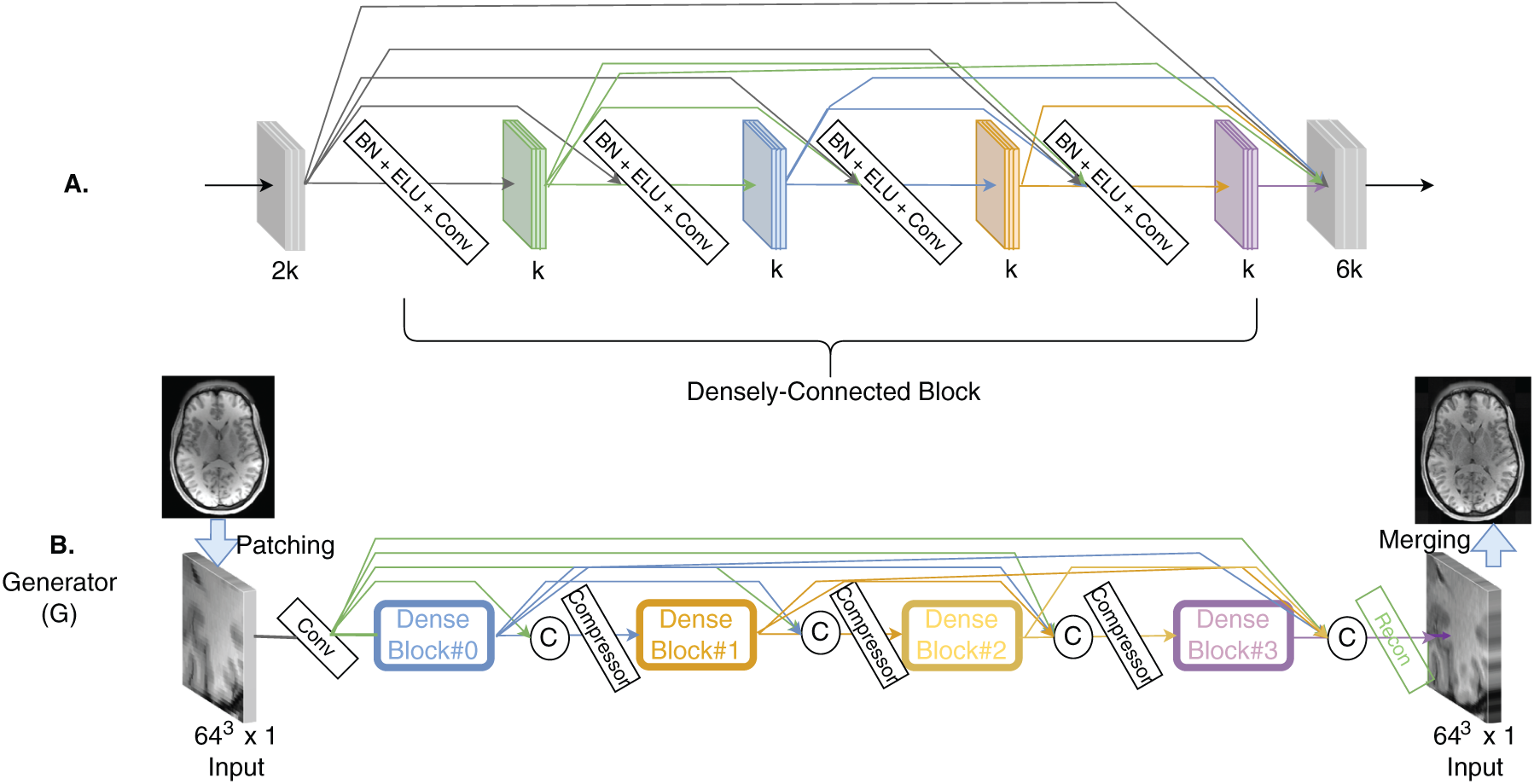
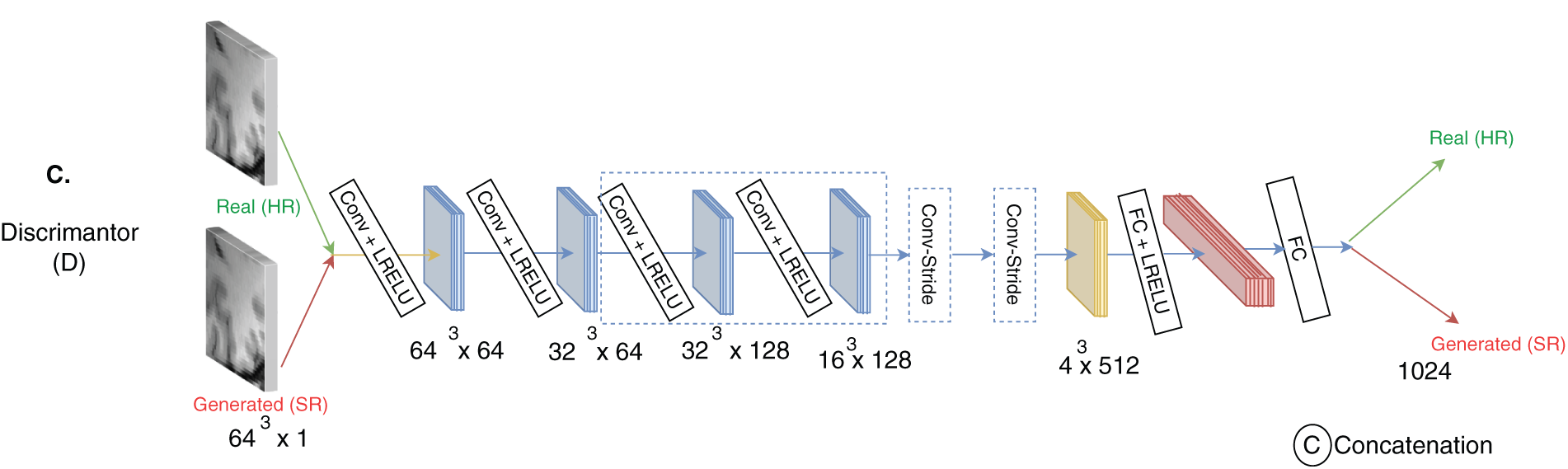
Figure 12: The architecture consists of (A) a
Additionally, conditional
Ahn et al. in [152] utilized
A novel framework named
Synthetic pterygium images were produced using the default configuration of the
The discriminator also followed the
Despite the architectural strengths of
These outcomes suggest that
Understanding how diseases evolve over time is essential for early detection and effective treatment planning. This is particularly important for severe conditions like Idiopathic Pulmonary Fibrosis
Deep learning models typically require large datasets, making it difficult to apply them in situations where limited data is available. One common solution is data augmentation, which involves creating training examples by generating new data. This approach involves basic techniques like random rotations, flipping, cropping, and adding noise. However, such transformations often fall short when applied to complex datasets like medical images. To address this, researchers have developed more strategies for medical imaging. The primary aim is to produce synthetic data that closely mirrors the original distribution. The emergence of Generative Adversarial Networks
GAN based data augmentation involves training a generator network to produce synthetic images from a latent space, thereby increasing the dataset’s diversity and variability beyond simple transformations. This method is especially advantageous in situations with limited labeled data or class imbalance, where creating additional samples of underrepresented classes can greatly enhance classifier performance [156]. For instance, in medical image analysis,
A data augmentation technique was presented by Frid et al. in [157] data augmentation approach that integrates traditional image perturbation techniques with the generation of synthetic liver lesions using Generative Adversarial Networks
Gan et al. introduced a generative adversarial network
Furthermore, datasets augmented with
In medical imaging applications of
Key datasets such as OASIS, SRPBS, and ABIDE are frequently employed in these studies, highlighting their critical role in advancing research within the field. Their extensive use demonstrates their value in bench marking and evaluating algorithm performance across a range of medical imaging applications. Ultimately, the diverse nature of medical imaging tasks requires a strategic approach to selecting algorithms. While certain GAN models excel in specific areas, the variation in results underscores the need for task-specific algorithm selection. This tailored approach is essential for advancing the capabilities and accuracy of medical image analysis.
4 Challenges, Ethics and Future Research Directions of GAN for Medical Images
Although
4.1 The Non Convergence Problem
In
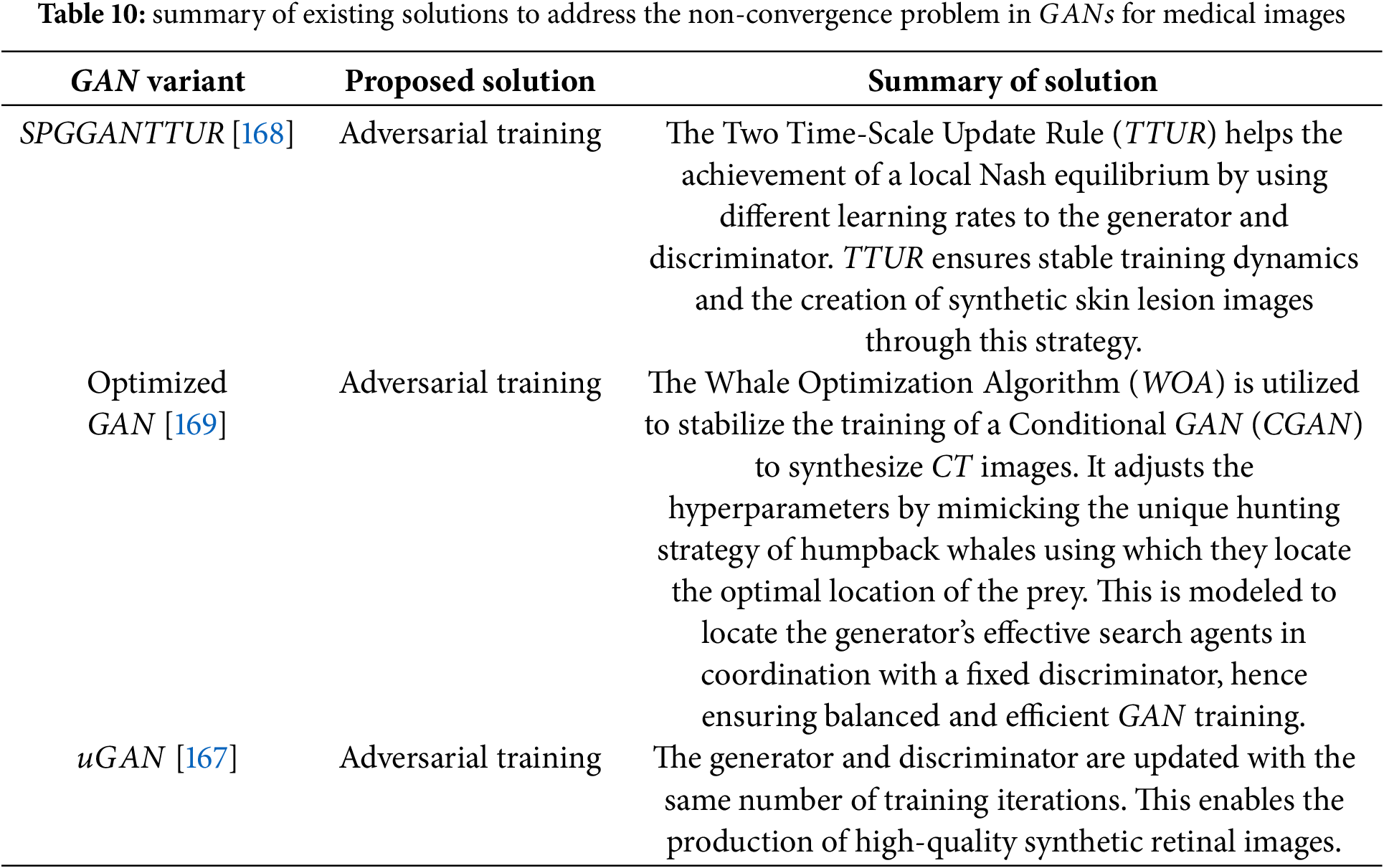
The issue of non-convergence is a significant challenge encountered during the training of
• optimization of update algorithms [167]
• adversarial learning [168]
• tuning of hyperparameters [169].
4.1.1 Optimization of Update Algorithms
The evolution of updating algorithms has been examined across different GAN architectures, including the original Vanilla GAN [42], the Wasserstein GAN (WGAN) introduced by [40], and the more recent
Achieving balance in GAN training is closely linked to adjusting the learning rates of the generator and discriminator. This approach was adopted by [168] to mitigate non-convergence issues in biomedical image synthesis. The underlying concept of stabilizing GAN training through learning rate control was originally proposed by [171], who introduced the Two Time-scale Update Rule (TTUR). TTUR employs separate learning rates for the generator and discriminator, enabling the model to approach a local Nash equilibrium without relying on multiple update steps. In their study, Abdelhalim et al. incorporated both TTUR and a custom discriminator update strategy into the SPGGAN framework for synthesizing skin lesion images. Specifically, they updated the discriminator five times for each generator update, promoting greater training stability. This adjustment aimed to slow down discriminator learning just enough to allow the generator to keep pace and improve image quality without mode collapse.
4.1.3 Tuning of Hyperparameters
Selecting suitable hyperparameters for controlling the generator and discriminator in
• Encircling strategy: The lead whale locates the prey and simulates encircling it. Analogously, the generator’s candidate solutions (search agents) evaluate a fitness function during each iteration and refine their positions accordingly.
• Distance-based updating: The proximity between the prey (optimal solution) and each search agent is calculated, and agent positions are adjusted based on this measure.
• Exploration through random search: Unlike the first rule which focuses on the best-known position, this rule updates the agents’ positions based on a randomized strategy to encourage exploration of the solution space.
The use of this optimized strategy enhances both the generator’s performance and the discriminator’s ability to distinguish between real and synthetic images. As a result, the model achieves adaptive loss tuning, leading to the generation of higher-quality and more diverse images. In terms of performance, the optimized GAN outperformed the baseline CGAN in classification tasks using the synthesized and original images. Specifically, it achieved an
Researchers have explored the use of Jensen-Shannon

4.2 Mode Collapse & Hallucinated Features
Mode collapse represents a significant challenge during the training of
The mode collapse problem can be alleviated by using different methods such as
• regularization
• modified architectures
• adversarial training.
In deep learning, minimizing the loss function is a primary objective; however, achieving this becomes difficult when the model contains excessively large weight values. Large weights can lead to overfitting, where the model performs well on training data but generalizes poorly to new, unseen data. To counteract this, regularization techniques are employed to constrain the size of the weights or limit the overall capacity of the model [42].
In the context of
1. Spectral Normalization [177]
2. Batch Normalization [38]
3. Self-Normalization [178]
Among these, spectral normalization has proven particularly effective in stabilizing GAN training by controlling the Lipschitz constant of the discriminator network.
Xu et al. [179] addressed the problem of mode collapse in
In the context of
Creating segmentation masks and corresponding ground truth images separately using
4.3 Metrics for Quantitative Evaluation
In addressing the training challenges of
4.4 Privacy Concerns and Ethics
Medical imaging concerns handling highly sensitive patient data, which raises important privacy concerns during both data collection and application. While
Federated learning presents a promising approach to reduce privacy concerns while using
Fig. 13 gives an illustration of the steps in federated learning. In the first phase, each participant independently calculates the model gradients on their local data. To ensure data confidentiality, cryptographic methods such as homomorphic encryption are applied before transmitting the encrypted gradients to the central server. In the second phase, the central (master) server performs secure aggregation of the encrypted gradients. In the third phase, the aggregated results are sent back to the participants. During the fourth phase, each participant decrypts the aggregated gradients and updates their local model accordingly. This cycle is repeated iteratively until either the loss function reaches convergence or a predefined number of iterations is completed. Throughout this process, the participants’ data remains stored locally, maintaining privacy and offering an advantage over centralized approaches such as those based on Hadoop. Federated learning facilitates collaborative model training across multiple databases without the need to centralize the data, enabling scalability with growing datasets while minimizing communication overhead, as only gradients—not raw data—are shared.
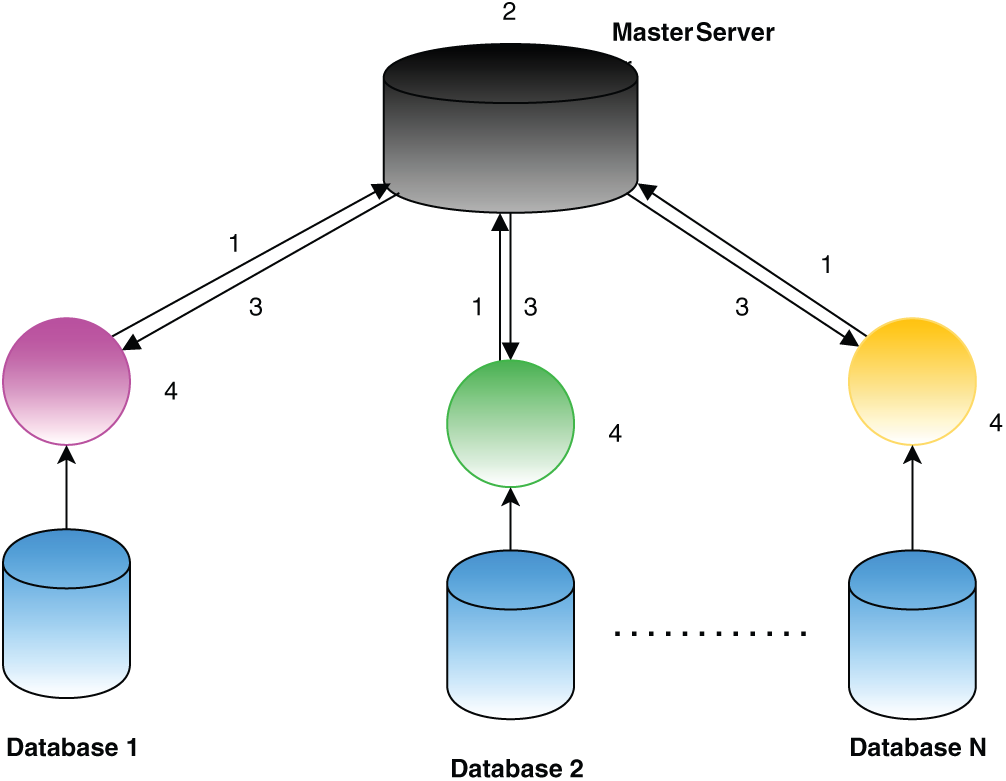
Figure 13: Steps in federated learning [165]
4.5 Need for Human in the Loop Studies
GAN based image generation models can produce photorealistic images; however, in medical applications, caution is essential as these images play a critical role in disease detection and diagnosis. It is imperative for medical professionals, as the end users, to thoroughly evaluate and validate GAN generated images to ensure their reliability and clinical utility. Trust and acceptance from doctors are crucial for integrating such models into healthcare workflows. While PSNR and SSIM are standard for evaluating image processing techniques [192], they often fail to reflect perceptual quality or clinical relevance, particularly in the presence of subtle distortions. This underscores the importance of expert-driven evaluation, as automated metrics alone may not capture diagnostic integrity. In medical imaging, radiologists and clinicians are best positioned to perform this qualitative assessment, as their expertise ensures that reconstructed images are not only visually plausible but also diagnostically accurate.
Realistic full-field digital mammograms were generated using a progressive GAN architecture [193], achieving high resolution and realism indistinguishable from real images, even by domain experts. Despite the specialized nature of medical imaging, both experts and non-experts in a reader study showed random success probabilities, emphasizing the critical role of human validation in ensuring clinical applicability. Similarly, the progressive growing GAN (PGGAN) [194] was employed to generate high-resolution chest radiographs (
Further advancing GAN applications, GANCS [195] presents a compressed sensing framework that models the low-dimensional manifold of high-quality MR images using a least-squares GAN (LSGAN) to capture fine textures, combined with
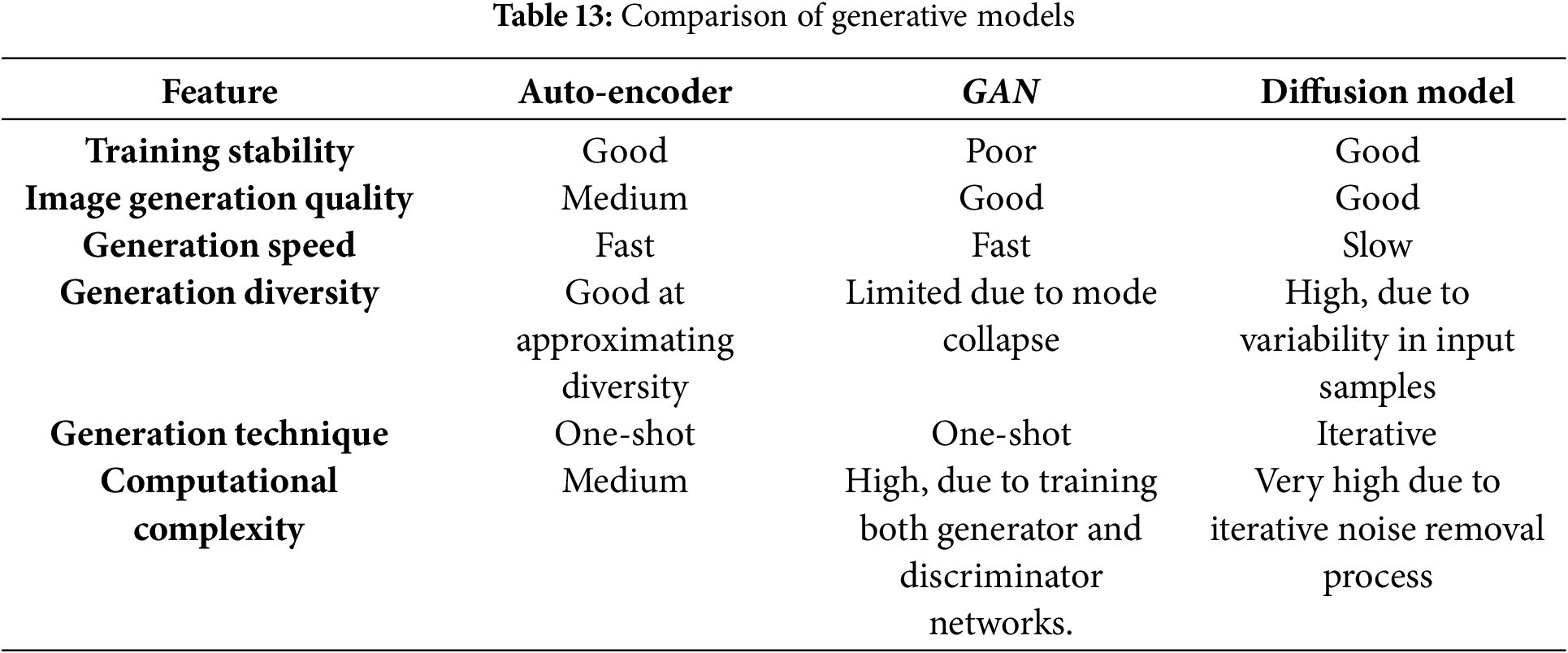
This section provides an insight into various generative models, including auto-encoders and diffusion models, each tailored for specific tasks in data generation and representation learning. The auto-encoder [197] has encoder–decoder network maps input data to a low-dimensional latent space, enabling the decoder to accurately reconstruct the input. This latent space also facilitates systematic analysis and manipulation of input properties, making the architecture essential for biomedical tasks like image reconstruction, data augmentation, and modality transfer. Diffusion models, a class of deep generative models [198], learn the prior probability distribution of images (e.g., brain PET or cardiac MRI) from training data and generate new samples by sampling from this distribution. Recently, they have emerged as state-of-the-art in generative modelling, producing higher-fidelity samples compared to auto-encoders and normalizing flows. The comparison between the generative models are shown in Table 13. Diffusion models are extensively applied in medical image processing tasks such as reconstruction, registration, classification, image-to-image translation, segmentation, denoising, and image generation. A detailed explanation regarding this is discussed in the following sub-section.
Diffusion Models
Emerging as promising alternatives to
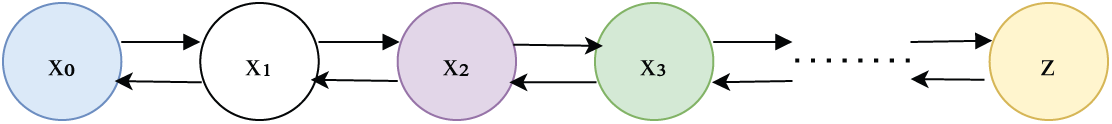
Figure 14: Diffusion model [165]
Generative Adversarial Networks
Despite the advancements in
Looking to the future, research should focus on building more stable and explainable GAN architectures tailored specifically to medical applications. Developing robust validation frameworks that incorporate expert clinical feedback will be vital to ensuring safety and effectiveness. Future studies should focus on interpretability, reliability, and adherence to clinical and regulatory standards to confirm the practical applicability of GAN based tools in healthcare. There is also potential for
Acknowledgement: The authors extend their appreciation to the Deanship of Research and Graduate Studies at King Khalid University for funding this work through Large Research Project under grant number RGP2/540/46.
Funding Statement: The research was supported by Deanship of Research and Graduate Studies at King Khalid University for funding this work through Large Research Project under grant number RGP2/540/46.
Author Contributions: The authors confirm contribution to the paper as follows: Conceptualization, Sameera V. Mohd Sagheer and U. Nimitha; methodology, P. M. Ameer; software, Sameera V. Mohd Sagheer; validation, Sameera V. Mohd Sagheer, U. Nimitha and P. M. Ameer; formal analysis, P. M. Ameer; investigation, Muneer Parayangat; resources, Mohamed Abbas; writing—original draft preparation, Sameera V. Mohd Sagheer; writing—review and editing, U. Nimitha; visualization, P. M. Ameer; supervision, Muneer Parayangat; project administration, Mohamed Abbas; funding acquisition, Mohamed Abbas and Krishna Prakash Arunachalam. All authors reviewed the results and approved the final version of the manuscript.
Availability of Data and Materials: Data openly available in a public repository.
Ethics Approval: Not applicable.
Conflicts of Interest: The authors declare no conflicts of interest to report regarding the present study.
Supplementary Materials: The supplementary material is available online at https://www.techscience.com/doi/10.32604/cmes.2025.067108/s1.
References
1. Yu Y, Feng T, Qiu H, Gu Y, Chen Q, Zuo C, et al. Simultaneous photoacoustic and ultrasound imaging: a review. Ultrasonics. 2024;139:107277. doi:10.1016/j.ultras.2024.107277. [Google Scholar] [PubMed] [CrossRef]
2. Dangoury S, marghichi El M, Sadik M, Fail A. The design of an efficient low-cost FPGA-based unit for generation ultrasound beamforming. Pertanika J Sci Technol. 2023;31(6):3077–92. doi:10.47836/pjst.31.6.24. [Google Scholar] [CrossRef]
3. Ali M, Magee D, Dasgupta U. Signal processing overview of ultrasound systems for medical imaging. In: White paper. Vol. SPRAB12. Dallas, TX, USA: Texas Instruments; 2008. [Google Scholar]
4. Tao Z, Tagare HD, Beaty JD. Evaluation of four probability distribution models for speckle in clinical cardiac ultrasound images. IEEE Transact Med Imag. 2006;25(11):1483–91. doi:10.1109/tmi.2006.881376. [Google Scholar] [PubMed] [CrossRef]
5. Weng L, Reid JM, Shankar PM, Soetanto K. Ultrasound speckle analysis based on the K distribution. J Acoust Soc America. 1991;89(6):2992–5. doi:10.1121/1.400818. [Google Scholar] [PubMed] [CrossRef]
6. Sagheer SVM, George SN. An approach for despeckling a sequence of ultrasound images based on statistical analysis. Sens Imag. 2017;18(1):29. doi:10.1007/s11220-017-0181-8. [Google Scholar] [CrossRef]
7. Henkelman RM. Measurement of signal intensities in the presence of noise in MR images. Medical Physics. 1985;12(2):232–3. doi:10.1118/1.595711. [Google Scholar] [PubMed] [CrossRef]
8. Zeng Y, Zhu J, Wang J, Parasuraman P, Busi S, Nauli SM, et al. Functional probes for cardiovascular molecular imaging. Quantit Ima Med Surg. 2018;8(8):838. doi:10.21037/qims.2018.09.19. [Google Scholar] [PubMed] [CrossRef]
9. Yang Q, Li N, Zhao Z, Fan X, Chang EIC, Xu Y. MRI cross-modality image-to-image translation. Scient Rep. 2020;10(1):3753. doi:10.1038/s41598-020-60520-6. [Google Scholar] [PubMed] [CrossRef]
10. Mredhula L, Dorairangasamy M. An extensive review of significant researches on medical image denoising techniques. Int J Comput Applicat. 2013;64(14):1–12. doi:10.5120/10699-1551. [Google Scholar] [CrossRef]
11. Ding Q, Long Y, Zhang X, Fessler JA. Statistical image reconstruction using mixed Poisson-Gaussian noise model for X-ray CT. arXiv:1801.09533. 2018. [Google Scholar]
12. Borsdorf A, Raupach R, Flohr T, Hornegger J. Wavelet based noise reduction in CT-images using correlation analysis. IEEE Transact Med Imag. 2008;27(12):1685–703. doi:10.1109/tmi.2008.923983. [Google Scholar] [PubMed] [CrossRef]
13. Rahiman MF, Rahim RA, Zakaria Z. Design and modelling of ultrasonic tomography for two-component high-acoustic impedance mixture. Sens Actuat A: Phys. 2008;147(2):409–14. doi:10.1016/j.sna.2008.05.024. [Google Scholar] [CrossRef]
14. Chen Y, Shi L, Feng Q, Yang J, Shu H, Luo L, et al. Artifact suppressed dictionary learning for low-dose CT image processing. IEEE Transact Med Imag. 2014;33(12):2271–92. doi:10.1109/isbi.2014.6868073. [Google Scholar] [CrossRef]
15. Ollinger J, Fessler J. Positron-emission tomography. IEEE Signal Process Mag. 1997;14(1):43–55. doi:10.1109/79.560323. [Google Scholar] [CrossRef]
16. Coxson PG. Consequences of using a simplified kinetic model for dynamic PET data. J Nucl Med. 1995;38(4):660–7. [Google Scholar]
17. Abdallah NG, Rashdan M, Khalaf AA. High resolution time-to-digital converter for pet imaging. In: International Conference on Innovative Trends in Communication and Computer Engineering (ITCE); 2020 Feb 8–9; Aswan, Egypt. p. 295–8. [Google Scholar]
18. Shabbir J, Anwer T. Artificial intelligence and its role in near future. arXiv:1804.01396. 2018. [Google Scholar]
19. Bengio Y, Courville A, Vincent P. Representation learning: a review and new perspectives. IEEE Transact Patt Analy Mach Intell. 2013;35(8):1798–828. doi:10.1109/tpami.2013.50. [Google Scholar] [PubMed] [CrossRef]
20. Zhang C, Bengio S, Hardt M, Recht B, Vinyals O. Understanding deep learning requires rethinking generalization. arXiv:1611.03530. 2016. [Google Scholar]
21. Schmidhuber J. Deep learning in neural networks: an overview. Neur Netw. 2015;61(3):85–117. doi:10.1016/j.neunet.2014.09.003. [Google Scholar] [PubMed] [CrossRef]
22. Talaei Khoei T, Ould Slimane H, Kaabouch N. Deep learning: systematic review, models, challenges, and research directions. Neu Comput Applicat. 2023;35(31):23103–24. doi:10.1007/s00521-023-08957-4. [Google Scholar] [CrossRef]
23. Pan Z, Yu W, Yi X, Khan A, Yuan F, Zheng Y. Recent progress on generative adversarial networks (GANsa survey. IEEE Access. 2019;7:36322–33. doi:10.1109/access.2019.2905015. [Google Scholar] [CrossRef]
24. Aljuaid A, Anwar M. Survey of supervised learning for medical image processing. SN Comput Sci. 2022;3(4):292. doi:10.1007/s42979-022-01166-1. [Google Scholar] [PubMed] [CrossRef]
25. Rani V, Kumar M, Gupta A, Sachdeva M, Mittal A, Kumar K. Self-supervised learning for medical image analysis: a comprehensive review. Evolv Syst. 2024;15(4):1607–33. doi:10.1007/s12530-024-09581-w. [Google Scholar] [CrossRef]
26. Halevy A, Norvig P, Pereira F. The unreasonable effectiveness of data. IEEE Intell Syst. 2009;24(2):8–12. doi:10.1109/mis.2009.36. [Google Scholar] [CrossRef]
27. Raza K, Singh NK. A tour of unsupervised deep learning for medical image analysis. Current Med Imag Rev. 2021;17(9):1059–77. doi:10.2174/1573405617666210127154257. [Google Scholar] [PubMed] [CrossRef]
28. Ganeshkumar M, Sowmya V, Gopalakrishnan E, Soman K. Unsupervised deep learning-based disease diagnosis using medical images. In: Cognitive and soft computing techniques for the analysis of healthcare data. Amsterdam, The Netherlands: Elsevier; 2022. p. 203–20. doi:10.1016/b978-0-323-85751-2.00011-6. [Google Scholar] [CrossRef]
29. Croitoru FA, Hondru V, Ionescu RT, Shah M. Diffusion models in vision: a survey. IEEE Transact Pattern Anal Mach Intell. 2023;45(9):10850–69. doi:10.1109/tpami.2023.3261988. [Google Scholar] [PubMed] [CrossRef]
30. Goodfellow I, Pouget-Abadie J, Mirza M, Xu B, Warde-Farley D, Ozair S, et al. Generative adversarial networks. Communicat ACM. 2020;63(11):139–44. doi:10.1145/3422622. [Google Scholar] [CrossRef]
31. De Souza VLT, Marques BAD, Batagelo HC, Gois JP. A review on generative adversarial networks for image generation. Comput Graph. 2023;114:13–25. [Google Scholar]
32. Gui J, Sun Z, Wen Y, Tao D, Ye J. A review on generative adversarial networks: algorithms, theory, and applications. IEEE Transact Know Data Eng. 2023;35(4):3313–32. doi:10.1109/tkde.2021.3130191. [Google Scholar] [CrossRef]
33. Krichen M. Generative adversarial networks. In: 2023 14th International Conference on Computing Communication and Networking Technologies (ICCCNT); 2023 Jul 6–8; Delhi, India. p. 1–7. [Google Scholar]
34. Saxena D, Cao J. Generative adversarial networks (GANs) challenges, solutions, and future directions. ACM Comput Surv (CSUR). 2021;54(3):1–42. doi:10.1145/3446374. [Google Scholar] [CrossRef]
35. Aggarwal A, Mittal M, Battineni G. Generative adversarial network: an overview of theory and applications. Int J Inform Manag Data Insights. 2021;1(1):100004. doi:10.1016/j.jjimei.2020.100004. [Google Scholar] [CrossRef]
36. Creswell A, White T, Dumoulin V, Arulkumaran K, Sengupta B, Bharath AA. Generative adversarial networks: an overview. IEEE Signal Process Magaz. 2018;35(1):53–65. doi:10.1109/msp.2017.2765202. [Google Scholar] [CrossRef]
37. Bisht A, Rawat K, Bhati JP, Vats S, Sharma V, Singh S. Generating images using vanilla generative adversarial networks. In: 2024 4th International Conference on Technological Advancements in Computational Sciences (ICTACS). IEEE; 2024 Nov 13–15; Tashkent, Uzbekistan: IEEE. p. 463–8. [Google Scholar]
38. Radford A, Metz L, Chintala S. Unsupervised representation learning with deep convolutional generative adversarial networks. arXiv:1511.06434. 2015. [Google Scholar]
39. Kim HJ, Lee D. Image denoising with conditional generative adversarial networks (CGAN) in low dose chest images. Nuclear Instrum Meth Phy Res Sec A Accel Spectrom Detect Assoc Equip. 2020;954(8):161914. doi:10.1016/j.nima.2019.02.041. [Google Scholar] [CrossRef]
40. Gulrajani I, Ahmed F, Arjovsky M, Dumoulin V, Courville AC. Improved training of wasserstein gans. In: Advances in neural information processing systems. Cambridge, MA, USA: The MIT Press; 2017. [Google Scholar]
41. Chu C, Zhmoginov A, Sandler M. Cyclegan, a master of steganography. arXiv:1712.02950. 2017. [Google Scholar]
42. Goodfellow I, Pouget-Abadie J, Mirza M, Xu B, Warde-Farley D, Ozair S, et al. Generative adversarial. arXiv:1406.2661. 2014. [Google Scholar]
43. Karras T, Aila T, Laine S, Lehtinen J. Progressive growing of gans for improved quality, stability, and variation. arXiv:1710.10196. 2017. [Google Scholar]
44. Motamed S, Rogalla P, Khalvati F. RANDGAN: randomized generative adversarial network for detection of COVID-19 in chest X-ray. Scient Rep. 2021;11(1):8602. doi:10.1038/s41598-021-87994-2. [Google Scholar] [PubMed] [CrossRef]
45. Kossen T, Subramaniam P, Madai VI, Hennemuth A, Hildebrand K, Hilbert A, et al. Synthesizing anonymized and labeled TOF-MRA patches for brain vessel segmentation using generative adversarial networks. Comput Biol Med. 2021;131:104254. doi:10.1016/j.compbiomed.2021.104254. [Google Scholar] [PubMed] [CrossRef]
46. Ahmad B, Sun J, You Q, Palade V, Mao Z. Brain tumor classification using a combination of variational autoencoders and generative adversarial networks. Biomedicines. 2022;10(2):223. doi:10.3390/biomedicines10020223. [Google Scholar] [PubMed] [CrossRef]
47. Krithika alias Anbu Devi M, Suganthi K. Review of medical image synthesis using GAN techniques. In: ITM Web of Conferences. Vol. 37. Les Ulis, France: EDP Sciences; 2021. 01005 p. [Google Scholar]
48. Zhou T, Li Q, Lu H, Cheng Q, Zhang X. GAN review: models and medical image fusion applications. Inform Fus. 2023;91(11):134–48. doi:10.1016/j.inffus.2022.10.017. [Google Scholar] [CrossRef]
49. Ma Y, Liu J, Liu Y, Fu H, Hu Y, Cheng J, et al. Structure and illumination constrained GAN for medical image enhancement. IEEE Transact Med Imag. 2021;40(12):3955–67. doi:10.1109/tmi.2021.3101937. [Google Scholar] [PubMed] [CrossRef]
50. Tightiz L, Jung MH, Song I, Lee K. Trustworthy TAVR navigator system, I: a generative adversarial network-driven medical twin approach for Post-TAVR pacemaker implantation prediction. Expert Syst Applicat. 2025;275(1):126973. doi:10.1016/j.eswa.2025.126973. [Google Scholar] [CrossRef]
51. Kazeminia S, Baur C, Kuijper A, van Ginneken B, Navab N, Albarqouni S, et al. GANs for medical image analysis. Artif Intelli Med. 2020;109(12):101938. doi:10.1016/j.artmed.2020.101938. [Google Scholar] [PubMed] [CrossRef]
52. Fu B, Zhang X, Wang L, Ren Y, Thanh DN. A blind medical image denoising method with noise generation network. J X-Ray Sci Technol. 2022;30(3):531–47. doi:10.3233/xst-211098. [Google Scholar] [PubMed] [CrossRef]
53. Li Y, Zhang K, Shi W, Miao Y, Jiang Z. A novel medical image denoising method based on conditional generative adversarial network. Computat Mathem Meth Med. 2021;2021(2):9974017. doi:10.1155/2021/9974017. [Google Scholar] [PubMed] [CrossRef]
54. Geng M, Meng X, Yu J, Zhu L, Jin L, Jiang Z, et al. Content-noise complementary learning for medical image denoising. IEEE Transact Med Imag. 2021;41(2):407–19. doi:10.1109/tmi.2021.3113365. [Google Scholar] [PubMed] [CrossRef]
55. Gondara L. Medical image denoising using convolutional denoising autoencoders. In: 2016 IEEE 16th International Conference on Data Mining Workshops (ICDMW); 2016 Dec 12–15; Barcelona, Spain. p. 241–6. [Google Scholar]
56. Xue Y, Xu T, Zhang H, Long LR, Huang X. SegAN: adversarial network with multi-scale L1 loss for medical image segmentation. Neuroinformatics. 2018;16(3–4):383–92. doi:10.1007/s12021-018-9377-x. [Google Scholar] [PubMed] [CrossRef]
57. Han Z, Wei B, Mercado A, Leung S, Li S. Spine-GAN: semantic segmentation of multiple spinal structures. Med Image Anal. 2018;50(1):23–35. doi:10.1016/j.media.2018.08.005. [Google Scholar] [PubMed] [CrossRef]
58. Trinh DH, Luong M, Dibos F, Rocchisani JM, Pham CD, Nguyen TQ. Novel example-based method for super-resolution and denoising of medical images. IEEE Transact Image Process. 2014;23(4):1882–95. doi:10.1109/tip.2014.2308422. [Google Scholar] [PubMed] [CrossRef]
59. Sánchez I, Vilaplana V. Brain MRI super-resolution using 3D generative adversarial networks. arXiv:1812.11440. 2018. [Google Scholar]
60. Wang TC, Liu MY, Zhu JY, Tao A, Kautz J, Catanzaro B. High-resolution image synthesis and semantic manipulation with conditional gans. In: Proceedings of the 2018 IEEE Conference on Computer Vision and Pattern Recognition; 2018 Jun 18–23; Salt Lake City, UT, USA. p. 8798–807. [Google Scholar]
61. Dar SU, Yurt M, Karacan L, Erdem A, Erdem E, Cukur T. Image synthesis in multi-contrast MRI with conditional generative adversarial networks. IEEE Transact Med Imag. 2019;38(10):2375–88. doi:10.1109/tmi.2019.2901750. [Google Scholar] [PubMed] [CrossRef]
62. Platscher M, Zopes J, Federau C. Image translation for medical image generation: ischemic stroke lesion segmentation. Biomed Signal Process Cont. 2022;72(4):103283. doi:10.1016/j.bspc.2021.103283. [Google Scholar] [CrossRef]
63. Upadhyay U, Chen Y, Hepp T, Gatidis S, Akata Z. Uncertainty-guided progressive GANs for medical image translation. In: Medical Image Computing and Computer Assisted Intervention-MICCAI 2021: 24th International Conference. Cham, Swizterland: Springer; 2021. p. 614–24. [Google Scholar]
64. Han C, Hayashi H, Rundo L, Araki R, Shimoda W, Muramatsu S, et al. GAN-based synthetic brain MR image generation. In: IEEE 15th International Symposium on Biomedical Imaging (ISBI 2018); 2018 Apr 4–7; Washington, DC, USA; 2018. p. 734–8. [Google Scholar]
65. Singh NK, Raza K. Medical image generation using generative adversarial networks: a review. In: Health informatics: a computational perspective in healthcare. Singapore: Springer; 2021. p. 77–96. [Google Scholar]
66. Fan C, Lin H, Qiu Y. U-Patch GAN: a medical image fusion method based on GAN. J Digit Imag. 2023;36(1):339–55. doi:10.1007/s10278-022-00696-7. [Google Scholar] [PubMed] [CrossRef]
67. Guo K, Chen J, Qiu T, Guo S, Luo T, Chen T, et al. MedGAN: an adaptive GAN approach for medical image generation. Comput Biol Med. 2023;163(7):107119. doi:10.1016/j.compbiomed.2023.107119. [Google Scholar] [PubMed] [CrossRef]
68. Chlap P, Min H, Vandenberg N, Dowling J, Holloway L, Haworth A. A review of medical image data augmentation techniques for deep learning applications. J Med Imag Radiat Oncol. 2021;65(5):545–63. doi:10.1111/1754-9485.13261. [Google Scholar] [PubMed] [CrossRef]
69. Garcea F, Serra A, Lamberti F, Morra L. Data augmentation for medical imaging: a systematic literature review. Comput Biol Med. 2023;152(1):106391. doi:10.1016/j.compbiomed.2022.106391. [Google Scholar] [PubMed] [CrossRef]
70. Kebaili A, Lapuyade-Lahorgue J, Ruan S. Deep learning approaches for data augmentation in medical imaging: a review. J Imag. 2023;9(4):81. doi:10.3390/jimaging9040081. [Google Scholar] [PubMed] [CrossRef]
71. Sun Y, Yuan P, Sun Y. MM-GAN: 3D MRI data augmentation for medical image segmentation via generative adversarial networks. In: 2020 IEEE International Conference on Knowledge Graph (ICKG); 2020 Aug 9–11; Nanjing, China. p. 227–34. [Google Scholar]
72. Goceri E. Medical image data augmentation: techniques, comparisons and interpretations. Artif Intell Rev. 2023;56(11):12561–605. doi:10.1007/s10462-023-10453-z. [Google Scholar] [PubMed] [CrossRef]
73. Tschuchnig ME, Gadermayr M. Anomaly detection in medical imaging—a mini review. In: International Data Science Conference. Cham, Switzerland: Springer; 2021. p. 33–8. [Google Scholar]
74. Xia X, Pan X, Li N, He X, Ma L, Zhang X, et al. GAN-based anomaly detection: a review. Neurocomputing. 2022;493:497–535. doi:10.1016/j.neucom.2021.12.093. [Google Scholar] [CrossRef]
75. Esmaeili M, Toosi A, Roshanpoor A, Changizi V, Ghazisaeedi M, Rahmim A, et al. Generative adversarial networks for anomaly detection in biomedical imaging: a study on seven medical image datasets. IEEE Access. 2023;11:17906–21. doi:10.1109/access.2023.3244741. [Google Scholar] [CrossRef]
76. Vyas B, Rajendran RM. Generative adversarial networks for anomaly detection in medical images. Int J Multidisci Innova Res Method. 2023;2(4):52–8. [Google Scholar]
77. Han C, Rundo L, Murao K, Noguchi T, Shimahara Y, Milacski ZÁ, et al. MADGAN: unsupervised medical anomaly detection GAN using multiple adjacent brain MRI slice reconstruction. BMC Bioinformatics. 2021;22(S2):31. doi:10.1186/s12859-020-03936-1. [Google Scholar] [PubMed] [CrossRef]
78. Zhang H, Guo W, Zhang S, Lu H, Zhao X. Unsupervised deep anomaly detection for medical images using an improved adversarial autoencoder. J Digital Imag. 2022;35(2):153–61. doi:10.1007/s10278-021-00558-8. [Google Scholar] [PubMed] [CrossRef]
79. Guan H, Liu M. Domain adaptation for medical image analysis: a survey. IEEE Transact Biomed Eng. 2022;69(3):1173–85. doi:10.1109/tbme.2021.3117407. [Google Scholar] [PubMed] [CrossRef]
80. Xie X, Chen J, Li Y, Shen L, Ma K, Zheng Y. MI 2 GAN: generative adversarial network for medical image domain adaptation using mutual information constraint. In: International Conference on Medical Image Computing and Computer-Assisted Intervention. Cham, Switzerland: Springer; 2020. p. 516–25. [Google Scholar]
81. Sun Y, Yang G, Ding D, Cheng G, Xu J, Li X. A GAN-based domain adaptation method for glaucoma diagnosis. In: 2020 International Joint Conference on Neural Networks (IJCNN); 2020 Jul 19–24; Glasgow, UK. p. 1–8. [Google Scholar]
82. Sagheer SVM, George SN. A review on medical image denoising algorithms. Biomed Signal Process Cont. 2020;61(11):102036. doi:10.1016/j.bspc.2020.102036. [Google Scholar] [CrossRef]
83. Isola P, Zhu JY, Zhou T, Efros AA. Image-to-image translation with conditional adversarial networks. In: Proceedings of the 2017 IEEE Conference on Computer Vision and Pattern Recognition; 2017 Jul 21–26; Honolulu, HI, USA. p. 1125–34. [Google Scholar]
84. Kitchenham B, Brereton OP, Budgen D, Turner M, Bailey J, Linkman S. Systematic literature reviews in software engineering-a systematic literature review. Informati Software Technol. 2009;51(1):7–15. doi:10.1016/j.infsof.2008.09.009. [Google Scholar] [CrossRef]
85. Ali M, Ali M, Hussain M, Koundal D. Generative adversarial networks (GANs) for medical image processing: recent advancements. Arch Comput Methods Eng. 2025;32(2):1185–98. doi:10.1007/s11831-024-10174-8. [Google Scholar] [CrossRef]
86. Tu H, Wang Z, Zhao Y. Unpaired image-to-image translation with diffusion adversarial network. Mathematics. 2024:3178. doi:10.21203/rs.3.rs-4502713/v1. [Google Scholar] [CrossRef]
87. Wang G, Shi H, Chen Y, Wu B. Unsupervised image-to-image translation via long-short cycle-consistent adversarial networks. Appl Intell. 2023;53(14):17243–59. doi:10.1007/s10489-022-04389-0. [Google Scholar] [CrossRef]
88. Sun L, Chen J, Xu Y, Gong M, Yu K, Batmanghelich K. Hierarchical amortized GAN for 3D high resolution medical image synthesis. IEEE J Biomed Health Inform. 2022;26(8):3966–75. doi:10.1109/jbhi.2022.3172976. [Google Scholar] [PubMed] [CrossRef]
89. Xun S, Li D, Zhu H, Chen M, Wang J, Li J, et al. Generative adversarial networks in medical image segmentation: a review. Comput Biol Med. 2022;140:105063. doi:10.1016/j.compbiomed.2021.105063. [Google Scholar] [PubMed] [CrossRef]
90. Armanious K, Jiang C, Fischer M, Küstner T, Hepp T, Nikolaou K, et al. MedGAN: medical image translation using GANs. Comput Med Imag Grap. 2020;79:101684. doi:10.1016/j.compmedimag.2019.101684. [Google Scholar] [PubMed] [CrossRef]
91. Zhang Q, Sun J, Mok GS. Low dose SPECT image denoising using a generative adversarial network. arXiv:1907.11944. 2019. [Google Scholar]
92. Mahapatra D, Bozorgtabar B, Garnavi R. Image super-resolution using progressive generative adversarial networks for medical image analysis. Comput Med Imag Grap. 2019;71(6):30–9. doi:10.1016/j.compmedimag.2018.10.005. [Google Scholar] [PubMed] [CrossRef]
93. You C, Li G, Zhang Y, Zhang X, Shan H, Li M, et al. CT super-resolution GAN constrained by the identical, residual, and cycle learning ensemble (GAN-CIRCLE). IEEE Transact Med Imag. 2020;39(1):188–203. doi:10.1109/tmi.2019.2922960. [Google Scholar] [PubMed] [CrossRef]
94. Yang Q, Yan P, Zhang Y, Yu H, Shi Y, Mou X, et al. Low-dose CT image denoising using a generative adversarial network with Wasserstein distance and perceptual loss. IEEE Transact Med Imag. 2018;37(6):1348–57. doi:10.1109/tmi.2018.2827462. [Google Scholar] [PubMed] [CrossRef]
95. Li Y, Shen L. cC-GAN: a robust transfer-learning framework for HEp-2 specimen image segmentation. IEEE Access. 2018;6:14048–58. doi:10.1109/access.2018.2808938. [Google Scholar] [CrossRef]
96. Hu B, Tang Y, Eric I, Chang C, Fan Y, Lai M, et al. Unsupervised learning for cell-level visual representation in histopathology images with generative adversarial networks. IEEE J Biomed Health Inform. 2018;23(3):1316–28. doi:10.1109/jbhi.2018.2852639. [Google Scholar] [PubMed] [CrossRef]
97. Nie D, Trullo R, Lian J, Wang L, Petitjean C, Ruan S, et al. Medical image synthesis with deep convolutional adversarial networks. IEEE Transact Biomed Eng. 2018;65(12):2720–30. [Google Scholar]
98. Bissoto A, Perez F, Valle E, Avila S. Skin lesion synthesis with generative adversarial networks. In: OR 2.0 context-aware operating theaters, computer assisted robotic endoscopy, clinical image-based procedures, and skin image analysis. Cham, Switzerland: Springer; 2018. p. 294–302. doi:10.1007/978-3-030-01201-4_32. [Google Scholar] [CrossRef]
99. Udrea A, Mitra GD. Generative adversarial neural networks for pigmented and non-pigmented skin lesions detection in clinical images. In: 2017 21st International Conference on Control Systems and Computer Science (CSCS); 2017 May 29–31; Bucharest, Romania. p. 364–8. [Google Scholar]
100. Wang Y, Yang N, Li J. GAN-based architecture for low-dose computed tomography imaging denoising. arXiv:2411.09512. 2024. [Google Scholar]
101. Yu X, Luan S, Lei S, Huang J, Liu Z, Xue X, et al. Deep learning for fast denoising filtering in ultrasound localization microscopy. Phy Med Biol. 2023;68(20):205002. doi:10.1088/1361-6560/acf98f. [Google Scholar] [PubMed] [CrossRef]
102. Ran M, Hu J, Chen Y, Chen H, Sun H, Zhou J, et al. Denoising of 3D magnetic resonance images using a residual encoder-decoder Wasserstein generative adversarial network. Med Image Anal. 2019;55(2):165–80. doi:10.1016/j.media.2019.05.001. [Google Scholar] [PubMed] [CrossRef]
103. Kaur A, Dong G. A complete review on image denoising techniques for medical images. Neural Process Lett. 2023;55(6):7807–50. doi:10.1007/s11063-023-11286-1. [Google Scholar] [CrossRef]
104. Yi X, Babyn P. Sharpness-aware low-dose CT denoising using conditional generative adversarial network. J Digital Imag. 2018;31(5):655–69. doi:10.1007/s10278-018-0056-0. [Google Scholar] [PubMed] [CrossRef]
105. Shan H, Zhang Y, Yang Q, Kruger U, Kalra MK, Sun L, et al. 3-D convolutional encoder-decoder network for low-dose CT via transfer learning from a 2-D trained network. IEEE Transact Med Imag. 2018;37(6):1522–34. doi:10.1109/tmi.2018.2878429. [Google Scholar] [CrossRef]
106. Wolterink JM, Leiner T, Viergever MA, Išgum I. Generative adversarial networks for noise reduction in low-dose CT. IEEE Transact Med Imag. 2017;36(12):2536–45. doi:10.1109/tmi.2017.2708987. [Google Scholar] [PubMed] [CrossRef]
107. Huang Z, Zhang J, Zhang Y, Shan H. DU-GAN: generative adversarial networks with dual-domain U-Net-based discriminators for low-dose CT denoising. IEEE Transact Instrument Measure. 2022;71:1–12. doi:10.1109/tim.2021.3128703. [Google Scholar] [CrossRef]
108. Zhu ML, Zhao LL, Xiao L. Image denoising based on GAN with optimization algorithm. Electronics. 2022;11(15):2445. doi:10.3390/electronics11152445. [Google Scholar] [CrossRef]
109. Wang R, Lei T, Cui R, Zhang B, Meng H, Nandi AK. Medical image segmentation using deep learning: a survey. IET Image Process. 2022;16(5):1243–67. doi:10.1049/ipr2.12419. [Google Scholar] [CrossRef]
110. Cai L, Fang H, Xu N, Ren B. Counterfactual causal-effect intervention for interpretable medical visual question answering. IEEE Trans Med Imaging. 2024;43(12):4430–41. doi:10.1109/TMI.2024.3425533. [Google Scholar] [PubMed] [CrossRef]
111. Song W, Wang X, Guo Y, Li S, Xia B, Hao A. Centerformer: a novel cluster center enhanced transformer for unconstrained dental plaque segmentation. IEEE Trans Multimedia. 2024;26:10965–78. doi:10.1109/TMM.2024.3428349. [Google Scholar] [CrossRef]
112. Dong X, Lei Y, Wang T, Thomas M, Tang L, Curran WJ, et al. Automatic multiorgan segmentation in thorax CT images using U-net-GAN. Med Phy. 2019;46(5):2157–68. doi:10.1002/mp.13458. [Google Scholar] [PubMed] [CrossRef]
113. Guo B, Yang Y, Yu Z, Zhu Y. Multi-modality medical image segmentation via adversarial learning with CV energy functional. Expert Syst Appl. 2025;283(8):127467. doi:10.1016/j.eswa.2025.127467. [Google Scholar] [CrossRef]
114. Wang G, Ma Q, Li Y, Mao K, Xu L, Zhao Y. A skin lesion segmentation network with edge and body fusion. Appl Soft Comput. 2025;170(1):112683. doi:10.1016/j.asoc.2024.112683. [Google Scholar] [CrossRef]
115. Gunasagar P, Durgaprasad V, Abhijith K, Sai RM, Ameer P. Enhancing facial expression synthesis through GAN with multi-scale dilated feature extraction and edge-enhanced facial features. In: TENCON 2023-2023 IEEE Region 10 Conference (TENCON); 2023 Oct 31–Nov 3; Chiang Mai, Thailand. p. 47–52. [Google Scholar]
116. Nimitha U, Ameer P. MRI super-resolution using similarity distance and multi-scale receptive field based feature fusion GAN and pre-trained slice interpolation network. Magn Reson Imag. 2024;110(5):195–209. doi:10.1016/j.mri.2024.04.021. [Google Scholar] [PubMed] [CrossRef]
117. Wang B, Chen W, Qian J, Feng S, Chen Q, Zuo C. Single-shot super-resolved fringe projection profilometry (SSSR-FPP100,000 frames-per-second 3D imaging with deep learning. Light Sci Appl. 2025;14(1):70. doi:10.1038/s41377-024-01721-w. [Google Scholar] [PubMed] [CrossRef]
118. Ledig C, Theis L, Huszár F, Caballero J, Cunningham A, Acosta A, et al. Photo-realistic single image super-resolution using a generative adversarial network. In: Proceedings of the 2017 IEEE Conference on Computer Vision and Pattern Recognition; 2017 Jul 21–26; Honolulu, HI, USA. p. 4681–90. [Google Scholar]
119. Zhao T, Lin Y, Xu Y, Chen W, Wang Z. Learning-based quality assessment for image super-resolution. IEEE Transact Multim. 2022;24:3570–81. doi:10.1109/tmm.2021.3102401. [Google Scholar] [CrossRef]
120. Chen Y, Shi F, Christodoulou AG, Xie Y, Zhou Z, Li D. Efficient and accurate MRI super-resolution using a generative adversarial network and 3D multi-level densely connected network. In: International Conference on Medical Image Computing and Computer-Assisted Intervention. Cham, Switzerland: Springer; 2018. p. 91–9. [Google Scholar]
121. Gurcan MN, Boucheron LE, Can A, Madabhushi A, Rajpoot NM, Yener B. Histopathological image analysis: a review. IEEE Rev Biomed Eng. 2009;2:147–71. doi:10.1109/rbme.2009.2034865. [Google Scholar] [PubMed] [CrossRef]
122. Vu CT, Phan TD, Chandler DM. A spectral and spatial measure of local perceived sharpness in natural images. IEEE Transact Image Process. 2012;21(3):934–45. doi:10.1109/tip.2011.2169974. [Google Scholar] [PubMed] [CrossRef]
123. Brenner DJ, Hall EJ. Computed tomography—an increasing source of radiation exposure. New Engl J Med. 2007;357(22):2277–84. doi:10.1056/nejmra072149. [Google Scholar] [PubMed] [CrossRef]
124. Gu Y, Zeng Z, Chen H, Wei J, Zhang Y, Chen B, et al. MedSRGAN: medical images super-resolution using generative adversarial networks. Multim Tools Applicat. 2020;79(29–30):21815–40. doi:10.1007/s11042-020-08980-w. [Google Scholar] [CrossRef]
125. Wang R, Fang Z, Gu J, Guo Y, Zhou S, Wang Y, et al. High-resolution image reconstruction for portable ultrasound imaging devices. EURASIP J Adv Signal Process. 2019;2019(1):1–12. doi:10.1186/s13634-019-0649-x. [Google Scholar] [CrossRef]
126. Luan S, Yu X, Lei S, Ma C, Wang X, Xue X, et al. Deep learning for fast super-resolution ultrasound microvessel imaging. Phy Med Biol. 2023;68(24):245023. doi:10.1088/1361-6560/ad0a5a. [Google Scholar] [PubMed] [CrossRef]
127. Nimitha U, Ameer P. Multi image super resolution of MRI images using generative adversarial network. J Ambient Intell Human Comput. 2024;15(4):2241–53. doi:10.1007/s12652-024-04751-9. [Google Scholar] [CrossRef]
128. Jia Y, Chen G, Chi H. Retinal fundus image super-resolution based on generative adversarial network guided with vascular structure prior. Scient Rep. 2024;14(1):22786. doi:10.1038/s41598-024-74186-x. [Google Scholar] [PubMed] [CrossRef]
129. Wang X, Xie L, Dong C, Shan Y. Real-esrgan: training real-world blind super-resolution with pure synthetic data. In: Proceedings of the 2021 IEEE/CVF International Conference on Computer Vision; 2021 Oct 11–17; Montreal, BC, Canada. p. 1905–14. [Google Scholar]
130. Zhao Y, Zheng Y, Liu Y, Zhao Y, Luo L, Yang S, et al. Automatic 2-D/3-D vessel enhancement in multiple modality images using a weighted symmetry filter. IEEE Transact Med Imag. 2017;37(2):438–50. doi:10.1109/tmi.2017.2756073. [Google Scholar] [PubMed] [CrossRef]
131. Jobson DJ, Rahman Z, Woodell GA. A multiscale retinex for bridging the gap between color images and the human observation of scenes. IEEE Transact Image Process. 1997;6(7):965–76. doi:10.1109/83.597272. [Google Scholar] [PubMed] [CrossRef]
132. Chen YS, Wang YC, Kao MH, Chuang YY. Deep photo enhancer: unpaired learning for image enhancement from photographs with gans. In: Proceedings of the 2018 IEEE Conference on Computer Vision and Pattern Recognition; 2018 Jun 18–23; Salt Lake City, UT, USA. p. 6306–14. [Google Scholar]
133. Alotaibi A. Deep generative adversarial networks for image-to-image translation: a review. Symmetry. 2020;12(10):1705. doi:10.3390/sym12101705. [Google Scholar] [CrossRef]
134. Lore KG, Akintayo A, Sarkar S. LLNet: a deep autoencoder approach to natural low-light image enhancement. Pattern Recognit. 2017;61(6):650–62. doi:10.1016/j.patcog.2016.06.008. [Google Scholar] [CrossRef]
135. Gatys LA. A neural algorithm of artistic style. arXiv:1508.06576. 2015. [Google Scholar]
136. Zhu JY, Park T, Isola P, Efros AA. Unpaired image-to-image translation using cycle-consistent adversarial networks. In: Proceedings of the 2017 IEEE International Conference on Computer Vision; 2017 Oct 22–29; Venice, Italy. p. 2223–32. [Google Scholar]
137. Yan S, Wang C, Chen W, Lyu J. Swin transformer-based GAN for multi-modal medical image translation. Front Oncol. 2022;12:942511. doi:10.3389/fonc.2022.942511. [Google Scholar] [PubMed] [CrossRef]
138. Liang J, Cao J, Sun G, Zhang K, Van Gool L, Timofte R. Swinir: image restoration using swin transformer. In: Proceedings of the 2021 IEEE/CVF iNternational Conference on Computer Vision; 2021 Oct 11–17; Montreal, BC, Canada. p. 1833–44. [Google Scholar]
139. Chen Y, Lin Y, Xu X, Ding J, Li C, Zeng Y, et al. Multi-domain medical image translation generation for lung image classification based on generative adversarial networks. Comput Meth Prog Biomed. 2023;229(20):107200. doi:10.1016/j.cmpb.2022.107200. [Google Scholar] [PubMed] [CrossRef]
140. Özbey M, Dalmaz O, Dar SU, Bedel HA, Özturk Ş, Güngör A, et al. Unsupervised medical image translation with adversarial diffusion models. IEEE Transact Med Imag. 2023;42(12):3524–39. doi:10.1109/tmi.2023.3290149. [Google Scholar] [PubMed] [CrossRef]
141. Kim KH, Do WJ, Park SH. Improving resolution of MR images with an adversarial network incorporating images with different contrast. Med Phy. 2018;45(7):3120–31. doi:10.1002/mp.12945. [Google Scholar] [PubMed] [CrossRef]
142. Shitrit O, Riklin Raviv T. Accelerated magnetic resonance imaging by adversarial neural network. In: Deep learning in medical image analysis and multimodal learning for clinical decision support. Cham, Switzerland: Springer; 2017. p. 30–8. doi:10.1007/978-3-319-67558-9_4. [Google Scholar] [CrossRef]
143. Seitzer M, Yang G, Schlemper J, Oktay O, Würfl T, Christlein V, et al. Adversarial and perceptual refinement for compressed sensing MRI reconstruction. In: Medical image computing and computer assisted intervention-MICCAI 2018. Cham, Switzerland: Springer; 2018. p. 232–40. doi:10.1007/978-3-030-00928-1_27. [Google Scholar] [CrossRef]
144. Armanious K, Mecky Y, Gatidis S, Yang B. Adversarial inpainting of medical image modalities. In: ICASSP 2019-2019 IEEE International Conference on Acoustics, Speech and Signal Processing (ICASSP); 2019 May 12–17; Brighton, UK. p. 3267–71. [Google Scholar]
145. Wang Y, Yu B, Wang L, Zu C, Lalush DS, Lin W, et al. 3D conditional generative adversarial networks for high-quality PET image estimation at low dose. Neuroimage. 2018;174(7):550–62. doi:10.1016/j.neuroimage.2018.03.045. [Google Scholar] [PubMed] [CrossRef]
146. Liao H, Huo Z, Sehnert WJ, Zhou SK, Luo J. Adversarial sparse-view CBCT artifact reduction. In: Medical image computing and computer assisted intervention-MICCAI 2018I. Cham, Switzerland: Springer; 2018. p. 154–62. doi:10.1007/978-3-030-00928-1_18. [Google Scholar] [CrossRef]
147. Quan TM, Nguyen-Duc T, Jeong WK. Compressed sensing MRI reconstruction using a generative adversarial network with a cyclic loss. IEEE Transact Med Imag. 2018;37(6):1488–97. doi:10.1109/tmi.2018.2820120. [Google Scholar] [PubMed] [CrossRef]
148. Mardani M, Gong E, Cheng JY, Vasanawala SS, Zaharchuk G, Xing L, et al. Deep generative adversarial neural networks for compressive sensing MRI. IEEE Transact Med Imag. 2018;38(1):167–79. doi:10.1109/tmi.2018.2858752. [Google Scholar] [PubMed] [CrossRef]
149. Kang E, Koo HJ, Yang DH, Seo JB, Ye JC. Cycle-consistent adversarial denoising network for multiphase coronary CT angiography. Med Phy. 2019;46(2):550–62. doi:10.1002/mp.13284. [Google Scholar] [PubMed] [CrossRef]
150. Bhadra S, Zhou W, Anastasio MA. Medical image reconstruction with image-adaptive priors learned by use of generative adversarial networks. In: Medical imaging 2020: physics of medical imaging. Vol. 11312. Bellingham, WA, USA: SPIE; 2020. p. 206–13. [Google Scholar]
151. Rashid ZM, Alsawaff ZH, Yousif AS, Al-Nima RRO. Optimization of PET image reconstruction for enhanced image quality in various tasks using a conventional PET scanner. J Elect Comput Eng. 2025;2025(1):8108611. doi:10.1155/jece/8108611. [Google Scholar] [CrossRef]
152. Ahn G, Choi BS, Ko S, Jo C, Han HS, Lee MC, et al. High-resolution knee plain radiography image synthesis using style generative adversarial network adaptive discriminator augmentation. J Orthop Res®. 2023;41(1):84–93. doi:10.1002/jor.25325. [Google Scholar] [PubMed] [CrossRef]
153. Zhu L, He Q, Huang Y, Zhang Z, Zeng J, Lu L, et al. DualMMP-GAN: dual-scale multi-modality perceptual generative adversarial network for medical image segmentation. Comput Biol Med. 2022;144(10):105387. doi:10.1016/j.compbiomed.2022.105387. [Google Scholar] [PubMed] [CrossRef]
154. Che Azemin MZ, Mohd Tamrin MI, Hilmi MR, Mohd Kamal K. Assessing the efficacy of StyleGAN 3 in generating realistic medical images with limited data availability. In: Proceedings of the 2024 13th International Conference on Software and Computer Applications; 2024 Feb 1–3; Bali Island, Indonesia. p. 192–7. [Google Scholar]
155. Zhao A, Xu M, Shahin AH, Wuyts W, Jones MG, Jacob J, et al. 4D VQ-GAN: synthesising medical scans at any time point for personalised disease progression modelling of idiopathic pulmonary fibrosis. arXiv:2502.05713. 2025. [Google Scholar]
156. Antoniou A, Storkey A, Edwards H. Data augmentation generative adversarial networks. arXiv:1711.04340. 2017. [Google Scholar]
157. Frid-Adar M, Klang E, Amitai M, Goldberger J, Greenspan H. Synthetic data augmentation using GAN for improved liver lesion classification. In: IEEE 15th International Symposium On Biomedical Imaging (ISBI 2018); 2018 Apr 4–7; Washington, DC, USA; 2018. p. 289–93. [Google Scholar]
158. Gan HS, Ramlee MH, Al-Rimy BAS, Lee YS, Akkaraekthalin P. Hierarchical knee image synthesis framework for generative adversarial network: data from the osteoarthritis initiative. IEEE Access. 2022;10:55051–61. doi:10.1109/access.2022.3175506. [Google Scholar] [CrossRef]
159. Bowles C, Chen L, Guerrero R, Bentley P, Gunn R, Hammers A, et al. Gan augmentation: augmenting training data using generative adversarial networks. arXiv:1810.10863. 2018. [Google Scholar]
160. Kong L, Lian C, Huang D, Hu Y, Zhou Q. Breaking the dilemma of medical image-to-image translation. Adv Neural Inform Process Syst. 2021;34:1964–78. [Google Scholar]
161. Du W, Tian S. Transformer and GAN-based super-resolution reconstruction network for medical images. Tsinghua Sci Technol. 2024;29(1):197–206. doi:10.26599/tst.2022.9010071. [Google Scholar] [CrossRef]
162. Zuo Q, Tian H, Li R, Guo J, Hu J, Tang L, et al. Hemisphere-separated cross-connectome aggregating learning via VAE-GAN for brain structural connectivity synthesis. IEEE Access. 2023;11:48493–505. doi:10.1109/access.2023.3276989. [Google Scholar] [CrossRef]
163. Zhang R, Lu W, Gao J, Tian Y, Wei X, Wang C, et al. RFI-GAN: a reference-guided fuzzy integral network for ultrasound image augmentation. Inform Sci. 2023;623(5):709–28. doi:10.1016/j.ins.2022.12.026. [Google Scholar] [CrossRef]
164. Zhang Y, Wang Z, Zhang Z, Liu J, Feng Y, Wee L, et al. GAN-based one dimensional medical data augmentation. Soft Comput. 2023;27(15):10481–91. doi:10.1007/s00500-023-08345-z. [Google Scholar] [CrossRef]
165. Showrov I, Aziz MT, Nabil HR, Jim JR, Kabir MM, Mridha M, et al. Generative adversarial networks (GANs) in medical imaging: advancements, applications and challenges. IEEE Access. 2024;12:35728–53. doi:10.1109/access.2024.3370848. [Google Scholar] [CrossRef]
166. Saad MM, O’Reilly R, Rehmani MH. A survey on training challenges in generative adversarial networks for biomedical image analysis. Artif Intell Rev. 2024;57(2):19. doi:10.1007/s10462-023-10624-y. [Google Scholar] [CrossRef]
167. Biswas S, Rohdin J, Drahanskỳ M. Synthetic retinal images from unconditional GANs. In: 2019 41st Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC); 2019 Jul 23–27; Berlin, Germany. p. 2736–9. [Google Scholar]
168. Abdelhalim ISA, Mohamed MF, Mahdy YB. Data augmentation for skin lesion using self-attention based progressive generative adversarial network. Expert Syst Applicat. 2021;165(3):113922. doi:10.1016/j.eswa.2020.113922. [Google Scholar] [CrossRef]
169. Goel T, Murugan R, Mirjalili S, Chakrabartty DK. Automatic screening of covid-19 using an optimized generative adversarial network. Cognit Computat. 2024;16(4):1666–81. doi:10.1007/s12559-020-09785-7. [Google Scholar] [PubMed] [CrossRef]
170. Wang Z, She Q, Ward TE. Generative adversarial networks in computer vision: a survey and taxonomy. ACM Comput Surv (CSUR). 2021;54(2):1–38. doi:10.1145/3439723. [Google Scholar] [CrossRef]
171. Heusel M, Ramsauer H, Unterthiner T, Nessler B, Hochreiter S. Gans trained by a two time-scale update rule converge to a local nash equilibrium. In: Advances in neural information processing systems. Cambridge, MA, USA: The MIT Press; 2017. doi:10.18034/ajase.v8i1.9. [Google Scholar] [CrossRef]
172. Mirjalili S, Lewis A. The whale optimization algorithm. Adv Eng Soft. 2016;95(12):51–67. doi:10.1016/j.advengsoft.2016.01.008. [Google Scholar] [CrossRef]
173. Dukler Y, Li W, Lin A, Montúfar G. Wasserstein of Wasserstein loss for learning generative models. In: The 36th International Conference on Machine Learning; 2019 Jun 10–15; Long Beach, CA, USA. p. 1716–25. [Google Scholar]
174. Laino ME, Cancian P, Politi LS, Della Porta MG, Saba L, Savevski V. Generative adversarial networks in brain imaging: a narrative review. J Imaging. 2022;8(4):83. doi:10.3390/jimaging8040083. [Google Scholar] [PubMed] [CrossRef]
175. Wolterink JM, Mukhopadhyay A, Leiner T, Vogl TJ, Bucher AM, Išgum I. Generative adversarial networks: a primer for radiologists. Radiographics. 2021;41(3):840–57. doi:10.1148/rg.2021200151. [Google Scholar] [PubMed] [CrossRef]
176. Lee M, Seok J. Regularization methods for generative adversarial networks: an overview of recent studies. arXiv:2005.09165. 2020. [Google Scholar]
177. Miyato T, Kataoka T, Koyama M, Yoshida Y. Spectral normalization for generative adversarial networks. arXiv:1802.05957. 2018. [Google Scholar]
178. Klambauer G, Unterthiner T, Mayr A, Hochreiter S. Self-normalizing neural networks. In: Advances in neural information processing systems. Cambridge, MA, USA: The MIT Press; 2017. [Google Scholar]
179. Xu L, Zeng X, Huang Z, Li W, Zhang H. Low-dose chest X-ray image super-resolution using generative adversarial nets with spectral normalization. Biomed Signal Process Cont. 2020;55(6):101600. doi:10.1016/j.bspc.2019.101600. [Google Scholar] [CrossRef]
180. Hoang Q, Nguyen TD, Le T, Phung D. MGAN: training generative adversarial nets with multiple generators. In: The 6th International Conference on Learning Representations; 2018 Apr 30–May 3; Vancouver, BC, Canada. p. 1–24. [Google Scholar]
181. Wu Y, Yue Y, Tan X, Wang W, Lu T. End-to-end chromosome Karyotyping with data augmentation using GAN. In: 2018 25th IEEE International Conference on Image Processing (ICIP); 2018 Oct 7–10; Athens, Greece. p. 2456–60. [Google Scholar]
182. Neff T, Payer C, Stern D, Urschler M. Generative adversarial network based synthesis for supervised medical image segmentation. In: Proceedings of the OAGM and ARW Joint Workshop; 2017 May 10–12; Wien, Austria. p. 140–5. [Google Scholar]
183. Saad MM, Rehmani MH, O’Reilly R. Addressing the intra-class mode collapse problem using adaptive input image normalization in gan-based x-ray images. In: 2022 44th Annual International Conference of the IEEE Engineering in Medicine & Biology Society (EMBC); 2022 Jul 11–15; Glasgow, UK. p. 2049–52. [Google Scholar]
184. Saad MM, Rehmani MH, O’Reilly R. A self-attention guided multi-scale gradient GAN for diversified x-ray image synthesis. In: Irish Conference on Artificial Intelligence and Cognitive Science; 2022 Dec 8–9; Munster, Ireland. p. 18–31. [Google Scholar]
185. Xue Y, Zhou Q, Ye J, Long LR, Antani S, Cornwell C, et al. Synthetic augmentation and feature-based filtering for improved cervical histopathology image classification. In: International Conference on Medical Image Computing and Computer-Assisted Intervention; 2019 Oct 13–17; Shenzhen, China: Springer. p. 387–96. [Google Scholar]
186. Kudo A, Kitamura Y, Li Y, Iizuka S, Simo-Serra E. Virtual thin slice: 3D conditional GAN-based super-resolution for CT slice interval. In: International Workshop on Machine Learning for Medical Image Reconstruction; 2019 Oct 17; Shenzhen, China. p. 91–100. doi:10.1007/978-3-030-33843-5_9. [Google Scholar] [CrossRef]
187. Segato A, Corbetta V, Di Marzo M, Pozzi L, De Momi E. Data augmentation of 3D brain environment using deep convolutional refined auto-encoding alpha GAN. IEEE Transact Med Robot Bionics. 2020;3(1):269–72. doi:10.1109/tmrb.2020.3045230. [Google Scholar] [CrossRef]
188. Kwon G, Han C, Kim DS. Generation of 3D brain MRI using auto-encoding generative adversarial networks. In: International Conference on Medical Image Computing and Computer-Assisted Intervention; 2019 Oct 13–17; Shenzhen, China. p. 118–26. [Google Scholar]
189. Qin Z, Liu Z, Zhu P, Xue Y. A GAN-based image synthesis method for skin lesion classification. Comput Meth Prog Biomed. 2020;195(2):105568. doi:10.1016/j.cmpb.2020.105568. [Google Scholar] [PubMed] [CrossRef]
190. Modanwal G, Vellal A, Mazurowski MA. Normalization of breast MRIs using cycle-consistent generative adversarial networks. Comput Meth Prog Biomed. 2021;208(4):106225. doi:10.1016/j.cmpb.2021.106225. [Google Scholar] [PubMed] [CrossRef]
191. Deng J, Dong W, Socher R, Li LJ, Li K, Fei-Fei L. Imagenet: a large-scale hierarchical image database. In: 2009 IEEE Conference on Computer Vision and Pattern Recognition; 2009 Jun 20–25; Miami, FL, USA. p. 248–55. [Google Scholar]
192. Sara U, Akter M, Uddin MS. Image quality assessment through FSIM, SSIM, MSE and PSNR—a comparative study. J Comput Communicat. 2019;7(3):8–18. doi:10.4236/jcc.2019.73002. [Google Scholar] [CrossRef]
193. Korkinof D, Harvey H, Heindl A, Karpati E, Williams G, Rijken T, et al. Perceived realism of high-resolution generative adversarial network-derived synthetic mammograms. Radi Artif Intell. 2021;3(2):e190181. doi:10.1148/ryai.2020190181. [Google Scholar] [PubMed] [CrossRef]
194. Wang Z, Lim G, Ng WY, Tan TE, Lim J, Lim SH, et al. Synthetic artificial intelligence using generative adversarial network for retinal imaging in detection of age-related macular degeneration. Front Med. 2023;10:1184892. doi:10.3389/fmed.2023.1184892. [Google Scholar] [PubMed] [CrossRef]
195. Mardani M, Gong E, Cheng JY, Vasanawala S, Zaharchuk G, Alley M, et al. Deep generative adversarial networks for compressed sensing automates MRI. arXiv:1706.00051. 2017. [Google Scholar]
196. Li Y, Yang H, Xie D, Dreizin D, Zhou F, Wang Z. POCS-augmented CycleGAN for MR image reconstruction. Appl Sci. 2021;12(1):114. doi:10.3390/app12010114. [Google Scholar] [PubMed] [CrossRef]
197. Ehrhardt J, Wilms M. Autoencoders and variational autoencoders in medical image analysis. In: Burgos N, Svoboda D, editors. Biomedical image synthesis and simulation: methods and applications. Cambridge, MA, USA: Academic Press; 2022. p. 129–62. doi:10.1016/b978-0-12-824349-7.00015-3. [Google Scholar] [CrossRef]
198. Kazerouni A, Aghdam EK, Heidari M, Azad R, Fayyaz M, Hacihaliloglu I, et al. Diffusion models in medical imaging: a comprehensive survey. Med Image Anal. 2023;88(1):102846. doi:10.1016/j.media.2023.102846. [Google Scholar] [PubMed] [CrossRef]
Cite This Article
 Copyright © 2026 The Author(s). Published by Tech Science Press.
Copyright © 2026 The Author(s). Published by Tech Science Press.This work is licensed under a Creative Commons Attribution 4.0 International License , which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.


 Submit a Paper
Submit a Paper Propose a Special lssue
Propose a Special lssue View Full Text
View Full Text Download PDF
Download PDF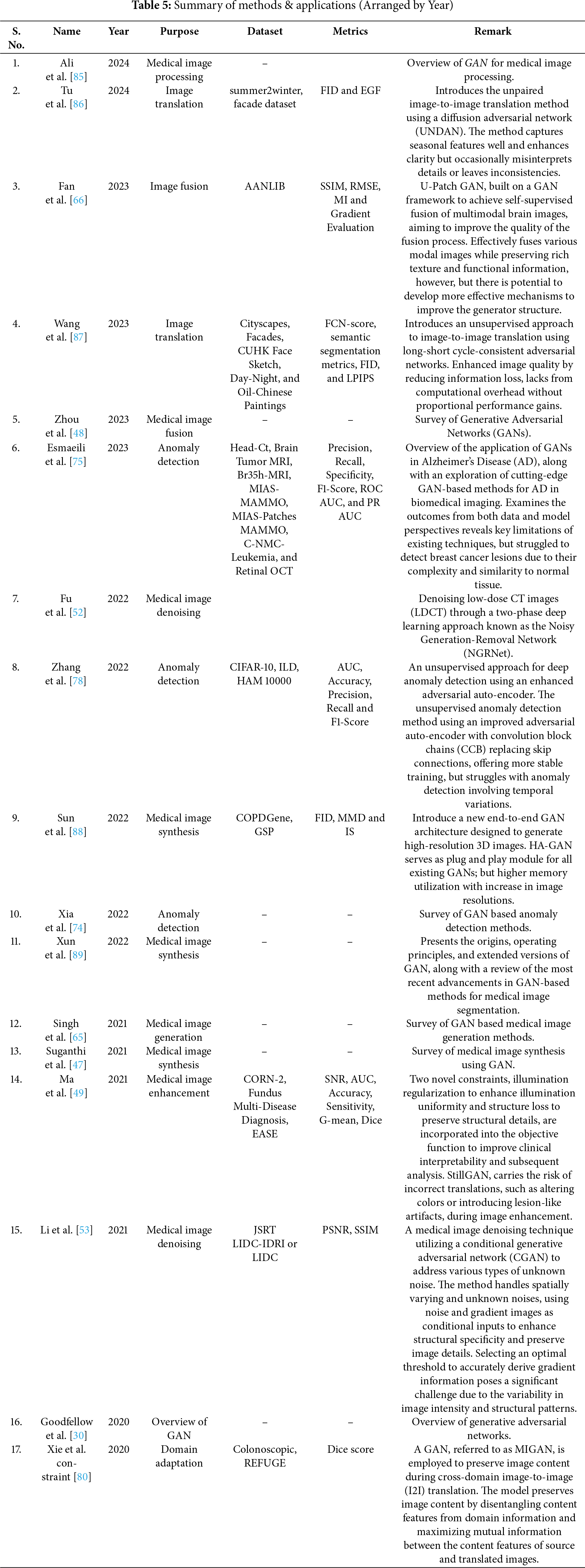
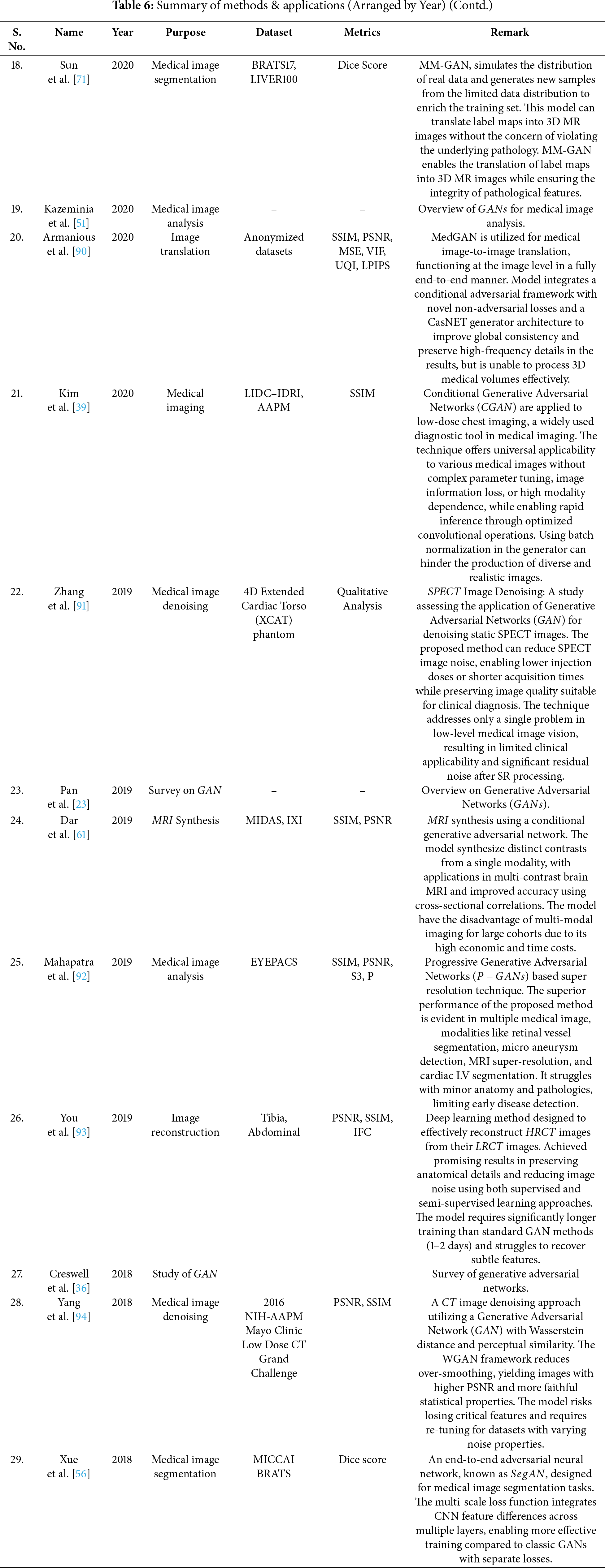

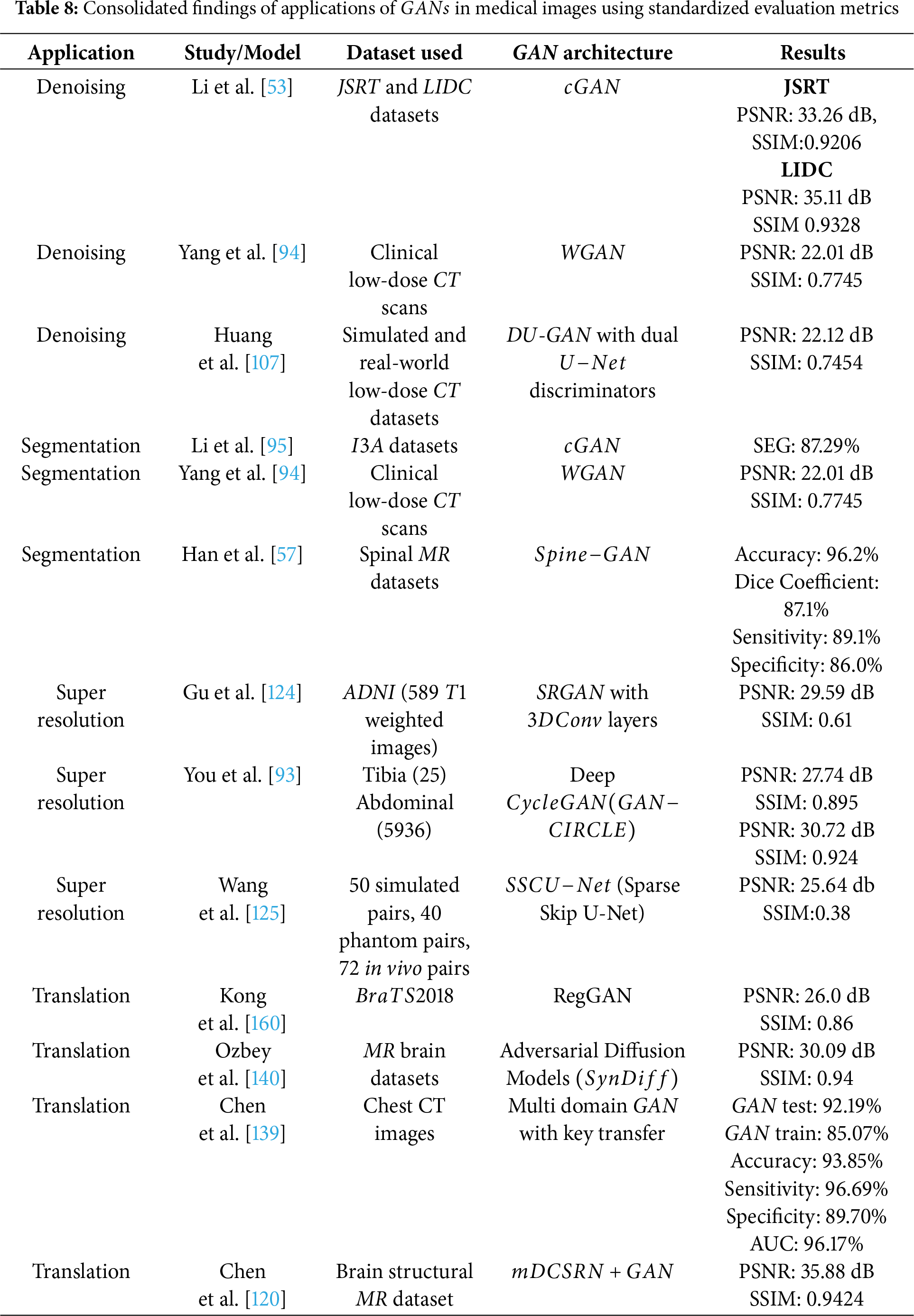
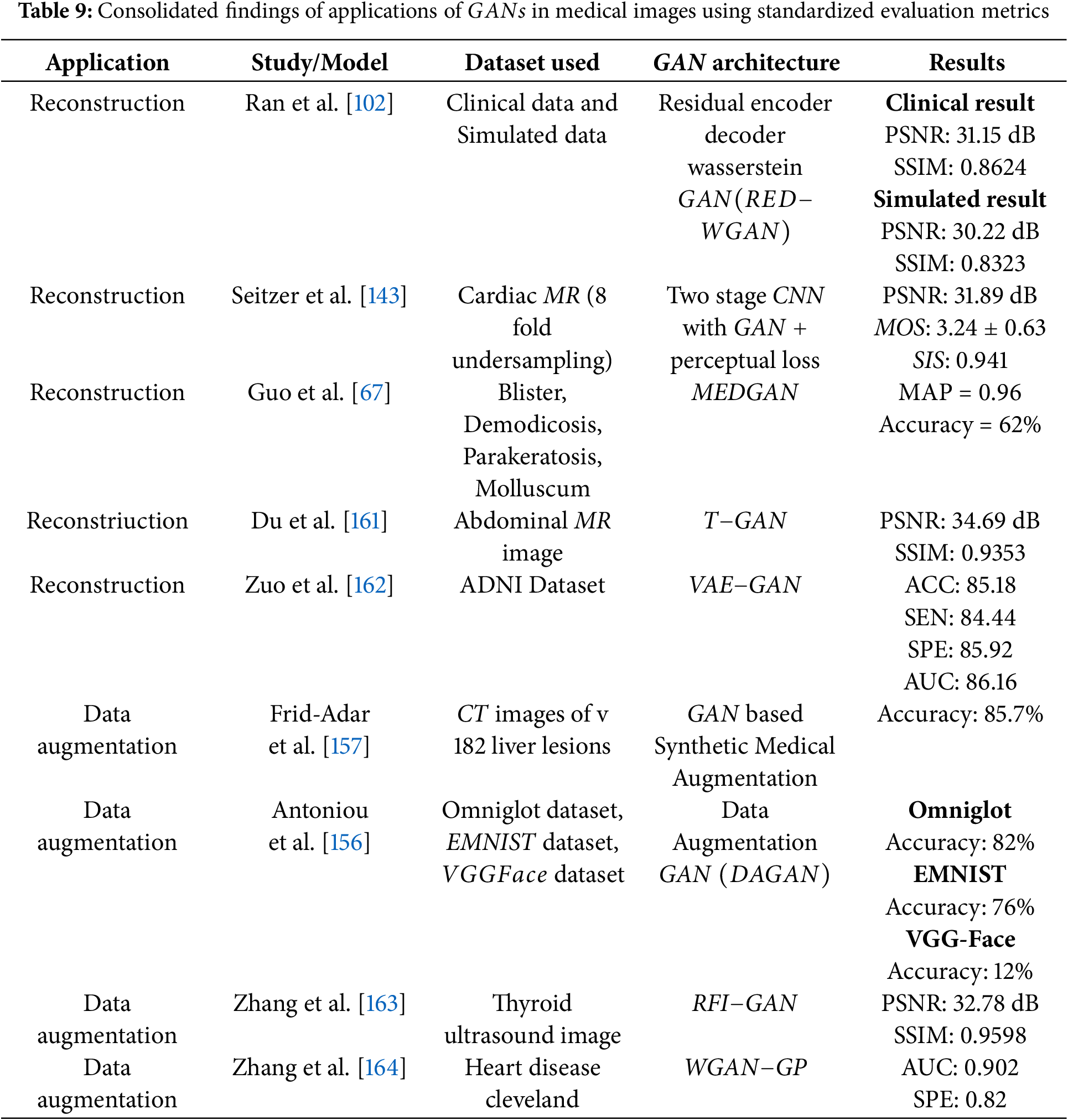
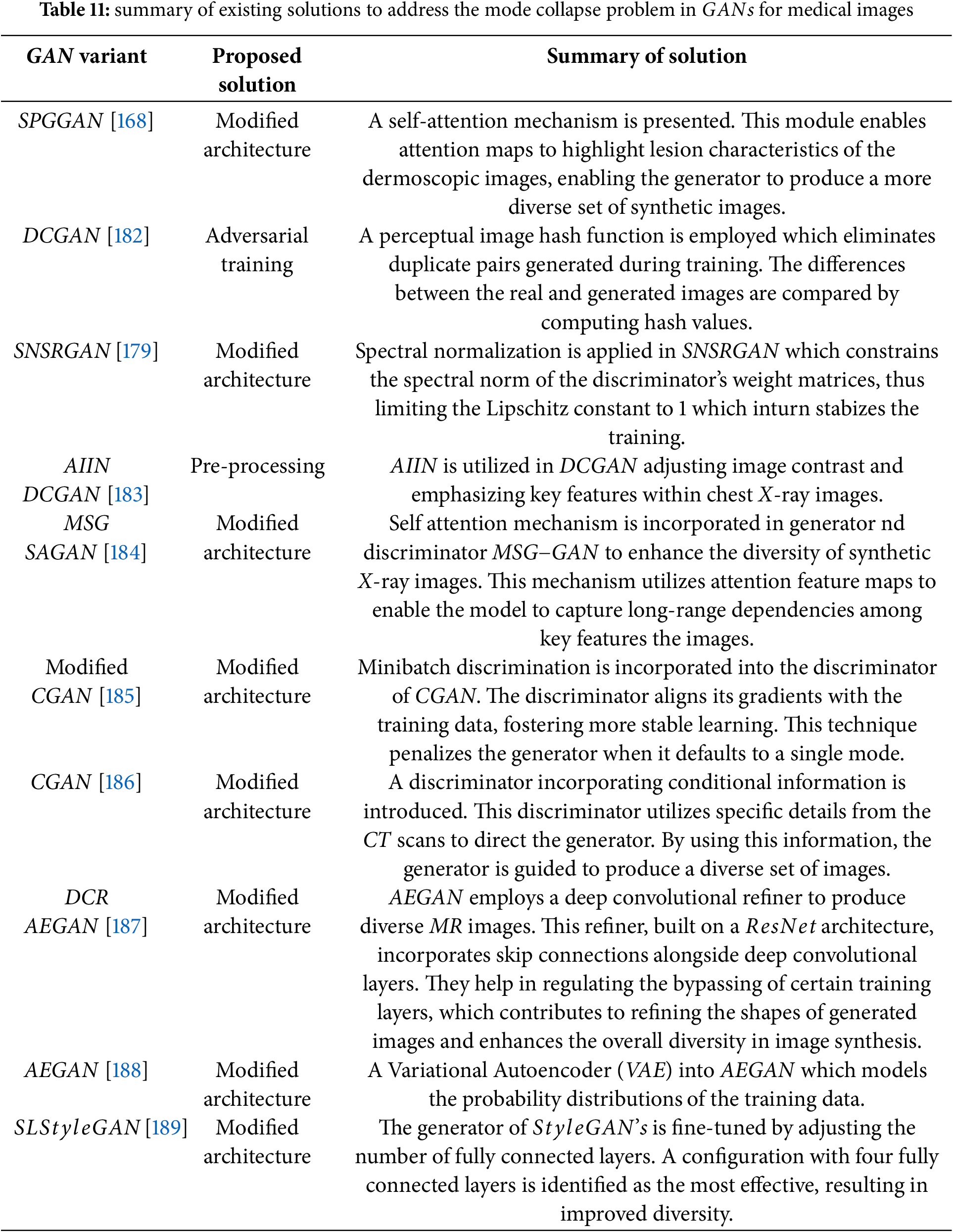

 Downloads
Downloads
 Citation Tools
Citation Tools