 Open Access
Open Access
ARTICLE
ETS1 Regulates Endothelial AQP1 Expression via Interaction with MEF2C in Endothelial Cells
School of Laboratory Medicine, Jilin Medical University, Jilin, China
* Corresponding Author: Yong Jiang. Email:
BIOCELL 2026, 50(4), 10 https://doi.org/10.32604/biocell.2026.075982
Received 12 November 2025; Accepted 20 January 2026; Issue published 21 April 2026
Abstract
Background: Aquaporin 1 (AQP1) plays a key role in myocardial ischemia-reperfusion (I/R) injury. This study aimed to elucidate the mechanisms by which erythroblast transformation-specific 1 (ETS1) and myocyte enhancer factor 2C (MEF2C) regulated AQP1 transcription. Methods: Human umbilical vein endothelial cells (HUVECs) and rats with coronary heart disease were employed for in vitro and in vivo experiments, respectively. Expressions of ETS1, MEF2C, and AQP1 were analyzed by western blotting and quantitative reverse transcription polymerase chain reaction (qRT-PCR). Chromatin immunoprecipitation (ChIP) and co-immunoprecipitation (Co-IP) assays were performed to confirm the interactions between ETS1 and MEF2C. Scratch wound healing and transwell assays were performed to assess the migration of HUVECs. Tube formation was conducted to assess the angiogenesis of HUVECs. HE and Masson staining were performed to assess the pathological changes and fibrosis of cardiac tissues; Immunofluorescence of platelet endothelial cell adhesion molecule (CD31) and AQP1 was performed to assess vascular endothelial cell dysfunction in cardiac tissues of rats with coronary heart disease. Results: ETS1 overexpression upregulated both MEF2C and AQP1 expressions, while ETS1 silencing induced the opposite results. ETS1 and MEF2C interacted with each other and bound to the proximal promoter of AQP1. MEF2C synergized with ETS1 to promote the migration and tube formation of HUVECs, which were abolished by AQP1 silencing. I/R injury caused structural damage and fibrosis of cardiac tissues in rats, which were relieved by ETS1 and MEF2C silencing. Also, the increased AQP1 and CD31 levels in cardiac tissues of rats induced by I/R injury were reversed by ETS1 and MEF2C knockdown. Conclusion: These results highlight the ETS1/MEF2C/AQP1 pathway as a potential therapeutic target for endothelial and vascular disorders.Graphic Abstract
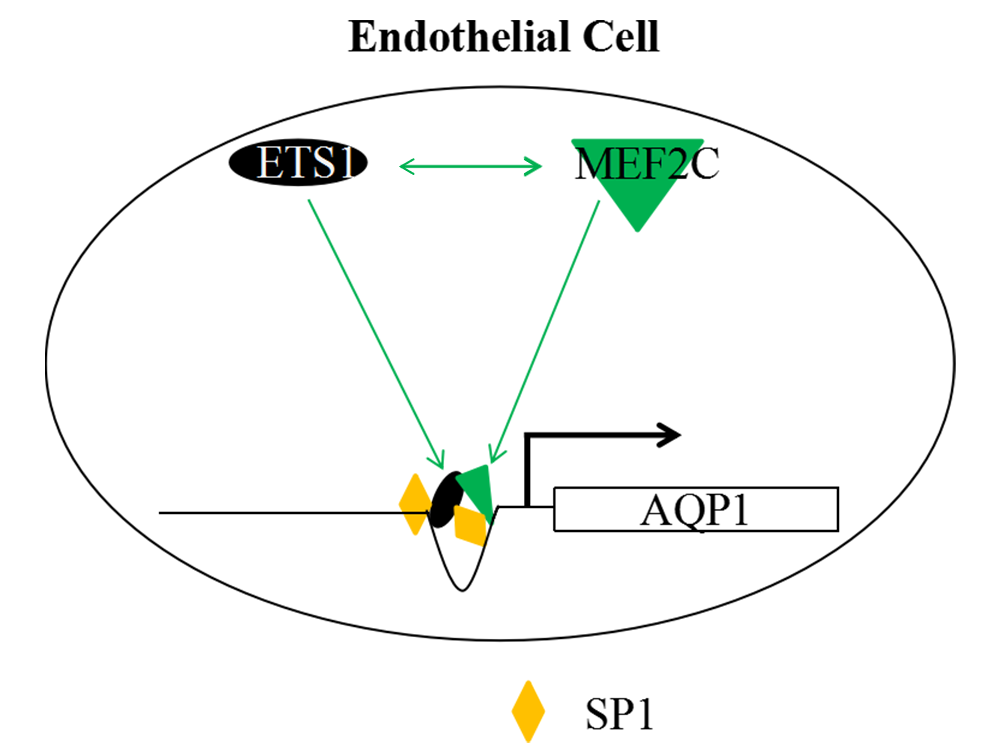
Keywords
Aquaporin 1 (AQP1), a member of the aquaporin family, plays a key role in controlling transcellular water movement in various tissues of the whole body [1]. Previous studies have demonstrated that AQP1 expression is increased in microvessels of the endometrium [2], lung [3], heart [4], and tumors [5,6]. AQP1 levels are up-regulated in most microvascular endothelial cells [7] with activities promoting angiogenesis [8]. Inhibition of AQP1 expression enhances the endothelial permeability of human pulmonary microvascular endothelial cells by repressing vascular endothelial (VE)-cadherin, epithelial sodium channel (ENaC), and Na-K ATPase [9]. AQP1 also exerts a vital role in regulating cellular response to osmotic stress and tissue water permeability [10].
Myocyte enhancer factor 2C (MEF2C) is a transcription factor belonging to the MEF2 family that exerts an essential impact on the differentiation, morphogenesis, and maintenance of muscle, heart, bone, blood vessels, nerves, blood, and the immune system by modulating differentiation, proliferation, migration, shape, and metabolism [11,12]. MEF2C is necessary for neuronal differentiation and appropriate neuronal distribution [13], craniofacial development [14], postnatal maturation of skeletal muscle and preservation of sarcomere integrity [15], and B-cell proliferation and survival are stimulated with antigen receptors [16]. During vascular development, MEF2C modulates endothelial cell organization and smooth muscle cell differentiation [17]. Moreover, MEF2C promotes human umbilical vein endothelial cell (HUVEC) tube formation by regulating AQP1 [18]. Also, MEF2C cooperated with SP1 to activate AQP1 transcription by binding to its proximal promoter in HUVECs [19].
Erythroblast transformation specific 1 (ETS1), a member of the ETS family of transcription factors, was originally characterized as an oncogene but was later found to have functional effects in normal cells [20]. Although most studies have explored the role of ETS1 in cancer [21,22], the effects of ETS1 on immunity [23], autoimmunity [24], and angiogenesis [25] have also been verified. Endothelial loss of ETS1 inhibits coronary vascular development [26], demonstrating the importance of endocardial ETS1 in cardiac development. A previous study has revealed that MEF2C serves as a direct transcriptional target of ETS factors via an evolutionarily conserved transcriptional enhancer, suggesting a direct link between MEF2C and ETS1 during endothelial cell development [27]. Therefore, this study aims to explore whether ETS1 interacts with MEF2C to modulate endothelial health by regulating AQP1 expression.
2.1 Cell Culture and Transfection
HUVECs were obtained from ATCC (Manassas, VA, USA) and cultured within complete endothelial medium (F-12K complemented with 10% FBS, 0.1 mg/mL heparin, and 30 μg/mL endothelial cell growth supplement (ECGS), all from Thermo Fisher Scientific, Waltham, MA, USA) at 37°C with 5% CO2. The HUVECs used in the experiments were between passages 25 and 35. All experiments were performed with mycoplasma-free HUVECs. Additionally, HUVECs were examined for short tandem repeats and were not contaminated with other cells. Subsequently, HUVECs were transfected with different vectors using Lipofectamine 3000 (cat. no. L3000150, Thermo Fisher Scientific).
The AQP1 promoter (1000 bp, pAQP1) was cloned into the pGL3-basic vector and co-transfected with pRL-TK (Promega E2241; Promega, Madison, WI, USA), in combination with ETS1 or MEF2C overexpression vector (pcDNA3.1-ETS1/MEF2C) or shRNA targeting ETS1 or MEF2C (pGPU6/GFP/Neo-shETS1/shMEF2C) into HUVECs via Lipofectamine 3000 (Thermo Fisher Scientific). HUVECs were harvested 48 h post-transfection and washed with 0.1 mol/L phosphate-buffered saline (PBS, PH7.4), and luciferase activity was measured using a dual-luciferase reporter assay system (cat. no. E1910, Promega, Madison, WI, USA).
2.3 Quantitative Reverse Transcription Polymerase Chain Reaction (qRT-PCR)
Total RNA was extracted from HUVECs and cardiac tissues of Sprague-Dawley (SD) rats using a RNeasy Plus Micro Kit (cat. no. 74034-50T, NUO YANG SHENG WU, Hangzhou, China) according to the manufacturer’s instructions. cDNA was prepared by reverse transcription using a TRUEscript One Step RT-PCR kit (PC3401, Aidlab Biotechnology Co., Ltd., Beijing, China) in line with specific instructions, and served as the starting material for qRT-PCR using the Step OnePlus Real-Time PCR system (Life Technologies Corp, Carlsbad, CA, USA). Primer Premier software 5.0 (Sangon Biotech Co., Ltd., Shanghai, China) was used to design all primer sequences. The 2−ΔΔCT method was normalized to GAPDH.
The following primers were used in the study:
AQP1(rat) forward: 5′-CTCTTCTGGAGGGCAGTGGTGG-3′
AQP1(rat) reverse: 5′-CCTGGACCGTCGTCTGGTTGTT-3′
MEF2C (rat) forward: 5′-ATTCGGACTGATGAAGAAG GCT-3′
MEF2C (rat) reverse: 5′-TTGTTGGTGCTGTTGAAGATGA-3′
ETS1 (rat) forward: 5′-CTCAAGCCGACTCTCACCATC-3′
ETS1 (rat) reverse: 5′-AACTCATTCACAGCCCACATC-3′
GAPDH (rat) forward: 5′-ACAGCAACAGGGTGGTGGAC-3′
GAPDH (rat) reverse: 5′-TTTGAGGGTGCAGCGAACTT-3′
AQP1(human) forward: 5′-GGCCTGGGCATCGAGATCAT-3′
AQP1(human) reverse: 5′-GGGCTACAGAGAGGCCGATG-3′
MEF2C (human) forward: 5′-GCGTGCTGTGTGACTGTGAG-3′
MEF2C (human) reverse: 5′-CATGCGGCTCGTTGTACTCC-3′
ETS1 (human) forward: 5′-CTGCAGCGTGTTCCCTCCTA-3′
ETS1 (human) reverse: 5′-CGGTCCCGCACATAGTCCTT-3′
GAPDH (human) forward: 5′-TGGGTGTGAACCATGAGAAGT-3′
GAPDH (human) reverse: 5′-TGAGTCCTTCCACGATACCAA-3′
HUVECs and cardiac tissues of SD rats were lysed via RIPA lysis buffer (cat. no. G2008, Servicebio Biotechnology Co., Ltd., Wuhan, China), and protein content was determined using a bicinchoninic acid (BCA) kit (cat. no. P0011, Beyotime Biotechnology, Shanghai, China). Proteins were separated using 10% SDS-polyacrylamide gels and then transferred onto a nitrocellulose membrane. The membrane was blocked with 5% non-fat milk for 1 h at room temperature, and incubated with the following primary antibodies overnight at 4°C: anti-AQP1 (1:5000, 20333-1-AP, Proteintech, Chicago, IL, USA), anti-MEF2C (1:500, 18290-1-AP, Proteintech), anti-ETS1 (1:5000, 66598-1-Ig, Proteintech), and anti-GAPDH (1:500, 13937-1-AP, Proteintech). GAPDH served as the loading control. Horseradish peroxidase (HRP)-labeled secondary antibodies (1:4000, SA00001-15/17, Proteintech) were used as secondary antibodies. Finally, an enhanced chemiluminescence kit (cat. no. WBKlS0100, Millipore, Burlington, MA, USA) was used to observe the protein blots, and ImageJ software version 4.3 (National Institutes of Health [NIH], Bethesda, MD, USA) was used for quantification.
2.5 Chromatin Immunoprecipitation (ChIP) Assay
ChIP assay was performed using a ChIP Assay Kit (cat. no. P2078, Beyotime, Shanghai, China) according to the manufacturer’s instructions. Briefly, formaldehyde (37%) was added to the HUVEC culture medium at a final concentration of 1%, and incubated at room temperature for 10 min. The digested chromatin was treated with an ultrasonic crusher (VCX500; Sonics, Newtown, CT, USA) for 3 min, ultrasonicated for 5 s, and stopped for 25 s. Chromatin DNA was fragmented into 100–200 bp fragments. Subsequently, 25 μg DNA was incubated with anti-ETS1 (5 μg, 1:2000, 12118-1-AP, Proteintech), anti-MEF2C (5 μg, 1:500, 18290-1-AP, Proteintech) or mouse IgG (1 mg/mL, ab18413, Abcam, Cambridge, UK) at 4°C for 2 h before adding protein A/G beads (cat. no. P2078, Beyotime, Shanghai, China) to each sample and rotating at 4°C overnight using a rotary instrument (80 rpm, VSZD-IIIAB, Woshin, Wuxi, China). The immunocomplexes were collected using a protein G agarose suspension and washed successively with low-salt, high-salt, and LiCl buffer once each, and TE buffer twice. Protein G was removed using the prepared elution buffer (10%–20% SDS, 20 mm NaHCO3, and 170 mL of sterilized water per 200 mL of buffer). The complexes were incubated at 65°C for 5 h to denature the cross-linking. The DNA was purified using a PCR purification kit (cat. no. D4013, Zymo Research, Irvine, CA, USA) and amplified by PCR with the following primers: forward (5′-gaggtctgtgtggtgt-3′)/reverse (5′-tcaattccctctgagag-3′) for the MEF2C binding site and forward (5′-gggagagagagagaga-3′)/reverse (5′-acaccacacagacctc-3′) for the ETS1 binding site. PCR amplification conditions: 95°C for 5 min, (94°C for 40 s, 53°C for 60 s, 72°C for 60 s) for 30 cycles, and finally 72°C for 10 min.
2.6 Co-Immunoprecipitation (Co-IP) Assay
HUVECs were cleaved into proteins using an IP lysate (cat. no. G2008; Servicebio, Wuhan, China). Proteins were incubated with each primary antibody or mouse IgG at 4°C for 2 h before adding protein A/G beads to each sample and rotating at 4°C overnight. The immunocomplexes were then washed, eluted, and reverse cross-linked. The proteins were extracted using 12% SDS-PAGE and then transferred to PVDF membranes (Millipore, Burlington, MA, USA). The membranes were incubated using 5% skimmed milk, followed by overnight incubation with primary antibodies under 4°C. After rinsing, the membranes were incubated for 1 h with an HRP-labeled secondary antibody (1:4000, SA00004-10, Proteintech, Wuhan, China) at ambient temperature. Finally, an enhanced chemiluminescence kit (cat. no. WBKlS0100; Millipore, Burlington, MA, USA) was used to observe the protein blots, and ImageJ software (NIH, version 4.3) was used for quantification. The primary antibodies used were anti-AQP1 (1:5000, 20333-1-AP, Proteintech), anti-MEF2C (1:500, 10056-1-AP, Proteintech), anti-ETS1 (1:5000, 66598-1-Ig, Proteintech), and anti-GAPDH (1:500, 13937-1-AP, Proteintech), with GAPDH as the loading control.
HUVECs were cultured at the appropriate density (5 × 105/well), and a wound was scratched with a 10 μL pipette gun when the cells reached 80% confluence and then cultured in serum-free medium in the incubator with 5% CO2 at 37°C. Photographs were taken at 0 and 48 h, respectively. An inverted microscope (Olympus BX63, Tokyo, Japan) was used to observe the cells at 0 and 48 h (200×). The migration rate of the cells was calculated using the following equation:
After Matrigel was liquefied at 4°C overnight, a 24-well plate was placed on ice for pre-cooling. Subsequently, 50 μL of Matrigel was added to each well, and the plate was placed flat and stationary at 4°C for 10 min. The plate with the matrix gel was then placed in the incubator with 5% CO2 at 37°C for 30 min. The solidified Matrigel was removed, and HUVECs (3 × 105/well) were inoculated into a plate pre-coated with Matrigel and incubated for 24 h. Subsequently, cells were incubated with 20 μm calcein (cat. no. Y271510, Beyotime, Shanghai, China) for 20 min, imaged via a fluorescence microscope (IX71, OLYMPUS, Tokyo, Japan), and the number of lumens was counted using ImageJ software.
2.9 Rat Coronary Heart Disease Model Induced by High-Fat Diet and Transient Ischemia/Reperfusion (I/R)
30 adult male SD rats (6 weeks, 180~220 g) were purchased from Cavens Laboratory Animal Co. (Changzhou, China) and acclimated for a week. The rats were allowed to access water and food ad libitum, housed under suitable temperature (22 ± 2°C) and humidity (65% ± 5%) with a 12/12-h light/dark cycle. The rats were fed a high-fat diet with 45% energy from lipids and 17% energy from sucrose (D12451 Research Diets, New Brunswick, NJ, USA) for 16 weeks.
Next, ligation of the left anterior descending artery (LAD) was performed. Anesthesia was induced using 5% isoflurane and maintained using 2% isoflurane. The rats were intubated with a 20-G IV catheter through the mouth and connected to a ventilator (KW-10; KEW BASIS, Nanjing, China). The chest was incised in the left lateral intercostal space between the 4th and 5th ribs. Ligation was performed using an 8-0 polypropylene suture on the same segment as the LAD. The suture was removed 30 min later, and the opening was closed. Rats were randomly divided into six groups (n = 5): NC, shETS1, shMEF2C, shETS1+shMEF2C, Model, Model+shETS1+shMEF2C. For shRNA delivery, AAV9-shETS1 or AAV9-MEF2C (Gene Pharma, Suzhou, China) was injected into the tail vein 24 h prior to LAD ligation. Rats were euthanized 7 days after LAD ligation following anesthesia with 5% isoflurane. Heart tissues were harvested for subsequent experiments. This study adhered to the recommendations outlined in ARRIVE Guidelines 2.0, and the experiments were approved by the Institutional Animal Care and Usage Committee of Jilin Medical University (JMU2020-0037).
2.10 Hematoxylin-Eosin (HE) and Masson Staining
After fixation with 4% formaldehyde, the cardiac tissues were embedded in paraffin and cut into 5-μm sections. Pathological changes in the sections were assessed using an HE staining kit (G1120, Solarbio, Beijing, China), and cardiac fibrosis was assessed using a Masson staining kit (G1340, Solarbio, Beijing, China). Images were captured using a light microscope (OLYMPUS BX63; Tokyo, Japan).
2.11 Immunofluorescence of AQP1 and CD31
Briefly, cardiac sections were dehydrated and immersed in a PBS blocking solution containing 1.5% normal goat serum and 0.2% Triton X-100. The sections were then incubated overnight with AQP1 antibody (1:1000, ab300463, Abcam) and CD31 antibody (1:1000, ab9498, Abcam) at 4°C, and then incubated with FITC-labeled goat anti-rabbit IgG (1:100, SA00003-1, Proteintech) or Cy3-labeled secondary antibody (1:200, SA00009-2, Proteintech) at room temperature for 1 h. Then, sections were incubated with 5 μg/mL Hoechst 33258 (C1017, Beyotime) for 5 min in the dark, and washed with 0.1% phosphate buffered saline Tween (PBST) for 4 times. Finally, the samples were incubated in a humid chamber for 15 min under light-protected conditions, and images were captured using a light microscope (OLYMPUS BX63; Tokyo, Japan).
All experiments were repeated three times or more. Data were analyzed using GraphPad Prism 5.0 (GraphPad Software, Dotmatics, Waltham, MA, USA) and are presented as mean ± SD. Differences between the groups were compared by ANOVA and Tukey’s POC host analysis. Statistical significance was set at p < 0.05.
3.1 AQP1 Expression in HUVECs Was Regulated by ETS1 and MEF2C
ETS1 activates MEF2C in the early stage of vasculogenesis and thereafter vascular development. Furthermore, MEF2C is a critical regulator of endothelial AQP1 expression [19,28]. Therefore, we examined whether ETS1 regulated AQP1 expression in endothelial cells. As shown in Fig. 1A, AQP1 and MEF2C protein levels were correlated with ETS1 levels. Notably, ETS1 overexpression significantly increased AQP1 and MEF2C protein levels, whereas silencing ETS1 markedly reduced their protein levels (Fig. 1A–D). Similarly, ectopic ETS1 promoted, whereas silencing ETS1 inhibited the transcriptional activity of the AQP1 promoter (Fig. 1E). Interestingly, silencing MEF2C dampened the ETS1-induced increase in AQP1 promoter activity (Fig. 1E).

Figure 1: ETS1 upregulated AQP1 expression. (A–D) HUVECs were transfected with an ETS1 overexpressing or shRNA silencing vector, and specified protein levels were detected by western blotting. (E) HUVECs were co-transfected with pAQP1 and pcDNA3.1-ETS1/MEF2C or pGPU6/GFP/Neo-shETS1/shMEF2C, and luciferase activity was analyzed to determine the regulation of ETS1/MEF2C on pAQP1. *p < 0.05, **p < 0.01.
3.2 ETS1 and MEF2C Cooperatively Regulated AQP1 Transcription in Endothelial Cells
Next, we investigated how ETS1 and MEF2C regulated AQP1 transcription. SP1 was a widely existing transcription factor that bound to the GC-rich sequences in the promoter region of genes, thereby regulating gene expression. It has been reported that SP1 regulates AQP1 expression as a transcription factor [29]. As expected, the protein complex containing ETS1, MEF2C, and SP1 was precipitated using antibodies against ETS1 or MEF2C (Fig. 2A). Moreover, both ETS1 (Fig. 2C) and MEF2C (Fig. 2B) antibodies enriched AQP1 promoter sequences containing either ETS1- or MEF2C-binding site.

Figure 2: ETS1 and MEF2C cooperatively activated AQP1 transcription. (A) ETS1 and MEF2C physically interacted with each other. Immunocomplexes were detected by western blotting using antibodies against ETS1 and MEF2C. The AQP1 promoter was chromatin-immunoprecipitated using antibodies against MEF2C (B) or ETS1 (C) and PCR with primers amplifying either ETS1- or MEF2C-binding site.
3.3 ETS1 Enhanced Endothelial Cell Migration and Tube Formation through AQP1
AQP1 has been shown to be critical for the biological functions of endothelial cells, including migration and angiogenesis [30]. Therefore, we next examined whether the regulation of AQP1 expression by ETS1 affected these processes. ETS1 overexpression significantly increased the motility of HUVECs, which was similar to AQP1 and MEF2C overexpression (Fig. 3A,B,D,E). Silencing MEF2C partially inhibited ETS1-induced HUVEC migration, whereas MEF2C synergized with ETS1 to promote cell migration (Fig. 3A,B,D,E).

Figure 3: ETS1 modulates the functions of HUVECs through AQP1. Migration of HUVECs was assessed by scratch wound healing (50 μm, (A,D)) and transwell migration (20 μm, (B,E)) assays. (C,F) Tube formation assay was performed to assess the angiogenesis of HUVECs (50 μm). *p < 0.05 compared to pcDNA3.1 (control). *p < 0.05, **p < 0.01.
Both ETS1 and MEF2C strongly promoted angiogenesis in HUVECs, and the increase in tube formation was more prominent when ETS1 and MEF2C were overexpressed simultaneously (Fig. 3C,F). AQP1 knockdown strongly inhibited HUVEC lumen formation (Fig. 3C,F).
3.4 Silencing ETS1 and MEF2C Relieved I/R-Induced Cardiac Damage
AQP1 is an intrinsic membrane protein located on the cell membrane that mediates water flux between blood arteries and the interstitium and plays a significant role in myocardial I/R injury [31]. The impact of ETS1 and MEF2C regulation of AQP1 on I/R-induced cardiac damage was the subject of our further research. Transient LAD ligation induced the disarrangement of a large number of cardiomyocytes with shrinkage or disappearance of some nuclei, and the interstitial space was significantly widened with fibrous tissue filling, which was relieved by silencing of ETS1 and MEF2C (Fig. 4A,B). Moreover, ETS1 and/or MEF2C silencing alone induced the shrinkage and disarrangement of a small number of myocardial cells (Fig. 4A) and a small amount of fibrous tissue filling the interstitial space of the heart (Fig. 4B) in control rats. Studies on cardiac tissue have mainly focused on the myocardial vascular endothelium. In the immunofluorescence experiments shown in Fig. 4C, the part that expressed CD31 was the endothelium (Red fluorescence), and the part that expressed both CD31 (Red fluorescence) and AQP1 (Green fluorescence) was AQP1 expressed by endothelial cells. Transient LAD ligation (ischemia/reperfusion) induced the upregulation of AQP1 and increased the number of CD31-positive microvessels, which was inhibited by the knockdown of ETS1 and MEF2C (Fig. 4D,E).

Figure 4: ETS1 and MEF2C are critical for cardiac health. Myocardial I/R injury was induced in SD rats via left anterior descending artery (LAD) ligation, and cardiac tissues were collected. (A) HE staining was performed to assess the pathological changes of cardiac tissues in rats (20 μm). (B) Masson staining was performed to assess cardiac fibrosis in rats (20 μm). (C–E) Immunofluorescence of AQP1 and CD31 was performed to assess angiogenesis in rats (50 μm). Green fluorescence represents CD31, red fluorescence represents AQP1, and blue fluorescence represents nuclei. **p < 0.01.
3.5 AQP1 Expression in Rat Cardiac Tissues Was Regulated by ETS1 and MEF2C
ETS1, MEF2C, and AQP1 mRNA (Fig. 5A–C) and protein levels (Fig. 5D–G) were substantially elevated in the rat hearts after transient LAD ligation. Silencing of ETS1 and MEF2C abrogated the upregulation of AQP1 induced by I/R injury (Fig. 5).


Figure 5: ETS1 regulated AQP1 expression in rat hearts. Myocardial I/R injury was induced in SD rats via left anterior descending artery (LAD) ligation, and cardiac tissues were collected. (A–C) AQP1, ETS1, and MEF2C mRNA levels of cardiac tissues in different rats were evaluated by real-time PCR. (D–G) The protein levels of AQP1, ETS1, and MEF2C of cardiac tissues in different rats were assessed by western blotting. **p < 0.01.
AQP1 plays a critical role in the adaptation of endothelial cells to environmental stimuli [32,33]; therefore, the regulation of its expression is important for maintaining endothelial function in health and disease, and this study elucidated the complexity of the regulation of AQP1 expression in endothelial cells. Here, we confirm that ETS1 cooperates with MEF2C to activate AQP1 transcription in HUVECs, and it promotes MEF2C expression. Overexpression of ETS1 promoted the migration and tube formation of HUVECs, which were diminished by MEF2C knockdown and abolished by AQP1 silencing. Moreover, I/R injury induced the upregulation of ETS1, MEF2C, and AQP1 in the rat heart and increased the number of microvessels. ETS1 and MEF2C knockdown relieved I/R injury, inhibited AQP1 expression, and reduced the number of microvessels in rat cardiac tissues.
Various genetic and physiological factors have been shown to affect AQP1 levels in endothelial cells. MEF2C induces AQP1 transcription in HUVECs, which is enhanced by SP1 [19]. Hypoxia reduces AQP1 levels by increasing miR-214 in human and mouse hearts [34] but up-regulates AQP1 expression in human retinal vascular endothelial cells via hypoxia-inducible factor 1 (HIF-1) binding to AQP1 promoter [35]. AQP1 transcription in the endothelial cells can be activated by Krüppel-like factor 2 (KLF2) [36]. MEF2C activates AQP1 transcription and promotes miR-133a-3p.1 expression which post-transcriptionally regulates AQP1 expression in HUVECs [28]. The current study further demonstrated that ETS1 was not only an activator of MEF2C transcription in endothelial cells but also promoted AQP1 transcription by forming a protein complex containing at least ETS1, MEF2C, and SP1 on the AQP1 proximal promoter. These results greatly expanded our understanding of the regulatory network governing AQP1 expression in endothelial cells.
Our data showing that ETS1 modulated endothelial cell function through AQP1 expression, added a new facet to the role of ETS1 in vascular biology. ETS1 acetylation and chromatin occupancy can be increased by VEGF to promote global transcription in endothelial cells [37]. Upregulation of miR-145 in adipose-derived stromal cells inhibits their angiogenic potential by targeting ETS1, which is reversed by ETS1 overexpression [38]. ETS1 is upregulated by RE1-silencing transcription factor (REST) to promote vascular growth in medulloblastoma [39]. ETS1 has been shown to recruit BRG1 to the promoter of protein phosphatase 2A (PP2A) structural subunit a (encoded by PR65A) to activate PR65A transcription in endothelial cells, and PP2A plays a role in controlling the bioavailability of NO in the vascular system [40]. Taken together, these results suggest that ETS1 influences various aspects of endothelial and vascular health.
However, there were some shortcomings. The rescue effect of AQP1 on the synergistic effect of MEF2C and ETS1 in vivo was unknown; the specific interaction between MEF2C and ETS1 in vitro was unclear, which all required the further investigation.
In summary, ETS1 activated the transcription of MEF2C in endothelial cells, and both ETS1 and MEF2C bound to the proximal region of the AQP1 promoter to stimulate AQP1 transcription. Moreover, ETS1 promoted HUVEC migration and tube formation, which were abrogated by silencing AQP1 expression. AQP1 expression and the number of microvessels in the rat heart significantly increased after transient I/R injury, which was inhibited by the silencing of ETS1 and MEF2C. These results demonstrate the potential of targeting the ETS1/MEF2C/AQP1 pathway to treat endothelial and vascular disorders.
Acknowledgement: None.
Funding Statement: This work was supported by the Natural Science Fund Program of Science and Technology Department of Jilin Province (No. YDZJ202401002ZYTS); Scientific Research Project of the Education Department of Jilin Province (No. JJKH20261153KJ); Jilin Province College Students Innovation Training Project (No. JLMU25150, S202313706012).
Author Contributions: Study conception and design: Yong Jiang; data collection: Yong Jiang, Rui Ma, Yu-ge Wu, Yi-ming Huo, Han-zhu Zhou, Jun-xuan Zhang; analysis and interpretation of results: Yong Jiang, Rui Ma, Yu-ge Wu, Yi-ming Huo, Han-zhu Zhou, Jun-xuan Zhang; draft manuscript preparation: Yong Jiang. All authors reviewed and approved the final version of the manuscript.
Availability of Data and Materials: The data that support the findings of this study are available from the corresponding author [Yong Jiang] upon reasonable request.
Ethics Approval: This study was approved by the Institutional Animal Care and Usage Committee of Jilin Medical University (JMU2020-0037).
Conflicts of Interest: The authors declare no conflicts of interest.
References
1. Devuyst O. Aquaporin-1 and osmosis: from physiology to precision in peritoneal dialysis. J Am Soc Nephrol. 2024;35(11):1589–99. doi:10.1681/ASN.0000000000000496. [Google Scholar] [PubMed] [CrossRef]
2. Feng C, Sun CC, Wang TT, He RH, Sheng JZ, Huang HF. Decreased expression of endometrial vessel AQP1 and endometrial epithelium AQP2 related to anovulatory uterine bleeding in premenopausal women. Menopause. 2008;15(4):648–54. doi:10.1097/gme.0b013e31816086ef. [Google Scholar] [PubMed] [CrossRef]
3. Müllertz KM, Strøm C, Trautner S, Amtorp O, Nielsen S, Christensen S, et al. Downregulation of aquaporin-1 in alveolar microvessels in lungs adapted to chronic heart failure. Lung. 2011;189(2):157–66. doi:10.1007/s00408-010-9276-x. [Google Scholar] [PubMed] [CrossRef]
4. Bıçakçı H, Sarsılmaz M, Ocaklı S, Uysal M, Irmak Sapmaz H, Acar T, et al. Investigation of the effects of aging on the expression of aquaporin 1 and aquaporin 4 protein in heart tissue. Anatol J Cardiol. 2017;17(1):18–23. doi:10.14744/AnatolJCardiol.2016.7033. [Google Scholar] [PubMed] [CrossRef]
5. Yang WY, Tan ZF, Dong DW, Ding Y, Meng H, Zhao Y, et al. Association of aquaporin-1 with tumor migration, invasion and vasculogenic mimicry in glioblastoma multiforme. Mol Med Rep. 2018;17(2):3206–11. doi:10.3892/mmr.2017.8265. [Google Scholar] [PubMed] [CrossRef]
6. Xu S, Huang S, Li D, Zou Q, Yuan Y, Yang Z. The expression of aquaporin-1 and aquaporin-3 in extrahepatic cholangiocarcinoma and their clinicopathological significance. Am J Med Sci. 2022;364(2):181–91. doi:10.1016/j.amjms.2021.11.004. [Google Scholar] [PubMed] [CrossRef]
7. Kim J, Jung Y. Different expressions of AQP1, AQP4, eNOS, and VEGF proteins in ischemic versus non-ischemic cerebropathy in rats: potential roles of AQP1 and eNOS in hydrocephalic and vasogenic edema formation. Anat Cell Biol. 2011;44(4):295–303. doi:10.5115/acb.2011.44.4.295. [Google Scholar] [PubMed] [CrossRef]
8. Qiao R, Cui X, Hu Y, Wei H, Xu H, Zhang C, et al. Hypoxia reduces mouse urine output via HIF1α-mediated upregulation of renal AQP1. Kidney Dis. 2024;10(6):504–18. doi:10.1159/000542087. [Google Scholar] [PubMed] [CrossRef]
9. Gao C, Tang J, Li R, Huan J. Specific inhibition of AQP1 water channels in human pulmonary microvascular endothelial cells by small interfering RNAs. J Trauma Acute Care Surg. 2012;72(1):150–61. doi:10.1097/TA.0b013e318230e25d. [Google Scholar] [PubMed] [CrossRef]
10. Jiang Y, Wang C, Ma R, Zhao Y, Ma X, Wan J, et al. Aquaporin 1 mediates early responses to osmotic stimuli in endothelial cells via the calmodulin pathway. FEBS Open Bio. 2021;11(1):75–84. doi:10.1002/2211-5463.13020. [Google Scholar] [PubMed] [CrossRef]
11. Pon JR, Marra MA. MEF2 transcription factors: developmental regulators and emerging cancer genes. Oncotarget. 2016;7(3):2297–312. doi:10.18632/oncotarget.6223. [Google Scholar] [PubMed] [CrossRef]
12. Taylor MV, Hughes SM. Mef2 and the skeletal muscle differentiation program. Semin Cell Dev Biol. 2017;72:33–44. doi:10.1016/j.semcdb.2017.11.020. [Google Scholar] [PubMed] [CrossRef]
13. Liu B, Ou WC, Fang L, Tian CW, Xiong Y. Myocyte enhancer factor 2A plays a central role in the regulatory networks of cellular physiopathology. Aging Dis. 2023;14(2):331–49. doi:10.14336/AD.2022.0825. [Google Scholar] [PubMed] [CrossRef]
14. Adrião A, Mariano S, Mariano J, Gavaia PJ, Cancela ML, Vitorino M, et al. mef2ca and mef2cb double mutant zebrafish show altered craniofacial phenotype and motor behaviour. Biomolecules. 2023;13(5):805. doi:10.3390/biom13050805. [Google Scholar] [PubMed] [CrossRef]
15. Potthoff MJ, Arnold MA, McAnally J, Richardson JA, Bassel-Duby R, Olson EN. Regulation of skeletal muscle sarcomere integrity and postnatal muscle function by Mef2c. Mol Cell Biol. 2007;27(23):8143–51. doi:10.1128/MCB.01187-07. [Google Scholar] [PubMed] [CrossRef]
16. Wilker PR, Kohyama M, Sandau MM, Albring JC, Nakagawa O, Schwarz JJ, et al. Transcription factor Mef2c is required for B cell proliferation and survival after antigen receptor stimulation. Nat Immunol. 2008;9(6):603–12. doi:10.1038/ni.1609. [Google Scholar] [PubMed] [CrossRef]
17. Lin Q, Lu J, Yanagisawa H, Webb R, Lyons GE, Richardson JA, et al. Requirement of the MADS-box transcription factor MEF2C for vascular development. Development. 1998;125(22):4565–74. doi:10.1242/dev.125.22.4565. [Google Scholar] [PubMed] [CrossRef]
18. Liang W, Guo L, Liu T, Qin S. MEF2C alleviates acute lung injury in cecal ligation and puncture (CLP)-induced sepsis rats by up-regulating AQP1. Allergol Immunopathol. 2021;49(5):117–24. doi:10.15586/aei.v49i5.477. [Google Scholar] [PubMed] [CrossRef]
19. Jiang Y, Liu H, Liu WJ, Tong HB, Chen CJ, Lin FG, et al. Endothelial aquaporin-1 (AQP1) expression is regulated by transcription factor Mef2c. Mol Cells. 2016;39(4):292–8. doi:10.14348/molcells.2016.2223. [Google Scholar] [PubMed] [CrossRef]
20. Shiu YT, Jaimes EA. Transcription factor ETS-1 and reactive oxygen species: role in vascular and renal injury. Antioxidants. 2018;7(7):84. doi:10.3390/antiox7070084. [Google Scholar] [PubMed] [CrossRef]
21. Liu Y, Shen L, Li Y, Sun X, Liang L, Jiang S, et al. ETS1-mediated regulation of SOAT1 enhances the malignant phenotype of oral squamous cell carcinoma and induces tumor-associated macrophages M2-like polarization. Int J Biol Sci. 2024;20(9):3372–92. doi:10.7150/ijbs.93815. [Google Scholar] [PubMed] [CrossRef]
22. Chen Y, Peng C, Chen J, Chen D, Yang B, He B, et al. WTAP facilitates progression of hepatocellular carcinoma via m6A-HuR-dependent epigenetic silencing of ETS1. Mol Cancer. 2019;18(1):127. doi:10.1186/s12943-019-1053-8. [Google Scholar] [PubMed] [CrossRef]
23. Yang Y, Han X, Sun L, Shao F, Yin Y, Zhang W. ETS transcription factors in immune cells and immune-related diseases. Int J Mol Sci. 2024;25(18):10004. doi:10.3390/ijms251810004. [Google Scholar] [PubMed] [CrossRef]
24. Battaglia M, Sunshine AC, Luo W, Jin R, Stith A, Lindemann M, et al. Ets1 and IL17RA cooperate to regulate autoimmune responses and skin immunity to Staphylococcus aureus. Front Immunol. 2023;14:1208200. doi:10.3389/fimmu.2023.1208200. [Google Scholar] [PubMed] [CrossRef]
25. Qin J, Zou L, Lu F, Liu F, Min Q, Zhu L. METTL3 promotes immature dental pulp stem cells-induced angiogenesis by regulating ETS1 mRNA stability in an m6A-HuR-dependent manner. Odontology. 2025;113(1):305–17. doi:10.1007/s10266-024-00977-3. [Google Scholar] [PubMed] [CrossRef]
26. Wang L, Lin L, Qi H, Chen J, Grossfeld P. Endothelial loss of ETS1 impairs coronary vascular development and leads to ventricular non-compaction. Circ Res. 2022;131(5):371–87. doi:10.1161/CIRCRESAHA.121.319955. [Google Scholar] [PubMed] [CrossRef]
27. De Val S, Anderson JP, Heidt AB, Khiem D, Xu SM, Black BL. Mef2c is activated directly by Ets transcription factors through an evolutionarily conserved endothelial cell-specific enhancer. Dev Biol. 2004;275(2):424–34. doi:10.1016/j.ydbio.2004.08.016. [Google Scholar] [PubMed] [CrossRef]
28. Jiang Y, Ma R, Zhao Y, Li GJ, Wang AK, Lin WL, et al. MEF2C/miR-133a-3p.1 circuit-stabilized AQP1 expression maintains endothelial water homeostasis. FEBS Lett. 2019;593(18):2566–73. doi:10.1002/1873-3468.13516. [Google Scholar] [PubMed] [CrossRef]
29. Xiong Z, Zhou X, Li Y, Yang L. PACAP inhibits sepsis-associated acute lung injury by inhibiting the Sp1/AQP1 pathway. Peptides. 2025;191:171411. doi:10.1016/j.peptides.2025.171411. [Google Scholar] [PubMed] [CrossRef]
30. Palethorpe HM, Tomita Y, Smith E, Pei JV, Townsend AR, Price TJ, et al. The aquaporin 1 inhibitor bacopaside II reduces endothelial cell migration and tubulogenesis and induces apoptosis. Int J Mol Sci. 2018;19(3):653. doi:10.3390/ijms19030653. [Google Scholar] [PubMed] [CrossRef]
31. Xiang S, Tang X. Interfering with AQP1 alleviates ferroptosis, improves mitochondrial function and energy metabolic disorder in hypoxia/reoxygenation-induced H9c2 cardiomyocytes via Wnt/β-catenin pathway. Microvasc Res. 2025;160:104821. doi:10.1016/j.mvr.2025.104821. [Google Scholar] [PubMed] [CrossRef]
32. Shabanian K, Shabanian T, Karsai G, Pontiggia L, Paneni F, Ruschitzka F, et al. AQP1 differentially orchestrates endothelial cell senescence. Redox Biol. 2024;76:103317. doi:10.1016/j.redox.2024.103317. [Google Scholar] [PubMed] [CrossRef]
33. Maltaneri RE, Schiappacasse A, Chamorro ME, Nesse AB, Vittori DC. Aquaporin-1 plays a key role in erythropoietin-induced endothelial cell migration. Biochim Biophys Acta Mol Cell Res. 2020;1867(1):118569. doi:10.1016/j.bbamcr.2019.118569. [Google Scholar] [PubMed] [CrossRef]
34. Rutkovskiy A, Bliksøen M, Hillestad V, Amin M, Czibik G, Valen G, et al. Aquaporin-1 in cardiac endothelial cells is downregulated in ischemia, hypoxia and cardioplegia. J Mol Cell Cardiol. 2013;56:22–33. doi:10.1016/j.yjmcc.2012.12.002. [Google Scholar] [PubMed] [CrossRef]
35. Tanaka A, Sakurai K, Kaneko K, Ogino J, Yagui K, Ishikawa K, et al. The role of the hypoxia-inducible factor 1 binding site in the induction of aquaporin-1 mRNA expression by hypoxia. DNA Cell Biol. 2011;30(8):539–44. doi:10.1089/dna.2009.1014. [Google Scholar] [PubMed] [CrossRef]
36. Fontijn RD, Volger OL, van der Pouw-Kraan TC, Doddaballapur A, Leyen T, Baggen JM, et al. Expression of nitric oxide-transporting aquaporin-1 is controlled by KLF2 and marks non-activated endothelium in vivo. PLoS One. 2015;10(12):e0145777. doi:10.1371/journal.pone.0145777. [Google Scholar] [PubMed] [CrossRef]
37. Chen J, Fu Y, Day DS, Sun Y, Wang S, Liang X, et al. VEGF amplifies transcription through ETS1 acetylation to enable angiogenesis. Nat Commun. 2017;8(1):383. doi:10.1038/s41467-017-00405-x. [Google Scholar] [PubMed] [CrossRef]
38. Arderiu G, Peña E, Aledo R, Juan-Babot O, Crespo J, Vilahur G, et al. microRNA-145 regulates the differentiation of adipose stem cells toward microvascular endothelial cells and promotes angiogenesis. Circ Res. 2019;125(1):74–89. doi:10.1161/CIRCRESAHA.118.314290. [Google Scholar] [PubMed] [CrossRef]
39. Shaik S, Maegawa S, Haltom AR, Wang F, Xiao X, Dobson T, et al. REST promotes ETS1-dependent vascular growth in medulloblastoma. Mol Oncol. 2021;15(5):1486–506. doi:10.1002/1878-0261.12903. [Google Scholar] [PubMed] [CrossRef]
40. Chen B, Zhao Q, Xu T, Yu L, Zhuo L, Yang Y, et al. BRG1 activates PR65A transcription to regulate NO bioavailability in vascular endothelial cells. Front Cell Dev Biol. 2020;8:774. doi:10.3389/fcell.2020.00774. [Google Scholar] [PubMed] [CrossRef]
Cite This Article
 Copyright © 2026 The Author(s). Published by Tech Science Press.
Copyright © 2026 The Author(s). Published by Tech Science Press.This work is licensed under a Creative Commons Attribution 4.0 International License , which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.


 Submit a Paper
Submit a Paper Propose a Special lssue
Propose a Special lssue View Full Text
View Full Text Download PDF
Download PDF Downloads
Downloads
 Citation Tools
Citation Tools