 Open Access
Open Access
ARTICLE
Cardiac Arrhythmia Disease Classifier Model Based on a Fuzzy Fusion Approach
1 College of Technological Innovation, Zayed University, Dubai, UAE
2 Department of Information Systems, College of Computer and Information Sciences, Jouf University, Sakaka, Saudi Arabia
3 Department of Electronics and Communications Engineering, Delta Higher Institute for Engineering & Technology, Mansoura, Egypt
4 College of Computing and Informatics, University of Sharjah, Sharjah, United Arab Emirates
5 Department of Information Systems, Mansoura University, Mansoura, 35516, Egypt
6 Department of Teacher Preparation, Faculty of Specific Education, Mansoura University, Mansoura, Egypt
* Corresponding Author: Abdulaziz Shehab. Email:
Computers, Materials & Continua 2023, 75(2), 4485-4499. https://doi.org/10.32604/cmc.2023.036118
Received 17 September 2022; Accepted 08 February 2023; Issue published 31 March 2023
Abstract
Cardiac diseases are one of the greatest global health challenges. Due to the high annual mortality rates, cardiac diseases have attracted the attention of numerous researchers in recent years. This article proposes a hybrid fuzzy fusion classification model for cardiac arrhythmia diseases. The fusion model is utilized to optimally select the highest-ranked features generated by a variety of well-known feature-selection algorithms. An ensemble of classifiers is then applied to the fusion’s results. The proposed model classifies the arrhythmia dataset from the University of California, Irvine into normal/abnormal classes as well as 16 classes of arrhythmia. Initially, at the preprocessing steps, for the miss-valued attributes, we used the average value in the linear attributes group by the same class and the most frequent value for nominal attributes. However, in order to ensure the model optimality, we eliminated all attributes which have zero or constant values that might bias the results of utilized classifiers. The preprocessing step led to 161 out of 279 attributes (features). Thereafter, a fuzzy-based feature-selection fusion method is applied to fuse high-ranked features obtained from different heuristic feature-selection algorithms. In short, our study comprises three main blocks: (1) sensing data and preprocessing; (2) feature queuing, selection, and extraction; and (3) the predictive model. Our proposed method improves classification performance in terms of accuracy, F1 measure, recall, and precision when compared to state-of-the-art techniques. It achieves 98.5% accuracy for binary class mode and 98.9% accuracy for categorized class mode.Keywords
Cardiac diseases are one of the major health challenges worldwide. Recently, according to a report released by the World Health Organization, cardiovascular disease is the number one disease that affects people’s health seriously and causes death globally. It has high mortality rates, with nearly 17.9 million people dying each year, accounting for about 31% of all global deaths [1]. The early diagnosis and careful treatment of such diseases can reduce unnecessary deaths. Delays in the detection of cardiac diseases may aggravate the severity of the disease and delay its treatment. In fact, the lack of financing in the healthcare infrastructure of developed countries restricts exposure to screening and thus increases the risk of undiagnosed cardiovascular disease. Therefore, the early diagnosis and detection of heart disease play a significant role in reducing the rate of death and determining appropriate treatment. Cardiac arrhythmia is an irregular type of heartbeat, which is either too quick (tachycardia) or too slow (bradycardia). Arrhythmias may be of various types, which are harmful, and may, unless detected and monitored in time, cause cardiac arrest or sudden death [2]. Arrhythmias are important cardiovascular diseases because they foretell anomalies in the electrophysiology of the heart. These heartbeat variations from the norm are distinguished utilizing electrocardiogram (ECG) signals. An ECG signal is a bioelectrical signal that records the electrical activity of the heart. ECG signals are utilized as a parameter to distinguish different heart infections. The traditional method of ECG analysis requires cardiologists to spend hours monitoring a patient’s status by examining and interpreting their heart activity on a long-term basis to discover abnormalities in ECG signals. Manual ECG analysis is time-consuming, requires great effort by cardiologists, and is practically difficult [3].
Delays in detecting irregular heartbeat may increase the seriousness of the disease and reduce the rate of successful care. To address these obstacles, a timely classification and an accurate diagnosis based on the automatic detection of various types of arrhythmias is essential and expected to play a major role in saving patients’ lives, enabling them to obtain effective care and sustain a healthy life. The automatic detection and classification of arrhythmia are great benefits to physicians also [4,5].
In recent decades, artificial intelligence (AI) and machine learning have been used widely in the healthcare field. AI can assist in the detection of arrhythmia from the enormous volumes of clinical information, providing a timely and accurate classification of arrhythmias. It can also assist in the early diagnosis of arrhythmia, provide an appropriate treatment plan for patients, and help patients avoid future clinical problems that may be life-threatening. A strong model classifier is needed to reliably distinguish between the different types of arrhythmia classes and a salient feature extractor is needed to extract meaningful information from the large volume of raw data [6].
In the last few years, the use of machine-learning techniques to build automatic systems for the classification and diagnosis of human diseases has received unprecedented attention. These systems play a vital role in assisting physicians to accomplish biomedical and healthcare tasks that were difficult to solve by other laboratory methods or human experts. Machine-learning techniques contribute to the early detection and better diagnosis of diseases and help save expensive medical costs and tests. Many techniques for the automated classification of arrhythmia have been created in the last decades to improve the automatic detection and classification of the disease. These techniques may be categorized into two main categories: (1) binary classification, which classifies patients into healthy or unhealthy; however, this form of classification can be used only in basic systems without diagnostic and control purposes; and (2) multi-class categorization, which aims to classify patients based on the different types of arrhythmias [3,7,8].
Feature extraction and selection are one of the main steps prior to the classification process. A large number of features in the data to be analyzed is one of the main challenges in any diagnosis model. This might affect the accuracy of the process and lead to difficulties in data interpretation [7]. Therefore, feature reduction is inevitable in the search for an optimal subset of features. The removal of irrelevant features often enhances the performance of the classifier. In general, in the literature, feature-selection techniques can be divided into two main categories: (1) wrapper techniques and (2) heuristic techniques [7]. The first technique estimates the features’ relevance using different machine-learning algorithms wrapped in the search process. Subsequently, these selected features are expected to improve the classification results. The second technique, on the other hand, estimates the features’ score according to heuristics, based on the statistical analysis of the data [9]. In this study, we utilize four classic heuristic approaches, namely, (1) information gain, (2) information gain retrieval, (3) Gini index, and (4) Relief-f, to select the most relevant features of cardiac arrhythmia. Accordingly, we experimentally conducted a feature-selection algorithm based on a neuro-fuzzy fusion approach to differentiate valuable features of the arrhythmia dataset.
The proposed work is aiming to integrate various types of Electronic Health Records (EHRs) data (structured data, unstructured data, and graphic data), sensory data, and user input data all over time to predict and pre-empt illnesses. It also helps improve patient care and treatment and eases the burden of clinicians by providing timely and assistive recommendations. Finally, it is serving an aging population that needs long-term care, and rising expectations from patients. The main contribution of our proposed model is to improve the efficiency and effectiveness of the healthcare services related to cardiac diseases through the usage of different Machine Learning techniques. Besides, it intends to apply modern information and communication technologies in all healthcare fields such as collecting, storing, restoring, and analyzing cardiac Arrhythmia information. This paper is organized as follows: Section 2 presents the literature review; Section 3 presents the details of our proposed model. The experimental results are presented in Section 4. Finally, the conclusion and future works are presented in Section 5.
Currently, various automatic system-based machine-learning approaches are proposed for arrhythmia classification. In general, the automatic systems for arrhythmia classification are composed of three main steps: preprocessing, feature extraction and selection, and classification. In this section, we present a systematic review of the current state-of-the-art methods used to detect cardiac arrhythmia, focusing on feature extraction, feature selection, and classification approaches, as well as the dataset used for automatic detection and diagnosis processes.
Ayar et al. [10] presented a model for both binary and multi-class ECG arrhythmia classification. Their proposed model used genetic algorithms (GA) for optimal feature selection and a decision tree (DT) with the C4.5 algorithm for data classification. Experiments were conducted using the University of California at Irvine (UCI) arrhythmia dataset; the average accuracy was 86.96% for binary classification and 78.76% for multi-class classification. Assodiky et al. [11] utilized deep learning to classify the arrhythmia. The results demonstrated that deep learning accomplished the best classification rate, with 76.51% accuracy. Moreover, experiments also showed that particle swarm optimization execution in feature reduction was superior to the GA. Kadam et al. [12] proposed a novel ECG arrhythmia classification model that incorporates a GA to select features and the Soft-Margin support vector machine (SVM) to classify arrhythmia into healthy and unhealthy classes. The model yielded an 87.83% exactness rate, with ten overlapping cross-approval strategies using the arrhythmia dataset. Darwaish et al. [13] used discrete wavelet transform for the preprocessing of the ECG signal and undecimated wavelet transform for extracting nine features. The probabilistic mode named the Bayesian network classifier was trained using the extracted features on the UCI arrhythmia dataset. Dalal et al. [14] utilized the kernel extreme learning machine classifier with the GA for arrhythmia classification. This experimentation was performed on the UCI repository arrhythmia and Polypyrimidine Tract-binding Protein (PTB) Diagnostic ECG Database. In this approach, cumulants are utilized to solve the problem of the missing value in the arrhythmia database; an 86.67% accuracy rate was achieved. Lang et al. [15] proposed a k-Nearest Neighbor (KNN) classifier method, which is called the difference-weighted k-nearest neighbor. Their proposed classifier could recognize unbalanced cardiac arrhythmia data after employing the principal component analysis method to deal with missing data in the arrhythmia dataset. It achieved an average accuracy of 70.80%. Pandey et al. [7] proposed a machine-learning-based system to detect and diagnose arrhythmia diseases based on ECG analysis. Their proposed system contains three phases: the first phase is preprocessing, in which data are normalized and missing values are removed; the second phase employs principal component analysis for feature selection; and finally, eight classifiers are applied to the various data splits for arrhythmia classification. Their experiments, which were conducted on the UCI cardiac arrhythmia dataset, showed that SVMs and Naive Bayes yielded a maximum accuracy of 89.74%. Singh et al. [9] proposed a model to diagnose and classify cardiac arrhythmia diseases. The authors treated the missing value problem in the arrhythmia dataset by replacing numerical attributes by their means and replacing nominal attributes by their modes. Their proposed model utilized the chi-square test, symmetrical uncertainty, and gain ratio for the feature-selection process, while the proposed model employed three classifiers for normal and abnormal classification. They achieved a high accuracy of 85.58%, by using a random forest (RF) classifier with gain ratio feature selection. Shandri et al. [16] presented a multi-class arrhythmia clustering model using fuzzy robust kernel c-means. The highest accuracy obtained was 90.48% on the UCI repository arrhythmia database. Jadhav et al. [17] presented a feature elimination–based approach to diagnosing arrhythmia by classifying it into normal and abnormal classes. The random subspace (RS) ensemble classifier is used for arrhythmia classification, yielding an accuracy of 91.11% with ensemble sizes of 15 and 20. Khan et al. [18] presented a two-stage cascade structure method to classify cardiac arrhythmia. The first stage employed logistic regression (LR), DT, and RF for cardiac arrhythmia classification, while the second stage employed Multi-Layer Perceptron (MLP) and then long short-term memory (LSTM) deep architectures for multi-class and binary classification arrhythmia. The experiments of the first stage yielded accuracies of 83.0%, 85.4%, and 87.5% for LR, DT, and RF, respectively, while the second phase yielded accuracies of 89.0% and 94.8% for MLP and LSTM, respectively. Yadav et al. [19] propose a framework for the classification of cardiac arrhythmia patients by focusing on the factors that are more related to their diagnosis. The authors use a database that was processed on the Health Sciences and Technology (HEST) which is categorized into a multi-class and a single-class. Itzhak et al. [20] propose a study to detect and classify three arrhythmia types, namely atrial fibrillation, bradycardia, and tachycardia. The authors relied on logistic regression, SVM, and random forest classifiers. Their best finding was a performance with a sensitivity of 0.92 and a specificity of 0.86 for a multiclass random forest classifier.
Most machine-learning algorithms are influenced highly by the miss-valued attributes and those containing several redundant values [9]. There are several schemes for imputing such missing values. Addressing these issues along with many others can be crucial, especially when one wants to improve model performance and generalize the ability of the model. As the dataset utilized in this study had such difficulties, we had to preprocess the data in an intelligent way that could help learners (classifiers) later. For the miss-valued attributes, we used the average value in the linear attributes group by the same class and the most frequent value for nominal attributes. While the imputation can be done randomly by assigning values to the data based on the domain knowledge, there were the right steps to ensure that the transformed data guarantee the model optimality attaining the maximum efficiency. However, some classes did not have any certain value for an attribute. Hence, the overall average of all averages was calculated per class. On the other hand, regarding data redundancy, we faced attributes with zero or a constant value and some attributes with correlated behaviors. As a result, we eliminated all such attributes that might bias the results of utilized classifiers. The preprocessing step led to 161 out of 279 attributes (features).
Fusing conflict evidence is one of the fundamental challenges in the decision-making field because of the fusion of ever-increasing uncertain data [21,22]. In this study, a novel fuzzy-based feature-selection fusion method is proposed to fuse high-ranked features obtained from different heuristic feature-selection algorithms. Fig. 1 shows the process flow of our proposed framework. In brief, it contains three main blocks: (1) sensing data and preprocessing; (2) feature queuing, selection, and extraction; and (3) the predictive model.
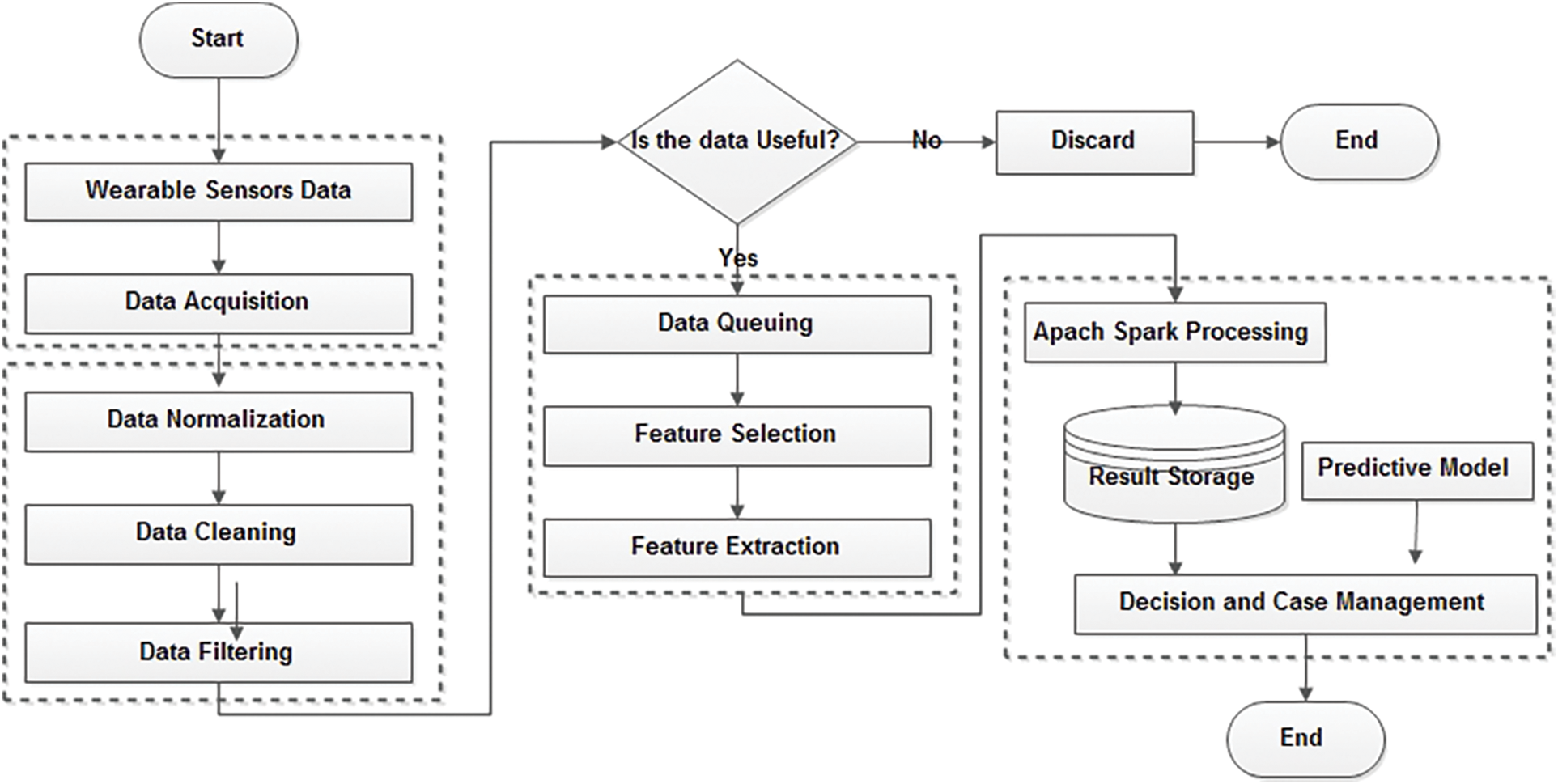
Figure 1: Working steps of the proposed model
Data normalization, which generalizes the effect of the different data acquired from sensors, could be defined in many different forms. Given the data from the sensors noted as DS and of size n, the normalized data DS* is obtained by applying the normalization approach. The normalization used in the experiment is done by the maximum and minimum values, which are illustrated in Eq. (1):
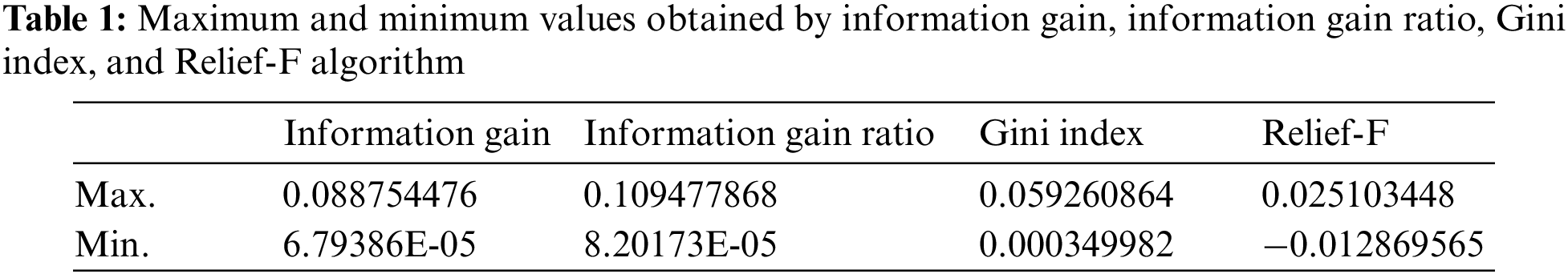
Many metrics approaches are utilized for such purposes of feature selection (information gain, information gain ratio, Gini index, and Relief-F). The Gini index, for instance, measures the inequality among values of a frequency distribution that range from 0 (complete equality) to 1 (complete inequality). The Gini coefficient is often calculated according to the Brown formula in Eq. (2), which G denotes the Gini coefficient,

Fuzzy set theory is an inference scheme that targets defective data. It relies on partial set membership, which provides fuzzy inference rather than crisp inference [23]. A fuzzy set A ⊆ U is defined by the function μA(u) in the interval [0, 1], as shown in Eq. (4):
Each element u in U is mapped into a membership degree μA(u) on [0, 1]. Fuzzy set theory is based mainly on the membership function, which is usually determined through experience and statistics. Occasionally, three types of membership functions are used widely: Gaussian, triangular, and trapezoidal membership functions [23]. In our model, the trapezoidal membership function was utilized, which is illustrated in Eq. (5):
where b and c are the abscissas of the two upper vertices, and a and d are the abscissas of the two lower vertices of the trapezoidal.
Fuzzy rules are implemented manually, based on criteria of different expertise and knowledge of the four feature-selection algorithms utilized. The criteria used to devise the fuzzy rules are summarized as follows: first, if the four algorithms have the same value with respect to classifying a specific sample, then we keep the same fuzzy value; second, if two classifiers yield the same linguistic value with respect to classifying a specific sample and the linguistic value of the different opinion classifier represents a major value, then, consequently, the linguistic value of the different opinion classifier is obtained; third, if two classifiers yield the same linguistic value with respect to classifying a specific sample and the linguistic value of the different opinion classifier represents a lower linguistic value, then, consequently, the linguistic value of the two classifiers with the same opinion is obtained; and fourth, if the three classifiers differ in opinion, then, consequently, the higher linguistic value is obtained. We apply these three criteria to form all the fuzzy rules for the fuzzy system to combine all the output results of the classifiers in the basic module unit for the hybrid model.
To validate our approach, we used the UCI cardiac arrhythmia dataset [17]. It contains the ECG signal data, which are composed of 279 attributes and 452 instances. Most of these attributes are linear (206) and the rest (73) are nominal. The dataset is categorized into 16 different classes. Class 01 belongs to normal people, classes 02–15 belong to different types of arrhythmia, and class 16 belongs to unclassified or unknown arrhythmia types. In general, for the purpose of binary classification, class 01 is called the normal class, and classes 02–16 are seen as the abnormal class. The normal class contains 245 instances, while the abnormal class contains 207 instances. Table 2 details the class types and their instances in the dataset. Classes 11, 12, and 13 contain no instances.

To evaluate the proposed model, Eqs. (6)–(9) were employed, namely, the accuracy, recall, precision, and F1 score. All the following metrics are expressed as percentages. For all, TP refers to a true positive, FP is a false positive, TN is a true negative, and FN is a false negative. We also used the receiver operating characteristics and the area under the curve.
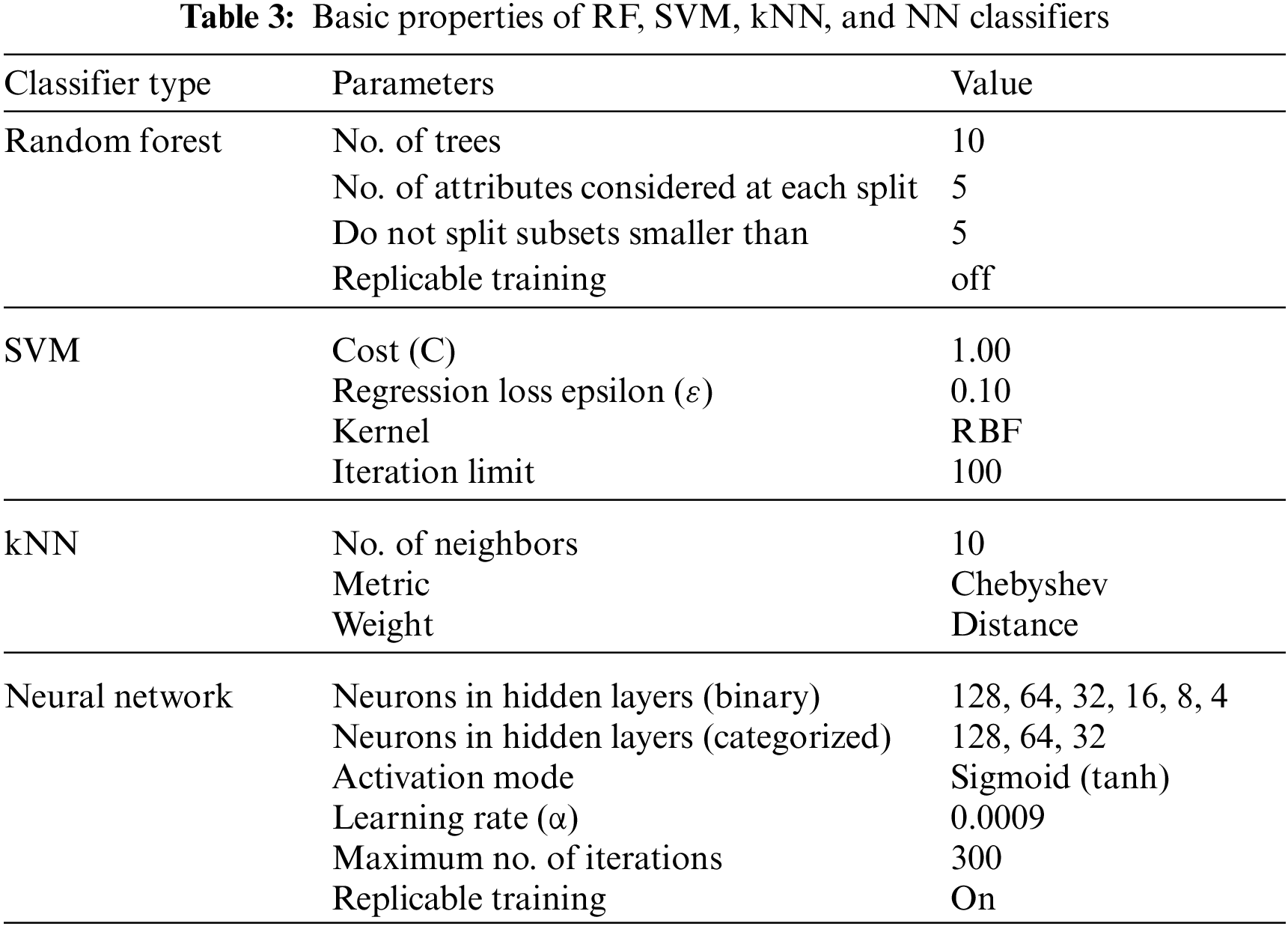
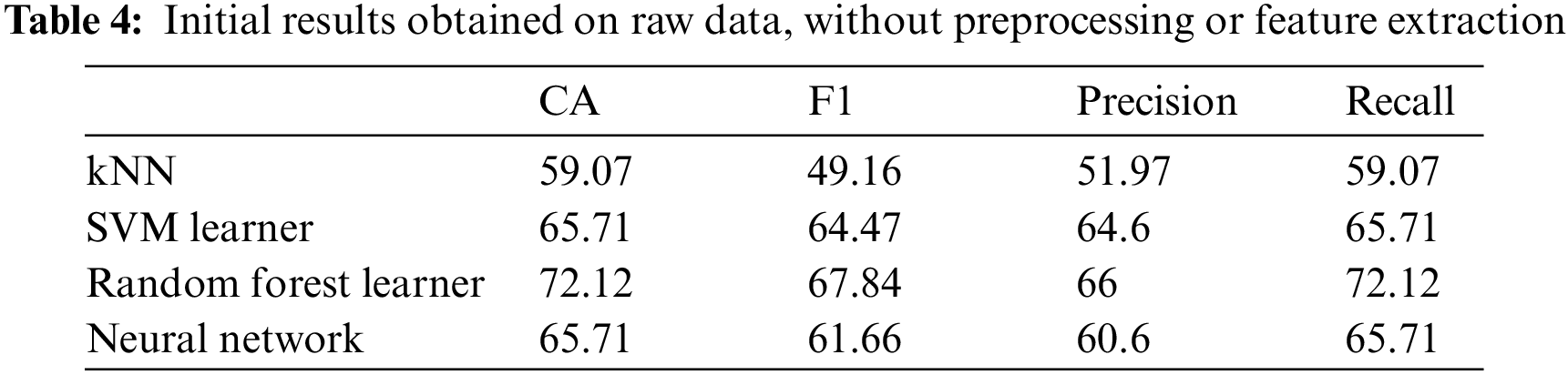
Initially, we performed a pilot experiment on the raw data, without any preprocessing actions and without feature extraction (i.e., using all 279 features). Table 3 summarizes the basic configuration of the four main classifiers used in the experiment, namely, RF, SVM, kNN, and NN. Every classifier gets the same set of heartbeat vectors and provides an output that represents the specific classification of the heartbeat. Table 4 shows these results, and it is noticeable that the accuracy is so weak because both the preprocessing of data and feature selection were ignored.
In this study, the results of classifying cardiac arrhythmia data into normal or abnormal classes (binary classification) are explored. The results of different classifiers with different feature extraction algorithms are used. Besides, the results of our fused approach were compared with those of a study using state-of-the-art methods. Table 5 summarizes the average accuracy measured for KNN, SVM, NN, and RF binary classifiers for 10, 25, 50, and 100 features selected using information gain, information gain ratio, Gini index, and Relief-F algorithms. All the results obtained were measured using tenfold cross-validation.
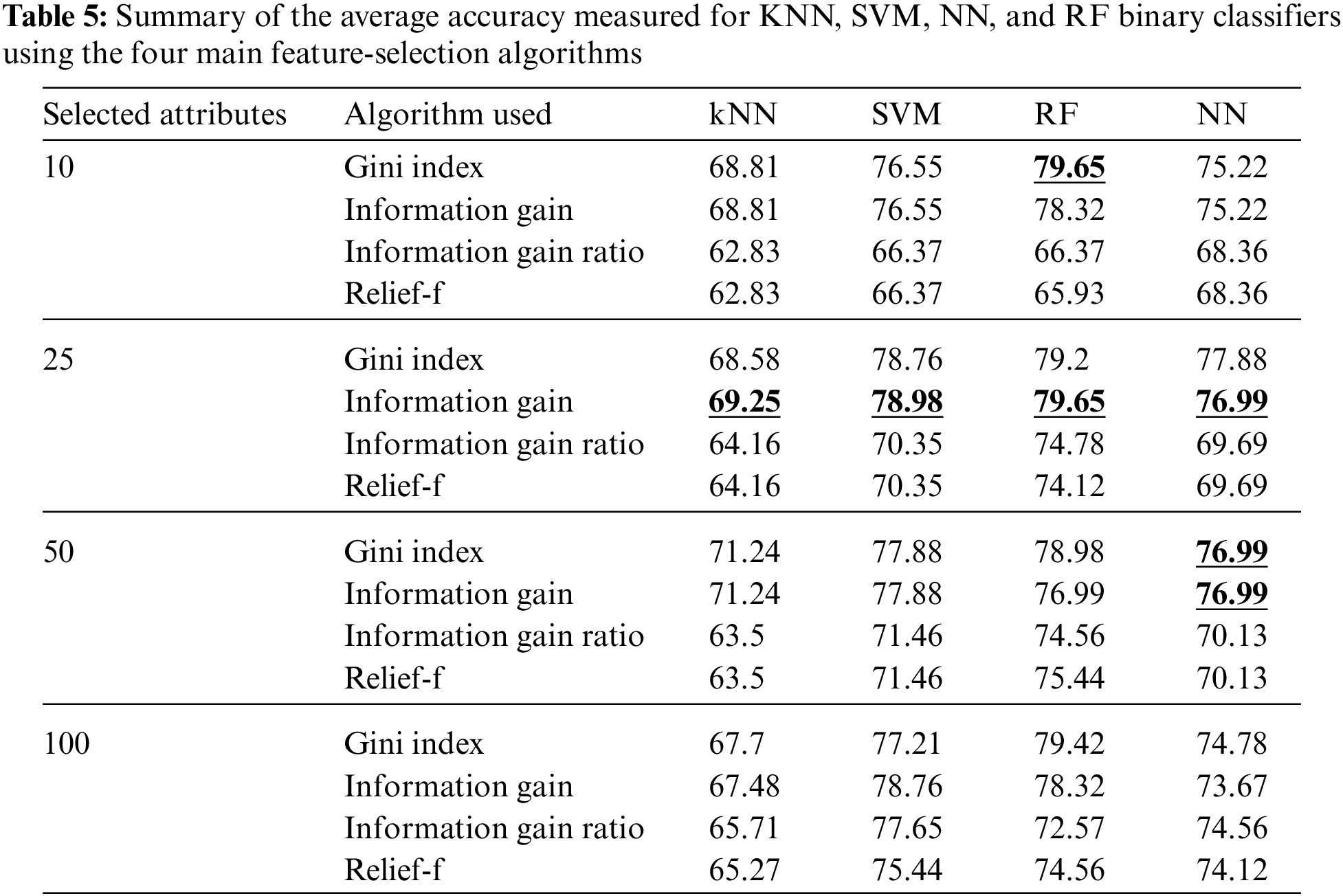
As noticed from Table 5, the information gain algorithm achieved the maximum average accuracy, using the 25 features selected, measured for KNN, SVM, NN, and RF, which were 69.25%, 78.98%, 79.65%, and 76.99%, respectively. The Gini index achieved the maximum average accuracy for the Relief-F algorithm using ten features, and both the Gini index and information gain achieved the maximum accuracy for neural networks using 50 features. In summary, the most suitable feature-selection algorithm was noticed to be information gain and the most suitable number of features was 25.
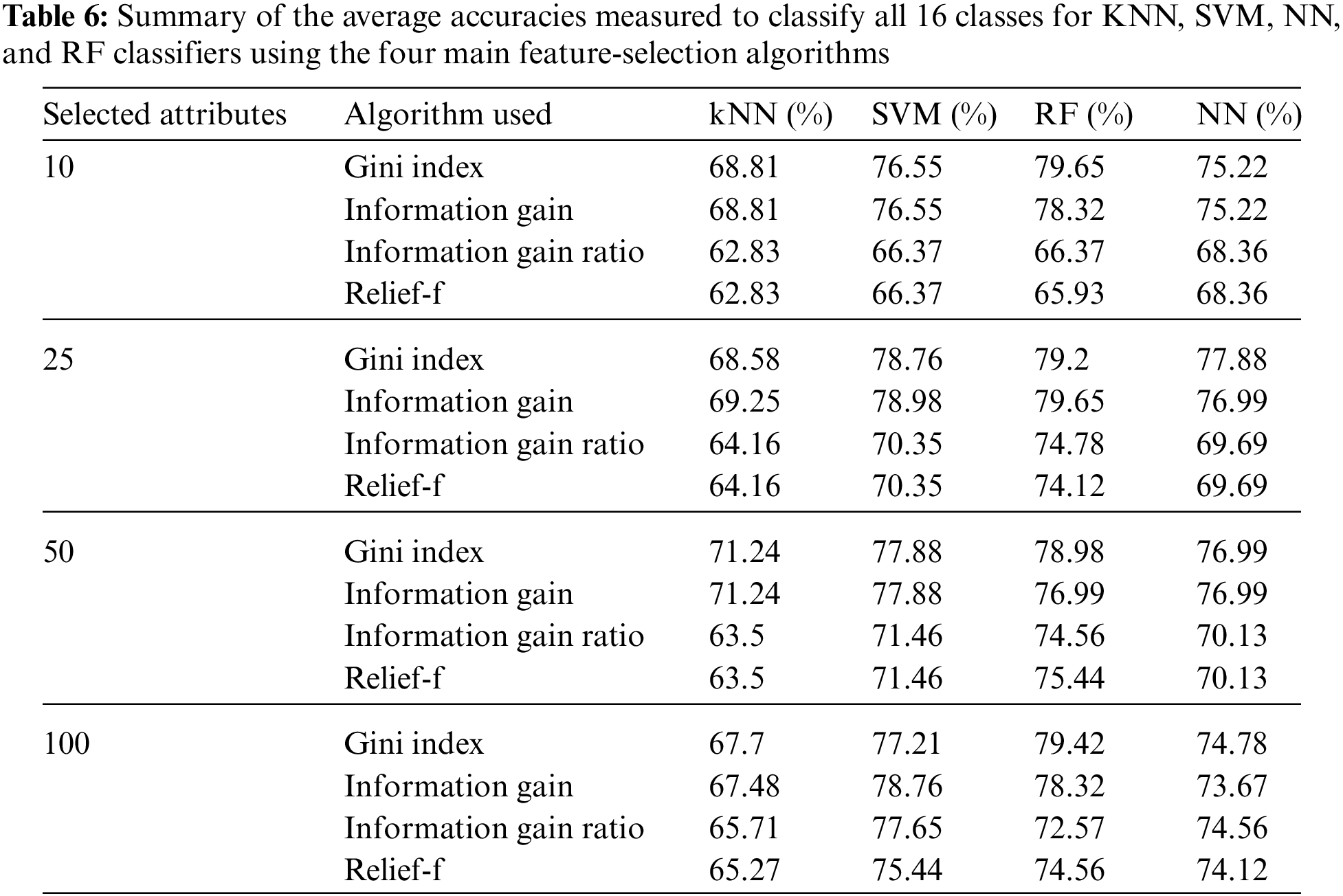
Besides, the results of the categorized classification mode are reported. As mentioned above, there were 16 classes (one normal class, 14 arrhythmia classes, and one unknown class) of data in the UCI cardiac arrhythmia dataset. Table 6 summarizes the average accuracies measured for KNN, SVM, NN, and RF classifiers to classify all 16 classes.
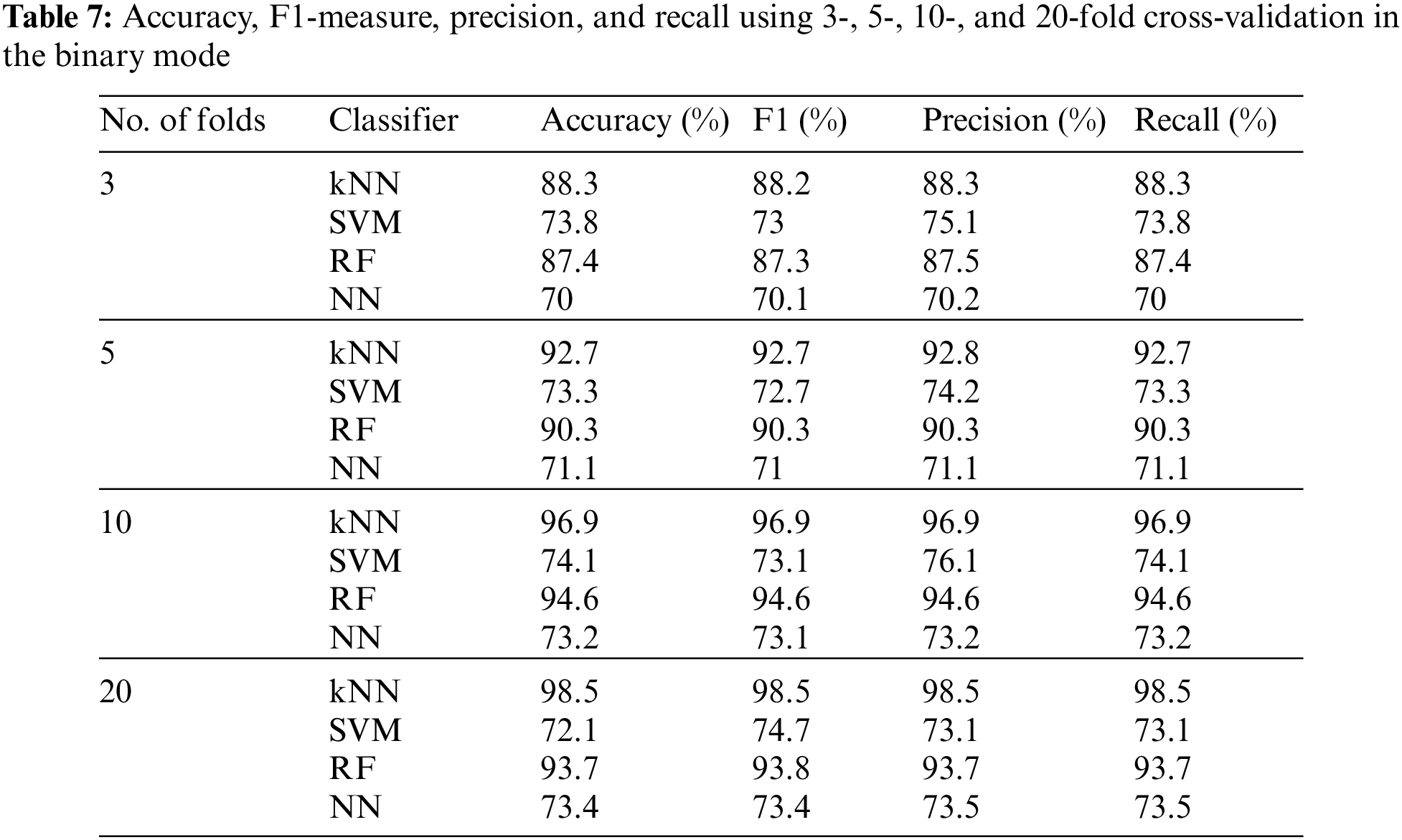
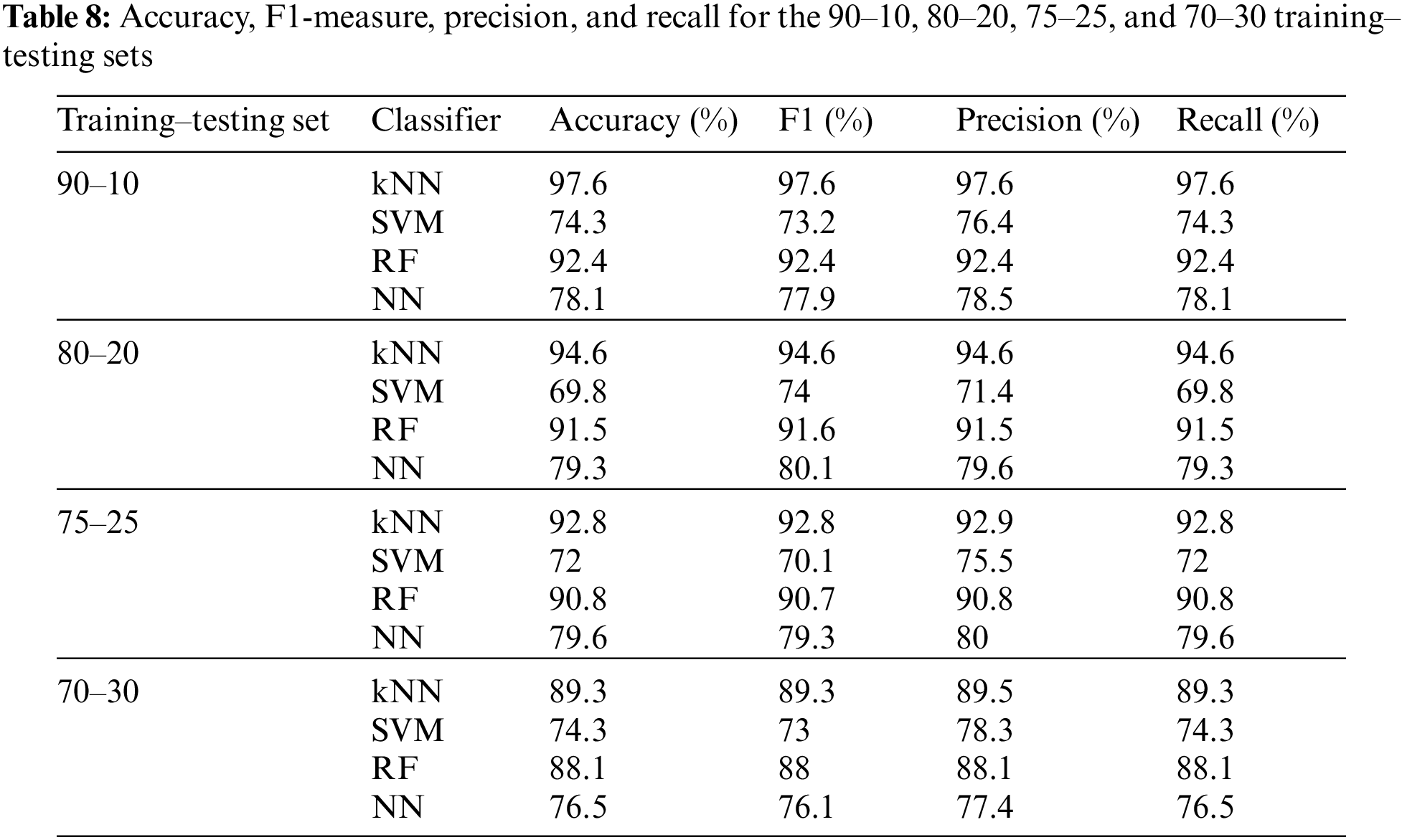
To test our proposed fused approach results, we used n-fold cross-validation, fixing the number of features at 20 for the binary mode. Table 7 shows the accuracy, F1-measure, precision, and recall using 3-, 5-, 10-, and 20-fold cross-validation and the four main classifiers. Besides, the binary mode was also tested using the training–testing sets 90–10, 80–20, 75–25, and 70–30 while fixing the number of features at 20. Table 8 summarizes these results. The maximum achievable accuracy was 97.6% for the kNN classifier at 20-fold.
The influence of changing the number of features was also studied in the categorized version using tenfold cross-validation, as shown in Table 9. In the categorized mode, the proposed method achieved 98.9% for the accuracy parameter for the kNN classifier with 50 selected features, which is higher than the highest accuracy value obtained using the binary mode.
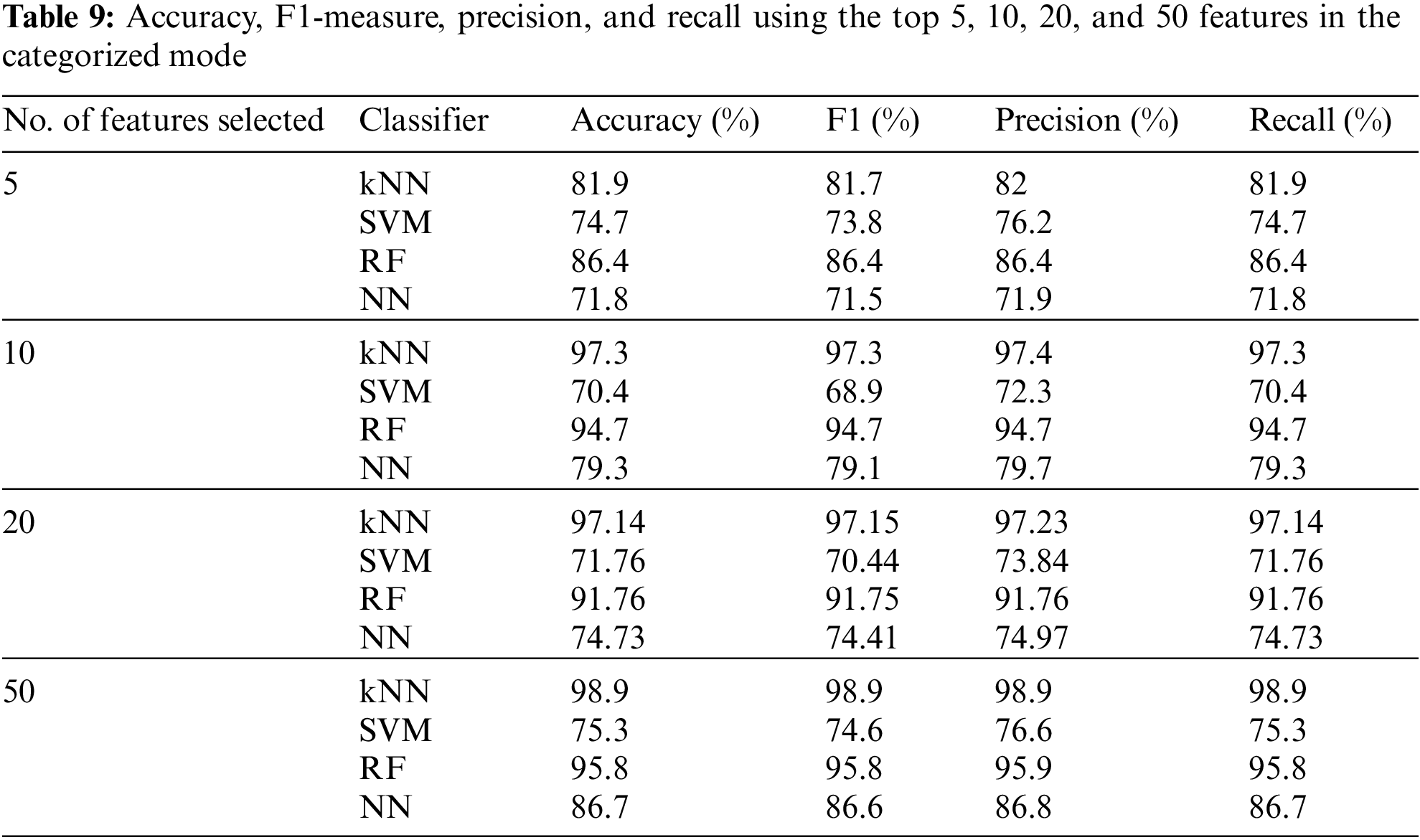
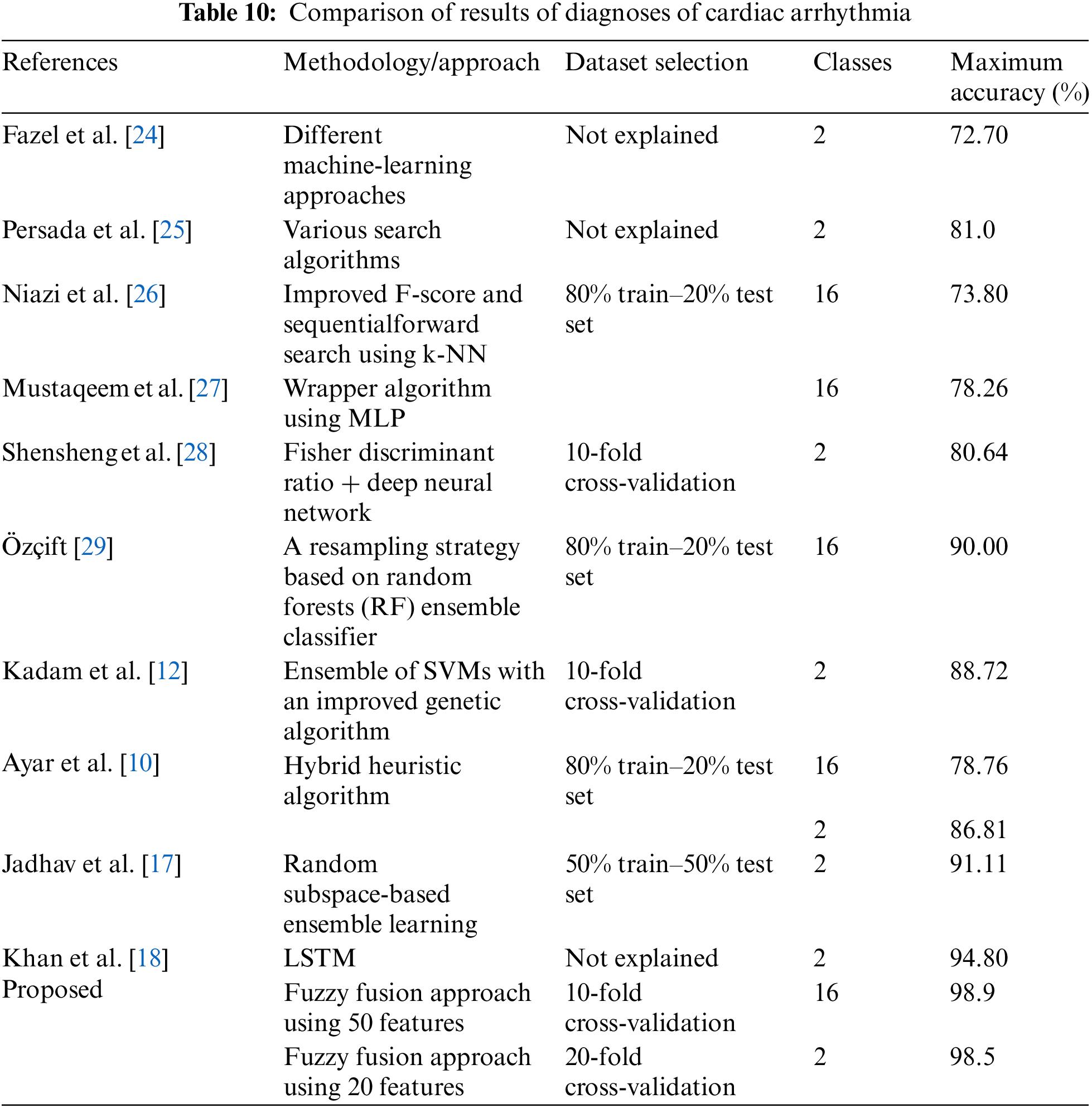
Table 10 presents a comparison of the diagnosis of cardiac arrhythmia results. It compares the proposed study with other leading studies using the same dataset. It concludes the maximum classification accuracy for ECG arrhythmia of the proposed method compared to the classification accuracies obtained by other methods in the literature. Table 10 shows that our proposed fuzzy fusion approach achieved superior accuracy compared to other methods. As shown in the table, Fazel et al. [24] achieved an accuracy of 72.7% using several machine-learning techniques. Persada et al. [25] achieved 81% maximum accuracy with the Best-First search using numerous search methods. Niazi et al. [26] achieved an accuracy of 73.8% using SVM and kNN. Mustaqeem et al. [27] achieved 78.26% accuracy by selecting the best features using a wrapper algorithm and various machine-learning classifiers. Shensheng et al. [28] achieved an accuracy of 80.6% using the Fisher discriminant ratio and deep neural networks using tenfold cross-validation. Özçift [29] proposed a resampling strategy based on RFs using an ensemble of classifiers and achieved 90% accuracy for 16 classes. Kadam et al. [12] used SVM with the aid of an improved GA to achieve 88.72% accuracy for only two classes. Ayar et al. [10], using a hybrid heuristic algorithm, achieved accuracies of 78.76% and 86.81% for 2 and 16 classes, respectively. Jadhav et al. [17] proposed an RS-based ensemble learning using a 50% train–50% set test and achieved an accuracy of 91.11%. Khan et al. [18] achieved 94.80% accuracy using the LSTM network using the binary mode. However, most of these state-of-the-art studies employed either a binary classifier (normal/abnormal) or 16 classes (categorized mode) for the classification problem, whereas our proposed model works for both modes. Our proposed study using the fuzzy fusion approach achieved accuracies of 98.5% and 98.9% for 2 and 16 classes, respectively. The results prove the superiority of our proposed model compared to other studies with cardiac arrhythmia datasets in terms of classification accuracy.
In this paper, an efficient fuzzy fusion approach is proposed to classify cardiac arrhythmia diseases. As the UCI arrhythmia dataset has 279 features, it was essential to fuse the top-ranked features generated from an ensemble of feature extraction algorithms. Therefore, in the proposed model, the fuzzy fusion approach was used to improve the feature-selection process. As a result, an ensemble of classifiers was used to classify arrhythmia diseases. The highest accuracies of 98.50% and 98.90% were obtained for the binary and categorized modes, respectively. These results were superior compared to those of state-of-the-art methods. The proposed model could help diagnose heart rhythm disease for students in an early stage with accuracy similar to professional doctors in this field. In the future, the model will be evaluated on some other standard datasets. Besides, we can also use deep learning models to improve our results.
Funding Statement: The authors received no specific funding for this study.
Conflicts of Interest: The authors declare that they have no conflicts of interest to report regarding the present study.
References
1. S. Sahoo, M. Dash, S. Behera and S. Sabut, “Machine learning approach to detect cardiac arrhythmias in ECG signals: A survey,” IRBM, vol. 86, pp. 10423, 2020. [Google Scholar]
2. S. Raj and K. C. Ray, “Sparse representation of ECG signals for automated recognition of cardiac arrhythmias,” Expert Systems with Applications, vol. 105, pp. 49–64, 2018. [Google Scholar]
3. T. Tuncer, S. Dogan, P. Pławiak and U. R. Acharya, “Automated arrhythmia detection using novel hexadecimal local pattern and multilevel wavelet transform with ECG signals,” Knowledge-Based Systems, vol. 186, pp. 104923, 2019. [Google Scholar]
4. H. Alshammari, S. A. El-Ghany and A. Shehab, “Big IoT healthcare data analytics framework based on fog and cloud computing,” Journal of Information Processing Systems, vol. 16, pp. 1238–1249, 2020. [Google Scholar]
5. A. Ismail, A. Shehab and I. El-Henawy, “Healthcare analysis in smart big data analytics: Reviews, challenges and recommendations,” in Security in Smart Cities: Models, Applications, and Challenges, Cairo, Egypt: Springer, Chapter 2, pp. 27–45, 2019. [Google Scholar]
6. V. Kadam, S. Jadhav and S. Yadav, “Bagging based ensemble of support vector machines with improved elitist GA-SVM features selection for cardiac arrhythmia classification,” International Journal of Hybrid Intelligent Systems, vol. 16, pp. 25–33, 2020. [Google Scholar]
7. S. K. Pandey, V. R. Sodum, R. R. Janghel and A. Raj, “ECG arrhythmia detection with machine learning algorithms,” in Data Engineering and Communication Technology, London, United Kingdom: Springer, Chapter 4, pp. 409–417, 2020. [Google Scholar]
8. K. -C. Chang, P. -H. Hsieh, M. -Y. Wu, Y. -C. Wang, J. -Y. Chen et al., “Usefulness of machine learning-based detection and classification of cardiac arrhythmias with 12-lead electrocardiograms,” Canadian Journal of Cardiology, vol. 37, pp. 94–104, 2021. [Google Scholar] [PubMed]
9. N. Singh and P. Singh, “Cardiac arrhythmia classification using machine learning techniques,” in Engineering Vibration, Communication and Information Processing, New York, USA: Springer, Chapter 11, pp. 469–480, 2019. [Google Scholar]
10. M. Ayar and S. Sabamoniri, “An ECG-based feature selection and heartbeat classification model using a hybrid heuristic algorithm,” Informatics in Medicine Unlocked, vol. 13, pp. 167–175, 2018. [Google Scholar]
11. H. Assodiky, I. Syarif and T. Badriyah, “Deep learning algorithm for arrhythmia detection,” in 2017 Int. Electronics Symp. on Knowledge Creation and Intelligent Computing (IES-KCIC), Keio, Surabaya, vol. 13, pp. 26–32, 2017. [Google Scholar]
12. V. J. Kadam, S. S. Yadav and S. M. Jadhav, “Soft-margin SVM incorporating feature selection using improved elitist GA for arrhythmia classification,” in Int. Conf. on Intelligent Systems Design and Applications, Vellor, India, vol. 76, pp. 965–976, 2018. [Google Scholar]
13. A. Darwaish, F. Naït-Abdesselam and A. Khokhar, “Detection and prediction of cardiac anomalies using wireless body sensors and bayesian belief networks,” ArXiv preprint arXiv:1904.07976, 2019. [Google Scholar]
14. S. Dalal and V. P. Vishwakarma, “GA based KELM optimization for ECG classification,” Procedia Computer Science, vol. 167, pp. 580–588, 2020. [Google Scholar]
15. J. Lang and F. Yang, “An improved classification method for arrhythmia electrocardiogram dataset,” in 2019 IEEE 2nd Int. Conf. on Information Communication and Signal Processing (ICICSP), Weihai, China, pp. 338–341, 2019. [Google Scholar]
16. N. Shandri and Z. Rustam, “Clustering arrhythmia multiclass using fuzzy robust kernel C-means (FRKCM),” in 2018 Int. Conf. on Applied Information Technology and Innovation (ICAITI), Padang, Indonesia, pp. 145–148, 2018. [Google Scholar]
17. S. Jadhav, S. Nalbalwar and A. Ghatol, “Feature elimination based random subspace ensembles learning for ECG arrhythmia diagnosis,” Soft Computing, vol. 18, pp. 579–587, 2014. [Google Scholar]
18. M. Khan, M. Karim and Y. Kim, “A two-stage big data analytics framework with real world applications using spark machine learning and long short-term memory network,” Symmetry, vol. 10, pp. 485, 2018. [Google Scholar]
19. S. S. Yadav and S. M. Jadhav, “Detection of common risk factors for diagnosis of cardiac arrhythmia using machine learning algorithm,” Expert Systems with Applications, vol. 163, pp. 113807, 2021. [Google Scholar]
20. S. B. Itzhak, S. S. Ricon, S. Biton, J. A. Behar and J. A. Sobel, “Effect of temporal resolution on the detection of cardiac arrhythmias using HRV features and machine learning,” Physiological Measurement, vol. 43, pp. 045002, 2022. [Google Scholar]
21. A. Z. Abualkishik, R. Almajed and S. A. Almutairi, “Early detection of cardiovascular diseases using deep learning feature fusion and MRI image analysis,” Fusion: Practice and Applications, vol. 8, pp. 16–24, 2022. [Google Scholar]
22. A. Koura and S. Hany, “Data mining algorithms for kidney disease stage prediction,” Journal of Cybersecurity and Information Management, vol. 1, pp. 21–29, 2020. [Google Scholar]
23. J. Dombi, “Membership function as an evaluation,” Fuzzy Sets and Systems, vol. 35, pp. 1–21, 1990. [Google Scholar]
24. A. Fazel, F. Algharbi and B. Haider, “Classification of cardiac arrhythmias patients,” CS229 Final Project Report, 2014. [Google Scholar]
25. A. G. Persada, N. A. Setiawan and H. A. Nugroho, “Comparative study of attribute reduction on arrhythmia classification dataset,” in 2013 Int. Conf. on Information Technology and Electrical Engineering (ICITEE), Padang, Indonesia, pp. 68–72, 2013. [Google Scholar]
26. K. A. K. Niazi, S. A. Khan, A. Shaukat and M. Akhtar, “Identifying best feature subset for cardiac arrhythmia classification,” in 2015 Science and Information Conf. (SAI), London, United Kingdom, pp. 494–499, 2015. [Google Scholar]
27. A. Mustaqeem, S. M. Anwar, M. Majid and A. R. Khan, “Wrapper method for feature selection to classify cardiac arrhythmia,” in 2017 39th Annual Int. Conf. of the IEEE Engineering in Medicine and Biology Society (EMBC), Hong Kong, Korea, pp. 3656–3659, 2017. [Google Scholar]
28. S. Shensheng Xu, M. -W. Mak and C. -C. Cheung, “Deep neural networks versus support vector machines for ECG arrhythmia classification,” in 2017 IEEE Int. Conf. on Multimedia & Expo Workshops (ICMEW), Hong Kong, Korea, pp. 127–132, 2017. [Google Scholar]
29. A. Özçift, “Random forests ensemble classifier trained with data resampling strategy to improve cardiac arrhythmia diagnosis,” Computers in Biology and Medicine, vol. 41, pp. 265–271, 2011. [Google Scholar]
Cite This Article
 Copyright © 2023 The Author(s). Published by Tech Science Press.
Copyright © 2023 The Author(s). Published by Tech Science Press.This work is licensed under a Creative Commons Attribution 4.0 International License , which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.


 Submit a Paper
Submit a Paper Propose a Special lssue
Propose a Special lssue View Full Text
View Full Text Download PDF
Download PDF Downloads
Downloads
 Citation Tools
Citation Tools