 Open Access
Open Access
ARTICLE
DeepClassifier: A Data Sampling-Based Hybrid BiLSTM-BiGRU Neural Network for Enhanced Type 2 Diabetes Prediction
1 School of Digital Science, Universiti Brunei Darussalam, Jalan Tungku Link, ABS Building, Bandar Seri Begawan, Brunei Darussalam
2 Department of Computer Science, Ahmadu Bello University, Zaria, Nigeria
3 Department of Computing, Universiti Teknologi PETRONAS, Sri Iskandar, Malaysia
4 School of Computer Science, Faculty of Computing, Digital and Data, Technological University (TU), Dublin, Ireland
5 Department of Electrical and Computer Engineering, Western University, London, ON, Canada
6 PAPRSB Institute of Health Sciences, Universiti Brunei Darussalam, Jalan Tungku Link, Gadong, Bandar Seri Begawan, Brunei Darussalam
* Corresponding Author: Abdullahi Abubakar Imam. Email:
(This article belongs to the Special Issue: Artificial Intelligence Models in Healthcare: Challenges, Methods, and Applications)
Computer Modeling in Engineering & Sciences 2026, 146(3), 37 https://doi.org/10.32604/cmes.2026.076187
Received 15 November 2025; Accepted 21 January 2026; Issue published 30 March 2026
Abstract
Artificial Intelligence (AI) in healthcare enables predicting diabetes using data-driven methods instead of the traditional ways of screening the disease, which include hemoglobin A1c (HbA1c), oral glucose tolerance test (OGTT), and fasting plasma glucose (FPG) screening techniques, which are invasive and limited in scale. Machine learning (ML) and deep neural network (DNN) models that use large datasets to learn the complex, nonlinear feature interactions, but the conventional ML algorithms are data sensitive and often show unstable predictive accuracy. Conversely, DNN models are more robust, though the ability to reach a high accuracy rate consistently on heterogeneous datasets is still an open challenge. For predicting diabetes, this work proposed a hybrid DNN approach by integrating a bidirectional long short-term memory (BiLSTM) network with a bidirectional gated recurrent unit (BiGRU). A robust DL model, developed by combining various datasets with weighted coefficients, dense operations in the connection of deep layers, and the output aggregation using batch normalization and dropout functions to avoid overfitting. The goal of this hybrid model is better generalization and consistency among various datasets, which facilitates the effective management and early intervention. The proposed DNN model exhibits an excellent predictive performance as compared to the state-of-the-art and baseline ML and DNN models for diabetes prediction tasks. The robust performance indicates the possible usefulness of DL-based models in the development of disease prediction in healthcare and other areas that demand high-quality analytics.Keywords
The phenomenon of diabetes in the contemporary world has become a significant health issue in the present days. The hormone that regulates the level of glucose in the blood, insulin, is poorly synthesized and used, which is the primary cause of diabetes [1]. Diabetes, if left untreated, may damage a wide range of bodily organs gradually [2]. Over the last ten years, diabetes has been diagnosed in over 400 million individuals, with most of those diagnosed residing in low- and middle-income nations [3]. Additionally, diabetes is responsible for causing 1.5 million fatalities annually. It has been projected that this figure will increase by almost 65% to surpass 700 million within a few decades. Specifically, reported research findings indicated that between 2030 and 2045, the number of people living with diabetes is anticipated to increase to 25% and 51% of the world population, respectively [3,4]. Based on prevalence, diabetes has two types, type 1 and type 2. The body’s immune system mistakenly damages insulin-producing beta cells in the pancreas in type 1, thereby resulting in the destruction of the insulin manufacturing process [5,6]. Type 2 diabetes is the most common, and it is strongly associated with unhealthy lifestyles such as bad eating habits, lack of physical activity, and poor mental health. Insulin resistance and insufficient insulin production by the pancreas characterize this type. As a result, the body loses its ability to regulate blood glucose levels; to keep the diabetic condition under control, patients with type 2 diabetes are placed on continuous medication and advised to maintain a healthy lifestyle due to its chronic nature [5,7].
The symptoms of diabetes may vary greatly from one kind of diabetes to another, depending on the specific kind of diabetes [6]. Some of the usual symptoms include, but are not limited to, increased appetite, frequent urination, weariness, impaired vision, unexplained weight loss, numbness, and recurring infections. One of the most common early symptoms of diabetes is an increase in urination and frequent dehydration [4]. Also, diabetic individuals can lose weight as they gain appetite. High blood sugar levels impact the lens of the eye, causing blurred vision. High blood sugar levels can impair the immune system, making people prone to infections and wound and injury healing. Such conditions result in serious complications, such as cardiovascular disease, kidney failure, neuropathy, and retinopathy, which significantly affect the quality of life of patients and burden healthcare systems across the globe with a heavy load [8,9]. These negative outcomes can be reduced only after being identified at the early stages and managed, and this is where the importance of efficient predictive and diagnostic methods lies [1,10,11].
The blood glucose can be controlled in a number of ways, depending on the type of diabetes. Insulin treatment is normally used to control type 1 diabetes, but type 2 diabetes might require oral medication or insulin injection, depending on the severity of the disease [4]. Diabetics can maintain normal blood glucose levels by regularly checking their blood glucose levels and, when necessary, by changing the doses of certain drugs. In this case, nutrition is also important. The management of diabetes requires adopting a healthy and well-balanced diet that is closely followed in terms of carbohydrate intake and the amount of carbohydrates that one consumes. The advantages of regular exercise include weight loss, increased insulin sensitivity, and normal blood sugar levels [12,13].
Historically, several clinical tests, such as hemoglobin A1c (HbA1c), oral glucose tolerance test (OGTT), and fasting plasma glucose (FPG), have been utilized in the prediction and diagnosis of diabetes [14,15]. Although these tests are accurate, they are usually invasive, time-consuming, and need access to health services, hence they are not as viable in conducting a mass screening in a diverse population. Moreover, the predictive capacity of the traditional risk assessment models relying on demographic, lifestyle, and genetic factors may be limited and often fail to explain the complicated interaction of factors leading to the development of diabetes. In addition, medical data analysis is critical in the diagnosis and understanding of diabetes, which needs to be done correctly [4,7,16]. Therefore, the application of artificial intelligence (AI) to predict diabetes with robust performance is highly in demand [5,17].
The implementation of AI has transformed many aspects, including health care. It is a very broad term and includes several technologies, including machine learning (ML) and deep neural networks (DNN), that are capable of processing large volumes of data with high accuracy [18]. DNN, which is a subset of AI, has become an effective technology that can overcome the shortcomings of traditional ML models [19]. DNNs are composed of successive layers of connected neurons that are capable of learning hierarchical data representations. This multi-layer architecture enables the modeling of complex and non-linear relationships and interactions among data, making it especially suited for predictive tasks in healthcare. DNNs have been quite successful in other medical field applications, such as medical imaging, genomics, and disease prediction, among others [20,21]. An example is the Convolutional Neural Networks (CNNs) that are effective in image-based problems, including the detection of diabetic retinopathy based on retinal images [22]. Similarly, a PPG-based digital biomarker framework that integrates multiset spatiotemporal feature fusion with explainable artificial intelligence (XAI) has been proposed to improve interpretability and predictive performance [23], and Recurrent Neural Networks (RNNs) that demonstrated their effectiveness in time-series analysis that could be utilized in monitoring glucose levels and forecasting future trends [24]. These successes highlight the possibility of DNNs to improve diabetes prediction algorithms.
Even though the DL-based diabetes prediction frameworks were quite successful, there are still a number of methodological gaps. Most of the existing research utilizes individual recurrent models such as Gated Recurrent Unit (GRU), and Long Short-Term Memory (LSTM), which can be less effective but reveal the various temporal relationships, including the heterogeneous clinical data. [24–26]. Furthermore, many hybrid deep learning approaches reported in the literature primarily combine convolutional and recurrent networks, designs that are often better suited to image or sensor-based data rather than structured clinical and tabular datasets commonly used for diabetes prediction [20,22]. Due to these limitations, the existing models lag to effectively generalize across datasets with varying feature characteristics and temporal distributions in real-world healthcare scenarios.
Moreover, the bidirectional recurrent neural networks have proved the better performance by learning the context-related information not only from past but future states as well [24]. Also, existing studies rarely explore the complementary strengths of different gated recurrent mechanisms within a unified framework. BiLSTM networks are well known for their ability to model long-term dependencies but often incur higher computational complexity, whereas BiGRU models offer faster convergence and reduced parameterization at the potential cost of representational depth [19,24]. This shows the lack of research on integrating these bidirectional networks into a combined hybrid network as a crucial gap, especially for type-2 diabetes prediction tasks involving complex clinical and temporal variables.
The use of DNNs in diabetes prediction is based on their ability to combine and process diverse data types, such as electronic health records (EHRs), genetic data, lifestyle, and continuous glucose monitoring data [7,25,26]. Through the training of large and multi-dimensional data, DNNs can detect minor patterns and risk factors that may contribute to the development of diabetes that cannot be detected by conventional ML approaches. This potential can result in improved and precise predictions that can be used to intervene early and manage the disease better.
With architectural limitations, dataset imbalance is also a significant issue; however, numerous studies on the topic implicitly assume balanced datasets, despite the imbalanced nature of medical data, which has an extensive literature on class imbalance. This assumption may lead to biased predictions and reduced clinical reliability. The need to mitigate the architectural weaknesses of current recurrent models, as well as address the issue of data imbalance, is crucial in developing robust and clinically relevant prediction systems. These limitations in the current research motivate the development of a hybrid bidirectional recurrent framework that not only enhances temporal feature learning but also introduces a data sampling method into the prediction process to address the crucial issue of latent class imbalance, a common phenomenon in medical datasets.
The major contributions of this research are as follows:
1. A novel bidirectional hybrid DNN model integrating BiLSTM and BiGRU to benefit the crucial temporal learning capabilities for type-2 diabetes prediction.
2. Strong generalizability across heterogeneous datasets, validating the model’s robustness and adaptability to varying demographic and clinical data distributions.
3. Comprehensive benchmarking of the proposed model against state-of-the-art ML and DL approaches using standard performance metrics, including accuracy, area under the curve (AUC), receiver operating curve (ROC), and F1-score.
4. Public release of the source code and datasets to support reproducibility and future research.
This study is further organized into the literature review Section 2, the methodology Section 3, the experimental findings Section 4, and the threats to validity Section 5 and the conclusion Section 6.
Prior to the adoption of computational intelligence in healthcare, diabetes diagnosis relied primarily on clinical expertise, observable symptoms, and patient medical history [27]. Despite the advantages of such expert based assesments, which were based on domain knowledge, they were subjective and susceptible to inter-practitioner variation by nature [28]. The first computational attempts were to formalize clinical reasoning using rule-based systems based on expert knowledge and medical guidelines [29]. Although these systems were consistent and interpretable, they were also rigid, limiting their ability to adapt to complex and dynamic patient situations and diverse clinical phenotypes, as well as to heterogeneous clinical data, and phenotypic variation of patients [30].
Statistical learning methods then became an alternative that was more data-oriented. The ease of use, interpretability, and binary classification applications made logistic regression (LR) popular among researchers [31,32]. On the same note, Linear Discriminant Analysis (LDA) and Bayesian classifiers were investigated to predict diabetes [33,34]. Although these methods are analytically transparent, they are highly reliant on strong assumptions about data distribution and linear separability as well, which are less effective when used with nonlinear or high-dimensional clinical data.
2.1 Machine Learning for Type-2 Diabetes Prediction
As the field of artificial intelligence advanced, machine learning (ML) techniques emerged as a popular approach in diabetes prediction because of their higher predictive accuracy and the flexibility of application [35]. Several types of ML algorithms, such as Support Vector Machines (SVM), Random Forests (RF), Decision Trees (DT), k-Nearest Neighbor (KNN), Naive Bayes (NB), and ensemble classifiers, are highly tested, especially on benchmark datasets, including PIMA [36,37]. The accuracy values in these studies have a range of 70–85% percent, and the performance of the studies is strongly dependent on the preprocessing strategies, feature selection strategies, and the nature of the datasets.
A number of studies have investigated hybrid ML systems and sampling techniques to improve the predictive accuracy. Minimum Redundancy Maximum Relevance (mRMR), Boruta, Principal Component Analysis (PCA) and tree-based importance measures are some of the feature selection methods that have been applied with varying success rates in feature selection in the most effective way possible [38–41]. Sampling-based methods such as shuffled and stratified sampling have shown a gain in performance over some datasets but tend to be weak over most datasets in general [42]. Despite some of the research claiming extremely high accuracies, up to 95% percent in some cases, the high results are often recorded on small or skewed data without a strict validation, which is of concern to overfitting and external validity [39,41].
Ensemble learning methods, particularly Gradient Boosting Machines (GBM), XGBoost, and LightGBM, have further improved classification performance by aggregating multiple weak learners [11,43,44]. While ensemble models generally outperform individual classifiers, they introduce increased computational complexity and reduced interpretability. Moreover, many ensemble-based studies rely on single train-test splits and do not explicitly address class imbalance, which is a common characteristic of medical datasets.
Hence, the machine ML methods have shown moderate diabetes prediction success, and the reported accuracy is usually between 70% and 85% on regular datasets used in the field of study [36,44]. These models, however, have a number of repetitive limitations. Their predictive performance is very sensitive to feature engineering, preprocessing options, and dataset-specific attributes, and thus, the predictive performance is not consistent across studies. Besides, the majority of ML-based approaches are based on shallow decision boundaries and fail to represent more complex nonlinear interactions among clinical variables. Though ensemble methods and sampling approaches can help enhance performance in part, they also tend to become more complex and computationally intensive, as well as not robust enough when used on heterogeneous or unbalanced medical data. These drawbacks limit the scalability and generalizability of the ML-based diabetes prediction systems to a real-life clinical context, thereby motivating the exploration of more expressive and sequence-aware deep learning architectures.
2.2 Deep Learning for Type-2 Diabetes Prediction
DL models have gained increasing attention in diabetes prediction due to their ability to automatically learn hierarchical feature representations without extensive manual feature engineering. The most popular architectures of DL used in this field are CNNs, deep belief networks (DBNs) and DNNs. Although these models have been proven to have a better representational ability than traditional machine learning models, they tend to be limited in terms of the flexibility of their architecture and the ability to utilize temporal dependencies inherent in clinical data.
Recent literature has discussed interpretable DL models to enhance clinical trust and transparency. As an example, DiaXplain combines deep neural networks with explainability techniques in order to diagnose type 2 diabetes in patients [45]. Despite the fact that DiaXplain increases interpretability, it is less focused on the temporal sequence model or the imbalance of classes, which restricts its use in longitudinal or heterogeneous clinical data. Likewise, CNN-based methods [46] can be used to process the fixed feature representation well, but cannot be used to extract sequential dependence because they were initially developed in the spatial feature extraction. As a result, their use in predicting diabetes is frequently based on transformed or image-like forms of tabular data, and thus may obscure temporal trends.
The ability of DBNs and DNNs to capture nonlinear relationships has also attracted a lot of research. DBN-based models, to learn deep feature hierarchies, have been found to use stacked Restricted Boltzmann machines, whilst DNNs rely on several fully connected layers to learn more intricate interactions between clinical variables. Though such models have high reported predictive accuracies up to 98% these results are often achieved with small datasets, fixed train-test sets, or highly curated samples, which have been questioned due to overfitting and generalizability to the real world. Additionally, the majority of DNN-based methods are based on fixed feature vectors, and they do not directly capture the temporal dynamics of patient health records [47–50].
More recent work on DL has started to experiment with sequence-aware and hybrid medical prediction architectures. Recurrent neural networks (RNNs) and their variants, including LSTM and GRU, have shown better results in the modeling of temporal dependencies in clinical data [24,48]. However, the majority of these works employ either LSTM or GRU individually, limiting their ability to balance long-term dependency learning with computational efficiency. Bidirectional recurrent models, which process sequences in both forward and backward directions, remain underutilized in diabetes prediction despite their demonstrated success in other healthcare applications.
In addition, several recent works have explored hybrid and attention-based DL frameworks for diabetes risk assessment. Transformer-inspired architectures [51], attention-enhanced RNN models [52], and attention-enhanced deep belief networks [53] have shown promise in improving prediction accuracy. Nonetheless, these studies are often evaluated on a single dataset, predominantly PIMA, and rarely address class imbalance or cross-dataset robustness. Furthermore, the computational complexity of such models may limit their deployment in resource-constrained clinical environments.
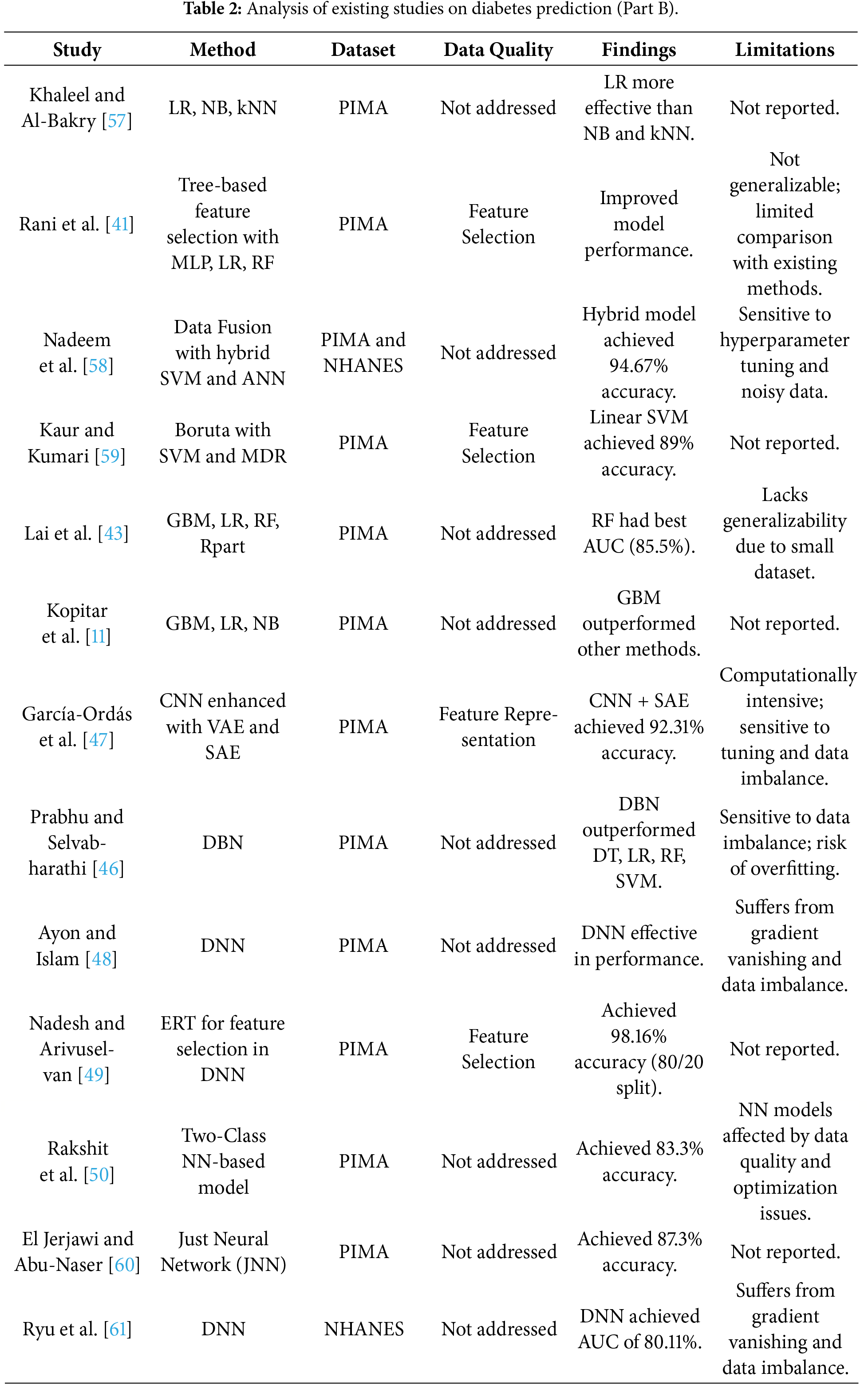
Although the DL models have a better representational ability, current studies in diabetes prediction using DL have significant limitations. Much of the literature on models is on single-architecture, e.g., CNNs or feedforward DNNs, which are frequently not trained on sequential clinical data [46] but instead on image-based or static representations of features. Recurrent models with the ability to capture time-related effects are relatively underutilized, with the ones that exist being generally used as single LSTM or GRU networks [24,46]. Moreover, the high predictive accuracies reported in some of the studies have often been attained on small or highly curated sets of data, which is a cause of concern about overfitting and practical implications in the real world [48,49]. These aspects point to the necessity of architectures with a balance in terms of temporal modeling functionality and computational efficiency. These shortcomings make the current research relevant, suggesting a hybrid BiLSTM-BiGRU model with imbalance-aware sampling to increase the predictive robustness, generalizability, and clinical usability. Tables 1 and 2 below give the detailed analysis of related studies that were reviewed.
In this section, the research methodology and experimental approach are outlined. It also gives a detailed analysis of the datasets under study, the proposed DNN model, the performance evaluation metrics used, and the overall experiments performed.
3.1 Dataset Source and Description
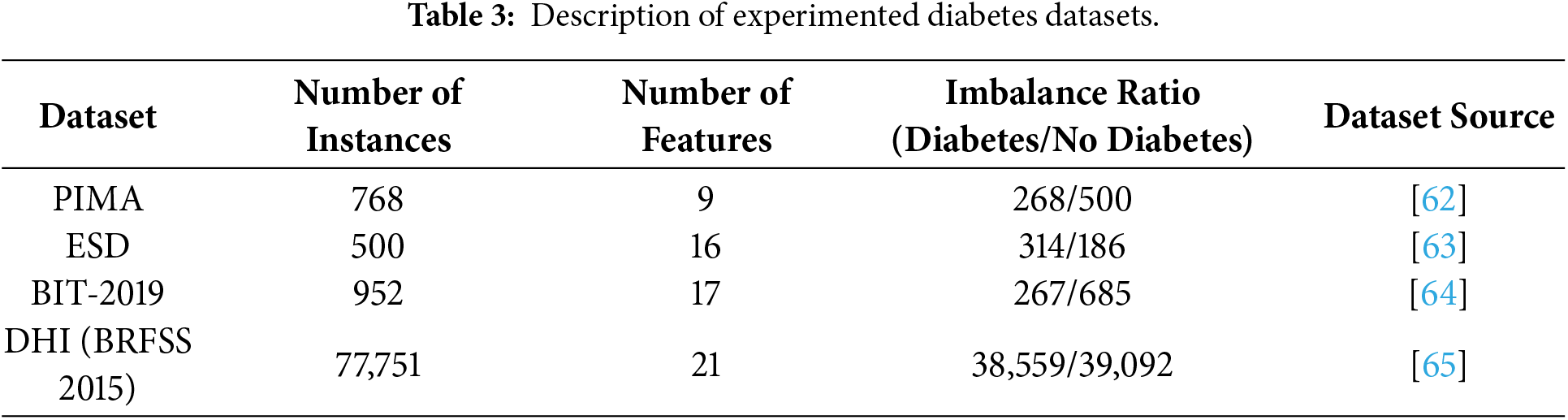
This experiment utilizes four different popular Diabetes datasets. They include PIMA, ESD John dataset, BIT-2019 dataset and DHI (BRFSS 2015) dataset. Most of these datasets are presented by the National Institute of Diabetes and Digestive and Kidney Diseases [39] and can be publicly accessible. In general, these datasets consist of features such as age, glucose, pregnancies, gender, skin thickness, junk food, stress, blood pressure, family diabetes history, BMI, insulin, sleep duration, alcohol intake, diabetes pedigree function, urine frequency, height, weight, location, cholesterol, waist/hip ratio, and outcome. Table 3 provides a numerical description of these datasets. The datasets are significantly different in terms of scale, composition of features and ratio of imbalance, which gives a diverse assessment environment of the robustness of models.
3.2 Data Harmonization and Preprocessing
To enhance the quality of the input datasets, various data preprocessing techniques were applied. Although most of the datasets, as shown in Table 4 are standard and popular, they still need preprocessing to enhance their quality before using them for the experiments. Most of the datasets have instances where values are null or zero (0). For instance, in the datasets, one can find the insulin or blood pressure values to be with 0 or null values, while records like that are not possible for a living human being. Consequently, this research considered these irregularities, such as null or zero (0) instances, as outliers and cleaned them before utilizing the datasets. One of the most adopted methods in cleaning the dataset is the mean imputation method, where the missing values are calculated and imputed. For missing value instances that are not obtainable using the available methods, are categorized as outliers and subsequently removed from the affected dataset, as part of data processing. Similarly, for better interpretation and improved readability, the experimental outcomes are either converted into categorical variables with 1 as Diabetic and 0 as Non–Diabetic, or with Diabetes and No Diabetes.
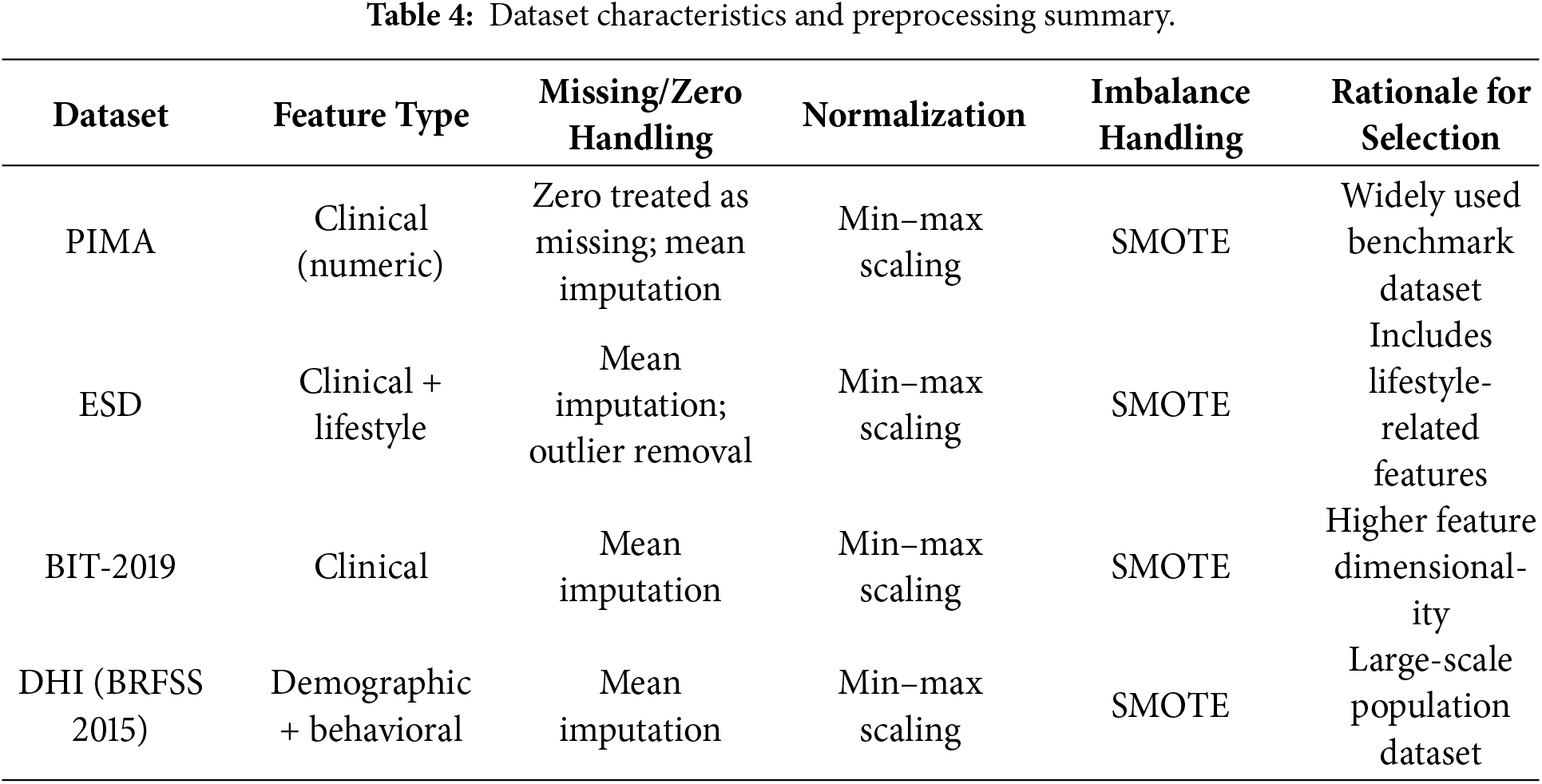

Furthermore, this study has minimized the problem of data imbalance, where the classes within each dataset and between the datasets are not equally represented. The mean imputation method is augmented with the Synthetic Minority Oversampling Technique (SMOTE) to balance the data to have a balanced representation of the class label. This process is crucial to guarantee fairness and robustness for the developed models. The dataset characteristics and preprocessing summary are shown in the Table 4.
3.3 Proposed Deep Classifier: A Data Sampling-Based Hybrid BiLSTM-BiGRU Neural Network
The DeepClassifier represents an advanced approach to Type 2 diabetes prediction, leveraging an advanced hybrid neural network architecture. Fundamentally, the model employs both BiLSTM and BiGRU, two strong versions of RNN, which have been known to learn both complex temporal dependencies in sequence data.
The model has an input layer, which is connected to a dense layer of 128 neurons, then there exists a batch normalization layer and a dropout layer of 0.2 to avoid overfitting. The resulting pre-processed input is then split into parallel BiLSTM and BiGRU layers having 64 units each. These bi-directional recurrent layers enable the model to analyse the input both forward and backward, as this will allow the model to capture a broader context of the data. The results of the BiLSTM and BiGRU layers are then summed together to make a rich representation, which is the combination of the advantages of both recurrent architectures. This combined output is flattened and sent through a sequence of dense layers. The model uses three thick layers that are of diminishing size, separated by batch normalization and dropout layers to ensure regularization in the network.
The final layer of the model consists of a single neuron with a sigmoid activation function, tailoring the output for binary classification-in this case, predicting the presence or absence of Type 2 diabetes. This architecture’s depth and complexity allow it to learn intricate patterns and relationships within the input data, leading to more accurate and reliable predictions of diabetes risk. Fig. 1 contains a full architectural description of the proposed DeepClassifier, presenting the data flow, parallel BiLSTM-BiGRU branches, and then dense layers, which allows the complete reproducibility of the model.
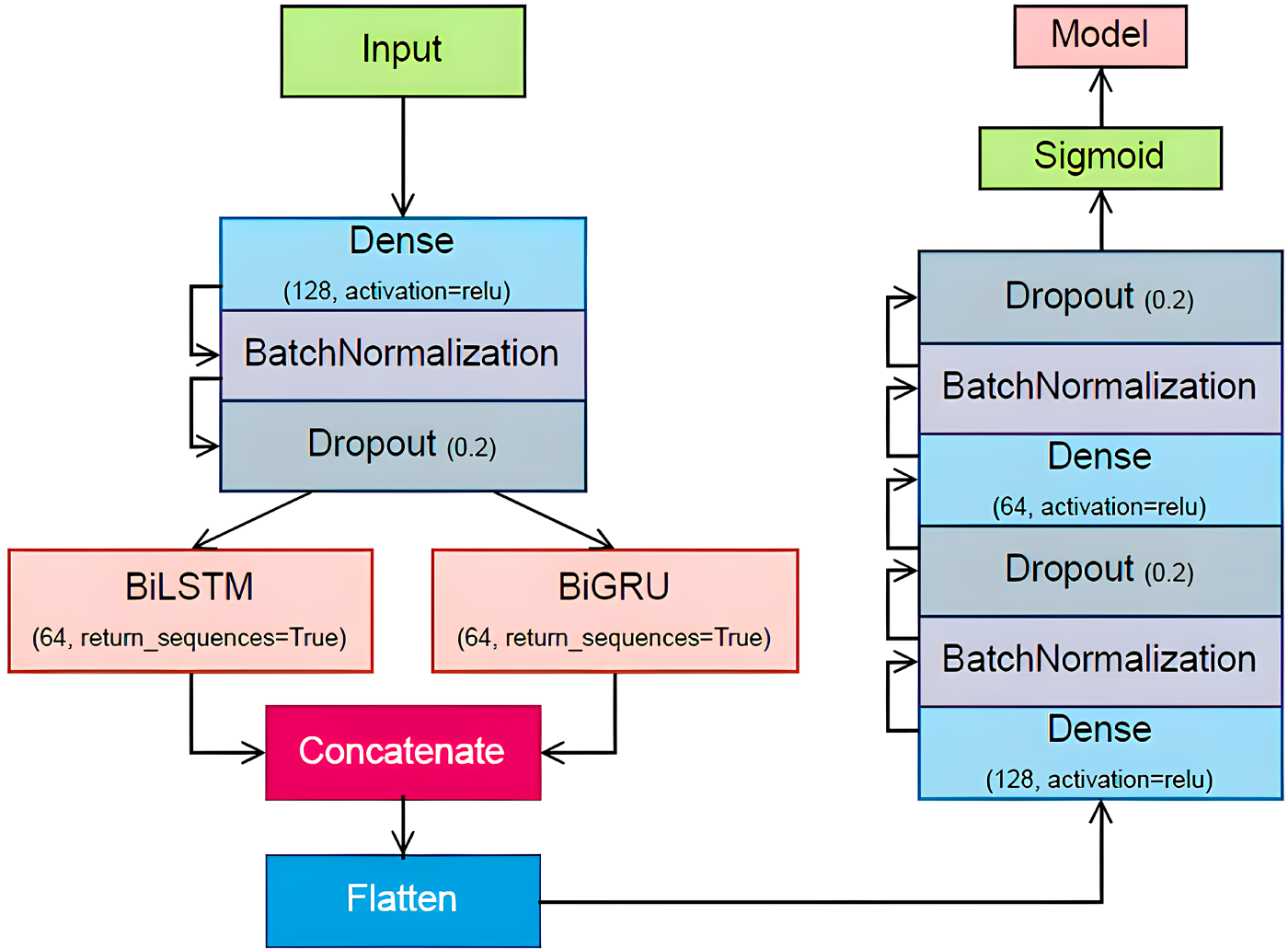
Figure 1: Illustration of the proposed DeepClassifier methodology.
The following section provides details about each of these layers and the working principle of the DeepClassifier’s methodology.
The first stage of a neural network is the input layer, which is the point of entry of the data to be processed in the network. It simply forwards the data to the first hidden layer, where the data is multiplied by the weights of the first hidden layer. The neurons of the input layer are associated with a feature of the input data so that each bit of information is analyzed separately by the network.
A layer with all its neurons fully connected to all the neurons of the preceding layer is called a dense layer (also referred to as a fully connected layer). This layer does a linear transformation of the data and then a non-linear activation function. The process entails the multiplication of the input data with a weight matrix and a bias term, and the result is subjected to the activation function to give the output.
The linear transformation can be defined as
where y is the output vector of the dense layer, W is the weight matrix, x is the input vector and b is the bias vector.
3.3.3 Batch Normalization Layer
The batch normalization is a method of enhancing the stability and speed of neural network training. It standardizes the input layer by adjusting and scaling the activations. In training, the mean and the variance of the mini-batch are calculated and used to normalize the data through batch normalization. This is done to minimize the internal covariate shift, which is the reduction in the distribution of network activations as a result of the change in network parameters as training progresses. The batch normalization is also a regularizer, which minimizes the use of other regularizations.
In the case of a mini-batch whose inputs are denoted as
where m is the number of samples in the mini batch. The normalization step is computed as:
where
where
Dropout can be expressed using a binary mask. During training, each unit is kept with a probability p (typically set to 0.5 for the hidden layers). If
where
3.3.5 Bidirectional LSTM (BiLSTM) Network
LSTM is a special kind of RNNs introduced to deal with the long-term dependencies by solving the problem of vanishing gradient. This problem occurs when information about the input diminishes as it passes through multiple layers, making it difficult to handle long-term dependencies. LSTM addresses this issue by incorporating a gating mechanism that regulates information flow within the network. LSTM comprises several critical layers, each performing distinct operations to manage and update the cell state, thereby enabling the network to maintain long-term dependencies. The overall structure of LSTM and BiLSTM is shown in Fig. 2.
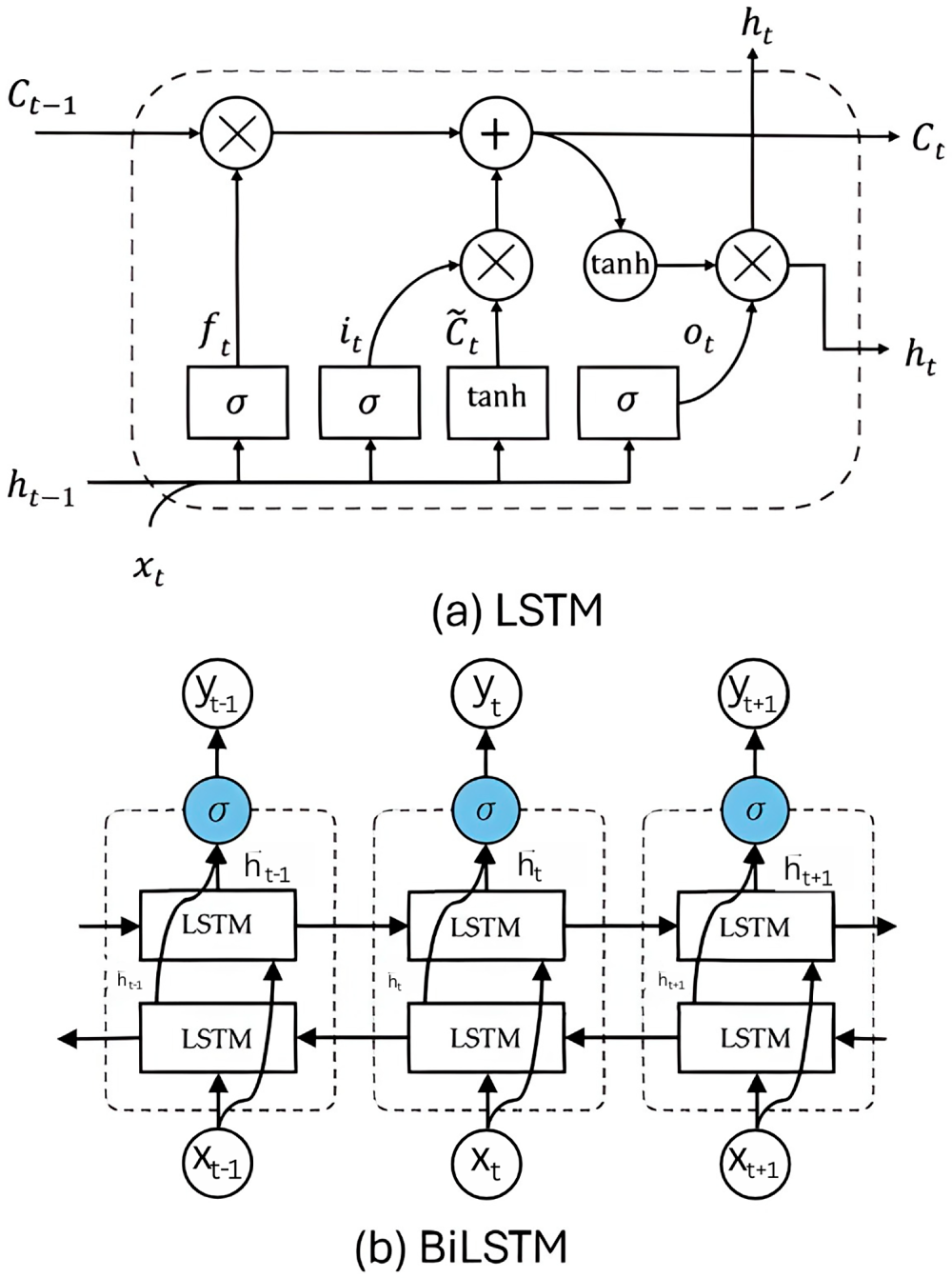
Figure 2: Structure of (a) LSTM and (b) BiLSTM.
As depicted in Eq. (7), the forget gate layer
As shown in Eqs. (13)–(15), the output of a BiLSTM layer is a vector representing the combined information from both forward and backward LSTM layers at each time step. This vector is typically obtained by applying a combination function, such as concatenation, to the hidden states of the two LSTM layers. In the context of diabetes prediction, the BiLSTM layer produces a final output: a vector that predicts for several future time points, the probability of a patient having diabetes.
3.3.6 Bidirectional GRU (BiGRU) Network
Gated Recurrent Unit (GRU) is a simplified version of the LSTM network that adaptively captures dependencies of different time scales by combining forget and input gates into a single update gate and lacks an independent memory cell. The GRU network uses update and reset gates to control the flow of information in the hidden states. These gates determine which data should be preserved for subsequent calculations and which can be discarded. The overall cell structure is depicted in Fig. 3.
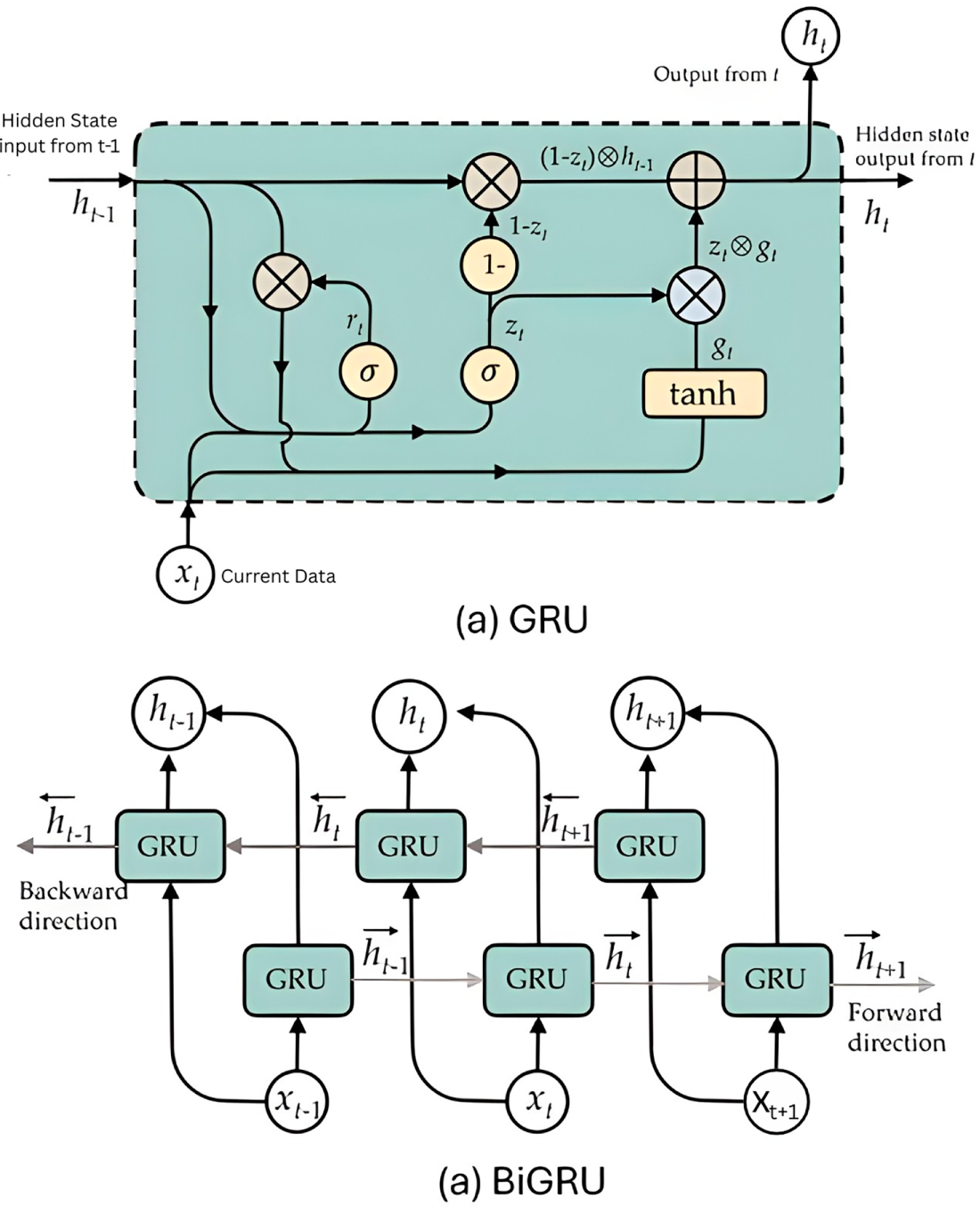
Figure 3: Structure of (a) GRU cell and (b) BiGRU.
As it is shown in Eqs. (16)–(19), A GRU calculates the hidden state
The update gate
The Bidirectional GRU (BiGRU) works in both forward and backward directions, and this enables the unit to capture both past and future contexts. Forwards and backward GRUs are independent, and only at the end of the process are their outputs combined. This implies that the positive neuron’s performance of the forward GRU has no correlation with the negative neuron’s performance of the backward GRU, and the opposite holds. The required computation is shown in the following equations:
3.3.7 Concatenation of BiLSTM and BiGRU
The study involves applying a combined BiLSTM and BiGRU model to predict the occurrence of diabetes in patients. Although typically used for sequential data, these models can also be applied to structured datasets to capture complex patterns and interactions among features. The required computations can be represented in the following equation:
where
Algorithm 1 considers the four datasets


3.4 Summary of Modal Architecture and Hyperparameter Tuning
To facilitate easier clarity and reproducibility, Table 5 summarizes all the hyperparameters used in this study in architecture and training.
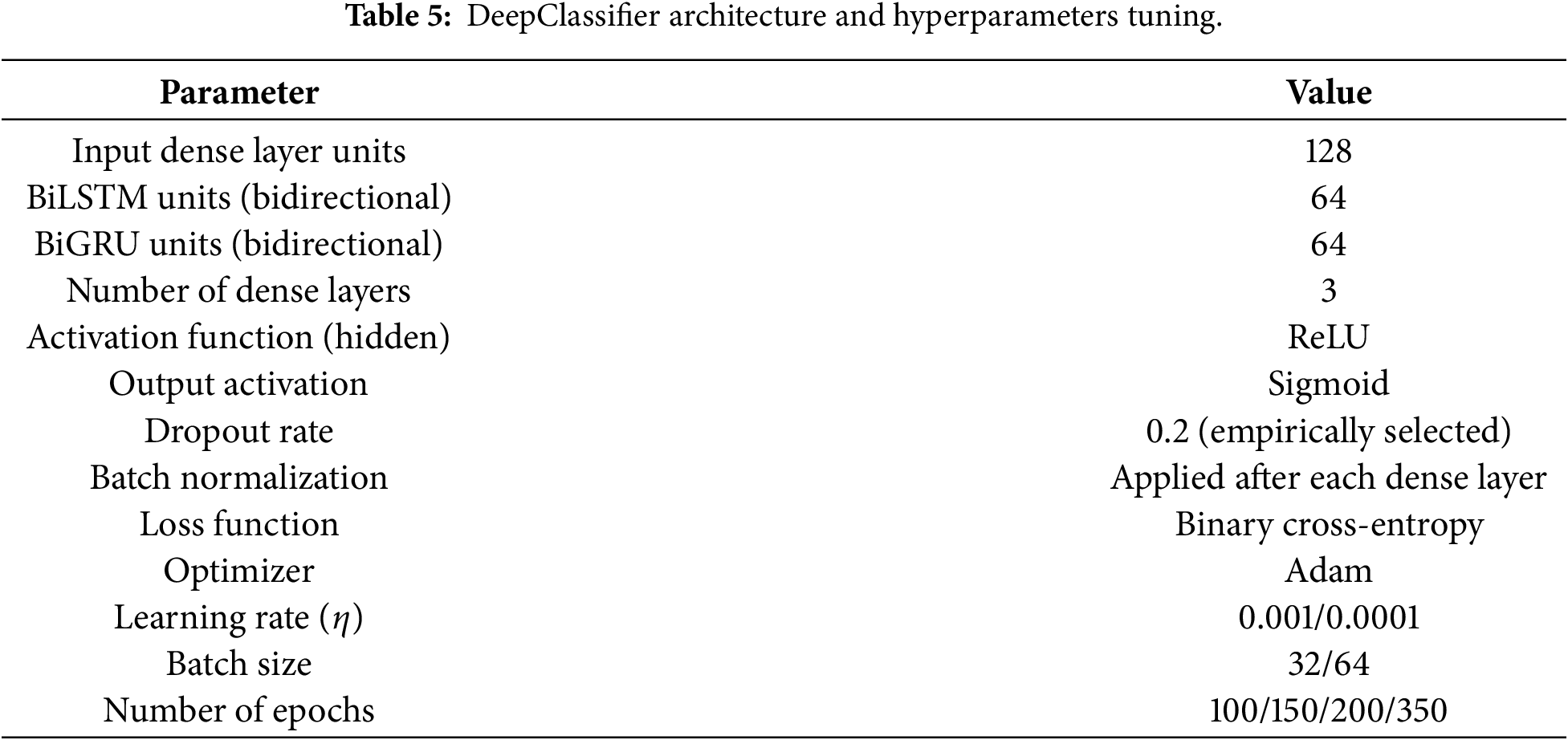

The optimum BiLSTM-BiGRU model in this study was achieved based on systematic empirical testing and not automated hyperparameter optimization methods. The main architectural parameters, such as the recurrent units, dense layers, dropout rate, batch size, and learning rate, were gradually tuned using the validation performance and training stability of various datasets. Although other more sophisticated optimization techniques, like Bayesian optimization or evolutionary algorithm may further improve parameter selection, their computational cost is prohibitive at the scale of heterogeneous datasets with very different scales of heterogeneity. The chosen architecture is a balanced one that had a consistent, stable convergence and a good generalization performance and was hence used as the final model structure in all experiments.
The proposed study involves a comprehensive ML pipeline. Initially, relevant patient data is collected and undergoes rigorous preprocessing to ensure data consistency and quality. Subsequently, the dataset is split into train and test sets for model development and evaluation. The proposed DeepClassifier model is trained extensively on the training data and carefully evaluated on test data. Fig. 4 shows the complete experimental workflow.
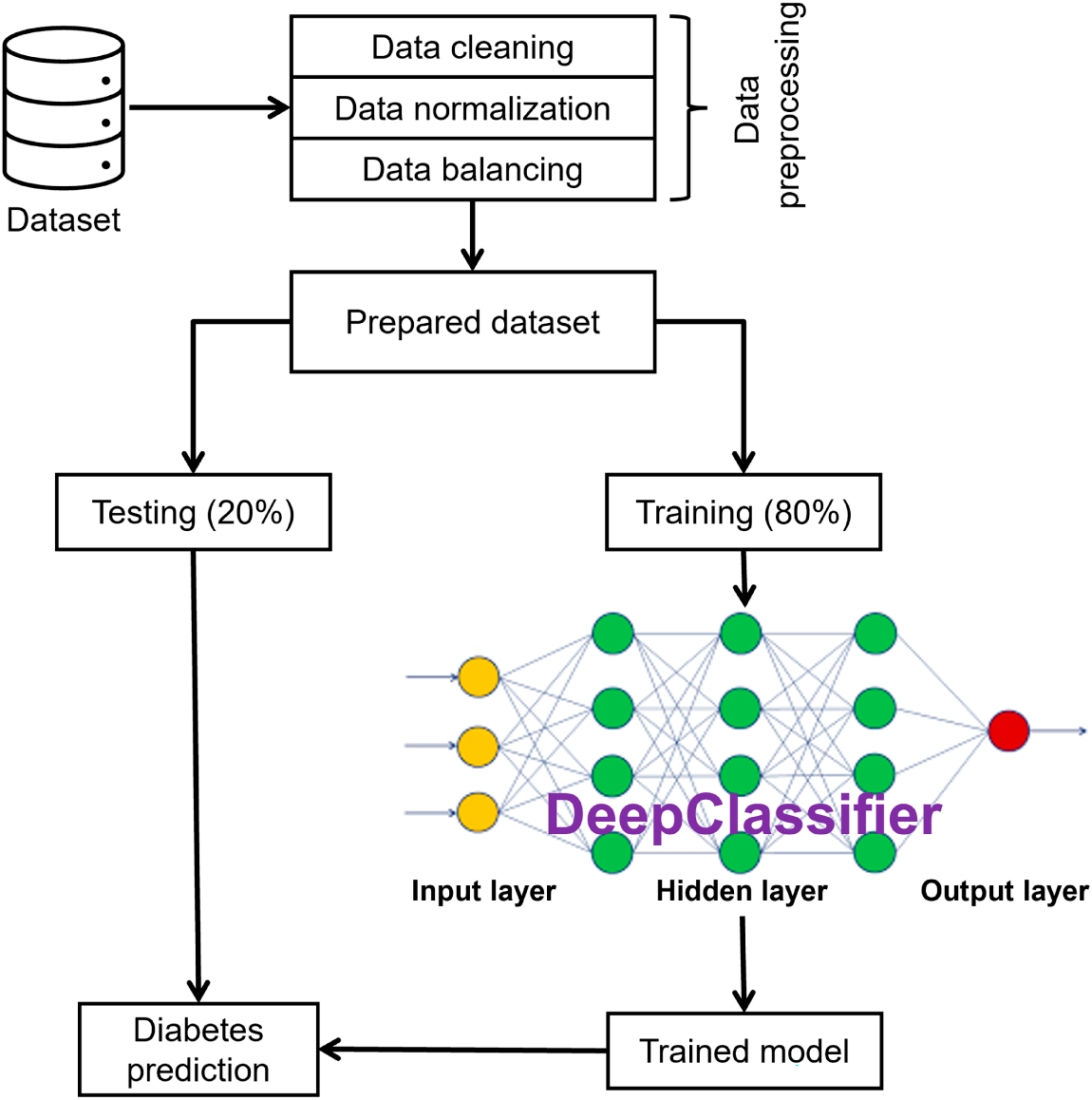
Figure 4: Experimental workflow.
The AMD Ryzen 7 4800H with Redeon Graphics processor and 128 GB RAM-based system was used for these experiments. Python 3.8 and 3.10 versions are used to implement the proposed model. The deep learning frameworks, TensorFlow, Pytorch, and Keras, were applied in two phases.
To compare and fully evaluate the performance of implemented models in terms of predictive performance, accuracy, precision, recall, f-1 score, false positive rate (FPR), true positive rate (TPR) and AUC measures were used. The choice of these metrics was predetermined by their popularity in the existing literature, as well as by the fact that they can provide a complete analysis of the model performance, both strong and weak aspects, under different prediction conditions.
In particular, the addition of TPR and FPR. The importance of AUC is particularly essential, since they display the association between actual and predicted classifications, despite skewed distributions of classes in the dataset. Through these measures, this study will have a reliable, multi-dimensional assessment plan that will be effective in describing the performance of the proposed models. Such a holistic method contributes to the improvement of understanding of the results and makes a significant comparison with the existing models, which makes a significant contribution to the field of health analytics.
Each dataset is experimented on for several epochs, relative to the size of the dataset and the learning rates. For instance, BIT-2019 was executed through 256 epochs, while the PIMA dataset on 100 epochs. Moreover, the ESD dataset increased the number of epochs by 50 against the BIT-2019, while DHI (BRFSS 2015) on 350 epochs due to its large size and variations in the dataset.
Each dataset, i.e., BIT, ESD, PIMA, and DHI (BRFSS 2015) is experimented on the proposed DeepClassifier. At first, the proposed DeepClassifier is trained using 80% of the dataset in question, after which the remaining 20% is used to test the prediction on multiple performance metrics.
The observed cumulative number of patients from across all datasets shares very similar attributes, such as age, height, BMI, weight, and gender. However, some datasets contain more attributes than others, with some attributes having dissimilar structures. This is considered a big problem that affects many of the existing homogeneous solutions. For instance, a solution could be highly compatible with the PIMA dataset while scoring very low on the BIT dataset. Despite this lack of uniformity in the samples and the dataset’s structural complexities, DeepClassifier was able to exhibit its flexible characteristics by adjusting its learning abilities while maintaining closely promising results across all the participating datasets.
Fig. 5 describes the results obtained from the experiment on each of the four datasets. As can be observed, DeepClassifier performs significantly well on all datasets, having the lowest accuracy value of 86% on the DHI (BRFSS 2015) dataset. This has demonstrated the good indiscriminative ability of DeepClassifier. Discrimination was, however, observed in other state-of-the-art solutions as reported in Table 1 and 2. In DeepClassifier, this is minimized with intercept recalibration during training and logistic recalibration. Indicating the overall model performance.
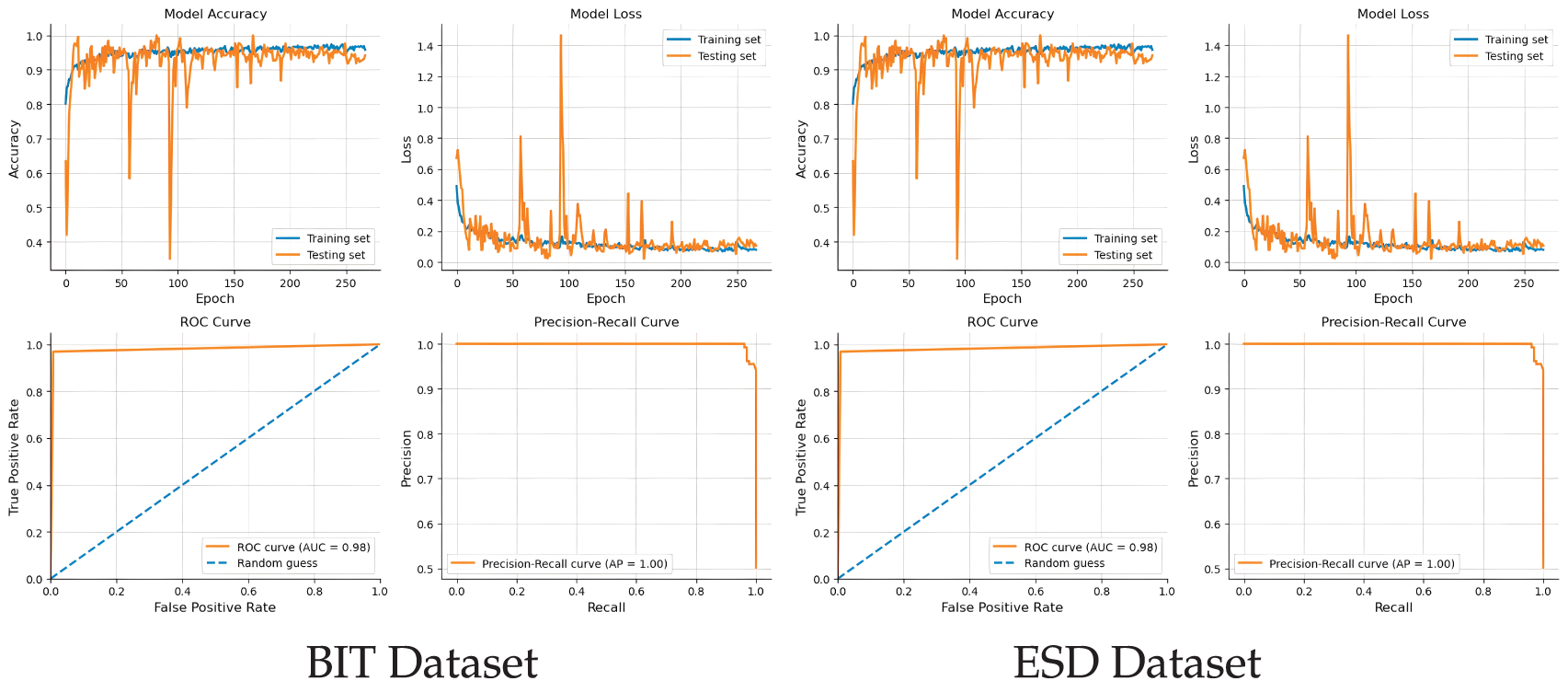
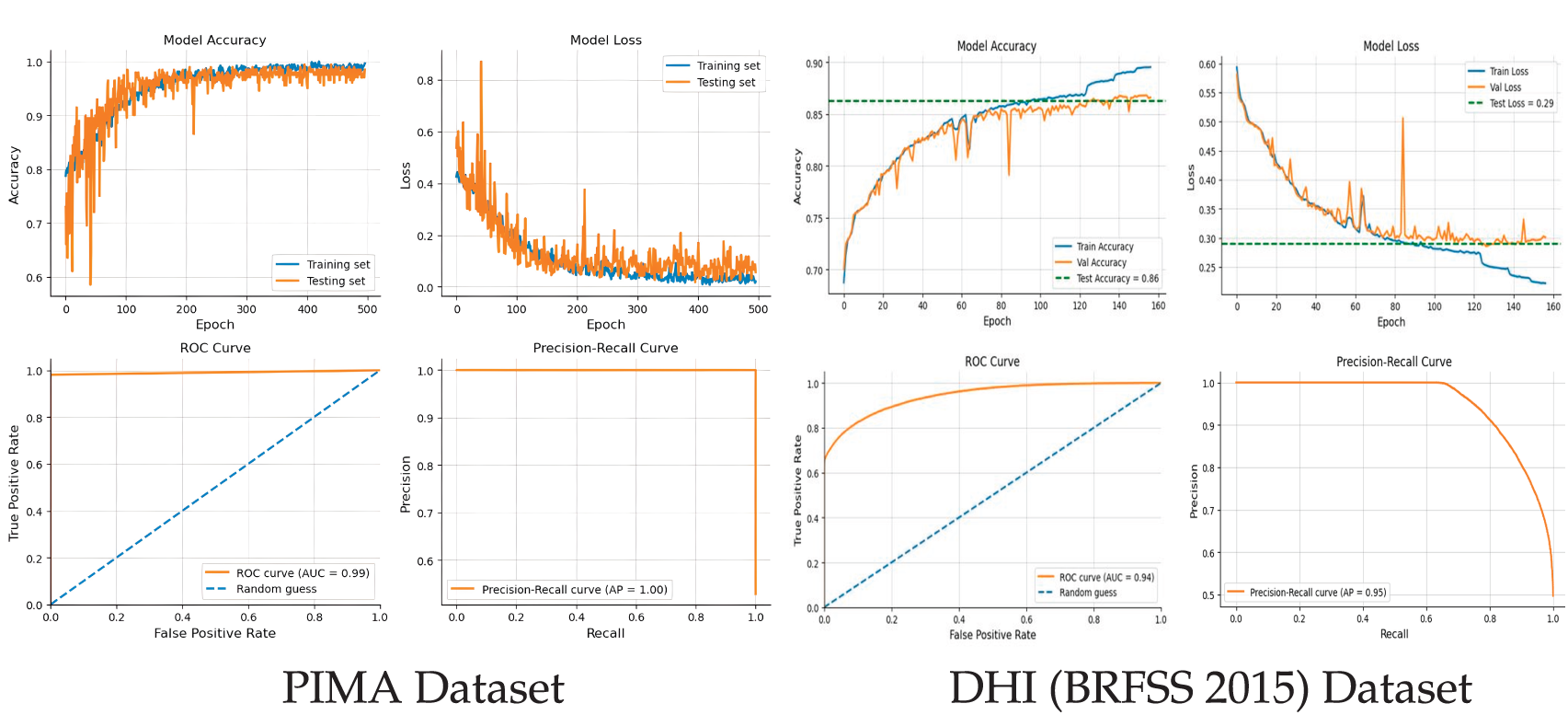
Figure 5: Performance analysis of the proposed model on experimented datasets.
While DeepClassifier did not exhibit similar performance on the ESD dataset, the gap is not considered significant. On the ESD dataset, DeepClassifier can predict correctly with 99% accuracy while having 98% recall, 99% f1-score, and a precision of 100%. This is considered a big achievement, especially since the ESD datasets are not of the same structure. Such achievement did not stop at ESD only. Similarly, the performance of DeepClassifier on PIMA datasets is parallel to that of ESD datasets. With PIMA, the accuracy obtained is 99%, precision of 98%, and f1-score of 99%. In addition, the recall value recorded on PIMA is up to 100%. Moreover, a consistent and similar performance is observed from the results obtained on the ESD and PIMA datasets. However, on PIMA, the precision is 98% while it’s up to 100% on ESD. The opposite is observed on recall value, having 98% for ESD and 100% for PIMA.
Furthermore, DeepClassifier continues to produce good results even on a more complex data structure, such as BIT datasets. On this dataset, DeepClassifier recorded a value of 98% accuracy and f1-score, while scoring higher on precision and recall with values of 97% and 99%, respectively. The performance of DeepClassifier was also measured on a large-sized DHI (BRFSS 2015) dataset, with recorded accuracy of 86% and promising precision and recall values. This consistent performance across all datasets is one of the important qualities of DeepClassifier. The following section describes the distribution of the results on ground truth, prediction, and comparisons.
With ESD, the ground truth test data shows a more distinct distribution than the ground truth of the PIMA dataset. This is consciously observed on prediction distribution with negative and positive cases. By comparison, it can be concluded that there are more false positives in ESD datasets than in PIMA. This is partly attributed to the data preprocessing techniques that are adopted by DeepClassifier.
Contrariwise, in the BIT and DHI (BRFSS 2015) dataset comparative distribution, there are more false negatives and false positives than in any other dataset. The overall results distributions of the proposed model are shown in Fig. 6.
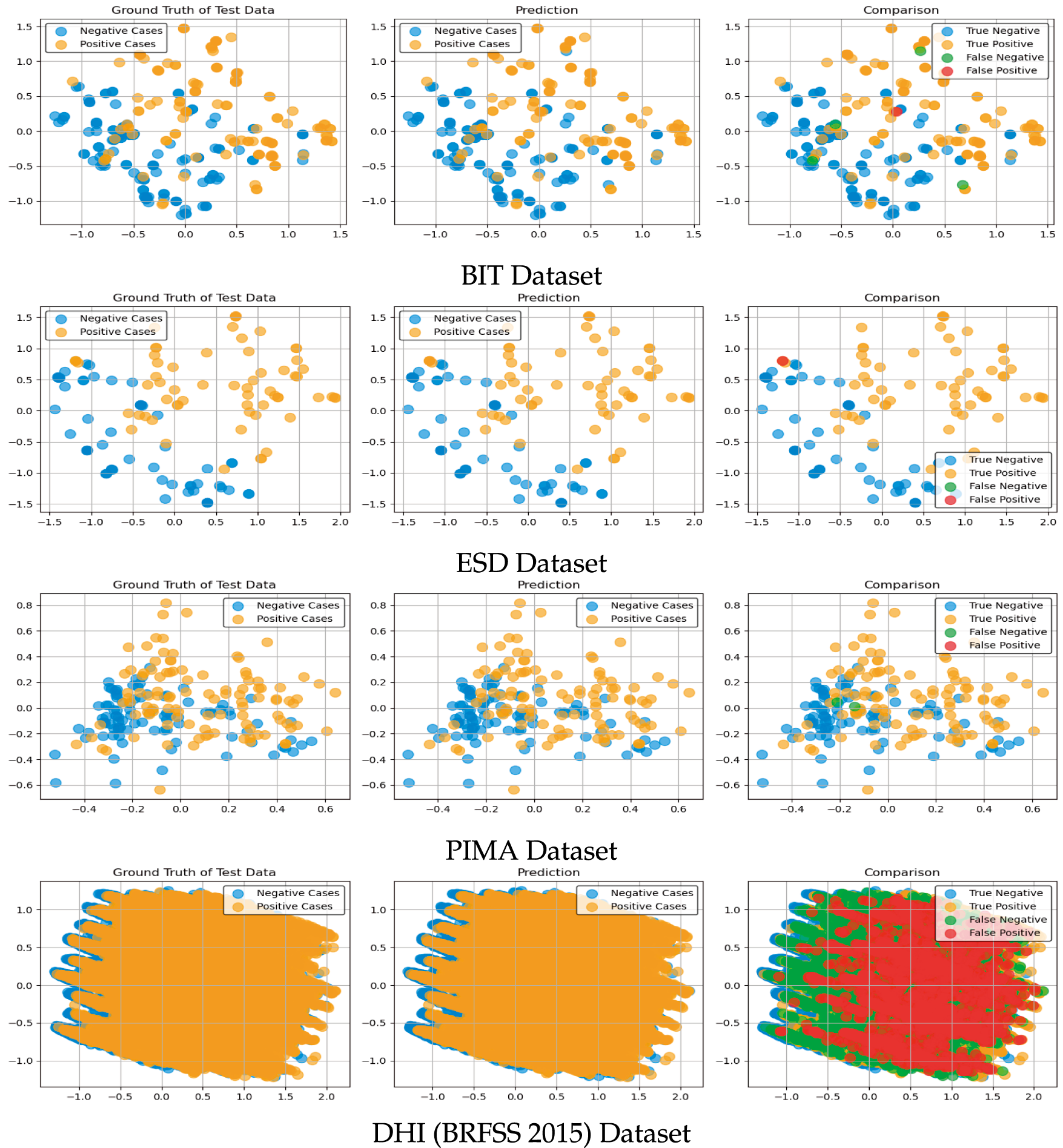
Figure 6: Results distribution of the proposed model on the experimented datasets.
4.3 Further Experimental Result Analysis
This study demonstrates that Type 2 diabetes can be accurately predicted with a heterogeneous dataset using a novel data sampling-based hybrid DNN model. At its core, the proposed model can capture and accommodate complex temporal dependencies in sequential data. This functionality is one of the key abilities of the proposed model. That is, DeepClassifier can predict Type 2 diabetes across different structural datasets.
This section aims to provide a more detailed clarity of the results beyond what performance measurement metrics offer. Using a confusion matrix such as True Positive (TP), True Negative (TN), False Positive (FP), and False Negative (FN), we will be able to identify the exact, correctly predicted patients and those that are predicted wrongly. These values should be able to correspond to the results presented in the previous section. The following subsections present the confusion matrix for all four datasets.
With the BIT dataset, the proposed DeepClassifier was able to correctly classify 127 patients (TP) and 125 patients as true negatives (TN), as depicted in Fig. 7 below. However, with the BIT dataset, DeepClassifier wrongly classified 1 patient as a False Positive (FP), and 4 patients as False Negative (FN). This wrong classification makes the BIT dataset structure to be the list compatible structure with the proposed DeepClassifier. However, this performance may only be considered low when compared to high performance with datasets, as may be observed in the following sections.
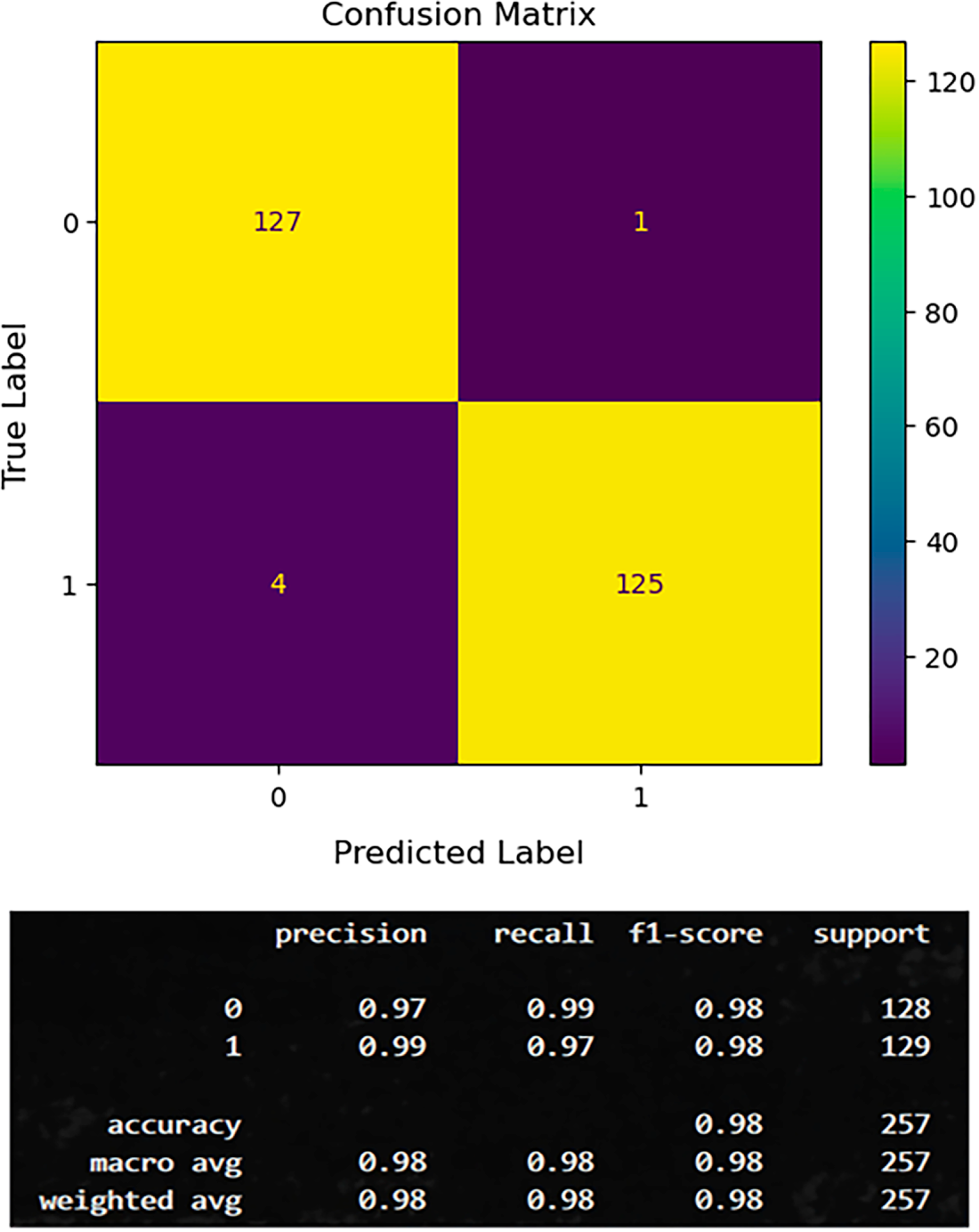
Figure 7: Confusion metrics of the proposed model on BIT-2019 datasets.
Unlike in the previous section, where DeepClassifier wrongly classifies 5 patients using BIT dataset, DeepClassifier proves to be more compatible with the ESD dataset, as only 1 patient is wrongly classified. As shown in Fig. 8, up to 65 patients are correctly classified as TP, while up to 62 patients are correctly classified as TN. Referring to Fig. 8, we can observe that while DeepClassifier records one false positive in its prediction, none were recorded for false negatives. This is still considered a lesser risk compared to recording a false negative, like with the PIMA dataset. This achievement clearly explains why the ESD dataset’s structure is considered more compatible with DeepClassifier than the BIT and PIMA datasets.
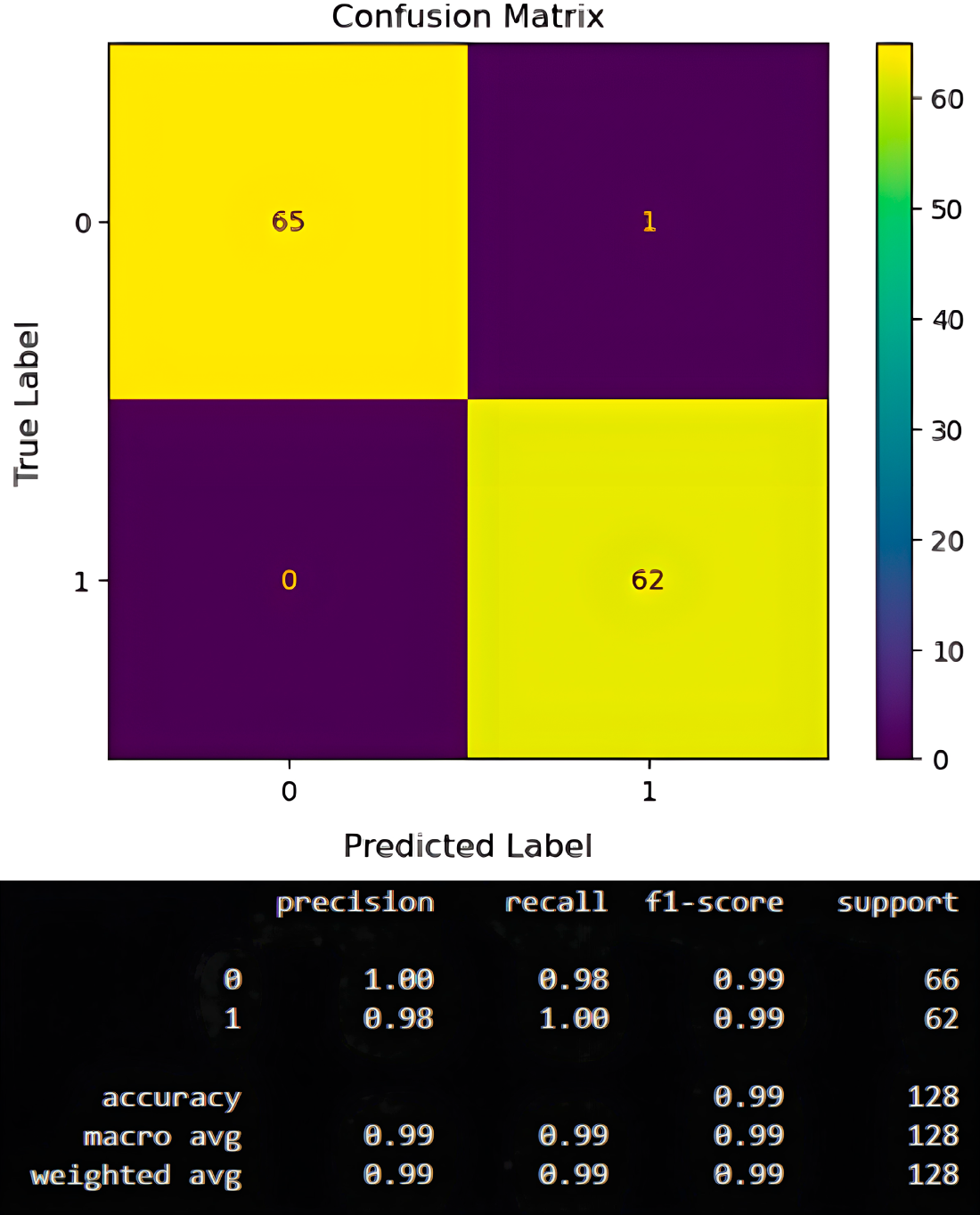
Figure 8: Confusion metrics of the proposed model on ESD john datasets.
To minimize dataset structural dependencies, PIMA was also applied to DeepClassifier while performing hyperparameter tuning. DeepClassifier was able to correctly predict up to 94 patients as true positives and 104 patients as true negatives. This is depicted in Fig. 9 below.
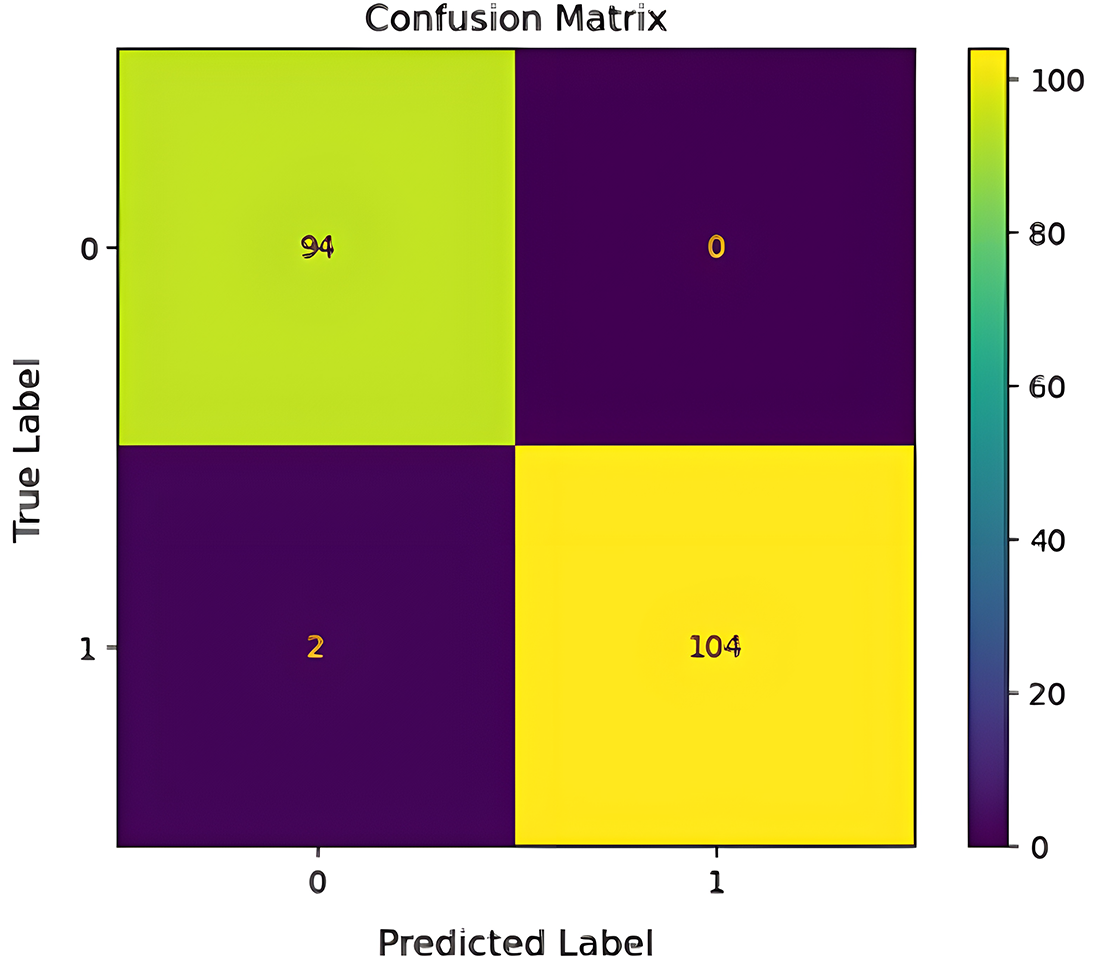
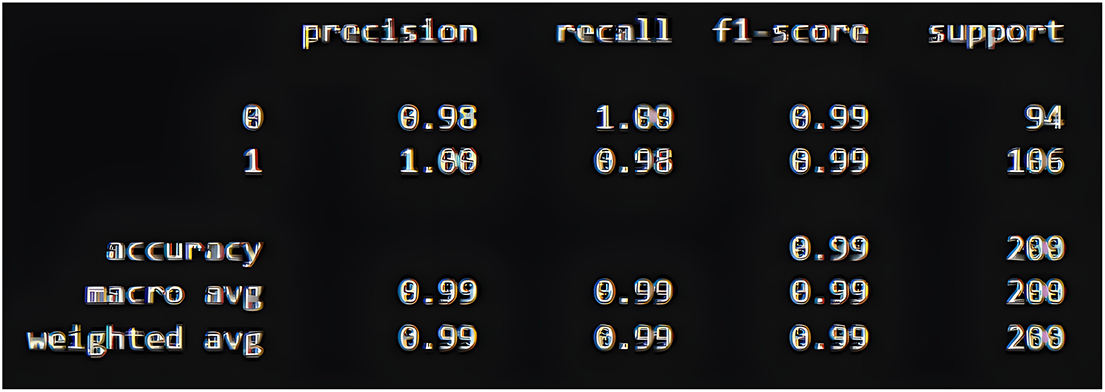
Figure 9: Confusion metrics of the proposed model on PIMA datasets.
4.3.4 DHI (BRFSS 2015) Dataset
The confusion matrix for the DHI dataset shows that the DeepClassifier correctly classified 29,359 diabetic and 37,320 non-diabetic individuals. At the same time, 9300 diabetic cases were missed, and 1772 non-diabetic cases were wrongly flagged. Overall, the model demonstrates reliable performance, though the presence of false negatives indicates a need for further refinement in the dataset to minimize missed diabetic cases as shown in Fig. 10 below.
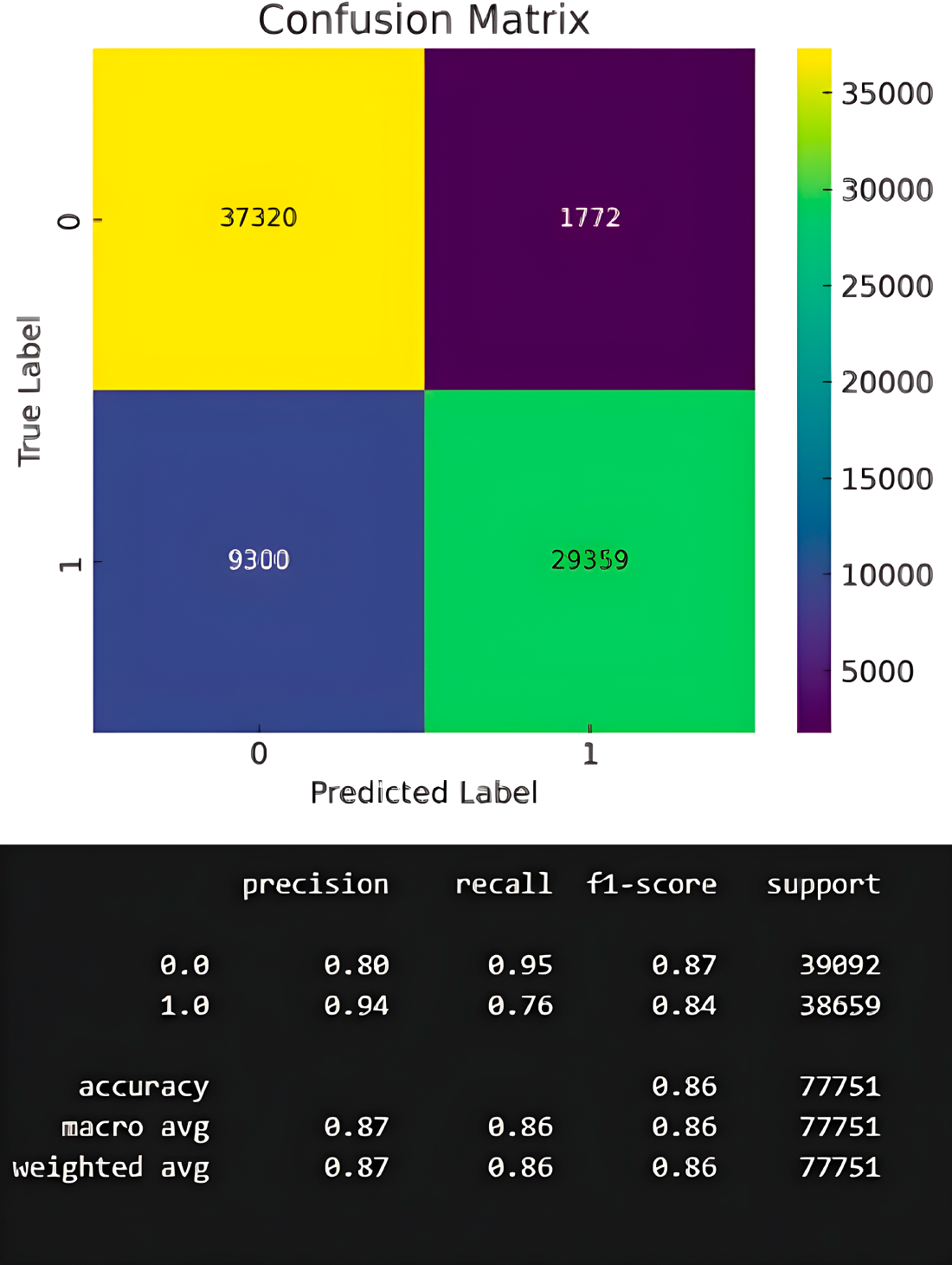
Figure 10: Confusion metrics of the proposed model on DHI datasets.
While this is considered an excellent performance for the DeepClassifier, the false negative prediction slightly negates this conclusion. Up to two false negatives are recorded using the PIMA dataset, which is slightly lower than what is obtainable using the BIT dataset.
In summary, the results are interpreted in the context of high accuracy and generality of model application amongst heterogeneous datasets. The various data preprocessing measures used as variable predictors to prepare all the datasets may be subject to computational errors. However, DeepClassifier was designed to accept all forms of related data regardless of its structural deformity and unlike patterns, therefore, it is highly unlikely that such peculiarities can meaningfully affect model performance.
Particularly, the use of multiple datasets, which are obtained from different sources, having different parameters and values, has provided additional generalizability; data structural errors are unlikely to differ significantly from a combination of all the datasets we have used in this study. In addition, evidence-based risk reduction, which is based on an Australian study, was also adopted to assume uptake and adherence are uniform across all datasets. Consequently, the chances of having a dataset with completely dissimilar parameters and structure are considered highly unlikely; thus, proved model generalizability of the proposed DeepClassifier.
4.4 Comparative Analysis with Existing Studies
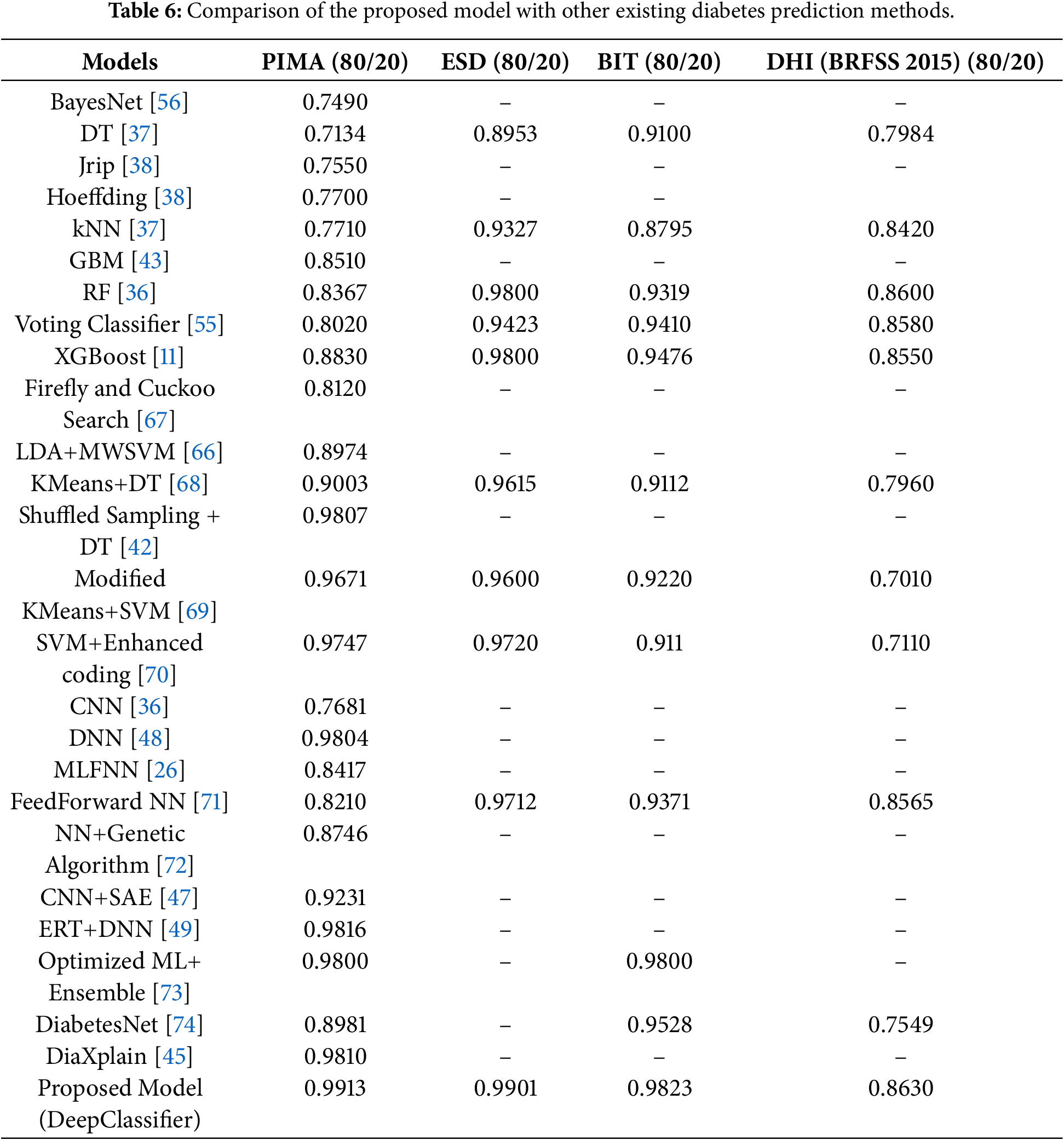
Table 6 emphasizes the comparative analysis of the proposed DeepClassifier model and the current methods based on the prediction accuracy. The experimental findings take advantage of benchmark research, such as methods described in [13,24,25,34,46,55,65,66] to put in perspective the performance of the proposed model. These available methods include a set of computational methods, most of which have been known to have good predictive capabilities.
For instance, several baseline ML models such as BayesNet [56], DT [37], kNN [37] and Jrip [38] with varying computational characteristics have been successfully deployed in existing studies with commendable results. Their aggregate predictive performance at best ranges between 70% to approximately 80% in accuracy values. Although the accuracy values are commendable, there is still room for improvement. The proposed model significantly outperformed the baseline MLs in terms of prediction accuracy.
Some advanced methods, such as hybrid ML models and ensemble methods, have also been deployed. Particularly, ensemble and hybrid models are known to be superior in performance to baseline ML models. Hybrid ML models, as proposed by [42,65–68,69] were better than the baseline ML models. Similar trends were noticed in the case of ensemble-based models presented in [11,36,43,55]. Regardless, the proposed models still achieved a higher predictive accuracy value than the advanced ML models. Besides, most of the existing studies lack generalizability, as they were primarily conducted on PIMA, which is a drawback.
Still from Table 6, further comparisons were presented as the proposed model is compared with existing DL-based models. Several existing studies with diverse DL architectures ranging from single NN models [36,48,70] to stacked autoencoders [47] and, in some cases, hybrid DL structures [26,49,71]. The proposed models surpassed these recent DL-based methods, and this further affirms the proposed model’s robustness and practical applicability in diabetes prediction.
Explainable deep networks, hybrid networks, and ensemble models are recent architectures of deep learning used to predict diabetes. Although these methods report good performance, they have commonly been tested on a single dataset, usually PIMA, and use simple train-test splits, which do not test their generalizability. Conversely, the proposed DeepClassifier is compared to classical and more recent deep learning models on a variety of heterogeneous data and supported by cross-validation and analysis. This more comprehensive analysis offers a more credible point of view of strength and points to the practical benefits of the proposed hybrid BiLSTM-BiGRU architecture in predicting diabetes.
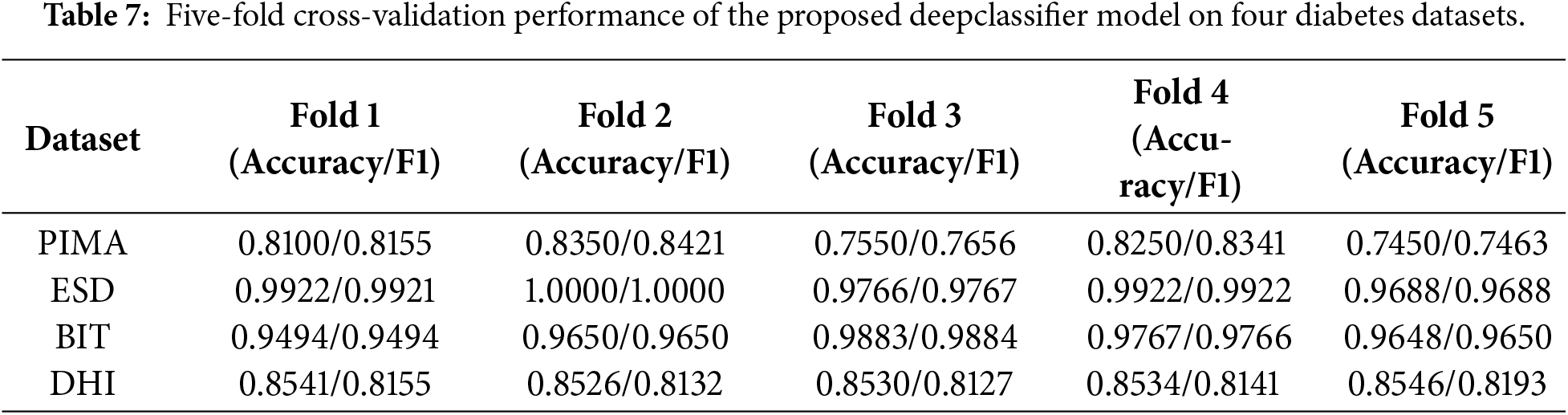
Furthermore, Table 6 shows that the prior studies’ results are evaluated using an 80/20 train/test split without cross-validation, so we compared the results of our model with these baselines. As those studies did not report fold metrics or raw estimates, formal significance testing could not be performed. In order to verify our model’s robustness and generalizability, we applied 5-fold cross-validation, which provides a more stable performance estimate, as shown in Tables 7 and 8.
4.5 Interpretation of Cross-Validation and t-Test Results
In order to test the technical rigor of our proposed DeepClassifier model, further applied 5-fold cross-validation and a t-test to check the model’s performance on four datasets. The 5-fold cross-validation experimental results are shown in the Table 7.
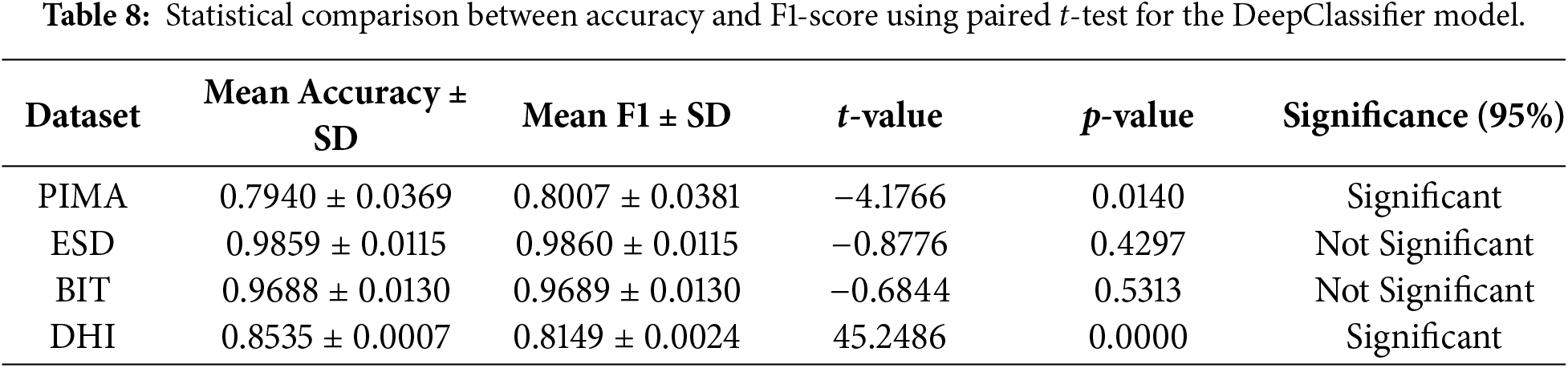
The results of the t-test applied to compare the significant difference between mean accuracy and mean F1-score are shown in Table 8. The experiment findings on the four diabetes datasets show that there are interesting variations in the performance metrics of accuracy and F1-score.
The mean accuracy (0.7940 − 0.0369) and F1-score (0.8007 − 0.0381) are very close in the case of the PIMA dataset, but the t-test (t = −4.1766, p = 0.0140) shows that the difference is statistically significant. This indicates that both measures exhibit similar tendencies, though slightly different variations in the class balance or model precision-recall behaviour influence the F1-score more significantly, resulting in a quantifiable deviation.
Conversely, the performance of the two metrics is significantly similar in the ESD and BIT data sets. The means of their accuracy and F1-score are within the range of less than 0.001, and the t-tests (
Accuracy and F1-score have a significant difference (
In general, all these results suggest that the accuracy and F1-score usually change in the same direction, but the sensitivity of these metrics to class imbalance and consistency of predictions differ among datasets. In the case of datasets such as PIMA and DHI, both measures will allow a more detailed view on the performance of the model, but in the case of ESD and BIT, one of the metrics is likely to be adequate to describe the reliability of the classifier.
Even though the results of statistical hypothesis testing cannot be applied in the cases of comparison of the results obtained in independent studies, Table 8 shows the evident performance improvements of the proposed DeepClassifier that were obtained in relation to the previously published techniques. In datasets of similar scale, the suggested method always achieves a superior or competitive accuracy and, at the same time, increases F1-score, meaning a more balanced classification performance. Conversely, other previous studies have shown high accuracy but with relatively lower F1-scores, especially in imbalanced datasets. This is an indicator of the effectiveness of the hybrid BiLSTM-BiGRU model with imbalance-aware sampling, which allows detecting the cases of minority classes more reliably than the current ML and DL methods.
In addition to effectively addressing key challenges in health analytics and diabetes prediction, such as class imbalance, cost-sensitive classification, and optimal performance, the proposed model can also be seamlessly integrated into contemporary EHRs. The model has the advantage of predictive capabilities that can aid early identification of individuals at risk for Type 2 diabetes, providing actionable insights for clinicians.
Similar to numerous other studies of the past, the current research has a number of limitations that can affect the interpretation of the results obtained and their generalizability. The two primary threats to validity are external and internal validity; the former is concerned with the applicability of the results to the population outside of the dataset and situation in which the research was conducted, and the latter is related to the correctness of the causal inferences made in the research. External validity is particularly important, as it allows knowing how effective the predictive models can be used in broader populations or other systems. As an illustration, the open-source tools used to create a model can produce results that are highly situation-dependent and can not be reliably applied to different datasets or different platforms. Also, the historical data that was utilized in this study also restricts the generalizability due to the fact that the data sets that were utilized could not be a complete reflection of the variability that could occur in the real world.
The potential confounding variables are also a challenge to internal validity and may obscure the true relationship between independent and dependent variables. Such problems as missing data or non-cumulative numerical values should be addressed (with attention) to prevent the biasing of the analysis. The reliability of the study conclusions might also be endangered by misinterpretation of the causal relations between the software metrics and defect occurrences.
The proposed prediction model is also specific to certain conditions of the operation, and it is most efficient in conditions such as those in which it has been developed. Their efficiency can be lowered when used on systems of a different nature. To surmount these issues, subsequent studies must involve larger experimental replications based on different data sets and platforms. These would assist in strengthening, increasing the flexibility, and external validity of the models in general, thus alleviating fears surrounding external validity.
In spite of these shortcomings, the research has provided valuable information on the application of DL models to predict the reliability. In particular, it shows how the characteristics of historical datasets affect predictive performance. Such results can be used to base further research on enhancing the scalability of the models, their cross-domain applicability, and reliability in the long term.
This study proves that enhanced Type 2 diabetes can be accurately predicted by performing data sampling techniques on heterogeneous datasets and by using two powerful variants of recurrent neural networks, such as Bidirectional Long Short-Term Memory (BiLSTM) and Bidirectional Gated Recurrent Unit (BiGRU). Through our experiments, we discovered that, with careful hyperparameter tuning, a hybridized model can enhance generalization and consistency across diverse datasets, enabling early intervention and improved disease management. DeepClassifier not only outperformed state-of-the-art models, but it has also demonstrated similar behaviour across dissimilar datasets.
At its core, DeepClassifier recorded while obtaining up to 86% at its lowest performance against a less adaptable dataset. The cross-dataset compatibility is one of the main distinctions of DeepClassifier. The study is limited by class imbalances and missing clinically relevant features in the datasets, which may affect our proposed model’s performance. In the future, we hope to decrease this gap and increase the model’s compatibility on various Type 2 diabetes data collected through different invasive methods, and emphasize real-world validation and interpretability.
Acknowledgement: None.
Funding Statement: This research paper was supported by the School of Digital Science, Universiti Brunei Darussalam, Brunei.
Author Contributions: Conceptualization, Sahalu Balarabe Junaidu, Abdullahi Abubakar Imam, Abdullateef Oluwagbemiga Balogun, Hussaini Mamman and Luiz Fernando Capretz; data curation, Abdullateef Oluwagbemiga Balogun, Ganesh Kumar, Hanif Abdul Rahman, Shuib Basri and Asmah Husaini; formal analysis, Abdullateef Oluwagbemiga Balogun, Sunder Ali Khowaja, Luiz Fernando Capretz and Usman Ali; funding acquisition, Sahalu Balarabe Junaidu, Abdullahi Abubakar Imam, Sunder Ali Khowaja and Ganesh Kumar; investigation, Sahalu Balarabe Junaidu, Abdullahi Abubakar Imam, Abdullateef Oluwagbemiga Balogun, Shuib Basri and Asmah Husaini; methodology, Sahalu Balarabe Junaidu, Abdullahi Abubakar Imam, Abdullateef Oluwagbemiga Balogun, Hussaini Mamman and Sunder Ali Khowaja; project administration, Sahalu Balarabe Junaidu, Abdullahi Abubakar Imam, Luiz Fernando Capretz, Usman Ali and Fatoumatta Conteh; resources, Sahalu Balarabe Junaidu, Abdullahi Abubakar Imam, Hanif Abdul Rahman, Shuib Basri, Abdullateef Oluwagbemiga Balogun and Sahalu Balarabe Junaidu; software, Usman Ali, Abdullahi Abubakar Imam and Ganesh Kumar; supervision, Abdullahi Abubakar Imam, Luiz Fernando Capretz and Hanif Abdul Rahman; validation, Abdullahi Abubakar Imam, Hanif Abdul Rahman, Ganesh Kumar, Hussaini Mamman and Usman Ali; visualization, Abdullahi Abubakar Imam, Abdullateef Oluwagbemiga Balogun, Fatoumatta Conteh, Shuib Basri and Asmah Husaini; writing original draft, Abdullahi Abubakar Imam, Abdullateef Oluwagbemiga Balogun, Fatoumatta Conteh and Asmah Husaini; writing review & editing, Shuib Basri, Fatoumatta Conteh, Hussaini Mamman and Sunder Ali Khowaja. All authors reviewed and approved the final version of the manuscript.
Availability of Data and Materials: The datasets analyzed during the current study are available in the PIMA repository, https://www.kaggle.com/datasets/uciml/pima-indians-diabetes-database (accessed on 10 September 2025); ESD repository, https://www.kaggle.com/datasets/abdelazizsami/early-stage-diabetes-risk-prediction?resource=download (accessed on 10 September 2025); BIT repository, https://www.kaggle.com/datasets/tigganeha4/diabetes-dataset-2019 (accessed on 10 September 2025); DHI repository, https://www.cdc.gov/brfss/annual_data/annual_2015.html (accessed on 10 September 2025).
Ethics Approval: Not applicable.
Conflicts of Interest: The authors declare no conflicts of interest.
References
1. Guan Z, Li H, Liu R, Cai C, Liu Y, Li J, et al. Artificial intelligence in diabetes management: advancements, opportunities, and challenges. Cell Rep Med. 2023;4(10):101213. doi:10.1016/j.xcrm.2023.101213. [Google Scholar] [PubMed] [CrossRef]
2. Tasin I, Nabil TU, Islam S, Khan R. Diabetes prediction using machine learning and explainable AI techniques. Healthc Technol Lett. 2023;10(1–2):1–10. doi:10.1049/htl2.12039. [Google Scholar] [PubMed] [CrossRef]
3. Kumar A, Gangwar R, Ahmad Zargar A, Kumar R, Sharma A. Prevalence of diabetes in India: a review of IDF diabetes atlas. Current Diab Rev. 2024;20(1):e130423215752. [Google Scholar]
4. Holt RI, Flyvbjerg A. Textbook of diabetes. Hoboken, NJ, USA: John Wiley & Sons; 2024. [Google Scholar]
5. Ramachandran A, Snehalatha C, Raghavan A, Nanditha A. Classification and diagnosis of diabetes. In: Textbook of diabetes. Hoboken, NJ, USA: John Wiley & Sons; 2024. p. 22–7. doi:10.1002/9781119697473.ch2. [Google Scholar] [CrossRef]
6. Quattrin T, Mastrandrea LD, Walker LS. Type 1 diabetes. The Lancet. 2023;401(10394):2149–62. doi:10.1016/s0140-6736(23)00223-4. [Google Scholar] [PubMed] [CrossRef]
7. Abdul Basith Khan M, Hashim MJ, King JK, Govender RD, Mustafa H, Al Kaabi J. Epidemiology of type 2 diabetes—global burden of disease and forecasted trends. J Epidemiol Global Health. 2020;10(1):107–11. doi:10.2991/jegh.k.191028.001. [Google Scholar] [PubMed] [CrossRef]
8. Rosengren A, Dikaiou P. Cardiovascular outcomes in type 1 and type 2 diabetes. Diabetologia. 2023;66(3):425–37. doi:10.1007/s00125-022-05857-5. [Google Scholar] [PubMed] [CrossRef]
9. Gregg EW, Buckley J, Ali MK, Davies J, Flood D, Mehta R, et al. Improving health outcomes of people with diabetes: target setting for the WHO Global Diabetes Compact. The Lancet. 2023;401(10384):1302–12. doi:10.1016/s0140-6736(23)00001-6. [Google Scholar] [PubMed] [CrossRef]
10. Ahmad HF, Mukhtar H, Alaqail H, Seliaman M, Alhumam A. Investigating health-related features and their impact on the prediction of diabetes using machine learning. Appl Sci. 2021;11(3):1173. doi:10.3390/app11031173. [Google Scholar] [CrossRef]
11. Kopitar L, Kocbek P, Cilar L, Sheikh A, Stiglic G. Early detection of type 2 diabetes mellitus using machine learning-based prediction models. Sci Rep. 2020;10(1):11981. doi:10.1038/s41598-020-68771-z. [Google Scholar] [PubMed] [CrossRef]
12. Yedjou CG, Grigsby J, Mbemi A, Nelson D, Mildort B, Latinwo L, et al. The management of diabetes mellitus using medicinal plants and vitamins. Int J Mol Sci. 2023;24(10):9085. doi:10.3390/ijms24109085. [Google Scholar] [PubMed] [CrossRef]
13. Popoviciu MS, Kaka N, Sethi Y, Patel N, Chopra H, Cavalu S. Type 1 diabetes mellitus and autoimmune diseases: a critical review of the association and the application of personalized medicine. J Pers Med. 2023;13(3):422. doi:10.3390/jpm13030422. [Google Scholar] [PubMed] [CrossRef]
14. Reaven PD, Newell M, Rivas S, Zhou X, Norman GJ, Zhou JJ. Initiation of continuous glucose monitoring is linked to improved glycemic control and fewer clinical events in type 1 and type 2 diabetes in the veterans health administration. Diabetes Care. 2023;46(4):854–63. doi:10.2337/dc22-2189. [Google Scholar] [PubMed] [CrossRef]
15. Herz D, Haupt S, Zimmer RT, Wachsmuth NB, Schierbauer J, Zimmermann P, et al. Efficacy of fasting in type 1 and type 2 diabetes mellitus: a narrative review. Nutrients. 2023;15(16):3525. doi:10.3390/nu15163525. [Google Scholar] [PubMed] [CrossRef]
16. Anand A, Shakti D. Prediction of diabetes based on personal lifestyle indicators. In: Proceedings of the 2015 1st International Conference on Next Generation Computing Technologies (NGCT). Piscataway, NJ, USA: IEEE; 2015. p. 673–6. [Google Scholar]
17. Ramesh J, Aburukba R, Sagahyroon A. A remote healthcare monitoring framework for diabetes prediction using machine learning. Healthcare Technol Lett. 2022;8(3):45–57. doi:10.1049/htl2.12010. [Google Scholar] [PubMed] [CrossRef]
18. Junaid SB, Imam AA, Balogun AO, de Silva LC, Surakat YA, Kumar G, et al. Recent advancements in emerging technologies for healthcare management systems: a survey. Healthcare. 2022;10(10):1940. doi:10.3390/healthcare10101940. [Google Scholar] [PubMed] [CrossRef]
19. Miikkulainen R, Liang J, Meyerson E, Rawal A, Fink D, Francon O, et al. Evolving deep neural networks. In: Artificial intelligence in the age of neural networks and brain computing. 2nd ed. London, UK: Academic Press; 2024. p. 269–87. doi:10.1016/b978-0-323-96104-2.00002-6. [Google Scholar] [CrossRef]
20. Rajput MS, Shah J, Patel V, Rajput NS, Kumar D. Deep learning tactics for neuroimaging genomics investigations in alzheimer’s disease. In: Artificial intelligence and machine learning in drug design and development. Hoboken, NJ, USA: John Wiley & Sons; 2024. p. 451–71. doi:10.1002/9781394234196.ch14. [Google Scholar] [CrossRef]
21. Alam A, Muqeem M. An optimal heart disease prediction using chaos game optimization-based recurrent neural model. Int J Inform Technol. 2024;16(5):3359–66. doi:10.1007/s41870-023-01597-w. [Google Scholar] [CrossRef]
22. Thanikachalam V, Kabilan K, Erramchetty SK. Optimized deep CNN for detection and classification of diabetic retinopathy and diabetic macular edema. BMC Med Imaging. 2024;24(1):227. doi:10.1186/s12880-024-01406-1. [Google Scholar] [PubMed] [CrossRef]
23. Ali M, Li J, Nie Z. PPG based digital biomarker for diabetes detection with multiset spatiotemporal feature fusion and XAI. Comput Model Eng Sci. 2025;145(3):4153–77. doi:10.32604/cmes.2025.073048. [Google Scholar] [CrossRef]
24. Alshehri OS, Alshehri OM, Samma H. A comparative study. In: Proceedings of the 2024 IEEE International Conference on Advanced Systems and Emergent Technologies (IC_ASET). Piscataway, NJ, USA: IEEE; 2024. p. 1–5. [Google Scholar]
25. Alhassan Z, McGough AS, Alshammari R, Daghstani T, Budgen D, Al Moubayed N. Type-2 diabetes mellitus diagnosis from time series clinical data using deep learning models. In: Proceedings of the Artificial Neural Networks and Machine Learning–ICANN 2018: 27th International Conference on Artificial Neural Networks; 2018 Oct 4–7; Rhodes, Greece. p. 468–78. [Google Scholar]
26. Kumar S, Bhusan B, Singh D, kumar Choubey D. Classification of diabetes using deep learning. In: Proceedings of the 2020 International Conference on Communication and Signal Processing (ICCSP). Piscataway, NJ, USA: IEEE; 2020. p. 0651–5. [Google Scholar]
27. Jaiswal V, Negi A, Pal T. A review on current advances in machine learning based diabetes prediction. Primary Care Diabetes. 2021;15(3):435–43. doi:10.1016/j.pcd.2021.02.005. [Google Scholar] [PubMed] [CrossRef]
28. Girardi D, Kueng J, Holzinger A. A domain-expert centered process model for knowledge discovery in medical research: putting the expert-in-the-loop. In: Proceedings of the International Conference on Brain Informatics and Health. Cham, Switzerland: Springer; 2015. p. 389–98. [Google Scholar]
29. Karthikeyan R, Geetha P, Ramaraj E. Rule based system for better prediction of diabetes. In: Proceedings of the 2019 3rd International Conference on Computing and Communications Technologies (ICCCT). Piscataway, NJ, USA: IEEE; 2019. p. 195–203. [Google Scholar]
30. Aamir KM, Sarfraz L, Ramzan M, Bilal M, Shafi J, Attique M. A fuzzy rule-based system for classification of diabetes. Sensors. 2021;21(23):8095. doi:10.3390/s21238095. [Google Scholar] [PubMed] [CrossRef]
31. Ye Y, Xiong Y, Zhou Q, Wu J, Li X, Xiao X. Comparison of machine learning methods and conventional logistic regressions for predicting gestational diabetes using routine clinical data: a retrospective cohort study. J Diabetes Res. 2020;2020:4168340. doi:10.1155/2020/4168340. [Google Scholar] [PubMed] [CrossRef]
32. Wilson PW, Meigs JB, Sullivan L, Fox CS, Nathan DM, D’Agostino RB. Prediction of incident diabetes mellitus in middle-aged adults: the Framingham Offspring Study. Arch Internal Med. 2007;167(10):1068–74. doi:10.1001/archinte.167.10.1068. [Google Scholar] [PubMed] [CrossRef]
33. Choubey DK, Kumar M, Shukla V, Tripathi S, Dhandhania VK. Comparative analysis of classification methods with PCA and LDA for diabetes. Current Diab Rev. 2020;16(8):833–50. doi:10.2174/1573399816666200123124008. [Google Scholar] [PubMed] [CrossRef]
34. Xie J, Liu Y, Zeng X, Zhang W, Mei Z. A Bayesian network model for predicting type 2 diabetes risk based on electronic health records. Mod Phys Lett B. 2017;31(19–21):1740055. doi:10.1142/s0217984917400553. [Google Scholar] [CrossRef]
35. Das K, Behera RN. A survey on machine learning: concept, algorithms and applications. Int J Innovat Res Comput Communicat Eng. 2017;5(2):1301–9. doi:10.59256/ijire.2023040224. [Google Scholar] [CrossRef]
36. Yahyaoui A, Jamil A, Rasheed J, Yesiltepe M. A decision support system for diabetes prediction using machine learning and deep learning techniques. In: 2019 1st International Informatics and Software Engineering Conference (UBMYK). Piscataway, NJ, USA: IEEE; 2019. p. 1–4. [Google Scholar]
37. Sarwar MA, Kamal N, Hamid MA, Shah WV. Prediction of diabetes using machine learning algorithms in healthcare. In: Proceedings of the 2018 24th International Conference on Automation and Computing (ICAC). Piscataway, NJ, USA: IEEE; 2018. p. 1–6. [Google Scholar]
38. Zou Q, Qu K, Luo Y, Yin D, Ju Y, Tang H. Predicting diabetes mellitus with machine learning techniques. Front Genet. 2018;9:515. doi:10.3389/fgene.2018.00515. [Google Scholar] [PubMed] [CrossRef]
39. Haq AU, Li JP, Khan J, Memon MH, Nazir S, Ahmad S, et al. Intelligent machine learning approach for effective recognition of diabetes in e-healthcare using clinical data. Sensors. 2020;20(9):2649. doi:10.3390/s20092649. [Google Scholar] [PubMed] [CrossRef]
40. Rubaiat SY, Rahman MM, Hasan MK. Important feature selection & accuracy comparisons of different machine learning models for early diabetes detection. In: Proceedings of the 2018 International Conference on Innovation in Engineering and Technology (ICIET). Piscataway, NJ, USA: IEEE; 2018. p. 1–6. [Google Scholar]
41. Rani DVV, Vasavi D, Kumar K. Significance of multilayer perceptron model for early detection of diabetes over ml methods. J Univ Shanghai Sci Technol. 2021;23(8):148–60. doi:10.51201/jusst/21/08358. [Google Scholar] [CrossRef]
42. Naz H, Ahuja S. Deep learning approach for diabetes prediction using PIMA Indian dataset. J Diab Metab Disord. 2020;19(1):391–403. doi:10.1007/s40200-020-00520-5. [Google Scholar] [PubMed] [CrossRef]
43. Lai H, Huang H, Keshavjee K, Guergachi A, Gao X. Predictive models for diabetes mellitus using machine learning techniques. BMC Endocr Disord. 2019;19(1):1–9. doi:10.1186/s12902-019-0436-6. [Google Scholar] [PubMed] [CrossRef]
44. Birjais R, Mourya AK, Chauhan R, Kaur H. Prediction and diagnosis of future diabetes risk: a machine learning approach. SN Appl Sci. 2019;1(9):1112. doi:10.1007/s42452-019-1117-9. [Google Scholar] [CrossRef]
45. Singh S, Wani NA, Kumar R, Bedi J. DiaXplain: a transparent and interpretable artificial intelligence approach for Type-2 diabetes diagnosis through deep learning. Comput Electr Eng. 2025;126:110470. doi:10.1016/j.compeleceng.2025.110470. [Google Scholar] [CrossRef]
46. Prabhu P, Selvabharathi S. Deep belief neural network model for prediction of diabetes mellitus. In: Proceedings of the 2019 3rd International Conference on Imaging, Signal Processing and Communication (ICISPC). New York, NY, USA: ACM; 2019. p. 138–42. [Google Scholar]
47. García-Ordás MT, Benavides C, Benítez-Andrades JA, Alaiz-Moretón H, García-Rodríguez I. Diabetes detection using deep learning techniques with oversampling and feature augmentation. Comput Methods Programs Biomed. 2021;202(1):105968. doi:10.1016/j.cmpb.2021.105968. [Google Scholar] [PubMed] [CrossRef]
48. Ayon SI, Islam MM. Diabetes prediction: a deep learning approach. Int J Inform Eng Elect Business. 2019;11(2):21–7. doi:10.5815/ijieeb.2019.02.03. [Google Scholar] [CrossRef]
49. Nadesh RK, Arivuselvan K. Type 2: diabetes mellitus prediction using deep neural networks classifier. Int J Cognit Comput Eng. 2020;1:55–61. doi:10.1016/j.ijcce.2020.10.002. [Google Scholar] [CrossRef]
50. Rakshit S, Manna S, Biswas S, Kundu R, Gupta P, Maitra S, et al. Prediction of diabetes type-II using a two-class neural network. In: Proceedings of the Computational Intelligence, Communications, and Business Analytics: First International Conference, CICBA 2017; 2017 Mar 24–25; Kolkata, India. p. 65–71. [Google Scholar]
51. Yang Z, Mitra A, Liu W, Berlowitz D, Yu H. TransformEHR: transformer-based encoder-decoder generative model to enhance prediction of disease outcomes using electronic health records. Nat Commun. 2023;14(1):7857. doi:10.21203/rs.3.rs-2922823/v1. [Google Scholar] [CrossRef]
52. Thangaraju T, Sharma OP. Hybird RNN based feature extraction for early prediction of CVDs using ECG signals for type 2 diabetic patients. J Appl Res Technol. 2023;21(3):424–32. doi:10.22201/icat.24486736e.2023.21.3.1844. [Google Scholar] [CrossRef]
53. Olabanjo OA, Wusu AS, Olabanjo OO, Asokere M, Afisi OT, Akinnuwesi BA. A novel deep learning model for early diabetes risk prediction using attention-enhanced deep belief networks with highly imbalanced data. Int J Inform Technol. 2025;17:1933–55. doi:10.1007/s41870-025-02459-3. [Google Scholar] [CrossRef]
54. Wilson P. Diabetes mellitus and coronary heart disease. Am J Kidney Dis. 1998;32(5 Suppl 3):S89–100. doi:10.1053/ajkd.1998.v32.pm9820468. [Google Scholar] [PubMed] [CrossRef]
55. Abdulhadi N, Al-Mousa A. Diabetes detection using machine learning classification methods. In: Proceedings of the 2021 International Conference on Information Technology (ICIT). Piscataway, NJ, USA: IEEE; 2021. p. 350–4. [Google Scholar]
56. Mercaldo F, Nardone V, Santone A. Diabetes mellitus affected patients classification and diagnosis through machine learning techniques. Procedia Comput Sci. 2017;112(C):2519–28. doi:10.1016/j.procs.2017.08.193. [Google Scholar] [CrossRef]
57. Khaleel FA, Al-Bakry AM. Diagnosis of diabetes using machine learning algorithms. Mat Today Proc. 2023;80(Pt 3):3200–3. doi:10.1016/j.matpr.2021.07.196. [Google Scholar] [CrossRef]
58. Nadeem MW, Goh HG, Ponnusamy V, Andonovic I, Khan MA, Hussain M. A fusion-based machine learning approach for the prediction of the onset of diabetes. Proc Healthcare. 2021;9(10):1393. doi:10.3390/healthcare9101393. [Google Scholar] [PubMed] [CrossRef]
59. Kaur H, Kumari V. Predictive modelling and analytics for diabetes using a machine learning approach. Appl Comput Inform. 2022;18:90–100. doi:10.1016/j.aci.2018.12.004. [Google Scholar] [CrossRef]
60. El_Jerjawi NS, Abu-Naser SS. Diabetes prediction using artificial neural network. Amsterdam, The Netherlands: Elsevier; 2018. [Google Scholar]
61. Ryu KS, Lee SW, Batbaatar E, Lee JW, Choi KS, Cha HS. A deep learning model for estimation of patients with undiagnosed diabetes. Appl Sci. 2020;10(1):421. doi:10.3390/app10010421. [Google Scholar] [CrossRef]
62. Kaggle. Pima Indians Diabetes Database. [cited 2025 Sep 10]. Available from: https://www.kaggle.com/datasets/uciml/pima-indians-diabetes-database. [Google Scholar]
63. Kaggle. U.M.L. Early-Stage Diabetes Risk Prediction. [cited 2025 Sep 10]. Available from: https://www.kaggle.com/datasets/abdelazizsami/early-stage-diabetes-risk-prediction?resource=download. [Google Scholar]
64. online DDA. Kaggle. [cited 2025 Sep 10]. Available from: https://www.kaggle.com/datasets/tigganeha4/diabetes-dataset-2019. [Google Scholar]
65. Centers for Disease Control and Prevention. Behavioral risk factor surveillance system survey data; 2015. Atlanta, GA, USA: U.S. Department of Health and Human Services, Centers for Disease Control and Prevention. [cited 2025 Sep 10]. Available from: https://www.cdc.gov/brfss/annual_data/annual_2015.html. [Google Scholar]
66. Çalisir D, Dogantekin E. An automatic diabetes diagnosis system based on LDA-Wavelet Support Vector Machine Classifier. Expert Syst Appl. 2011;38(7):8311–5. doi:10.1016/j.eswa.2011.01.017. [Google Scholar] [CrossRef]
67. Haritha R, Babu DS, Sammulal P. A hybrid approach for prediction of type-1 and type-2 diabetes using firefly and cuckoo search algorithms. Int J Appl Eng Res. 2018;13(2):896–907. [Google Scholar]
68. Chen W, Chen S, Zhang H, Wu T. A hybrid prediction model for type 2 diabetes using K-means and decision tree. In: Proceedings of the 2017 8th IEEE International Conference on Software Engineering and Service Science (ICSESS). Piscataway, NJ, USA: IEEE; 2017. p. 386–90. [Google Scholar]
69. Yilmaz N, Inan O, Uzer MS. A new data preparation method based on clustering algorithms for diagnosis systems of heart and diabetes diseases. J Med Syst. 2014;38(5):48. doi:10.1007/s10916-014-0048-7. [Google Scholar] [PubMed] [CrossRef]
70. Ribeiro ÁC, Barros AK, Santana E, Príncipe JC. Diabetes classification using a redundancy reduction preprocessor. Res Biomed Eng. 2015;31(2):97–106. doi:10.1590/1517-3151.0608. [Google Scholar] [CrossRef]
71. Zhang Y, Lin Z, Kang Y, Ning R, Meng Y. A feed-forward neural network model for the accurate prediction of diabetes mellitus. Int J Scient Technol Res. 2018;7(8):151–5. [Google Scholar]
72. Dadgar SMH, Kaardaan M. A hybrid method of feature selection and neural network with genetic algorithm to predict diabetes. Int J Mechatron Electr Comput Technol. 2017;7:3397–404. doi:10.1007/s00500-024-10386-x. [Google Scholar] [CrossRef]
73. Abousaber I, Abdalla HF, El-Ghaish H. Robust predictive framework for diabetes classification using optimized machine learning on imbalanced datasets. Front Artif Intell. 2025;7:1499530. doi:10.3389/frai.2024.1499530. [Google Scholar] [PubMed] [CrossRef]
74. Zhang Z, Ahmed KA, Hasan MR, Gedeon T, Hossain MZ. DiabetesNet: a deep learning approach to diabetes diagnosis. arXiv:2403.07483. 2024. [Google Scholar]
Cite This Article
 Copyright © 2026 The Author(s). Published by Tech Science Press.
Copyright © 2026 The Author(s). Published by Tech Science Press.This work is licensed under a Creative Commons Attribution 4.0 International License , which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.


 Submit a Paper
Submit a Paper Propose a Special lssue
Propose a Special lssue View Full Text
View Full Text Download PDF
Download PDF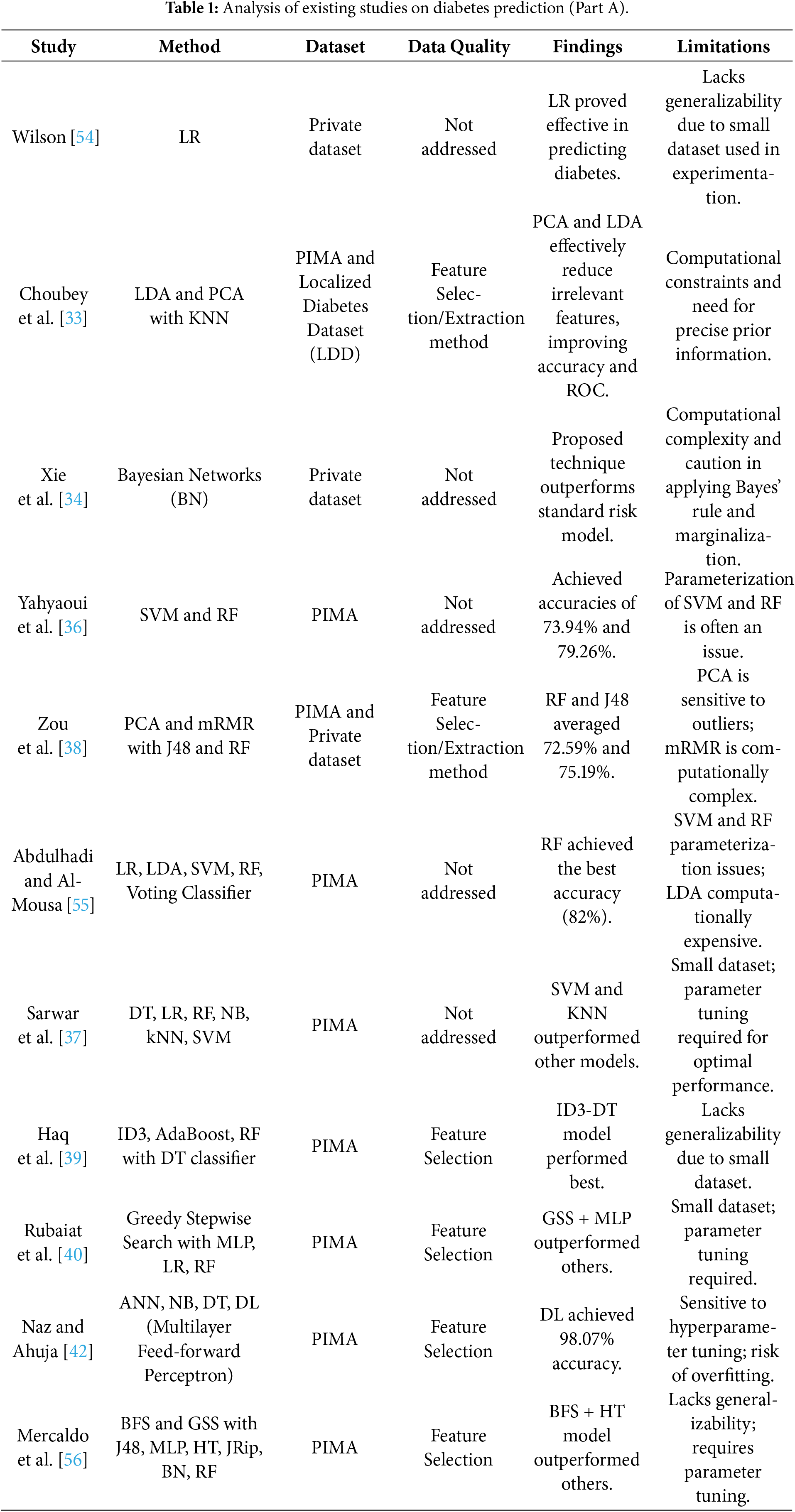
 Downloads
Downloads
 Citation Tools
Citation Tools