 Open Access
Open Access
ARTICLE
Galectin 2 regulates JAK/STAT3 signaling activity to modulate oral squamous cell carcinoma proliferation and migration in vitro
1 Medical School, University of Electronic Science and Technology of China, Chengdu, 610072, China
2 Department of Stomatology, Sichuan Provincial People’s Hospital, University of Electronic Science and Technology of China, Chengdu, 610072, China
* Corresponding Author: LI XIAO. Email:
(This article belongs to the Special Issue: Subcellular Organelles and Cellular Molecules: Localization, Detection, Prediction, and Diseases)
BIOCELL 2024, 48(5), 793-801. https://doi.org/10.32604/biocell.2024.048395
Received 06 December 2023; Accepted 18 February 2024; Issue published 06 May 2024
Abstract
Background: Galectin 2 (LGALS2) is a protein previously reported to serve as a mediator of disease progression in a range of cancers. The function of LGALS2 in oral squamous cell carcinoma (OSCC), however, has yet to be explored, prompting the present study to address this literature gap. Methods: Overall, 144 paired malignant tumor tissues and paracancerous OSCC patient samples were harvested and the LGALS2 expression levels were examined through qPCR and western immunoblotting. The LGALS2 coding sequence was introduced into the pcDNA3.0 vector, to enable the overexpression of this gene, while an LGALS2-specific shRNA and corresponding controls were also obtained. The functionality of LGALS2 as a regulator of the ability of OSCC cells to grow and undergo apoptotic death in vitro was assessed through EdU uptake and CCK-8 assays, and flow cytometer, whereas a Transwell system was used to assess migratory activity and invasivity. An agonist of the Janus Kinase 2 (JAK2)/Signal Transducer and Activator of Transcription 3 (STAT3) pathway was also used to assess the role of this pathway in the context of LGALS2 signaling. Results: Here, we found that lower LGALS2 protein and mRNA expression were evident in OSCC tumor tissue samples, and these expression levels were associated with clinicopathological characteristics and patient survival outcomes. Silencing LGALS2 enhanced proliferation in OSCC cells while rendering these cells better able to resist apoptosis. The opposite was instead observed after LGALS2 was overexpressed. Mechanistically, the ability of LGALS2 to suppress the progression of OSCC was related to its ability to activate the JAK/STAT3 signaling axis. Conclusion: Those results suggest a role for LGALS2 as a suppressor of OSCC progression through its ability to modulate JAK/STAT3 signaling, supporting the potential utility of LGALS2 as a target for efforts aimed at treating OSCC patients.Keywords
Oral squamous cell carcinoma (OSCC) is the most frequently diagnosed head and neck malignancy [1], with approximately 54,540 new diagnoses throughout the globe in 2023 alone [2]. While any patients can be effectively treated through surgical tumor resection in combination with radiotherapy and chemotherapy, the 5-year survival rate for individuals with OSCC remains below 50% and has improved little substantially over the past two decades [3]. Metastatic progression is one of the primary drivers of mortality in OSCC, with tumors often disseminating to the lungs and other distant tissues such that patients with metastatic disease generally survive for less than one year [3,4]. The research focused on clarifying the processes that drive the proliferative and metastatic activity of OSCC cells thus have the potential to inform new efforts to prevent or treat progressive forms of this cancer.
Members of the galectin (LGALS) glycoprotein protein family exhibit galactoside-binding activity and harbor carbohydrate recognition domains (CRDs) that are approximately 130 amino acids in length and highly conserved [5]. The 15 mammalian LGALS family proteins (LGALS1-15) exhibit varying patterns of expression in different tumor types and healthy tissues [6]. Functionally, they have been shown to shape important processes including differentiation, cell adhesion, inflammatory activity, signaling within cells, and metastatic progression [7–9]. The dysregulated expression of LGALS proteins has been linked to invasivity and metastasis in several tumors [10,11]. Thijssen et al. [12] reported close associations between LGALS levels and key tumor-related parameters including clinical staging, grade, and lymph node status. There are also multiple reports highlighting the prognostic value of LGALS members in particular forms of cancer [13,14]. No research to date, however, has fully documented OSCC expression patterns for particular LGALS proteins or verified their prognostic relevance.
In papillary thyroid carcinoma and breast cancer, LGALS2 upregulation was recently found to be linked with better outcomes [15,16]. In vivo CRISPR screening has further established LGALS2 as a candidate immune escape-related regulatory protein in triple-negative breast cancer (TNBC) such that antibody-mediated LGALS2 blockade was successful as a means of reversing immunosuppression and arresting tumor growth [17]. In a separate whole-genome CRISPR knockout screening efforts, LGALS2 enrichment was detected in cells exposed to sublethal H2O2 challenge, with its overexpression suppressing STAT3 activation in response to H2O2 and impairing human colon tumor epithelial cell proliferative activity [18]. Given the absence of any studies focused on the prognostic or functional relevance of LGALS2 in OSCC to date, this study was developed in an effort to more fully understand how this protein functions in this cancer type.
An in vitro strategy was employed to assess the functions played by LGALS2, evaluating OSCC cell proliferative activity, migration, invasivity, and apoptosis following its silencing or overexpression. The role of the JAK/STAT3 signaling axis was also assessed in the context of LGALS2 overexpression. Together, these analyses revealed the ability of LGALS2 to suppress the growth of OSCC tumors via regulating JAK2/STAT3 signaling.
Pairs of tumors and paracancerous tissue samples were collected from 144 OSCC patients undergoing surgical treatment, and corresponding clinical and follow-up data for these individuals were obtained. The Ethics Committee of The Sichuan Provincial People’s Hospital approved the present study (Lunshen (Yan)2022-50), which conformed to the 2013 Declaration of Helsinki. All patients gave written informed consent for study participation.
SCC-25 and SCC-9 cells from the Cell Bank (Shanghai, China) were cultured in dulbecco’s modified eagle medium (DMEM) containing 10% Fetal Bovine Serum (FBS, A5669401, Gibco, MD, USA) with 1% penicillin-streptomycin solution (BL505A, Biosharp, Beijing, China). Every 6 days, cells were split 1:3, exchanging media every second day. To knock down LGALS2, shRNA (5′-AAUGAGUUGCAGACAAUGGUG-3′) and control plasmids were purchased from Genepharma (Shanghai, China), whereas LGALS2 (NM_006498.3) was overexpressed by preparing a pcDNA3.0 vector harboring its coding sequence. Lipofectamine 2000 (11668500, Invitrogen, CA, USA) was employed to transfect cells at 80%–90% confluency as directed, and media was replaced at 6 h post-transfection, collecting cells for downstream use after a further 24–48 h.
A CCK-8 kit (V13154, Invitrogen, CA, USA) was employed as a tool to assess viability. Briefly, SCC-25 and SCC-9 cells were seeded into a 96-well plates with 5,000 cells per well and then 100 μL of DMEM (Thermo Fisher Scientific, USA) containing 10% CCK-8 reagent was added per well for 2 h. A NanoDrop 2000c microplate reader (Thermo Fisher Scientific) was subsequently employed to measure absorbance at 450 nm.
An EdU (5-ethynyl-20-deoxyuridine) assay kit (C10310, Ribobio, Guangzhou, China) was utilized as a tool to quantify proliferation. After adding cells into confocal plates (10 × 105/well), EdU buffer (50 μM) was added followed by a 2 h incubation at 37°C. Following a 30 min fixation with 4% formaldehyde, cells were permeabilized using 0.1% Triton X-100 for 20 min, stained with EdU solution, and Hoechst was used for nuclear counterstaining prior to imaging under a fluorescence microscope.
After transfection, OSCC cell lines were cultured until 90% confluent in 6-well plates, collected, and stained using 10 μL of dye from an Annexin V-fluorescein isothiocyanate (FITC)/Propidium Iodide (PI) apoptosis detection kit (AP101, Lianke Biotech Co., Ltd., Hangzhou, China) for 15 min while protected from light at room temperature. Cells were subsequently analyzed through the use of a flow cytometer (FACS Calibur, BD Biosciences, USA).
Following a 24 h fixation step in 4% paraformaldehyde (G1101, Servicebio, Wuhan, China), 4 μm Paraffin sectioning was performed in a sequence involving de-paraffination, antigen retrieving, endogenous-peroxidase inactivation, block-step using goat serum (BL1097A, Biosharp, Hefei, China) IHC staining of OSCC tumor and healthy adjacent tissues was performed with anti-LGALS2 (Solarbio, K006747P; 1:100), imaging stained samples under a light microscope (Olympus, Tokyo, Japan).
After extracting RNA using TRIzol (15596026, Invitrogen, CA, USA), cDNA was prepared and qPCR was analyzed with SYBR Green (TaKaRa, Otsu, Shiga, Japan) and the following settings: 40 cycles of 94°C for 30 s, 55°C for 30 s, and 72°C for 90 s. Relative gene expression was computed with the 2−ΔΔCt technique. These analyses utilized the following primers: Homo sapiens-LGALS2-F 5′-GCGAATCCACCATTGTCTGC-3′, Homo sapiens-LGALS2-R 5′-GGCTGAAGCACAGGTGATCT-3′, Homo sapiens-GAPDH-F 5′-GACAGTCAGCCGCATCTTCT-3′, and Homo sapiens-GAPDH-R 5′-GCGCCCAATACGACCAAATC-3′.
For transwell migration assay, in total, 3 × 103 cells in 100 μL were transferred into the upper portion of Transwell insert (354234, Corning, NY, USA), filling the lower chamber using 600 μL of media containing 20% FBS. Following a 24 h incubation period, methanol was used to fix cells for 15 min, after which crystal biolet staining was conducted for 20 min, and cells were imaged with a microscope (ICX41, SunGrant, Suzhou, China), counting cells in five random 200x fields of view. For transwell invasion assay, in total, 3 × 103 cells in 100 μL were transferred into the upper portion of a Matrigel coated Transwell insert (354234, Corning, NY, USA), and then cultured and followed with transwell migration assay.
Following RIPA buffer (G2002, Servicebio, Wuhan, China)-mediated protein extraction and quantification through the use of a BCA assay (Beyotime, Shanghai, China), proteins were separated through electrophoresis prior to transfer onto PVDF membranes (Millipore, MA, USA). Blots were blocked with 5% skim milk for 2 h, probed with primary (LGALS2 (Solarbio, K006747P; 1:1000); JAK2 (Solarbio, K002264P; 1:1000); p-JAK2 (Solarbio, K009379P; 1:1000); STAT3 (Solarbio, K000233M; 1:1000); p-STAT3 (Solarbio, K009371P; 1:1000)) and secondary antibodies HRP-conjugated Affinipure Goat Anti-Mouse IgG(H+L) (Cat No. SA00001-1, proteintech, Wuhan, China) or HRP-conjugated Affinipure Goat Anti-Rabbit IgG(H+L) (Cat No. SA00001-2, proteintech, Wuhan, China), imaged via enhanced chemiluminescence (ECL) (BL520B, Biosharp, Beijing, China), and analyzed using ImageJ (NIH, MD, USA).
GraphPad Prism 7.0 (GraphPad, CA, USA) was employed for analyses of data based on at least three replicate experiments. Results are means ± standard deviation (SD), and were compared with one-way ANOVAs or t-tests. Chi-square tests were used to probe relationships between the expression of LGALS2 and clinicopathological characteristics. p < 0.05 was selected as the cut-off when defining significance.
LGALS2 is a prognostic factor that is downregulated in OSCC
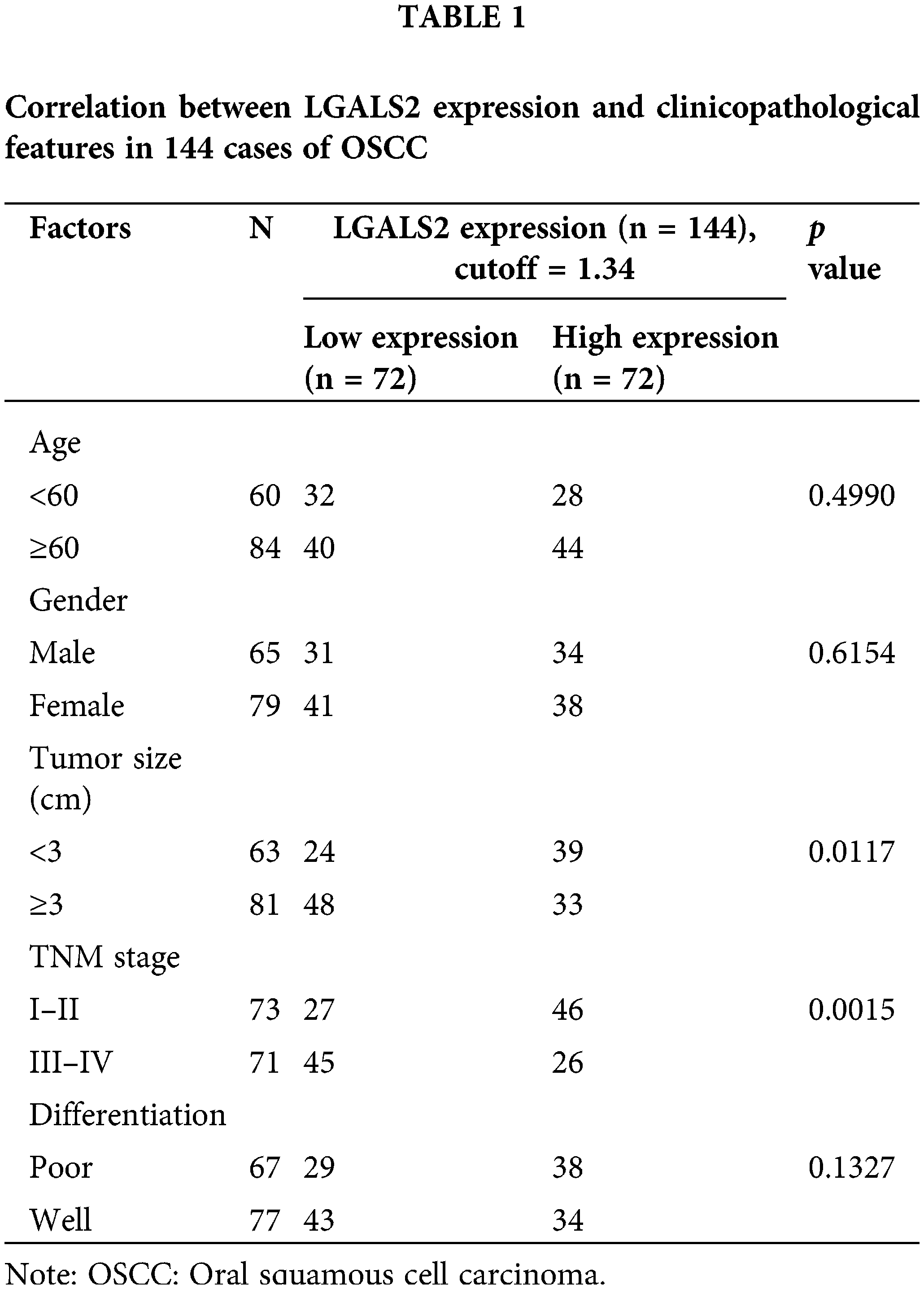
To determine how effectively LGALS2 can be employed to aid in diagnosing OSCC, 144 pairs of tumors and adjacent healthy tissue from individuals with OSCC were analyzed, with qPCR revealing significant LGALS2 mRNA downregulation in tumors as compared to matched healthy tissue (p < 0.001; Fig. 1A). Similar protein-level results were also detected through western immunoblotting (Figs. 1B, 1C), and IHC staining also demonstrated a pronounced drop in tumor-associated LGALS2 expression relative to paracancerous samples when analyzing these 144 pairs of OSCC patient-derived tissues (Figs. 1D, 1E). A strong correlation between LGALS2 levels and the TNM staging and tumor size of these patients was also observed (Table 1). When overall survival was compared between those patients in this cohort expressing high and low levels of LGALS2, the expression of higher levels of this gene was confirmed to be associated with better patient outcomes (Fig. 1F). The downregulation of LGALS2 may thus be an effective prognostic biomarker for OSCC.
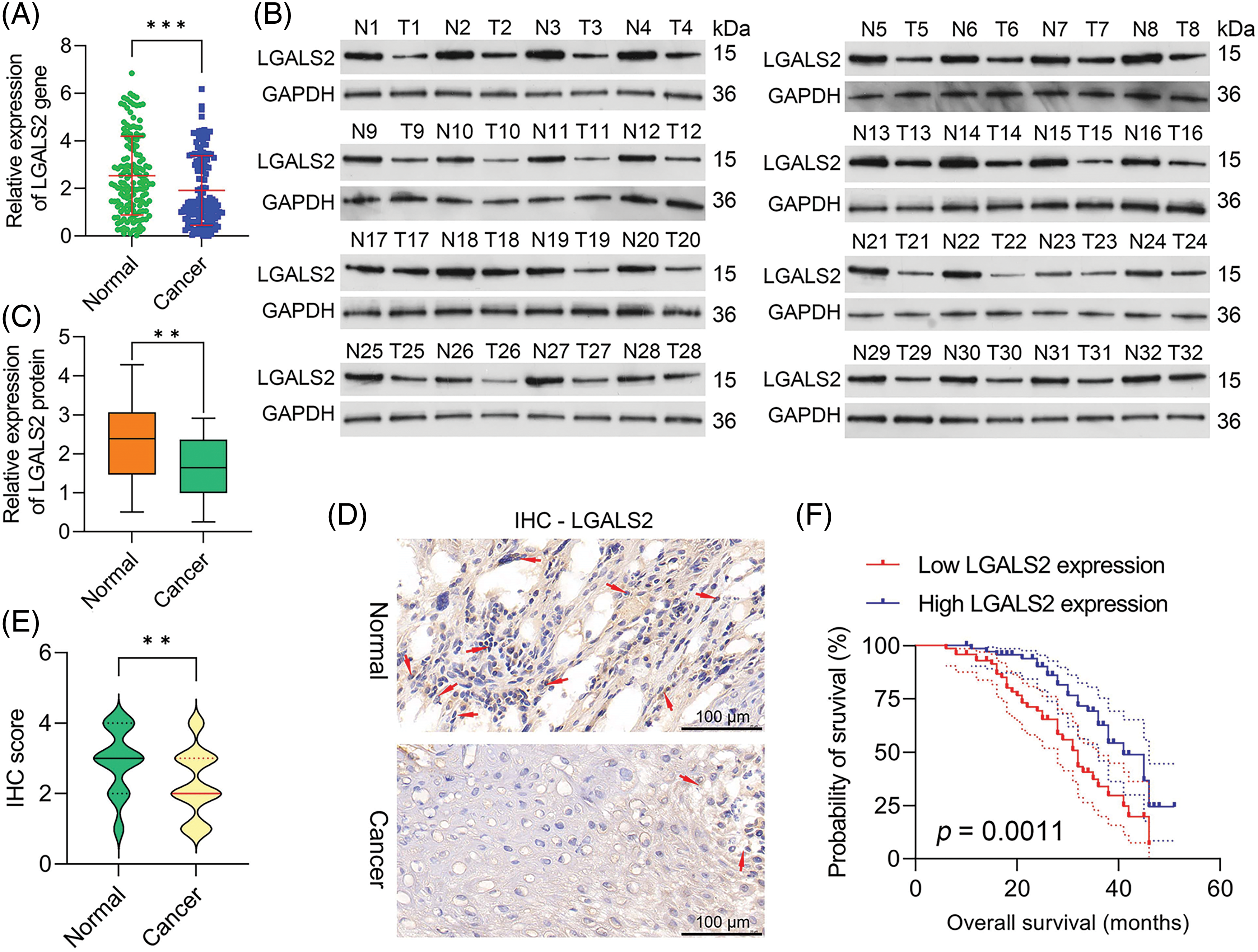
Figure 1: LGALS downregulation in OSCC tumor tissues is predictive of a poor prognosis. (A) qPCR was used to compare LGALS2 levels within tumors and paracancerous tissues for OSCC patients, normalizing the results to the indicated control group. (B) Analyses of 32 paired tumor tissue samples revealed a significant reduction in LGALS2 protein content within the tumors of OSCC patients relative to adjacent healthy tissue. (C) LGALS2 protein levels in tumors and adjacent tissues from 32 OSCC patients were quantified. (D and E) Representative IHC images (Scale bar: 100 μm) and corresponding IHC score-based quantification of LGALS2 levels in tumors and paracancerous tissues. (F) Kaplan-Meier curves were used to assess relationships between LGALS2 expression and overall survival. Results from three repeated experiments are given as means ± standard deviation. **p < 0.01; ***p < 0.001.
LGALS2 downregulation facilitates OSCC cell proliferation in vitro
To elucidate the functional roles that LGALS2 plays in OSCC, it was next silenced or overexpressed in SCC-25 and SCC-9 cells, with the efficiency of knockdown/overexpression having successfully been confirmed through qPCR and western immunoblotting (Figs. 2A–2C). Using a CCK-8 approach, overexpressing LGALS2 was found to suppress the ability of OSCC cells to proliferate as compared to control construct transfection (Fig. 2D), while the opposite was observed upon LGALS2 silencing. The same phenotypes were also detected in EdU uptake assays (Figs. 2E, 2F). Flow cytometry also revealed that silencing LGALS2 helped protect against apoptotic OSCC cell death, whereas apoptosis rates were higher following the overexpression of this protein (Figs. 2G, 2H). These findings support a functional role for LGALS2 as a tumor suppressor in patients with OSCC.
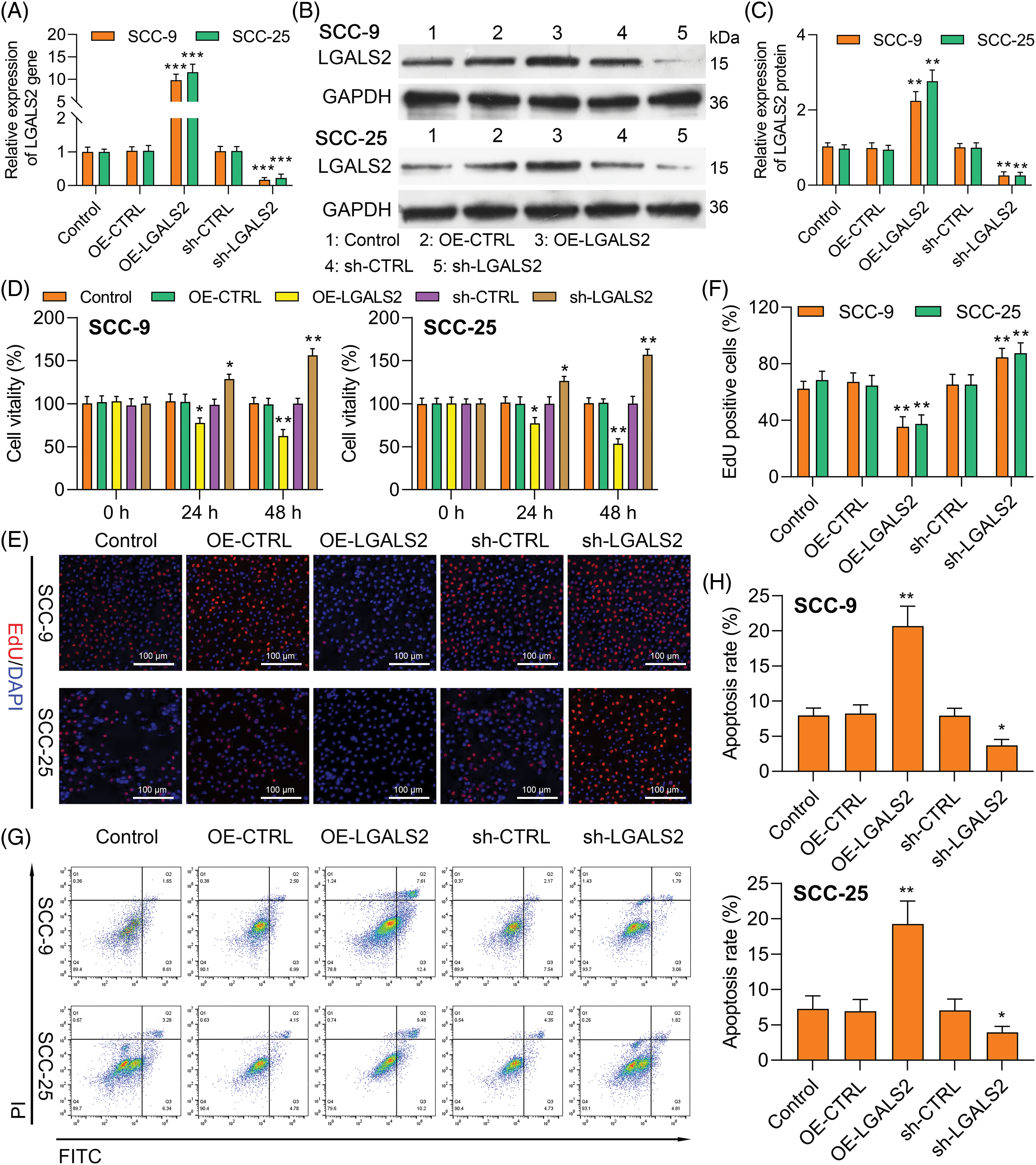
Figure 2: LGALS2 knockdown enhances proliferative activity and induces apoptosis (A–C) SCC-25 and SCC-9 cells were transfected with the OE-LGALS2, sh-LGALS2, or corresponding vector controls after which LGALS2 levels were analyzed via qPCR and Western immunoblotting. (D) Upregulating and silencing LGALS2 respectively impairs and enhances SCC-25 and SCC-9 cell proliferation. (E and F) EdU uptake assays were employed to quantify the proliferation of cells transfected as in (A). Scale bar: 100 μm. (G and H) Flow cytometry was used to assess the apoptotic death of cells transfected as in (A). Results from three repeated experiments are given as means ± standard deviation. *p < 0.05; **p < 0.01; ***p < 0.001. OE: Overexpression.
LGALS2 suppresses the invasivity and migration of OSCC cells through the JAK/STAT3 signaling axis
In Transwell assays, overexpressing LGALS2 was found to suppress the ability of SCC-25 and SCC-9 cells to migrate and invade through assay membranes (Figs. 3A–3D). In light of evidence suggesting the ability of LGALS2 to inhibit proliferation and survival in OSCC cells by suppressing JAK/STAT3 signaling, Western immunoblotting was next employed to quantify p-JAK2, JAK2, p-STAT3, and STAT3 levels within OSCC cells. LGALS2 overexpression was found to suppress levels of both STAT3 and JAK2 phosphorylation, whereas the opposite was observed upon LGALS2 knockdown (Fig. 3E and Suppl. Fig. S1). LGALS2 is therefore capable of suppressing OSCC progression at least in part via a JAK2/STAT3-dependent signaling mechanism.
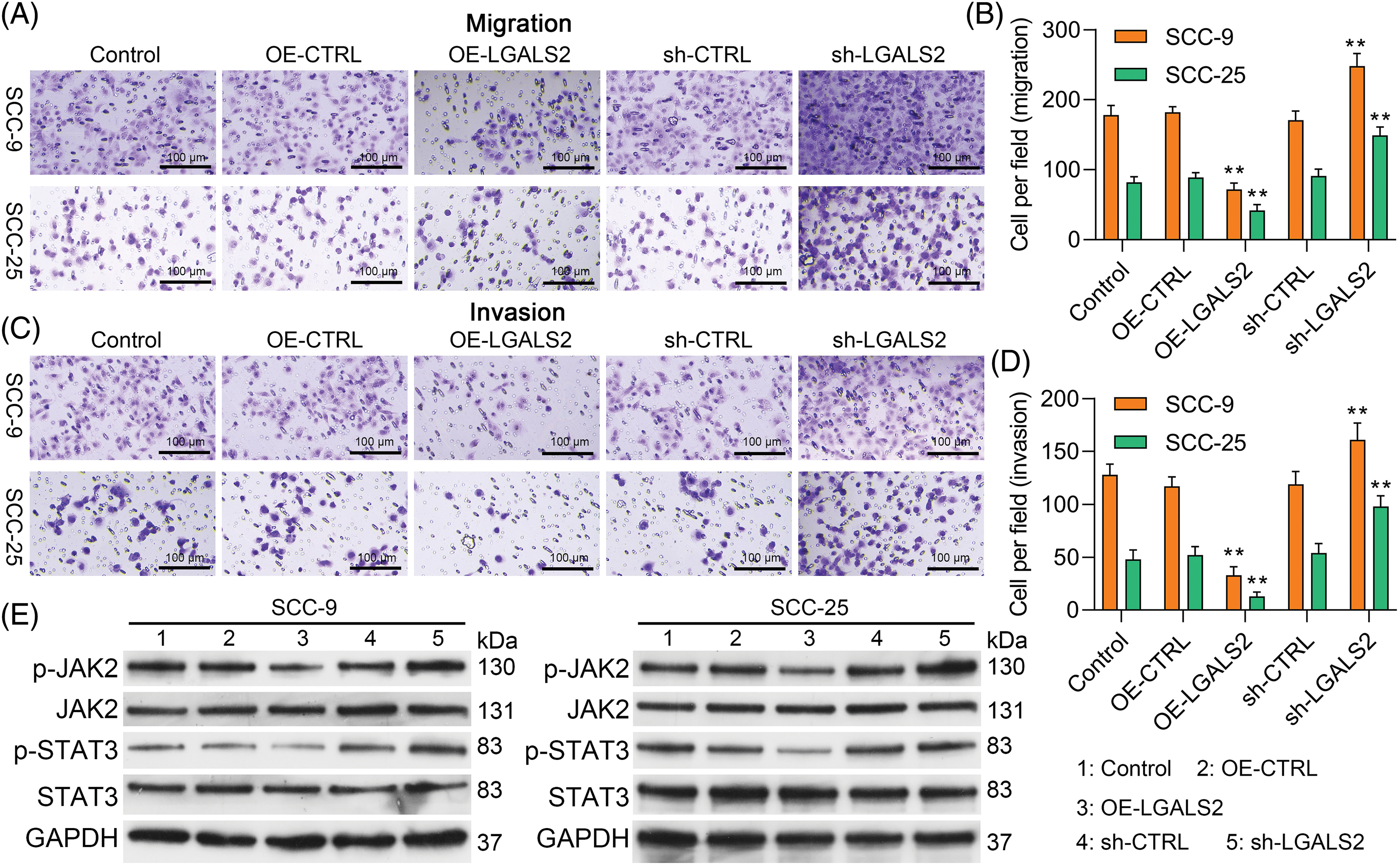
Figure 3: LGALS2 modulates JAK/STAT3 signaling to suppress migratory activity and invasivity in OSCC cells. (A–D) LGALS2 was found to suppress OSCC cell invasivity in a transwell assay. Representative 100× images and corresponding quantification are profiled (Scale bar: 100 μm). (E) Western blotting was used to detect p-JAK2, JAK2, p-STAT3, and STAT3 levels within the indicated cell lines following transfection with OE-CTRL, OE-LGALS2, sh-CTRL, and sh-LGALS2 constructs. GAPDH served as a normalization control for densitometric quantification efforts. Results are from three repeated experiments. **p < 0.01. OE: Overexpression.
JAK2/STAT3 pathway hyperactivation overcomes LGALS2-mediated inhibition of the proliferation of OSCC cells
Treatment with the 1 μM of α7 nAchR-JAK2-STAT3 agonist (dissolved in dimethylsulfoxide with concentration of 10 mM, HY-146066, MedChemExpress, Shanghai, China) to promote the hyperactivation of this signaling axis was found to partially overcome the inhibitory impact that LGALS2 was found to exert on the growth of OSCC cells (Figs. 4A–4C). Flow cytometry further confirmed that STAT3 activation in response to HY-146066 treatment was associated with the inhibition of apoptotic cell death relative to that of cells in the OE-LGALS2 group (Figs. 4D, 4E). Finally, Transwell assays were conducted in which STAT3 hyperactivation was found to overcome the inhibitory effects that LGALS2 had on the migration and invasivity of OSCC cells (Fig. 5). These findings further confirmed the ability of LGALS2 to suppress the progressive growth of OSCC through the consequent inhibition of the STAT3 signaling axis.
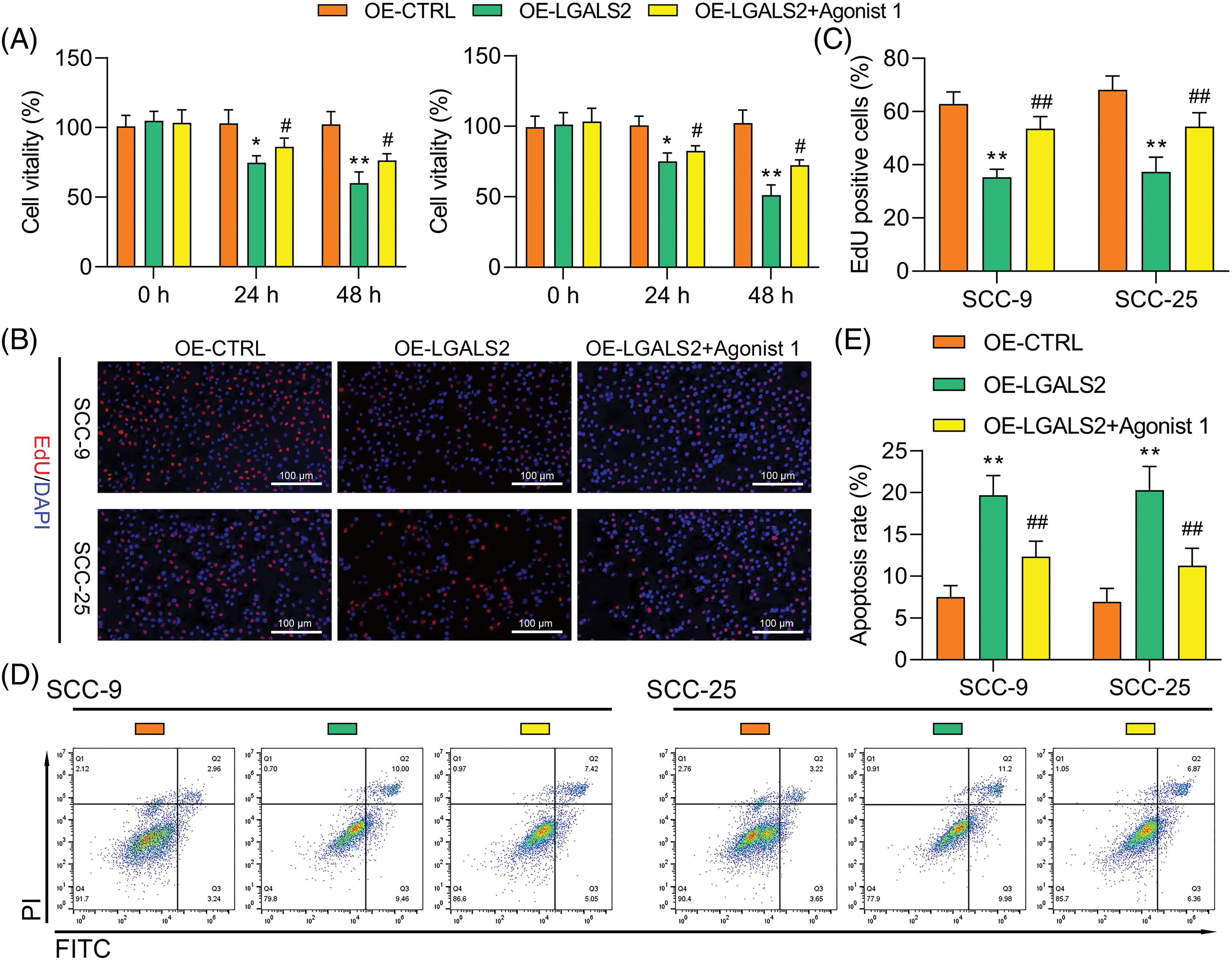
Figure 4: JAK2-STAT3 pathway activation partially reverses the ability of LGALS2 to suppress the growth of OSCC tumors. HY-146066 (50 ng/ml) was used to treat OSCC cells following LGALS2 overexpression plasmid transfection, after which these cells were evaluated through CCK-8 (A), EdU (B and C) and flow cytometry approaches (D and E). Results are from three repeated experiments. **p < 0.01 vs. OE-CTRL group; #p < 0.05, ##p < 0.01 vs. OE-LGALS2 group. OE: Overexpression.
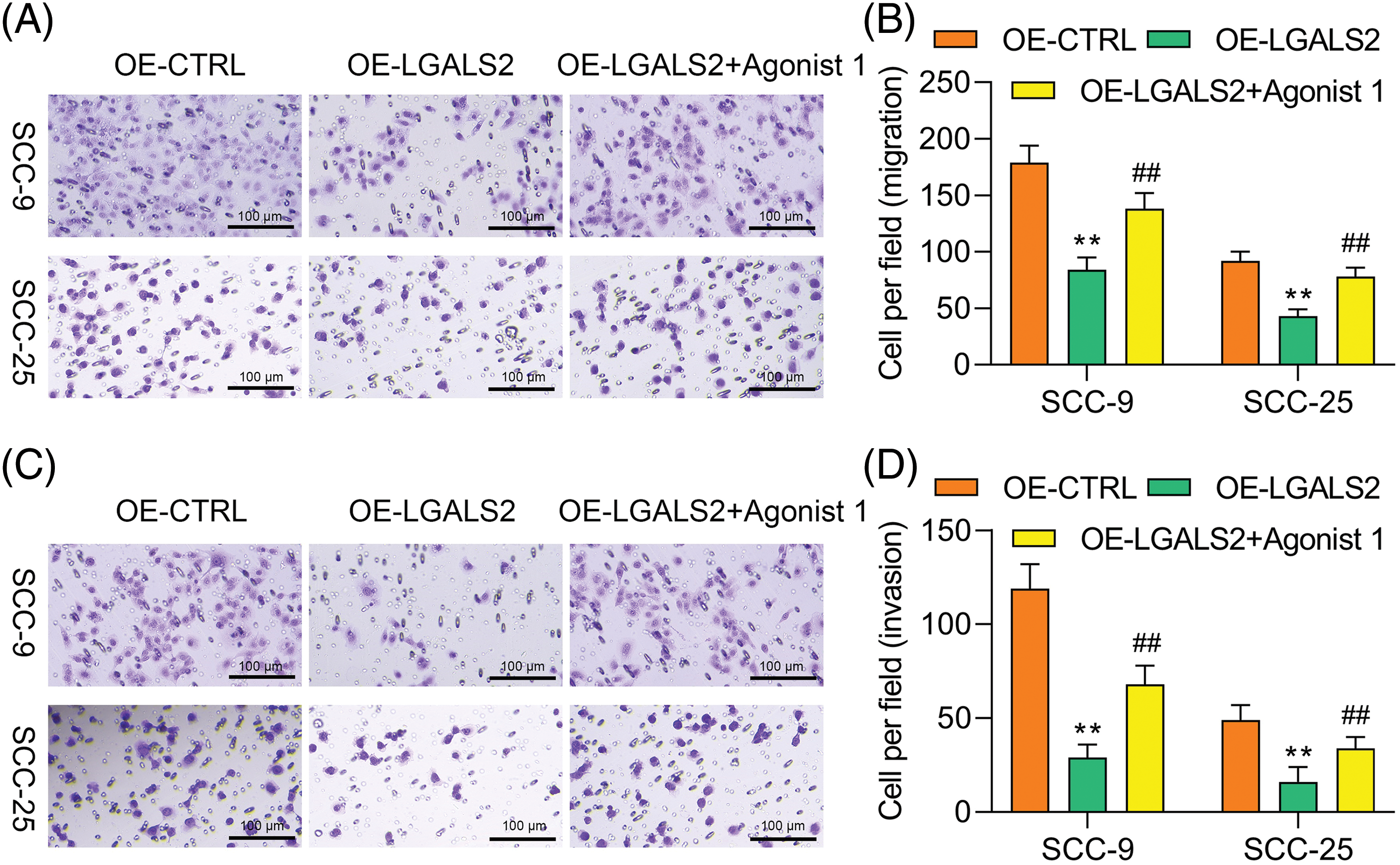
Figure 5: JAK2-STAT3 pathway activation overcomes the ability of LGALS2 to inhibit migratory and invasive activity in OSCC cells. HY-146066 (50 ng/ml) was used to treat OSCC cells following LGALS2 overexpression plasmid transfection, after which the ability of these cells to migrate (A and B) and invade (C and D) was assessed. Results are from three repeated experiments. Scale bar: 100 μm. **p < 0.01 vs. OE-CTRL group; ##p < 0.01 vs. OE-LGALS2 group. OE: Overexpression.
OSCC is characterized by high rates of metastatic progression, recurrence, and mortality. Owing to these factors and the absence of reliable treatments for most patients, OSCC continues to impose a profound burden on global public health [2,19]. Research focused on identifying key drivers of OSCC tumor metastasis and recurrence is essential in order to aid in the production of more reliable therapeutic tools for this disease.
LGALS proteins utilize conserved CRDs to bind to galactosidase-containing glycoproteins. A growing number of reports have identified LGALS proteins as key mediators of survival, inflammation, motility, and adhesion [5,7,20], with prior evidence highlighting the dysregulated expression of certain LGALS proteins in liver [21], bladder [22], prostate [22], thyroid [23], breast [24]. At a mechanistic level, LGALS proteins can interact with particular receptors to shape immunosuppressive activity by modulating cellular activation, differentiation, or costimulatory signaling [25,26]. Aberrant expression of certain LGALS proteins has been linked to worse prognostic outcomes, including clinical staging, tumor grade, and lymph node metastasis [27,28]. As the most heavily studied members of this family, LGALS1 and LGALS3 are known to be closely associated with cancer-related outcomes. Increased LGALS1 levels, for example, are related to worse ovarian cancer (OC) [29], gingival squamous cell cancer [30], nasopharyngeal cancer [31], and pancreatic ductal adenocarcinoma [32] patient outcomes. LGALS3, in contrast, exhibits variable tumor type-specific phenotypes. In their analysis, Ma et al. determined that anti-LGALS3 nanoparticles could effectively be used for drug delivery, thereby enhancing the concentrations of doxorubicin within thyroid cancer cells in vivo [33].
There have been past reports tying LGALS2 to gestational diabetes, which is a metabolic disorder of pregnancy [34]. With respect to its role in cancer, Jung et al. [32] found the downregulation of LGALS2 to be associated with gastric carcinoma lymph node progression and advanced clinical staging, suggesting that the loss of this protein may enable more robust tumorigenic activity. However, research regarding LGALS2 in human cancer remains limited. Lower LGALS2 levels have been shown to be correlated with superior overall survival outcomes in grade III breast cancer [17], and there have been similar findings in basal-like, luminal B, TP53-mutated, HER2-overexpressing, and wild-type breast cancer patients [35]. Chetry et al. [15] also determined that LGALS2 was predictive of more favorable outcomes for individuals with breast cancer. Despite this evidence that LGALS2 can serve as a cancer-related biomarker, its relevance in the context of OSCC has not previously been documented.
Systematic studies focused on the molecular pathways that drive the pathogenesis of OSCC have enabled the development of more effective therapeutic interventions for affected patients [36]. Signaling through the JAK/STAT3 pathway is one of the best-characterized mediators of protein synthesis growth, proliferative activity, metabolism, and the regulation of apoptosis [37]. Dysregulated receptor tyrosine kinase activation can provoke persistent JAK/STAT3 signaling conducive to enhanced vascularization and proliferative growth [38]. There is also clear evidence for the key role of such JAK/STAT3 signaling in the onset and progression of OSCC, with dysregulated pathway activity playing a role in changes in tumor cell sensitivity to specific therapeutic agents [39,40]. Additional research focused on the mechanistic interplay between LGALS2 and JAK/STAT3 signaling may inform efforts to treat patients with OSCC, as supported by the present findings indicating that activity in this pathway is closely associated with LGALS2 expression levels, sustaining the malignant growth and survival of OSCC cells.
Overall, the present results suggest that LGALS2 is capable of regulating JAK/STAT3 signaling to inhibit the progression of OSCC, consistent with a role for LGALS2 as a viable target for therapeutic intervention and prognostic biomarker for this cancer type.
There were several limitations in our present study. First, an in vivo study needs to be performed to confirm the role of LGALS2 in OSCC growth. Second, CRISPR/Cas9 has emerged as a powerful gene editing tool and has potential in the field of precision medicine, CRISPR/Cas9 could be used for knockout of LGALS2 to determine the potential role of LGALS2 in OSCC proliferation and migration. Finally, it is useful for high-throughput sequencing or gene microarray to analyze the down-stream signaling pathway of LGALS2.
Acknowledgement: None.
Funding Statement: The research is supported by grants from Key R&D Project of Science and Technology Foundation of Sichuan Province (2022YFS0290).
Author Contributions: Study conception and design: Li Xiao; data collection: Xinru Feng, Li Xiao; analysis and interpretation of results: Xinru Feng, Li Xiao; draft manuscript preparation: Li Xiao. All authors reviewed the results and approved the final version of the manuscript.
Availability of Data and Materials: The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
Ethics Approval: Informed consent was signed by each patient and the approval for conducting these experimental procedures was provided through the Ethics Committee of the Sichuan Provincial People’s Hospital (Lunshen (Yan)2022-50).
Conflicts of Interest: The authors declare that they have no conflicts of interest to report regarding the present study.
References
1. Chamoli A, Gosavi AS, Shirwadkar UP, Wangdale KV, Behera SK, Kurrey NK, et al. Overview of oral cavity squamous cell carcinoma: risk factors, mechanisms, and diagnostics. Oral Oncol. 2021;121:105451. [Google Scholar] [PubMed]
2. Siegel RL, Miller KD, Wagle NS, Jemal A. Cancer statistics, 2023. CA: Cancer J Clin. 2023;73(1):17–48. [Google Scholar] [PubMed]
3. Johnson DE, Burtness B, Leemans CR, Lui VWY, Bauman JE, Grandis JR. Head and neck squamous cell carcinoma. Nat Rev Dis Primers. 2020;6(1):92. [Google Scholar] [PubMed]
4. Vitorio JG, Duarte-Andrade FF, Dos Santos Fontes Pereira T, Fonseca FP, Amorim LSD, Martins-Chaves RR, et al. Metabolic landscape of oral squamous cell carcinoma. Metabolomics. 2020;16(10):105. [Google Scholar] [PubMed]
5. Xu WD, Huang Q, Huang AF. Emerging role of galectin family in inflammatory autoimmune diseases. Autoimmun Rev. 2021;20(7):102847. [Google Scholar] [PubMed]
6. Manero-Rupérez N, Martínez-Bosch N, Barranco LE, Visa L, Navarro P. The galectin family as molecular targets: hopes for defeating pancreatic cancer. Cells. 2020;9(3):689. [Google Scholar]
7. Shi Y, Tang D, Li X, Xie X, Ye Y, Wang L. Galectin family members: eemerging novel targets for lymphoma therapy? Front Oncol. 2022;12:889034. [Google Scholar] [PubMed]
8. Danguy A, Camby I, Kiss R. Galectins and cancer. Biochim Biophys Acta. 2002;1572(2–3):285–93. [Google Scholar] [PubMed]
9. Cai Y, Sun Z, Shao C, Wang Z, Li L. Role of galectin-3 in vascular calcification. Glycoconj J. 2023;40(2):149–58. [Google Scholar] [PubMed]
10. Shariq M, Quadir N, Alam A, Zarin S, Sheikh JA, Sharma N, et al. The exploitation of host autophagy and ubiquitin machinery by Mycobacterium tuberculosis in shaping immune responses and host defense during infection. Autophagy. 2023;19(1):3–23. [Google Scholar] [PubMed]
11. Ibrahim BA, Mohamed SH, Hassaan MMM, Sabbah NA. Associations of galectin-3 expression and LGALS-3 (rs4652) gene variant with coronary artery disease risk in diabetics. J Med Biochem. 2021;40(4):395–406. [Google Scholar] [PubMed]
12. Thijssen VL, Heusschen R, Caers J, Griffioen AW. Galectin expression in cancer diagnosis and prognosis: a systematic review. Biochim Biophys Acta. 2015;1855(2):235–47. [Google Scholar] [PubMed]
13. Morishita A, Oura K, Tadokoro T, Shi T, Fujita K, Tani J, et al. Galectin-9 in gastroenterological cancer. Int J Mol Sci. 2023;24(7):6174. [Google Scholar] [PubMed]
14. Marino KV, Cagnoni AJ, Croci DO, Rabinovich GA. Targeting galectin-driven regulatory circuits in cancer and fibrosis. Nat Rev Drug Discov. 2023;22(4):295–316. [Google Scholar] [PubMed]
15. Chetry M, Bhandari A, Feng R, Song X, Wang P, Lin J. Overexpression of galectin2 (LGALS2) predicts a better prognosis in human breast cancer. Am J Transl Res. 2022;14(4):2301–16. [Google Scholar] [PubMed]
16. Xu D, Guo L, Zhang S, Hou Q. LGALS2 suppresses the progression of papillary thyroid carcinoma by regulating the PI3K/AKT pathway. Gland Surg. 2022;11(9):1518–28. [Google Scholar] [PubMed]
17. Ji P, Gong Y, Jin ML, Wu HL, Guo LW, Pei YC, et al. In vivo multidimensional CRISPR screens identify Lgals2 as an immunotherapy target in triple-negative breast cancer. Sci Adv. 2022;8(26):eabl8247. [Google Scholar] [PubMed]
18. Li H, Zhao L, Lau YS, Zhang C, Han R. Genome-wide CRISPR screen identifies LGALS2 as an oxidative stress-responsive gene with an inhibitory function on colon tumor growth. Oncogene. 2021;40(1):177–88. [Google Scholar] [PubMed]
19. Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA: Cancer J Clin. 2021;71(3):209–49. [Google Scholar] [PubMed]
20. Rabinovich GA, Liu FT, Hirashima M, Anderson A. An emerging role for galectins in tuning the immune response: lessons from experimental models of inflammatory disease, autoimmunity and cancer. Scand J Immunol. 2007;66(2–3):143–58. [Google Scholar] [PubMed]
21. Setayesh T, Colquhoun SD, Wan YY. Overexpression of galectin-1 and galectin-3 in hepatocellular carcinoma. Liver Res. 2020;4(4):173–9. [Google Scholar] [PubMed]
22. Kaur M, Kaur T, Kamboj SS, Singh J. Roles of galectin-7 in cancer. Asian Pac J Cancer Prev. 2016;17(2):455–61. [Google Scholar] [PubMed]
23. Li J, Vasilyeva E, Wiseman SM. Beyond immunohistochemistry and immunocytochemistry: a current perspective on galectin-3 and thyroid cancer. Expert Rev Anticancer Ther. 2019;19(12):1017–27. [Google Scholar] [PubMed]
24. Yasinska IM, Sakhnevych SS, Pavlova L, Teo Hansen Selno A, Teuscher Abeleira AM, Benlaouer O, et al. The Tim-3-galectin-9 pathway and its regulatory mechanisms in human breast cancer. Front Immunol. 2019;10:1594. [Google Scholar] [PubMed]
25. Ledeen RW, Kopitz J, Abad-Rodriguez J, Gabius HJ. Glycan chains of gangliosides: functional ligands for tissue lectins (Siglecs/Galectins). Prog Mol Biol Transl Sci. 2018;156:289–324. [Google Scholar] [PubMed]
26. O’Sullivan JA, Carroll DJ, Bochner BS. Glycobiology of eosinophilic inflammation: ccontributions of siglecs, glycans, and other glycan-binding proteins. Front Med. 2017;4:116. [Google Scholar]
27. Dong R, Zhang M, Hu Q, Zheng S, Soh A, Zheng Y, et al. Galectin-3 as a novel biomarker for disease diagnosis and a target for therapy (Review). Int J Mol Med. 2018;41(2):599–614. [Google Scholar] [PubMed]
28. Pereira AR, Menezes Falcao L. Galectin-3, a prognostic marker--and a therapeutic target? Rev Port Cardiol. 2015;34(3):201–8. [Google Scholar] [PubMed]
29. Chen L, Yao Y, Sun L, Zhou J, Liu J, Wang J, et al. Clinical implication of the serum galectin-1 expression in epithelial ovarian cancer patients. J Ovarian Res. 2015;8(1):78. [Google Scholar] [PubMed]
30. Noda Y, Kishino M, Sato S, Hirose K, Sakai M, Fukuda Y, et al. Galectin-1 expression is associated with tumour immunity and prognosis in gingival squamous cell carcinoma. J Clin Pathol. 2017;70(2):126–33. [Google Scholar] [PubMed]
31. Chang SL, Li CF, Lin C, Lin YS. Galectin-1 overexpression in nasopharyngeal carcinoma: effect on survival. Acta Otolaryngol. 2014;134(5):536–42. [Google Scholar] [PubMed]
32. Zhao X, Li H, Lyu S, Zhai J, Ji Z, Zhang Z, et al. Single-cell transcriptomics reveals heterogeneous progression and EGFR activation in pancreatic adenosquamous carcinoma. Int J Biol Sci. 2021;17(10):2590–605. [Google Scholar] [PubMed]
33. Ma X, Li X, Shi J, Yao M, Zhang X, Hou R, et al. Host-guest polypyrrole nanocomplex for three-stimuli-responsive drug delivery and imaging-guided chemo-photothermal synergetic therapy of refractory thyroid cancer. Adv Healthc Mater. 2019;8(17):e1900661. [Google Scholar] [PubMed]
34. Hepp P, Unverdorben L, Hutter S, Kuhn C, Ditsch N, Groß E, et al. Placental galectin-2 expression in gestational diabetes: a systematic, histological analysis. Int J Mol Sci. 2020;21(7):2404. [Google Scholar] [PubMed]
35. Barrow H, Guo X, Wandall HH, Pedersen JW, Fu B, Zhao Q, et al. Serum galectin-2, -4, and -8 are greatly increased in colon and breast cancer patients and promote cancer cell adhesion to blood vascular endothelium. Clin Cancer Res. 2011;17(22):7035–46. [Google Scholar] [PubMed]
36. Xing M. Molecular pathogenesis and mechanisms of thyroid cancer. Nat Rev Cancer. 2013;13(3):184–99. [Google Scholar] [PubMed]
37. Johnson DE, O’Keefe RA, Grandis JR. Targeting the IL-6/JAK/STAT3 signalling axis in cancer. Nat Rev Clin Oncol. 2018;15(4):234–48. [Google Scholar] [PubMed]
38. Jin W. Role of JAK/STAT3 signaling in the regulation of metastasis, the transition of cancer stem cells, and chemoresistance of cancer by epithelial-mesenchymal transition. Cells. 2020;9(1):217. [Google Scholar] [PubMed]
39. Huang C, Li H, Zhou L, Li D. Circ_0005050 promotes the proliferation of oral squamous cell carcinoma and inhibits the apoptosis by activating JAK/STAT3 signaling pathway. Pathol Res Pract. 2022;238:154058. [Google Scholar] [PubMed]
40. Liu C, Zhou J, Zhang S, Fu J, Li Y, Hao Y, et al. Mesenchymal stem cells-derived IL-6 promotes invasion and metastasis of oral squamous cell carcinoma via JAK-STAT3 signalling. Oral Dis. 2023. doi:https://doi.org/10.1111/odi.14617. [Google Scholar] [PubMed] [CrossRef]
Supplementary Materials
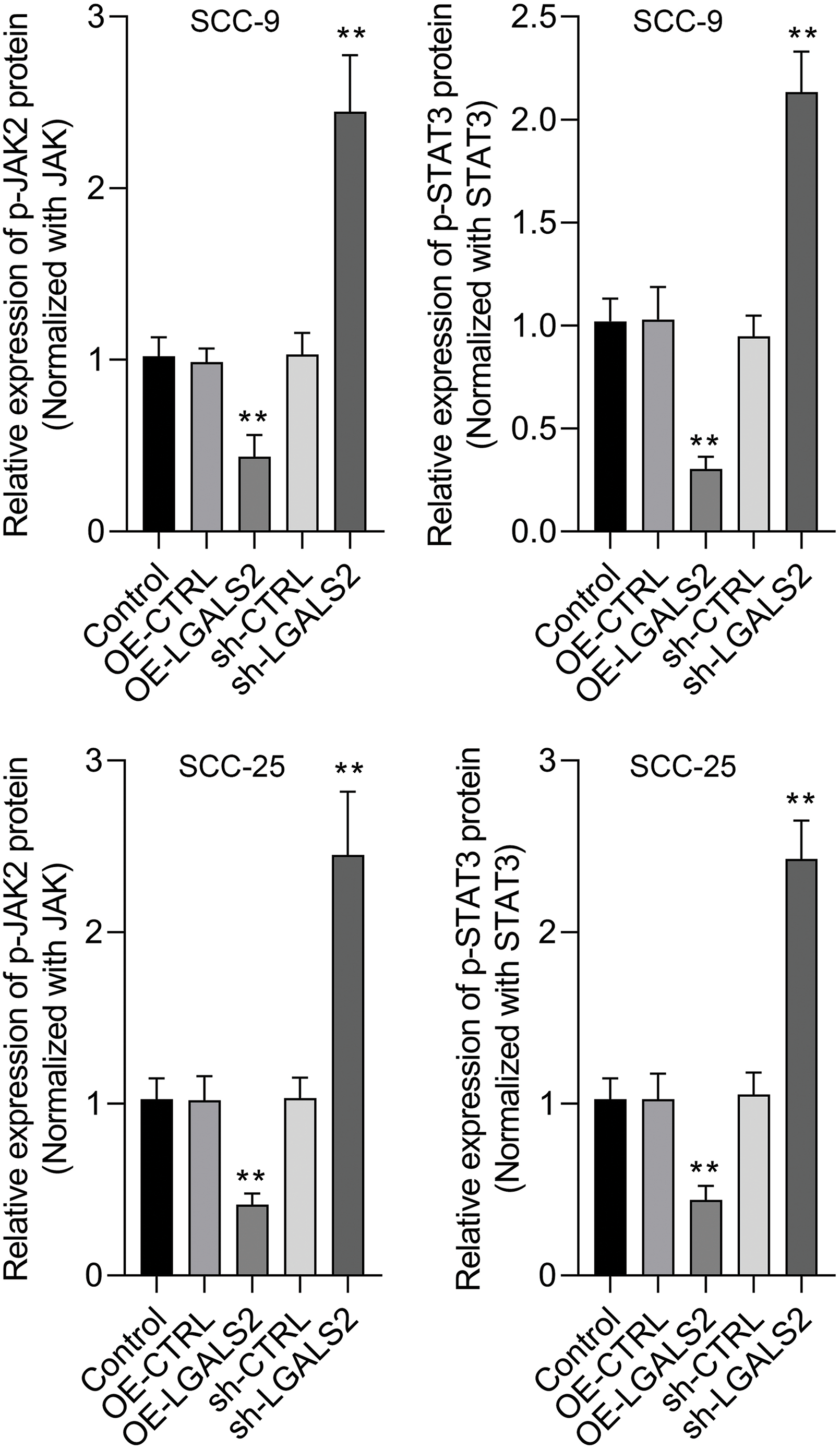
Figure S1: The protein expression levels were quantitatively analyzed. Overexpression of LGALS2 reduced the phosphorylation level of JAK2 and STAT3, while knockdown of LGALS2 induced the phosphorylation level of JAK2 and STAT3. **p < 0.01.
Cite This Article
 Copyright © 2024 The Author(s). Published by Tech Science Press.
Copyright © 2024 The Author(s). Published by Tech Science Press.This work is licensed under a Creative Commons Attribution 4.0 International License , which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.


 Submit a Paper
Submit a Paper Propose a Special lssue
Propose a Special lssue View Full Text
View Full Text Download PDF
Download PDF Downloads
Downloads
 Citation Tools
Citation Tools