 Open Access
Open Access
ARTICLE
Molecular Fingerprinting of Three Ex-Situ Cultivated Populations of Acalypha gaumeri Pax & K. Hoffm (Female and Male) and Evaluation of Their Antifungal Activity Against Phytopathogens
1 Centro de Investigación Científica de Yucatán A.C., 43 # 130, x 32 y 34, Chuburná de Hidalgo, Mérida, Yucatán, México
2 Tecnológico Nacional de México, campus Conkal, Avenida Tecnológico S/N, Conkal, Conkal, Yucatán, México
* Corresponding Authors: Daisy Pérez-Brito. Email: ; Marcela Gamboa-Angulo. Email:
(This article belongs to the Special Issue: Plant-Derived Antimicrobials: Phytochemical Defense, Plant Metabolism, and Ecological Roles)
Phyton-International Journal of Experimental Botany 2026, 95(2), 3 https://doi.org/10.32604/phyton.2026.072668
Received 01 September 2025; Accepted 05 January 2026; Issue published 28 February 2026
Abstract
Acalypha gaumeri (Euphorbiaceae) is the only endemic species of the genus in the Yucatan Peninsula. It is dioecious and has antifungal properties against various phytopathogens. In the present study, molecular identification of A. gaumeri was performed using the rbcL region, confirming its belonging to the Acalypha genus. Its genetic diversity was evaluated using 10 SPAR markers (ISSR and DAMD) from 60 individuals collected from female and male plants of the Kiuic, Tinum and Yaxcaba ex-situ populations. The results showed a high level of genetic polymorphism (PIC = 0.980) and significant differences among the populations. Ethanol and aqueous extracts from leaves, stems, and roots of both genders and three populations were evaluated against three phytopathogenic fungi. Only the ethanol extracts of the roots showed inhibitory antifungal activity, whereas Kiuic and Tinum, both male and female individuals, showed inhibitory effects at 1000 µg/mL against the three pathogens. The ethanol extract of the female flowering plant of Kiuic showed activity at minimum inhibitory concentrations of 250 µg/mL against Alternaria chrysanthemi CICY004 and 500 µg/mL against Colletotrichum gloeosporioides CICY002 and Penicillium oxalicum ITC25. Chromatographic profile of the ethanol extracts confirmed the presence of alkaloidal components in the ex-situ cultivated plants, which were analyzed by HPLC. The results revealed that the peaks at TR of 7.60, 7.88, and 8.49 min were the most abundant components (9.95%–21.93%), with differences between female and male plants of the three populations. This research confirms that A. gaumeri cultivated and genetically characterized is a potential raw material to develop an eco-friendly product for the control of fungal diseases in crops.Keywords
Plant-pathogenic fungi cause significant economic losses [1]. Species of the Colletotrichum genus are considered to be among the “Top 10” in terms of their economic and scientific importance [2]. Alternaria chrysanthemi, for example, is responsible for losses in chrysanthemum species, which are the second most productive cut flower in Mexico [3,4], while Penicillium oxalicum causes stem rot in tomato plants and is a secondary pathogen of postharvest fruit of Capsicum chinensis [5]. Synthetic fungicides, such as mancozeb, chlorothalonil, and difenoconazole, are generally used to control phytopathogens [6]. These fungicides are effective to a certain extent, but they have adverse effects on human health and the environment. They also generate resistance to the pathogens [7]. Among viable alternatives to the safe replacement of synthetic fungicides are aqueous plant extracts. In a search for antifungal plant extracts to control phytopathogens, a botanical screening effort revealed that Acalypha gaumeri exhibits broad-spectrum antifungal activity.
In the Yucatan Peninsula, 14 species of Acalypha (Euphorbiaceae) have been documented. Among these, A. gaumeri Pax & K. Hoffm. is the sole species classified as an endemic, small and dioecious shrub [8,9]. Extracts of A. gaumeri have demonstrated insecticidal [10], nematicidal [11], and antifungal [12,13] activity on phytopathogens and parasites. Ethanol extracts (EEs) of A. gaumeri root inhibited mycelial growth (IMG) of Alternaria tagetica (ATTC 53771), A. chrysanthemi (CICY004), Corynespora cassiicola (ITC03), Alternaria sp. (ITC02), Colletotrichum capsici (CC4), Colletotrichum gloeosporioides (CG4), Curvularia sp. (ITC10) and Helmintosporium sp. (ITC04) [12,13]. Furthermore, the aqueous extracts of the root of A. gaumeri are effective in controlling leaf blight in chrysanthemum, which is caused by A. chrysanthemi [14]. Samples from three different populations of A. gaumeri collected in Oxkutzcab, Tinum and Yaxcaba (Yucatan state, Mexico) showed antifungal activity against A. chrysanthemi (CICY004), C. gloeosporioides (CG4) and Pseudocercospora fijiensis [15]. The three populations of plants with female flowers and male flowers are maintained in ex-situ propagation in the experimental nursery of Centro de Investigación Científica de Yucatán A. C. (CICY). Therefore, these ex-situ cultivars should be evaluated for biological efficacy and their spectrum of activity against other phytopathogenic fungi expanded.
The species A. gaumeri (Fig. 1) has been identified morphologically by botanical characteristics (Pax, F. and K. Hoffmann) and molecularly by amplification of the ITS, ndhF, and trnL-F regions [16]. Molecular studies are relevant for species identification and have been used to complement accurate and reliable characterization [17,18,19]. This is the first step in ensuring the origin of an extract’s efficacy, quality and safety [20]. In recent years, single-primer amplification reaction (SPAR) methods have been used to detect genetic variation in plants, particularly in poorly studied species. This method includes targeted amplification of Deoxyribonucleic acid (DNA), Random amplification of polymorphic DNA (RAPD), inter-simple sequence repeat (ISSR), and Directed amplification of minisatellite DNA (DAMD) [21]. Genetic diversity studies have been conducted on various members of the family Euphorbiaceae. For example, the genetic diversity of 35 sun poinsettia (Euphorbia spp.) genotypes, collected in Morelos, México, were examined using RAPD molecular markers [22]. Sapium macrocarpum, collected in three distinct locations within the medium-subtropical evergreen forest of the Selva, El Ocote Biosphere Reserve (REBISO) in Chiapas, Mexico, exhibited significant genetic diversity. However, minimal differentiation was observed among the localities [23].
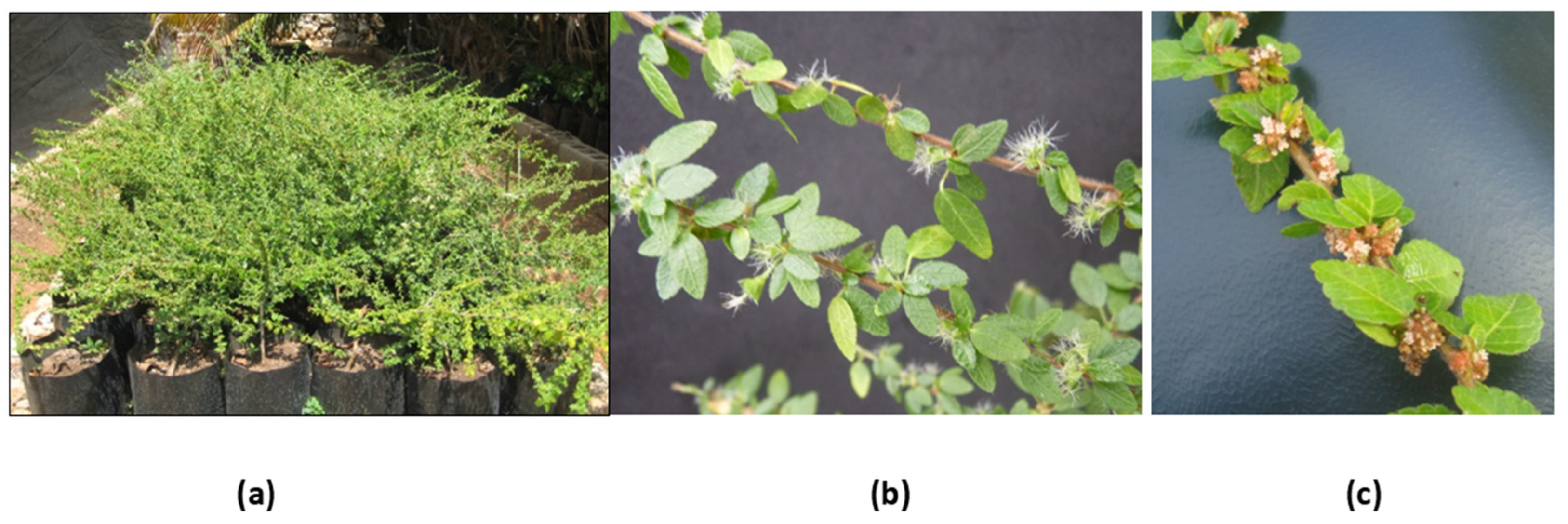

Figure 1: Acalypha gaumeri (a) Plants ex-situ cultivated, (b) female flowers and (c) male inflorescence.
This study contributes to ongoing research on A. gaumeri. The aims were the molecular identification of A. gaumeri collected in the state of Yucatan, the analysis of the genetic fingerprint of three Kiuic, Tinum and Yaxcaba populations (females and males) grown ex-situ using SPAR-type molecular markers and the evaluation of the antifungal activity of aqueous and ethanol extracts of the roots, stems and leaves against A. chrysanthemi, Colletotrichum gloeosporioides and P. oxalicum, and also the chromatographic profile of the ethanol extracts of these ex-situ plants cultivated.
This study utilised A. gaumeri plants (both male and female) from the Kiuic (Municipality of Oxkutzcab), Tinum, and Yaxcaba populations. These were grown in pots from cuttings at the experimental nursery, Centro de Investigación Científica de Yucatán A. C.
Sixty samples of young, symptomless leaves, preferably from the basal part, were collected from 10 male and 10 female plants from each population in November 2022. These fresh samples (3 g) were washed and frozen at −80°C for molecular identification.
Genomic DNA was extracted from the samples that had been individually deep-frozen using the method established by Tapia-Tussell et al. [24]. DNA concentration (ng/μL) and purity (A260/A280) were measured using a spectrophotometer (Nanodrop 2000, Madrid, Spain). DNA integrity was determined by 0.8% agarose gel electrophoresis. Samples were diluted to 20 and 5 ng/μL concentration with sterile ultrapure water and stored at −20°C for subsequent use.
2.3 Molecular Identification of Acalypha gaumeri
Genomic DNA obtained from leaves of a female Yaxcaba (YF) plant with the primers rbcL-R and rbcL-F was amplified [25]. The amplicons were sent to Macrogen in South Korea for sequencing. The sequences obtained were edited using BioEdit software version 7.0.5.3 [26] and compared with the National Centre for Biotechnology Information GenBank database (http://blast.ncbi.nlm.gov) using the BLAST program.
A phylogenetic tree was constructed using the Neighbour-Joining method [27] with MEGA software version 11 to identify samples at the species level. An in silico analysis was conducted to compare the restriction profiles of the A. gaumeri sample under study with those of a sequence of A. chamaedrifolia. This analysis utilized a total of 60 restriction enzymes and was performed using the pDRAW32 program edition 1.1.147 [28].
2.4 Analysis with ISSR and DAMD Markers
The genetic diversity of the three A. gaumeri populations was determined using 10 SPAR markers. The markers comprised 5 ISSR and 5 DAMD loci [27,29]. The ISSR markers were IS01, IS21, IS14, IS17 and IS19. The DAMD markers were HBV, M13, HVA, 33.6 and HVR (Table 1).
Table 1: List of SPAR markers used to obtain the genetic fingerprint of Acalypha gaumeri populations.
| Type of Primer | Primer | Sequence |
|---|---|---|
| ISSR | IS01 | 5′-GAC AGA CAG ACA GAC A-3′ |
| IS21 | 5′-GTC GTC GTC GTC GTC GTC-3′ | |
| IS14 | 5′-AGA GAG AGA GAG AGA GTA-3′ | |
| IS17 | 5′-TGT GTG TGT GTG TGT GGT-3′ | |
| IS19 | 5′-AGA GAG AGA GAG AGA GTC-3′ | |
| DAMD | HBV | 5′-GGT GTA GAG AGG GGT-3′ |
| M13 | 5′-GAG GGT GGC GGT TCT-3′ | |
| HVA | 5′-AGG ATG GAA AGG AGG C-3′ | |
| 33.6 | 5′-AGG GCT GGA GG-3′ | |
| HVR | 5′-CCT CCT CCC TCC T-3′ |
The polymerase chain reaction (PCR) reactions for the ISSR primers included a final volume of 25 µL for each reaction, containing 10 ng of genomic DNA and 1× PCR buffer (Invitrogen, Carslbad, CA, USA), 2.5 mM of MgCl2 (Invitrogen, Carslbad, CA, USA), 0.25 mM of dNTPs (Invitrogen, Carslbad, CA, USA), 1 µM primer and 1U of Taq DNA polymerase (Invitrogen, Carslbad, CA, USA). The amplification reactions were performed on a C1000 Touch (Bio-Rad, Hercules, California, USA). Amplification conditions included an initial denaturation at 94°C for 2 min, followed by 36 cycles of amplification, each at 94°C for 40 s, 54°C for 45 s, and 72°C for 1.5 min, and a final extension step at 72°C for seven min.
The PCR reactions for the DAMD primers were performed using the methodology outlined by Verma and Rana (2013). The final volume of each reaction was 25 µL containing 60 ng of genomic DNA, 1× PCR buffer (Invitrogen, Carslbad, CA, USA), 2 mM of MgCl2 (Invitrogen, Carslbad, CA, USA), 0.25 mM of DNTP’s (Invitrogen, Carslbad, CA, USA), 0.2 µm primer and 1U de Taq DNA polymerase (Invitrogen, Carslbad, CA, USA). Amplification included an initial denaturation for 5 min at 92°C, followed by 40 cycles of amplification, each at 94°C for 1 min, 55°C for 2 min, and 72°C for 2 min, and a final extension step for seven min at 72°C.
The fragments obtained with ISSR and DAMD markers were separated at 6% on polyacrylamide gels in a 29:1 ratio (acrylamide: bisacrylamide) (Millipore-Sigma, St. Louis, MO, USA). The gels were run at 200 V for 2.5 h in 1× Tris-glycine (TG) buffer (Millipore-Sigma, St. Louis, MO, USA). The DNA band sizes were subsequently determined relative to the 1 Kb molecular marker (Invitrogen, Carlsbad, CA, USA). Gel staining was performed using the silver staining technique [30].
Roots of adult plants of A. gaumeri were collected in September at the CICY nursery after two years of culture. Each population (3) was separated by the sex of the flowers (female or male) and the type of plant tissue (leaf, stem, and root). Roots were washed with plenty of water, dried for 4 weeks in natural light at 40°C, and then crushed using a Pagani mill (Pagani-Dycomet, SA de CV, Mexico DF, Mexico) with a mesh number of 5. Finally, the samples obtained were extracted with water and ethanol (3 × 2 × 3× 2), resulting in 36 extracts.
The dried and ground plant material of each A. gaumeri sample (30 g) was macerated with ethanol (300 mL) at 24°C for 2 days (3 times). The solvent with the sample was filtered on paper (Whatman no. 1) and concentrated under reduced pressure in a rotary evaporator (IKA RV-10, Staufen, Germany) until the dry ethanol extract was obtained. The samples were stored under refrigerated conditions (4°C) until they were used.
The dried and ground plant material (1.5 g) was extracted with distilled water (20 mL), heated to boiling for 3 min, and then cooled to room temperature. The mixture was then filtered using cotton gauze and Whatman no. 1 filter paper. The filtered volume was adjusted to 25 mL to obtain the aqueous extract at an initial concentration of 6% w/v. The aqueous extracts (AEs) were sterilized using a 0.22 µm Millipore filter (Merck-Millipore, Burlington, MA, USA), lyophilized, and stored frozen for evaluation.
The phytopathogenic fungi (Table 2) used to evaluate antifungal activity were obtained from the Biotechnology Unit’s stock at the CICY and correspond to fungal isolates collected from plants in the region. They were grown on potato dextrose agar (PDA) for 7 days. A. chrysanthemi (CICY004) was maintained at 18 ± 2°C in the dark, while C. gloeosporioides (CICY002) and P. oxalicum (ITC25) were maintained at 23 ± 2°C for seven days [13].
Table 2: Phytopathogenic strains tested for antifungal activity in Acalypha gaumeri extracts.
| Strain | Key | GenBank Number | Host |
|---|---|---|---|
| Alternaria chrysanthemi | CICY004 | MH846127.1 | Dendranthema grandiflorum |
| Colletotrichum gloeosporioides | CICY002 | HM562713.1 | Carica papaya |
| Penicillium oxalicum | ITC25 | KP868627 | Capsicum chinense |
4.1 Antifungal Assay by Microdilution
The spore suspension of A. chrysanthemi was prepared at 2.5 × 104 spores/mL, while those of C. gloeosporioides and P. oxalicum were prepared at 1 × 105 spores/mL from PDA medium.
The ethanol extracts were resuspended in a mixture of dimethyl sulfoxide (DMSO) and 0.5% Tween 20 at a concentration of 80 μg/μL (stock solution). The AEs were suspended with sterile distilled water at a concentration of 6% w/v (stock solution). A serial dilution was performed using the stock solution in Roswell Park Memorial Institute (RPMI) 1640 medium (1:1 v/v) to a final volume of 100 μL/per well. Subsequently, 100 μL of the spore suspension was added to each well, bringing the final volume to 200 μL. The final concentrations of the ethanol extracts were 2000, 1000, 500, 250, 125 and 62.5 μg/mL with 0.62% of 0.5%-DMSO-tween 20 solution. Concerning the aqueous extracts, each one (100 μL) was deposited in a microwell, after which the spore suspension (100 μL) was added at a final concentration of 3% w/v. The negative growth control was the blank, RPMI 1640 medium and spore suspension with 0.5% DMSO-tween 20 solution; and the positive control was the commercial fungicide prochloraz (CE45) at 0.11% (w/v) and 40% DMSO solution. All samples were replicated four times. As previously indicated, the microplates were incubated at different temperatures and photoperiod conditions. The assay was terminated at 96 h, and plate reading were performed at 24, 48, 72, and 96 h using the National Committee for Clinical Laboratory Standards [13,31] numerical scale. The percentages of mycelial growth inhibition (MGI) were determined using the Abbott formula [(%MGI in the negative control—%MGI in the treatment)/%MGI in the negative control] × 100 [32].
4.2 Fungicidal or Fungistatic Effect
Samples (10 μL) without fungal growth in the microwells were transferred to Petri dishes containing PDA medium. These samples were incubated under the same conditions previously outlined for each pathogen, and their progress was observed for 3 days. The effect of the extract was classified as fungicidal, indicated by the absence of mycelial growth, and fungistatic as evidenced by hyphal growth after several days to weeks of observation.
The presence or absence of each PCR amplicon was determined by assigning a value of “1” for the presence of the fragment and a value of “0” for its absence. Binary data obtained from ISSR and DAMD profiles were used to construct a similarity matrix using Jaccard coefficients [33] and the Neighbour-Joining method to generate a dendrogram with the Sequential Agglomerative Hierarchical Nested Agglomerative Clustering (SAHN) module of the NTSYS pc software, version 2.21q [34].
Principal component analysis (PCoA) was performed to differentiate the A. gaumeri accessions. Polymorphic information content (PIC) values were also calculated using the equation developed by Anderson et al. [35].
One-way analysis of variance was performed on the percentage inhibition of mycelial growth data, with the original data first being transformed using the formula y = arsin[√(y/100)]. Treatment means were compared using the Tukey multiple range test (p = 0.05). Analyses of variance were performed using SAS version 9.4 for Windows.
The presence of alkaloids in the ethanol extracts was detected using Dragendorff reagent with a butanol:acetic acid:water (7:1:2, v/v/v) eluent system, as previously reported by Vargas-Díaz et al. [15]. The orange spots detected on the plate were recorded as a frontal reference (FR).
4.5 Liquid Chromatography Analysis
The liquid chromatograms of the ethanol extracts were obtained using an Agilent Technologies 1260 chromatograph coupled with a multichannel UV/Visible detector. The six ethanol extracts of the roots of A. gaumeri ex-situ cultivated and a sample collected in the field were analyzed. The samples were diluted to 1 mg/mL in HPLC-grade methanol and then filtered through a 0.22 µm Millipore filter. A 20 μL aliquot of each sample was injected into a Grace C18 column with an internal diameter of 4.6 mm, a length of 250 mm and particles measuring 5 µm in thickness. The mobile phase consisted of a mixture of solvents A: 30 mM of sodium acetate buffer (80 mL), acetonitrile (20 mL) and acetic acid (1 mL); and B: acetonitrile (45 mL), methanol (45 mL), water (10 mL), and acetic acid (0.25 mL). The method starts with A:B 40:60 (6 min), 0:100 (2 min), 40:60 (2 min), 0:100 (1 min). The retention times (Rt) of the peaks (min) were recorded at a wavelength of 280 nm, and the percentage of absorption area was calculated.
Leaf samples of A. gaumeri (60) were obtained from female and male plants of the Kiuic (Oxkutzcab), Tinum and Yaxcaba populations ex-situ cultivated at CICY. The genomic DNA concentration of each sample ranged from 142 to 1500 ng/µL, with a mean of 821 ng/µL. The DNA purity value (A260/A280 ratio) ranged from 1.74 to 2.03. Furthermore, a 330 base pairs (bp) PCR product was obtained for all samples when amplified from the 16S ribosomal region of the chloroplast, indicating that there were no PCR reaction inhibitors in these DNA samples.
The sequence obtained with the rbcL primers was compared with sequences deposited in NCBI, showing identities values of 98–100% with different species of the genus Acalypha (Table 3). In order to corroborate the identification of the sample under study, a phylogenetic tree was constructed using the neighbour-joining method (Fig. 2). The phylogenetic analysis unequivocally demonstrated that the A. gaumeri specimen is distinctly categorized within the clade corresponding to the genus Acalypha. However, the sequence obtained was not directly grouped with any of the other species analyzed; instead, it was positioned on an independent branch, which demonstrates its molecular differentiation from the other species of the genus. The final outcome of the sequence obtained from A. gaumeri was deposited in GenBank with the accession number PP819641.1.
Table 3: The accession number and maximum identity of the Acalypha species.
| Specie | Accession | Percentage of Identity | Query Cover |
|---|---|---|---|
| A. gaumeri | PP819641.1 | 100 | 100 |
| A. chamaedrifolia | MH549716.1 | 100 | 99 |
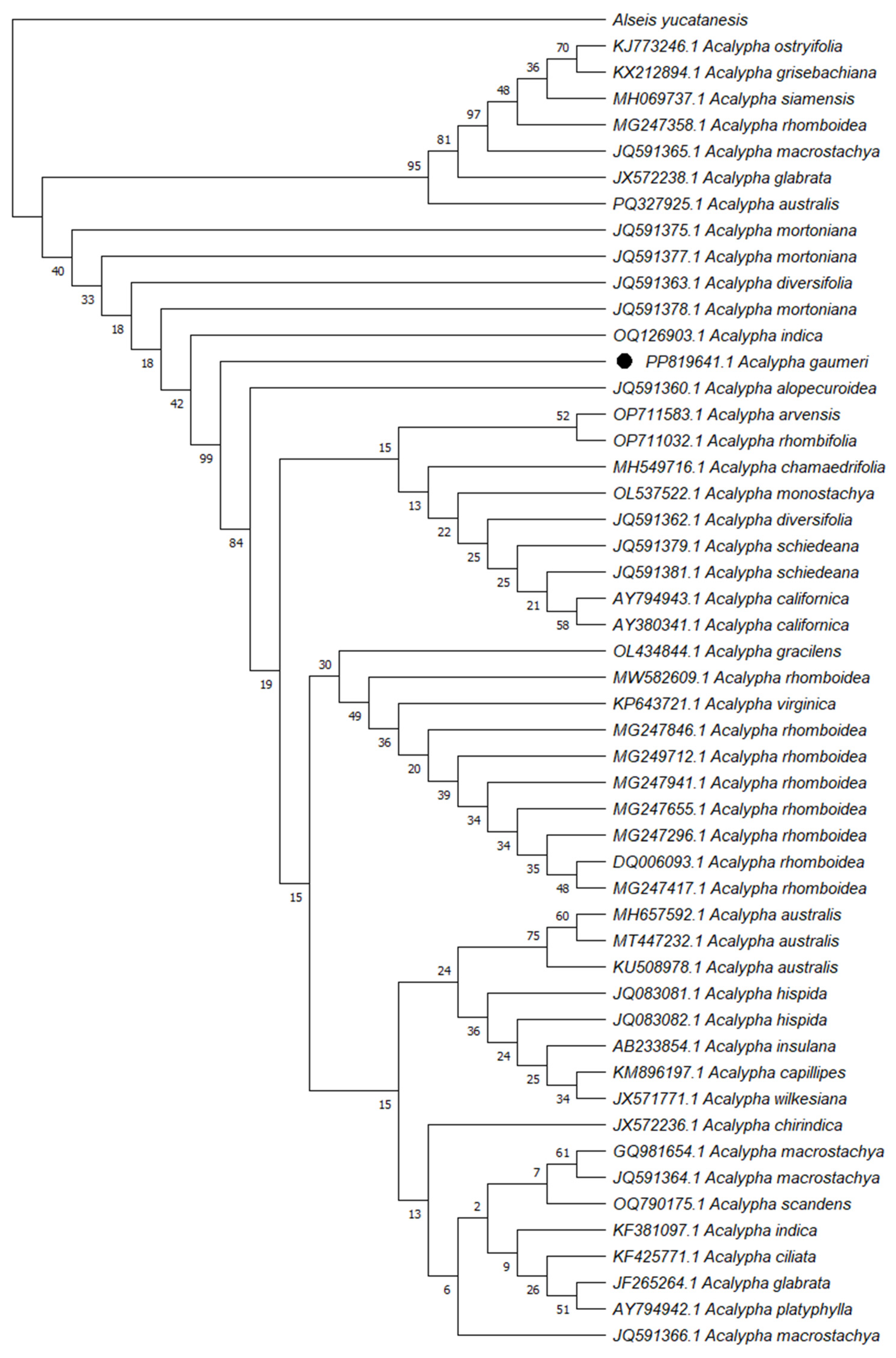
Figure 2: Phylogenetic analysis of Acalypha gaumeri was inferred using the neighbour-joining method with 1000 bootstrap replicates, performed in MEGA 11 with the external sequence of Alseis yucatanensis.
Among the 10 microsatellite PCRs, several samples did not amplify with the ISSR 14 and ISSR 21 markers and were therefore discarded from the analysis. With the remaining three ISSRs and the five DAMDs, a PIC value of 0.980 was obtained, indicating a highly significant correlation. Tests with DAMD primers demonstrated an average polymorphism value of 0.984, higher than that of ISSR primers (0.975). The HVR (0.999) and 33.6 (0.995) markers provided the most polymorphic information in this study.
A dendrogram of the 60 samples under study was generated (Fig. 3) with data from the absence-presence matrices of the 10 primers processed (5 ISSR + 5 DAMD). The dendrogram analysis revealed two large clades (Fig. 3, clade 2 and 3) and one small clade (clade 1). The 60 samples were almost equally divided as follows: 4 samples were allocated to clade 1; 33 samples to clade 2; and 21 to clade 3. A substantial genetic discrepancy was identified between the samples. The most genetically similar accessions, as indicated by a similarity coefficient of 0.44, were TF5 and TF6 from the propagated plants of the Tinum population in clade 2c. Accessions YF3 and YF4, which exhibited a similarity coefficient of 0.34, are derived from the Yaxcaba population and are classified within subclade 3a (Fig. 3).
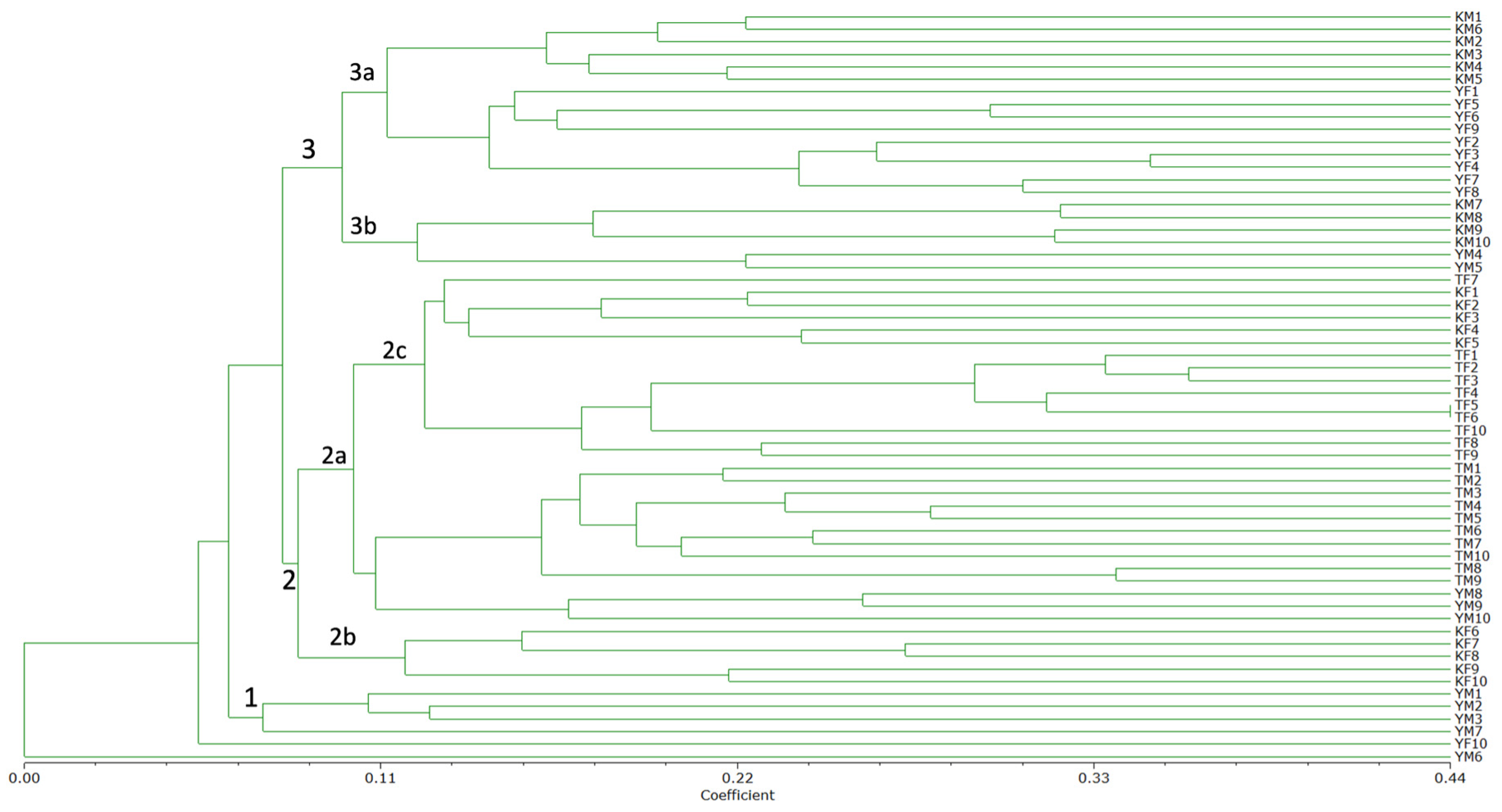
Figure 3: ISSR and DAMD consensus of the unweighted pair group arithmetic mean (UPGMA) dendrogram based on Jaccard’s coefficient, showing relationships among 60 Acalypha gaumeri samples determined using cumulative data. The numbers 1, 2, and 3 are indicative of the primary clades, while each number accompanied by a letter (a, b, or c) signifies the subclades that are formed according to their respective groupings.
The second clade (subclades 2a and 2b) comprises ten samples of female Kiuic, ten samples of female and male Tinum, and three samples of male Yaxcaba. Clade 3 comprises ten samples from the male Kiuic population, eight samples from the female Yaxcaba population, and two samples from the male Yaxcaba population. Conversely, clade 1 comprises four samples from the male Yaxcaba population and a single sample from the female Yaxcaba population (Fig. 3). The majority of the samples classified within clade 2 exhibited geographical proximity to specimens from the Tinum population. In contrast, individuals from clade 3 exhibited increased dispersion, as demonstrated by specimens from the Kiuic and Yaxcaba populations.
In the two-dimensional representation of the data (Fig. 4A), all clusters demonstrate a clear separation between clades. However, the three-dimensional graph provides a more detailed illustration of this separation between clades (Fig. 4B).
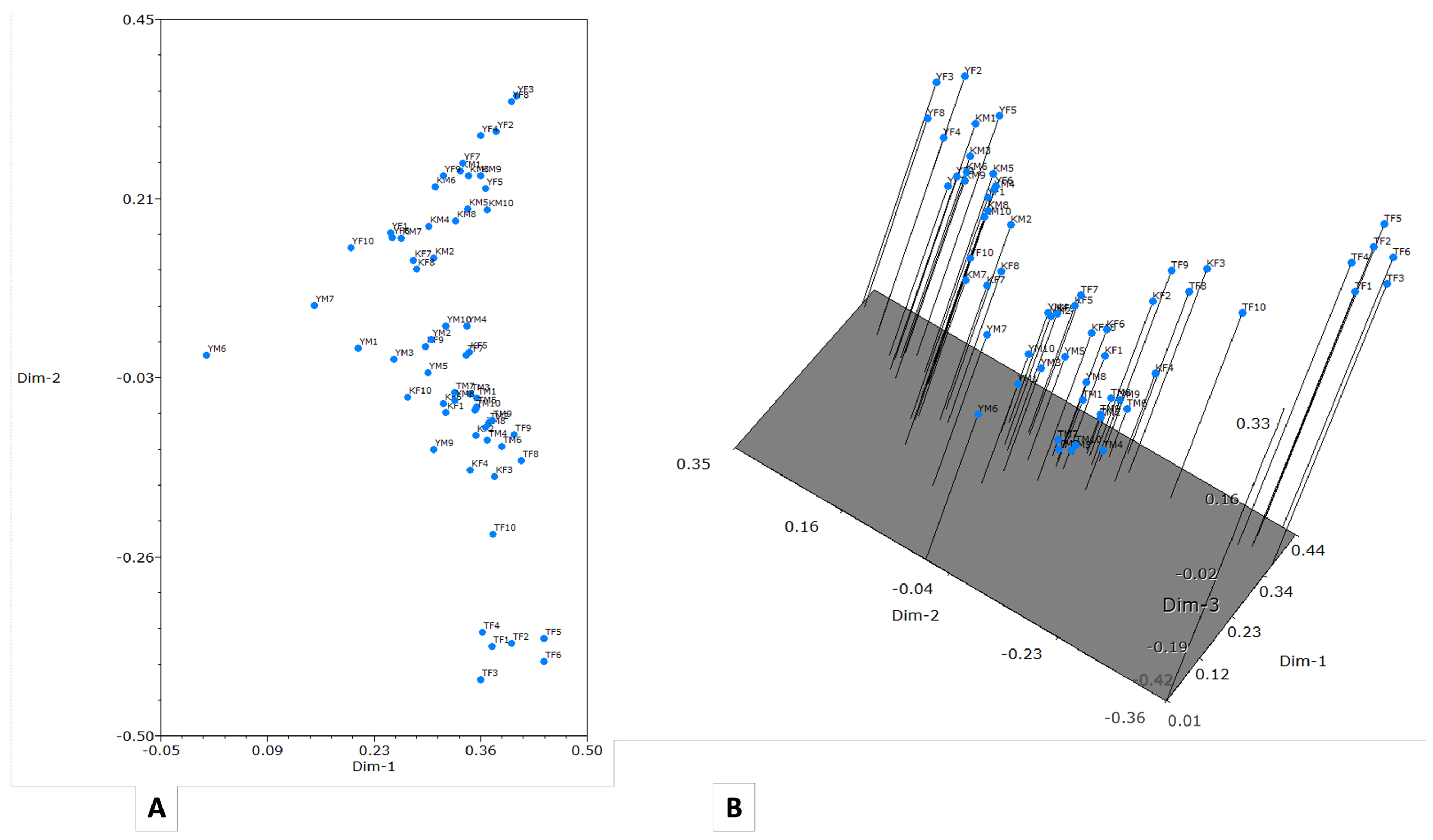
Figure 4: Principal component analysis obtained with NTSYS software, version 2.21q. (A) Two-dimensional plot; (B) Three-dimensional plot.
5.4 Extraction Yield of Dried and Ground Roots
The yield of the extracts is shown in Table 4. In general, the leaves yielded the highest ethanol extract, ranging from 9.27% to 16.61%. The stems yielded between 1.79% and 3.74%, while the roots yielded between 1.88% and 2.49%. The highest yield was obtained from the leaves of the Kiuic female population, at 16.61%.
Conversely, the aqueous extracts from the leaves of A. gaumeri across the three populations had higher solids yields after freeze-drying, ranging from 15.83% to 25.8%, compared to the stem (0.13% to 3.23%) or roots (0.56% to 11.26%). The highest yield was found in the female Kiuic leaves (25.53%), while the lowest yield was found in the male leaves of the same population (12.4% solids) (Table 4).
Table 4: Percentage yield of the ethanol and aqueous extracts from three populations of Acalypha gaumeri.
| Population | Plant Tissue | Flower Sex | Key | Yield EE (%) | Yield Lyophilized AE (%) |
|---|---|---|---|---|---|
| Kiuic | L | F | KLF | 16.61 | 25.53 |
| M | KLM | 11.79 | 12.4 | ||
| S | F | KSF | 3.12 | 0.13 | |
| M | KSM | 3.74 | 3.2 | ||
| R | F | KRF | 2.01 | 0.56 | |
| M | KRM | 2.31 | 2.36 | ||
| Tinum | L | F | TLF | 11.52 | 25.8 |
| M | TLM | 10.66 | 18.03 | ||
| S | F | TSF | 1.79 | 2.43 | |
| M | TSM | 3.00 | 3.23 | ||
| R | F | TRF | 2.35 | 2.96 | |
| M | TRM | 2.13 | 0.83 | ||
| Yaxcaba | L | F | YLF | 9.73 | 15.83 |
| M | YLM | 9.27 | 19.56 | ||
| S | F | YSF | 2.81 | 2.4 | |
| M | YSM | 2.81 | 1.83 | ||
| R | F | YRF | 2.49 | 3.23 | |
| M | YRM | 1.88 | 11.26 |
5.5 Antifungal Activity of Acalypha gaumeri Extracts
The antifungal activity of the 36 extracts (ethanol and aqueous) of leaves, stems, and roots of A. gaumeri, male and female specimens from three populations cultivated in ex-situ conditions, against A. chrysanthemi, C. gloeosporioides and P. oxalicum showed few differences between flower sex, but high differences between populations were found (Table 5). All ethanol extracts obtained from the roots of the three populations of A. gaumeri exhibited inhibitory effects against the three phytopathogens at 2000 µg/mL. However, only the ethanol extracts of female plant stems from Tinum (TSF; MGI = 50%), Yaxcaba (YSF), and plant leaves (YLF) exhibited a slight (MGI = 25%–50%) antifungal effect on at least one of the phytopathogens tested.
Table 5: Percentage of mycelial growth inhibition (MGI) of ethanol (2000 µg/mL) and aqueous (3% w/v) extracts from Acalypha gaumeri (leaves, stem, and roots) of three populations ex-situ cultivated against three fungal phytopathogens.
| Population/Controls | Plant Tissue | Flower Sex | Key/Control | Mycelial Growth Inhibition (%) | |||||
|---|---|---|---|---|---|---|---|---|---|
| Alternaria chrysanthemi | Colletotrichum gloeosporioides | Penicillium oxalicum | |||||||
| EE | AE | EE | AE | EE | AE | ||||
| Kiuic | L | F | KLF | 0 | 0 | 0 | 0 | 0 | 0 |
| M | KLM | 0 | 0 | 0 | 0 | 0 | 0 | ||
| S | F | KSF | 0 | 0 | 0 | 0 | 0 | 0 | |
| M | KSM | 0 | 0 | 0 | 0 | 0 | 0 | ||
| R | F | KRF | 100 | 0 | 100 | 0 | 100 | 0 | |
| M | KRM | 100 | 0 | 100 | 0 | 100 | 0 | ||
| Tinum | L | F | TLF | 0 | 0 | 0 | 0 | 0 | 0 |
| M | TLM | 0 | 0 | 0 | 0 | 0 | 0 | ||
| S | F | TSF | 0 | 0 | 50 | 0 | 0 | 0 | |
| M | TSM | 0 | 0 | 0 | 0 | 0 | 0 | ||
| R | F | TRF | 100 | 0 | 100 | 0 | 100 | 0 | |
| M | TRM | 100 | 0 | 100 | 0 | 100 | 0 | ||
| Yaxcaba | L | F | YLF | 25 | 0 | 0 | 0 | 0 | 0 |
| M | YLM | 0 | 0 | 0 | 0 | 0 | 0 | ||
| S | F | YSF | 25 | 0 | 0 | 0 | 0 | 0 | |
| M | YSM | 0 | 0 | 0 | 0 | 0 | 0 | ||
| R | F | YRF | 100 | 0 | 100 | 0 | 100 | 0 | |
| M | YRM | 100 | 0 | 100 | 0 | 100 | 0 | ||
| NC | 0 | 0 | 0 | 0 | 0 | ||||
| SS | 0 | 0 | 0 | 0 | 0 | ||||
| Blank | 0 | 0 | 0 | 0 | 0 | ||||
| PC | 100 | 100 | 100 | 100 | 100 | ||||
There are no aqueous extracts from A. gaumeri with activity against the three fungal pathogens evaluated at 3% w/v (Table 5). Therefore, the minimum inhibitory concentration (MIC) was determined using serial dilutions of the six ethanol extracts that were active against each evaluated fungus (Table 6, Table 7 and Table 8).
The antifungal effects of the six ethanol extracts were inhibitory at 2000 and 1000 µg/mL (Table 6). These same extracts showed a statistically significant difference in the percentage of MGI at 500, 250, and 125 µg/mL against A. chrysanthemi, with MGI values ranging from 0% to 100%. The lowest MIC, at 250 µg/mL, was observed with the KRF and TRM ethanol extracts, indicating a fungicidal effect. Followed by KRM and TRF with MIC values of 500 µg/mL and fungistatic effect (Table 6).
Table 6: Minimum inhibitory concentration (MIC) of ethanol extracts from the roots of female and male Acalypha gaumeri plants from three populations ex-situ cultivated against Alternaria chrysanthemi.
| Population | Sample | Alternaria chrysanthemi (% MGI) Ethanol Extract Concentration (μg/mL) | MIC (μg/mL) | |||||
|---|---|---|---|---|---|---|---|---|
| 2000 | 1000 | 500 | 250 | 125 | 62.5 | |||
| Kiuic | KRF | 100a | 100a | 100a | 100a | 50b | 0b | 250++ |
| KRM | 100a | 100a | 100a | 50c | 0d | 0b | 500+ | |
| Tinum | TRF | 100a | 100a | 100a | 75b | 25c | 0b | 500+ |
| TRM | 100a | 100a | 100a | 100a | 50b | 0b | 250++ | |
| Yaxcaba | YRF | 100a | 100a | 75b | 50c | 0d | 0b | 1000+ |
| YRM | 100a | 100a | 75b | 25d | 0d | 0b | 1000+ | |
| NC | 0b | 0b | 0c | 0e | 0d | 0b | ||
| SS | 0b | 0b | 0c | 0e | 0d | 0b | ||
| Blank | 0b | 0b | 0c | 0e | 0d | 0b | ||
| PC | 100a | 100a | 100a | 100a | 100a | 100a | ||
5.5.2 Colletotrichum gloeosporioides
The six ethanol extracts were inhibitory at concentrations of 2000, 1000, and 500 µg/mL, except for KRM, which had MGI values of 75% on C. gloeosporioides (Table 7). At 250 µg/mL, the six ethanol extracts showed statistically significant differences in their MGI percentages, ranging from 0% to 100%. Therefore, KRF, TRF, and TRM ethanol extracts showed the lowest MICs of 500 µg/mL against C. gloeosporioides, exhibiting a fungicidal effect (Table 7).
Table 7: Minimum inhibitory concentration (MIC) of ethanol extracts from the roots of female and male Acalypha gaumeri plants from three populations ex-situ cultivated against Colletotrichum gloeosporioides.
| Population | Sample | Colletotrichum gloeosporioides (% MGI) Ethanol Extract Concentration (μg/mL) | MIC (μg/mL) | |||||
|---|---|---|---|---|---|---|---|---|
| 2000 | 1000 | 500 | 250 | 125 | 62.5 | |||
| Kiuic | KRF | 100a | 100a | 100a | 50c | 0b | 0b | 500++ |
| KRM | 100a | 100a | 75b | 0e | 0b | 0b | 1000+ | |
| Tinum | TRF | 100a | 100a | 100a | 75b | 0b | 0b | 500++ |
| TRM | 100a | 100a | 100a | 0e | 0b | 0b | 500++ | |
| Yaxcaba | YRF | 100a | 100a | 0c | 0e | 0b | 0b | 1000+ |
| YRM | 100a | 100a | 0c | 25d | 0b | 0b | 1000+ | |
| NC | 0b | 0b | 0c | 0e | 0b | 0b | ||
| SS | 0b | 0b | 0c | 0e | 0b | 0b | ||
| Blank | 0b | 0b | 0c | 0e | 0b | 0b | ||
| PC | 100a | 100a | 100a | 100a | 100a | 100a | ||
The six ethanol extracts of A. gaumeri from the three populations exhibited inhibitory effects on P. oxalicum at 2000 and 1000 µg/mL. The serial dilution at 500–62.5 µg/mL indicated statistically significant differences in the MGI for the pathogenic fungi. The KRF extract was the only sample to exhibit the lowest MIC (500 µg/mL) against P. oxalicum and a fungicidal effect. In addition, only RKF induced MGI of 75% at 250 µg/mL (Table 8).
The ethanol extract of KRF demonstrated the capacity to inhibit the growth of the three test fungal pathogens, with MIC values ranging from 250 to 500 µg/mL. This extract exhibits a fungicidal effect against the three phytopathogens (Table 8).
Table 8: Minimum inhibitory concentration (MIC) of the ethanol extracts (EE) from the roots of female and male Acalypha gaumeri plants from three populations ex-situ cultivated against Penicillium oxalicum.
| Population | Sample | Penicillium oxalicum (% MGI) Ethanol Extract Concentration (μg/mL) | MIC (μg/mL) | |||||
|---|---|---|---|---|---|---|---|---|
| 2000 | 1000 | 500 | 250 | 125 | 62.5 | |||
| Kiuic | KRF | 100a | 100a | 100a | 75b | 50b | 25b | 500++ |
| KRM | 100a | 100a | 75b | 50c | 25c | 0c | 1000+ | |
| Tinum | TRF | 100a | 100a | 75b | 50c | 25c | 0c | 1000+ |
| TRM | 100a | 100a | 50c | 25d | 0d | 0c | 1000+ | |
| Yaxcaba | YRF | 100a | 100a | 75b | 50c | 25c | 0c | 1000+ |
| YRM | 100a | 100a | 75b | 50c | 25c | 0c | 1000+ | |
| NC | 0b | 0b | 0d | 0e | 0d | 0c | ||
| SS | 0b | 0b | 0d | 0e | 0d | 0c | ||
| Blank | 0b | 0b | 0d | 0e | 0d | 0c | ||
| PC | 100a | 100a | 100a | 100a | 100a | 100a | ||
All ethanol extracts tested positive in the Dragendorff test, producing several orange spots at different FR values of 0.27, 0.39, 0.47, 0.54 and 0.59.
The ethanol extracts from the roots of A. gaumeri ex-situ cultivated showed two to four peaks each with percentages of at least 8% (Table 9). The plants from Kiuic and Tinum exhibited a similar chromatographic profile, displaying four peaks (Rt of 4.5, 4.84, 6.5, and 7.88 min) in female plants and three (Rt of 4.5, 4.84, and 8.49 min) in male plants. The population of Yaxcaba exhibited a different chromatographic profile: with two peaks (Rt of 4.5 and 8.49 min) in female plants and three peaks (Rt of 4.3, 4.5, and 7.6 min) in male plants. The most abundant peak corresponded to the least polar compound with a TR of 8.49 in KRM (21.93%) and YRF (18.40%) min. The field sample from Yaxcaba revealed three prominent peaks (Rt of 4.8, 5.91 and 6.75 min) with 11.09–15.60% of area.
Table 9: Percentage of component area of the components in the liquid chromatograms of the ethanol extracts obtained from the ex-situ cultivated roots of Acalypha gaumeri from the Kiuic, Tinum and Yaxcaba populations.
| Population | Sample | Retention Time (min) | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 4.30 | 4.50 | 4.84 | 5.81 | 6.50 | 6.72 | 7.60 | 7.88 | 8.49 | ||
| Kiuic | KRF | 14.00 | 16.81 | 16.49 | ||||||
| KRM | 7.12 | 7.76 | 21.93 | |||||||
| Tinum | TRF | 15.63 | 10.29 | 15.16 | ||||||
| TRM | 7.57 | 10.05 | 9.95 | |||||||
| Yaxcaba | YRF | 11.01 | 18.40 | |||||||
| YRM | 6.99 | 7.00 | 17.24 | |||||||
| Yaxcaba field | 11.09 | 15.60 | 12.92 | |||||||
This research contributes to the comprehensive knowledge of A. gaumeri. Therefore, the first part of the work focused on molecular identification and characterization of the populations from Kiuic, Tinum, and Yaxcaba, which were maintained in our nurseries.
In Mexico, the Acalypha genus includes 108 species, of which 61% are catalogued as endemic to the country. Among these, A. gaumeri is currently the only species endemic to the Yucatan Peninsula and is classified as “near threatened” [8,9]. This research employed a dual approach to identifying A. gaumeri, integrating morphological characteristics with molecular identification from genomic material. The specimens used in this study were collected and propagated in the Yucatan Peninsula. Previous deposits of A. gaumeri in GenBank include sequences obtained with ITS (OM140897), ndh-F (OM066409) and trnL-F (OM066565) primers, using herbarium specimens [16]. In our study, ITS and matK primers were used with fresh leaf material from A. gaumeri without success. Therefore, this is the first report of the A. gaumeri sequence deposited in GenBank using the rbcL primer. The rbcL gene has been the most thoroughly characterized and is considered to be one of the most reliable and useful for molecular identification studies [36]. The results of the phylogenetic analysis demonstrate the separation of A. gaumeri from the other sequences analyzed, with a high bootstrap support value (99%), indicating a genetic difference between the species. In this study, the use of the rbcL primer is demonstrated to be an effective method for molecular species identification. In this context, it has been confirmed that the rbcL primer is suitable for molecular identification of A. gaumeri, as other alternative markers have limitations in terms of amplification. Consequently, it is recommended that subsequent studies evaluate additional markers, such as trnL-F and ndh-F, to consolidate molecular identification. This finding is consistent with previous reports indicating that one of the main regions of the plant DNA barcode is rbcL. When used in conjunction with complementary regions, it facilitates accurate identification in taxonomic studies [37].
As part of this study, we present the first report on genetic diversity in the ex-situ cultivated Kiuic, Tinum, and Yaxcaba populations of A. gaumeri. This documentation used ISSR and DAMD markers and revealed a high degree of polymorphism. Previously, a high degree of polymorphism was reported in genetic diversity studies of 54 varieties of Ricinus communis (Euphorbiaceae), with an average of 74.59% using ISSR markers [38]. On the other hand, DAMD markers showed greater polymorphism than ISSR markers. These data are consistent with the report on Jatropha curcas, in which DAMD and ISSR markers displayed PIC values of 0.872 and 0.86, respectively [39]. Furthermore, this report indicates that markers 33.6 and HVR had the highest PIC values (0.914 and 0.898, respectively) [39], consistent with those obtained for A. gaumeri. The findings of this molecular marker study showed that individuals within populations possess a high degree of genetic diversity. This behaviour is generally associated with geographic distribution, reproductive mode, fecundity, and seed dispersal [39]. Another critical factor influencing genetic diversity within ex-situ cultivated populations is the genetic variation present in the parental populations [40]. However, this study did not directly compare populations with their parental populations. Given that the individuals analyzed in each population descended from wild parents, it can be postulated that the genetic variability present in situ was maintained ex situ. This hypothesis is supported by the dendrogram, which shows genetic differentiation between populations from different locations and within populations, as evidenced by differences between male and female individuals. These results on genetic variability indicate that, despite ex-situ propagation, these populations maintain their genetic differences. Growers should take this information when selecting individuals for reproduction to produce antifungal extracts.
Among the 36 ethanol and aqueous extracts, 22% showed antifungal activity against the tested plant pathogens. The active extracts included ethanol extracts from roots (KRF, KRM, TRF, TRM, YRM, YRF) against the three phytopathogens, which were inhibitory at 1000 μg/mL. The most active ethanol extracts, as indicated by the lowest MIC, were from A. gaumeri roots collected at Kiuic from female plants (KRF) against A. chrysanthemi, C. gloeosporioides and P. oxalicum at 250 µg/mL, 500 µg/mL and 500 µg/mL, respectively. This is the first study of the antifungal activity of A. gaumeri cultivated outside its natural environment, confirming that the roots of cultivated plants continue to inhibit the mycelial growth of A. chrystanthemi and C. gloeosporioides when evaluated using the microdilution bioassay. In addition, separate evaluations of male and female specimens are provided. The ethanol extracts were found to be more efficacious than those reported from field-collected plant extracts, with MIC values of 500 µg/mL and 1000 µg/mL against A. chrysanthemi CICY004 and C. gloeosporioides CG4, respectively [15].
According to previous reports, ethanol root extract of A. gaumeri at 1 mg/mL induced 68%–97% IGM on A. chrysanthemi (CICY004), Alternaria tagetica (ATTC 53771), Alternaria sp. (ITC02), Colletotrichum capsici (CC2), C. gloeosporioides (GC4), C. cassiicola (ITC03), Curvularia sp. (ITC10) and Helminthosporium sp. (ITC04) using the agar dilution assay [12,13]. Also tested on C. gloeosporioides (CICY002) and Fusarium oxysporum (CICY003) without activity [12]. The spectrum of antifungal activity of A. gaumeri was enriched by the effect of root ethanol extracts against P. oxalicum ITC25 isolated from habanero pepper (Capsicum chinenese).
The aqueous extracts from A. gaumeri did not inhibit A. chrysanthemi CICY004, C. gloeosporioides CICY002 and P. oxalicum ITC25 at 3% w/v. Other studies conducted with aqueous extracts from A. gaumeri roots showed no effect on the MGI against A. alternata ITC24, Corynespora cassiicola ITC23, Curvularia lunata ITC22, and F. equiseti ITC32. However, these completely inhibited the sporulation of C. lunata ITC22 and the conidial germination of both C. lunata ITC22 and F. equiseti ITC32. In addition, aqueous extracts from A. gaumeri roots exhibited 69% growth inhibition on A. chrysanthemi CICY004 using 3% w/v microdilution assay [13]. Plant material was collected in the field (Yaxcaba), and the agar dilution assay was used [41]. In addition, weekly application of aqueous extracts (3%) to chrysanthemum plants for three weeks under field conditions reduced the severity of leaf blight disease, as did the fungicide Captan® [13].
The chromatographic profile of ethanol extracts from A. gaumeri reveals differences among populations and genera, as well as between field-collected and ex-situ cultivated plants. Field and nursery extracts showed the presence of two major alkaloidal components, as determined by the Dragendorff test in thin-layer chromatography [15] and by HPLC at 280 nm. But the difference in their retention time is due to structural differences. Only one peak was found in all samples at Rt of 4.5 min (7–15.63%), while peaks at Rt of 5.91 min (15.60%) and 6.75 min (12.92%) were only detected in the field sample. Among the female populations of Kiuic and Tinum, KRF was the most active extract, showing higher peak areas at Rt of 6.5 (16.81%) and 7.88 (16.49%) min than other extracts. However, among the male populations, the most active was TRM exhibited higher peak abundances at Rt 4.5 min (7.57%) and 4.84 min (10.05%). This is consistent with the activity observed in the roots of A. gaumeri from the Tinum population, which were collected in the field and showed MIC values of 500 and 1000 μg/mL against A. chrysanthemi and C. gloeosporioides [15].
It was also found that female individuals from Kiuic, belonging to clade 2, as well as male and female individuals from Tinum, were more active and had the lowest MIC. According to Wang et al. [42], these results may correlate with antifungal activity against the tested phytopathogenic fungi. Genetic variability within a species results in differences in the kinds and amounts of secondary metabolites, as evidenced by chromatographic profile and biological effects.
The demonstrated antifungal activity of A. gaumeri ex-situ propagated, and its complementary molecular identification are promising biorational product options. This finding highlights the need for ongoing evaluations against other phytopathogens. The chemical identification of alkaloid compounds in field-collected plants is being carried out to verify their activity and toxicity. Additionally, the purification and identification of the compounds detected in ex-situ cultivated plants must be carried out. Seasonal monitoring of activity and chemical profile is also required.
The molecular identification of A. gaumeri from Yucatan using the rbcL primer complements its taxonomy. The A. gaumeri plants sampled from the Kiuic (Oxkutzcab), Tinum, and Yaxcaba, populations in Yucatan showed a higher genetic diversity than expected. The genetic fingerprints obtained for female individuals in the Kiuic population showed slight differences compared with those from the Tinum and Yaxcaba populations. The molecular markers ISSR and DAMD were useful for detecting polymorphisms within and between individuals and populations. Antifungal activity is maintained in specimens propagated and cultured outside their environment. The activity of A. gaumeri varies depending on the sex, organ, and population. The root is the main plant tissue of A. gaumeri that inhibits the growth of phytopathogenic fungi. The spectrum of antifungal activity of A. gaumeri is broadened by the sensitivity of P. oxalicum. The differences in the genetic fingerprint of the female population in Kiuic could be related to its significant biological activity and chemical profile. This information will enable important decisions on the selection of materials for further study, ensuring the sustainability of the raw materials used to produce botanical extracts.
Acknowledgement:
Funding Statement: This research was supported by the SECIHTI project PDCPN-2015-266, México.
Author Contributions: The authors confirm contribution to the paper as follows: Performed the experiments, collected, analyzed the data, and wrote the original draft, Christian Pérez-Chablé; performed antifungal experiment, Irma L. Medina-Baizabal; supervision and curation of molecular data, Anuar Magaña-Alvarez; provided fungal phytopathogens, Jairo Cristóbal-Alejo; conceived the project, resources, funding acquisition, project administration, supervised and writing—reviewed and edited, Marcela Gamboa-Angulo and Daisy Pérez-Brito. All authors participated in interpreting the data. All authors reviewed the results and approved the final version of the manuscript.
Availability of Data and Materials: The data that support the findings of this study are available from the Corresponding Author, Marcela Gamboa-Angulo, upon reasonable request.
Ethics Approval: Not applicable.
Conflicts of Interest: The authors declare no conflicts of interest to report regarding the present study.
References
1. El-Baky NA , Amara AA . Recent approaches towards control of fungal diseases in plants: An updated review. J Fungi. 2021; 7: 900– 16. doi:10.3390/jof7110900. [Google Scholar] [CrossRef]
2. Dean R , Van-Kan JA , Pretorius ZA , Hammond-Kosack KE , Di Pietro A , Spanu PD , et al. The Top 10 fungal pathogens in molecular plant pathology. Mol Plant Pathol. 2012; 13( 4): 414– 30. doi:10.1111/j.1364-3703.2011.00783.x. [Google Scholar] [CrossRef]
3. Olejnik A , Parkitna K , Kozak B , Florczak S , Matkowski J , Nowosad K . Assessment of the genetic diversity of chrysanthemum cultivars using SSR markers. Agron. 2021; 11: 2318. doi:10.3390/agronomy11112318. [Google Scholar] [CrossRef]
4. Secretaría de Agricultura y Desarrollo Rural. [cited 2025 Mar 5]. Available from: https://www.gob.mx/agricultura/articulos/las-flores-estan-en-el-campo-en-las-miradas-en-las-palabras. [Google Scholar]
5. Allende-Molar R , Picos-Muñoz PA , Márquez-Zequera I , Carrillo-Fasio JA , García-Estrada RS , León-Félix J . Identificación morfológica y molecular de Penicillium oxalicum causante de pudrición de tallos y frutos de tomate. Rev Mex Fitopatol. 2013; 31( 1): 13– 9. (In Spanish). [Google Scholar]
6. Park J , Kim S , Jo M , An S , Kim Y , Yoon J , et al. Isolation and identification of Alternaria alternata from potato plants affected by leaf spot disease in Korea: selection of effective fungicides. J Fungi. 2024; 10: 53. doi:10.3390/jof10010053. [Google Scholar] [CrossRef]
7. Romanazzi G , Sanzani SM , Bi Y , Tian S , Gutiérrez-Martínez P , Alkan N . Induced resistance to control postharvest decay of fruit and vegetables. Postharvest Biol Technol. 2016; 122: 82– 94. doi:10.1016/j.postharvbio.2016.08.003. [Google Scholar] [CrossRef]
8. Villaseñor JL . Checklist of the native vascular plants of Mexico. Rev Mex Biodivers. 2016; 87: 559– 902. doi:10.1016/j.rmb.2016.06.017. [Google Scholar] [CrossRef]
9. Fernández-Concha GC , Ramírez-Morillo I , Pérez-Sarabia JE , Tapia-Muñoz JL , Medina HE , Cetzal-Ix W , et al. Assessing the risk of extinction of vascular plants endemic to the Yucatán Peninsula Biotic Province by means of distributional data. Ann Missouri Bot Gard. 2021; 106( 1): 424– 57. doi:10.3417/2021661. [Google Scholar] [CrossRef]
10. Cruz-Estrada A , Gamboa-Angulo M , Borges-Argáez R , Ruiz-Sánchez E . Insecticidal effects of plant extracts on immature whitefly Bemisia tabaci Genn. (Hemiptera: Aleyroideae). Electron J Biotechnol. 2013; 16( 1): 6. http://dx.doi.org/10.2225/vol16-issue1-fulltext-6. doi:10.2225/vol16-issue1-fulltext-6. [Google Scholar] [CrossRef]
11. Cristóbal-Alejo J , Tun-Suárez JM , Moguel-Catzin S , Marbán-Mendoza N , Medina-Baizabal L , Simá-Polanco P , et al. In vitro sensitivity of Meloidogyne incognita to extracts from native Yucatecan plants. Nematropica. 2006; 36: 89– 98. https://journals.flvc.org/nematropica/article/view/69732. [Google Scholar]
12. Gamboa-Angulo M , Cristóbal-Alejo J , Medina-Baizabal IL , Chi-Romero F , Mendéz-González R , Sima-Polanco P , et al. Antifungal properties of selected plants from the Yucatán peninsula, Mexico. World J Microbiol Biotechnol. 2008; 24: 1955– 9. doi:10.1007/s11274-008-9658-x. [Google Scholar] [CrossRef]
13. Vargas A , Gamboa M , Medina I , Pérez D , Alejo JC , Ruiz E . Evaluación de extractos de plantas nativas Yucatecas contra Alternaria chrysanthemi y espectro de actividad antifúngica de Acalypha gaumeri. Rev Mex Fitopatol. 2014; 32: 1– 11. (In Spanish). [Google Scholar]
14. Vargas-Díaz AA , Cristóbal-Alejo J , Canto-Canché B , Gamboa-Angulo MM . Aqueous extracts from Acalypha gaumeri and Bonellia flammea for leaf blight control in chrysanthemum (Chrysanthemum morifolium). Rev Mex Fitopatol. 2022; 40( 1): 40– 58. doi:10.18781/r.mex.fit.2109-1. [Google Scholar] [CrossRef]
15. Vargas-Diaz AA , Pérez-Brito D , Hernández-Carlos B , Cristóbal-Alejo J , Andrade-Canto S , Gamboa-Angulo M . Acalypha gaumeri: antifungal activity of three populations under edaphic and seasonal variations and ex-situ propagation. Phyton-Int J Exp Bot. 2025; 94( 9): 2839– 53. doi:10.32604/phyton.2025.066682. [Google Scholar] [CrossRef]
16. Levin GA , Cardinal-McTeague WM , Steinmann VW , Sagun VG . Phylogeny, classification, and character evolution of Acalypha (Euphorbiaceae: Acalyphoideae). Syst Bot. 2022; 47( 2): 477– 97. doi:10.1600/036364422X16512572275034. [Google Scholar] [CrossRef]
17. Acharya R , Roy S , Harisha CR , Hegde S , Ranade A . Anatomical and genetic identification of two Euphorbia species. J Indian Syst Med. 2021; 9: 252– 8. doi:10.4103/jism.jism_1_21. [Google Scholar] [CrossRef]
18. Agarwal M , Shrivastava N , Padh H . Advances in molecular marker techniques and their applications in plant sciences. Plant Cell Rep. 2008; 27: 617– 31. doi:10.1007/s00299-008-0507-z. [Google Scholar] [CrossRef]
19. Kumar H , Priya P , Singh N , Kumar M , Choudhary BK , Kumar L , et al. RAPD and ISSR marker-based comparative evaluation of genetic diversity among Indian germplasms of Euryale ferox: an aquatic food plant. Appl Biochem Biotechnol. 2016; 180: 1345– 60. doi:10.1007/s12010-016-2171-z. [Google Scholar] [CrossRef]
20. Rabeh M , El-Hawary S , El-Tantawy M , Abdelmohsen U , Badr W . Survey the genetic diversity of eight Opuntia species from Egypt using random amplified polymorphic DNA (RAPD) technique. J Adv Biomed Pharm Sci. 2020; 3: 40– 4. doi:10.21608/jabps.2019.20728.1062. [Google Scholar] [CrossRef]
21. Adhikari S , Biswas A , Saha S , Biswas A , Ghosh P . SPAR methods reveal high genetic diversity within populations and moderate gene flow of pointed gourd (Trichosanthes dioica Roxb.) germplasm. Biocatal Agric Biotechnol. 2020; 29: 101760. doi:10.1016/j.bcab.2020.101760. [Google Scholar] [CrossRef]
22. Galindo-García DV , Alia-Tejacal I , Núñez-Colin CA , Andrade-Rodríguez M , Canul-Ku J , Colinas-León MT , et al. Diversidad genética de nochebuena de sol (Euphorbia spp.) en Morelos, México, con marcadores moleculares RAPD. Rev Chapingo Ser Hortic. 2019; 25( 2): 113– 27. (In Spanish). doi:10.5154/r.rchsh.2018.06.012. [Google Scholar] [CrossRef]
23. Ruiz-Montoya LR , López-López MZ , Lorenzo C , García-Bautista M , Ramírez-Marcial N . Variación genética de cuatro especies de árboles tropicales de la Reserva de la Biosfera Selva El Ocote, Chiapas, México. Acta Bot Mex. 2021; 128: 8. (In Spanish). doi:10.21829/abm128.2021.1847. [Google Scholar] [CrossRef]
24. Tapia-Tussell R , Quijano-Ramayo A , Rojas-Herrera R , Larque-Saavedra A , Perez-Brito D . A fast, simple, and reliable high-yielding method for DNA extraction from different plant species. Mol Biotechnol. 2005; 31: 137– 9. doi:10.1385/MB:31:2:137. [Google Scholar] [CrossRef]
25. Zechman FW . Phylogeny of the Dasycladales (Chlorophyta, Ulvophyceae) based on analyses of rubisco large subunit (rbcL) gene sequences. J Phycol. 2003; 39: 819– 27. doi:10.1046/j.1529-8817.2003.02183.x. [Google Scholar] [CrossRef]
26. Hall TA . BioEdit: a user-friendly biological sequence alignment editor and analysis program for windows 95/98/NT. Nucleic Acids Symp Ser. 1999; 41: 95– 8. [Google Scholar]
27. Weiguo Z , Zhihua Z , Xuexia M , Sibao W , Lin Z , Yile P , et al. Genetic related among cultivated and wild mulberry (Moraceae: Morus) as revealed by inter-simple sequence repeat analysis in China. Can J Plant Sci. 2005; 86: 251– 7. doi:10.4141/P04-110. [Google Scholar] [CrossRef]
28. AcaClone . pDRAW32, edition 1.1.147. Software. 2021 [cited 2025 Mar 5]. Available from: https://www.acaclone.com. [Google Scholar]
29. Verma S , Rana TS . Genetic relationship among wild and cultivated accesions of curry leaf plant (Muraya koenigii (L.) Spreng.), as revealed by DNA Fingerprinting methods. Mol Biotechnol. 2013; 53: 139– 49. doi:10.1007/s12033-012-9500-4. [Google Scholar] [CrossRef]
30. Hoisington D , Khairallah M , Gonzalez-de-Leon D . Laboratory protocols: CIMMYT applied molecular genetics laboratory. 2nd ed. Texcoco, Mexico: CIMMYT; 1994. p. 51– 2. [Google Scholar]
31. Clinical and Laboratory Standards Institute . Reference method for broth dilution antifungal susceptibility testing of filamentous fungi. Approved Standard M38-A; Wayne, PA, USA: National Committee for Clinical Laboratory Standards. 2002 [cited 2025 Mar 5]. Available from: https://cdn.bfldr.com/YLD4EVFU/at/m59q93gqrnpqvqc7735z6j/m38ed3_sample.pdf. [Google Scholar]
32. Abbott WS . A method of computing the effectiveness of an insecticide. J Econ Entomol. 1925; 18: 256– 67. doi:10.1093/jee/18.2.265a. [Google Scholar] [CrossRef]
33. Jaccard P . Nouvelles recherché sur la distribution florale. Bull Soc Vaudoise Sci Nat. 1908; 44: 223– 70. (In Spanish). [Google Scholar]
34. Rohlf FJ . NTSYS-pc: numerical taxonomy and multivariate analysis system. Setauket, NY, USA: Exeter Software; 2000. [Google Scholar]
35. Anderson JA , Churcill GA , Autrique JE , Tanksley SD , Sorrels ME . Optimizing parental selection for genetic linkage maps. Genome. 1992; 36: 181– 6. doi:10.1139/g93-024. [Google Scholar] [CrossRef]
36. Fazekas AJ , Burgess KS , Kesanakurti PR , Graham SW , Newmaster SG , Husband BC , et al. Multiple multilocus DNA barcodes from the plastid genome discriminate plant species equally well. PLoS One. 2008; 3( 7): e2802. doi:10.1371/journal.pone.0002802. [Google Scholar] [CrossRef]
37. Fazekas AJ , Kuzmina ML , Newmaster SG , Hollingsworth PM . DNA barcoding methods for land plants. In: Kress W , Erickson D , editors. DNA barcodes. Methods in molecular biology. Totowa, NJ, USA: Humana Press; 2012. Volume 858. doi:10.1007/978-1-61779-591-6_11. [Google Scholar] [CrossRef]
38. Kim H , Lei P , Wang A , Liu S , Zhao Y , Huang F , et al. Genetic diversity of Castor bean (Ricinus communis L.) revealed by ISSR and RAPD markers. Agronomy. 2021; 11: 457. doi:10.3390/agronomy11030457. [Google Scholar] [CrossRef]
39. Gautam Murty S , Patel F , Punwar BS , Patel M , Singh AS , Fougat RS . Comparison of RAPD, ISSR, and DAMD markers for genetic diversity assessment between accessions of Jatropha curcas L. and its related species. J Agric Sci Technol. 2013; 15( 5): 1007– 22. [Google Scholar]
40. Chen C , Chang H , Pang X , Liu Q , Xue L , Yin C . Genetic diversity analysis and conservation strategy recommendations for ex situ conservation of Cupressus chengiana. BMC Plant Biol. 2025; 25: 552. doi:10.1186/s12870-025-06581-z. [Google Scholar] [CrossRef]
41. Moo-Koh FA , Cristóbal-Alejo J , Tun-Suárez JM , Medina-Baizabal IL , Arjona-Cruz AA , Gamboa-Angulo M . Activity of aqueous extracts from native plants of the Yucatan Peninsula against fungal pathogens of tomato in vitro and from Croton chichenensis against Corynespora cassiicola on tomato. Plants. 2022; 11( 21): 2821. doi:10.3390/plants11212821. [Google Scholar] [CrossRef]
42. Wang H , Ma C , Ma L , Du Z , Wang H , Ye H , et al. Secondary metabolic profiling and artemisinin biosynthesis of two genotypes of Artemisia annua. Planta Med. 2009; 75( 15): 1625– 33. doi:10.1055/s-0029-1185814. [Google Scholar] [CrossRef]
Cite This Article
 Copyright © 2026 The Author(s). Published by Tech Science Press.
Copyright © 2026 The Author(s). Published by Tech Science Press.This work is licensed under a Creative Commons Attribution 4.0 International License , which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.


 Submit a Paper
Submit a Paper Propose a Special lssue
Propose a Special lssue View Full Text
View Full Text Download PDF
Download PDF Downloads
Downloads
 Citation Tools
Citation Tools