 Open Access
Open Access
ARTICLE
In vitro study of emodin-induced nephrotoxicity in human renal glomerular endothelial cells on a microfluidic chip
National Institute for Nutrition and Health, Chinese Center for Disease Control and Prevention, Beijing, 100050, China
* Corresponding Author: Jiyong Yin,
# These authors contributed equally to this work
(This article belongs to the Special Issue: Cellular Biomechanics in Health and Diseases)
BIOCELL 2023, 47(1), 125-131. https://doi.org/10.32604/biocell.2023.022937
Received 01 April 2022; Accepted 21 June 2022; Issue published 26 September 2022
Abstract
Emodin is an effective component of rhubarb with positive pharmacological effects on human health. However, it is also toxic to different cells or tissues to varying degrees. The effects of emodin on glomerular endothelial cells (GECs) remain to be tested, and the documented works were always performed in vitro and hardly reflect the real physiological situation. To study the effects of emodin on GECs in a biomimetic environment, we utilized a microfluidic chip to assess the physiological reaction of human renal glomerular endothelial cells to various concentrations of emodin in this work. The results showed that emodin caused cytotoxicity, impaired glomerular filtration barrier integrity to macromolecules, and increased barrier permeability in a dose-dependent manner. With the increase in emodin concentration, the concentration of the pro-inflammatory cytokine tumor necrosis factor-α, interleukin (IL)-6, transforming growth factor-β1, and monocyte chemoattractant protein (MCP-1) increased while the production of inflammatory cytokine IL-6 first increased and then decreased with the increase in emodin concentration. Our findings shed new light on emodin-induced nephrotoxicity and provide insights for the application of microfluidic chip devices to reveal drug-cell interactions.Keywords
Emodin is the main active ingredient of total rhubarb anthraquinones (TRAs), which are extracted from the roots of the plant Rheum palmatum (Wang et al., 2008). Years of research have shown that emodin has many medically valuable properties, including anti-inflammatory, antibacterial, antiviral, antitumor, and immunosuppressive properties (Shrimali et al., 2013; Cui et al., 2020; Han et al., 2015). However, its toxicity, including hepatotoxicity, nephrotoxicity, genotoxicity, reproductive toxicity and phototoxicity, have also been confirmed by studies both in vivo and in vitro (Dong et al., 2016; Li et al., 2020; He et al., 2012). TRAs have been shown to cause nephrotoxicity in Sprague Dawley rats, resulting in swelling and degeneration of tubular epithelial cells (Yan et al., 2006). (Wang et al., 2007, 2015) found that emodin induces apoptosis through cathepsin B and caspase 3 activation in HK-2 cells. However, the exact mechanisms of the emodin cytotoxicity are unknown, which limits the effective prevention of its nephrotoxicity.
The blood filtration process by renal endothelium is critical for the maintenance of bodily homeostasis (Molema and Aird, 2012). Most studies on emodin nephrotoxicity relied on animal models, in which differences from humans cannot be eliminated. Moreover, animal models are not conducive to high-throughput quantitative studies. By contrast, the simple and high-throughput assays of renal cell cultures have gained popularity. However, only a few models have investigated key representative features of the glomerular barrier, which blocks the passage of large protein molecules, such as albumin, between the circulatory system and urinary system, enabling the passage of water and small metabolites. The glomerular barrier is mainly composed of podocytes, fused basement membranes, and glomerular endothelial cells (GECs).
The design of complex and controllable micro-devices relying on the recent advances in engineering technology mimics the environment close to in vivo physiological conditions (Chang et al., 2016; Maschmeyer et al., 2015; Li et al., 2016). Many studies have used living cells to model kidney tissue, such as renal tubules (Jang and Suh, 2010; Weber et al., 2016; Batchelder et al., 2015). In vitro simulations have helped investigators develop micro-scale kidney models that sufficiently enhance cell polarization to rearrange cell junctions. However, all of these studies focused on renal tubular tissues. Here, we constructed a microfluidic device lined with human GECs on simulated glomeruli to study emodin exposure-induced nephrotoxicity. Cell viability, expression of tight junction proteins (TJPs), lactate dehydrogenase (LDH) leakage, and permeability changes were examined in GECs to explore the toxic effects of different concentrations of emodin. This renal model has the potential to serve as an effective platform for other drug toxicology studies, helping provide evaluation and testing parameters closer to the human physiological environment than animal models.
Human renal glomerular endothelial cells (HRGECs) were obtained from BeNa Culture Collection (Beijing, China). The medium contained 1% penicillin/streptomycin, 1% endothelial cell growth supplement, and 5% FBS. Cells were cultured in a humidified atmosphere containing 5% CO2 at 37°C. HRGECs were treated with emodin (TCI, Shanghai, China) from a 10 mM stock solution in dimethyl sulfoxide.
Culturing human renal glomerular endothelial cells on a chip
HRGECs were observed by fluorescence microscopy (EVOS M7000). The perfusion of media with cells and compounds was controlled by Microfluidics Platform and CellASIC ONIX M04S Microfluidics Switching Plates. The HRGECs were centrifuged and re-suspended to a density of 1.0–5.0 × 106 cells/mL. These cells were first loaded into the imaging chamber and incubated overnight to allow binding of the cells, after which perfused the treatment compounds into the chamber for 48 h (3 psi).
Assessment of cellular toxicity
The emodin concentration gradient was set to 0–200 μM. After the cells were treated for 48 h, the concentration of LDH released from the cells was monitored using an LDH assay Kit (Nanjing Jiancheng Bioengineering Institute, Nanjing, China). Cell death was assessed by live/dead cell staining Kit (Calcein AM/PI, KeyGEN). The live cells were green colored, and the dead cells were red colored under the fluorescence microscope. Quantitative cell viability data were obtained by the CCK-8 assay (Dojindo), as described by the manufacturer. Briefly, CCK-8 reagent was added to the cell culture (1:10), and measure the absorbance at 450 nm after 4 h of incubation.
Real-time polymerase chain reaction (RT-qPCR)
Total RNA was extracted using RNAprep pure Micro Kit (TIANGEN), and then reverse transcription, and real-time fluorescence quantitative PCR was conducted through the LightCycler 480 system (Roche) according to the protocol of FastKing One Step RT-qPCR Kit (TIANGEN). The program for reverse transcription was one cycle of 50°C for 30 min and denaturation at 95°C for 3 min, then 40 cycles of 95°C for 15 s and 60°C for 30 s. The primers for amplification of ZO-1 and GAPDH are listed in Table 1. Average Ct values were calculated to the Ct values for GAPDH, and the normalized values were subjected to formula 2−ΔΔCt to estimate the fold change between the experimental groups and control. All reactions were performed in triplicates.

Cells were washed with PBS, fixed with 4% paraformaldehyde for 20 min, permeabilized with 0.2% Triton-X100, and then incubated with the rabbit primary monoclonal antibody for ZO-1 (1:50; Cell Signaling Technology) at 37°C for 2 h. An Alexa 488 conjugated goat secondary antibody (1:500; Cell Signaling Technology) was then added and incubated at room temperature for 1 h. The nuclei were counterstained with 4′,6-diamidino-2-phenylindole (DAPI) for 5 min after washing the samples with PBS. Finally, the stained cells were photographed using a fluorescence microscope.
Enzyme-linked immunosorbent assay (ELISA)
According to the user’s guidelines, the levels of tumor necrosis factor (TNF)-α, interleukin (IL)-6, transforming growth factor (TGF)-β1, and monocyte chemoattractant protein (MCP)-1 in the cell culture fluid of HRGECs were measured using ELISA Kits (Beyotime). Measure A450 of the samples for three times.
Permeability testing of the glomerular endothelial cells layer
HRGECs were seeded in culture plate inserts of a transwell chamber (0.4 μm polycarbonate membranes) at densities of 1 × 105/mL, and 0.5 and 1.5 mL serum-containing medium was added to the inside and outside chambers, respectively. HRGEC monolayers were cultured to confluence for 2 days to assess the permeability of the GEC layer. After treatment with the indicated concentrations of emodin (0, 25, 50, 100, and 200 μM) for 48 h, the chambers were washed with PBS, and 250 μM of 150 kDa FITC-IgG (Solarbio, Beijing, China) in 1 mL PBS was added to the top chamber and 1.5 mL culture medium to the bottom chamber. Filtrates were removed from the bottom chamber and replaced with 1.5 mL fresh culture medium after 15, 30, 45, and 60 min for analysis in a fluorometer (Molecular Devices) with excitation at 485 nm and emission at 510 nm. The relative permeability of IgG at each time was calculated by dividing the fluorescence intensity through the cell barrier by the total fluorescence intensity. The data represent the means of three replication.
Statistical analysis was conducted using GraphPad Prism 8.0. The data are presented as means ± SD. One-way analysis of variance (ANOVA) was used to assess the differences between groups, and a P-value < 0.05 indicated a significant difference.
Establishment of the monolayer human renal glomerular endothelial cells model on a chip
The CellASIC® ONIX M04S-03 microfluidic plate is a 4-chamber cell culture plate that enables continuous and controlled solution switching when used in combination with the corresponding microfluidic system to simulate the dynamic micro-environment of the kidney. This biomimetic plate provides a controllable and dynamic microenvironment for cell culture (Fig. 1A). The suspensions comprising 1–5 × 106 cells/mL were perfused into the cell chamber pre-coated with the extracellular matrix containing type I collagen and cultured for 72 h. A confluent cell monolayer was formed that mimicked the glomerular endothelial barrier by continuous perfusion of the flow culture medium at the indicated flow rate (0.5–1 psi), providing adequate nutrition and minimum pressure similar to those experienced by human glomerular lining cells in vivo. After 72 h of culture, immunofluorescence assay of HRGECs revealed a well-defined confluent endothelial monolayers with the tight junction protein ZO-1 (TJP1) having a continuous linear distribution (Fig. 1B). The HRGECs area ratio was about 98.3%.

Figure 1: Plate configuration and characterization of HRGECs on a chip. (A) The M04S microfluidic plate has four independent culture units, each with one gravity flow inlet (1), four solution inlets (2–5), one cell inlet (6), and two outlets (7 and 8). The image on the right is an enlarged image of that on the left. There are six inlets connecting to one culturing chamber, including one gravity flow inlet, four solution inlets, and one cell inlet. Four independent dosing tests were carried out simultaneously. (B) Immunofluorescence staining of the ZO-1 (green) 3 days after the formation of a confluent cell monolayer.
Analysis of emodin-induced cytotoxicity in human renal glomerular endothelial cells on-chip
The infiltration of HRGECs by exogenous toxicants can lead to nephrotoxicity and endothelial barrier dysfunction. To evaluate the effects of emodin on HRGECs, the evaluation indexes, including ZO-1 expression, LDH leakage and cell viability, were analyzed after exposing the cells to emodin at different concentrations (50 to 200 μM) for 48 h. In the preliminary experiment, we set a wider range of experimental concentrations. The experimental results showed that when the drug concentration was lower than 50 μM, emodin had no significant toxic effect on HRGECs, which were almost dead when emodin concentration was higher than 200 μM. Emodin showed a good dose-response relationship at concentrations only in the range of 50–200 μM. Therefore, the experimental concentration range was set at 50–200 μM (Data omitted). Cell viability was determined by dead/live staining and the CCK-8 assay. According to Fig. 2A, as the emodin concentration increased, the number of live cells decreased, accompanied by a concomitant gradual increase in the number of dead cells. Similarly, after treatment with 50 to 200 μM emodin, cell viability declined in a dose-dependent manner (Fig. 2B). When the concentration was 50 μM, cell viability decreased slightly; at 100 and 200 μM, cell viability showed a significant decline with time. Cell viability decreased to 29.3% following exposure to 200 μM emodin. The leakage of LDH was examined, which is a recognized marker of cellular membrane injury. Under the same conditions, emodin exposure increased LDH leakage, indicating the aggravation of cell damage and a dose-dependent effect of emodin toxicity (Fig. 2C).

Figure 2: Emodin toxicity measured on a chip. (A) Viability of HRGECs after exposure to emodin (0, 50, 100, and 200 μM). The dead cells were stained red, while the live cells were stained green. (B) CCK-8 assay was conducted to determine HRGECs viability at different points. (C) LDH leakage assay in HRGECs for different durations. Data are presented as means ± SD. n = 3, *P < 0.05.
The expression of ZO-1 contributes to the maintenance of the glomerular endothelial barrier function. To evaluate the effect of emodin on TJP ZO-1 expression, RT-qPCR was performed. As shown in Fig. 3A, the transcriptional level of the ZO-1 gene decreased with the increase in emodin concentration. When the effect of emodin on the expression of ZO-1 in endothelial cells was analyzed by immunofluorescence staining, the expression of ZO-1 in HRGECs was found to decrease significantly compared to untreated control, and the connections between cells were relaxed and destroyed following emodin exposure (Fig. 3B), in a concentration-dependent manner. These results suggest that emodin may impair the integrity of the glomerular barrier by reducing the expression of ZO-1.

Figure 3: Expression of ZO-1 in human renal glomerular endothelial cells after exposure to emodin. (A) Real-time quantitative polymerase chain reaction analysis. (B) Immunofluorescence staining analysis. DAPI-stained cells (blue), ZO-1 (green).
Detection of pro-inflammatory cytokines
The cell culture supernatant was collected, and a commercial ELISA Kit was used to detect the expression of IL-6 as a pro-inflammatory cytokine that reflects the degree of cell damage caused by emodin. The results showed that low doses of emodin could induce cytokine IL-6, while high doses of emodin had a strong cytotoxic effect, killing cells without inducing their activation, so the release of cytokine IL-6 was significantly reduced (Fig. 4A). These results indicated that emodin has a concentration threshold for the induction of pro-inflammatory cytokines.

Figure 4: The effect of emodin on the levels of interleukin (IL)-6 (A), tumor necrosis factor (TNF)-α (B), transforming growth factor (TGF)-β1 (C), and monocyte chemoattractant protein (MCP)-1 (D). Data are presented as means ± SD. n = 3, *P < 0.05.
The effect of emodin on the pro-inflammatory cytokine TNF-α was also assessed using a commercial ELISA Kit. The concentration of TNF-α increased with the increase in emodin concentration (Figs. 4B–4D). This result again indicated that high doses of emodin are toxic to cells, causing the release of TNF-α. Interestingly, the trend of the release of TNF-α was different from that of IL-6, which may be due to the different regulatory networks controlling the cytokines.
The level of other pro-inflammatory cytokine, MCP-1, and TGF-β1, regulating inflammatory processes was also determined. These two factors showed the same trend as TNF-α with the change in emodin concentrations (Figs. 4C and 4D). These results again confirmed the cytotoxic effect of emodin.
Effect of emodin on barrier permeability of human renal glomerular endothelial cells
We hypothesized that the shrinkage of HRGECs resulted in the passage of macromolecules from the paracellular space through the cell layer. Thus, we performed an assay using fluorescent labeled IgG as an indicator in a transwell chamber to assess the permeability of the glomerular endothelial barrier to macromolecules. The measured fluorescence intensities through the HRGEC barrier are shown in Table 2. The permeability for IgG increased with the increase in emodin concentration, suggesting that the HRGEC layer was damaged after exposure to emodin in a dose-dependent manner (Fig. 5).
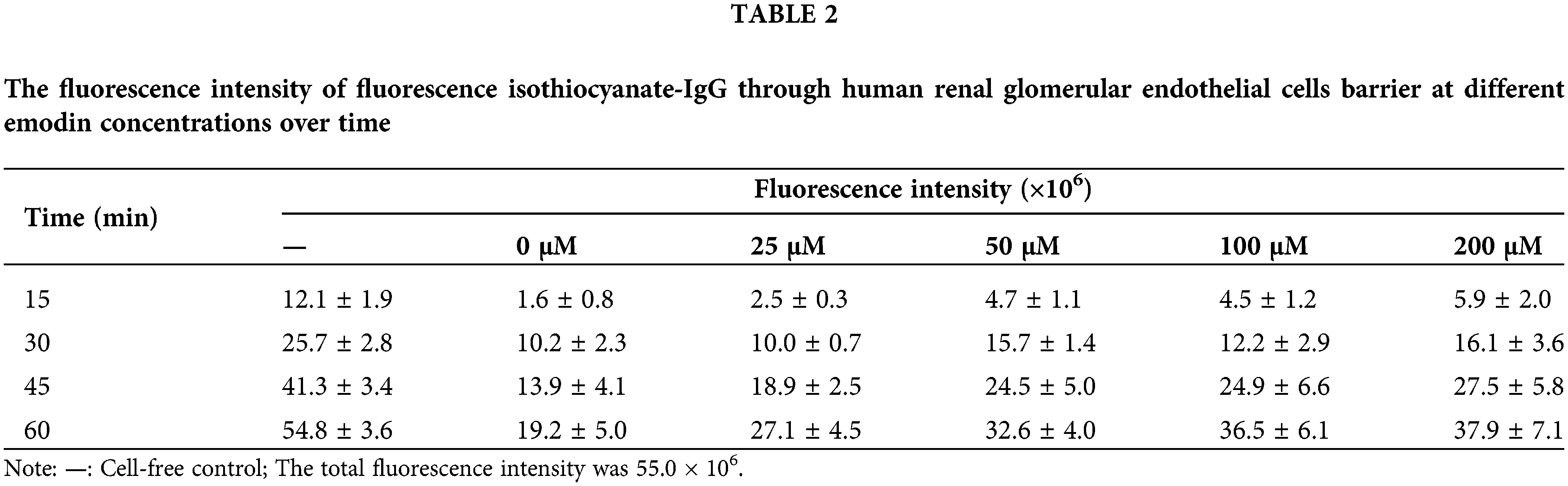

Figure 5: Effect of emodin on the permeability of the human renal glomerular endothelial cells (HRGEC) barrier. HRGEC barrier permeability to IgG over 60 min. —: Cell-free control.
Emodin exhibits positive pharmacological effects on human health but also shows toxicity to multiple tissues or organs (Dong et al., 2016). Nephrotoxicity is one of the most severe negative effects of emodin. Many studies have determined different mechanisms of emodin-induced nephrotoxicity. Nesslany et al. conducted a mouse in vivo comet assay and reported that aloe-emodin induces primary DNA damage in the kidneys and liver (Nesslany et al., 2009). (Wang et al., 2015) demonstrated that emodin caused decreased cell viability, cleavage and activation of caspase 3, and loss of mitochondrial membrane potential in an HK-2 cell assay. Recently, (Cao et al., 2019) reported that emodin damages the cell morphology and cytoskeleton to varying degrees in HK-2 cells. Nevertheless, some studies showed abrogation of nephrotoxicity or amelioration of renal tubular cell apoptosis by emodin in rats and in vitro cultured cells, respectively (Ali et al., 2013; Liu et al., 2016). These results remain contrary to the above-described negative effects. The problem here is the experimental system employed in most studies, i.e., animal and human cell models. Although animal models can mimic the real internal environment, they are not conducive to high throughput screening, which can be achieved using human cell models. However, cell models cannot closely mimic the renal filtration process.
The development of microfluidic devices brings hope for solutions to the contradiction between animal and cell models. When passing through the kidney, drugs are concentrated in the renal tubules. (Li et al., 2017) modeled cadmium-induced nephrotoxicity using a kidney-on-a-chip device. The device enables direct visualization and quantitative analysis of cadmium-induced nephrotoxicity in real time. In this study, we established an emodin-induced nephrotoxicity model by growing HRGECs monolayers on a chip. In this model, cell viability decreased as the concentration of emodin increased, which was also evidenced by LDH leakage. Moreover, the relative expression of the cell junction protein ZO-1 decreased with emodin exposure in a dose-dependent manner. However, there were differences in the expression trends of inflammatory cytokines. At a low dose of emodin, inflammatory cytokine IL-6 production increased while it decreased at higher doses. By contrast, TNF-α, TGF-β1, and MCP-1expression increased at all emodin doses. It seems that emodin has a concentration threshold for the induction of IL-6 but not for TNF-α, TGF-β1 and MCP-1. These results suggest that emodin may exert its cytotoxic effects via a complicated mechanism. Further studies are needed to explain it well. Overall, this study further confirms the nephrotoxicity of emodin and reveals the real-time reaction of cells to emodin to some extent.
In addition to the animal models, organ-on-a-chip devices have significantly contributed to the study of various drug-organ interactions, greatly improving the drug screening and discovery processes (Zheng et al., 2016; Zhang et al., 2018). The organ-on-a-chip device can overcome the tissue barrier, which is a significant conventional problem in vitro systems. Like emodin, the effective components of traditional Chinese medicine may have some negative effects on human cells. In vitro cell assays may be useful to demonstrate some patterns, but they cannot mimic the complex environment of real tissues or organs and therefore, can only provide a qualitative analysis. With organ-on-a-chip devices, we are more likely to obtain reliable quantitative results, which will be important for a detailed understanding of the effects of traditional Chinese medicinal materials. Our work provides insights into the use of microfluidic chips for evaluating drug toxicity and cell activity in the real-time. However, the human body is a complex, dynamically regulated system, and engineered devices cannot completely mimic the environment inside the body. Therefore, we must be careful in drawing conclusions and continue improving the process. Bionics and 3D printing technology might better mimic the natural conditions of human organs, which will play an important role in the near future.
Availability of Data and Materials: All data generated or analyzed during this study are included in this published article (and its supplementary information files).
Author Contribution: ZY designed the study. ZY, DC, WQ, JW, and LW performed the experiments. ZY analyzed and interpreted data. ZY wrote, reviewed, and revised the manuscript. JH, QZ, and JY provided administrative, technical, and material support.
Ethics Approval: Not applicable.
Funding Statement: This work was supported by the National Key R&D Program of China (Project No. 2018YFC1602103), Ministry of Science and Technology of China.
Conflicts of Interest: The authors declare that they have no conflicts of interest to report regarding the present study.
References
Ali BH, Al-Salam S, Al Husseini IS, Al-Lawati I, Waly M, Yasin J, Fahim M, Nemmar A (2013). Abrogation of cisplatin-induced nephrotoxicity by emodin in rats. Fundamental & Clinical Pharmacology 27: 192–200. DOI 10.1111/j.1472-8206.2011.01003.x. [Google Scholar] [CrossRef]
Batchelder CA, Martinez ML, Tarantal AF (2015). Natural scaffolds for renal differentiation of human embryonic stem cells for kidney tissue engineering. PLoS One 10: e0143849. DOI 10.1371/journal.pone.0143849. [Google Scholar] [CrossRef]
Cao C, Hui L, Li C, Yang Y, Zhang J, Liu T, Hao R, Zhang Y (2019). In vitro study of the nephrotoxicity of total Dahuang (Radix Et Rhizoma Rhei Palmati) anthraquinones and emodin in monolayer human proximal tubular epithelial cells cultured in a transwell chamber. Journal of Traditional Chinese Medicine 39: 609–623. DOI 10.19852/j.cnki.jtcm.2019.05.002. [Google Scholar] [CrossRef]
Chang SY, Weber EJ, Ness KV, Eaton DL, Kelly EJ (2016). Liver and kidney on chips: Microphysiological models to understand transporter function. Clinical Pharmacology & Therapeutics 100: 464–478. DOI 10.1002/cpt.436. [Google Scholar] [CrossRef]
Cui Y, Chen LJ, Huang T, Ying JQ, Li J (2020). The pharmacology, toxicology and therapeutic potential of anthraquinone derivative emodin. Chinese Journal of Natural Medicines 18: 425–435. DOI 10.1016/s1875-5364(20)30050-9. [Google Scholar] [CrossRef]
Dong X, Fu J, Yin X, Cao S, Li X, Lin L, Huyiligeqi, Ni J (2016). Emodin: A review of its pharmacology, toxicity and pharmacokinetics. Phytotherapy Research 30: 1207–1218. DOI 10.1002/ptr.5631. [Google Scholar] [CrossRef]
Han JW, Shim DW, Shin WY, Heo KH, Kwak SB, Sim EJ, Jeong JH, Kang TB, Lee KH (2015). Anti-inflammatory effect of emodin via attenuation of NLRP3 inflammasome activation. International Journal of Molecular Sciences 16: 8102–8109. DOI 10.3390/ijms16048102. [Google Scholar] [CrossRef]
He Q, Liu K, Wang S, Hou H, Yuan Y, Wang X (2012). Toxicity induced by emodin on zebrafish embryos. Drug and Chemical Toxicology 35: 149–154. DOI 10.3109/01480545.2011.589447. [Google Scholar] [CrossRef]
Jang KJ, Suh KY (2010). A multi-layer microfluidic device for efficient culture and analysis of renal tubular cells. Lab on a Chip 10: 36–42. DOI 10.1039/B907515A. [Google Scholar] [CrossRef]
Li Q, Gao J, Pang X, Chen A, Wang Y (2020). Molecular mechanisms of action of emodin: As an anti-cardiovascular disease drug. Frontiers in Pharmacology 11: 559607. DOI 10.3389/fphar.2020.559607. [Google Scholar] [CrossRef]
Li Z, Guo Y, Yu Y, Xu C, Xu H, Qin J (2016). Assessment of metabolism-dependent drug efficacy and toxicity on a multilayer organs-on-a-chip. Integrative Biology 8: 1022–1029. DOI 10.1039/C6IB00162A. [Google Scholar] [CrossRef]
Li Z, Jiang L, Tao T, Su W, Guo Y, Yu H, Qin J (2017). Assessment of cadmium-induced nephrotoxicity using a kidney-on-a-chip device. Toxicology Research 6: 372–380. DOI 10.1039/C6TX00417B. [Google Scholar] [CrossRef]
Liu H, Gu LB, Tu Y, Hu H, Huang YR, Sun W (2016). Emodin ameliorates cisplatin-induced apoptosis of rat renal tubular cells in vitro by activating autophagy. Acta Pharmacologica Sinica 37: 235–245. DOI 10.1038/aps.2015.114. [Google Scholar] [CrossRef]
Maschmeyer I, Lorenz AK, Schimek K, Hasenberg T, Ramme AP et al. (2015). A four-organ-chip for interconnected long-term co-culture of human intestine, liver, skin and kidney equivalents. Lab on a Chip 15: 2688–2699. DOI 10.1039/c5lc00392j. [Google Scholar] [CrossRef]
Molema G, Aird WC (2012). Vascular heterogeneity in the kidney. Seminars in Nephrology 32: 145–155. DOI 10.1016/j.semnephrol.2012.02.001. [Google Scholar] [CrossRef]
Nesslany F, Simar-Meintières S, Ficheux H, Marzin D (2009). Aloe-emodin-induced DNA fragmentation in the mouse in vivo comet assay. Mutation Research/Genetic Toxicology and Environmental Mutagenesis 678: 13–19. DOI 10.1016/j.mrgentox.2009.06.004. [Google Scholar] [CrossRef]
Shrimali D, Shanmugam MK, Kumar AP, Zhang J, Tan BK, Ahn KS, Sethi G (2013). Targeted abrogation of diverse signal transduction cascades by emodin for the treatment of inflammatory disorders and cancer. Cancer Letters 341: 139–149. DOI 10.1016/j.canlet.2013.08.023. [Google Scholar] [CrossRef]
Wang C, Jiang Z, Yao L, Wu X, Sun L et al. (2008). Participation of cathepsin B in emodin-induced apoptosis in HK-2 cells. Toxicology Letters 181: 196–204. DOI 10.1016/j.toxlet.2008.05.013. [Google Scholar] [CrossRef]
Wang C, Wu X, Chen M, Duan W, Sun L, Yan M, Zhang L (2007). Emodin induces apoptosis through caspase 3-dependent pathway in HK-2 cells. Toxicology 231: 120–128. DOI 10.1016/j.tox.2006.11.064. [Google Scholar] [CrossRef]
Wang C, Wu X, Chen M, Duan W, Sun L, Yan M, Zhang L (2015). Involvement of PPARγ in emodin-induced HK-2 cell apoptosis. Toxicology in Vitro 29: 228–233. DOI 10.1016/j.tiv.2014.10.021. [Google Scholar] [CrossRef]
Weber EJ, Chapron A, Chapron BD, Voellinger JL, Lidberg KA et al. (2016). Development of a microphysiological model of human kidney proximal tubule function. Kidney International 90: 627–637. DOI 10.1016/j.kint.2016.06.011. [Google Scholar] [CrossRef]
Yan M, Zhang LY, Sun LX, Jiang ZZ, Xiao XH (2006). Nephrotoxicity study of total rhubarb anthraquinones on Sprague Dawley rats using DNA microarrays. Journal of Ethnopharmacology 107: 308–311. DOI 10.1016/j.jep.2006.03.031. [Google Scholar] [CrossRef]
Zhang B, Korolj A, Lai BFL, Radisic M (2018). Advances in organ-on-a-chip engineering. Nature Reviews Materials 3: 257–278. DOI 10.1038/s41578-018-0034-7. [Google Scholar] [CrossRef]
Zheng F, Fu F, Cheng Y, Wang C, Zhao Y, Gu Z (2016). Organ-on-a-chip systems: Microengineering to biomimic living systems. Small 12: 2253–2282. DOI 10.1002/smll.201503208. [Google Scholar] [CrossRef]
Cite This Article
 Copyright © 2023 The Author(s). Published by Tech Science Press.
Copyright © 2023 The Author(s). Published by Tech Science Press.This work is licensed under a Creative Commons Attribution 4.0 International License , which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.


 Submit a Paper
Submit a Paper Propose a Special lssue
Propose a Special lssue View Full Text
View Full Text Download PDF
Download PDF Downloads
Downloads
 Citation Tools
Citation Tools