 Open Access
Open Access
ARTICLE
A Transcriptional Suppressor NTL8 Interacts with CONSTANS to Control Photoperiod-Mediated Flowering Time in Arabidopsis thaliana
1 Shanghai Key Laboratory of Bio-Energy Crops, School of Life Sciences, Shanghai University, Shanghai, China
2 Department of Agronomy, University of Almeria, Almeria, Spain
* Corresponding Authors: García-Caparrós Pedro. Email: ; Xiangyang Hu. Email:
(This article belongs to the Special Issue: Plant Secondary Metabolism and Functional Biology Volume II)
Phyton-International Journal of Experimental Botany 2026, 95(3), 21 https://doi.org/10.32604/phyton.2026.067796
Received 13 May 2025; Accepted 28 January 2026; Issue published 31 March 2026
Abstract
Plants perceive rhythmic photoperiodic signals to modulate flowering time. In Arabidopsis thaliana, long-day light conditions accelerate flowering through CONSTANS (CO)-activated FLOWERING LOCUS T (FT) signal pathway. The CO protein abundance presents circadian oscillation, enabling precise regulation of FT transcription. NTL8 belongs to the NAC transcription factor family and is reported to control leaf trichome development. Here, we reported that NTL8 regulated flowering time in Arabidopsis, because overexpressing NTL8 significantly delayed flowering time, whereas loss-of-function mutant of NTL8 accelerated flowering time. NTL8 also presented circadian expression, maintaining elevated transcript levels during the daytime. Biochemical and genetic analyses revealed that NTL8 physically interacted with CO in planta, thereby antagonizing CO activity and repressing FT expression to delay the flowering time. Furthermore, overexpressing NTL8 reduced the protein stability of CO, particularly by attenuating CO accumulation in the morning time and promoting CO degradation during the night. Collectively, our findings indicated that NTL8 delayed photoperiod-dependent flowering time by suppressing CO-mediated activation of FT expression and destabilizing CO protein. This study uncovers a previously unrecognized role for NTL8 in coordinating circadian and photoperiodic signals to fine-tune flowering time in Arabidopsis thaliana.Keywords
Supplementary Material
Supplementary Material FileFlowering at the appropriate time leads to successful fertilization and seed production, thereby ensuring optimal crop yield [1,2,3,4,5]. Plant flowering time is accurately controlled by a complicated etwork of endogenous physiological signals as well as external environmental cues [4,6]. Among these, plants sense the day length via the photoperiodic pathway, also called photoperiod-mediated flowering time, which is controlled by the activity of the B-box protein CONSTANS (CO) in leaf phloem companion cells [6,7]. Transcription of CO is governed by the circadian clock, resulting in a rhythmic oscillation of its transcript level [4,8]. The proper functioning of CO also relies on its degradation through the ubiquitin-proteasome system at nighttime and dawn, allowing maximal CO levels to be reached near sunset in long photoperiods (16-h illumination/8-h darkness) [8,9,10,11]. The accumulated CO subsequently triggers the transcription of the flowering hormone gene FLOWERING LOCUS T (FT) within the vascular tissues of leaves [12,13,14]. The florigenic FT peptide subsequently functions as a mobile developmental cue, moving to the shoot apex where it induces downstream flowering genes including APETALA1 (AP1) and LEAFY (LFY), ultimately triggering floral bud development in the reproductive apex [5,14,15]. Multiple transcriptional regulators influence CO abundance and activity. FLOWERING bHLH (FBH) and TEOSINTE BRANCHED 1/CYCLOIDEA/PROLIFERATING CELL NUCLEAR ANTIGEN FACTOR (TCP) TCP4 cooperate with GIGANTEA (GI) to promote CO gene activation [16,17], while the E3 ligases CONSTITUTIVE PHOTOMORPHOGENIC 1 (COP1) and HIGH EXPRESSION OF OSMOTICALLY RESPONSIVE GENE1 (HOS1) specifically target CO for breakdown during nighttime and dawn phases, respectively [8,9]. Additional factors like FLAVIN-BINDING, KELCH REPEAT, and F BOX 1 (FKF1) facilitate the removal of CYCLING DOF FACTOR 1 (CDF1) to maintain rhythmic CO production [18,19,20]. The regulation of CO protein stability and transcriptional activity involves additional components including PHYTOCHROME-ASSOCIATED LATE-FLOWERING (PHL) and EARLY ACTIVATION TARGET 1 (TOE1), which collectively fine-tune floral transition timing [18,21,22]. Earlier work revealed that ABI5-INTERACTING PROTEIN 2 (AFP2) promotes CO turnover through direct interaction, resulting in delayed flowering [23]. These observations establish that CO abundance and activity integrate diverse environmental signals at both transcriptional and post-translational levels. The NAC (NAM/ATAF/CUC) family represents one of the most extensive groups of transcriptional regulators in plants [24,25], with members governing fundamental developmental processes ranging from flower morphogenesis and cell division to cuticle formation and hair cell differentiation [25]. Emerging research underscores their pivotal roles in abiotic stress adaptation. Arabidopsis contains 117 NAC genes, including 14 membrane-associated members designated as NTL proteins [26,27], which exhibit specialized functions. NAC TRANSMEMBRANE MOTIF1 (NTM1) enhances pathogen defense [28], whereas NTL4 mediates drought-associated leaf aging [29]. ANAC089/NTL14 modulates ER stress-related cell death and flowering time via FLC regulation [30,31], while ANAC013/NTL1 participates in oxidative stress responses [32]. Particularly, NTL8 influences flowering through BFT interaction and regulates salt-stress germination [33]. Subsequent findings demonstrated its role in epidermal patterning through TRY and TCL1 targeting [34], along with contributions to cold adaptation and salinity tolerance during germination [35]. However, the detailed pathways through which NTL8 modulates floral induction require comprehensive exploration.
In this study, we examined how NTL8 influences floral timing, discovering its mRNA abundance follows daily rhythmic changes with maximal accumulation preceding sunset and gradual reduction during nocturnal and early daylight periods, closely resembling CO’s transcriptional rhythm. Experimental data demonstrated NTL8 delays floral transition in extended photoperiods through FT repression, while protein interaction studies established its physical association with CO, indicating their cooperative regulation of FT levels to adjust flowering to day length. These results uncover a previously unknown role for NTL8 in floral timing control and provide mechanistic insights into how the NTL8-CO partnership operates within the light cycle-dependent flowering network.
The plant materials employed in this research, comprising the Arabidopsis thaliana T-DNA insertion lines ntl8-1 (WiscDsloxHs159_07E) and ntl8-2 (SM_3_16309), were generously supplied by Prof. Shucai Wang from Linyi University [34], while the ft-10 line was obtained from Dr. Leiyin Zheng at the Chinese Academy of Sciences. Hybrid plants containing multiple mutations or transgenes were created through genetic crosses followed by isolation of pure-breeding offspring. For cultivation, sterilized seeds were placed on 0.5× MS plates containing 8 g/L agar and 10 g/L sucrose (adjusted to pH 5.7), with young plants maintained at 22°C under either extended (16-h illumination at 100 μmol·m−2·s−1/8-h dark) or reduced (8-h light/16-h dark) daily light periods. Mature specimens were transplanted into a 3:1 soil-vermiculite mixture under controlled temperature conditions. The timing of floral transition was assessed by enumerating basal leaves when the primary stem elongated to 1–3 cm, with phenotypic analysis performed on 20–30 individuals per genetic variant.
2.2 Plasmid Construction and Transgenic Plants
The 35S::CO-Flag fusion was created by PCR amplification of the complete CO coding region with PrimeSTAR DNA polymerase (Takara, Japan), followed by ligation to a Flag tag sequence. This amplicon was subsequently inserted into the pRI101-AN plasmid through In-Fusion HD cloning (Clontech, USA). The engineered vector was transferred into Arabidopsis Col-0 ecotype via Agrobacterium transformation, with transformants identified by growth on 0.5× MS plates supplemented with 50 μg/mL kanamycin [36]. Oligonucleotide sequences employed for molecular cloning appear in Supplementary Table S1.
2.3 Protein Immunoblots Analysis
Plant tissue proteins were isolated from young plants according to established protocols [37,38]. Cellular proteins were obtained with a lysis solution comprising 50 mM Tris-HCl (pH 7.5), 150 mM sodium chloride, 1 mM EDTA (pH 8.0), 0.1% Triton X-100, 10 mM sodium fluoride, and 5% glycerol, enhanced with phosphatase inhibitors (Roche) and 1 mM PMSF (Sigma). After centrifugation at 15,000× g for 10 min at 4°C, supernatants were collected for quantification with Bradford assay (Invitrogen). Protein samples (15 μg per lane) were resolved on 12% SDS-PAGE gels and electroblotted onto PVDF membranes, which were subsequently incubated with specific primary antibodies, including CO antibody (1:1000, Cat. 220746, Orizymes) generated against CO-His fusion protein, HA antibody (1:3000 dilution, Roche), FLAG antibody (1:3000, Sigma), GFP antibody (1:3000, Clontech), or actin antibody (1:1000, Sigma), followed by HRP-linked anti-mouse or anti-rat IgG (1:3000, Promega). Immunoreactive bands were visualized using an enhanced chemiluminescence detection system (Genescript).
To conduct yeast two-hybrid analysis, the gene fragment containing NTL8 and CO were PCR-amplified and subsequently inserted into pGBKT7 (DNA-binding domain vector) and pGADT7 (activation domain vector) respectively through In-Fusion cloning technology (Clontech) [23,38]. Y187 and AH109 yeast cells were genetically modified with the activation domain and DNA-binding domain constructs correspondingly via polyethylene glycol transformation following standard procedures. Following selection on tryptophan-deficient (630413, Clontech) or leucine-deficient (630414, Clontech) agar plates, three separate colonies per plasmid combination were crossed and cultured on double dropout medium (630417, Clontech) for 72 h to verify mating efficiency. The resulting diploid strains were then plated on quadruple dropout selection plates (630428, Clontech) to assess potential molecular interactions through colony formation.
To examine protein interactions in vitro, complete or partial coding sequences of NTL8 and CO were PCR-amplified and subsequently inserted into pET28a (Merck) and pGEX-4T-1 (Pharmacia) expression vectors, creating pET28a-CO and pGST-NTL8 recombinant plasmids [23]. Oligonucleotides employed for amplification are provided in Supplementary Table S1. For bacterial protein production, these plasmids were introduced into E. coli Rosetta cells, with recombinant protein synthesis initiated by IPTG treatment. The expressed GST-fused NTL8 was purified using glutathione-coupled resin (GE Healthcare), while GST protein served as negative control. In binding experiments, 2 μg of His-tagged CO was mixed with either GST or GST-NTL8 bound to beads in interaction solution (50 mM Tris-HCl, pH 8.0, 100 mM NaCl, 1 mM EDTA) at 4°C for 12 h. Following incubation, captured complexes were rigorously rinsed with wash buffer (50 mM Tris-HCl, pH 7.4, 100 mM NaCl, 0.6% Triton X-100). Resolved proteins from 8% SDS-PAGE were transferred to membranes and analyzed by western blotting with His-specific (Abmart) or GST-specific (Abmart) antibodies, then visualized using HRP-conjugated anti-mouse IgG (1:5000; Promega).2.6 Co-immunoprecipitation.
To investigate protein interactions in planta, NTL8-HA and CO-Flag lines were crossed to create dual-expressing transgenic plants. For co-immunoprecipitation experiments with CO-FLAG as the target, 14-day-old NTL8-HA/CO-Flag seedlings were cryogenically pulverized [23]. Cellular proteins were isolated with extraction buffer containing 100 mM MOPS (pH 7.6), 150 mM sodium chloride, 0.1% NP-40, 1% Triton X-100, 20 mM iodoacetamide, 1 mM PMSF, protease inhibitors (2 μg/mL aprotinin, 5 μg/mL leupeptin, 1 μg/mL pepstatin) and phosphatase inhibitors (Roche). After centrifugation at 22,000× g (4°C, 10 min) and Miracloth filtration, 1 mL cleared lysate was mixed with HA-conjugated magnetic beads (Sigma) for 2 h at 4°C with rotation. Following four washes with 50 mM Tris (pH 8.0), 150 mM NaCl, 0.1% Triton X-100, captured complexes were heat-eluted (95°C, 10 min) in SDS sample buffer (100 mM Tris-HCl pH 6.8, 200 mM DTT, 2% SDS, 20% glycerol, 0.2% bromophenol blue). Immunodetection was performed using FLAG (1:3000, Sigma) or HA (1:3000, Roche) antibodies, with NTL8-HA/GFP-Flag plants serving as negative controls.
Total RNA was isolated from two-week-old seedlings with TRIzol (Invitrogen), followed by cDNA synthesis from 1.5 μg DNase-digested RNA in 20 μL reactions employing M-MuLV reverse transcriptase (Fermentas) and oligo(dT)18 primers [23,38]. Quantitative PCR analysis was conducted using diluted cDNA (2–10 ng/μL) with SYBR Green Master Mix on a LightCycler 480 system (Roche) according to standard protocols, with PP2A serving as the reference gene. Three technical replicates were performed for each assay, with primer sequences provided in Supplementary Table S1.
2.8 Protoplast Transient Expression Assay
To analyze transient gene expression in Arabidopsis protoplasts, a 2.7-kb FT promoter fragment was ligated into pGreenII 0800-LUC to create FTpro::LUC reporter plasmids, while NTL8 and CO coding sequences were subcloned into pGreenII 62-SK downstream of the 35S promoter (primer details in Supplementary Table S1). Leaf mesophyll protoplast isolation and transformation followed established protocols with slight adjustments. Luminescence measurements employed the dual-luciferase system (Promega), where firefly LUC activity was normalized to 35S-driven Renilla luciferase (REN) from the same vector, with LUC/REN ratios determining relative expression levels.
2.9 Luciferase Complementation Imaging (LCI) Assay
To examine protein-protein interactions through luciferase complementation imaging, NTL8 and CO coding sequences were inserted into pCAMBIA1300-nLUC and pCAMBIA1300-cLUC plasmids respectively, then introduced into Agrobacterium GV3101. Bacterial suspensions carrying these constructs were pressure-infiltrated into Nicotiana benthamiana foliage, with luminescence signals captured 48 h post-infiltration using a Tanon 5200 cooled-CCD imaging platform (Shanghai), following established methodology [39].
2.10 Chromatin Immunoprecipitation Following by qPCR Analysis (CHIP-qPCR)
To analyze histone modifications, 14-day-old LD-grown seedlings were collected at ZT16 (light-off transition) and immediately vacuum-infiltrated for 15 min with fixation solution containing 10 mM Tris-HCl (pH 8.0), 0.4 M sucrose, 1 mM EDTA, 1 mM PMSF, 0.25% Triton X-100, and 1% formaldehyde. The reaction was stopped with 0.125 M glycine under vacuum for 10 min, followed by three washes with chilled water. Frozen tissues were pulverized in liquid nitrogen, and chromatin was isolated, sonicated to 300–500 bp fragments, then incubated with Protein A/G magnetic beads coupled with anti-acetyl-H3 antibody (Abcam #ab47915) recognizing multiple lysine residues (K9/14/18/23/27). After 5-h elution at 65°C, immunoprecipitated DNA was purified (NEB kit) and quantified by qPCR (LightCycler 480, SYBR Green), with enrichment values calculated by normalizing against input DNA and wild-type controls (primers in Supplementary Table S1).
3.1 Circadian Expression of NTL8
To investigate the expression pattern of NTL8 during both daytime and nighttime, two-week-old seedlings were subjected to long-day (LD) photoperiodic conditions (16-h light, 8-h darkness), and the transcriptional level of NTL8 was measured at 4-h intervals. As shown in Fig. 1A, the expression of NTL8 showed rhythmic oscillations, with a gradual increase during the daytime, reaching a peak at ZT16, coinciding with dusk, and then decreasing during the nighttime. Similarly, when seedlings were grown under short-day (SD) photoperiodic conditions (8-h light, 16-h darkness), rhythmic expression of NTL8 was still observed, with a peak at ZT8, followed by sustained low expression level during the nighttime. Notably, the maximum transcriptional level of NTL8 under SD conditions was lower than that observed under LD conditions during the daytime. These data suggest the circadian oscillation of NTL8, and indicate that light conditions induced the expression of NTL8, while darkness suppressed the expression. To further test this hypothesis, we shifted seedlings from constant dark (cD) conditions to light conditions (L), resulting in the induced expression of NTL8 (Fig. 1B). Conversely, when shifting the seedlings from constant light conditions (cL) to dark conditions (D), NTL8 expression was downregulated (Fig. 1B). These data confirm that the expression of NTL8 is light-inducible and presents a circadian rhythm.
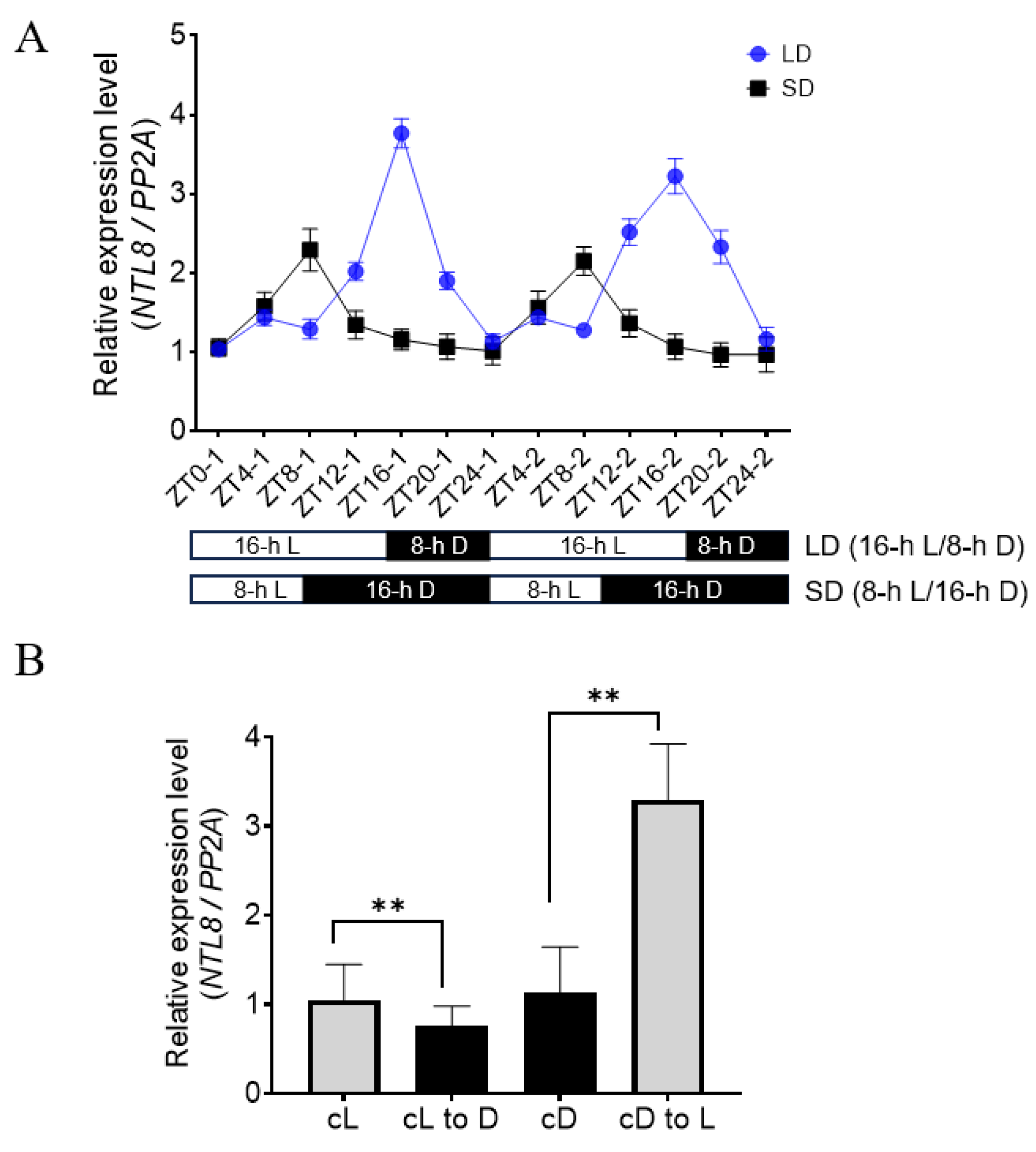
Figure 1: The circadian expression of NTL8 by RT-qPCR analysis. (A) Two-week-old wild-type Col seedlings grown under long-day (LD) or short-day (SD) conditions were sampled for RT-qPCR analysis. PP2A is used as an internal control (n = 3, mean ± SD). (B) Light-responsive NTL8 expression was examined in Col seedlings subjected to four light treatments: continuous light (cL, 24 h), continuous darkness (cD, 24 h), light-to-dark transition (12 h darkness after cL), and dark-to-light transition (12 h light after cD), followed by RT-qPCR analysis (n = 3, mean ± SD, **p < 0.01 by Student’s t-test).
3.2 NTL8 Regulates Flowering Time
Most circadian-regulated genes are known to influence flowering time [1,6,7,12]. To investigate the potential role of NTL8 in flowering time regulation, we obtained two T-DNA insertion lines, in which T-DNA insertions disrupted the second and third exons, respectively, thereby impairing its functional transcript (Supplementary Fig. S1A,B). These two mutant lines were named ntl8-1 and ntl8-2, respectively. In parallel, we obtained several independent transgenic lines overexpressing an NTL8-HA fusion gene under the control of constitutive 35S promoter (35S: NTL8-HA, abbreviated as NTL8-HA, Supplementary Fig. S1C). Under long day (LD) conditions (16-h light/8-h dark), the flowering time of ntl8 mutant lines was slightly earlier compared to wild-type Col line, while the flowering time of NTL8-HA was significantly later than that in the wild-type Col line (Fig. 2A). In contrast, no significant differences in flowering time were observed among Col, ntl8 mutants, and NTL8-HA transgenic lines under short day (SD) conditions (8-h light/16-h dark, Fig. 2A). These findings suggest that NTL8 delays the flowering time via the photoperiodic pathway.
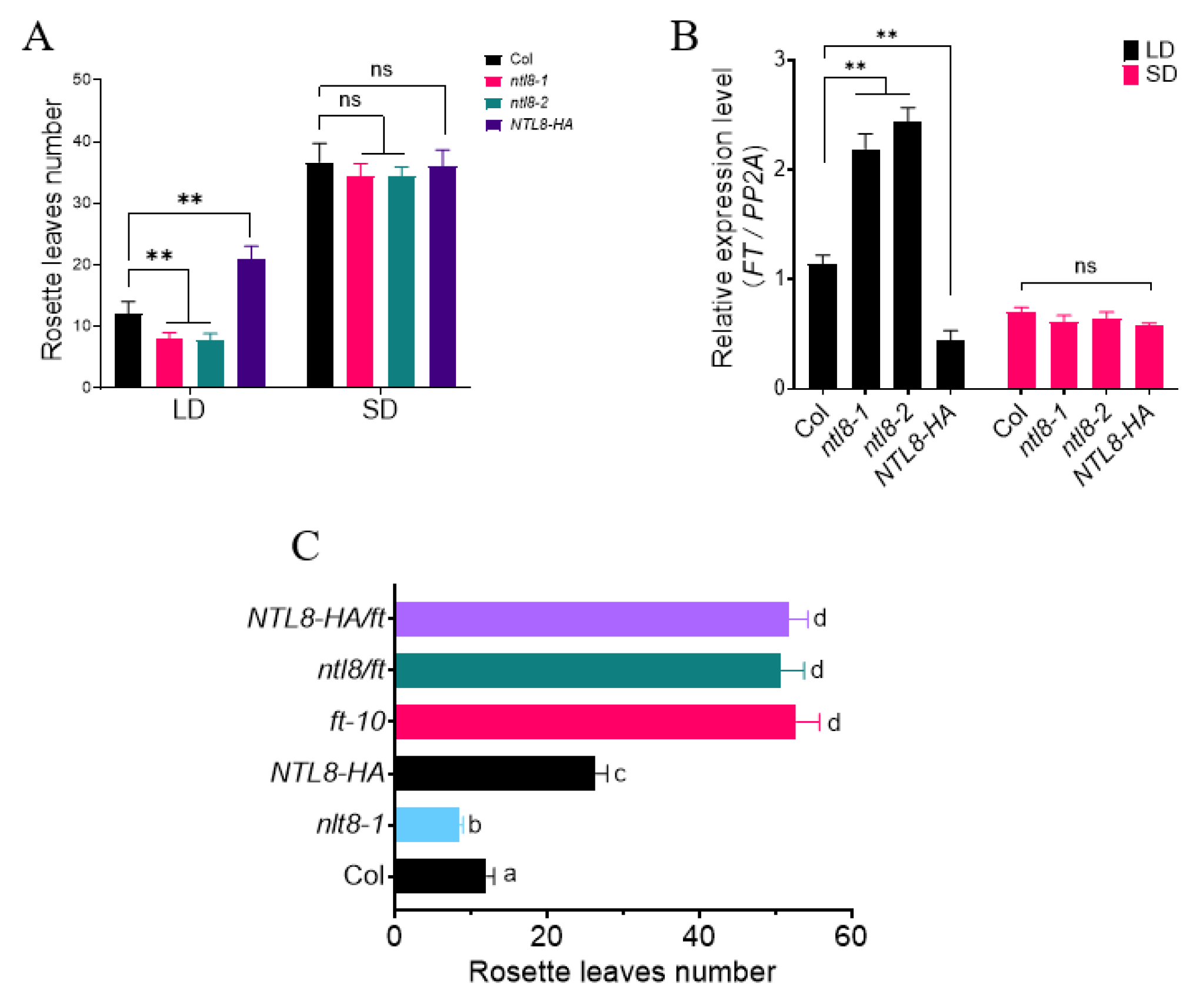
Figure 2: NTL8 controls the flowering time through FT signal pathway. (A) Flowering time variation in Col, ntl8 and NTL8-HA under LD/SD conditions. Plants were grown under LD (16 h light/8 h dark) or SD (8 h light/16 h dark) conditions, with flowering time determined by total rosette leaves number at bolting) (n = 3, mean ± SD; **p < 0.01, t-test). (B) FT expression differences in Col, ntl8 and NTL8-HA under LD/SD. Two-week-old seedlings grown under LD or SD were sampled at ZT16 (LD) or ZT8 (SD) for RT-qPCR analysis, PP2A was used as internal control; n = 3, mean ± SD; **p < 0.01, t-test). (C) Flowering time comparison among Col, ntl8 and NTL8-HA under LD. Flowering was assessed by total rosette leaves number at bolting (n = 3, mean ± SD). Different letters indicate significant differences (p < 0.05, ANOVA with Tukey’s test).
FLOWERING LOCUS T (FT) acts as a mobile florigen that promotes flowering in Arabidopsis thaliana [1,2,14]. To determine whether NTL8 acted through the regulation of FT we compared the expression levels of FT in wild-type Col, ntl8 mutants (ntl8-1 and ntl8-2), and the NTL8-HA transgenic line (the line #1 was used hereafter) under long-day (LD) conditions. As shown in Fig. 2B, the transcriptional level of FT in both ntl8-1 and ntl8-2 mutants were higher than that in NTL8-HA line, compared to the wild-type Col line. These expression dynamics correlated with the observed flowering time phenotype of ntl8 mutant and NTL8-HA lines, which supported the hypothesis that NTL8 affected flowering time by regulating FT expression. To further validate this relationship, we crossed ntl8-1 mutant and NTL8-HA line with the late-flowering ft-10 mutant to obtain the ntl8/ft and NTL8-HA/ft lines. Under LD conditions, both lines exhibited a late-flowering phenotype, similar to ft-10 mutant (Fig. 2C). Thus, these data confirm our previous hypothesis that NTL8 requires FT signal to control flowering time under LD conditions.
CO controls the expression of FT to accelerate flowering during photoperiod-dependent flowering pathway [1,6,7]. Given that NTL8 also modulates flowering time via the photoperiodic pathway, we sought to investigate the potential interaction between NTL8 and CO. To this end, we first checked their physical interaction. In yeast two-hybrid assays, yeast cells co-transformed with NTL8 and CO constructs were able to grow on the selective medium lacking Trp/Leu/His/Ade (-LWHA), suggesting a direct interaction between the two proteins in yeast (Fig. 3A). This interaction was further supported by in vitro pull-down assays, in which NTL8-GST was co-immunoprecipitated with His-CO protein extracted from bacterial lysates, whereas GST alone did not interact with His-CO (Fig. 3B). To confirm this interaction in planta, we also checked the interaction between NTL8 and CO in plant cells. As shown in Fig. 3C, we crossed the NTL8-HA transgenic line with CO-Flag line to obtain NTL8-HA/CO-Flag plants. As a negative control, we crossed the transgenic GFP-Flag line with NTL8-HA to obtain GFP-Flag/NTL8-HA plants. Total protein was extracted from the NTL8-HA/CO-Flag line and immunoprecipitated using anti-HA resin. A strong immunoblotting signal for CO-Flag was detected in the immunoprecipitated complex from the NTL8-HA/CO-Flag using an anti-Flag antibody, whereas GFP protein was not detected in the control immunoprecipitate from the GFP-Flag/NTL8-HA line. Thus, these results indicated that NTL8 interacted with CO in vitro and in vivo. To further confirm this interaction, we performed a luciferase complementation imaging (LCI) assay in Nicotiana benthamiana leaves. A strong bioluminescent signal was observed in leaves co-expressing CO-nLuc, and cLuc-NTL8, but not in those co-expressing the combination of CO-nLuc with cLUC, or nLUC with CO-cLuc (Fig. 3D). Collectively, these data provided strong evidence of the interaction of NTL8 and CO in planta.
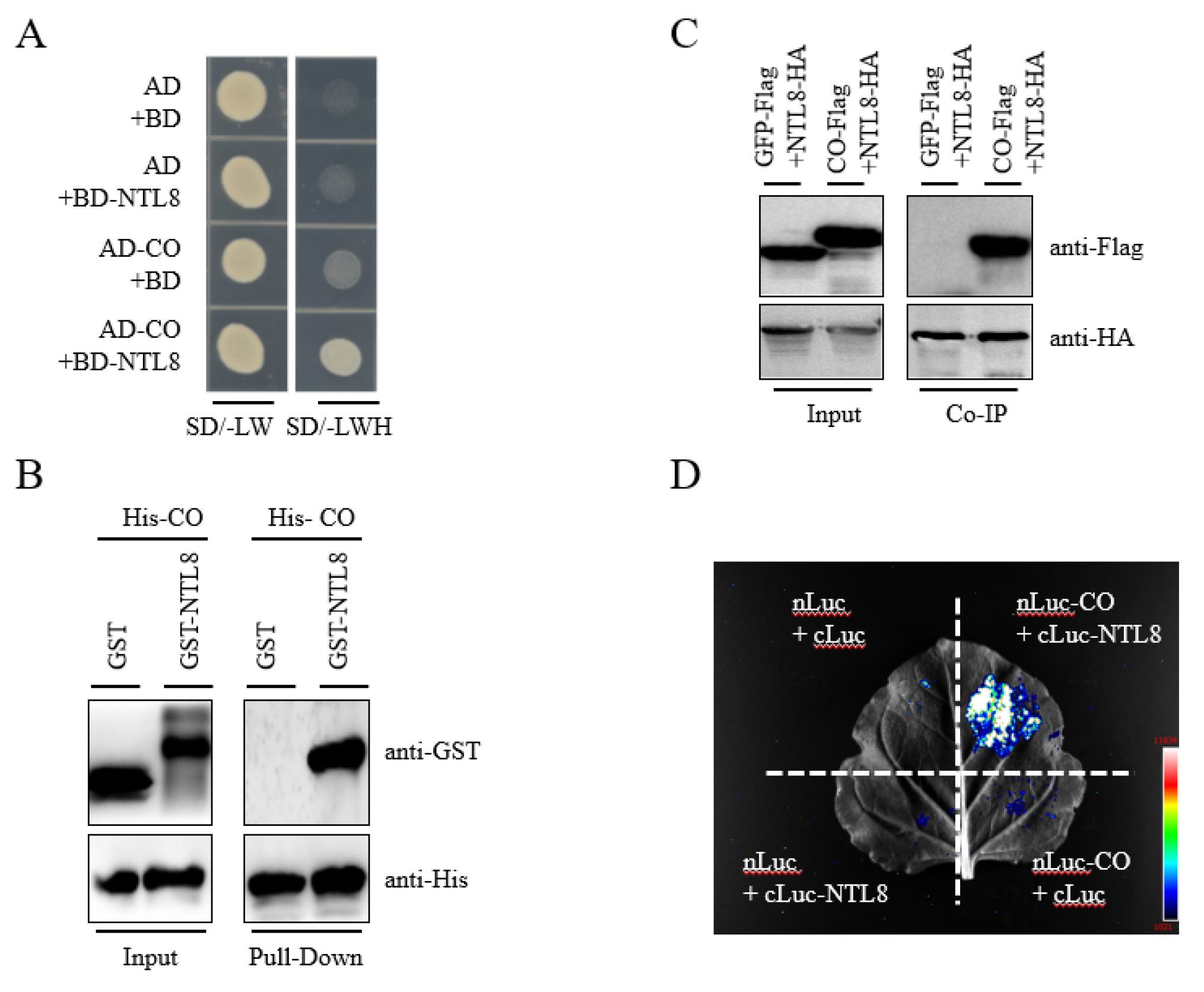
Figure 3: Interaction between NTL8 and CO in vitro and in planta. (A) Y2H assay for NTL8-CO interaction. Yeast carrying AD/BD constructs were grown on -LW or -LWH medium. (B) In vitro pull-down of His-CO with GST-NTL8. GST served as control; bound proteins detected by anti-His/GST. (C) Co-IP of NTL8-CO in Arabidopsis. HA-resin precipitated proteins from CO-Flag/NTL8-HA seeds, detected by anti-Flag/HA. GFP-Flag/NTL8-HA as control. (D) LCI assay in N. benthamiana. CO-nLUC and NTL8-cLUC were co-expressed; luminescence imaged at 48 h post-infiltration (n = 3).
3.4 NTL8 Affects the Protein Stability of CO
Given that NTL8 interacts with CO and delays flowering time, we hypothesized that NTL8 might influence CO protein stability under LD conditions, thereby modulating the flowering time. To test this possibility, we checked the circadian accumulation of CO protein in the wild-type Col line and NTL8-HA genetic background. Using a commercial anti-CO antibody, we monitored the circadian expression of CO protein levels throughout the diurnal cycle, observing a characteristic circadian expression pattern: CO protein accumulated during the daytime, peaking prior to nightfall, and was rapidly degraded during the nighttime (Fig. 4A,B). Compared to the pattern observed in Col plants, the protein abundance of CO in the NTL8-HA plants was lower at several key time points under LD conditions: ZT0 (dawn), ZT4 (dawn) and ZT20 (nighttime). Notably, at ZT4 or ZT20 points, the protein levels of CO in NTL8-HA plants were markedly lower than those in Col. However, CO protein levels at ZT12 and ZT16 did not show obvious differences between Col and NTL8-HA lines. We further compared CO protein accumulation among Col, ntl8-1 mutants, and NTL8-HA plants across different time points (Fig. 5A,B). Under LD conditions, CO protein levels in NTL8-HA plants at ZT0, ZT4 and ZT20 were significantly lower than those in ntl8-1 mutants or Col. In addition, a statistically significant differences in CO accumulation between ntl8-1 mutants and Col plants was detected at ZT16 and ZT20 (Fig. 5A,B). These results suggested that NTL8 regulates the protein stability of CO, thereby modulating flowering time under LD conditions. This finding was consistent with the observed flowering time phenotype and FT expression patterns in ntl8-1 mutants and NTL8-HA line under LD conditions. Furthermore, we assessed CO accumulation pattern among Col, ntl8-1 mutant, and NTL8-HA lines under short-day (SD) conditions. No significant differences were detected at different time points, although CO protein abundance in NTL8-HA plants was slightly lower than that in Col or ntl8-1 mutants at ZT12 (Supplementary Fig. S2). These data further support the conclusion that NTL8 affects CO-mediated flowering predominantly under LD conditions, in line with the absence of flowering time differences among the lines under SD conditions (Fig. 2A).
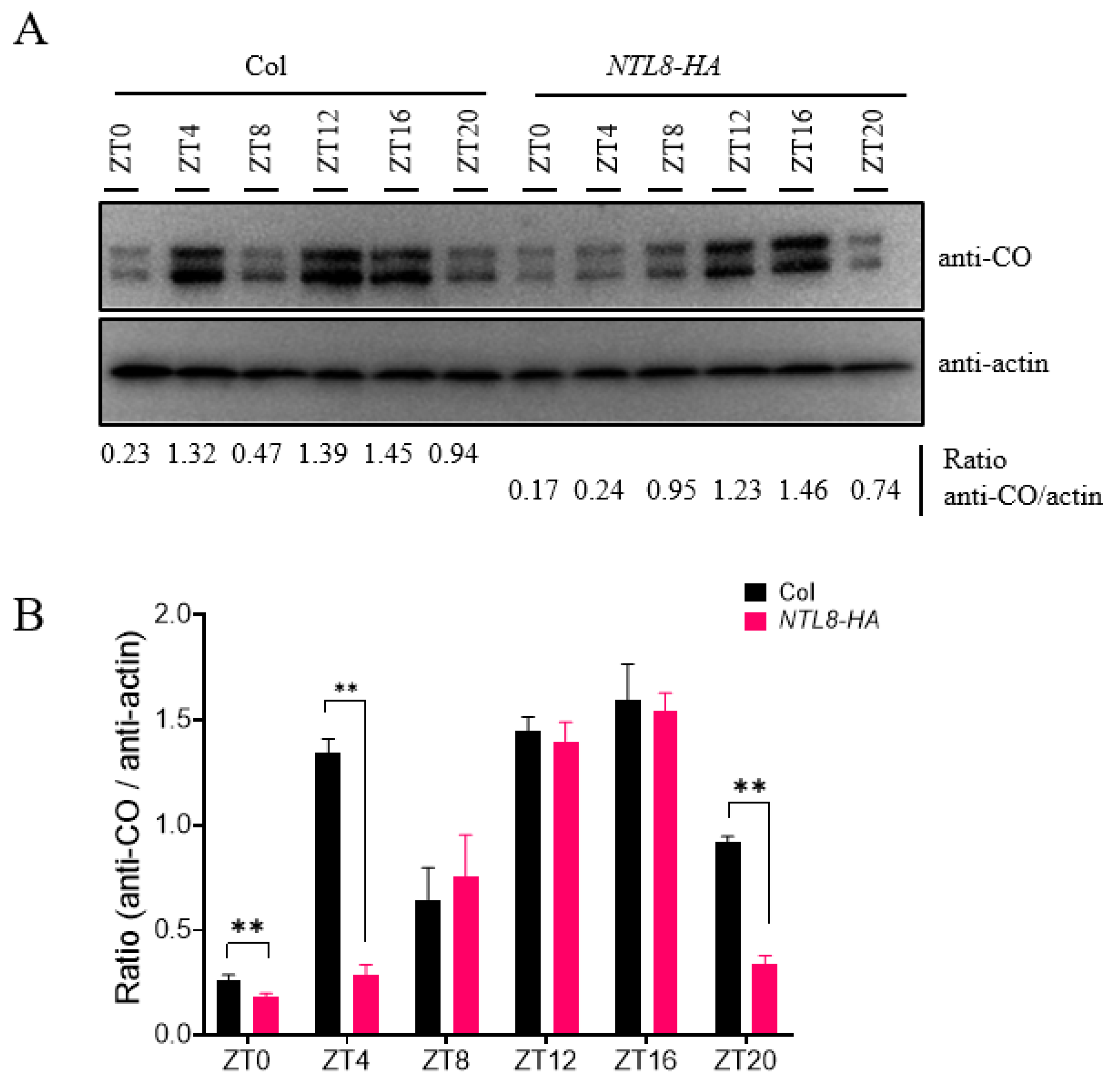
Figure 4: Diurnal oscillation of CO protein abundance in Col and NTL8-HA lines under LD conditions. (A) CO protein oscillation in Col and NTL8-HA under LD. Samples collected every 4 h (ZT0-24) for immunoblot (anti-CO/actin). The ratio of CO to Actin was evaluated by the immunoblot signal of anti-CO and anti-actin antibodies, and is listed at the bottom of the figure. (B) Quantified CO/Actin ratios in Col vs. NTL8-HA. Data from three replicates (mean ± SD; **p < 0.01, t-test).
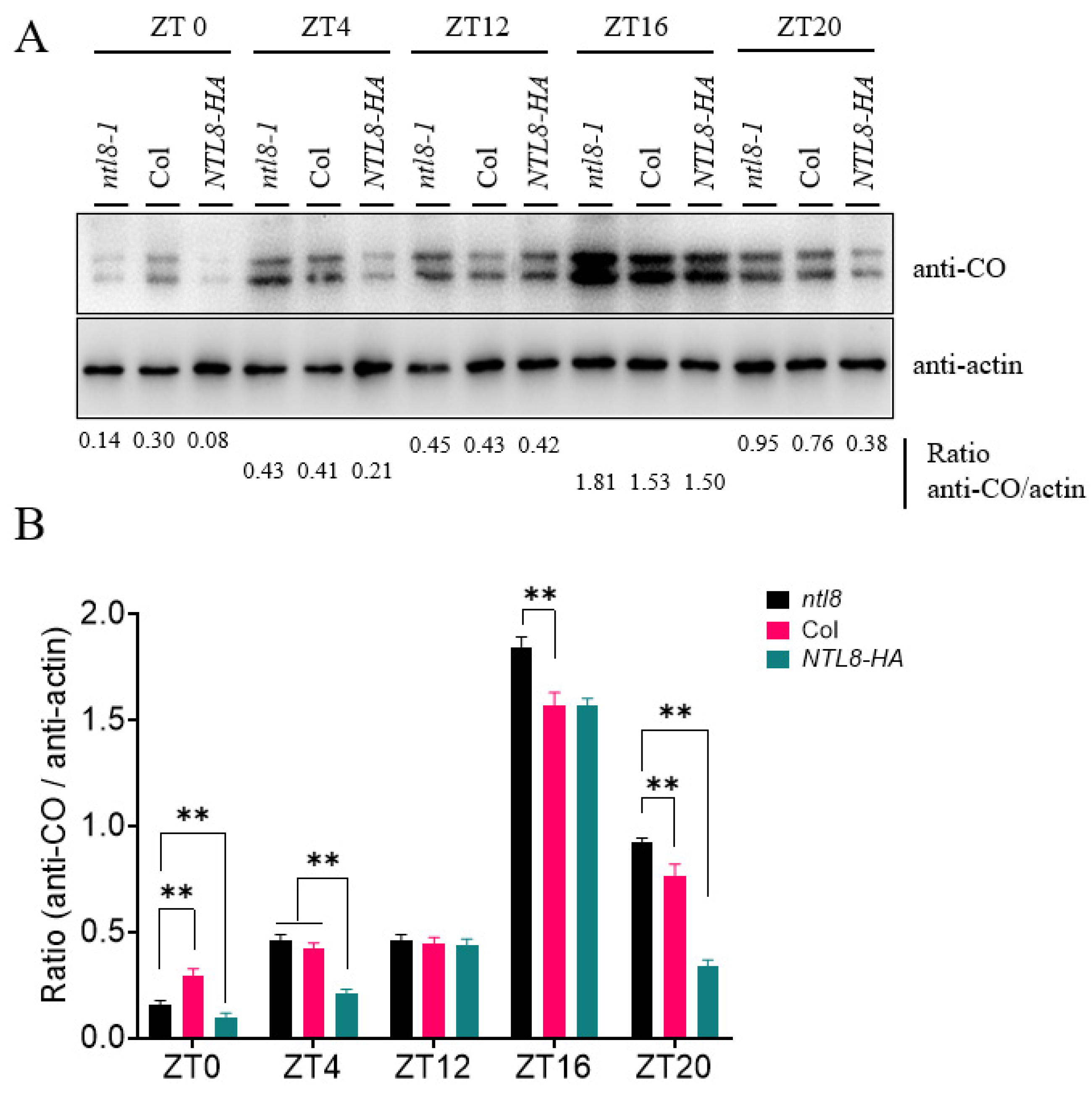
Figure 5: Differential oscillation of CO protein abundance among Col, ntl8-1 and NTL8-HA lines under LD conditions. (A) CO protein levels in Col, ntl8-1 and NTL8-HA at ZT4/12/16/22 (LD). The ratio of CO to Actin was evaluated by the immunoblot signal for anti-CO or anti-actin antibody, and is listed at the bottom of the figure. (B) Quantified CO/Actin ratios (Col vs. mutants). Data are presented as mean ± standard deviation (SD). Statistical differences were analyzed by Student’s t-test analysis (**p < 0.01).
3.5 NTL8 Antagonizes the Effect of CO on Activating FT Expression
To better understand the genetic relationship between NTL8 and CO, we generated CO-Flag/NTL8-HA and CO-Flag/ntl8 lines through reciprocal genetic crosses. As previously observed, the NTL8-HA line showed a late flowering phenotype characterized by an increased the rosette leaf number, whereas the ntl8-1 mutant showed an early flowering phenotype with fewer rosette leaves (Fig. 2A). Consistent with previous results, the CO-Flag line also showed an early flowering phenotype with reduced rosette leaf number. In this study, we found that overexpression of NTL8-HA in the CO-Flag/NTL8-HA line partially delayed the early flowering phenotype of the CO-Flag line (Fig. 6A). Correspondingly, the FT expression level in CO-Flag/NTL8-HA plants at ZT16 was slightly lower than that in CO-Flag plants (Fig. 6B). These results suggest that NTL8 antagonizes the early flowering phenotype of CO-Flag by suppressing FT expression. Furthermore, we observed that both of CO-Flag/ntl8 and ntl8-1 lines showed similar earlier flowering time (Fig. 6A), with elevated expression of FT at ZT16 under LD conditions, significantly higher than in ntl8 plants. These findings indicated that inactivation of NTL8 in ntl8 mutant line further enhanced the effect of CO-Flag in promoting FT expression, supporting the notion of an antagonistic relationship between CO and NTL8 in regulating FT expression.
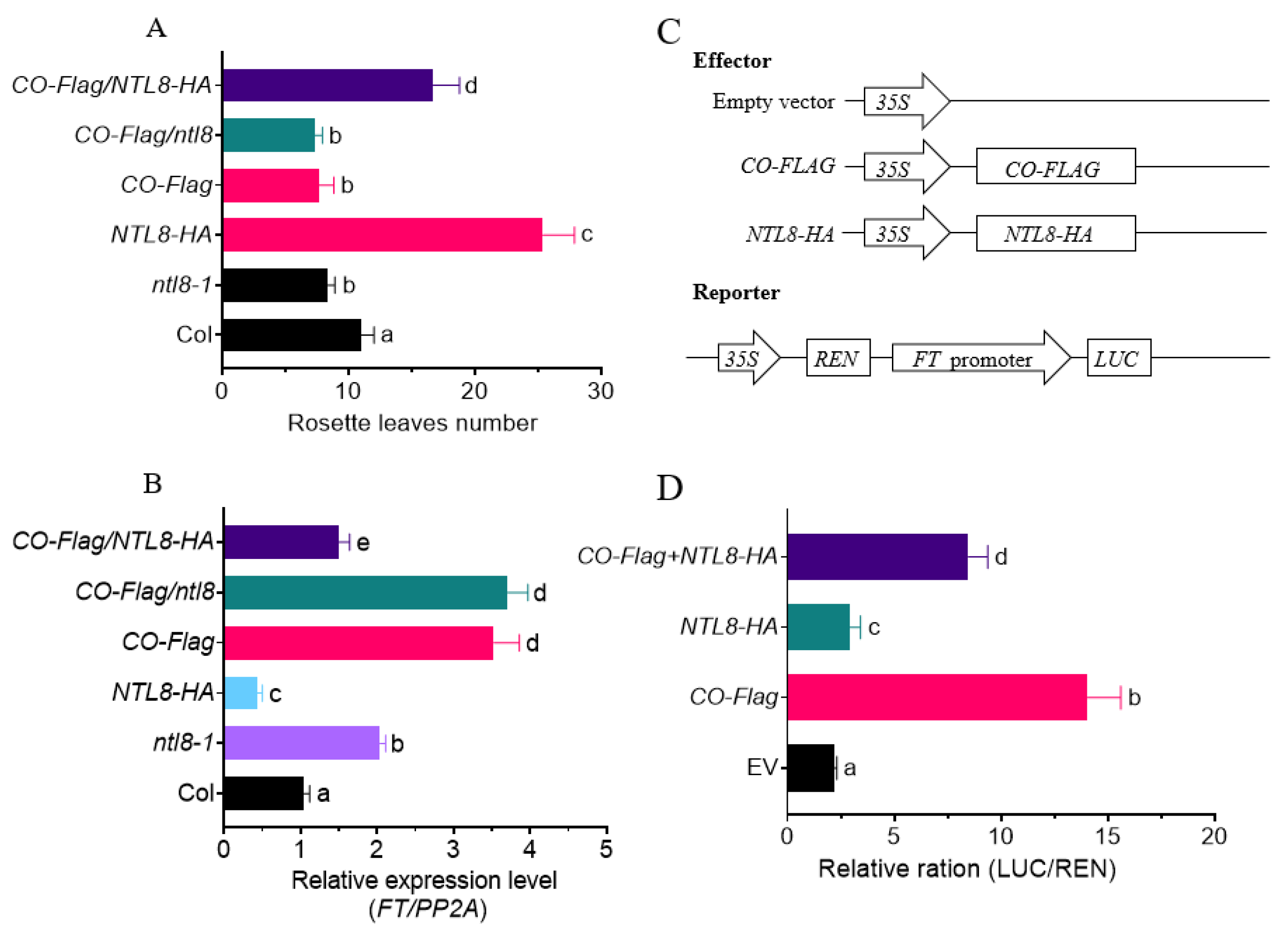
Figure 6: NTL8 and CO antagonistically regulate the expression of FT and flowering time. Bars labeled with different letters are significantly different at p < 0.05 (Tukey’s test). (A) Different flowering time comparison among Col, ntl8-1, NTL8-HA and different crossed line under LD. (n = 3, mean ± SD). (B) FT expression levels among Col, ntl8-1, NTL8-HA and different crossed line under LD at ZT16 by RT-qPCR analysis. (n = 3, mean ± SD). PP2A was used as the internal control. (C) Schematic diagram of the effector and reporter constructs used in the transient transactivation assay. The reporter contains FT-promoter-driven Luciferase genes. (D) NTL8 modulates CO-activated FT expression by transient expression assay. The firefly luciferase and Renilla luciferase (LUC: REN) ratio represents FTpro:LUC activity relative to the internal control (35Spro: REN). (n = 3, mean ± SD).
Since CO directly binds to the promoter of FT to activate its expression, whereas NTL8 suppresses the expression of FT, this suggests an antagonistic relationship between CO and NTL8 in regulating FT expression. To test this hypothesis, we performed transient protoplast transformation assays to evaluate the effect of NTL8 on CO-mediated activation of FT expression in Arabidopsis leaf protoplasts. As shown in Fig. 6C,D, CO efficiently activated FT expression; however, this activation was attenuated when NTL8-HA was co-expressed with CO-Flag effector. As a control, expression of NTL8-HA alone with the FTpro::LUC reporter also partially suppressed LUC expression (Fig. 6D). Collectively, these results confirmed that NTL8 antagonized CO to suppress FT expression, thereby delaying flowering time.
Previous studies have demonstrated that CO upregulates histone acetylation at the FT locus to epigenetically activate FT expression [1,2,3,4,5,6,7]. Given that NTL8 counteracts CO to suppress FT expression, we speculated that NTL8 may attenuate CO-mediated histone acetylation at the FT locus. To test this possibility, we performed ChIP-qPCR analysis to measure three different regions of the FT locus, including the P1 region near the transcription start site (TSS) (FT-P, −297 bp–128 bp upstream of the start codon), the P2 region within the first exon close to the start codon (FT-E, +17 bp to +221 bp downstream of start codon), and the P3 region within the first intron of FT genome (FT-I, +568 bp to +785 bp downstream of the start codon) (Fig. 7). These regions were previously reported to show increased histone acetylation upon CO overexpression. Consistent with these findings, we observed a relatively higher level of histone acetylation at the FT locus in the CO-Flag line compared with Col wild type at ZT16 under LD conditions. In contrast, histone acetylation levels were lower in the NTL8-HA line and higher in the ntl8-1 mutant line, which was consistent with the FT expression pattern and their different flowering time phenotypes. Furthermore, compared with CO-Flag line, overexpressing NTL8 in CO-Flag/NTL8-HA reduced histone acetylation levels. On the contrary, silencing NTL8 in the CO-Flag/ntl8 line increased histone acetylation levels. These findings suggest that NTL8 suppresses the expression of FT by modulating CO-mediated H3 acetylation levels at FT locus.
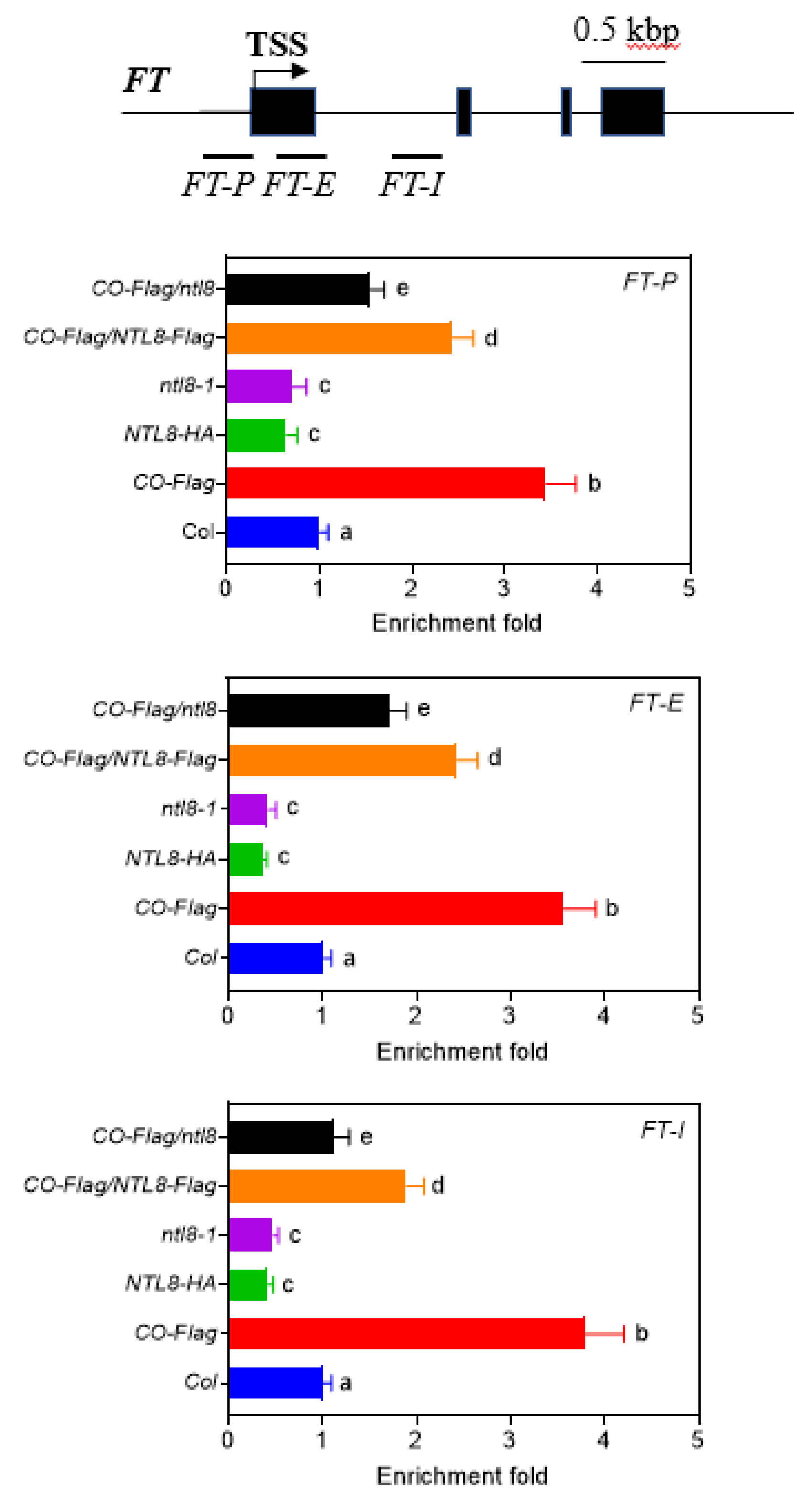
Figure 7: NTL8 represses CO-mediated upregulation of histone H3 acetylation level at the FT locus by ChIP-PCR analysis at the indicated regions (FT-P, FT-E, and FT-L fragments within the FT locus). The Col, CO-Flag, NTL8-HA, ntl8-1, CO-Flag/NTL8-HA and CO-Flag/ntl8 lines were used. The seedlings were grown under LD conditions for 2 weeks and leaf samples were collected at ZT16 for ChIP-PCR analysis. An antibody against acetylated histone H3 was used for immunoprecipitation. ACTIN2 was used as an internal control, and relative histone H3 acetylation levels at the FT locus were normalized to those of ACTIN2. The detailed positions of FT-P, FT-E, and FT-L are indicated by bars below the FT gene. Exons are shown as solid boxes, and introns as solid lines. Data are presented as mean ± standard deviation (SD) of three independent experiments. Bars with different letters indicate values that are significantly different at p < 0.05 (Tukey’s test).
4.1 NTL8 Acts As the Novel Modulator to Control Photoperiodic Flowering Time
Previous studies have reported that saline stress induced the expression of NTL8 [33,36]. In the present study, we additionally demonstrated that NTL8 was light-responsive and presented a circadian rhythm, with elevated transcript levels during the daytime and a marked reduction during the nighttime, similar to CO, the core regulator of the photoperiodic flowering pathway (Fig. 1 and Fig. 2). Consistent with prior findings, our data confirmed that NTL8 negatively controlled flowering time under long-day (LD) conditions, as overexpressing NTL8 delayed flowering time, whereas loss-of-function mutant of ntl8 showed earlier flowering than the wild type (Fig. 2). FT encodes a mobile florigen that promotes flowering time, and CO binds to the promoter of FT to activate its expression, thereby promoting flowering [1,14]. In line with this, we observed that FT transcript levels in ntl8 were higher than that in NTL8-HA line (Fig. 2D). Genetic analysis further supported the conclusion that NTL8 acted upstream of FT to regulate flowering time. Specifically, both of ntl8/ft double mutant and NTL8-HA/ft line showed later flowering time similar to that of the ft-10 mutant (Fig. 2E), indicating that NTL8 mediated photoperiodic flowering through targeting FT. Consistent with our findings, previous studies have shown that long-term cold treatment, or vernalization, efficiently suppresses the flowering time of the NTL8-HA line, similar to the wild-type Col line. Furthermore, a dominant mutation of NTL8 was found to constitutively activate the expression of VIN3, leading to the silencing of FLC during vernalization, suggesting a complex role for NTL8 in controlling flowering time in response to cold conditions [36]. Given that both NTL8 and NTL14 contain transmembrane domain (TMD) and cooperatively modulate seed dormancy [40], and that NTL14 has also been reported to control flowering time by modulating FLC, the negative component for flowering time [31], it is plausible that NTL8 may similarly influence the expression of FLC to control flowering time. Moreover, ANAC060, which contained TMD and is clustered into the same phylogenetic clade as NTL8, ANAC060 suppresses the expression of ABI5, which is the essential signal transmitter of ABA signaling [39,40,41]. Since ABI5 also plays a role in modulating flowering time, it would be interesting to investigate whether NTL8 controls flowering time through an ABI5-dependent pathway [39,41]. Similar to cold treatment, GA treatment reversed the later-flowering phenotype of NTL8-HA line. Interestingly, GA treatment can reduce the expression of NTL8, and the seed germination of ntl8 showed increased tolerance to GA biosynthesis inhibitors [33,36], suggesting distinct functions of NTL8 in seed germination and flowering time regulation. In contrast, saline treatment induces the expression of NTL8, and overexpressing NTL8 delayed the flowering time, consistent with the late-flowering phenotype of NTL8-HA transgenic plant under salt stress. As previous studies reported that saline stress reduces FT expression, we also observed lower FT expression level in the NTL8-HA line (Fig. 2), supporting the hypothesis that salt stress delays flowering time by activating NTL8 and subsequently reducing FT signaling. Furthermore, since salt stress has been shown to induce the expression of BFT, which competes with FT for FD bindings and suppresses the activity of FT [36], it will be important to further investigate whether saline-induced flowering delay involves NTL8-mediated regulation of BFT.
4.2 NTL8 Interacts with CO to Influence Its Transactivation to FT Expression
CO acts as a central hub in the photoperiodic flowering regulatory network [1,6]. Its transcription presented a circadian rhythm, and its protein abundance also showed rhythmic oscillation, accumulating during the daytime to efficiently activate FT expression and subsequently being degraded by the E3 ubiquitin ligase COP1 during the night [8]. Our previous research showed that the ABA associated protein AFP2 epigenetically repressed the transcriptional level of CO and promoted its proteasomal degradation, thereby delaying flowering time in Arabidopsis [39]. In the present study, multiple lines of evidence, including yeast two hybrid, in vitro pull-down assays, and in vivo CO-IP, confirmed the physical interaction between CO and NTL8 (Fig. 3). Genetic analysis further revealed that the ntl8 mutant exhibited an early-flowering phenotype, while the NTL8-HA line showed a delayed-flowering phenotype under LD conditions, but not under SD conditions (Fig. 2), suggesting a photoperiodic-dependent regulatory role. Moreover, overexpressing NTL8 delayed the early-flowering phenotype of CO-Flag (Fig. 6), further supporting an antagonistic interaction between NTL8 and CO in controlling photoperiodic flowering time. Consistent with this, transient protoplast transformation assays showed that overexpressing NTL8 attenuated the transcriptional action of CO on FT expression (Fig. 6), providing mechanistic insights into how NTL8 antagonizes the acceleration effect of CO on FT expression. In addition, our results showed the circadian oscillation of NTL8 transcript, with high transcript levels during the daytime and declining during the nighttime (Fig. 1). This diurnal expression pattern of NTL8 suggests that it may fine-tune CO activity by repressing FT expression during the day, thereby contributing to the precise temporal regulation of flowering time.
The NTL8 homolog ANAC089 has been reported to bind to the promoter of FLC to suppress its expression, thereby promoting earlier flowering [31]. Similarly, another homolog, ANAC060 binds to the promoter of ABI5 to suppress its expression and attenuate ABA signaling [40]. However, in the case of NTL8, we found no direct evidence of binding of NTL8 to FT promoter, although NTL8 clearly influenced FT expression. One possible explanation for this discrepancy lies in the complex and highly variable presence of cis-elements within the promoter of FT. Previous studies have reported that key enhancer elements of FT can reside more than 10 kb upstream from the start codon [42], suggesting that our inability to detect the direct binding of NTL8 may be due to not targeting the correct cis-regulatory region within the FT promoter. Additionally, previous studies have demonstrated that histone deacetylase (HDAC) complexes, such as those containing SAP30 function-related proteins 1/2 (AFR1/2) epigenetically decreases FT transcription at the end of day by reducing acetylation level at FT locus, Furthermore, CO was shown to interact with MRG1/2, which then recruits the histone deacetylase HADC to reduce the histone acetylation level at the FT locus, thereby reducing its expression. These findings raise the possibility that NTL8 suppresses the expression of FT through a similar epigenetic mechanism. In support of this hypothesis, we observed that overexpression of NTL8 in CO-Flag genetic background reduced histone acetylation level at the FT locus, whereas silencing NTL8 in the CO-Flag genetic background line led to an increase in acetylation levels at the FT locus (Fig. 7). Collectively, these results suggest that NTL8 may recruit HDAC complexes to the FT locus to epigenetically reduce its expression at dusk, providing a mechanistic explanation for the antagonistic relationship between NTL8 and CO in regulating FT expression and flowering time.
In addition to the circadian rhythm observed at the transcript level, the protein abundance of CO also presented a circadian pattern, characterized by accumulation during the daytime, and degradation during the nighttime. Several E3 ubiquitin ligases, such as HOS1 and COP1, have been reported to mediate CO degradation [8,11]. Our previous study showed that AFP2 interacts with CO and promotes its degradation during the night [23]. In the present study, we observed a relatively lower protein abundance of CO in the NTL8-HA line, particularly at ZT20 during the nighttime, where CO levels were markedly lower than those in wild-type Col plants. These findings suggest that overexpression of NTL8 induced the protein degradation of CO during the night. It is possible that NTL8 synergistically enhanced the activity of E3 ligases, such as COP1, thereby promoting the degradation of CO during the nighttime. Further investigation will be necessary to elucidate whether, and through which mechanisms, NTL8 influences the protein abundance and stability of CO via specific E3 ligases such as COP1. Notably, the reduced protein abundance of CO observed at ZT4 (dawn) in the NTL8-HA transgenic line may explain the reduced FT expression observed in these plants. This finding suggested that NTL8 may also suppress CO accumulation during the daytime. Previous studies have reported that the E3 ligase HOS1 also affects CO accumulation during dawn. Therefore, it is plausible that NTL8 influences CO accumulation in the morning through modulation of HOS1 activity. Nevertheless, further investigation will be required to clarify these potential regulatory mechanisms.
In summary, this study expanded our understanding of the role of NTL8 in the regulation of flowering time. Our results showed that NTL8 interacted with CO to suppress FT expression, thereby delaying flowering under long-day conditions. A combination of physiological, biochemical, and genetic analyses further supported that NTL8 antagonized CO function to suppress FT transcription. In addition, we found that NTL8 influenced the protein stability of CO during the nighttime, acting at a post-transcriptional level. Collectively, our findings contribute to a deeper understanding of the mechanisms underlying the CO-mediated photoperiodic flowering pathway, and suggest that the NTL8-CO module represents a novel regulatory mechanism that requires further attention, especially for its potential applications in the genetic modification of flowering time in horticultural research.
In this study, we performed genetic analysis to confirm that NTL8 negatively regulates flowering time in Arabidopsis, as overexpressing NTL8 delays flowering time but silencing NTL8 shows early flowering time. Further analysis shows NTL8 itself presents circadian expression, and NTL8 also interacts with CO, thereby destabilize CO stability to repress FT expression, resulting into late flowering. Thus, we propose a uncovered mechanism of NTL8 in coordinating flowering time through antagonizing CO to reduce FT expression.
Acknowledgement:
Funding Statement: This work was supported by the National Natural Science Foundation of China (31970289).
Author Contributions: Yue Jin: investigation, methodology, resources, writing–original draft. Xiaoying Tang: investigation, methodology, conceptualization. Bingke Wei: methodology, validation. Lulu Zhi: investigation, methodology. Yan Mao: investigation, methodology. García-Caparrós Pedro: writing—original draft, writing—review & editing. Xiangyang Hu: conceptualization, methodology, resources, writing—original draft, writing—review & editing, supervision, funding acquisition, project administration. All authors reviewed and approved the final version of the manuscript.
Availability of Data and Materials: The authors confirm that the data supporting the findings of this study are available within the article.
Ethics Approval: Not applicable.
Conflicts of Interest: The authors declare no conflicts of interest.
Supplementary Materials: The supplementary material is available online at https://www.techscience.com/doi/10.32604/phyton.2026.067796/s1. Supplementary Table S1: List of primers used in this study. Supplementary Figure S1: Identification of ntl8 mutant or NTL8-HA lines. (A,B) Verification of the ntl8 mutant and its overexpressed lines. The genomic structure of NTL8 gene and its T-DNA insertion (A). The location of T-DNA in the ABI4 locus was detected by specific primers pairs (B);The expression of NTL8-HA in the transgenic line overexpressing NTL8-HA was detected by western blotting analysis using anti-HA antibody (C). Supplementary Figure S2: The dynamic change of CO abundance of Col, ntl8-1 and NTL8-HA line under SD condition (8-h D/16-h L). (A) The leave samples from wild-type Col, ntl8 mutant and the transgenic NTL8-HA backgrounds were collected at ZT4, ZT12, ZT16 and ZT22 under SD conditions. The total proteins were extracted for CO protein abundance analysis and anti-actin was used as the internal control (upper panel). The ratio of CO to Actin was evaluated by the immunoblot signal for anti-CO and anti-actin, and is listed at the bottom of the figure. (B) The ratio of CO to Actin in the extracted protein wild-type Col, ntl8 mutant and the transgenic NTL8-HA lines were evaluated by the immunoblot signal for anti-CO and anti-actin based on three individual western blotting experiments, and the quantification degree were presented. Three biological replications were performed. Data are presented as mean ± standard deviation (SD). Statistical differences were analyzed by t-test analysis, ns indicates that the result is not statistically significant.
References
1. Freytes SN , Canelo M , Cerdán PD . Regulation of flowering time: When and where? Curr Opin Plant Biol. 2021; 63: 102049. doi:10.1016/j.pbi.2021.102049. [Google Scholar] [CrossRef]
2. Johansson M , Staiger D . Time to flower: Interplay between photoperiod and the circadian clock. J Exp Bot. 2015; 66( 3): 719– 30. doi:10.1093/jxb/eru441. [Google Scholar] [CrossRef]
3. Jackson SD . Plant responses to photoperiod. New Phytol. 2009; 181( 3): 517– 31. doi:10.1111/j.1469-8137.2008.02681.x. [Google Scholar] [CrossRef]
4. Imaizumi T . Arabidopsis circadian clock and photoperiodism: Time to think about location. Curr Opin Plant Biol. 2010; 13( 1): 83– 9. doi:10.1016/j.pbi.2009.09.007. [Google Scholar] [CrossRef]
5. Susila H , Nasim Z , Ahn JH . Ambient temperature-responsive mechanisms coordinate regulation of flowering time. Int J Mol Sci. 2018; 19( 10): 3196. doi:10.3390/ijms19103196. [Google Scholar] [CrossRef]
6. Shim JS , Kubota A , Imaizumi T . Circadian clock and photoperiodic flowering in Arabidopsis: CONSTANS is a hub for signal integration. Plant Physiol. 2017; 173( 1): 5– 15. doi:10.1104/pp.16.01327. [Google Scholar] [CrossRef]
7. Putterill J , Robson F , Lee K , Simon R , Coupland G . The CONSTANS gene of Arabidopsis promotes flowering and encodes a protein showing similarities to zinc finger transcription factors. Cell. 1995; 80( 6): 847– 57. doi:10.1016/0092-8674(95)90288-0. [Google Scholar] [CrossRef]
8. Liu LJ , Zhang YC , Li QH , Sang Y , Mao J , Lian HL , et al. COP1-mediated ubiquitination of CONSTANS is implicated in cryptochrome regulation of flowering in Arabidopsis. Plant Cell. 2008; 20( 2): 292– 306. doi:10.1105/tpc.107.057281. [Google Scholar] [CrossRef]
9. Lazaro A , Valverde F , Piñeiro M , Jarillo JA . The Arabidopsis E3 ubiquitin ligase HOS1 negatively regulates CONSTANS abundance in the photoperiodic control of flowering. Plant Cell. 2012; 24( 3): 982– 99. doi:10.1105/tpc.110.081885. [Google Scholar] [CrossRef]
10. Cheng Z , Zhang X , Huang P , Huang G , Zhu J , Chen F , et al. Nup96 and HOS1 are mutually stabilized and gate CONSTANS protein level, conferring long-day photoperiodic flowering regulation in Arabidopsis. Plant Cell. 2020; 32( 2): 374– 91. doi:10.1105/tpc.19.00661. [Google Scholar] [CrossRef]
11. Jung JH , Seo PJ , Park CM . The E3 ubiquitin ligase HOS1 regulates Arabidopsis flowering by mediating CONSTANS degradation under cold stress. J Biol Chem. 2012; 287( 52): 43277– 87. doi:10.1074/jbc.m112.394338. [Google Scholar] [CrossRef]
12. Takagi H , Hempton AK , Imaizumi T . Photoperiodic flowering in Arabidopsis: Multilayered regulatory mechanisms of CONSTANS and the florigen FLOWERING LOCUS T. Plant Commun. 2023; 4( 3): 100552. doi:10.1016/j.xplc.2023.100552. [Google Scholar] [CrossRef]
13. Zhang B , Feng M , Zhang J , Song Z . Involvement of CONSTANS-like proteins in plant flowering and abiotic stress response. Int J Mol Sci. 2023; 24( 23): 16585. doi:10.3390/ijms242316585. [Google Scholar] [CrossRef]
14. Wigge PA . FT, a mobile developmental signal in plants. Curr Biol. 2011; 21( 9): R374– 8. doi:10.1016/j.cub.2011.03.038. [Google Scholar] [CrossRef]
15. Miskolczi P , Singh RK , Tylewicz S , Azeez A , Maurya JP , Tarkowská D , et al. Long-range mobile signals mediate seasonal control of shoot growth. Proc Natl Acad Sci U S A. 2019; 116( 22): 10852– 7. doi:10.1073/pnas.1902199116. [Google Scholar] [CrossRef]
16. Ito S , Song YH , Josephson-Day AR , Miller RJ , Breton G , Olmstead RG , et al. FLOWERING BHLH transcriptional activators control expression of the photoperiodic flowering regulator CONSTANS in Arabidopsis. Proc Natl Acad Sci U S A. 2012; 109( 9): 3582– 7. doi:10.1073/pnas.1118876109. [Google Scholar] [CrossRef]
17. Kubota A , Ito S , Shim JS , Johnson RS , Song YH , Breton G , et al. TCP4-dependent induction of CONSTANS transcription requires GIGANTEA in photoperiodic flowering in Arabidopsis. PLoS Genet. 2017; 13( 6): e1006856. doi:10.1371/journal.pgen.1006856. [Google Scholar] [CrossRef]
18. Song YH , Smith RW , To BJ , Millar AJ , Imaizumi T . FKF1 conveys timing information for CONSTANS stabilization in photoperiodic flowering. Science. 2012; 336( 6084): 1045– 9. doi:10.1126/science.1219644. [Google Scholar] [CrossRef]
19. Imaizumi T , Tran HG , Swartz TE , Briggs WR , Kay SA . FKF1 is essential for photoperiodic-specific light signalling in Arabidopsis. Nature. 2003; 426( 6964): 302– 6. doi:10.1038/nature02090. [Google Scholar] [CrossRef]
20. Imaizumi T , Schultz TF , Harmon FG , Ho LA , Kay SA . FKF1 F-box protein mediates cyclic degradation of a repressor of CONSTANS in Arabidopsis. Science. 2005; 309( 5732): 293– 7. doi:10.1126/science.1110586. [Google Scholar] [CrossRef]
21. Endo M , Kudo D , Koto T , Shimizu H , Araki T . Light-dependent destabilization of PHL in Arabidopsis. Plant Signal Behav. 2014; 9( 3): e28118. doi:10.4161/psb.28118. [Google Scholar] [CrossRef]
22. Zhang B , Wang L , Zeng L , Zhang C , Ma H . Arabidopsis TOE proteins convey a photoperiodic signal to antagonize CONSTANS and regulate flowering time. Genes Dev. 2015; 29( 9): 975– 87. doi:10.1101/gad.251520.114. [Google Scholar] [CrossRef]
23. Chang G , Wang C , Kong X , Chen Q , Yang Y , Hu X . AFP2 as the novel regulator breaks high-temperature-induced seeds secondary dormancy through ABI5 and SOM in Arabidopsis thaliana. Biochem Biophys Res Commun. 2018; 501( 1): 232– 8. doi:10.1016/j.bbrc.2018.04.222. [Google Scholar] [CrossRef]
24. Mathew IE , Agarwal P . May the fittest protein evolve: Favoring the plant-specific origin and expansion of NAC transcription factors. Bioessays. 2018; 40( 8): e1800018. doi:10.1002/bies.201800018. [Google Scholar] [CrossRef]
25. Chakraborty R , Roy S . Evaluation of the diversity and phylogenetic implications of NAC transcription factor members of four reference species from the different embryophytic plant groups. Physiol Mol Biol Plants. 2019; 25( 2): 347– 59. doi:10.1007/s12298-018-0581-9. [Google Scholar] [CrossRef]
26. Kim SY , Kim SG , Kim YS , Seo PJ , Bae M , Yoon HK , et al. Exploring membrane-associated NAC transcription factors in Arabidopsis: Implications for membrane biology in genome regulation. Nucleic Acids Res. 2007; 35( 1): 203– 13. doi:10.1093/nar/gkl1068. [Google Scholar] [CrossRef]
27. Kim SG , Lee S , Seo PJ , Kim SK , Kim JK , Park CM . Genome-scale screening and molecular characterization of membrane-bound transcription factors in Arabidopsis and rice. Genomics. 2010; 95( 1): 56– 65. doi:10.1016/j.ygeno.2009.09.003. [Google Scholar] [CrossRef]
28. Breeze E , Vale V , McLellan H , Pecrix Y , Godiard L , Grant M , et al. A tell tail sign: A conserved C-terminal tail-anchor domain targets a subset of pathogen effectors to the plant endoplasmic reticulum. J Exp Bot. 2023; 74( 10): 3188– 202. doi:10.1093/jxb/erad075. [Google Scholar] [CrossRef]
29. Lee S , Seo PJ , Lee HJ , Park CM . A NAC transcription factor NTL4 promotes reactive oxygen species production during drought-induced leaf senescence in Arabidopsis. Plant J. 2012; 70( 5): 831– 44. doi:10.1111/j.1365-313X.2012.04932.x. [Google Scholar] [CrossRef]
30. Albertos P , Tatematsu K , Mateos I , Sánchez-Vicente I , Fernández-Arbaizar A , Nakabayashi K , et al. Redox feedback regulation of ANAC089 signaling alters seed germination and stress response. Cell Rep. 2021; 35( 11): 109263. doi:10.1016/j.celrep.2021.109263. [Google Scholar] [CrossRef]
31. Li J , Zhang J , Wang X , Chen J . A membrane-tethered transcription factor ANAC089 negatively regulates floral initiation in Arabidopsis thaliana. Sci China Life Sci. 2010; 53( 11): 1299– 306. doi:10.1007/s11427-010-4085-2. [Google Scholar] [CrossRef]
32. De Clercq I , Vermeirssen V , Van Aken O , Vandepoele K , Murcha MW , Law SR , et al. The membrane-bound NAC transcription factor ANAC013 functions in mitochondrial retrograde regulation of the oxidative stress response in Arabidopsis. Plant Cell. 2013; 25( 9): 3472– 90. doi:10.1105/tpc.113.117168. [Google Scholar] [CrossRef]
33. Kim SG , Lee AK , Yoon HK , Park CM . A membrane-bound NAC transcription factor NTL8 regulates gibberellic acid-mediated salt signaling in Arabidopsis seed germination. Plant J. 2008; 55( 1): 77– 88. doi:10.1111/j.1365-313X.2008.03493.x. [Google Scholar] [CrossRef]
34. Tian H , Wang X , Guo H , Cheng Y , Hou C , Chen JG , et al. NTL8 regulates trichome formation in Arabidopsis by directly activating R3 MYB genes TRY and TCL1. Plant Physiol. 2017; 174( 4): 2363– 75. doi:10.1104/pp.17.00510. [Google Scholar] [CrossRef]
35. Zhao Y , Antoniou-Kourounioti RL , Calder G , Dean C , Howard M . Temperature-dependent growth contributes to long-term cold sensing. Nature. 2020; 583( 7818): 825– 9. doi:10.1038/s41586-020-2485-4. [Google Scholar] [CrossRef]
36. Clough SJ , Bent AF . Floral dip: A simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 1998; 16( 6): 735– 43. doi:10.1046/j.1365-313x.1998.00343.x. [Google Scholar] [CrossRef]
37. Ying S , Jing S , Cheng L , Sun H , Tian Y , Zhi L , et al. Allantoin alleviates seed germination thermoinhibition in Arabidopsis. Phyton. 2022; 91( 9): 1893– 904. doi:10.32604/phyton.2022.022679. [Google Scholar] [CrossRef]
38. Lu S , Hu Y , Chen Y , Yang Y , Jin Y , Li P , et al. Putrescine enhances seed germination tolerance to heat stress in Arabidopsis thaliana. Phyton. 2022; 91( 9): 1879– 91. doi:10.32604/phyton.2022.022605. [Google Scholar] [CrossRef]
39. Chang G , Yang W , Zhang Q , Huang J , Yang Y , Hu X . ABI5-BINDING PROTEIN2 coordinates CONSTANS to delay flowering by recruiting the transcriptional corepressor TPR2. Plant Physiol. 2019; 179( 2): 477– 90. doi:10.1104/pp.18.00865. [Google Scholar] [CrossRef]
40. Song S , Willems LAJ , Jiao A , Zhao T , Eric Schranz M , Bentsink L . The membrane associated NAC transcription factors ANAC060 and ANAC040 are functionally redundant in the inhibition of seed dormancy in Arabidopsis thaliana. J Exp Bot. 2022; 73( 16): 5514– 28. doi:10.1093/jxb/erac232. [Google Scholar] [CrossRef]
41. Wang Y , Li L , Ye T , Lu Y , Chen X , Wu Y . The inhibitory effect of ABA on floral transition is mediated by ABI5 in Arabidopsis. J Exp Bot. 2013; 64( 2): 675– 84. doi:10.1093/jxb/ers361. [Google Scholar] [CrossRef]
42. Cao S , Kumimoto RW , Gnesutta N , Calogero AM , Mantovani R , Holt BF 3rd . A distal ccaat/nuclear factor y complex promotes chromatin looping at the flowering locus t promoter and regulates the timing of flowering in Arabidopsis. Plant Cell. 2014; 26( 3): 1009– 17. doi:10.1105/tpc.113.120352. [Google Scholar] [CrossRef]
Cite This Article
 Copyright © 2026 The Author(s). Published by Tech Science Press.
Copyright © 2026 The Author(s). Published by Tech Science Press.This work is licensed under a Creative Commons Attribution 4.0 International License , which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.


 Submit a Paper
Submit a Paper Propose a Special lssue
Propose a Special lssue View Full Text
View Full Text Download PDF
Download PDF Downloads
Downloads
 Citation Tools
Citation Tools