 Open Access
Open Access
ARTICLE
CANNSkin: A Convolutional Autoencoder Neural Network-Based Model for Skin Cancer Classification
1 SDAIA-KFUPM Joint Research Center for Artificial Intelligence, King Fahd University of Petroleum and Minerals, Dhahran, 31261, Saudi Arabia
2 Department of Computer Engineering, King Fahd University of Petroleum and Minerals, Dhahran, 31261, Saudi Arabia
3 Information Systems Department, Imam Mohammad Ibn Saud Islamic University (IMSIU), Riyadh, 11432, Saudi Arabia
4 Department of Electrical Engineering, Imam Mohammad Ibn Saud Islamic University (IMSIU), Riyadh, 11432, Saudi Arabia
5 School of Computing, Edinburgh Napier University, Merchiston Campus, Edinburgh, EH10 5DT, UK
* Corresponding Authors: Abdul Jabbar Siddiqui. Email: ; Abdul Khader Jilani Saudagar. Email:
(This article belongs to the Special Issue: Exploring the Impact of Artificial Intelligence on Healthcare: Insights into Data Management, Integration, and Ethical Considerations)
Computer Modeling in Engineering & Sciences 2026, 146(2), 40 https://doi.org/10.32604/cmes.2026.074283
Received 07 October 2025; Accepted 31 December 2025; Issue published 26 February 2026
Abstract
Visual diagnosis of skin cancer is challenging due to subtle inter-class similarities, variations in skin texture, the presence of hair, and inconsistent illumination. Deep learning models have shown promise in assisting early detection, yet their performance is often limited by the severe class imbalance present in dermoscopic datasets. This paper proposes CANNSkin, a skin cancer classification framework that integrates a convolutional autoencoder with latent-space oversampling to address this imbalance. The autoencoder is trained to reconstruct lesion images, and its latent embeddings are used as features for classification. To enhance minority-class representation, the Synthetic Minority Oversampling Technique (SMOTE) is applied directly to the latent vectors before classifier training. The encoder and classifier are first trained independently and later fine-tuned end-to-end. On the HAM10000 dataset, CANNSkin achieves an accuracy ofKeywords
Developing an efficient system to automatically categorize skin cancer is essential, considering the increasing incidence of skin cancer and the need for early identification [1,2]. Skin cancer is broadly classified into Melanoma and Non-melanoma, with 324,635 and 1,198,073 new cases reported globally in 2020, respectively [3]. Skin melanoma ranks 17th in the world in terms of frequency of malignancy. Statistics on non-melanoma skin cancer are frequently withheld and often underreported due to its high prevalence and frequent misdiagnosis [3]. But when the illness spreads below the skin, there is little chance of survival. Furthermore, the accuracy of the diagnosis usually depends on the dermatologist’s experience because early signs of skin cancer may not be obvious [4]. For more accurate diagnoses, an automated system is a vital tool that can aid detection and diagnosis. In addition, it is very difficult and rarely generalizable to diagnose skin cancer with the unaided eye [5]. Thus, it is imperative to develop an automatic skin cancer classification system that is effective and efficient in terms of speed, cost, and precision [6].
However, it can be difficult to achieve automatic skin cancer categorization due to the complexity and diversity of images depicting skin diseases [5]. First, a misclassification could occur due to the many interclass similarities across various skin lesions. Second, skin lesions from the same class also differ in feature, size, colour, and location, making it difficult to classify skin lesions from the same class. Traditionally, handcrafted feature extraction methods like Menzies Method and ABCD Rule [7], are used to extract features from skin disease images. These features are fed into machine learning algorithms like Support Vector Machines (SVM), K-Nearest Neighbors (KNN), and XGBoost for classification [8,9]. However, it is ineffective to distinguish each type of skin cancer only by hand-crafted features because of the difficulty in generalizing to a wider range of disease types and the limited number of specified features. Skin cancer detection and classification as a computer vision problem has received attention from researchers around the globe. Their works implemented several deep learning models, especially using CNN and some machine learning-based models, including SVM, KNN, DT, etc. [7], each with their corresponding individual strengths and weaknesses. However, skin cancer datasets are typically highly imbalanced datasets, and properly addressing the issue is a key factor that affects the model performance. Similarly, while diverse metrics are used to assess the models, evaluating the models on metrics that provide insight into the overall as well as class-wise performance can reveal if the models perform well across the multi-class lesions or if there are specific challenges for certain classes.
Moreover, the robustness of the model to classify skin lesions from different datasets on which it was not trained further determines its generalization ability. In this work, we propose an improved method to enable enhanced skin cancer classification and diagnosis, demonstrating remarkable results on a popular dataset. The overall contributions made by this paper can be outlined as follows:
1. Propose a deep learning-based novel method for automated skin cancer classification and diagnosis based on convolutional autoencoders and neural networks.
2. Address the issue of class imbalance by leveraging latent embeddings from the proposed model and SMOTE, demonstrating enhanced performance in detecting minority skin cancer classes.
3. Conducted a comprehensive performance evaluation of the proposed model on a publicly available dataset using different performance metrics and demonstrated superiority over state-of-the-art competitive methods.
The succeeding sections of the paper are organized as follows: Section 2 discusses the existing works related to skin disease classification. Section 3 depicts the methodology used in the proposed work for skin disease classification. Section 4 discusses the Experimental setup; and Section 5 shows the results and detailed discussion on it, and finally, Section 6 concludes the paper.
Skin cancer poses a significant threat to one’s life, especially when it remains undetected and untreated during its initial stages, and over the years, researchers have developed various models to help medical professionals in its detection and classification. These approaches could be broadly grouped into: (i) classical machine learning-based approaches, and (ii) deep learning-based approaches.
2.1 Classical Machine Learning-Based Approaches
In the past, researchers have used various means to extract the features from the image datasets before feeding these into conventional machine learning models. These features main were extracted based on handcrafted methods, image processing and feature engineering [10], rule-based approach (e.g., ABCD rule (Asymmetry, Border, Color, and Diameter)) [11,12], statistical histogram-based approach [13,14], and others.
For example, some works utilized features like homogeneity, contrast, energy, correlation, and entropy derived from the Gray-Level Co-occurrence matrix (GLCM) feature extractor, which were fed into the KNN, Artificial Neural Networks (ANN), Naïve Bayes, Decision Tree, and SVM machine learning classifiers.
2.2 Deep Learning-Based Approaches
With the coming of deep learning and computer vision, the approach to skin cancer detection and classification has shifted significantly.
Efforts have also been made to study efficient deep learning models for skin cancer classification. For example, reference [15] developed eight EfficientNet models with varying depth to capture complex features of the dataset. Their work performed transfer learning by finetuning the baseline EfficientNet through removing top 3 layers and replacing them with a dense layer, batch normalization, dropout layer, and employing the swish activation function. However, reduced discriminating power was noted in low-level EfficientNet models B0-B3, and mid-level EfficientNet (B4 and B5) have outstanding performance that makes intuitive sense. High-level models (B6–B7) were overfitting the data. Hence, it was observed that there is a trade-off between model depth and performance. The performance of the best model, B4, which has an overall accuracy of 87.9% (HAM10000 skin cancer dataset), is heavily impaired by the class imbalance issue; hence, the model performed poorly in classifying the minority classes.
Some works explored hybrid approaches, such as the work of [7], who proposed the use of transfer learning and ensemble learning to automate and improve skin cancer classification. Pretrained deep learning models were used as feature extractors for machine learning models before their classifications were ensembled into a logistic regression model using Stacking. The authors proposed the use of three pretrained deep neural networks, ResNet50, VGC19, and Xception networks. Their features were fed into machine learning models like Random Forest, KNN, SVM, Gaussian NB, AdaBoost, decision tree, and regression. These models served as the base model for the stacking algorithm. Each prediction of these base models was ensembled using stacking and fed into a neural node for classification.
To enhance the learning of fine-grained lesion information across different spatial scales, prior work has explored various attention-driven and clinically inspired strategies. For example, reference [16] incorporated multi-scale triplet attention modules into pretrained CNN backbones to emphasize salient lesion regions through spatial and channel attention. Although this improves representation quality, imbalance is still handled only through basic image augmentations such as rotation, shear, and zoom, which may increase sample count but do not correct feature-space skewness.
More recently, CI-Net [17] has introduced a clinically inspired architecture that explicitly models the diagnostic behavior of dermatologists by simulating zoom, observe, compare, and distinguish steps. Through lesion-area attention, feature extraction, lesion feature attention, and a distinguishing module, CI-Net captures domain-relevant cues and achieves strong performance across multiple ISIC datasets. However, despite its interpretability and attention mechanisms, CI-Net still relies on conventional augmentation to address data imbalance.
While these approaches effectively enrich lesion representations and integrate expert knowledge, they continue to depend heavily on pixel-space augmentation and attention gating. These do not directly tackle the core issue of class imbalance at the representation level.
With the advent of transformer-based models, researchers have studied their application to the skin cancer classification problem. One such work is of [18], which used a vision transformer model to perform skin cancer classification. It is aimed at solving the dimensionality reduction problem associated with CNN-based models. The authors employed extensive data augmentation to tackle the effect of class imbalance associated with datasets used in training skin cancer classification models. The authors proposed a preprocessing data augmentation module that augments data classes in the dataset that have a lower number of images. Different augmentation techniques, like rotation, flip, contrast, and scaling, were performed. The vision transformer proposed in their paper consists of the embedding layer, encoder layer, and classifier. At the embedding layer, the patches are embedded with positional embedding and concatenated with a learnable class token. These embedded patches are then fed into the encoder layer. The encoder layer contains a series of multi-head self-attention modules and a multilayer perceptron. The classifier layer classifies the patches into different classes. The transformer-based model in [18] performed excellently well for each class’ classification and also outperformed the state-of-the-art models studied in the paper. However, their model is characterized by a large model size and a large number of parameters, making it difficult to implement on resource-constrained devices.
Some efforts have been made to perform segmentation in addition to classification ok skin cancer. For instance, reference [19] proposed a two-stage network approach involving segmentation and classification using atrous residual convolutional networks. Their work subjected the input images to patching in order to enhance inter-class classification performance. Their model was trained and evaluated on the HAM10000 image dataset and has shown classification performance better than some pretrained models including UNet, PSPNet, and LinkNet.
Recent reviews in broader oncology, such as Colorectal-focused AI surveys [8] and systematic XAI analyses across cancer types [20] have emphasized that data imbalance and lack of representation-level correction remain dominant barriers to robust generalization, even in high-performing deep models. They highlight that attention mechanisms and XAI visualizations (e.g., Grad-CAM, SHAP) improve transparency but do not inherently remedy skewed feature manifolds. Overall, it is evident that the intrinsic imbalance of the skin cancer datasets is not fully addressed in prior works. While data augmentation is used in some works to increase the number of samples [21], the approach may not be readily effective and applicable to skin cancer classification, as some of the transformations, such as contrast change, may distort the naturally occurring form of the disease. In this work, we develop an enhanced deep learning-based model for skin cancer classification that strategically integrates convolutional autoencoder and neural networks, as shown in Fig. 1, effectively addressing the issue of class imbalance. Furthermore, we leverage the Grad-CAM methods for interpretability of the model’s predictions.
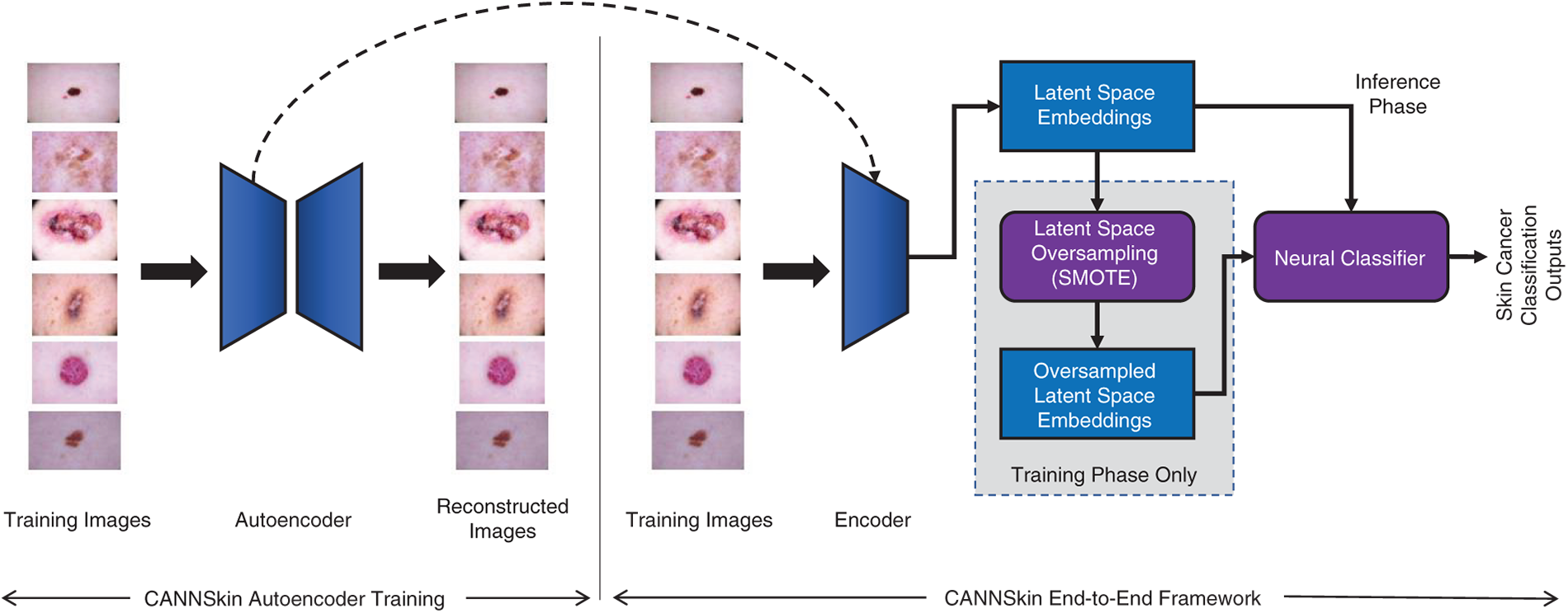
Figure 1: The proposed CANNSkin’s architecture and framework
Skin cancer detection has always been challenging for dermatologists; however, with the aid of deep neural networks, early detection of this disease could be possible and consequently make a positive impact to increasing the survival rate of the cancer patients. With this in mind, this work proposes a novel convolutional autoencoder and neural network-based model for skin cancer classification, which addresses the data imbalance issue.
The proposed method comprises three main components: (i) convolutional encoder, (ii) latent space oversampling (for data balancing), and (iii) neural classifier, as depicted in Fig. 1. The convolutional encoder is trained and extracted from an autoencoder model that is trained to regenerate the input skin images. Once trained, the encoder produces latent embeddings of the skin images. The data balancing module, i.e., the latent space oversampling, is utilized in the training phase only. Specifically, the latent embeddings of the training set’s skin images are used with SMOTE [22] to generate more samples of the minority classes. The resulting balanced set of latent embeddings is then used to train the neural classifier.
In the training phase, the convolutional autoencoder and the neural classifier are first independently trained. Then, the convolutional encoder is integrated with the neural classifier for end-to-end fine-tuning. As depicted in Fig. 1, in inference, the input skin images are transformed into latent embeddings by the encoder, and the neural classifier uses these embeddings for skin cancer classification.
Autoencoders [23] are neural networks that are primarily designed to encode input data in a way that allows for efficient and accurate reconstruction. By learning to compress the input data into a latent-space representation and then reconstructing the output from this representation, autoencoders are effective in applications such as dimensionality reduction, feature learning, and anomaly detection. Fig. 2 shows a simplified architecture of an autoencoder.
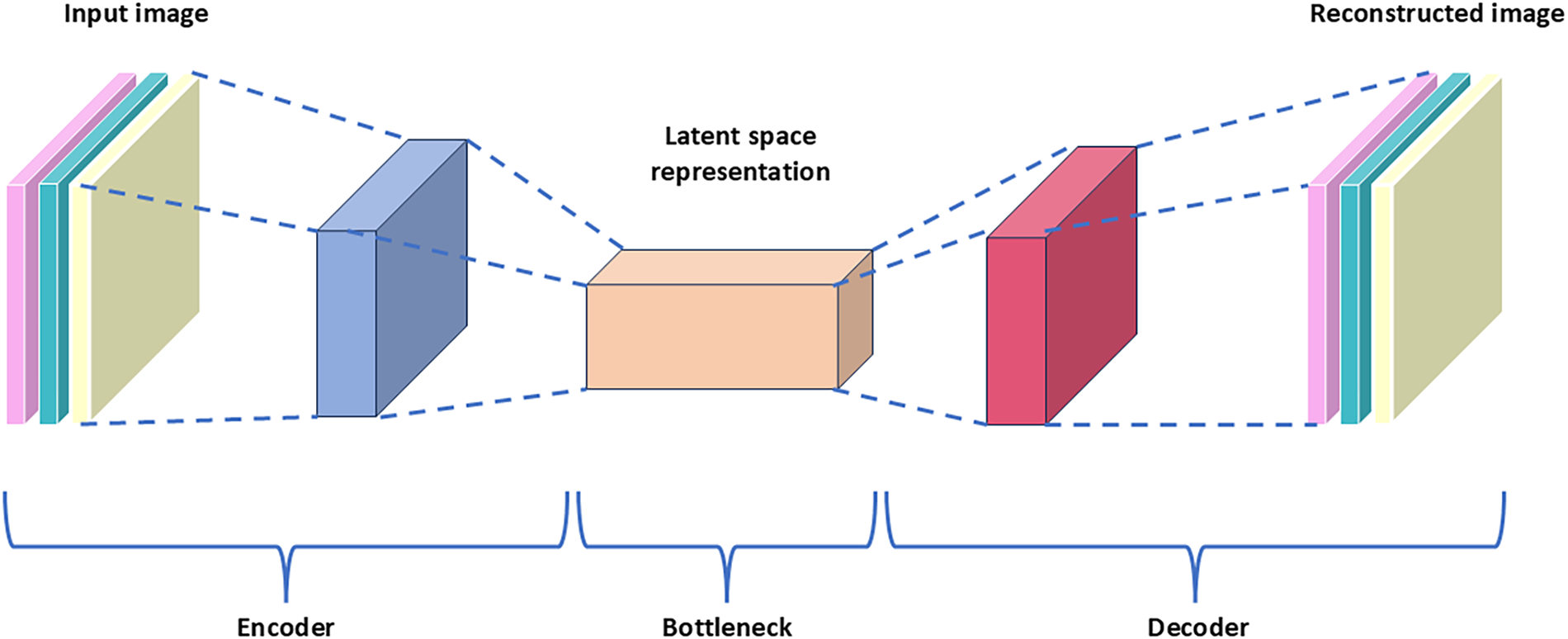
Figure 2: Simplified architecture of an autoencoder
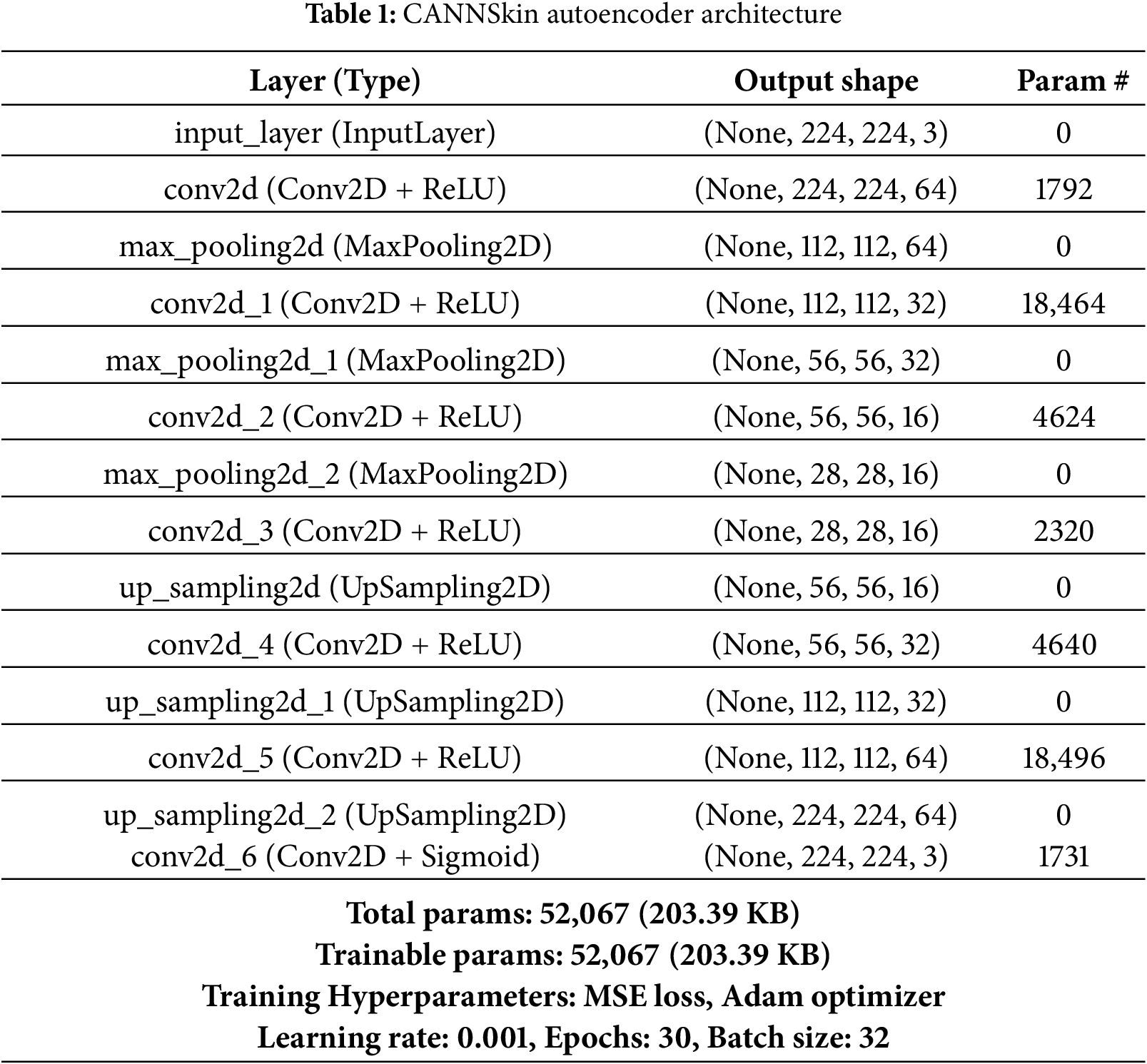
The encoder part of the autoencoder compresses the input images into a latent space representation of the input. This is achieved by using a series of layers that progressively reduce the dimensionality of the input images. The latent space generated is a compressed lower-dimensional representation of the input data, and it contains essential features that can represent the input images. The decoder part of the autoencoder reconstructs the input data from the latent space. During training, the goal of the autoencoder is to minimize the difference between the input data and the reconstructed output. Typically, mean squared error is used as a loss function for training. In this work, we employ the architecture in Table 1 for the CANNSkin’s autoencoder.
In order to develop a skin cancer classification model while combating the issue of class imbalance plaguing automatic medical image diagnosis training, we propose training a deep autoencoder that could reconstruct skin lesion images and propose oversampling of latent space embeddings of the skin images. The latent space represents the compressed, lower dimension representation of the input images. Leveraging the latent space embeddings from the encoder of the autoencoder as features, oversampling techniques could be used to generate oversampled latent space embeddings of minority classes. In this work, we proposed applying the Synthetic Minority Over-sampling Technique (SMOTE) [22] to oversample the latent space embeddings of each image in order to balance the training data of the neural classifier to address the issue of class imbalance. SMOTE is an oversampling technique that works by generating synthetic samples for the minority class(es) to balance the class distribution. It identifies the minority class samples in the dataset and finds k nearest neighbors of individual samples using the Euclidean distance in the feature space. The flowchart of SMOTE is shown in Fig. 3.
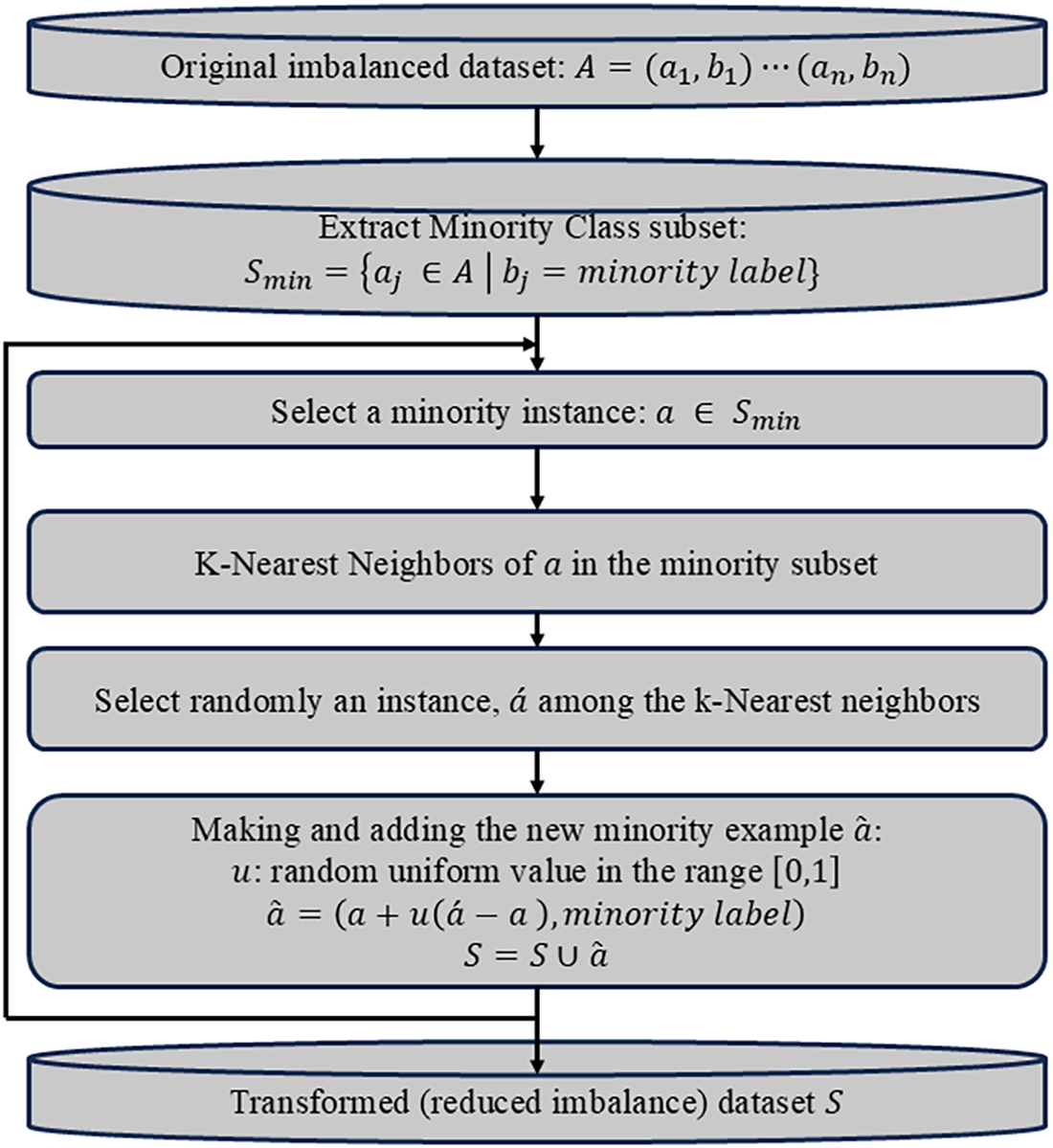
Figure 3: Flowchart outlining the key steps of SMOTE
Traditional oversampling techniques, such as SMOTE, operate by interpolating between existing samples in the feature space. When applied directly in the pixel domain, interpolation between flattened images produces unrealistic samples because natural images lie on a highly non-linear low-dimensional manifold. Also, pixel-wise linear combinations distort spatial structure, break local correlations, and introduce artifacts that do not correspond to valid dermatological patterns. This limitation has been widely reported in imbalanced-learning studies, where SMOTE is shown to be unsuitable for raw image tensors.
In contrast, the latent space produced by our convolutional encoder is a compact, semantically meaningful representation in which Euclidean interpolation is more appropriate. The encoder extracts high-level lesion characteristics such as texture, color distribution, border irregularity, and structural patterns. Therefore, synthetic embeddings generated by SMOTE remain on the manifold of plausible lesion features and preserve clinically relevant characteristics. This makes latent-space oversampling theoretically consistent with the assumptions of SMOTE and avoids the geometric distortions that occur in the image domain.
Moreover, common image-level data augmentation (rotation, flipping, scaling) cannot correct class imbalance, as it increases all classes equally, and may introduce artificial invariances that are clinically unrealistic. Applying SMOTE on Latent-space embeddings directly increases minority-class density in the feature manifold and thus provides a principled way to balance the dataset for the classifier.
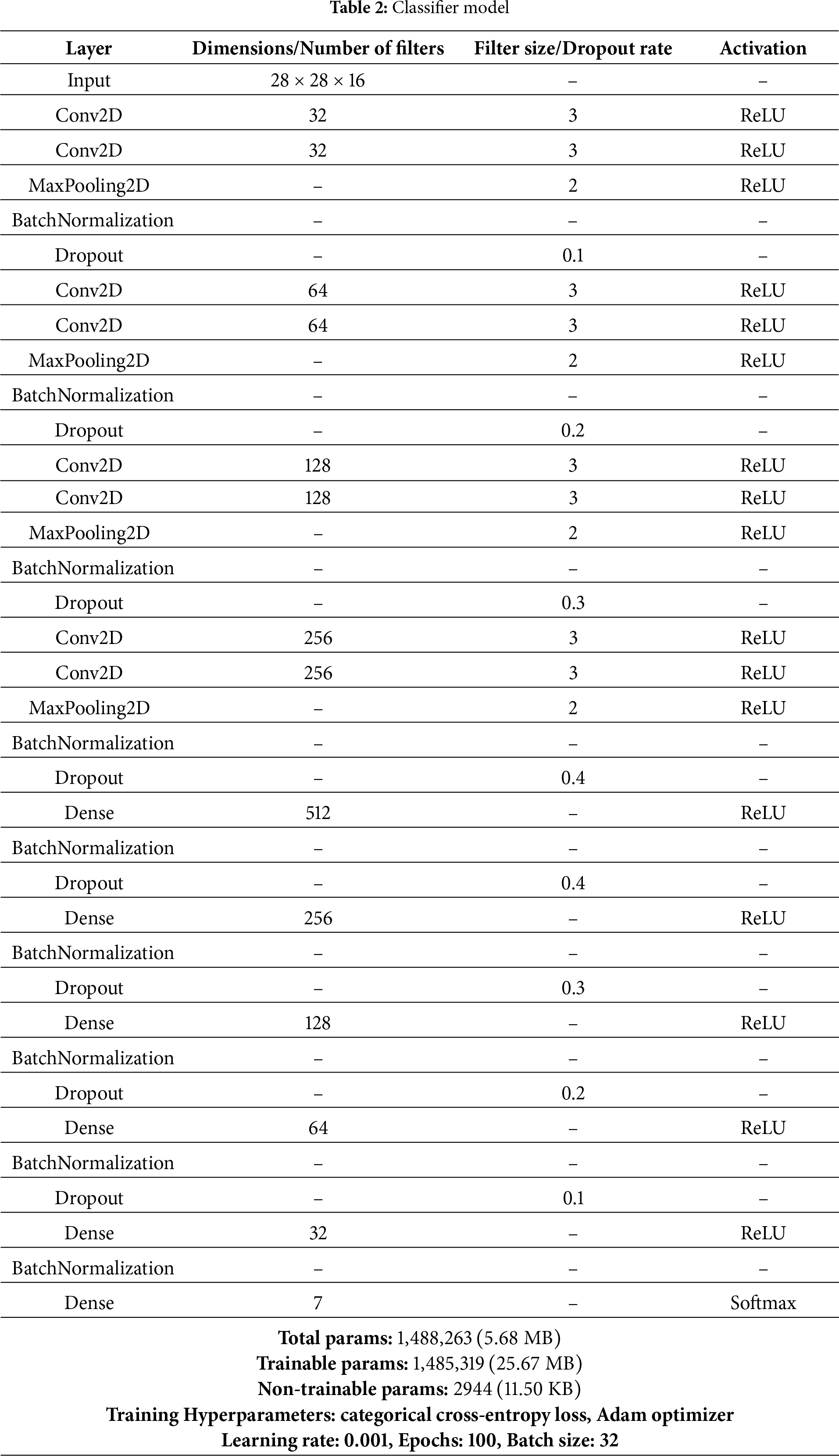
The classifier model in the CANNSkin framework proposed in this study is a convolutional neural network. It is a supervised deep learning model that uses labeled data to train a model for classification. The oversampled latent space embeddings of skin images and their associated ground truth are fed into the series of fully connected neural network layers. In addition, categorical cross-entropy is utilized as a loss function for the multiclass classification of skin cancer. The architecture of the proposed neural classifier is presented in Table 2.
HAM10000 Dataset
In this work, we used the images of skin lesions from the HAM10000 dataset of [24]. The “Human Against Machine” (HAM10000) dataset has made a substantial contribution to the medical image analysis and dermatological fields. It is made up of 10,015 dermatoscopic pictures of typical pigmented skin lesions, which are essential for melanoma research and skin cancer detection.
The goal of the dataset is to improve the precision and effectiveness of dermatological diagnostics by supporting the development and assessment of machine learning algorithms for automated skin lesion classification. The classes present in the dataset are shown in Fig. 4 and they include melanocytic nevi (nv), benign keratosis-like lesions (bkl), basal cell carcinoma (bcc), melanoma (mel), dermatofibroma (df), vascular lesions (vasc), and actinic keratoses and intraepithelial carcinoma (akiec).
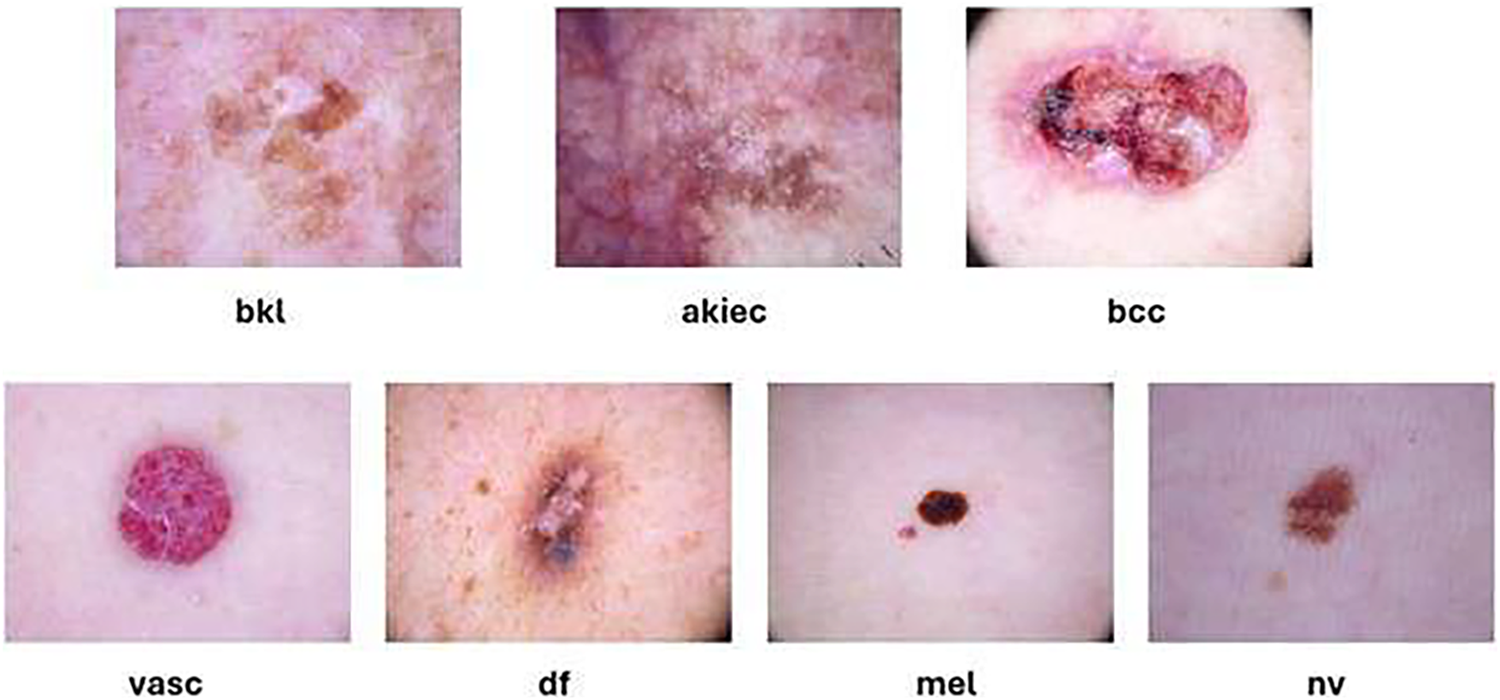
Figure 4: Samples from the HAM10000 dataset
The dataset also includes the metadata of the skin lesion images, which include the localisation, unique identifier, age, and sex of the patients. The localization explains where the skin lesion picture was taken from the patients’ bodies. Fig. 5 shows the distribution of the different classes of skin lesions in the dataset. Nonetheless, just like other skin cancer datasets, HAM10000 [24] classes are highly imbalanced as shown in Fig. 2. To ensure a consistent and reproducible evaluation pipeline, we adopted a fixed train–validation–test split of the HAM10000 dataset. Specifically, 90% of the images were allocated to the training/validation pool, and the remaining 10% were held out as an unseen test set. The training portion was further divided into 90% for training the autoencoder and 10% for validation, yielding an effective 81/9/10 split. This same partitioning strategy was used throughout all stages of the CANNSkin framework. All images were preprocessed using a standardized pipeline to ensure input consistency. Each image was converted to RGB format, resized to
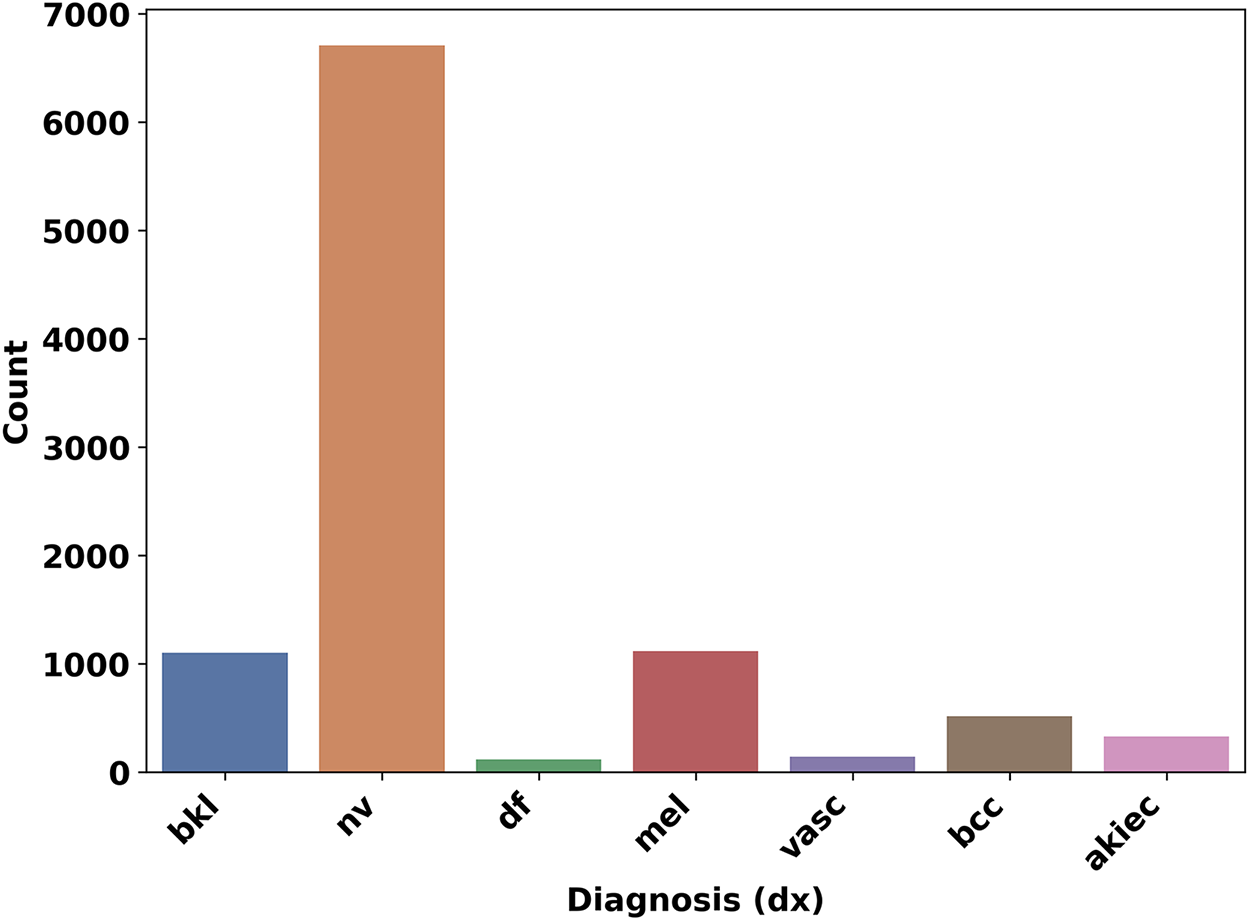
Figure 5: Dataset Classes and their count
Data augmentation was intentionally not used for class balancing, as it increases all classes uniformly and does not address dataset imbalance. Instead, minority-class oversampling was performed exclusively in the latent space using SMOTE. This preprocessing strategy ensures that both the autoencoder and classifier receive consistent and undistorted inputs.
ISIC 2019 Dataset
To further evaluate the robustness of the proposed CANNSkin framework, we additionally utilized the ISIC 2019 dataset [25], a large-scale dermoscopic benchmark released as part of the International Skin Imaging Collaboration (ISIC) challenge. The dataset contains 25,331 dermoscopic images spanning eight diagnostic categories, including melanoma, melanocytic nevus, basal cell carcinoma, actinic keratosis, benign keratosis, dermatofibroma, vascular lesions, and squamous cell carcinoma. Compared to HAM10000, ISIC 2019 is substantially more heterogeneous, incorporating images from multiple clinical centers, devices, and acquisition settings, and displays more pronounced class imbalance.
To ensure consistency across datasets, we applied the same preprocessing pipeline and train–validation–test strategy we used with HAM10000. All images were converted to RGB format, resized to
The architecture of the autoencoder used, as shown in Table 1 was initialized and trained on the train set for 30 epochs while validating with the validation set. Rectified Linear Unit (ReLU) was used in all convolution layers except the output layer, where the Sigmoid function was used. Moreover, the training and validation losses were monitored over the epochs (see Fig. 6). The loss function, mean squared error (MSE), Batch size of 32, and learning rate of 0.001, were used for training, and it was optimized using Adam’s optimizer. In order to check the reconstruction performance of the autoencoder, structural similarity (SSIM) and peak signal-to-noise ratio (PSNR) were used to evaluate it. SSIM considers changes in structural information, luminance, and contrast. These three components are combined to produce a similarity index value that ranges from -1 to 1, where 1 indicates identical images. Its equation is given as
where
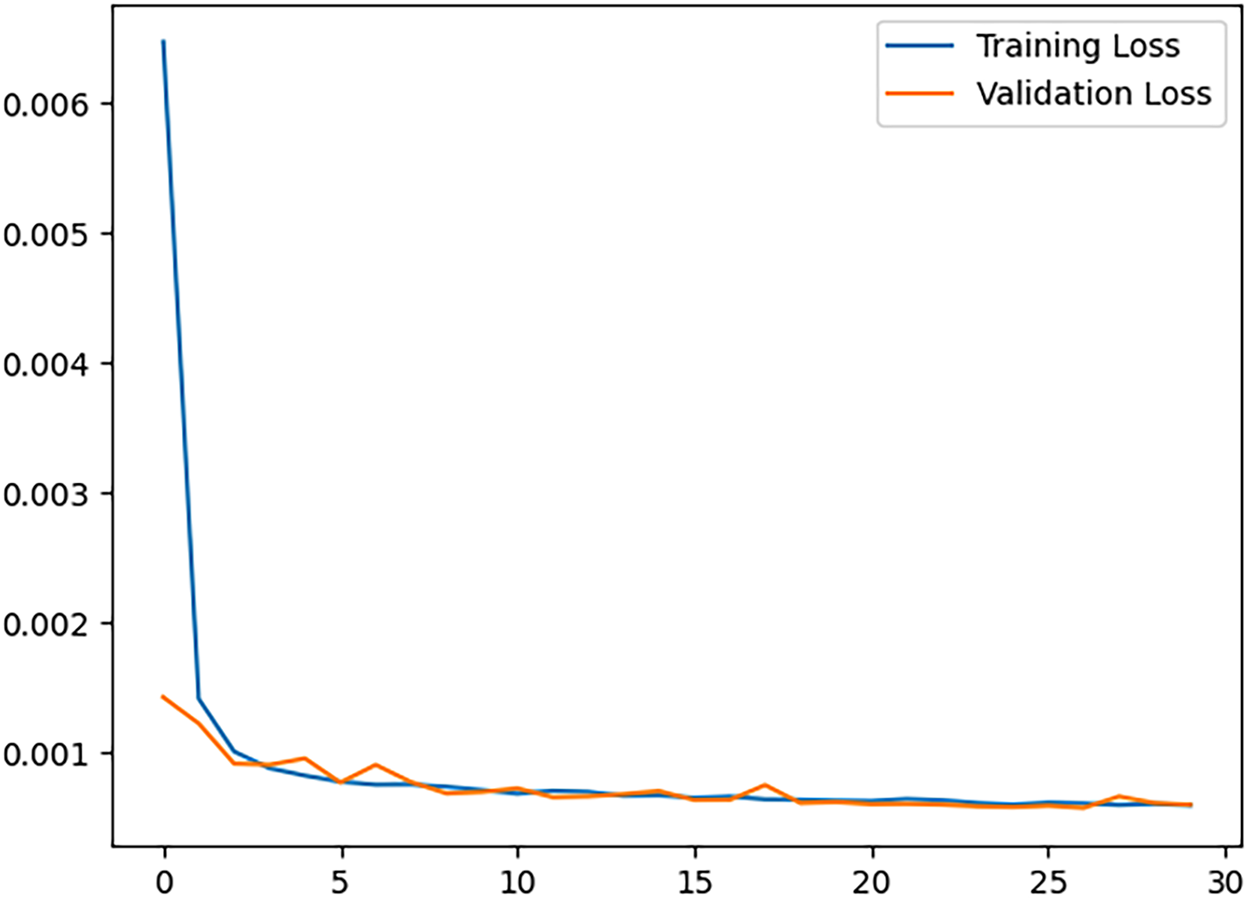
Figure 6: Convolutional autoencoder training and validation loss
The trained encoder is used to generate latent space representations of the whole image dataset, which was divided into training, validation, and testing samples. Then, the training samples were fed into the SMOTE algorithm, using the scikit learn library [3], and the test set was used for evaluating the models.
The oversampled dataset is divided into 81% train set, 9% validation set, and 10% test set. The architecture of the classifier model is shown in Table 2. SoftMax was used for classification into 7 classes and the Adam optimizer was employed for training. The classifier model was trained for 60 epochs, ensuring there is no overfitting.
In order to effectively evaluate our model, we divided our work into four experiments. Using the trained encoder (described in Section 3.2), the first two experiments involve training classifier models independently of the trained encoder model, i.e., the latent space representations of the dataset are generated and obtained before training begins. Depending on how the classifier models are trained, there are four versions: M1, M2, M3, and M4.
• Model M1: classifier is trained on imbalanced latent space representations of the skin cancer dataset; the encoder weights are not fine-tuned.
• Model M2: classifier is trained on the balanced dataset of latent space embeddings after oversampling is done using SMOTE on the training samples; the encoder weights are not fine-tuned.
• Model M3: The model M3 refers to the model obtained by training using the CANNSkin end-to-end framework to fine-tune the trained encoder model and the trained classifier model M1 in an end-to-end manner.
• Model M4: The model M4 was developed through integrated fine-tuning of the trained encoder model and the trained classifier model M2 in an end-to-end fashion.
To develop M3 and M4, the trained encoder model and the trained classifier models were integrated and re-trained (fine-tuned) as one end-to-end model, as depicted in Fig. 1 (right). In training the M1 and M2, the independently trained encoder model was frozen (no training of its weights was performed).
All experiments were done in a Google Colab Pro environment using the TPUv2. Important libraries used include TensorFlow, Matplotlib, Scikit Learn, Pandas, and Numpy.
Popular evaluation metrics in any classification problem are expressed in the form of True Positive (TP), False Negative (FN), true negative (TN) and false positive (FP), and balanced multiclass accuracy. A false negative denotes the amount of incorrect classification of the negative class. True positive is the number of correctly classified positive cases. True Negative means the number of correctly detected negative classes. A false positive is the amount of incorrect classification of the positive class.
• Precision denotes rate of true positive to the sum of positives.
• Accuracy (ACC) means rate of the total correctly classified data samples to the sum of data samples.
• Recall (True positive rate) signifies the rate of correctly classified malicious samples to sum of malicious data samples.
This is also the class-wise accuracy of the multiclass classification.
• False Positive Rate signifies the rate of incorrectly classified benign samples to the sum of all benign data.
• F1 score is expressed as:
5.1 Evaluating the CANNSkin Autoencoder
In this work, we developed an efficient autoencoder capable of reconstructing the input skin cancer images. The autoencoder was evaluated using SSIM, PSNR, and visual inspection. High SSIM and PSNR values of 97% and 31 dB were recorded over the entire test set, and visual inspection of the test images signifies there are no many visual differences between the input images and the reconstructed ones. Table 3 presents the results of PSNR per individual classes of the image dataset, while Fig. 7 shows some reconstructed images by the autoencoder together with their corresponding original images. The trained encoder part of the autoencoder is used for subsequent experiments.
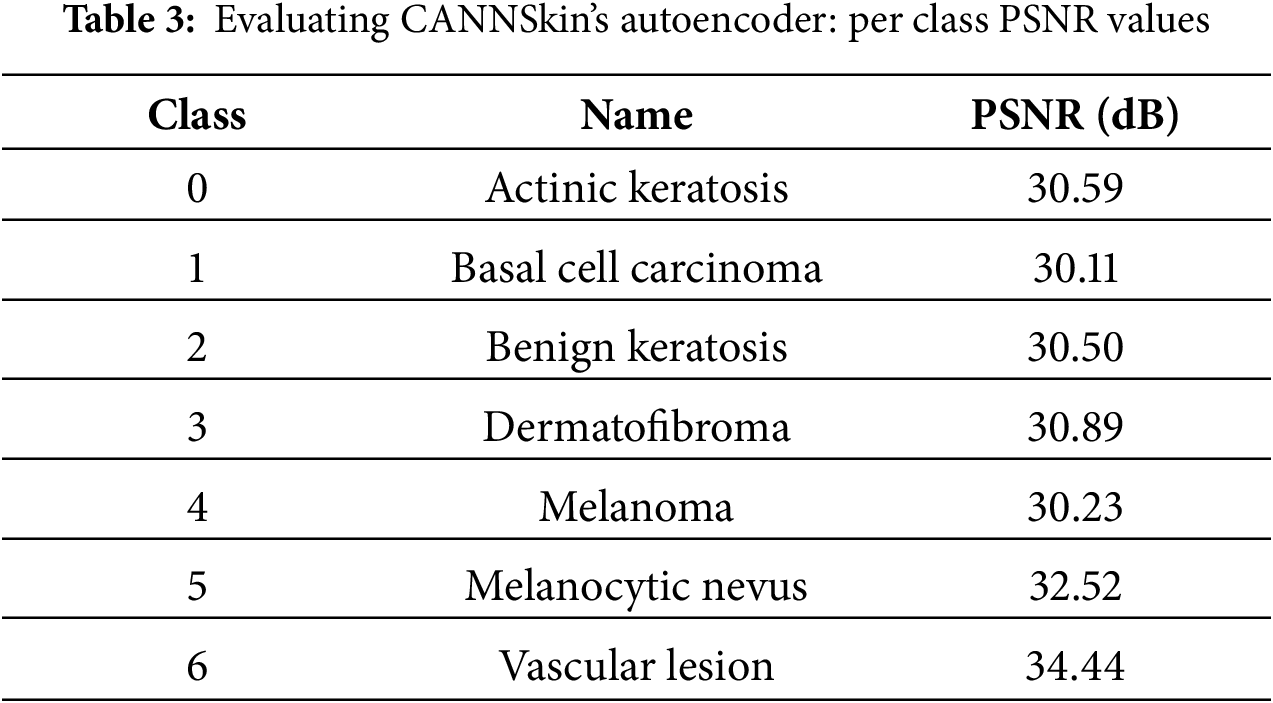
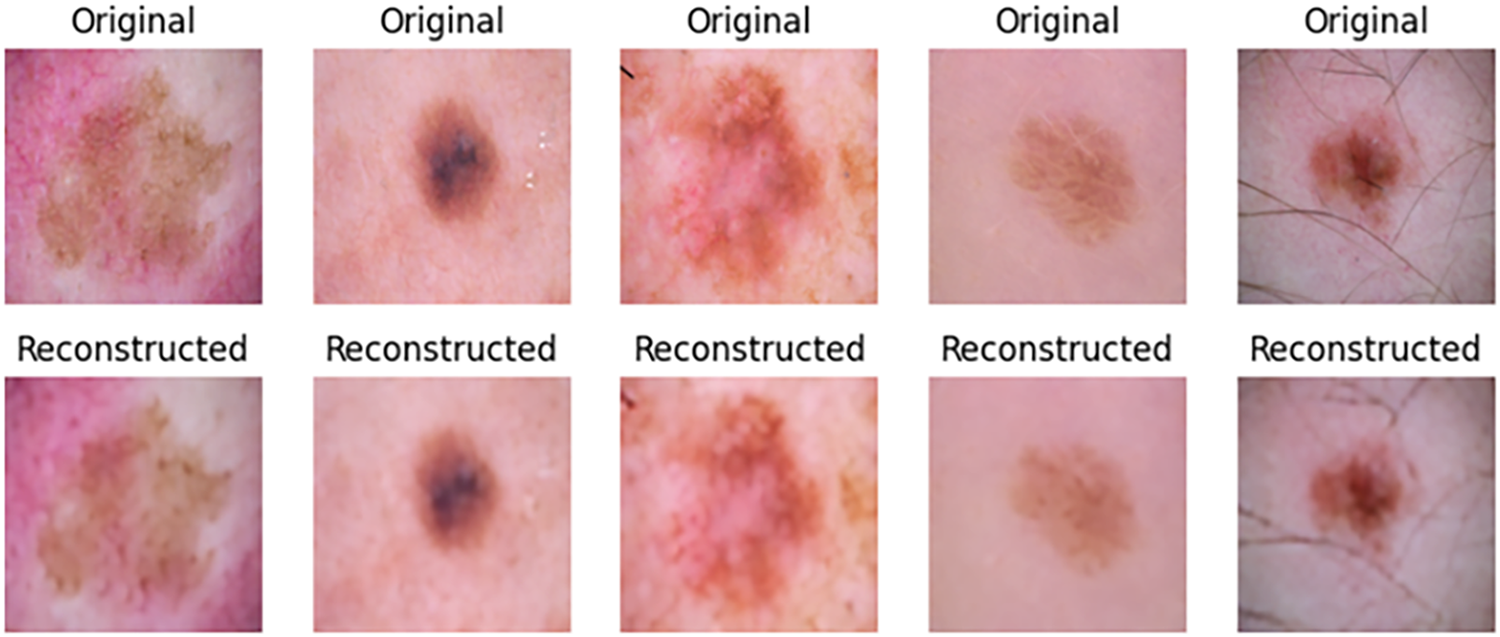
Figure 7: Samples of reconstructed images from CANNSkin’s trained convolutional autoencoder
5.2 Inter-Class Classification Performance
Models M1 and M3 trained on imbalanced latent vectors and imbalanced data samples, respectively, were biased towards the class with the highest number of samples, as shown respectively in Tables 4 and 5 and the confusion matrices shown respectively in Fig. 8a,c. For example, the minority class (dermatofibroma), having the least number of samples during both training and testing, demonstrated the lowest influence, which is evident from the performance results shown in both the tables and the confusion matrices. This is because the models M1 and M3 focused heavily on the majority class during training, thereby leading to high misclassification in the other (minority) classes. The model M1 gives an overall accuracy of 77% with low precision, recall, and f1-score, and model M3 gives an overall accuracy of 71% with lower precision, recall, and f1-score as compared to model M1. However, both models recorded an average performance, which calls for the need to appropriately address the class imbalance present in the dataset, supporting the proposed CANNSkin’s methods.
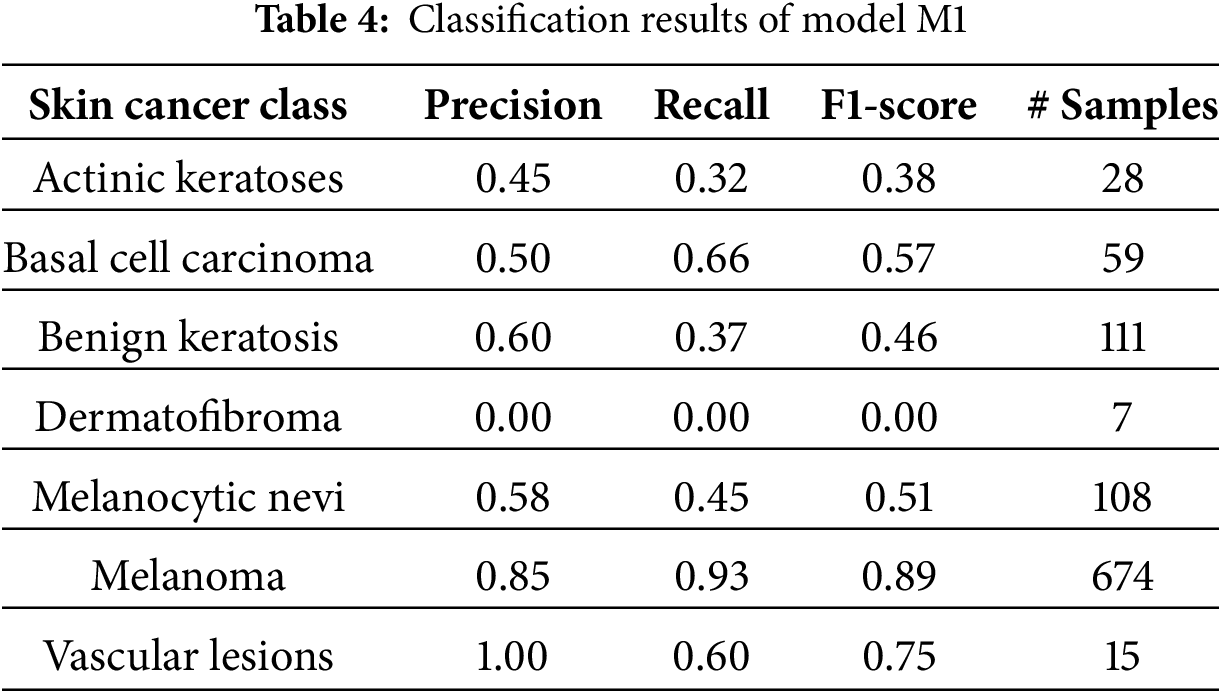
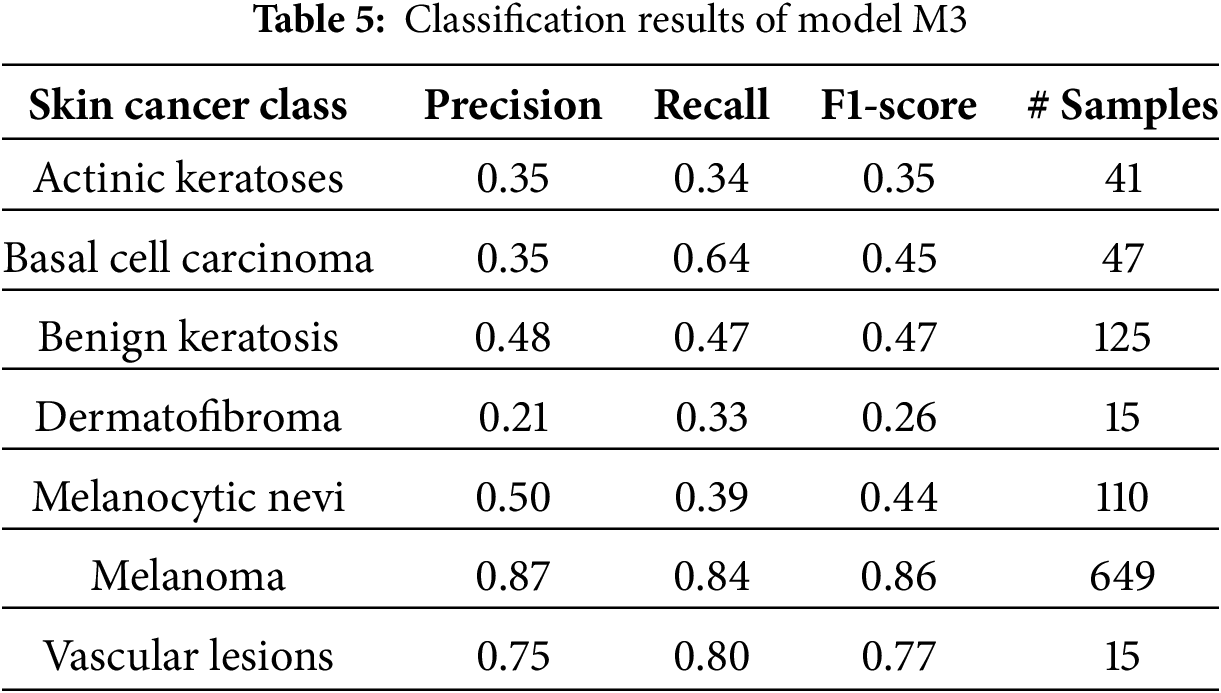
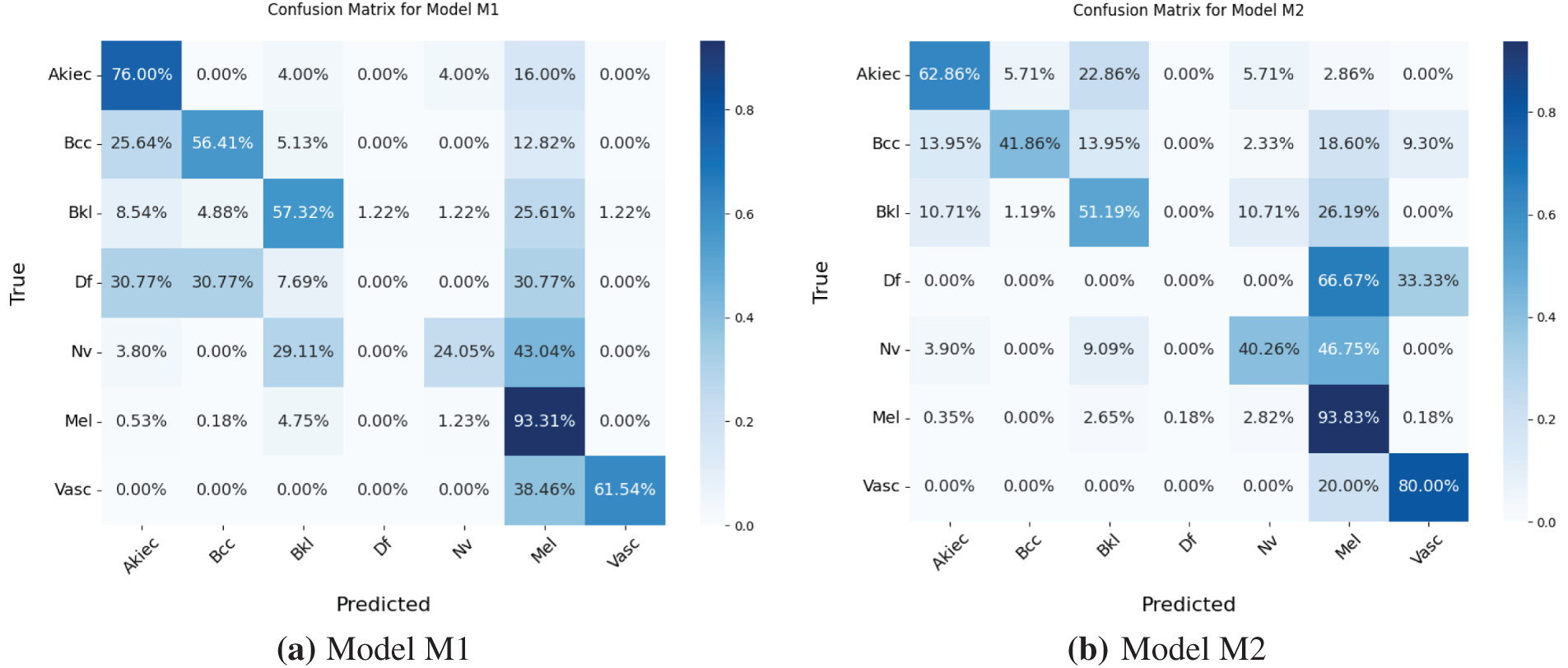
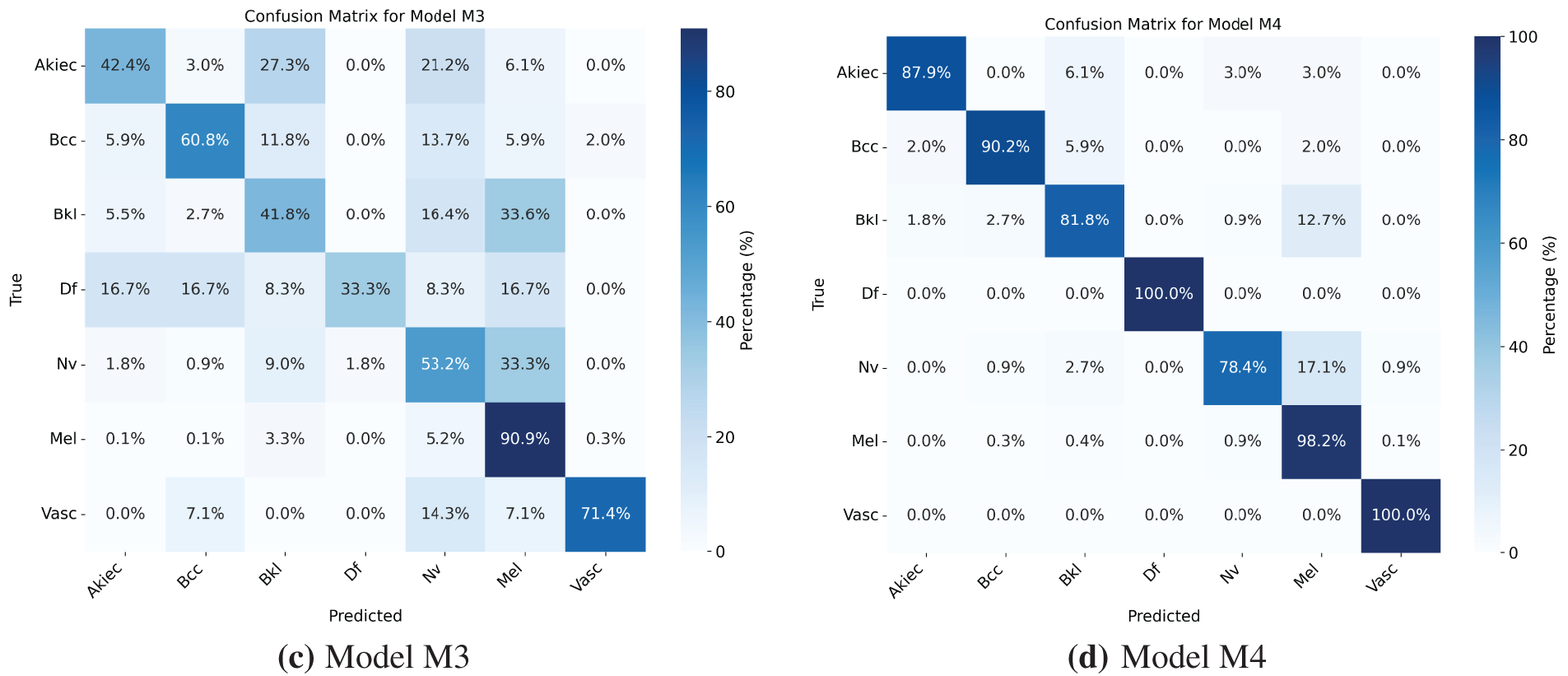
Figure 8: (a–d) Confusion matrices for Models M1–M4
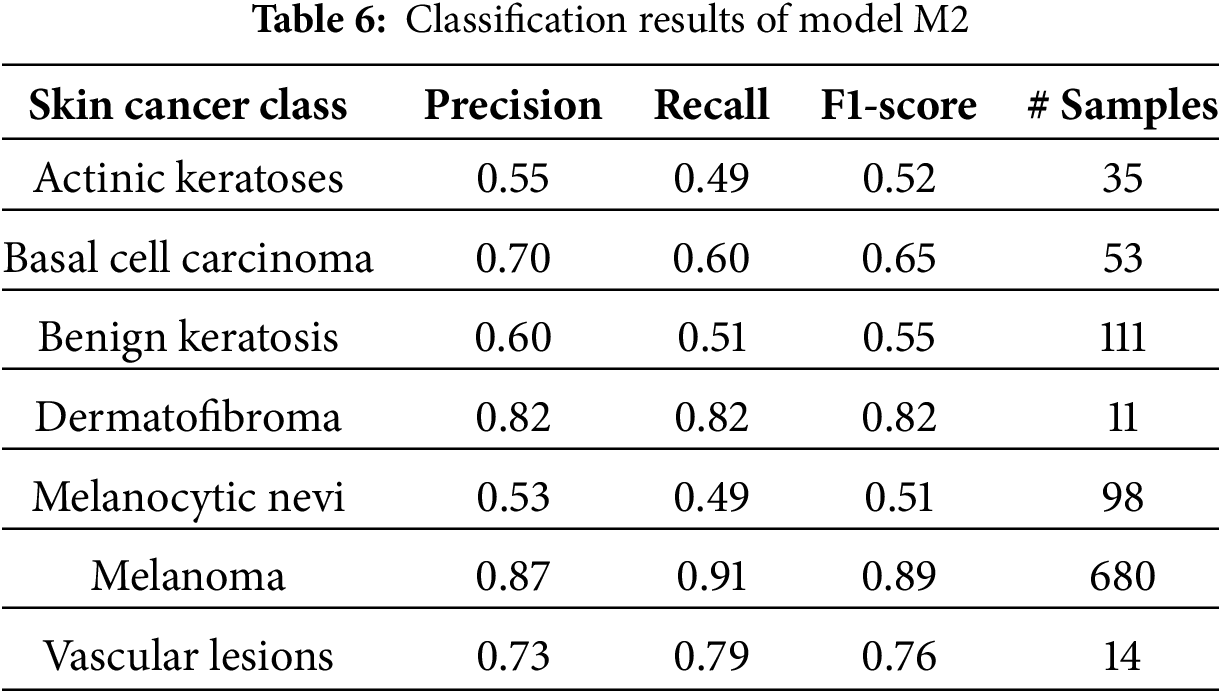
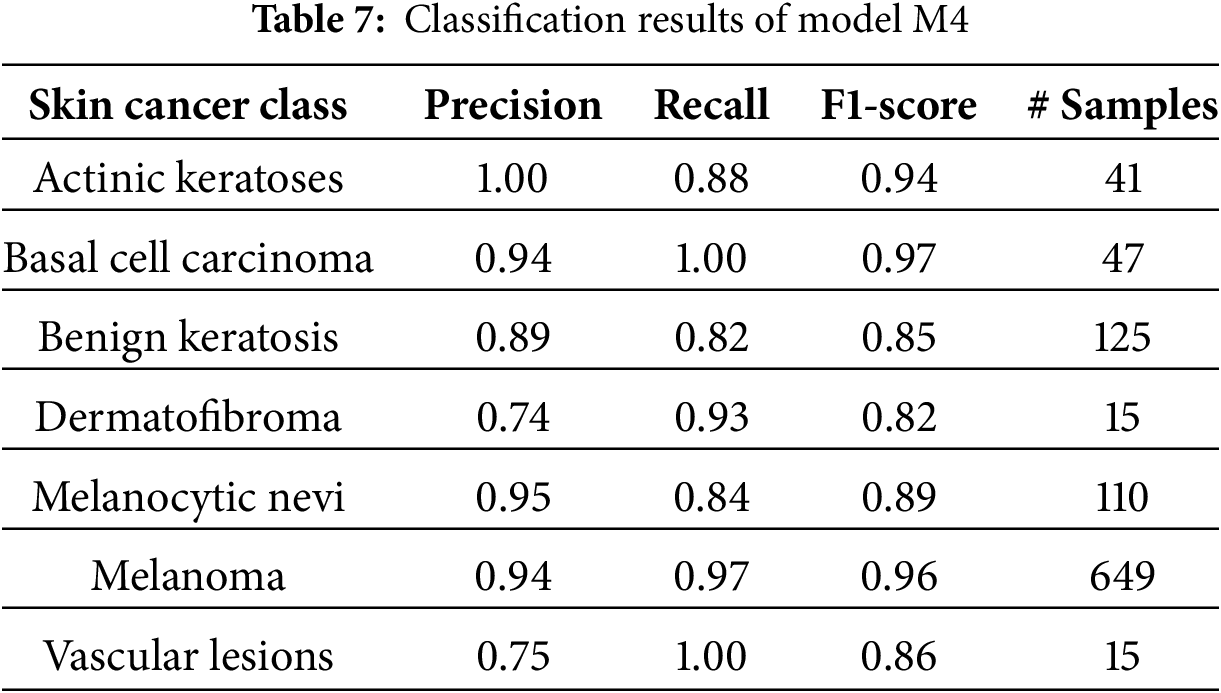
The model M2 trained on the oversampled latent embeddings gives an accuracy of 79% and high precision, recall, and F1-score of 78%, 79%, and 79%, respectively. Similarly, Table 6 and Fig. 8b show the classification results and confusion matrix for model M2, respectively. From the result obtained, it shows that model M2 can effectively classify each of classes, however, it lacks robustness in identifying new data samples. On the other hand, model M4 demonstrated the highest performance compared to the previous models, proving the superiority of the proposed CANNSkin framework. It gives an accuracy, precision, recall, and f1-score of 93%. This model can effectively classify each of the classes as shown in the classification report and confusion matrix in Table 7 and Fig. 8d, respectively. Moreover, it is clear that the effect of the class imbalance has been addressed by M4 as seen in the performance improvement of the dermatofibroma class (minority), which had the least performance by models M1 and M3. This implies that the model can recognize the different inter-class similarities and intra-class differences that is associated with skin cancer images. Also, it further indicates that fine-tuning the encoder model integrated with the classifier model (as in M4) aids in effectively capturing representative features present in the images, which are essential for classifying the input images correctly.
Moving further on the evaluation of the models, we used the Receiver Operating Characteristics Area Under Curve (ROC AUC) to further ascertain the efficacy of the developed models in discriminating between the multiple classes of the image dataset. Investigating models M1 and M3, the ROC curves shown in Fig. 9a,c indicate high misclassification among classes of the skin cancer lesions. On the other hand, the ROC curve of model M2 in Fig. 9b shows better classification of the skin cancer classes with less misclassification. The ROC curve, Fig. 9d, of model M4 shows the superiority of model M4 in classifying each of the different classes. This indicates that the model M4 is better at learning the similarities between different classes of skin cancer.
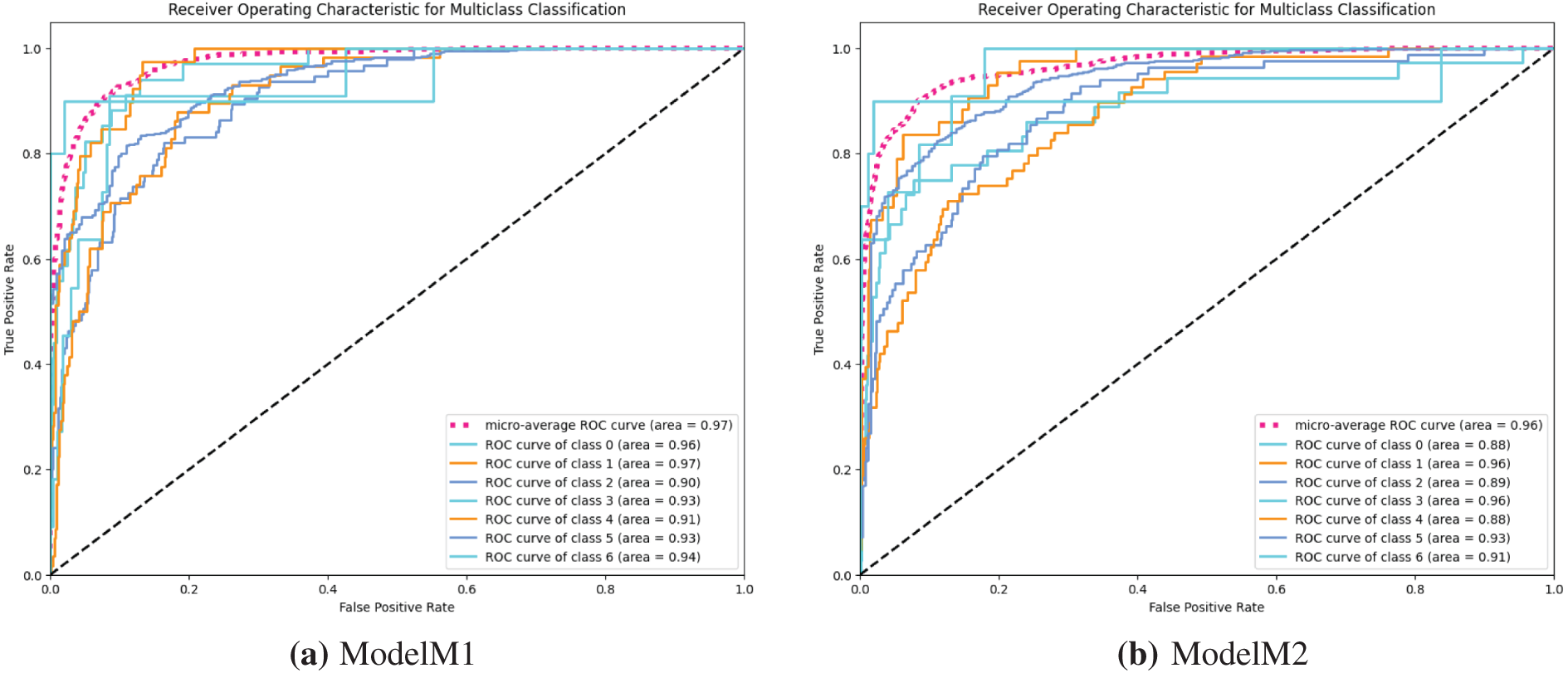
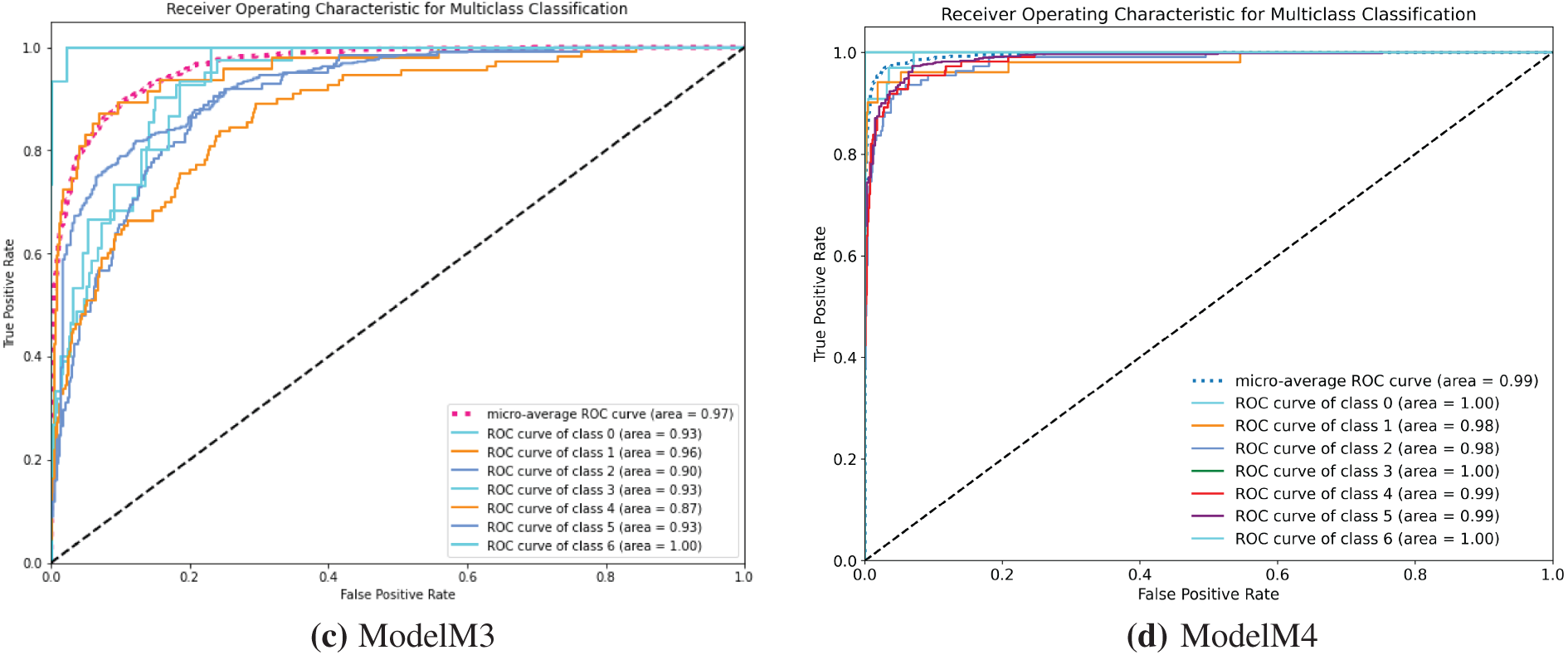
Figure 9: (a–d) receiver operating characteristic (ROC) curves for models M1–M4
To further assess the stability and statistical reliability of the proposed CANNSkin framework, we conducted an evaluation on the held-out test set of the HAM10000 dataset, in which the test set was split into 10 subsets, evaluating the model on each subset to obtain the mean and standard deviations. This analysis is essential for validating that the performance of the model is not dependent on a particular train–test split and that the classifier maintains consistent predictive behavior across different data partitions.
Across the 10 test subsets, CANNSkin demonstrated high accuracy with low variability. The model achieved an average accuracy of 0.9301 with a standard deviation of 0.0268, indicating stable performance across folds. Macro precision and macro recall were recorded as 0.9025
Importantly, the model produced a mean ROC–AUC of 0.9844
Balanced multiclass accuracy provides an unbiased measure of performance across all lesion categories by giving equal weight to minority and majority classes, as reported in Table 8. The results clearly show that Models M1 and M3, which were trained on imbalanced data, achieve relatively low balanced accuracies (53.50% and 56.10%), indicating that they remain strongly biased toward majority classes despite achieving reasonable overall accuracy. Model M2 shows a modest improvement to 56.84%, demonstrating that applying SMOTE on the latent embeddings helps mitigate imbalance at the feature level.
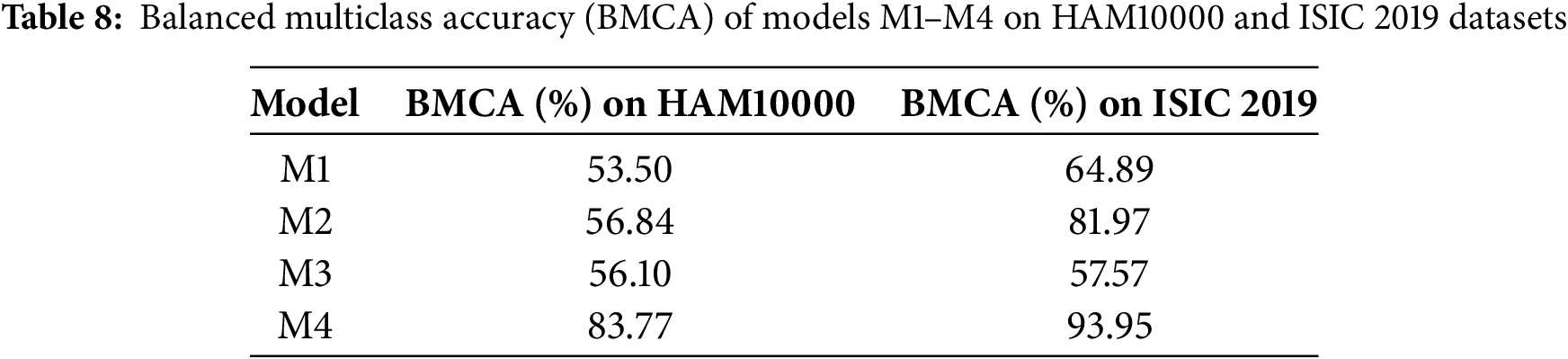
The most substantial improvement is observed in Model M4, which attains a balanced accuracy of 83.77%. This leap of more than 25 percentage points over M1 and M3 confirms that combining latent-space oversampling with end-to-end fine-tuning enables the model to learn more robust and equitable decision boundaries across all seven skin lesion types. Importantly, this demonstrates that the proposed CANNSkin framework not only increases overall classification accuracy but also significantly enhances fairness and discriminative ability for minority classes, addressing a critical limitation in prior skin lesion classification research.
Moreover, Gradient Class Activation Map (Grad-CAM) is a visual technique that explains the predictions of computer vision models. It computes the importance map by finding the derivative of a layer’s output per class. Grad-CAM is usually obtained from the convolutional feature map and shows the important features that contribute greatly to a model’s prediction for a class. In other words, it is a type of deep neural network explainability tool that highlights what the model looks at in the input for its predictions [20]. In this work, Grad-CAM was used to further affirm the model’s performance and robustness in predicting the different classes of skin cancer lesions. From Fig. 10, it shows that model M4 focuses on the right areas that are distinctive of the various classes. Moreover, this gives high confidence and further indicates that the high performance of the model in terms of accuracy, precision, recall, and ROC is reflective of the true capability for skin cancer lesions classification and not based on irrelevant or common areas from the input images.
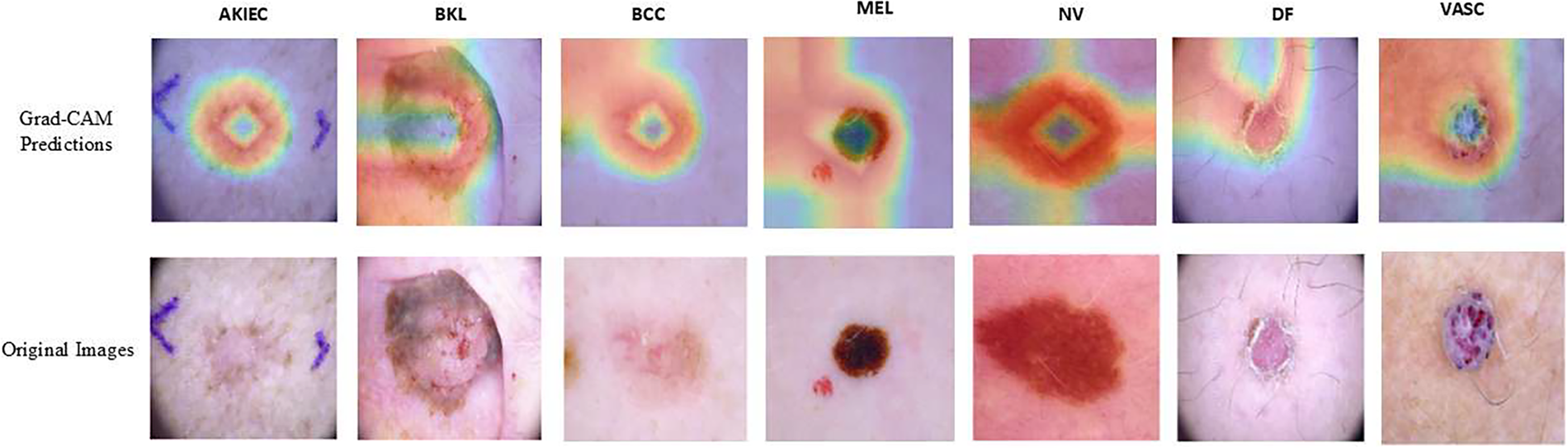
Figure 10: Grad-CAM interpretation of the model M4 predictions
5.3 Computational Efficiency and Inference Latency
To assess the computational efficiency of the proposed model, we measured the total parameter count, memory footprint, communication cost, and inference latency. Table 9 summarizes these metrics for CANNSkin.
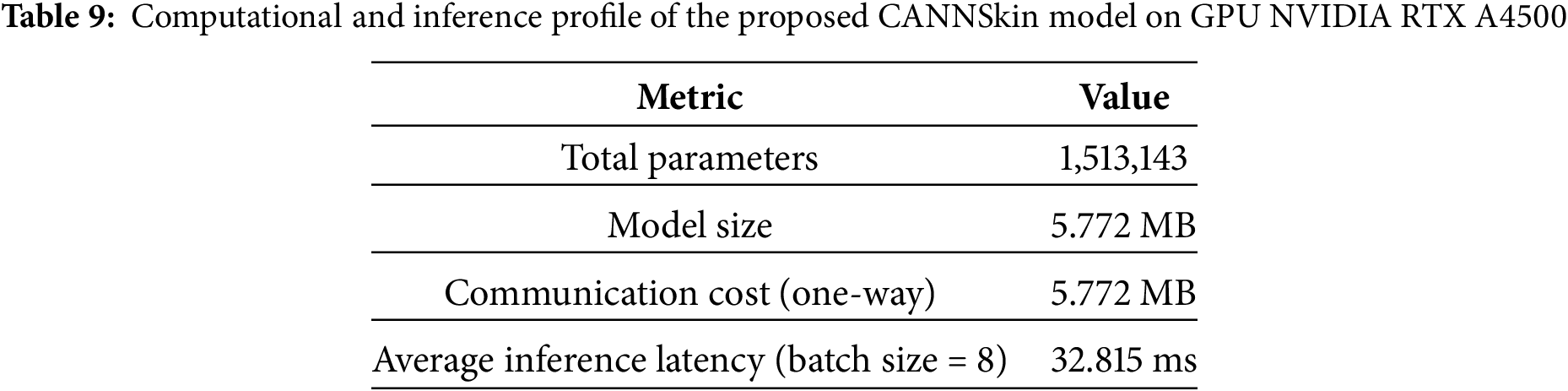
Table 9 shows that CANNSkin is computationally efficient, with only 1.5 M parameters and a compact 5.77 MB model size; significantly smaller than typical architectures such as ResNet [19] or EfficientNet [15]. This low footprint reduces memory demand and enables deployment on resource-limited devices.
The communication cost is also modest (5.77 MB one-way), making the model well-suited for federated or distributed learning scenarios. Inference is fast, with an average latency of 32.8 ms per batch (approximately 4 ms per image), enabling near–real-time prediction. Overall, the results demonstrate that CANNSkin provides strong performance while remaining lightweight and efficient for practical clinical use.
5.4 Evaluation on ISIC 2019 Dataset
To further validate the robustness and generalization capability of the proposed CANNSkin framework, we conducted an additional evaluation on the ISIC 2019 dataset [25], a large-scale dermoscopic benchmark comprising 25,331 images across eight diagnostic categories. Compared to HAM10000, the ISIC 2019 dataset presents higher heterogeneity, more severe class imbalance, and greater intra-class variability, making it a more challenging benchmark for automated lesion classification.
As before, all images were resized to
Autoencoder Reconstruction Performance
Before evaluating classification, we first assessed whether the CANNSkin autoencoder can reliably encode the ISIC 2019 image manifold. High-quality reconstruction is essential because our proposed method relies on the assumption that the latent space preserves meaningful, semantically consistent lesion features on which SMOTE can operate. On ISIC 2019, the autoencoder achieved an average SSIM of 0.86 and an average PSNR of 35.03 dB, demonstrating that the encoder consistently captures the structural and chromatic characteristics of dermoscopic lesions, despite differences in acquisition conditions. These results confirm that the latent space is suitable for interpolation and synthetic sample generation.
Classification Performance on ISIC 2019
Table 10 presents the performance of all four CANNSkin model variants on ISIC 2019. Consistent with observations from HAM10000, models trained on imbalanced latent representations (M1 and M3) achieve lower performance due to overfitting toward majority lesion categories. In contrast, models trained with latent-space SMOTE balancing (M2 and M4) demonstrate substantial improvements.
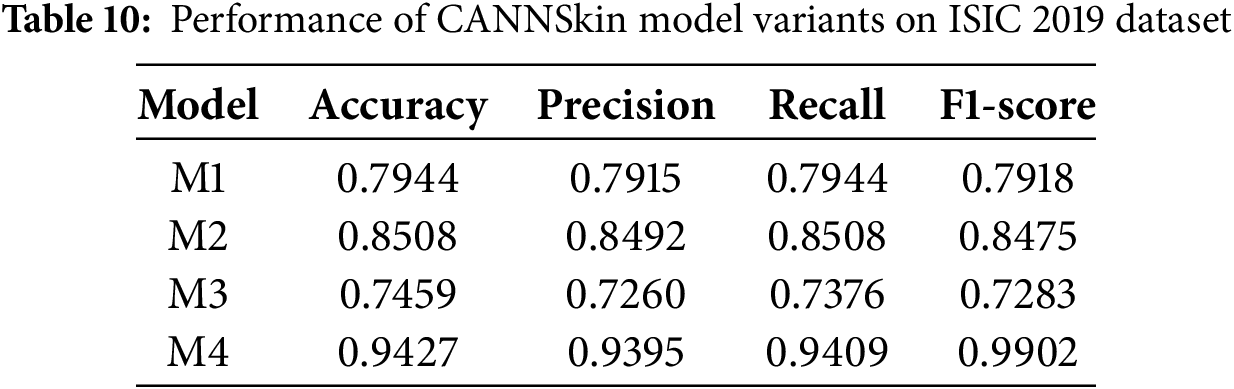
Notably, Model M4 achieves the highest performance, with an exceptional accuracy of 94.27% precision of 93.95%, a recall of 94.09%, and an F1-score of 99.02%, demonstrating strong discriminative capability across all lesion categories. This large performance gain highlights the importance of combining latent-space oversampling with end-to-end fine-tuning under a stabilized low-learning-rate regime.
Furthermore, to assess the stability and reliability of the proposed CANNSkin framework on the ISIC 2019 dataset, we conducted an additional evaluation by dividing the held-out test set into ten folds and evaluating the final M4 model across all partitions. The model demonstrated consistently strong performance with minimal variance, achieving an average accuracy of 0.9467
To further analyze class-wise behavior across lesion categories, we report the detailed classification performance of the best-performing model (M4) in Table 11. The results indicate consistently strong precision, recall, and F1-scores across all eight diagnostic categories, including clinically underrepresented classes such as AK, DF, VASC, and SCC. The overall BMCA of CANNSkin (M4) on the ISIC 2019 dataset was 0.9395.
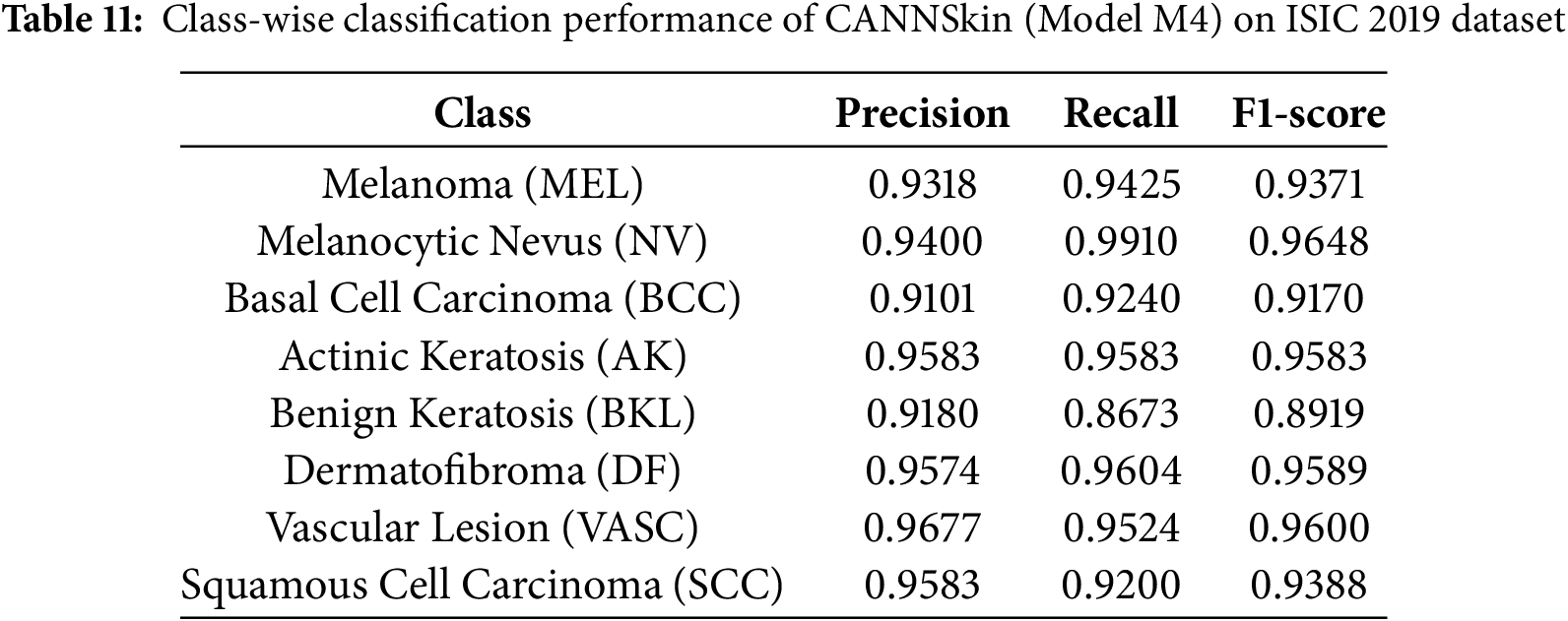
The improvement confirms that latent-space oversampling enhances the model’s ability to capture minority lesion characteristics and mitigates the imbalance-driven degradation observed in the baseline variants. These results reinforce the central finding of this work: applying SMOTE within the learned latent feature space produces balanced, semantically meaningful representations that lead to more robust and effective skin cancer classification.
5.5 Comparison with Existing Studies
From the aforementioned experiments, it is evident that the model M4, obtained by the integrated training of the pretrained encoder model and M2, which was trained on oversampled latent space representations of the training dataset, has the best classification performance. To further assess the proposed model’s superiority, Table 12 compares CANNSkin’s results with the reported results of some previously existing models ([15,19,26,27]) using the HAM10000 dataset.
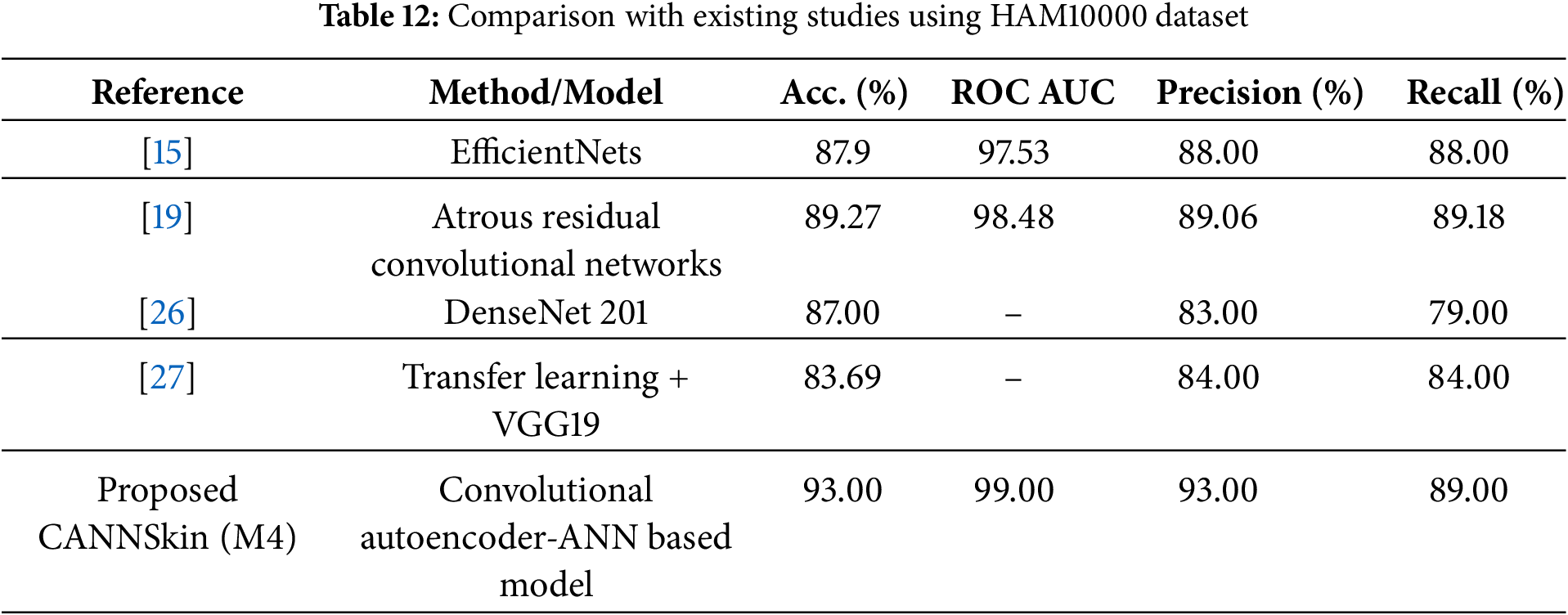
From the works compared against, while reference [15] indicated the presence of the dataset imbalance, they did not explicitly state how they addressed it, except highlighting that they evaluated the model with metrics that take into account the class imbalance. However, references [19,26], and [27] performed image data augmentation to address the issue of the class imbalance. From the results of the comparison, it is clear that our proposed approach of using SMOTE to balance the imbalanced latent embeddings has a significant effect towards improving the skin cancer classification performance. Consequently, from Table 12, model M4 outperformed the previous models. This can be attributed to leveraging CANNSkin’s integrated encoder and classifier models and its balancing of classes using the synthetic oversampling technique.
The proposed CANNSkin framework can also be contrasted with the recent MTA-Net model [16], which introduces a multi-scale triplet attention module on top of a pre-trained CNN and reports strong performance on both HAM10000 and ISIC 2019. On HAM10000, MTA-Net achieves an accuracy of 91.51% and a balanced multiclass accuracy (BMCA) of 87.18% [16]. Our CANNSkin model (M4) attains a superior overall accuracy of 93.0% and a macro F1-score of 91.0%, with a macro ROC–AUC of 99.0%, indicating excellent ranking and discrimination capability across classes. While MTA-Net reports a slightly higher BMCA, CANNSkin offers competitive or superior global metrics with a substantially simpler architecture that focuses on latent-space class balancing rather than sophisticated attention modules.
On the more challenging ISIC 2019 benchmark, MTA-Net reports 78.4% accuracy and 66.7% BMCA [16]. On the other hand, our proposed CANNSkin (M4) achieves an accuracy of 94.27%, demonstrating superiority over MTA-Net. In future work, it would be worth investigating how the strengths of the two methods could be synergized as complementary agents: the former emphasizes multi-scale attention for fine-grained feature learning, whereas CANNSkin emphasizes latent-space oversampling for principled class-imbalance mitigation, yielding state-of-the-art performance on HAM10000 and ISIC 2019.
This study proposed the CANNSkin framework for enhanced skin cancer classification using clinical skin images. This work focuses on tackling the class imbalance associated with skin cancer classification models. Notable publicly available datasets are known to contain high class imbalance; as such, most of the models developed in the field are either overfitting or biased towards the majority class(es). Though oversampling techniques have been developed, they may not be suitable for image datasets. This work presented CANNSkin, a convolutional autoencoder–based framework enhanced with latent-space oversampling for robust multi-class skin cancer classification. By applying SMOTE to the encoder’s latent representations and fine-tuning the integrated encoder–classifier network, the proposed method achieved strong and consistent performance on HAM10000 and demonstrated excellent generalization on the more complex ISIC 2019 dataset, where the fully trained model (M4) reached 94.27% accuracy with high reconstruction fidelity (PSNR 35.03 dB, SSIM 0.86). These results highlight the effectiveness of latent-space balancing and end-to-end refinement for improving discrimination across both majority and minority lesion classes.
A limitation of the current study includes the lack of uncertainty-awareness in the predictions, which may be needed for trustworthiness. Future work will explore extending CANNSkin through transfer-learning strategies to further strengthen feature representation, as well as federated or privacy-preserving learning setups to support deployment across multiple clinical centers without compromising patient data. Additional directions include continual learning to accommodate new lesion types and uncertainty-aware prediction to enhance clinical decision support.
Acknowledgement: The authors extend their appreciation to the Deanship of Scientific Research at Imam Mohammad Ibn Saud Islamic University (IMSIU) for funding this work through (grant number IMSIU-DDRSP2601).
Funding Statement: This work was supported and funded by the Deanship of Scientific Research at Imam Mohammad Ibn Saud Islamic University (IMSIU) (grant number IMSIU-DDRSP2601).
Author Contributions: The authors confirm contribution to the paper as follows: Abdul Jabbar Siddiqui: Conceptualization, methodology, original draft, analysis, reviewing and editing; Saheed Ademola Bello: Conceptualization, methodology, original draft, analysis; Muhammad Liman Gambo: Conceptualization, methodology, original draft, analysis; Abdul Khader Jilani Saudagar: Conceptualization, methodology, analysis, reviewing and editing, funding; Mohamad A. Alawad: Original draft, analysis, reviewing and editing; Amir Hussain: Conceptualization, original draft, analysis, reviewing and editing. All authors reviewed the results and approved the final version of the manuscript.
Availability of Data and Materials: The dataset used in this study is publicly accessible at: Harvard Dataverse. https://doi.org/10.7910/DVN/DBW86T (2018) [24].
Ethics Approval: Not applicable.
Conflicts of Interest: The authors declare no conflicts of interest to report regarding the present study.
References
1. Hameed M, Zameer A, Raja MAZ. A comprehensive systematic review: advancements in skin cancer classification and segmentation using the ISIC dataset. Comput Model Eng Sci. 2024;140(3):2131–64. doi:10.32604/cmes.2024.050124. [Google Scholar] [CrossRef]
2. Akram T, Almarshad F, Alsuhaibani A, Naqvi SR. BioSkinNet: a bio-inspired feature-selection framework for skin lesion classification. Comput Model Eng Sci. 2025;143(2):2333–59. doi:10.32604/cmes.2025.064079. [Google Scholar] [CrossRef]
3. WCRF International. Skin cancer statistics 2020. 2020. [cited 2024 Feb 29]. Available from: https://www.wcrf.org/cancer-trends/skin-cancer-statistics. [Google Scholar]
4. Siddiqui AM, Abbas H, Asim M, Ateya AA, Abdallah HA. SGO-DRE: a squid game optimization-based ensemble method for accurate and interpretable skin disease diagnosis. Comput Model Eng Sci. 2025;144(3):3135–68. doi:10.32604/cmes.2025.069926. [Google Scholar] [CrossRef]
5. Ahmad B, Usama M, Huang CM, Hwang K, Hossain MS, Muhammad G. Discriminative feature learning for skin disease classification using deep convolutional neural network. IEEE Access. 2020;8:39025–33. doi:10.1109/access.2020.2975198. [Google Scholar] [CrossRef]
6. Reddy DA, Roy S, Kumar S, Tripathi R. A scheme for effective skin disease detection using optimized region growing segmentation and autoencoder based classification. Procedia Comput Sci. 2023;218:274–82. doi:10.1016/j.procs.2023.01.009. [Google Scholar] [CrossRef]
7. Bassel A, Abdulkareem AB, Alyasseri ZAA, Sani NS, Mohammed HJ. Automatic malignant and benign skin cancer classification using a hybrid deep learning approach. Diagnostics. 2022;12(10):2472. doi:10.3390/diagnostics12102472. [Google Scholar] [PubMed] [CrossRef]
8. Farhoudian A, Heidari A, Shahhosseini R. A new era in colorectal cancer: artificial Intelligence at the forefront. Comput Biol Med. 2025;196:110926. doi:10.1016/j.compbiomed.2025.110926. [Google Scholar] [PubMed] [CrossRef]
9. Victor A, Ghalib MR. Automatic detection and classification of skin cancer. Int J Intell Eng Syst. 2017;10:444–51. doi:10.22266/ijies2017.0630.50. [Google Scholar] [CrossRef]
10. Viknesh CK, Kumar PN, Seetharaman R, Anitha D. Detection and classification of melanoma skin cancer using image processing technique. Diagnostics. 2023;13(21):3313. doi:10.3390/diagnostics13213313. [Google Scholar] [PubMed] [CrossRef]
11. Alquran H, Qasmieh IA, Alqudah AM, Alhammouri S, Alawneh E, Abughazaleh A, et al. The melanoma skin cancer detection and classification using support vector machine. In: 2017 IEEE Jordan Conference on Applied Electrical Engineering and Computing Technologies (AEECT). Piscataway, NJ, USA: IEEE; 2017. p. 1–5. [Google Scholar]
12. Thanh D, Prasath VBS, Hieu L, Hien NN. Melanoma skin cancer detection method based on adaptive principal curvature, colour normalisation and feature extraction with the ABCD rule. J Digit Imaging. 2020;33:574–85. doi:10.1007/s10278-019-00316-x. [Google Scholar] [PubMed] [CrossRef]
13. Alfed N, Khelifi F. Bagged textural and color features for melanoma skin cancer detection in dermoscopic and standard images. Expert Syst Applicat. 2017;90:101–10. doi:10.1016/j.eswa.2017.08.010. [Google Scholar] [CrossRef]
14. Javed R, Rahim M, Saba T, Fati S, Rehman A, Tariq U. Statistical histogram decision based contrast categorization of skin lesion datasets dermoscopic images. Comput Mat Cont. 2020;67(2):2337–52. doi:10.32604/cmc.2021.014677. [Google Scholar] [CrossRef]
15. Ali K, Shaikh ZA, Khan AA, Laghari AA. Multiclass skin cancer classification using EfficientNets–a first step towards preventing skin cancer. Neurosci Inform. 2022;2(4):100034. doi:10.1016/j.neuri.2021.100034. [Google Scholar] [CrossRef]
16. Gajera HK, Nayak DR, Zaveri MA. MTA-Net: multi-scale triplet attention-aware network for multiclass skin lesion classification. Comput Biol Med. 2025;196:110729. doi:10.1016/j.compbiomed.2025.110729. [Google Scholar] [PubMed] [CrossRef]
17. Liu Z, Xiong R, Jiang T. CI-Net: clinical-inspired network for automated skin lesion recognition. IEEE Trans Med Imag. 2023;42(3):619–32. doi:10.1109/tmi.2022.3215547. [Google Scholar] [PubMed] [CrossRef]
18. Aladhadh S, Alsanea M, Aloraini M, Khan T, Habib S, Islam M. An effective skin cancer classification mechanism via medical vision transformer. Sensors. 2022;22(11):4008. doi:10.3390/s22114008. [Google Scholar] [PubMed] [CrossRef]
19. Ramamurthy K, Muthuswamy A, Mathimariappan N, Kathiresan GS. A novel two-staged network for skin disease detection using atrous residual convolutional networks. Concurr Comput. 2023;35(26):e7834. doi:10.1002/cpe.7834. [Google Scholar] [CrossRef]
20. Toumaj S, Heidari A, Jafari Navimipour N. Leveraging explainable artificial intelligence for transparent and trustworthy cancer detection systems. Artif Intell Med. 2025;169:103243. doi:10.1016/j.artmed.2025.103243. [Google Scholar] [PubMed] [CrossRef]
21. Nguyen KD, Zhou YH, Nguyen QV, Sun MT, Sakai K, Ku WS. SILP: enhancing skin lesion classification with spatial interaction and local perception. Expert Syst Appl. 2024;258:125094. doi:10.1016/j.eswa.2024.125094. [Google Scholar] [CrossRef]
22. Chawla NV, Bowyer KW, Hall LO, Kegelmeyer WP. SMOTE: synthetic minority over-sampling technique. J Artif Intell Res. 2002;16:321–57. doi:10.1613/jair.953. [Google Scholar] [CrossRef]
23. Wang Y, Yao H, Zhao S. Auto-encoder based dimensionality reduction. Neurocomputing. 2016;184(4):232–42. doi:10.1016/j.neucom.2015.08.104. [Google Scholar] [CrossRef]
24. Tschandl P, Rosendahl C, Kittler H. The HAM10000 dataset, a large collection of multi-source dermatoscopic images of common pigmented skin lesions. Sci Data. 2018;5(1):180161. doi:10.1038/sdata.2018.161. [Google Scholar] [PubMed] [CrossRef]
25. Codella N, Rotemberg V, Tschandl P, Celebi ME, Dusza S, Gutman D, et al. Skin lesion analysis toward melanoma detection 2018: a challenge hosted by the international skin imaging collaboration (isic). arXiv:1902.03368. 2019. [Google Scholar]
26. Rajesh AVP, Rao KN, Sai GNV, Kumar KD, Karthik KRS. Skin cancer detection and intensity analysis using deep learning. In: 2024 International Conference on Emerging Systems and Intelligent Computing (ESIC). Cham, Switzerland: Springer; 2024. p. 376–81. [Google Scholar]
27. Swetha-R N, Shrivastava VK, Parvathi K. Multiclass skin lesion classification using image augmentation technique and transfer learning models. Int J Intell Unmanned Syst. 2024;12(2):220–8. doi:10.1108/ijius-02-2021-0010. [Google Scholar] [CrossRef]
Cite This Article
 Copyright © 2026 The Author(s). Published by Tech Science Press.
Copyright © 2026 The Author(s). Published by Tech Science Press.This work is licensed under a Creative Commons Attribution 4.0 International License , which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.


 Submit a Paper
Submit a Paper Propose a Special lssue
Propose a Special lssue View Full Text
View Full Text Download PDF
Download PDF Downloads
Downloads
 Citation Tools
Citation Tools