 Open Access
Open Access
ARTICLE
Smad8 is involvement in follicular development via the regulation of granulosa cell growth and steroidogenesis in mice
1 College of Biotechnology and Pharmaceutical Engineering, West Anhui University, Lu’an, 237012, China
2 Lu’an Academy of Agricultural Sciences, Lu’an, 237001, China
3 College of Life Sciences, Anhui Normal University, Wuhu, 241000, China
4 College of Animal Science and Technology, Nanjing Agricultural University, Nanjing, 210095, China
* Corresponding Authors: SHAN LU. Email: ; YAFEI CAI. Email:
(This article belongs to the Special Issue: Cellular Signal Transduction in Biological Activities)
BIOCELL 2024, 48(1), 139-147. https://doi.org/10.32604/biocell.2023.045884
Received 11 September 2023; Accepted 23 November 2023; Issue published 30 January 2024
Abstract
Background: SMAD family proteins (SMADs) are crucial transcription factors downstream of transforming growth factor beta (TGF-ß)/SMAD signaling pathways that have been reported to play a pivotal role in mammalian reproduction. However, the role of SMAD family member 8 (SMAD8, also known as SMAD9), a member of the SMAD family, in mammalian reproduction remains unclear. Methods: We employed RNA interference techniques to knock down Smad8 expression in mouse granulosa cells (GCs) to investigate the effects of Smad8 on GC growth and steroidogenesis. Results: Our findings revealed a significant decrease in the proliferative capacity and a substantial increase in the apoptosis rate of GCs after transfection with Smad8-siRNA for 48 h. Subsequent hormone assays demonstrated a significant decrease in estradiol (E2) levels, whereas progesterone (P4) remained unchanged. Further mechanistic analysis showed that the mRNA expression of proliferating cell nuclear antigen (Pcna), Cyclin D2, cell cycle-dependent kinase 4 (Cdk4), B-cell lymphoma-2 (Bcl-2), estrogen receptor (Er), luteinizing hormone receptor (Lhr) and cytochrome P450 family 19 subfamily A member 1 (Cyp19a1) significantly decreased. Conversely, the mRNA of cysteine aspartate proteinase 3 (Caspase 3) significantly increased, wheras Bcl2-associated X (Bax), follicle-stimulating hormone receptor (Fshr) and cytochrome P450 family 11 subfamily A member 1 (Cyp11a1) remained unchanged compared to the controls. Conclusion: This study indicates that Smad8 knockdown inhibits cell proliferation, promotes apoptosis, reduces Er and Lhr transcription, and decreases E2 production in mouse GCs. These findings suggest that Smad8 may serve as a novel genetic marker for mammalian reproduction.Keywords
Enhancing the fertility of female mammals holds great significance in both production and research domains. Numerous studies have revealed the pivotal role of ovarian follicle growth and development in determining the number of offspring a female mammal can produce. However, the process of follicular development is intricately regulated by multiple factors, and the final regulatory outcome may lead to follicular ovulation or atresia. An integral mechanism within follicular development and ovulation is the interaction between GCs and oocytes. GCs can provide essential nutrients and stimulatory factors required for oocyte development. Consequently, the development and normal function of GCs play a key role in oocyte maturation and ovulation.
Bone morphogenetic proteins (BMPs) are integral members of TGF-ß superfamily. Prior research has demonstrated the involvement of BMPs in ovarian and follicle development through the regulation of GC growth and steroidogenesis [1,2]. For example, BMP-2 is involved in follicular atresia [3], whereas BMP-4 and BMP-7 participate in the primordial-primary follicle transition [1,4], the BMP-6 participated plays a role in the progesterone synthesis [5] and the growth differen-tiation factor 9 (GDF-9), highly homologous with the BMP-15 [6], inhibits Fshr expression, affecting further follicular development [7], among other functions.
SMAD proteins are crucial transcription factors in the TGF-ß/SMAD signaling pathways. These proteins are categorized into three types by function: receptor-regulated SMAD1/2/3/5/8 (R-SMADs), common-SMAD4 (Co-SMAD), and inhibitory SMAD6/7 (I-SMADs), all of which mediate intracellular TGF-ß/SMAD signal transduction. Studies have shown that the knockout of Smad genes can affect mammalian reproductive ability. For instance, knockout of Smad1 or Smad5 leads to a marked reduction in primordial germ cells [8,9], whereas Smad3 knockout results in defects in follicular growth, atresia, and differentiation [10]. In the case of Smad4 knockout mice, it leads to a reduction in the number of ovulations and defects in GC development [11]. Therefore, BMP/SMAD signaling pathways play a pivotal role in the ovary and follicle development in mammals.
SMAD8 is the most recently identified member of the SMAD protein family, and there has been limited research on it to date. SMAD8 possesses a distinctive functional domain that sets it apart from other R-SMADs [12]. Recent research has indicated that Smad8 exerts a dominant negative role in regulating gene transcription in cultured C2C12 cells [13]. Subsequent studies have also implicated Smad8 in the initiation of follicles in geese [14]; however, the exact mechanism underlying its regulation of follicular development remains unknown. This knowledge gap encourages further exploration into the potential mechanisms by which Smad8 affects follicular development. To address this, in the present study, we initially localized the expression of SMAD8 protein in mouse follicles. The siRNA interference sequence of the Smad8 gene was designed and synthesized and transfected into mouse GCs in vitro. The efficiency of Smad8 silencing was assessed using quantitative polymerase chain reaction (qPCR) and western blot analysis. Furthermore, we evaluated and analyzed the proliferation and steroid synthesis capacity of GCs post-silencing. Concurrently, we observed and analyzed the expression of reproductive hormone receptor genes, cell proliferation, and apoptosis genes, aiming to explore the internal mechanisms through which the Smad8 gene impacts follicle development.
Animals, ovarian tissue harvest, GC isolation, and culture
All animal experiments conducted in this research received approval from the Welfare and Ethics Committee of West Anhui University. In the experiment, 10 healthy eight-week-old female Kunming mice (Suzhou Xishan Experimental Animal Co., Ltd., Suzhou, China) were housed in groups of five per cage with free access to food and water. To induce ovarian growth and follicular development, the mice received an intraperitoneal injection of 10 IU of pregnant mare serum gonadotropin (PMSG; Ningbo Sansheng Biological Technology, Ningbo, China) per mouse. After 48 h of PMSG treatment, the mice were euthanized, and their ovaries were collected.
The ovaries were placed in a 35-mm dish filled with pre-warmed phosphate-buffered saline (PBS), and mouse GCs were carefully punctured and released from follicles larger than 200 µm in diameter using a syringe needle under a dissecting microscope. Mouse GCs were cultured following the method introduced by Liu et al. [15]. GCs were plated in Dulbecco’s modified Eagle medium/F12 medium (Invitrogen, Carlsbad, USA) with 10% (v/v) fetal bovine serum (Gibco, NY, USA) and 100 IU/mL penicillin/streptomycin. The culture plates were maintained in a 5% (v/v) CO2 atmosphere at 37°C, and cells exhibiting exponential growth were used for subsequent experiments.
Localization of SMAD8 by immunohistochemistry and immunofluorescence
For immunohistochemistry, ovarian tissues were sectioned to a thickness of 5 µm after embedding in paraffin. Transverse paraffin sections were incubated with anti-SMAD8 primary antibody (Abcam, MA, USA) overnight at 4°C. Subsequently, the sections were treated with secondary antibodies conjugated with horseradish peroxidase (HRP), followed by diaminobenzidine staining and counterstaining with hematoxylin. Images were captured using a digital optical microscope (Nikon, TKY, Japan).
For immunofluorescence, mouse GCs were cultured in a six-well plate and subsequently fixed with 4% (w/v) paraformaldehyde. The GCs were incubated with SMAD8 for overnight at 4°C after blocking with 5% for 30 min at 37°C. After rinsing three times, GCs were incubated with secondary antibody goat anti-rabbit IgG (Beijing ZSGB-Bio, Beijing, China) at room temperature for 1 h. Next, the cells were stained with DAPI (Boster Biological Technology, Wuhan, China) at room temperature for 10 min and images were acquired using a fluorescent microscope (Olympus, TKY, Japan).
Design, screening, and transfection of siRNA targeting Smad8
The mouse Smad8 cDNA sequence (GenBank No. XM_006501728) was analyzed to select suitable siRNA target sites. These fragments were pre-experimentally screened to confirm their specificity to Smad8. Ultimately, a pair of oligonucleotides was designed and synthesized by Shanghai GenePharma Co., Ltd., Shanghai, China. The siRNA fragments were as follows: forward, 5′-UGUUGCCUACUACGAACUAAATT-3′, and reverse, 5′-UGUUGCCUACUACGAACUAAATT-3′. In addition, a pair of nonsense sequences were used as a control (NC-siRNA), which did not match any known sequence. The siRNA sequences targeting Smad8 and the nonsense sequences were named Smad8-siRNA and negative control siRNA (NC-siRNA), respectively.
Smad8-siRNA and NC-siRNA were transfected into the cultured mouse GCs at a dose of 16.7 nmol/L using siRNA-mate (Shanghai GenePharma, Shanghai, China) following the manufacturer’s protocol, GCs and culture medium were harvested to analyses after transfection for analysis at 24, 48 and 72 h post-transfection. The study comprised three repetitions per set, with three independent repetitions of the same treatment.
Total RNA extraction and qRT-PCR
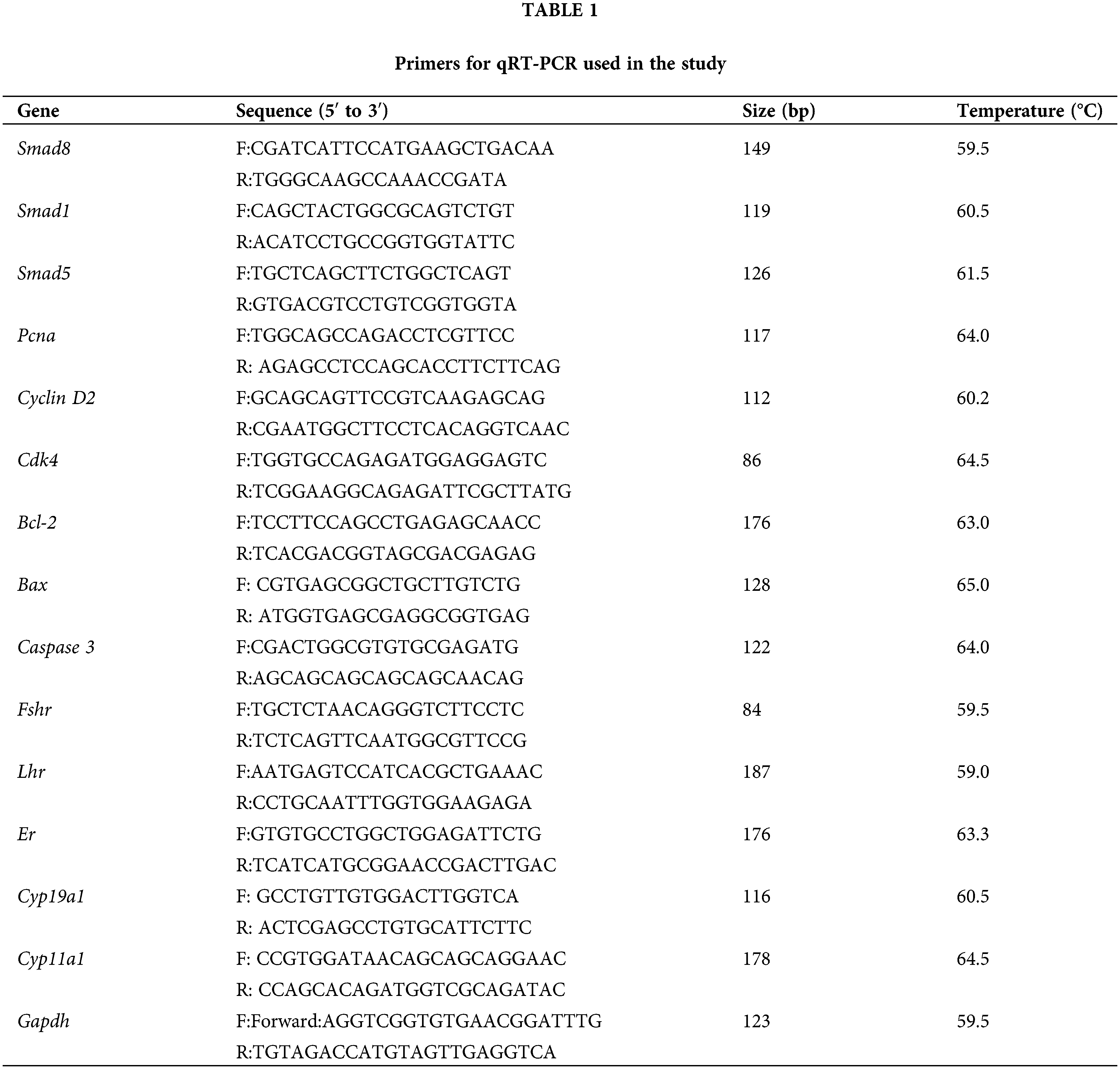
Total RNA was extracted from treated mouse GCs using Trizol reagent (Invitrogen, Carlsbad, USA) following the manufacturer’s instructions. The quality of the extracted RNA was assessed using an ultra-trace nucleic acid analyzer (Shanghai Jiapeng Technology Co., Ltd., Shanghai, China), with all samples exhibiting OD260/280 ratios between 1.8 and 2.1. The RNA was reverse-transcribed into cDNA using a reverse transcription kit (Tiangen, Beijing, China) according to the protocol of the kit. Quantitative real-time PCR (qRT-PCR) was performed with an MX3000p system (Agilent, CA, USA) using the SuperReal PreMix Plus kit with SYBR Green (Tiangen, Beijing, China) following the protocol of the kit. The sequences of the target genes and the reference gene, glyceraldehyde-3-phosphate dehydrogenase (Gapdh), were designed based on mouse mRNA sequences (Table 1). Relative RNA expression was determined using the 2−ΔΔCt analysis method [16].
Total protein extraction and western blot
Total protein was extracted from treated mouse GCs using radioimmunoprecipitation assay lysis buffer (Beyotime Biotechnology, Shanghai, China), and the protein sample concentration was measured with the BCA kit (Boster, Wuhan, China) following the protocol of the kit. Approximately 15 µg of protein was dissolved in loading buffer electrophoresed on 12% (w/v) SDS mini-gels, and transferred onto PVDF membranes (Beyotime Biotechnology, Shanghai, China). The membranes were blocked with 5% (w/v) non-fat milk in TBST buffer for 1 h at room temperature and then probed with primary antibodies against SMAD8 (Abcam, 1:1600) or GAPDH (Beijing ZSGB-Bio, Beijing, China; 1:2000) overnight at 4°C. After being washed them three times in TBST, the membranes were incubated for 1 h with secondary antibodies (goat anti-rabbit or goat anti-mouse) conjugated with HRP at room temperature and then washed again in TBST. The blot bands were visualized using an electrochemiluminescence assay (Boster Biological Technology, Wuhan, China). Each protein experiment was was repeated three times.
Measurement of cell proliferation using the cell counting Kit 8 (CCK-8) assay
Mouse GCs were cultured in a 96-well plates at a density of 3−5 × 103 cells per well. Cell proliferation of treated cells was assessed at 12, 24, 36, 48, 60 and 72 h using a CCK-8 assay kit (Beyotime Biotechnology, Shanghai, China) following the manufacturer’s instructions.
Measurement of cell apoptosis by flow cytometry
Apoptotic cells were assessed using a apoptosis detection kit (KeyGEN Biotechnology, Nanjing, China) with Annexin V-FITC/PI following the manufacturer’s protocol. Briefly, mouse GCs were digested at 48 h after treatment, washed twice for 5 min with cold PBS buffer at 600 × g and then incubated with 5 µL of annexin V-FITC/PI in a 1 ×105 cell culture for 10 min. Apoptosis rates were analyzed by FACScan (Beckman, FL, USA).
Measurement of E2 and P4 by enzyme-linked immunosorbent assay (ELISA)
Mouse GCs were cultured at a density of 3−5 × 103 cells per well in 24-well plates. The culture medium was collected at 48 h after transfection for analysis. E2 and P4 concentrations were determined in the medium using a mouse ELISA kit (Halin Biotechnology, Shanghai, China) for E2 and P4 according to the protocol of the kit. Concentrations of E2 < 20 ng/mL and P4 < 1 ng/mL were considered negligible.
Results were presented as the mean ± standard deviation (SD) of triplicate values. Data analysis and evaluation of significant differences were conducted using Student’s t-test or one-way analysis of variance, followed by multiple comparisons using Duncan’s test in Statistical Product and Service Solutions 20.0 software (SPSS Inc., Shanghai, China). Statistical significance was accepted at p < 0.05 and is denoted in figures and tables by * or different superscript lowercase letters.
Location of SMAD8 expression in mouse follicles
To elucidate the potential role of Smad8 in ovarian and follicular development, we initially investigated the localization of SMAD8 expression in the mouse ovary. As illustrated in Fig. 1, immunohistochemical results revealed specific SMAD8 expression in the follicular GCs (Fig. 1A) and no signal was detected elsewhere in the mouse ovary. Further subcellular localization as determined by immunofluorescence, indicated that SMAD8 was primarily localized in the cytoplasm of mouse GCs (Fig. 1B).
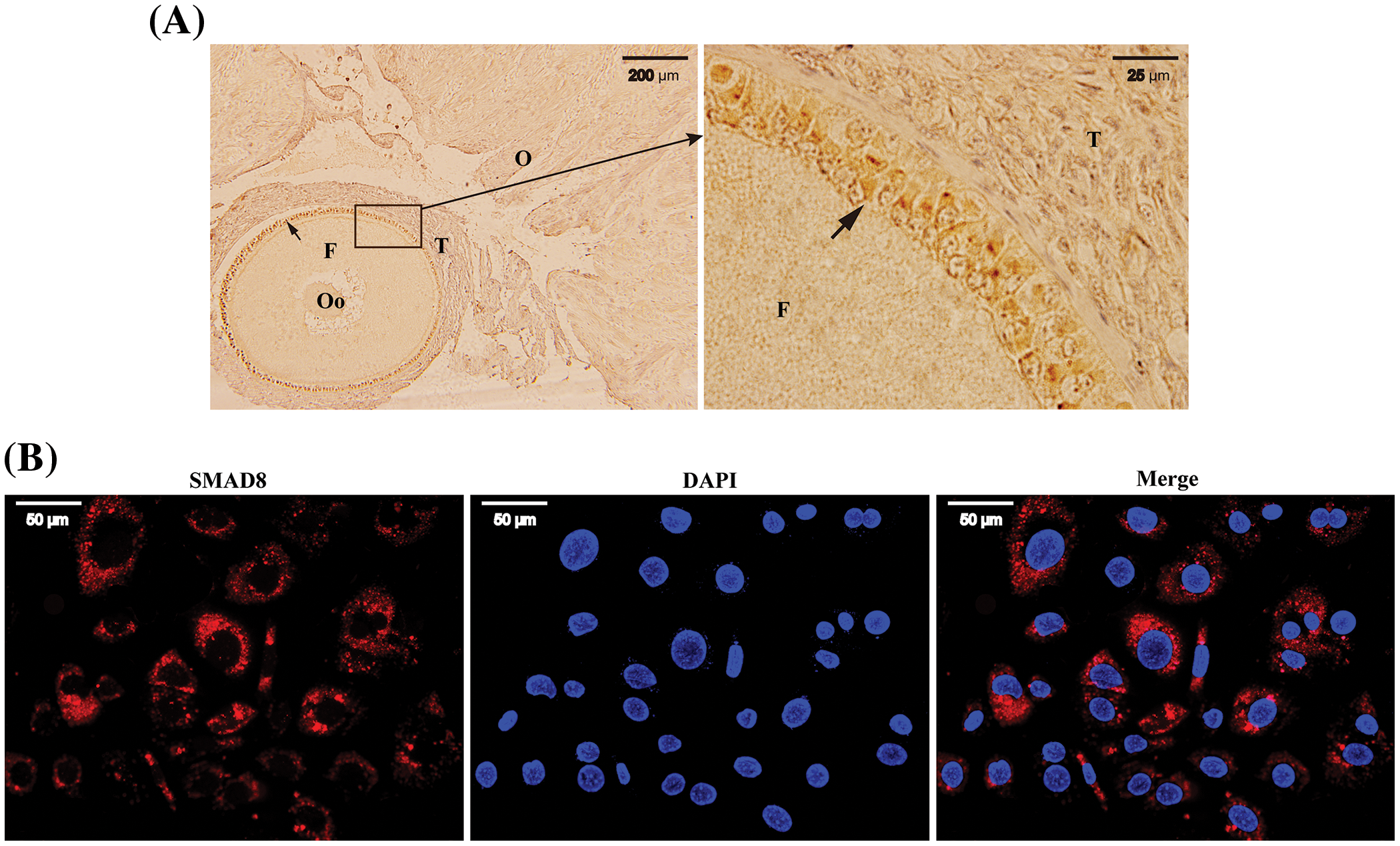
Figure 1: The location of SMAD8 in mice ovaries and follicles. Immunohistochemistry revealed that SMAD8 was expressed in the follicular GCs. The letters O, F, T and Oo represents the ovaries, the follicles, the theca interna, and the oocytes, respectively. Arrows indicate the SMAD8 signaling (left × 10; right × 100) (A). Immunofluorescence results indicated (×40) that SMAD8 was expressed in the cytoplasm of GCs (B). Red color indicates SMAD8 signaling and blue color indicates the nucleus.
Inhibition of Smad8 expression with Smad8-siRNA
We assessed the abundance of Smad8 mRNA and protein expression using qRT-PCR and western blot following transfection of Smad8-siRNA or NC-siRNA into mouse GCs. As shown in Fig. 2A, the mRNA expression of Smad8 significantly decreased at 48 and 72 h compared to the controls (blank and NC-siRNA; p < 0.05) after Smad8-siRNA transfection. The results presented in Fig. 2B demonstrated a significant reduction in SMAD8 protein levels at 48 h after transfection with Smad8-siRNA (p < 0.05). These findings suggest that the 48-h time point post-Smad8-siRNA transfection was suitable for subsequent studies. Additionally, to eliminate the potential effects of interfering with Smad8 expression on other R-SMADs, we measured the mRNA expression of Smad1 and Smad5 via qRT-PCR. The results indicated that Smad8 interference did not affect the expression of Smad1 and Smad5 (Figs. 2C and 2D). These results confirm the effectiveness of the designed interference sequence for the Smad8 gene while demonstrating no potential impact on the expression of other R-SMADs.
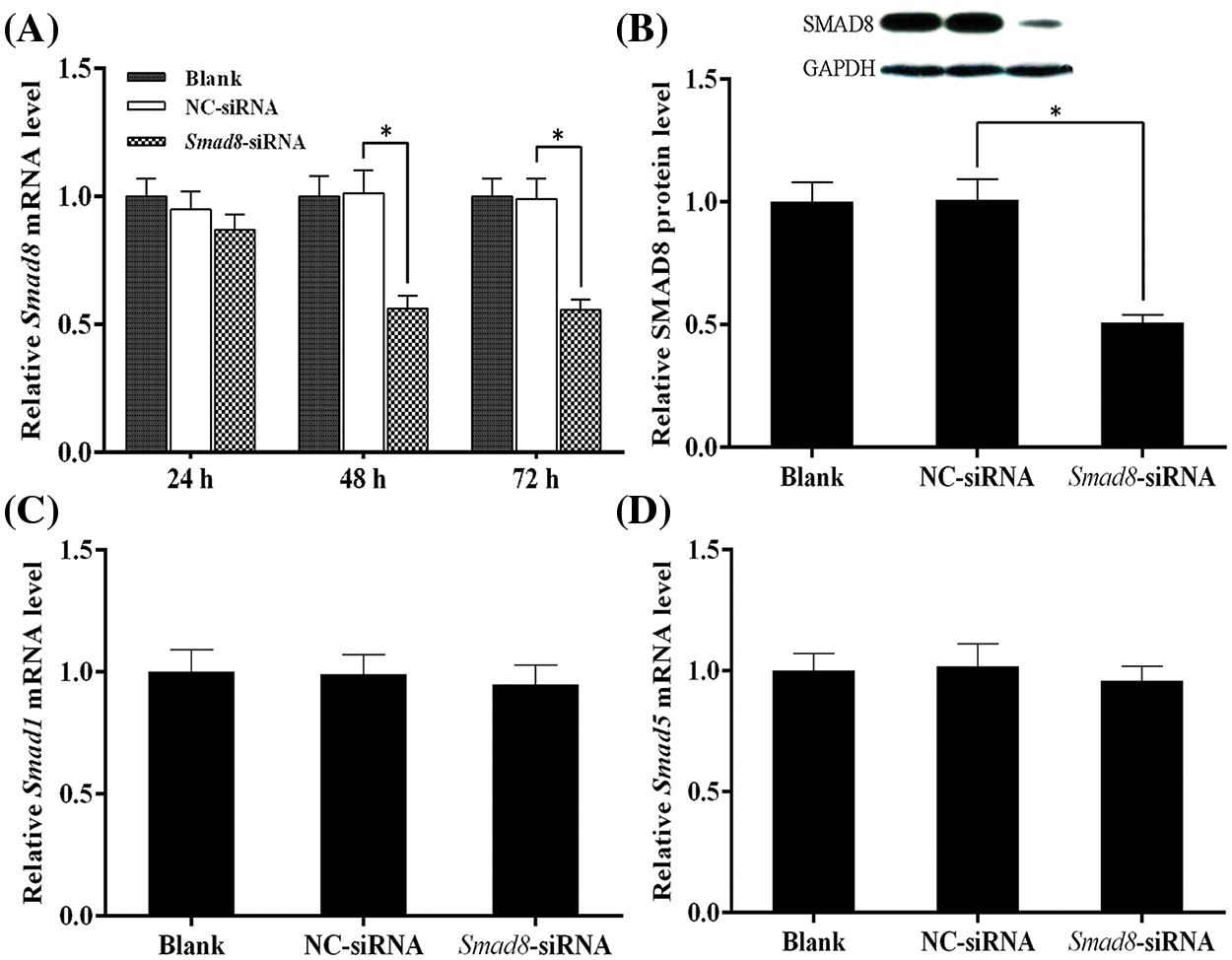
Figure 2: Analysis of Smad8 knockdown efficiency and Smad1/5 mRNA expression. The results are presented as mean ± standard deviation (n = 3). * represents p < 0.05. Analysis of Smad8 expression using qRT-PCR and western blotting (A–B). Analysis of Smad1/5 mRNA expression using qRT-PCR (C–D).
Smad8 knockdown reduces the proliferation ability of mouse GCs
To gain a deeper insight into the impact of Smad8 knockdown on mouse GCs, we employed CCK-8 analysis to measure cell proliferation ability. As anticipated, the results presented in Fig. 3 indicated a significant decrease in cell proliferation ability in mouse GCs following Smad8 silencing compared to the controls (p < 0.05). This outcome suggests that Smad8 plays a role in the proliferation of GCs.
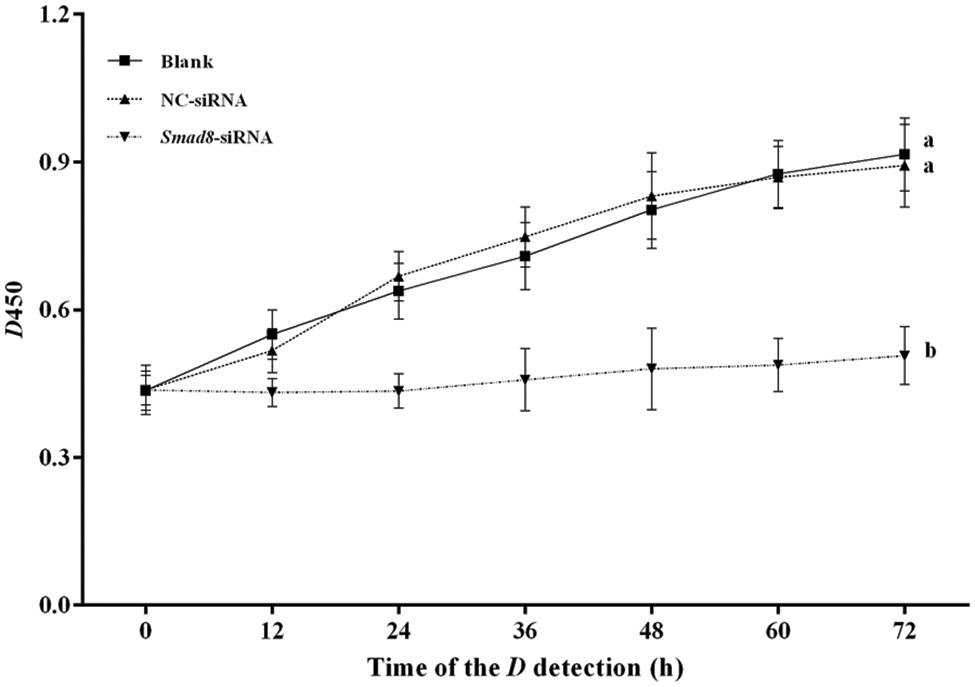
Figure 3: Analysis of GC proliferation using CCK-8 assay. The results are presented as mean ± standard deviation (n = 3). Significant differences between the results are indicated by different superscript lowercase letters on the curve. The difference between the same letters means that the difference is not significant (p > 0.05), and the difference between different letters means that the difference is significant (p < 0.05).
Smad8 knockdown enhances apoptosis in mouse GCs
To assess the effect of Smad8 silencing, we determined the apoptosis rate of treated mouse GCs using flow cytometry analysis. The results shown in Fig. 4 revealed a significant increase in the apoptosis rate in the Smad8-siRNA group compared to the controls (p < 0.05). These results demonstrate that Smad8 is involved in the apoptosis process of GCs.

Figure 4: Analysis of the GC apoptosis. The results are presented as mean ± standard deviation (n = 3). ** represents p < 0.01. Detection of apoptosis in GCs using flow cytometry (A). Analysis of apoptosis in GCs (B).
Effects of silencing Smad8 on the expression of proliferation and apoptosis-related genes
To elucidate the mechanisms underlying the inhibition of cell proliferation and the increase in apoptosis rate following Smad8 silencing, we examined the expression of genes associated with cell growth. Pcna, a marker gene of cell proliferation, is involved in DNA synthesis [17]. Cdk4 and Cyclin D2 are functional markers of the cell cycle [18]. Caspase 3, Bcl-2, and Bax are markers of apoptosis [19]. The mRNA levels of these genes associated with cell growth were measured using qRT-PCR. In the Smad8-siRNA group, Pcna, Cdk4, Cyclin D2, and Bcl-2 mRNA level significantly decreased compared to the blank group (p < 0.05), whereas Caspase 3 mRNA significantly increased (p < 0.05), with Bax mRNA remaining unchanged (Fig. 5). These results reveal that Smad8 is involved in the proliferation and apoptosis of GCs by regulating the expression of genes related to proliferation and apoptosis.
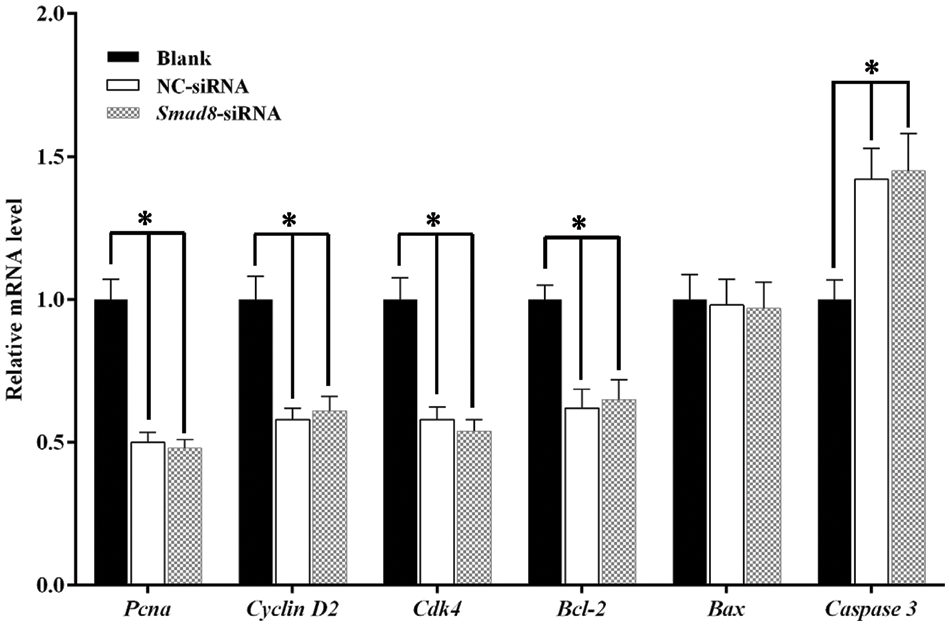
Figure 5: Gene expression related to cell proliferation and apoptosis in GCs. The results are presented as mean ± standard deviation (n = 3). * represents p < 0.05.
Smad8 knockdown reduces E2 expression with no effect on P4
E2 and P4 levels were measured in the culture medium using ELISA. As presented in Table 2, E2 concentration significantly decreased in the Smad8-siRNA group compared to the controls (p < 0.05), whereas P4 concentration exhibited a slight reduction that was not statistically significant.
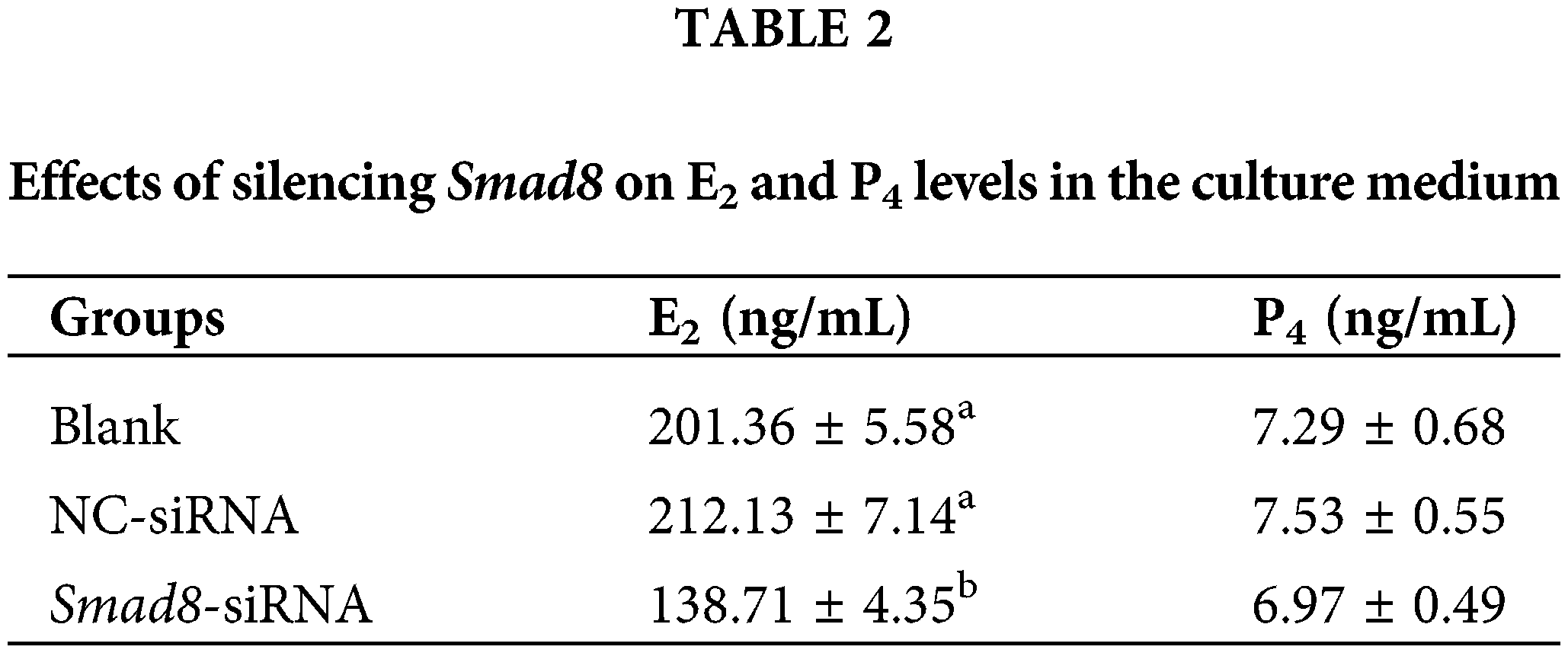
The values represent the average of at least three repetitions (n = 3). In the same column, different superscript lowercase letters represent a significant difference compared to the blank control group (p < 0.05).
Effect of Smad8 knockdown on steroidogenic enzyme and hormone receptor-related genes
To further investigate whether Smad8 knockdown altered the expression of genes, related to steroidogenesis and hormone receptors, we focused on Cyp19a1 and Cyp11a1, which are pivotal steroidogenic enzyme genes regulating E2 and P4 production [20]. Additionally, we examined Er and Fshr as well as Lhr, which are essential factors for normal follicular development [21,22]. The results demonstrated that the mRNA levels of Er, Lhr and Cyp19a1 significantly decreased in the Smad8-siRNA group compared to the controls (p < 0.05), whereas Cyp11a1 and Fshr mRNA levels remained unchanged (Fig. 6). These findings further highlight the involvement of Smad8 in the regulation of GC development through the modulation of steroid hormone synthases and reproductive hormone receptor-related genes. Additionally, a summary of the changes in key genes within the signaling pathways that regulate cell growth and steroidogenesis after Smad8 silencing is depicted in the Fig. 7.
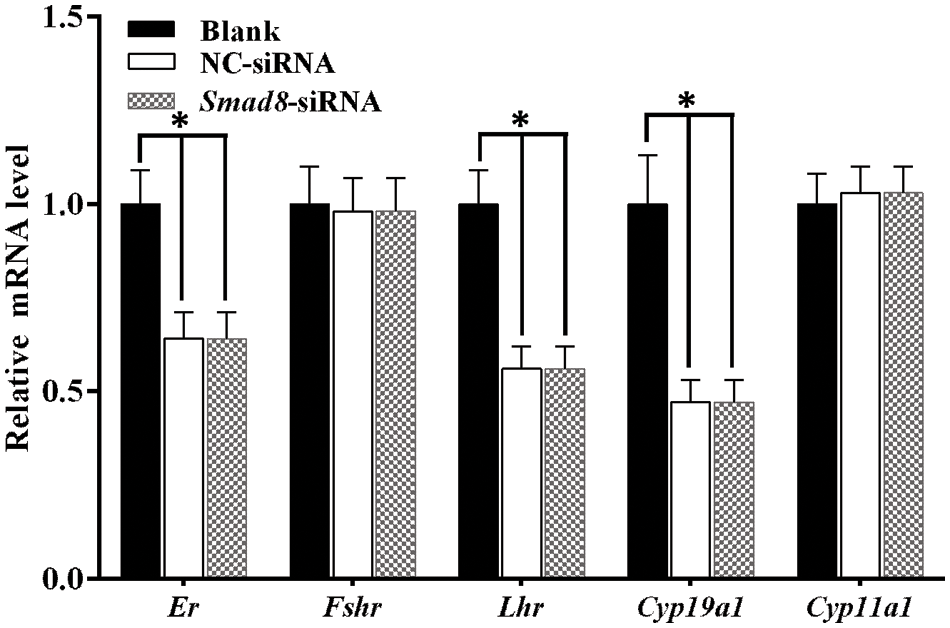
Figure 6: Expression of genes related to steroidogenic enzymes and hormone receptors in GCs. The results are presented as mean ± standard deviation (n = 3). *represents p < 0.05.
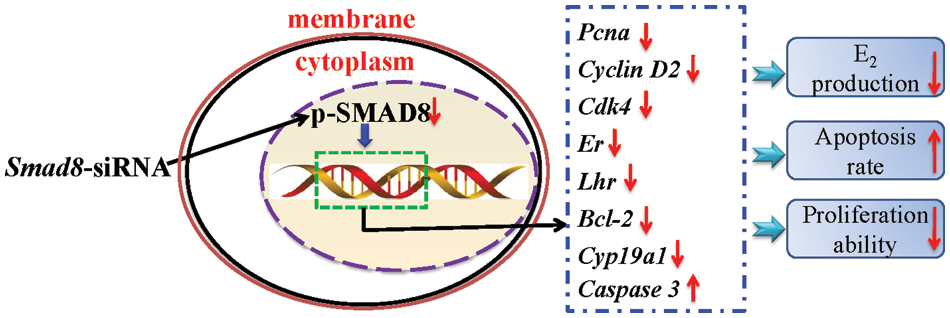
Figure 7: The mechanism underlying the upregulation of the growth and steroidogenesis by Smad8 in mouse GCs. The upward arrows next to the gene indicate a significant increase, and the downward arrows indicate a significant decrease.
The BMP/SMAD signaling transduction pathway plays a vital role in regulating mammalian reproduction, particularly during ovarian and follicular development and maturation [23,24]. Several Smad gene knockout experiments have demonstrated that SMADs can affect ovarian function in mammals and the growth and differentiation of GCs [9,12]. SMAD8 is one of important signaling molecules downstream of BMP/SMAD signaling pathway, mediating the intracellular signal transduction. However, there is limited research on Smad8 in the context of animal reproduction. To further understand the potential role of Smad8 in animal reproduction, we used RNA interference to silence Smad8 in mouse GCs in vitro. We found that the silencing Smad8 blocked the BMP/SMAD signaling pathway, resulting in the inhibition of GCs proliferation, induction of GC apoptosis, and reduction of the ability of GC to synthesize E2. These results indicate the pivotal role of Smad8 in regulating the development of follicular GC in mice, thereby influencing the follicle development process.
SMADs are widely expressed in various tissues and organs, contributing to various biological functions, with inactive R-SMADs typically residing in the cytoplasm [25,26]. Unexpectedly, our immunohistochemical results suggest that SMAD8 is exclusively expressed in the follicular GCs of mouse ovaries. Furthermore, subcellular localization via immunofluorescence demonstrates that SMAD8 signal is localized within the cytoplasm of GCs. Prior research has indicated that SMAD8 becomes activated following phosphorylation, with the activated p-SMAD8 translocating to the nucleus to regulate the expression of related genes [13]. These experimental outcomes strongly imply that Smad8 is intricately involved in the growth and differentiation of follicular GCs.
Reference [27] proposed a crucial approach for investigating the function of target gene. To comprehend the role of Smad8 in GC development, we conducted an interference experiment targeting Smad8. In our study, Smad8-siRNA significantly suppressed the expression of Smad8, as confirmed by mRNA and protein expression analyses. Due to the high homology among R-SMADs, we also assessed the expression of other R-SMADs, revealing that the Smad8-siRNA did not affect the Smad1 and Smad5 mRNA expression. Therefore, it is feasible to use Smad8-siRNA for the specific knockdown of Smad8 in mouse GCs in vitro.
The GCs are pivotal factors influencing follicular maturation and ovulation, making them indispensable for follicular development [28]. It is well-established that cell cycle progression and cell proliferation ability are two important determinants of cell growth. In our study, we examined the proliferation ability and apoptosis rate of mouse GCs and analyzed the underlying mechanisms. Our results indicate that the proliferation ability of GCs significantly weakened following Smad8 silencing. Furthermore, key factors associated with cell proliferation, such as Pcna (involved in DNA synthesis) [29], Cyclin D2, and Cdk4 (associated with cell cycle progression) [19,30] exhibited significant decreases. The apoptosis rate of GCs increased significantly, accompanied by an upregulation of Caspase 3, which promotes apoptosis, and a decrease in Bcl-2, which inhibits apoptosis [31], However, the expression of Bax, a promoter of apoptosis [32–34], remained unchanged in this study. These results suggest that the decline in cell viability after Smad8 silencing is attributable to changes in gene expression related to cell proliferation and apoptosis. Smad8 gene plays a role in the proliferation and apoptosis of GCs and has a direct impact on follicle development, maturation, and ovulation.
Mammalian follicular development is regulated by multiple endocrine hormones, including pituitary gonadotropins and steroid hormones. For instance, the combination of E2 and FSH can promotes the differentiation and selection [35], whereas P4 and LH promote follicular cell proliferation and ovulation [36]. Consequently, these hormones are crucial for follicular development and ovulation. Our results showed a significant decrease in E2 production after Smad8 inhibition, whereas P4 remained unaffected. This led us to investigate the steroidogenic enzyme genes in the treated GCs. Aromatase, encoded by Cyp19a1, is known to catalyze the conversion of androgen to estrogen [37], whereas P450scc, encoded by Cyp11a1, is a mitochondrial enzyme that plays a crucial role in the hydrolysis of cholesterol side chains to produce P4 [38]. These enzymes are key components of the steroid synthesis pathways that regulate E2 and P4 production. Our research indicated a significant decrease in Cyp19a1 expression, which is associated with estrogen synthesis, after Smad8 interference, whereas the expression of Cyp11a1 remained unaffected. Hormones play a biological role in the transmission of information by binding to their corresponding receptors. The qRT-PCR results also revealed that Er and Lhr mRNA levels decreased significantly after Smad8 knockdown, whereas Fshr mRNA expression was not affected. We hypothesize that the decrease in E2 products following Smad8 silencing is the result of decreased Cyp19a1 expression in treated GCs. Simultaneously, hormone receptor Er and Lhr which are closely related to GC development, exhibited decreased expression in treated GCs. These receptors are crucial for GC development, ultimately affecting their growth and function. The development of follicles fundamentally hinges on the interaction between oocytes and GCs. Therefore, any factors affecting GC development will significantly impact follicular growth, determining whether the follicle matures and eventually ovulates. This study shows the close relationship between the Smad8 gene and GC development, suggesting that further investigation on this gene will help to elucidate the development mechanisms underlying mouse follicular development.
In conclusion, we used RNA interference technique to silence the expression of Smad8 in this study, leading to a decline in growth and steroidogenesis ability of mouse GCs. Therefore, it is concluded that Smad8 is closely related to the mammalian follicular GCs development and function maintenance, and may be a new genetic marker gene that affects mammalian reproduction.
Acknowledgement: None.
Funding Statement: This research was supported by the High-Level Talent Research Start-Up Funds of West Anhui University (No. WGKQ2021031); Key Project of Natural Science Foundation of Anhui Province of China (No. 2108085QC136); Key Project of Quality Engineering in Higher Education Institutions of Anhui Province (No. 2020jyxm2128); National College Student Innovation and Entrepreneurship Training Program (No. 202110370093). Innovation and Entrepreneurship Training Program for College Students of Anhui Province (No. S202010376114).
Author Contributions: The authors confirm their contribution to the paper as follows: Daolun Yu designed the experiments and writing. Deyong She and Lei Yang analyzed the data and prepared some figures. Ruina Zhang and Kai Ge performed the experiment and editing the article. Shan Lu and Yafei Cai supervised the project and reviewing, and editing the article. All authors have revised and agreed to publish this article. All authors reviewed and approved the manuscript.
Availability of Data and Materials: The datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request.
Ethics Approval: This research was approved by the Ethics Committee of the West Anhui University (Approval No. 202301001).
Conflicts of Interest: The authors declare that they have no conflicts of interest to report regarding the present study.
References
1. Kashino C, Hasegawa T, Nakano Y, Iwata N, Yamamoto K, Kamada Y, et al. Involvement of BMP-15 in glucocorticoid actions on ovarian steroidogenesis by rat granulosa cells. Biochem Biophys Res Commun. 2021;559:56–61. [Google Scholar] [PubMed]
2. Suyama A, Iwata N, Soejima Y, Nakano Y, Yamamoto K, Nada T, et al. Roles of NR5A1 and NR5A2 in the regulation of steroidogenesis by clock gene and bone morphogenetic proteins by human granulosa cells. Endocr J. 2021;68(11):1283–91. [Google Scholar] [PubMed]
3. Chakraborty P, Anderson RL, Roy SK. Bone morphogenetic protein 2- and estradiol-17β-induced changes in ovarian transcriptome during primordial follicle formation. Biol Reprod. 2022;107(3):800–12. [Google Scholar] [PubMed]
4. Yang D, Yang X, Dai F, Wang Y, Yang Y, Hu M, et al. The role of bone morphogenetic protein 4 in ovarian function and diseases. Reprod Sci. 2021;28(12):3316–30. [Google Scholar] [PubMed]
5. Mattar D, Samir M, Laird M, Knight PG. Modulatory effects of TGF-β1 and BMP6 on thecal angiogenesis and steroidogenesis in the bovine ovary. Reprod. 2020;159(4):397–408. [Google Scholar]
6. Paulini F, Melo EO. Effects of growth and differentiation factor 9 and bone morphogenetic protein 15 overexpression on the steroidogenic metabolism in bovine granulosa cells in vitro. Reprod Domest Anim. 2021;56(6):837–47. [Google Scholar] [PubMed]
7. Hlokoe VR, Tyasi TL, Gunya B. Chicken ovarian follicles morphology and growth differentiation factor 9 gene expression in chicken ovarian follicles: review. Heliyon. 2022;8(1):e08742. [Google Scholar] [PubMed]
8. Luo H, Chen B, Weng B, Tang X, Chen Y, Yang A, et al. miR-130a promotes immature porcine Sertoli cell growth by activating SMAD5 through the TGF-β-PI3K/AKT signaling pathway. Faseb J. 2020;34(11):15164–79. [Google Scholar] [PubMed]
9. Senft AD, Bikoff EK, Robertson EJ, Costello I. Genetic dissection of Nodal and Bmp signalling requirements during primordial germ cell development in mouse. Nat Commun. 2019;10(1):1089. [Google Scholar] [PubMed]
10. Shen H, Wang Y. Activation of TGF-β1/Smad3 signaling pathway inhibits the development of ovarian follicle in polycystic ovary syndrome by promoting apoptosis of granulosa cells. J Cell Physiol. 2019;234(7):11976–85. [Google Scholar] [PubMed]
11. Li M, Liang W, Zhu C, Qin S. Smad4 mediates Bmf involvement in sheep granulosa cell apoptosis. Gene. 2022;817:146231. [Google Scholar] [PubMed]
12. Wang Q, Xiong F. SMAD proteins in TGF-β signalling pathway in cancer: regulatory mechanisms and clinical applications. Diagnostics. 2023;13(17):2769. [Google Scholar] [PubMed]
13. Tsukamoto S, Mizuta T, Fujimoto M, Ohte S, Osawa K, Miyamoto A, et al. Smad9 is a new type of transcriptional regulator in bone morphogenetic protein signaling. Sci Rep. 2014;4:7596. [Google Scholar] [PubMed]
14. Yu D, Zhang L, Wang H, Chen F, Chen J, Zhang Z, et al. A potential role for SMAD9 in goose follicular selection through regulation of mRNA levels of luteinizing hormone receptor. Theriogenology. 2019;135:204–12. [Google Scholar] [PubMed]
15. Liu S, Jia Y, Meng S, Luo Y, Yang Q, Pan Z. Mechanisms of and potential medications for oxidative stress in ovarian granulosa cells: a review. Int J Mol Sci. 2023;24(11):9205. [Google Scholar] [PubMed]
16. Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods. 2001;25(4):402–8. [Google Scholar] [PubMed]
17. Fujii S, Fuchs RP. A comprehensive view of translesion synthesis in Escherichia coli. Microbiol Mol Biol Rev. 2020;84(3):e00002–20. [Google Scholar] [PubMed]
18. Pieters T, T’Sas S, Vanhee S, Almeida A, Driege Y, Roels J, et al. Cyclin D2 overexpression drives B1a-derived MCL-like lymphoma in mice. J Exp Med. 2021;218(10):e20202280. [Google Scholar] [PubMed]
19. Zhou Y, Bai L, Tian L, Yang L, Zhang H, Zhang Y, et al. Iridium(III)-BBIP complexes induce apoptosis via PI3K/AKT/mTOR pathway and inhibit A549 lung tumor growth in vivo. J Inorg Biochem. 2021;223:111550. [Google Scholar] [PubMed]
20. Wang Y, Chen M, Xu J, Liu X, Duan Y, Zhou C, et al. Core clock gene Bmal1 deprivation impairs steroidogenesis in mice luteinized follicle cells. Reprod. 2020;160(6):955–67. [Google Scholar]
21. Yang J, Gong Z, Shen X, Bai S, Bai X, Wei S. FSH receptor binding inhibitor depresses carcinogenesis of ovarian cancer via decreasing levels of K-Ras, c-Myc and FSHR. Anim Biotechnol. 2021;32(1):84–91. [Google Scholar] [PubMed]
22. Yurina A, Skovorodin E, Dolmatova I. Evaluation of reproductive traits and ovarian and uterine morphology of sows with different genotypes for the estrogen receptor, prolactin receptor, and follicle-stimulating hormone subunit beta genes. Can J Vet Res. 2021;85(3):186–92. [Google Scholar] [PubMed]
23. de Moraes FP, Missio D, Lazzari J, Rovani MT, Ferreira R, Gonçalves PBD, et al. Local regulation of antral follicle development and ovulation in monovulatory species. Anim Reprod. 2022;19(4):e20220099. [Google Scholar] [PubMed]
24. Magro-Lopez E, Muñoz-Fernández M. The role of BMP signaling in female reproductive system development and function. Int J Mol Sci. 2021;22(21):11927. [Google Scholar] [PubMed]
25. Peng Y, Shi XE, Huang KL, Yao XP, Chen FF, Li X, et al. Knock-down Sox5 suppresses porcine adipogenesis through BMP R-Smads signal pathway. Biochem Biophys Res Commun. 2020;527(2):574–80. [Google Scholar] [PubMed]
26. Sisto M, Ribatti D, Lisi S. SMADS-mediate molecular mechanisms in sjögren’s syndrome. Int J Mol Sci. 2021;22(6):3203. [Google Scholar] [PubMed]
27. Wang P, Zhou Y, Richards AM. Effective tools for RNA-derived therapeutics: siRNA interference or miRNA mimicry. Theranostics. 2021;11(18):8771–96. [Google Scholar] [PubMed]
28. Kulus M, Kranc W, Sujka-Kordowska P, Mozdziak P, Jankowski M, Konwerska A, et al. The processes of cellular growth, aging, and programmed cell death are involved in lifespan of ovarian granulosa cells during short-term IVC-study based on animal model. Theriogenology. 2020;148:76–88. [Google Scholar] [PubMed]
29. Fan L, Bi T, Wang L, Xiao W. DNA-damage tolerance through PCNA ubiquitination and sumoylation. Biochem J. 2020;477(14):2655–77. [Google Scholar] [PubMed]
30. Knudsen ES, Kumarasamy V, Nambiar R, Pearson JD, Vail P, Rosenheck H, et al. CDK/cyclin dependencies define extreme cancer cell-cycle heterogeneity and collateral vulnerabilities. Cell Rep. 2022;38(9):110448. [Google Scholar] [PubMed]
31. Kaloni D, Diepstraten ST, Strasser A, Kelly GL. BCL-2 protein family: attractive targets for cancer therapy. Apoptosis. 2023;28(1–2):20–38. [Google Scholar] [PubMed]
32. Aghaei M, KhanAhmad H, Aghaei S, Ali Nilforoushzadeh M, Mohaghegh MA, Hejazi SH. The role of Bax in the apoptosis of Leishmania-infected macrophages. Microb Pathog. 2020;139:103892. [Google Scholar] [PubMed]
33. Ganbarjeddi S, Azimi A, Zadi Heydarabad M, Hemmatzadeh M, Mohammadi S, Mousavi Ardehaie R, et al. Apoptosis induced by prednisolone occurs without altering the promoter methylation of BAX and BCL-2 genes in acute lymphoblastic leukemia cells CCRF-CEM. Asian Pac J Cancer Prev. 2020;21(2):523–9. [Google Scholar] [PubMed]
34. Niu H, Chen X, Fu X, Zhang J, Li G, Wang Y, et al. SOD1G93A induces a unique PSAP-dependent mitochondrial apoptosis pathway via Bax-Bak interaction. BIOCELL. 2021;45(4):963–70. doi:https://doi.org/10.32604/biocell.2021.015297. [Google Scholar] [CrossRef]
35. Grymowicz M, Rudnicka E, Podfigurna A, Napierala P, Smolarczyk R, Smolarczyk K, et al. Hormonal effects on hair follicles. Int J Mol Sci. 2020;21(15):5342. [Google Scholar] [PubMed]
36. Ding Z, Duan H, Ge W, Lv J, Zeng J, Wang W, et al. Regulation of progesterone during follicular development by FSH and LH in sheep. Anim Reprod. 2022;19(2):e20220027. [Google Scholar] [PubMed]
37. Sahlman H, Itkonen A, Lehtonen M, Keski-Nisula L, Rysä J. Altered activities of CYP1A1 and CYP19A1 enzymes in women using SSRI medication during pregnancy. Placenta. 2022;129:30–5. [Google Scholar] [PubMed]
38. Heidarzadehpilehrood R, Pirhoushiaran M, Abdollahzadeh R, Binti Osman M, Sakinah M, Nordin N, et al. A review on CYP11A1, CYP17A1, and CYP19A1 polymorphism studies: candidate susceptibility genes for polycystic ovary syndrome (PCOS) and infertility. Genes. 2022;13(2):302. [Google Scholar] [PubMed]
Cite This Article
 Copyright © 2024 The Author(s). Published by Tech Science Press.
Copyright © 2024 The Author(s). Published by Tech Science Press.This work is licensed under a Creative Commons Attribution 4.0 International License , which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.


 Submit a Paper
Submit a Paper Propose a Special lssue
Propose a Special lssue View Full Text
View Full Text Download PDF
Download PDF Downloads
Downloads
 Citation Tools
Citation Tools