 Open Access
Open Access
ARTICLE
Multimodal Graph-Enhanced Vision Transformer for Interpretable Skin Lesion Classification
1 Department of Mathematical Sciences, College of Science, Princess Nourah Bint Abdulrahman University, Riyadh, Saudi Arabia
2 Center of Excellence in Cyber Security (CYBEX), Prince Sultan University, Riyadh, Saudi Arabia
3 Computer Science Department, The National University of Computer and Emerging Sciences (NUCES-FAST), Islamabad, Pakistan
4 Institute of Computer Science, Khwaja Fareed University of Engineering and Information Technology, Abu Dhabi Road, Rahim Yar Khan, Punjab, Pakistan
5 Artificial Intelligence & Data Analytics Lab, College of Computer and Information Sciences, Prince Sultan University, Riyadh, Saudi Arabia
* Corresponding Author: Adil Ali Saleem. Email:
(This article belongs to the Special Issue: Advances in Deep Learning and Computer Vision for Intelligent Systems: Methods, Applications, and Future Directions)
Computer Modeling in Engineering & Sciences 2026, 147(1), 38 https://doi.org/10.32604/cmes.2026.080335
Received 07 February 2026; Accepted 18 March 2026; Issue published 27 April 2026
Abstract
The use of automated skin lesion classification is still a disadvantage, since there is a great visual similarity between benign and malignant lesions. The majority of deep learning methods utilize dermoscopic images only, without taking into account clinical metadata employed by dermatologists on a regular basis. The following paper proposes a vision-graph multimodal framework that links Image encoding to graph neural networks based on metadata representation through the fusion of learnable attention. The framework focuses on three limitations, which are underutilization of clinical context, absence of interpretability, and suboptimal incorporation of modalities. Gradient-weighted Class Activation Mapping++ (Grad-CAM++) is used to obtain dual explainability of visual attention, and SHapley Additive exPlanations (SHAP) to obtain feature importance. Examining the HAM10000 and Derm7pt datasets, statistically significant advances (p < 0.001) of 89.3% and 92.1% accuracy are obtained, which is 4.1% and 2.7% higher than baselines that can only use images. Focusing on weight analysis will provide metadata with 37.7% averaged variance with an error of 8.4%, which confirms the clinical importance of multimodal modeling. The study of ablation shows that graph-based metadata encoding is 1.4% better than standard multilayer perceptron encoding (p = 0.003).Graphic Abstract
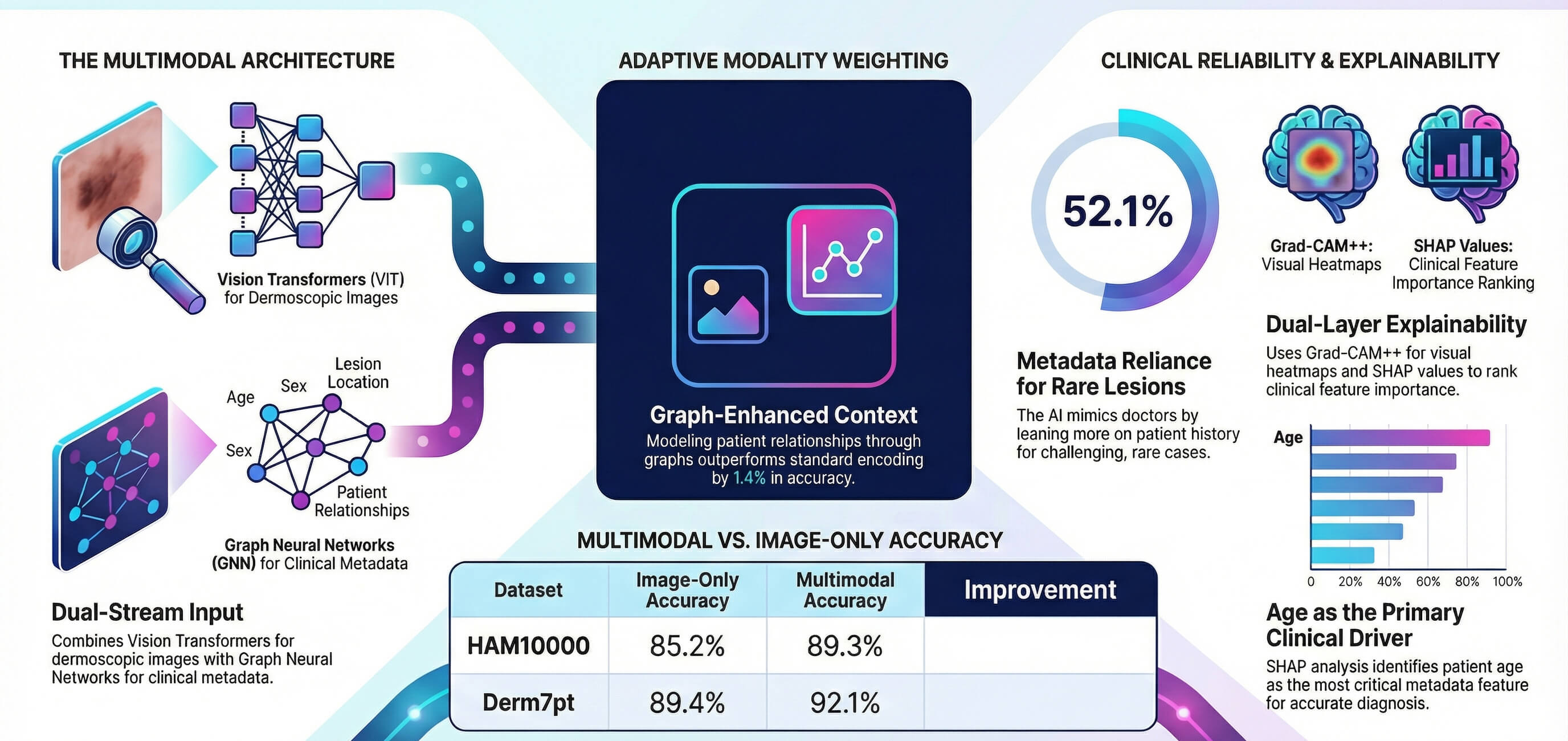
Keywords
Melanoma is the most lethal, and it is the commonest malignancy of the skin in the whole world. Early diagnosis is a useful measure to survive, although it is still challenging because significantly benign and malignant lesions look similar [1]. Dermatologists get 70 to 85 percent diagnosis when detecting melanoma at early stages [2]. The field of deep learning has demonstrated itself as useful in automated classification, but most methods view this as a purely computer vision problem, neglecting very useful clinical information such as patient age, sex, and the location of lesions, information that dermatologists often rely on in a clinical system to make specific diagnoses [3–5].
Vision Transformers (ViTs) have been shown to be more effective than Convolutional Neural Networks (CNNs) in medical imaging [6,7]. Nevertheless, current ViT-based dermatology systems do not incorporate metadata and may therefore lack vital diagnostic associations. Graph Neural Networks (GNNs) offer strong relational modelling capabilities that remain underexplored for encoding clinical metadata [8,9]. Simple fusion strategies such as concatenation yield modest improvements but lack principled modality weighting [10,11].
The clinical deployment of artificial intelligence (AI) diagnostic systems requires not only high accuracy but also interpretability. While gradient-based visualisation methods such as Gradient-weighted Class Activation Mapping (Grad-CAM) [12] provide spatial explanations, they do not address metadata contributions. Bidirectional explainability frameworks are needed to build clinician trust and support clinical adoption. Recent work has demonstrated the value of integrating clinical context into image-based classifiers for skin lesion analysis [13]. This study addresses that gap by proposing a multimodal graph-enhanced Vision Transformer that jointly predicts from dermoscopic images and clinical metadata with explanations spanning both modalities.
This work makes the following contributions:
• A multimodal architecture combining ViT image encoders with GNN metadata encoders, with empirical justification for graph-based encoding over standard multilayer perceptron (MLP) approaches through controlled ablation studies (1.4% improvement, p = 0.003).
• An attention-based fusion mechanism that learns optimal modality weighting adaptively per sample, demonstrating 1.9% improvement over concatenation-based fusion (p < 0.001) with statistical validation of learned weights.
• A dual explainability framework integrating Grad-CAM++ for visual attention with SHAP for metadata feature importance, providing comprehensive interpretability across both modalities.
• Extensive empirical validation on two benchmark datasets with comprehensive ablation studies, including statistical significance testing, confidence intervals, and per-class performance analysis.
The proposed approach bridges the gap between unimodal AI systems and the multimodal reasoning employed by dermatologists, achieving competitive performance while providing clinically meaningful explanations. The remainder of this paper is organized as follows. Section 2 presents the related work relevant to this study. Section 3 describes the proposed methodology in detail. The experimental setup and data is reported in Section 4 along with algorithms in Section 5. Section 6 provides the obtained results and Section 7 discusses those results, and finally, Section 8 concludes the paper.
Recent advances in automated skin lesion analysis have increasingly relied on deep learning with CNN-Transformer hybrids and large dermoscopic corpora. Najjar et al. combined VGG19 local features with a ViT backbone and rotation/shift feature-map augmentation (RSPDA) for robust global context modeling, achieving accuracies of 97.9%, 97.1%, and 98.67% on MSK10000, HAM10000, and PH2, respectively [14]. Gallazzi et al. constructed a large merged dermoscopic dataset to mitigate data scarcity and trained pure Transformer-based Deep Neural Networks (DNNs) end-to-end, reporting 86.37% test accuracy and highlighting the benefit of large-scale pretraining and long-range self-attention for multiclass lesion diagnosis [15].
Amin et al. proposed a two-stage pipeline with BASNet for boundary-aware segmentation and a compact convolutional–Transformer model (CCTM) for classification, using MED-NODE, PH2, ISIC-2019/2020, HAM10000, and DermNet, and reported over 98% classification accuracy by leveraging both local CNN and global Transformer features [16]. Xin et al. introduced SkinTrans, an improved ViT with multi-scale overlapping patch embedding and contrastive learning, reaching 94.3% and 94.1% accuracy on HAM10000 and a clinical dermoscopy dataset by emphasizing multi-scale context and separating similar lesion encodings [17]. Ozdemir and Pacal designed a ConvNeXtV2+separable self-attention hybrid trained on ISIC 2019, obtaining 93.48% accuracy and 91.82% F1-score, and showed that combining CNN fine-grained patterns with efficient attention improves multiclass robustness [18].
Arshed et al. compared off-the-shelf ViT against 11 CNN transfer-learning models on HAM10000, demonstrating that pre-trained ViT achieved 92.14% accuracy and outperformed CNNs under class-imbalance mitigation [19]. Khan et al. integrated deep pre-trained CNN features (ResNet101, DenseNet201), improved moth flame optimization for discriminative feature selection, and Kernel Extreme Learning Machine (KELM) classification, achieving 98.70% segmentation accuracy on PH2 and 90.67% classification accuracy on HAM10000 [20]. Yang et al. boosted ViT and EfficientNet via multi-scale attention maps and ensemble majority voting, reaching 95.05% accuracy on ISIC 2018, demonstrating that attention-guided focus on discriminative lesion regions improves performance [21].
Halawani et al. proposed a hybrid Enhanced ViT+DenseNet169 (EViT-Dens169) with a Spatial Detail Enhancement Block, fusing Transformer global context and CNN edge/texture cues, and obtained 97.1% accuracy and 99.29% specificity on ISIC 2018 across seven classes [22]. Hu et al. designed a multi-scale Transformer with feature fusion and optimized self-attention on ISIC 2017, surpassing ResNet50, VGG19, ResNeXt, and vanilla ViT in accuracy, AUC, and F1-score, while Grad-CAM visualizations indicated improved interpretability of attended lesion regions [23]. Alenezi et al. combined wavelet transforms, ResNet-based deep residual features, and ReLU-based Extreme Learning Machine, achieving 96.91% accuracy on ISIC 2017 and 95.73% on HAM10000, emphasizing frequency-enhanced representations for subtle texture patterns [24].
Multimodal and graph-based fusion has emerged to incorporate clinical context beyond images. Cai et al. introduced a multimodal Transformer with a ViT image encoder, a Soft Label Encoder for metadata, and a Mutual Attention decoder to fuse modalities, improving performance on a private dataset and ISIC 2018 compared to unimodal CNN/ViT baselines [25]. Shivasree and RaviSankar proposed SkinHarmoNet, combining EfficientNet-B4 image features, temporal sensor signals via Bidirectional Long Short-Term Memory (BiLSTM) with attention, and ClinicalBERT text embeddings, then aggregating them using multi-head cross-attention and a Graph Attention Network (GAT) over patient graphs; this multimodal graph-enhanced model achieved 89.6% accuracy and F1 = 0.886 for early skin disease detection [26]. Khurshid et al. developed DualRefNet, a multimodal framework fusing smartphone and dermoscopic images with metadata through dual-stage feature refinement, obtaining balanced accuracies of 0.845 on PAD-UFES20 and 0.815 on ISIC 2019, addressing class imbalance and heterogeneity [27]. Koparde et al. combined conditional generative adversarial networks with YOLOv5 for skin lesion localization and classification, demonstrating the benefit of integrated detection and classification pipelines for clinical deployment [13].
Other works systematically explore deep feature extraction and selection strategies. Benyahia et al. evaluated 17 pre-trained CNNs as feature extractors and 24 classical classifiers on ISIC 2019 and PH2, finding DenseNet201 with fine k-nearest neighbor (KNN) or cubic support vector machine (SVM) yielded 92.34% accuracy on ISIC 2019 and 99% on PH2, underscoring the strength of deep descriptors with non-deep classifiers [28]. Khan et al. fused optimized color features with DCNN-9 deep representations for joint segmentation and classification on ISBI 2016–2018, reaching 96.5% accuracy on ISBI 2017 and showing that handcrafted color and deep features can complement each other [29]. Khan et al. extracted and fused ResNet-50/101 features and applied kurtosis-controlled Principal Component Analysis (PCA) with SVM-Radial Basis Function (SVM-RBF), achieving 95.60% accuracy on ISBI 2017, 90.20% on ISBI 2016, and 89.8% on HAM10000, demonstrating effective dimensionality reduction for multiclass lesions [30]. Srinivasu et al. combined MobileNetV2 and Long Short-Term Memory (LSTM) on HAM10000, using gray-level co-occurrence matrix statistics to track disease progression, and reported over 85% accuracy with reduced computation suitable for mobile deployment [31].
Beyond pure dermoscopy, Xu et al. and Javed et al. introduced advanced segmentation frameworks, where Javed et al. combined region-based active contour with JSEG for improved lesion segmentation and classification, and Xu et al. proposed CCT-Net, a CNN–Transformer dual-branch encoder with attention-based feature fusion, demonstrating the effectiveness of combining local and global contextual features for precise lesion segmentation, achieving Dice scores up to 93.21% and IoU up to 87.7% on ISIC 2016–2018 and PH2 datasets [32,33]. Youssef et al. compared two ConvNeXt-ViT hybrids on HAM10000: a simple ConvNeXt+ViT fusion (94.5% accuracy) and an advanced variant with quantum-inspired feature selection and cross-attention fusion (97.3% accuracy and 0.98 AUC-ROC), illustrating that sophisticated cross-attention fusion can rival or surpass state-of-the-art CNN and ViT baselines [34].
Despite the strong performance of CNN-transformer hybrids, pure vision transformers, and multimodal fusion frameworks, several limitations remain evident in the literature. First, most multimodal approaches rely on simple concatenation or shallow attention mechanisms that fail to model population-level relationships among patients, thereby underutilizing clinical metadata. Second, many Transformer-based models emphasize visual features while implicitly assuming uniform relevance of metadata across samples, leading to modality dominance and reduced robustness in ambiguous or visually similar lesions. Third, interpretability is often limited to image-level attention, with minimal insight into the contribution of non-imaging clinical factors. Finally, class imbalance and dataset heterogeneity continue to challenge generalization across cohorts. These gaps motivate the proposed Multimodal Graph-Enhanced Vision Transformer, which explicitly models metadata dependencies through graph-based message passing, employs adaptive attention-based modality fusion to balance visual and clinical cues, and integrates dual-level explainability via Grad-CAM++ and SHAP to enhance clinical transparency while improving robustness under imbalanced settings.
The proposed architecture consists of three components: a Vision Transformer encoder
where

Figure 1: Multimodal architecture overview showing vision transformer encoder for images, graph neural network encoder for metadata, attention-based fusion module, and classification head.
The model is expressed as:
3.1 Vision Transformer Encoder
The image encoder employs Vision Transformer Base/16 pre-trained on ImageNet-21k. Input images are resized to
where LN denotes layer normalization and MHSA denotes multi-head self-attention with 12 heads. The Feed-Forward Network (FFN) in Eq. (4) applies two linear transformations with a Rectified Linear Unit (ReLU) activation. The final image representation is extracted from the class token:
3.2 Graph Neural Network Metadata Encoder
A graph convolutional network encodes metadata to capture population-level patterns through message passing. For a batch of B samples with metadata vectors
where
Each metadata vector is projected to hidden dimension 256, then two graph convolutional network layers perform message passing:
The final metadata representation is
Several architectural design choices require clarification. First, the batch-level fully connected graph is constructed to achieve a clinically meaningful objective: allowing each sample to interact with patients sharing similar demographic characteristics during training. The Gaussian similarity kernel encourages grouping according to age, sex, and anatomical location, while the learnable temperature parameter
Second, the metadata representation consists of heterogeneous feature types, including continuous variables (age), binary variables (sex), and one-hot encoded categorical variables (anatomical location, comprising up to 15 categories). The Euclidean distance defined in Eq. (5) treats all feature dimensions uniformly. In practice, this limitation is mitigated by normalizing age to the range
Third, during inference, the GNN encoder applies its learned parameters to a locally constructed graph formed from the available test-batch samples using the same procedure employed during training. In the limiting case of single-sample inference, no neighbouring nodes are available and the GNN reduces to its learned linear transformations without message passing. The relational inductive bias is therefore encoded within the learned GNN parameters rather than a fixed global graph structure, enabling generalization to previously unseen samples. Stable training convergence, illustrated in Section 6, together with consistent performance across five cross-validation folds with a standard deviation of
A learnable attention mechanism adaptively weights modalities. Given image features
The fused representation is:
This allows the model to emphasize image features for visually distinctive lesions while relying more on metadata for ambiguous cases.
The model is trained end-to-end using cross-entropy loss with class balancing:
where
Grad-CAM++ computes class activation maps by backpropagating gradients from the predicted class through the final Vision Transformer block. For target class
which is upsampled to original image resolution.
SHAP computes Shapley values for each metadata feature, quantifying each feature’s contribution to the prediction while accounting for interactions. SHAP values are computed for 1000 randomly sampled test cases and aggregated for correctly classified vs misclassified samples. Given that the metadata comprises at most 17 features (one continuous, one binary, and 15 one-hot anatomical location categories), exact Shapley value computation is tractable: for |F| features, evaluation requires
HAM10000 [35] contains 10,015 dermoscopic images across 7 diagnostic categories (melanocytic nevi, melanoma, benign keratosis, basal cell carcinoma, actinic keratosis, vascular lesions, dermatofibroma). Images were acquired using various dermatoscopic devices (Dermatoscope DermLite, 3Gen, HEINE) at multiple institutions. Metadata includes patient age (range: 5–85 years, mean: 52.3), sex (male: 56%, female: 44%), and anatomical location (encoded as 15 discrete categories). Class distribution is imbalanced: nevi 67.0%, melanoma 11.1%, benign keratosis 10.9%, basal cell carcinoma 5.1%, actinic keratosis 3.2%, vascular 1.4%, dermatofibroma 1.1%.
Derm7pt [36] comprises 2000 images with rich clinical annotations, including the 7-point checklist criteria: pigment network (typical, atypical, absent), dots/globules (regular, irregular, absent), streaks (present, absent), regression structures (present, absent), blue-whitish veil (present, absent), vascular structures (comma, hairpin, absent), and pigmentation (regular, irregular). Additional metadata includes patient demographics and anatomical location. Class distribution: melanoma 31%, melanocytic nevi 43%, basal cell carcinoma 14%, seborrheic keratosis 9%, miscellaneous 3%.
Images undergo preprocessing: hair removal using morphological operations, contrast-limited adaptive histogram equalization for illumination normalization, resize to
All experiments use PyTorch 2.0.1 with CUDA 11.7 on NVIDIA A100 graphics processing units (GPUs). Batch size is 32 for training and 64 for inference. The Vision Transformer encoder uses pre-trained weights from ImageNet-21k. Graph Neural Network layers use PyTorch Geometric 2.3.0.
Stratified 5-fold cross-validation is employed with 80% training, 10% validation, and 10% test per fold. Final results report mean and standard deviation across all 5 folds. Primary metrics include accuracy, macro F1 (unweighted mean of per-class F1), weighted F1, and macro Area Under the Curve (AUC). Statistical testing uses paired
Baselines include ResNet-50, DenseNet-121, Vision Transformer Base (image only), Vision Transformer with concatenated metadata (Vision Transformer+Concat), Vision Transformer with added metadata (Vision Transformer+Add), and Vision Transformer with multilayer perceptron-encoded metadata (Vision Transformer+MLP). All baselines use identical training procedures and hyperparameters. Key hyperparameters including learning rates, GNN hidden dimension (256), dropout rate (0.3), and batch size (32) were selected via grid search on the validation split of fold 1 and applied uniformly across all remaining folds, following standard practice. Inverse class frequency weighting was adopted as a principled and hyperparameter-free strategy for class imbalance; focal loss and re-sampling strategies are identified as directions for future investigation.
Algorithm 1 outlines the complete training procedure. The algorithm employs end-to-end optimization with layer-wise learning rate decay for the Vision Transformer encoder, where deeper layers receive smaller learning rates to preserve pretrained knowledge. Class imbalance is addressed through inverse frequency weighting in the cross-entropy loss. Early stopping with patience of 10 epochs prevents overfitting by monitoring validation macro F1-score. The training procedure integrates all three architectural components: Vision Transformer image encoding, Graph Neural Network metadata encoding with batch-level graph construction, and attention-based fusion for adaptive modality weighting.

Algorithm 2 describes the inference procedure with dual explainability. The algorithm first performs forward propagation through all model components to generate predictions. Visual explanations are then computed via Grad-CAM++, which uses second-order gradients to generate pixel-importance maps highlighting diagnostically relevant image regions. Simultaneously, SHAP values quantify the contribution of each metadata feature by computing Shapley values through systematic feature subset evaluation. This dual explainability framework provides comprehensive interpretability spanning both visual and clinical modalities, essential for building clinician trust and enabling clinical deployment. During inference, the GNN encoder

Algorithm 3 details the batch-level graph construction process. For each training batch, a fully-connected graph is constructed where nodes represent individual samples and edges encode metadata similarity. Edge weights are computed using a Gaussian kernel with learnable temperature parameter

Table 1 presents the main results. The proposed method achieves 89.3%

Table 2 presents per-class performance on HAM10000. The proposed method shows consistent improvements across all classes, with larger gains for minority classes. The largest F1 improvements occur for vascular lesions (+0.07) and dermatofibroma (+0.06), the two rarest classes, suggesting metadata provides crucial discriminative information when visual features alone are insufficient.

Table 3 presents comprehensive ablation studies. Image features alone achieve 85.2% accuracy, while metadata alone achieves only 71.4%, confirming visual information is primary but metadata provides substantial complementary value. Learned attention fusion (89.3%) outperforms fixed strategies: concatenation (87.4%), addition (86.8%). Graph Convolutional Network encoding (89.3%) outperforms 2-layer multilayer perceptron (87.9%) by 1.4% (p = 0.003), validating the hypothesis that graph-based encoding captures beneficial population patterns.

Table 4 reports computational complexity for all model variants. The proposed method introduces a modest overhead of approximately 0.8 million additional parameters and 1.3 ms per sample in inference latency relative to the ViT-Base backbone, corresponding to the 15% increase noted in the Discussion. GPU memory consumption increases by approximately 0.4 GB. These figures confirm that the proposed multimodal framework imposes a manageable computational cost relative to the performance gains achieved.

Fig. 2 presents confusion matrices for both datasets. The HAM10000 confusion matrix reveals the model correctly classifies the majority class (nevi) with 92% accuracy while maintaining reasonable performance on minority classes. Melanoma, the most critical class clinically, achieves 85% recall. Common misclassifications occur between visually similar categories such as benign keratosis and melanoma. The Derm7pt confusion matrix shows more balanced performance across classes due to the more uniform class distribution, with overall diagonal dominance indicating strong classification performance.

Figure 2: Confusion matrices on HAM10000 and Derm7pt datasets. Percentages represent the proportion of true labels predicted as each class. Strong diagonal values indicate high classification accuracy, with common misclassifications occurring between visually similar lesion types.
Table 5 shows per-class attention distribution. On average, image features receive 62.3% attention weight while metadata receives 37.7%. Rare and challenging classes show higher metadata attention, with vascular lesions receiving 52.1% metadata attention compared to 31.8% for nevi. Correlation analysis reveals that metadata attention negatively correlates with class frequency (Pearson r = −0.89, p = 0.007), demonstrating that the model adaptively relies more on metadata for rare and challenging cases. This correlation is computed across seven class-level observations and should be interpreted as an indicative pattern of adaptive fusion behavior rather than a definitive statistical relationship given the limited number of class-level data points.

Fig. 3 visualizes the learned attention weights. Panel (a) shows average modality attention weights with error bars representing standard deviation. Panel (b) presents per-class attention distribution, revealing that rare classes rely more heavily on metadata. Panel (c) demonstrates the correlation between metadata attention and per-class accuracy, showing that classes with higher metadata attention tend to have lower baseline accuracy, confirming adaptive fusion behavior.

Figure 3: Attention weight analysis showing (a) average modality attention weights, (b) per-class attention distribution, and (c) correlation between metadata importance and classification accuracy. The model adaptively increases metadata reliance for challenging classes.
Fig. 4 presents SHAP values for metadata features. Age is the most important feature (mean absolute SHAP value 0.42), followed by sex (0.18) and anatomical location features (0.15–0.01). Comparing correctly classified vs misclassified cases, misclassified cases show higher variance in SHAP values (standard deviation 0.31 vs. 0.24, p = 0.008), which may reflect uncertainty in multimodal integration, noise in the metadata signal, or model instability for ambiguous cases; a causal interpretation of this variance difference requires further controlled investigation.

Figure 4: SHAP-based metadata feature importance analysis for (a) correctly classified cases and (b) misclassified cases. Age emerges as the most important clinical feature, while misclassified cases show higher variance in feature importance.
Fig. 5 shows training dynamics on HAM10000. Panel (a) displays training and validation loss convergence, with validation loss stabilizing around epoch 35. Panel (b) shows training and validation accuracy curves, with the model achieving 89.3% validation accuracy. Early stopping was triggered at epoch 40 when validation F1-score stopped improving.

Figure 5: Training dynamics on HAM10000 dataset showing (a) training and validation loss curves and (b) training and validation accuracy curves over 50 epochs. Early stopping at epoch 40 prevents overfitting.
Fig. 6 presents Grad-CAM++ visualizations for selected cases. The model correctly focuses on lesion boundaries and internal structures for accurate predictions. For the misclassified melanoma case, the attention diffuses across the image rather than concentrating on diagnostic features, suggesting visual ambiguity. The visualizations demonstrate that the model learns clinically relevant attention patterns.

Figure 6: Grad-CAM++ visual explanations for selected cases showing original images.
Three principal findings emerge from this study. First, clinical metadata contributes meaningfully to diagnostic accuracy. The 4.1% accuracy improvement over the image-only ViT baseline confirms the value of incorporating clinical context, a result consistent with established dermatological practice in which patient history and lesion site inform diagnosis for ambiguous presentations. Attention weight analysis shows that metadata accounts for 37.7% of predictions on average, with substantially higher contribution for rare and diagnostically challenging classes. Second, graph-based metadata encoding outperforms standard isolated encoding. The Graph Convolutional Network (GCN) delivers a 1.4% accuracy gain over an equivalent parameterized MLP (p = 0.003), confirming that modelling inter-patient relationships through message passing captures population-level patterns that isolated feature extraction cannot. The learned adjacency weights indicate that the model groups patients by age and anatomical region, reflecting the clinically meaningful subpopulation structure used by dermatologists.
Third, adaptive fusion is more effective than fixed fusion techniques. Attention-based fusion outperforms concatenation by 1.9% (p < 0.001). In per-sample analysis, visually distinct lesions receive greater attention from the image modality (72.4% vs. 62.3% on average), whereas visually ambiguous lesions rely more heavily on metadata. This adaptive behaviour mirrors the reasoning process of dermatologists. Most misclassifications occur between visually similar lesion categories, as shown by the confusion matrix analysis. Melanoma is occasionally misclassified as benign keratosis, which represents a clinically significant error warranting further investigation. The model achieves 85% melanoma recall; while this represents a meaningful improvement over the ViT baseline (83%), it falls short of the thresholds typically required for primary screening deployment (>90% sensitivity). This indicates the model is better positioned as a clinical decision-support tool rather than a standalone screening system, and future work should target melanoma recall specifically through class-weighted hard example mining or class-specific attention mechanisms.
Receiver Operating Characteristic (ROC) curve analysis confirms good discriminative ability across all classes, with AUC values consistently above 0.82. The high AUC for melanoma (0.89) supports the model’s clinical utility for case prioritisation, while the lower AUC for vascular lesions (0.82) reflects the inherent challenge of rare-class diagnosis. The negative correlation between class frequency and metadata attention (
To assess sensitivity to the Gaussian kernel temperature parameter
There are several limitations that should be acknowledged. The method requires complete metadata, which may not always be available in real-world clinical settings. Preliminary tests using mean imputation show that accuracy declines by 2.1% at a 20% missing data rate; a systematic investigation across multiple imputation strategies (including multivariate imputation by chained equations and
Dataset limitations also exist. HAM10000 metadata contains only three features, potentially underestimating multimodal gains achievable with richer clinical annotations. Both datasets exhibit class imbalance, necessitating careful selection of validation metrics. Geographic and demographic biases are present, with HAM10000 predominantly containing samples from European populations. Performance on darker skin types therefore, requires targeted validation.
Clinical validation also remains preliminary. Prospective evaluation in real clinical workflows is essential. Future work should explore richer metadata integration, including family history, genetic markers, and previous diagnoses. Large vision-language models offering zero-shot recognition capabilities represent a promising direction for reducing annotation dependence in clinical deployment. Comparison with more expressive GNN architectures such as Graph Attention Networks and GraphSAGE is a natural extension; the primary objective of the current ablation is to establish the benefit of graph-based relational encoding over non-relational MLP encoding, which is demonstrated by the 1.4% gain.
Federated learning with differential privacy could enable multi-institutional training while preserving patient privacy. Model compression techniques could enable mobile deployment. Active learning strategies could reduce annotation burden. Prospective clinical trials with dermatologists are essential to validate real-world utility.
This study presents a multimodal architecture for skin lesion classification that combines dermoscopic images with clinical metadata using a Vision Transformer (ViT), Graph Neural Networks (GNNs), and attention-based fusion. The model achieves accuracy rates of 89.3% on HAM10000 and 92.1% on Derm7pt, with statistically significant improvements relative to image-only baselines. Ablation studies confirm the value of each architectural component: graph-based metadata encoding outperforms MLP encoding by 1.4%, and attention-based fusion outperforms concatenation by 1.9%. Attention weight analysis reveals that metadata contributes to 37.7% of predictions on average, with greater weight for challenging and rare classes, mirroring the clinical practice of dermatologists who give more weight to patient history when faced with ambiguous lesions. The dual explainability framework combining Grad-CAM++ for image attention and SHAP for metadata feature importance provides comprehensive interpretability across both modalities. Per-class results show steady improvements across all diagnostic categories, with particularly pronounced gains for the minority classes. Receiver Operating Characteristic (ROC) curve analysis demonstrates strong discriminative capacity with AUC values exceeding 0.82 across all classes. Confusion matrix analysis reveals that errors predominantly occur between visually similar lesion categories, identifying clear targets for future improvement. The validity of findings is supported by statistical significance testing and confidence intervals across five-fold cross-validation. The proposed framework bridges the gap between unimodal AI systems and the multimodal clinical reasoning of dermatologists, achieving competitive performance with clinically meaningful interpretability. Several limitations remain: the model depends on complete metadata at inference time, has been validated on European-predominant cohorts, and has not been evaluated prospectively in clinical workflow settings; addressing these constraints through richer metadata integration, demographically diverse datasets, and controlled clinical trials represents the primary agenda for future work.
Acknowledgement: This research is supported by Princess Nourah bint Abdulrahman University Researchers Supporting Project, Princess Nourah bint Abdulrahman University, Riyadh, Saudi Arabia. The authors are also thankful to the Artificial Intelligence and Data Analytics (AIDA) Laboratory, College of Computer and Information Sciences, Prince Sultan University, Riyadh, Saudi Arabia, for article processing charge support.
Funding Statement: This research is funded by Princess Nourah bint Abdulrahman University Researchers Supporting Project number (PNURSP2026R346), Princess Nourah bint Abdulrahman University, Riyadh, Saudi Arabia.
Author Contributions: Faten S. Alamri contributed to conceptualization, methodology, validation, investigation, formal analysis and writing of the original draft. Afia Zafar contributed to methodology, software development, formal analysis, validation, data curation, visualization, and writing—review and editing. Noor Ayesha contributed to visualization, investigation, methodology, data curation, formal analysis, and validation. Adil Ali Saleem contributed to methodology, software development, formal analysis, investigation, data curation, and writing of the original draft. Amjad R. Khan contributed to project administration, supervision, visualization, investigation, validation, and writing—review and editing. All authors reviewed and approved the final version of the manuscript.
Availability of Data and Materials: The datasets used in this study HAM10000: Publicly available at https://dataverse.harvard.edu/dataset.xhtml?persistentId=doi:10.7910/DVN/DBW86T. Derm7pt: Available upon request from authors [36].
Ethics Approval: Not applicable.
Conflicts of Interest: The authors declare no conflicts of interest.
References
1. Esteva A, Kuprel B, Novoa RA, Ko J, Swetter SM, Blau HM, et al. Dermatologist-level classification of skin cancer with deep neural networks. Nature. 2017;542(7639):115–8. doi:10.1038/nature21056. [Google Scholar] [PubMed] [CrossRef]
2. Haenssle HA, Fink C, Schneiderbauer R, Toberer F, Buhl T, Blum A, et al. Man against machine: diagnostic performance of a deep learning convolutional neural network for dermoscopic melanoma recognition in comparison to 58 dermatologists. Ann Oncol. 2018;29(8):1836–42. doi:10.1093/annonc/mdy166. [Google Scholar] [PubMed] [CrossRef]
3. Garib G, Mery D, Navarrete-Dechent C. Evaluation of the importance of metadata in skin lesion classification. Signal Image Video Process. 2025;19(11):887. doi:10.1007/s11760-025-04498-6. [Google Scholar] [CrossRef]
4. Melanoma Research Alliance. MRA-MIDAS: multimodal image dataset for AI-based skin cancer. Palo Alto, CA, USA: Center for Artificial Intelligence in Medicine & Imaging; 2023. doi:10.71718/15nz-jv40. [Google Scholar] [CrossRef]
5. Nasir S, Bilal M, Khalidi H. Detection and classification of skin cancer by using CNN-enabled cloud storage data access control algorithm based on blockchain technology. Int J Theor Appl Comput Intell. 2025:146–59. doi:10.65278/ijtaci.2025.31. [Google Scholar] [CrossRef]
6. Dosovitskiy A, Beyer L, Kolesnikov A, Weissenborn D, Zhai X, Unterthiner T, et al. An image is worth 16x16 words: transformers for image recognition at scale. arXiv:2010.11929v2. 2021. doi:10.48550/arXiv.2010.11929. [Google Scholar] [CrossRef]
7. Shi S, Liu W. B2-ViT net: broad vision transformer network with broad attention for seizure prediction. IEEE Trans Neural Syst Rehabil Eng. 2024;32:178–88. doi:10.1109/tnsre.2023.3346955. [Google Scholar] [PubMed] [CrossRef]
8. Kipf TN, Welling M. Semi-supervised classification with graph convolutional networks. arXiv:1609.02907. 2017. doi:10.48550/arXiv.1609.02907. [Google Scholar] [CrossRef]
9. Gan X, Li T, Gong C, Li D, Dong D, Liu J, et al. GraphCSR: a degree-equalized CSR format for large-scale graph processing. Proc VLDB Endow. 2025;18(11):4255–68. doi:10.14778/3749646.3749691. [Google Scholar] [CrossRef]
10. Adamu S, Alhussian H, Aziz N, Abdulkadir SJ, Alwadin A, Abubakar Imam A, et al. The future of skin cancer diagnosis: a comprehensive systematic literature review of machine learning and deep learning models. Cogent Eng. 2024;11(1):2395425. doi:10.1080/23311916.2024.2395425. [Google Scholar] [CrossRef]
11. Jaber NJF, Akbas A. Melanoma skin cancer detection based on deep learning methods and binary Harris Hawk optimization. Multimed Tools Appl. 2025;84(22):25709–22. doi:10.1007/s11042-024-19864-8. [Google Scholar] [CrossRef]
12. Selvaraju RR, Cogswell M, Das A, Vedantam R, Parikh D, Batra D. Grad-CAM: visual explanations from deep networks via gradient-based localization. In: Proceedings of the 2017 IEEE International Conference on Computer Vision (ICCV); 2017 Oct 22–29; Venice, Italy, p. 618–26. doi:10.1109/iccv.2017.74. [Google Scholar] [CrossRef]
13. Koparde S, Kotwal J, Deshmukh S, Adsure S, Chaudhari P, Kimbahune V. A conditional generative adversarial networks and Yolov5 Darknet-based skin lesion localization and classification using independent component analysis model. Inform Med Unlocked. 2024;47(7):101515. doi:10.1016/j.imu.2024.101515. [Google Scholar] [CrossRef]
14. Najjar FH, Khudhair ZN, Mohamed F, Rahim MSM, Chan VS, Ali AH. Transformer-aided skin cancer classification using VGG19-based feature encoding. Sci Rep. 2025;15(1):40204. doi:10.1038/s41598-025-24081-w. [Google Scholar] [PubMed] [CrossRef]
15. Gallazzi M, Biavaschi S, Bulgheroni A, Gatti TM, Corchs S, Gallo I. A large dataset to enhance skin cancer classification with transformer-based deep neural networks. IEEE Access. 2024;12:109544–59. doi:10.1109/access.2024.3439365. [Google Scholar] [CrossRef]
16. Amin J, Azhar M, Arshad H, Zafar A, Kim SH. Skin-lesion segmentation using boundary-aware segmentation network and classification based on a mixture of convolutional and transformer neural networks. Front Med. 2025;12:1524146. doi:10.3389/fmed.2025.1524146. [Google Scholar] [PubMed] [CrossRef]
17. Xin C, Liu Z, Zhao K, Miao L, Ma Y, Zhu X, et al. An improved transformer network for skin cancer classification. Comput Biol Med. 2022;149:105939. doi:10.1016/j.compbiomed.2022.105939. [Google Scholar] [PubMed] [CrossRef]
18. Ozdemir B, Pacal I. A robust deep learning framework for multiclass skin cancer classification. Sci Rep. 2025;15(1):4938. doi:10.1038/s41598-025-89230-7. [Google Scholar] [PubMed] [CrossRef]
19. Arshed MA, Mumtaz S, Ibrahim M, Ahmed S, Tahir M, Shafi M. Multi-class skin cancer classification using vision transformer networks and convolutional neural network-based pre-trained models. Information. 2023;14(7):415. doi:10.3390/info14070415. [Google Scholar] [CrossRef]
20. Khan MA, Sharif M, Akram T, Damaševičius R, Maskeliūnas R. Skin lesion segmentation and multiclass classification using deep learning features and improved moth flame optimization. Diagnostics. 2021;11(5):811. doi:10.3390/diagnostics11050811. [Google Scholar] [PubMed] [CrossRef]
21. Yang G, Luo S, Greer P. Boosting skin cancer classification: a multi-scale attention and ensemble approach with vision transformers. Sensors. 2025;25(8):2479. doi:10.3390/s25082479. [Google Scholar] [PubMed] [CrossRef]
22. Halawani HT, Senan EM, Asiri Y, Abunadi I, Mashraqi AM, Alshari EA. Enhanced early skin cancer detection through fusion of vision transformer and CNN features using hybrid attention of EViT-Dens169. Sci Rep. 2025;15(1):34776. doi:10.1038/s41598-025-18570-1. [Google Scholar] [PubMed] [CrossRef]
23. Hu J, Xiang Y, Lin Y, Du J, Zhang H, Liu H. Multi-scale transformer architecture for accurate medical image classification. In: Proceedings of the 2025 International Conference on Artificial Intelligence and Computational Intelligence; 2025 Feb 14–16; Kuala Lumpur Malaysia. doi:10.1145/3730436.3730505. [Google Scholar] [CrossRef]
24. Alenezi F, Armghan A, Polat K. Wavelet transform based deep residual neural network and ReLU based Extreme Learning Machine for skin lesion classification. Expert Syst Appl. 2023;213(8):119064. doi:10.1016/j.eswa.2022.119064. [Google Scholar] [CrossRef]
25. Cai G, Zhu Y, Wu Y, Jiang X, Ye J, Yang D. A multimodal transformer to fuse images and metadata for skin disease classification. Vis Comput. 2023;39(7):2781–93. doi:10.1007/s00371-022-02492-4. [Google Scholar] [PubMed] [CrossRef]
26. Shivasree Y, RaviSankar V. Design of an iterative hybrid multimodal deep learning method for early skin disease detection with cross-attention and graph-based fusions. MethodsX. 2025;15(1):103584. doi:10.1016/j.mex.2025.103584. [Google Scholar] [PubMed] [CrossRef]
27. Khurshid M, Singh R, Vatsa M. Multimodal dual-stage feature refinement for robust skin lesion classification. Sci Rep. 2025;15(1):37775. doi:10.1038/s41598-025-14839-7. [Google Scholar] [PubMed] [CrossRef]
28. Benyahia S, Meftah B, Lézoray O. Multi-features extraction based on deep learning for skin lesion classification. Tissue Cell. 2022;74(22):101701. doi:10.1016/j.tice.2021.101701. [Google Scholar] [PubMed] [CrossRef]
29. Khan MA, Sharif MI, Raza M, Anjum A, Saba T, Ali Shad S. Skin lesion segmentation and classification: a unified framework of deep neural network features fusion and selection. Expert Syst. 2019;39(7):e12497. doi:10.1111/exsy.12497. [Google Scholar] [CrossRef]
30. Khan MA, Javed MY, Sharif M, Saba T, Rehman A. Multi-model deep neural network based features extraction and optimal selection approach for skin lesion classification. In: Proceedings of the 2019 International Conference on Computer and Information Sciences (ICCIS); 2019 Apr 3–4; Sakaka, Saudi Arabia. doi:10.1109/iccisci.2019.8716400. [Google Scholar] [CrossRef]
31. Srinivasu PN, SivaSai JG, Ijaz MF, Bhoi AK, Kim W, Kang JJ. Classification of skin disease using deep learning neural networks with MobileNet V2 and LSTM. Sensors. 2021;21(8):2852. doi:10.3390/s21082852. [Google Scholar] [PubMed] [CrossRef]
32. Javed R, Rahim MSM, Saba T, Rashid M. Region-based active contour JSEG fusion technique for skin lesion segmentation from dermoscopic images. Biomed Res. 2019;30(6):1–10. [Google Scholar]
33. Xu Z, Guo X, Wang J. Enhancing skin lesion segmentation with a fusion of convolutional neural networks and transformer models. Heliyon. 2024;10(10):e31395. doi:10.1016/j.heliyon.2024.e31395. [Google Scholar] [PubMed] [CrossRef]
34. Youssef AA, Badr E, Veraldo N, Hamza D. A comprehensive analysis of hybrid ConvNeXt and vision transformer architectures for skin cancer classification. Int J Res Appl Sci Eng Technol. 2025;13(5):1808–15. doi:10.22214/ijraset.2025.70514. [Google Scholar] [CrossRef]
35. Tschandl P, Rosendahl C, Kittler H. The HAM10000 dataset, a large collection of multi-source dermatoscopic images of common pigmented skin lesions. Sci Data. 2018;5(1):180161. doi:10.1038/sdata.2018.161. [Google Scholar] [PubMed] [CrossRef]
36. Kawahara J, Daneshvar S, Argenziano G, Hamarneh G. Seven-point checklist and skin lesion classification using multitask multimodal neural nets. IEEE J Biomed Health Inform. 2019;23(2):538–46. doi:10.1109/jbhi.2018.2824327. [Google Scholar] [PubMed] [CrossRef]
Cite This Article
 Copyright © 2026 The Author(s). Published by Tech Science Press.
Copyright © 2026 The Author(s). Published by Tech Science Press.This work is licensed under a Creative Commons Attribution 4.0 International License , which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.


 Submit a Paper
Submit a Paper Propose a Special lssue
Propose a Special lssue View Full Text
View Full Text Download PDF
Download PDF Downloads
Downloads
 Citation Tools
Citation Tools