 Open Access
Open Access
ARTICLE
A Tritipyrum-Derived HVA22 Homolog Enhances Wheat Salt Tolerance
1 College of Agriculture, Guizhou University, Guiyang, China
2 Guizhou Subcenter of National Wheat Improvement Center, Guiyang, China
* Corresponding Authors: Suqin Zhang. Email: ; Guangdong Geng. Email:
# These authors contributed equally to this work.
(This article belongs to the Special Issue: Plant Responses to Abiotic Stress)
Phyton-International Journal of Experimental Botany 2026, 95(3), 8 https://doi.org/10.32604/phyton.2026.075984
Received 12 November 2025; Accepted 04 February 2026; Issue published 31 March 2026
Abstract
HVA22 is a gene induced by abscisic acid (ABA) and abiotic stress. Previous transcriptome data of salt-tolerant Tritipyrum “Y1805” revealed that HVA22 was significantly upregulated under salt stress. Gene TtHVA22 was successfully amplified from “Y1805”, with an open reading frame of 468 bp and encoding a protein of 156 amino acids. Gene TtHVA22 was transformed into bread wheat “1718” via coleoptile method. The relative expression level of TtHVA22 in roots was remarkably higher than in stems and leaves under salt stress. During the seedling stage, the TtHVA22 overexpression (OE) line exhibited less leaf wilting under salt stress than wild-type (WT) plants. Under salt stress and recovery conditions, TtHVA22 OE significantly increased root length, plant height, fresh weight, and dry weight compared to WT plants. Additionally, the levels of ABA, soluble sugars, soluble proteins, proline, pyruvate, and photosynthetic pigments and peroxidase activity were significantly higher in the OE lines than in the WT plants; however, their malondialdehyde content and relative conductivity were opposite. Two years of field data demonstrated that stem diameter and grain yield per plant were significantly greater in the OE lines than the WT plants. Therefore, wheat salt tolerance was improved in the TtHVA22 OE lines by osmotic regulation, antioxidation, and chlorophyll stabilization.Keywords
Supplementary Material
Supplementary Material FileMore than 1.38 billion hectares of land worldwide are impacted by salinization, accounting for 10.7% of the total global land area [1]. The total area of saline soil in China is 36 million hectares, approximately 4.88% of the total land area [2]. Soil salinization is one significant abiotic stress inhibiting global agricultural production, seriously affecting crop growth and yield. Common wheat (Triticum aestivum L.) is one of the most important food crops in the world [3], and understanding the salt-tolerance function of wheat and breeding salt-tolerant wheat cultivars are key measures to use salinized land and ensure food security.
Under adverse conditions such as salinization [4], drought [5], cold [6], and high temperature [7], the content of the stress hormone abscisic acid (ABA) in plants rises rapidly, thereby enhancing plant stress resistance [8]. Abscisic acid plays an important role in the response to salt stress, promoting stomatal closure [9], regulating ion absorption, transportation, and distribution in plants [10], inducing the synthesis of stress proteins such as late embryogenesis abundant (LEA) proteins [11] under adverse conditions, and eliminating excessive reactive oxygen species (ROS) [12]. The gene HVA22 (Hordeum vulgare ABA-responsive gene 22), initially discovered in barley (Hordeum vulgare L.), belongs to the LEA family and is induced by ABA and adverse stress [13]. Its promoter region contains ABA response elements and other stress response elements [14], and its expression is strongly induced by adverse stress, which is significantly upregulated under abiotic stresses (including high salt, drought, and low temperature) [15,16]. The overexpression (OE) of HVA22 in barley significantly improves plant drought resistance, and the mechanism is related to ABA-induced stomatal closure and water transpiration reduction [16]. In barley, the expression level of HVA22 increases under salt stress, drought, and cold, and the mRNA level of HVA22 in dormant barley grains remains high after water absorption, but drops to an undetectable level after 12 h of water absorption in non-dormant grains [17]. In rice (Oryza sativa L.), OsHLP1 (OsHVA22-Like Protein 1) interplays with OsATG8b and recruits the immune negative regulatory protein OsNTL6 into the autophagosome for degradation, thereby activating rice resistance to rice blast [18].
Research on the stress resistance related to HVA22 genes has mainly focused on Arabidopsis thaliana [19], barley [16], rice [18], cotton (Gossypium hirsutum L.) [20], tomato (Solanum lycopersicum L.) [21], and oilseed rape (Brassica napus L.) [22]. However, there are few reports concerning TtHVA22 salt tolerance in Tritipyrum. This study investigated the characteristics and salt-tolerant function of TtHVA22 using bioinformatics analysis, phenotypic identification, and physiological and biochemical measurements of OE lines, in order to breed salt-tolerant wheat.
The plant materials used were salt-tolerant octoploid (AABBDDEE) Tritipyrum “Y1805” and salt-sensitive common wheat “Chinese Spring” and “1718”. The “Y1805” is a pure progeny from a wide cross of common wheat (Triticum aestivum L.) and Thinopyrum elongatum, which not only includes the A, B, and D chromosomes from the wheat parent, but also a set of E group chromosomes originated from Th. elongatum. Tritipyrum “Y1805” has advantageous traits such as salt tolerance, early maturity, and resistance to wheat powdery mildew and rusts. Gene TtHVA22 was transformed into the salt-sensitive common wheat “1718” using the coleoptile method [23], and two out of 10 pure T3 OE lines were randomly chosen for this experiment. The seeds of these plant materials were preserved in Guizhou Subcenter of National Wheat Improvement Center. This study complies with relevant institutional, national, and international guidelines and regulations.
2.2 Plant Growth Conditions and Salt-Stress Treatment
Seeds of two OE lines and the wild-type (WT) “1718” were disinfected by 75% alcohol for 1 min, and then rinsed with water. They were placed in Petri dishes containing two layers of moist filter paper until seeds began germinating. The seedlings were cultured with 1/2 Hoagland solution under a light cycle of 16/8 h and temperature of 25/16°C (day/night). 250 mM NaCl was screened according to preliminary gradient tests, and salt-stress treatment (1/2 Hoagland solution with 250 mM NaCl) began at the two-leaf stage (about 14 d after germination). At 0 h (CK, control), 24 h of salt stress (T1), and 1 h after recovery (T2), the roots, stems, and leaves of 10 uniform seedlings were sampled, immediately frozen in liquid nitrogen, and stored at −80°C for gene cloning, real-time fluorescence quantitative PCR (qPCR), and physiological and biochemical analysis. The experiment included three biological replicates.
2.3 Screening and Cloning of TtHVA22
Total RNA of Tritipyrum “Y1805” and wheat “Chinese Spring” was extracted with an Plant RNA Kit (Omega, Guangzhou, China). After treatment with RNase-free DNase I (Takara, Dalian, China), the concentration and quality of total RNA were determined. The cDNA library construction and RNA-sequencing were carried out on the BGISEQ-500 platform at the Beijing Genomics Institute (BGI, ShenZhen, China). An average of 10.54 Gb of clean data per sample was generated, with three biological replicates in the experiment. Quality control and read trimming were conducted using SOAPnuke (BGI) and Trimmomatic, respectively. The clean reads were mapped to the reference genomes (T. aestivum, AABBDD and Th. elongatum, EE) by using HISAT2 (v2.1.0) software. RSEM (v1.2.8) was used to quantify gene expression levels. The differentially expressed gene was screened according to a false discovery rate threshold < 0.01, p-value < 0.01, and absolute log2fold change (FC) value > 1 between salt-treated and control samples using DESeq software [24]. The raw sequencing data have been deposited in the NCBI SRA under accession number PRJNA769794 (https://www.ncbi.nlm.nih.gov/sra/?term=PRJNA769794).
The specific primers according to the coding sequence of Tel5E01G132800 reference gene were designed (Table S1). Then, the target gene was amplified by PCR using “Y1805” cDNA as the template. The target fragment was recovered and purified using the DNA extraction and purification kit (Omega), and then ligated in the pEGOEPubi-H OE vector, which had been modified to have a green fluorescent protein gene for rapid transgenic detection. The inserted sequence was driven by a cauliflower mosaic virus (CaMV) 35S promoter.
2.4 TtHVA22 Bioinformatics Analysis
The amino acid sequences of HVA22 homologous proteins in the NCBI database were compared using Snapgene software. Open reading frames of the TtHVA22 protein were predicted using the online software ORF Finder (https://www.ncbi.nlm.nih.gov/gorf/gorf.html). Its physicochemical properties were analyzed using Expasy-PortParam (https://web.expasy.org/protparam/), subcellular localization determined through Wolf Psort (https://wolfpsort.hgc.jp/), signal peptide analyzed using SignalP 4.1 (https://services.healthtech.dtu.dk/services/SignalP-4.1/), hydrophilicity/hydrophobicity predicted using Expasy-ProtScale (https://web.expasy.org/protscale/), phosphorylation sites analyzed using NetPhos 3.1 (https://services.healthtech.dtu.dk/services/NetPhos-3.1/), and transmembrane domains analyzed using THHMM 2.0 (https://services.healthtech.dtu.dk/services/TMHMM-2.0/). Its protein domains were analyzed using Conserved Domains in the NCBI database (https://www.ncbi.nlm.nih.gov/Structure/cdd/wrpsb.cgi), secondary structure predicted using Sompa (https://npsa.lyon.inserm.fr/cgi-bin/npsa_automat.pl?page=/NPSA/npsa_sopma.html), tertiary structure predicted using Swiss Model (https://swissmodel.expasy.org/), homologous sequences retrieved in the NCBI database using Blast, and a phylogenetic tree constructed using MEGA-X software (with Bootstrap tests set at 1000).
2.5 TtHVA22 Genetic Transformation
Gene TtHVA22 was transformed into the salt-sensitive common wheat “1718” using the coleoptile method [23]. In brief, the OE vector harboring the TtHVA22 gene was introduced into Agrobacterium strain EHA105 (Takara). The meristem of coleoptile tips was cut and soaked in Agrobacterium inoculum under 15 KPa vacuum for 5 min infection. Then, these coleoptiles were kept humidity by covering a plastic bag and put in the dark for 3 days before peat culture. Positive transgenic plants were preliminarily screened by PCR, and then single-copy transgenic plants were detected by qPCR for the subsequent experiments.
2.6 Determination of TtHVA22 Copy Number by qPCR
The genomic DNAs from WT plants and plasmid DNA harboring the TtHVA22 gene were independently extracted, and the Pinb-D1 (single-copy) and TtHVA22 genes were amplified using the specific primers (Table S1). The reaction mixture was prepared according to the instructions of a 2× RealStar Fast SYBR qPCR Mix kit (Cas No. A301-10, GenStar, Beijing, China) (Table S2). The PCR program was as follows: 95°C for 30 s followed by 40 cycles of 95°C for 5 s, 60°C for 30 s, and 72°C for 15 s. qPCR standard curves for the Pinb-D1 and TtHVA22 genes were generated as reference [25,26]. Subsequently, the OE lines were detected for TtHVA22 copy number using the above methods. Each sample included three biological replications and three technical replications.
2.7 Analysis of TtHVA22 Expression Pattern
We ground the root, stem, and leaf samples of the TtHVA22 OE line and WT plants into powder in liquid nitrogen. Their total RNA was extracted with a Plant RNA Kit and then reverse transcribed into cDNA. The specific primers spanning the 35S–CDS junction were designed (Tables S1, S3 and S4) to detect TtHVA22 relative expression level under salt stress. The reaction mixture for qPCR was prepared as described in Section 2.6. The PCR program was as follows: 95°C for 2 min followed by 40 cycles of 95°C for 15 s, 60°C for 15 s, and 72°C for 15 s. Housekeeping 18S rRNA and β-actin were used as internal reference genes [27,28,29], and gene relative expression level was calculated by 2−ΔΔCT method [30].
2.8 Phenotypic and Physiological-Biochemical Analysis of TtHVA22
The samples of WT plants and the OE lines were collected during the CK, T1, and T2 stages. Root length, plant height, and plant fresh/dry weight of the seedlings were determined.
The contents of soluble sugar (via anthrone colorimetric reaction; Cas No. AKPL008M, Boxbio, Beijing, China), soluble protein (via Coomassie Brilliant Blue staining; Cas No. AKPR015, Boxbio), proline (by acidic ninhydrin colorimetric method; Cas No. AKAM003M, Boxbio), pyruvic acid (by 2,4-dinitrophenylhydrazine colorimetric method; Cas No. AKAC002M, Boxbio), and malondialdehyde (via thiobarbituric acid colorimetric assay; Cas No. AKFA013M, Boxbio) in the leaves, as well as peroxidase activity (via guaiacol method; Cas No. AKAO005M, Boxbio) were determined according to the appropriate kit’s instructions, and one unit (U) of enzyme activity was defined as a change in absorbance by 0.005 at 470 nm min−1 g−1 sample fresh weight at 25°C. The ABA content in leaves was measured using an ELISA kit (Cas No. SY-01051P1, Sinceyanbiomart, Yancheng, China). The contents of photosynthetic pigments in leaves were determined by ethanol extraction colorimetric method [31]. Root relative conductivity was determined according to Song et al. [32].
Data analysis and graph drawing were performed using GraphPad Prism 10.1.2 (GraphPad Software, San Diego, CA, USA). A two-way analysis of variance was employed after evaluating homogeneity of variances (by Brown-Forsythe test, p > 0.05) and normality of residuals [by Quartile-Quartile (Q-Q) plot test], followed by Tukey’s honestly significant difference (HSD) post-hoc tests.
3.1 Screening and Cloning of TtHVA22
Previous transcriptome analysis of Tritipyrum “Y1805” demonstrated that gene Tel5E01G132800 from Th. elongatum was significantly upregulated under salt stress (log2FC = 7.33). However, its expression level rapidly decreased after recovery (log2FC = 3.08) (Table S5). In contrast, Tel5E01G132800 showed no expression under salt stress and recovery conditions in salt-sensitive wheat “Chinese Spring.” Blast alignment results indicated that the amino acid sequence encoded by this gene contained the TB2-DP1 domain. Thus, it might belong to the HVA22 protein family. Based on the Tel5E01G132800 reference gene, specific primers were designed (Table S1). A cDNA fragment of 468 bp was successfully amplified from “Y1805” (Fig. 1A). This gene shared 97.05% homology with Tel5E01G132800 and was named TtHVA22 (Fig. 1B).
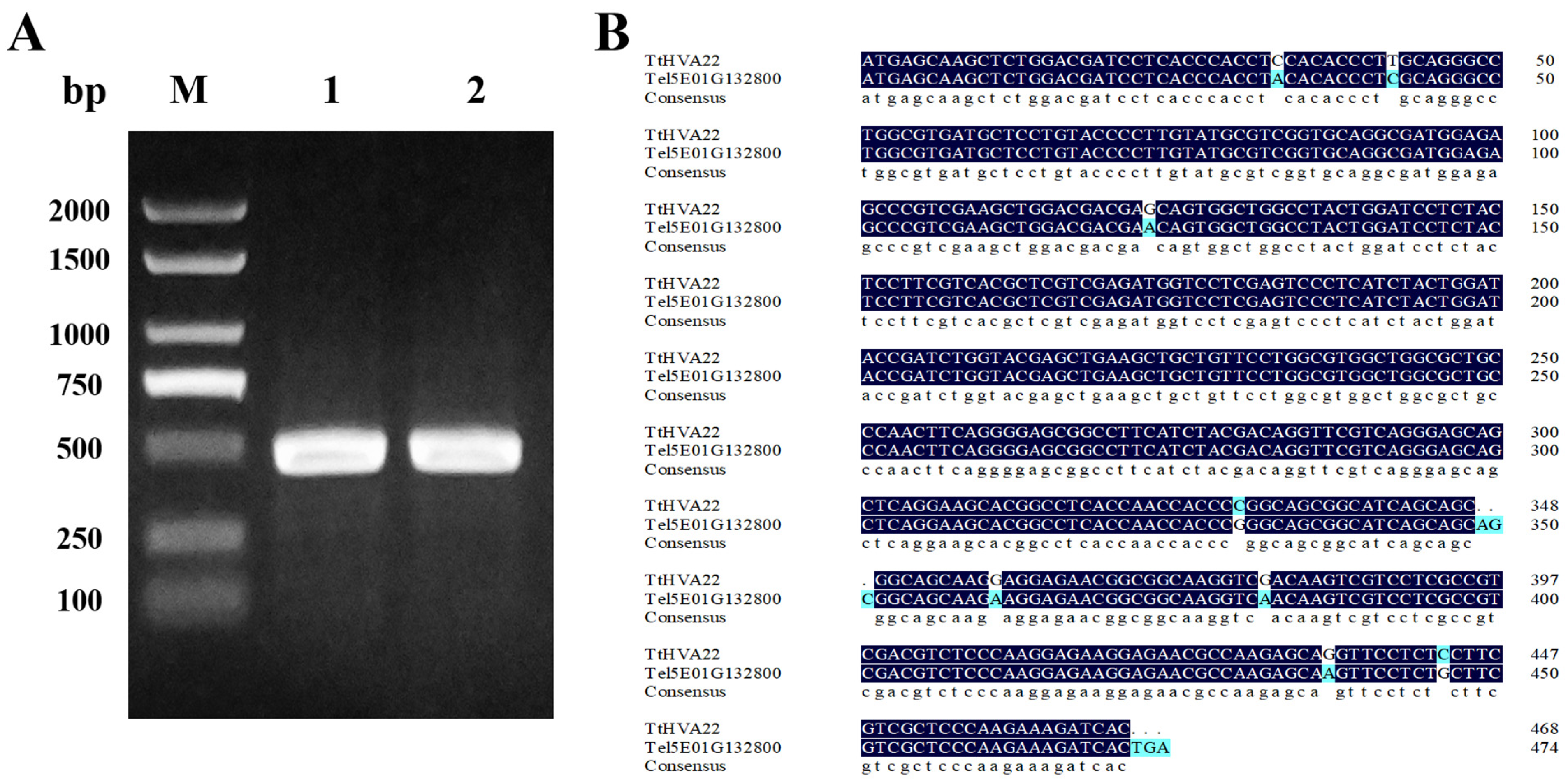
Figure 1: PCR product of TtHVA22, and the cDNA sequence comparison between genes TtHVA22 and Tel5E01G132800. (A) Lanes 1 and 2: target band of TtHVA22. M: D2000 marker. (B) Blue color indicates different bases.
3.2 Bioinformatics Analysis of TtHVA22 Protein
Bioinformatics analysis revealed that the TtHVA22 protein contained 156 amino acids, with a molecular formula of C827H1264N206O224S4, a relative molecular mass of 17.80 kDa, a theoretical isoelectric point of 8.76, an instability index of 34.91, and an aliphatic index of 93.78. The TtHVA22 was a dual-transmembrane protein, and had no signal peptide (Fig. 2A). The signal peptide sequence generally refers to the amino acid sequence at the N-terminal for transmembrane transfer. This sequence is usually present in secretory proteins, suggesting that TtHVA22 was a non-secretory protein. The average hydrophilicity index (GRAVY) value of TtHVA22 was −0.12 (Fig. 2B), suggesting that it was a hydrophilic protein. The phosphorylation site prediction showed that the serine (13), threonine (1), and tyrosine sites (1) had scores above 0.5 (Fig. 2C), suggesting that TtHVA22 contains predicted phosphorylation sites, primarily at serine residues, which may be involved in salt-stress signaling. The protein contained two transmembrane domains, located at amino acids 7–29 and 44–66 (Fig. 2D). The conserved domain (TB2-DP1-HVA22) of TtHVA22 was located at amino acids 24–98 (Fig. 2E). This protein had three types of secondary structures, including α-helices, random coils, and extended chains (Fig. 2F). Homology modeling showed that the similarity between the model and A0A1E5W487.1.A (an HVA22-like protein) was 84.62%, which met model standard (Fig. 2G). The phylogenetic tree of HVA22 protein sequences from diverse species clustered TtHVA22 with common wheat and Th. elongatum Tel5E01G132800 into one category (Fig. 2H). Therefore, TtHVA22 was evolutionarily and functionally similar to common wheat and Th. elongatum.
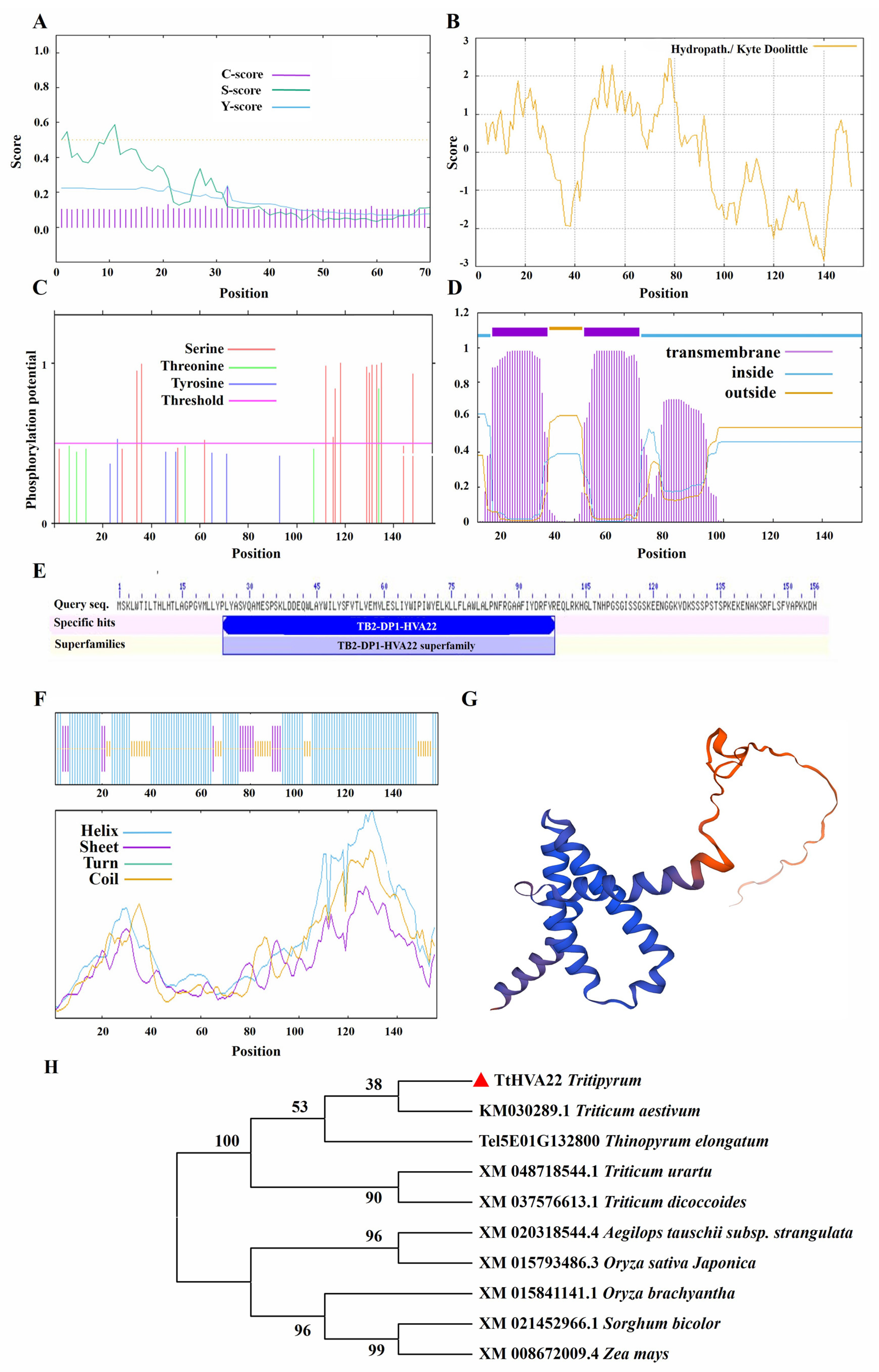
Figure 2: Bioinformatics analysis of TtHVA22 protein. (A) Signal peptide. (B) Hydrophilicity/hydrophobicity. (C) Phosphorylation site. (D) Transmembrane domain. (E) Protein domain. (F) Secondary structure. (G) Tertiary structure. (H) Phylogenetic tree.
3.3 Detection of TtHVA22 Copy Number in the OE Lines
Using the linear relationship between the Ct values and their corresponding logarithmic values of different initial template concentrations, we obtained the standard curves of the two genes (Fig. 3). They were y = −0.3527x + 23.638 (R2 = 0.9963) for Pinb-D1 and y = −0.3433x + 16.785 (R2 = 0.9933) for TtHVA22. The correlation coefficients of the two standard curves were almost 1, showing that they could be adopted to detect gene copy numbers of the OE lines. The two OE lines were single-copy (Table S6), which were used for the subsequent experiments.
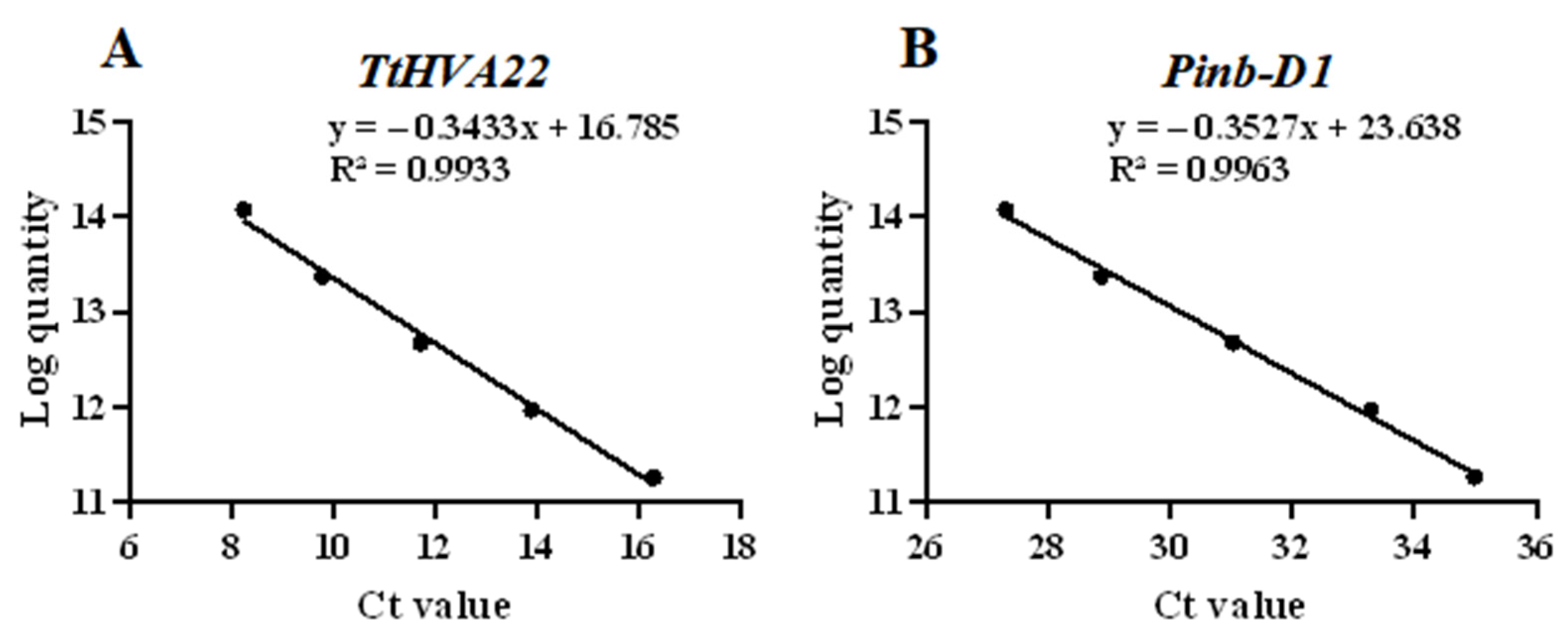
Figure 3: Standard curves of the TtHVA22 (A) and Pinb-D1 (B) genes. The logarithm of the initial template copy number are displayed on the y-axis and the Ct value on the x-axis.
3.4 Expression Pattern of TtHVA22
The response of TtHVA22 to salt stress was detected by qPCR. Under salt stress, the expression level of TtHVA22 in roots was the highest, followed by stems and leaves (Fig. 4A–D). Expression of TtHVA22 in roots of the two OE lines were 2.63- and 2.95-fold those of the WT plants (Fig. 4A). Expression of TtHVA22 in roots of OE line was the highest at 24 h of salt stress, and rapidly reduced in the recovery period. Thus, roots of the TtHVA22 OE lines were highly sensitive to salt stress and likely allowed for rapid adaptation to salt stress.
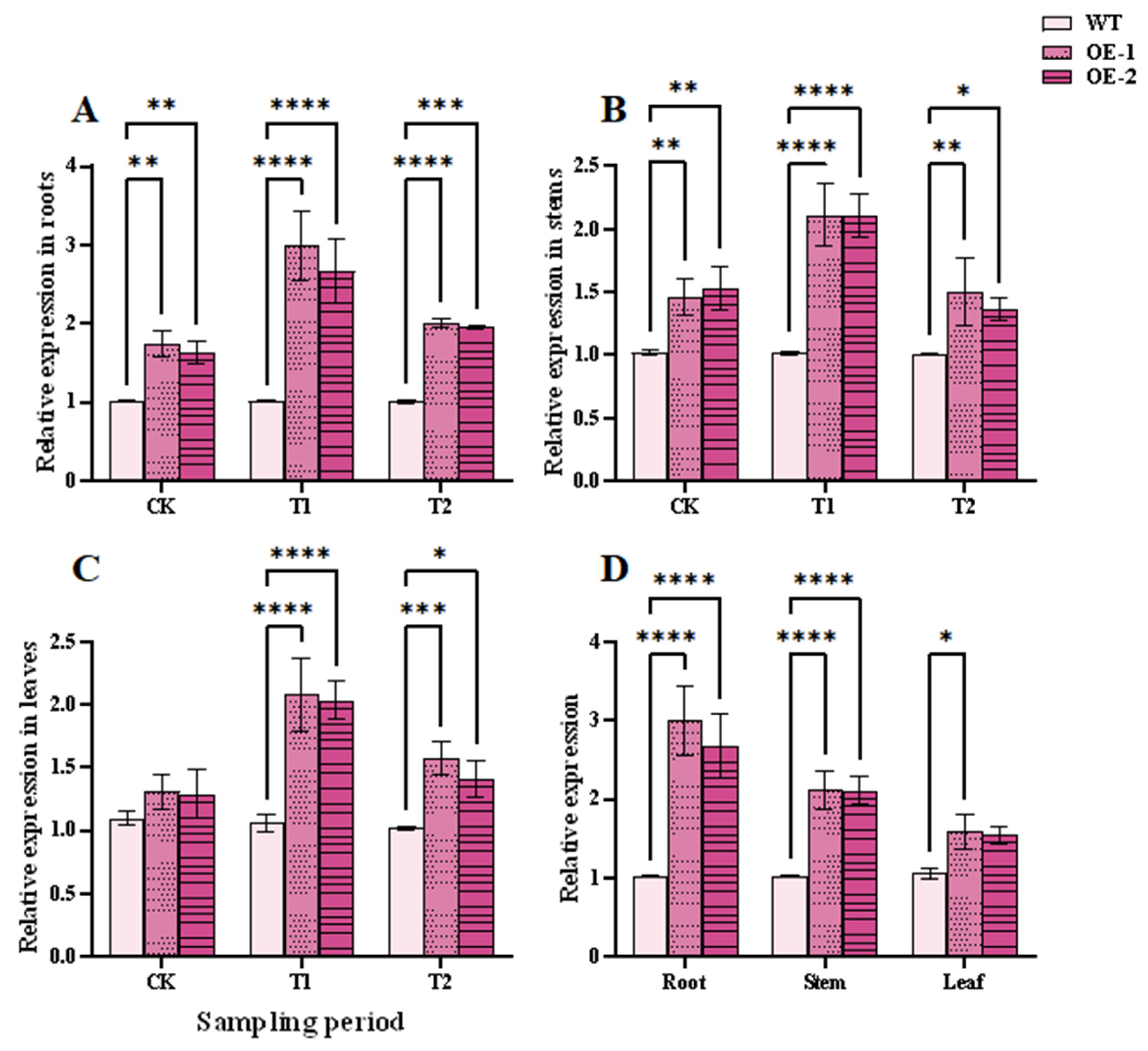
Figure 4: The qPCR determination of TtHVA22 expression levels in wild-type and overexpression plants. (A) TtHVA22 relative expression levels in roots. (B) TtHVA22 relative expression levels in stems. (C) TtHVA22 relative expression levels in leaves. (D) TtHVA22 relative expression levels in roots, stems, and leaves after 24 h of salt stress. CK, T1 and T2 represent the control, 24 h after salt stress, and 1 h after recovery, respectively. Data are presented as means ± SDs (n = 3). The significance between the WT plants and OE line was analyzed: *represents p < 0.05; **represents p < 0.01; ***represents p < 0.001; and ****represents p < 0.0001.
3.5 Phenotype of TtHVA22 OE Lines under Salt Stress
After 5 h and 24 h of salt stresses, some leaves of WT plants showed wilting and lodging, while those of the TtHVA22 OE lines remained more straighter, with no obvious changes (Fig. 5A–E). Under 24 h of salt stress (T1) and 1 h of recovery (T2) conditions, root length, plant height, and fresh/dry weight were significantly greater for the two OE lines than WT plants (Fig. 6). This indicated a stronger inhibitory effect of salt stress on growth of WT plants compared to OE lines. As a result, plant salt tolerance could be enhanced in the TtHVA22 OE lines.
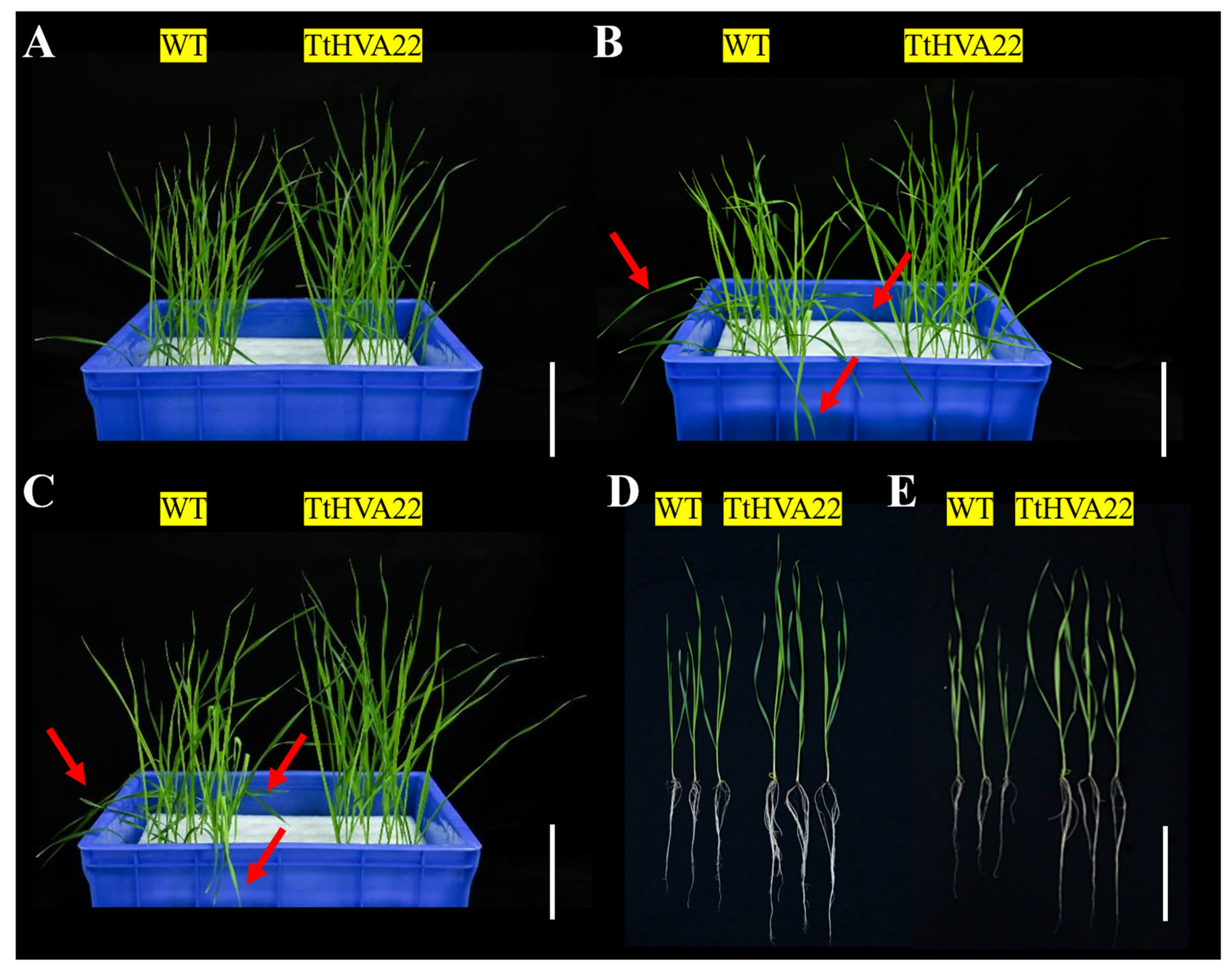
Figure 5: Effect of salt stress on wheat phenotype at the seedling stage. (A,D) Wild-type plants and TtHVA22 overexpression line under normal condition. (B) Wild-type plants and TtHVA22 overexpression line after 5 h of salt stress. (C,E) Wild-type plants and TtHVA22 overexpression plants after 24 h of salt stress. The red arrows indicate severe wilting of leaves. Scale bar = 5 cm.
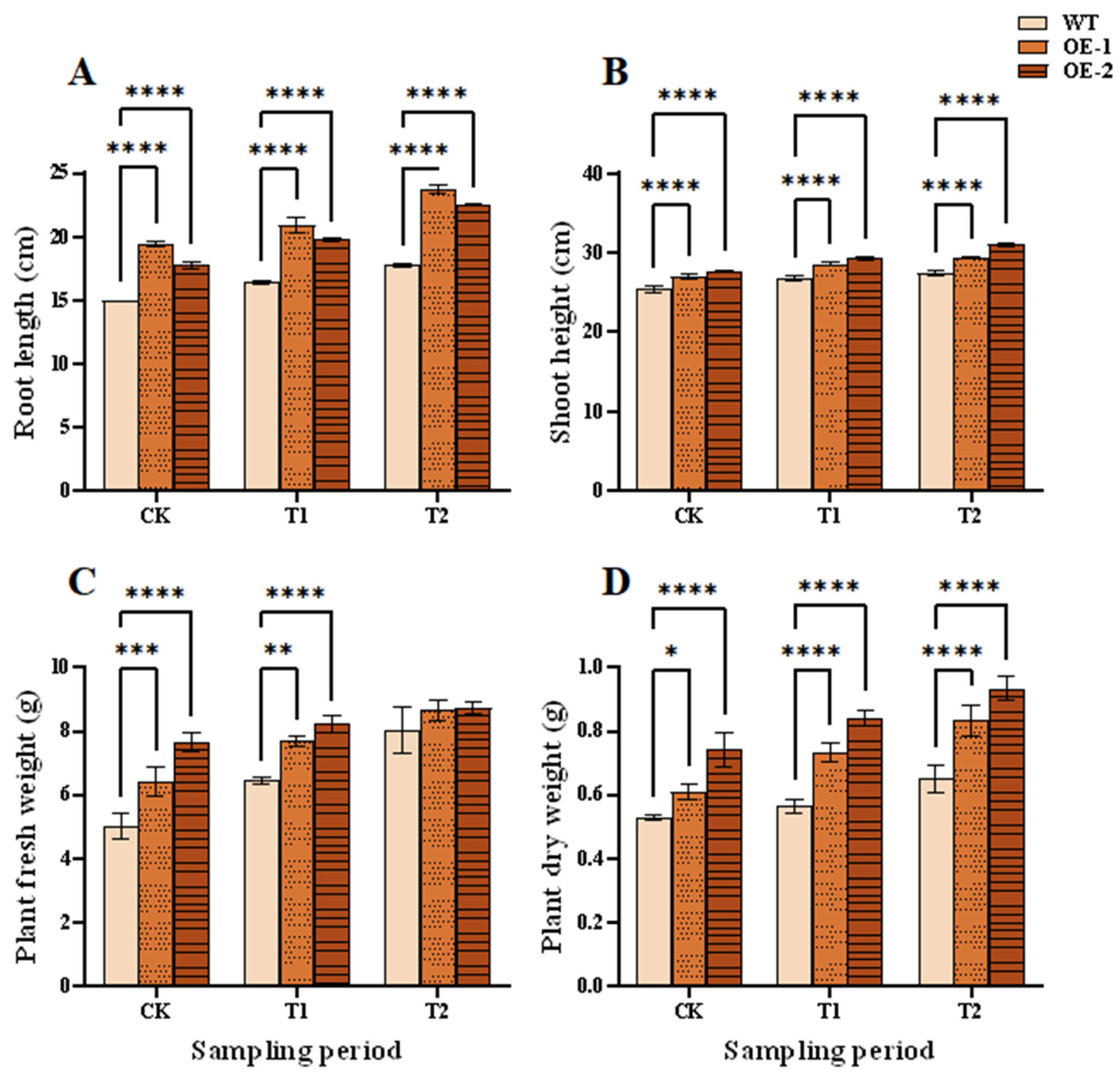
Figure 6: Effects of salt stress and recovery on growth indicators of wheat seedlings. (A) Root length. (B) Seedling height. (C) Plant fresh weight. (D) Plant dry weight. CK, T1 and T2 represent the control, 24 h after salt stress, and 1 h after recovery, respectively. Data are presented as means ± SDs (n = 3). The significance between the WT plants and OE lines was analyzed: *represents p < 0.05; **represents p < 0.01; ***represents p < 0.001; and ****represents p < 0.0001.
3.6 ABA and Osmolyte Levels in TtHVA22 OE Lines
Under adverse conditions, the ABA content in plants increases rapidly, as does their stress resistance [33]. In the normal (CK), T1, and T2 periods, the ABA content was markedly higher in the two OE lines than in WT plants (Fig. 7A). Thus, wheat ABA content increased and salt tolerance could be improved in the OE lines.
Osmolytes such as soluble sugar [34], soluble protein [35], and proline [36] can help plants cope with osmotic stress and ion toxicity caused by high salt environments, thereby maintaining normal growth and metabolism. Under normal, salt stress, and recovery conditions, the contents of soluble sugars, soluble proteins, and proline were markedly higher in the two OE lines than in WT plants (Fig. 7B–D). At the T1 stage, soluble sugar content in the OE-1 and OE-2 lines was 33.95% and 53.77% higher than in the WT plants, respectively; correspondingly, soluble protein content was 29.70% and 40.00% higher and proline content was 24.40% and 67.72% higher, respectively. We concluded that the OE lines could rapidly regulate cell osmolyte contents, enhance water absorption and retention capacity, and thereby improve plant salt tolerance.
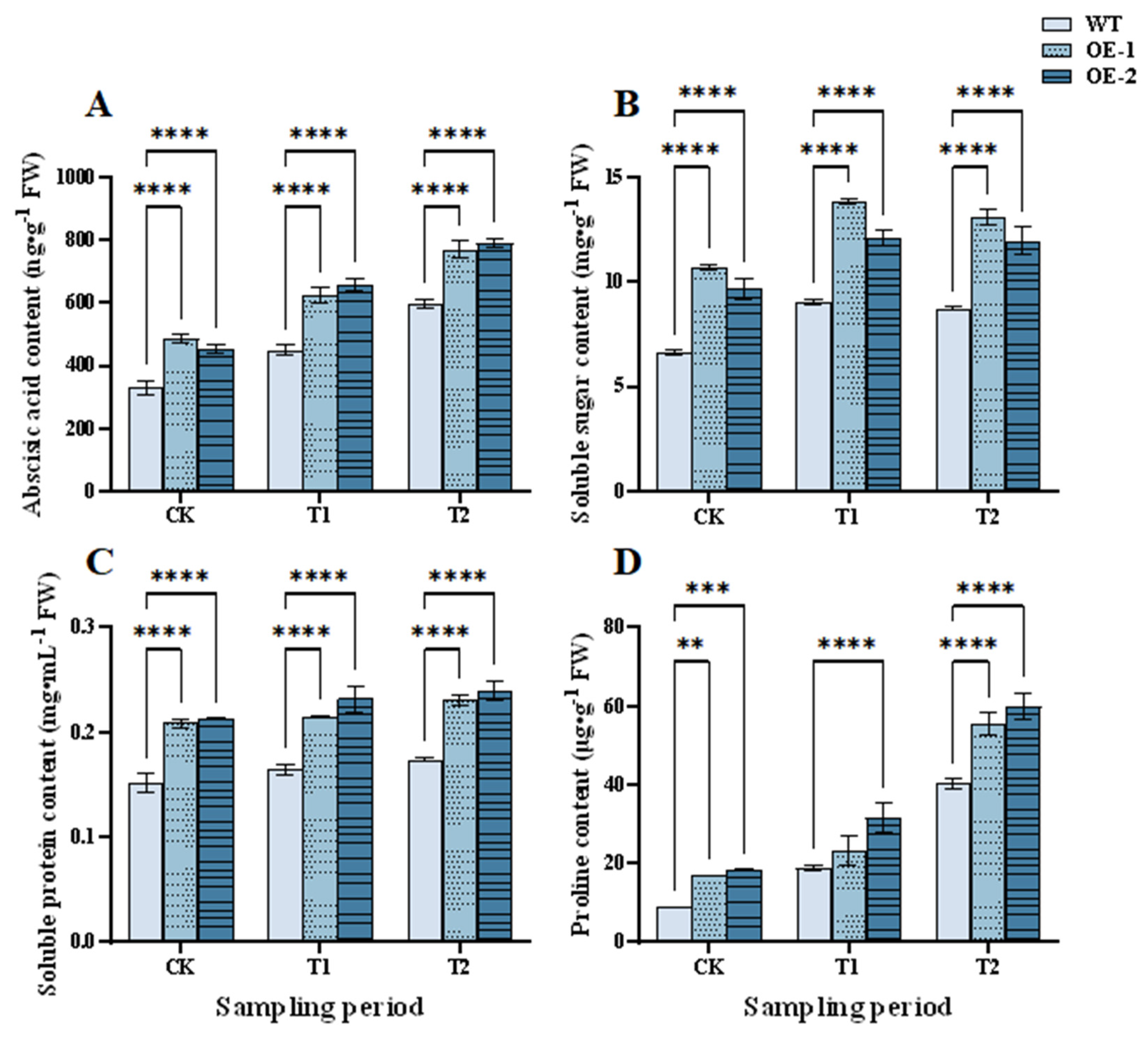
Figure 7: Effects of salt stress and recovery on contents of abscisic acid and osmolytes in the overexpression lines. (A) Abscisic acid. (B) Soluble sugars. (C) Soluble proteins. (D) Proline. CK, T1 and T2 represent the control, 24 h after salt stress, and 1 h after recovery, respectively. Data are presented as means ± SDs (n = 3). The significance between the WT plants and OE lines was analyzed: **represents p < 0.01; ***represents p < 0.001; and ****represents p < 0.0001.
3.7 Effect of TtHVA22 OE on Antioxidant Capacity
Peroxidase activity level can reflect the internal metabolic status of plants and can be an indicator of wheat adaptability and salt tolerance under salt stress [37]. At the T1 stage, peroxidase activity was significantly higher for the two OE lines than the WT plants, with the highest activity in OE-1, at 58.38% higher than WT plants. At the T2 stage, peroxidase activities of the OE lines continued to increase, and were 4922.67 and 4322.67 U/g FW higher than WT plants, respectively (Fig. 8A). Pyruvate content had a similar trend to peroxidase activity (Fig. 8B). Pyruvate participates in energy supply, material synthesis, and antioxidation [38]. The increase in peroxidase activity and pyruvate content in the OE lines could remove redundant ROS within cells, and enhance wheat salt tolerance under salt stress.
Accumulation of ROS causes membrane lipid peroxidation and permeability. This will enhance levels of malondialdehyde and relative conductivity [39,40]. In the CK, T1, and T2 periods, the levels of malondialdehyde and relative conductivity were significantly lower in the OE lines than the WT plants. The levels of malondialdehyde and relative conductivity in the OE lines and WT plants rose at the T1 stage, but fell at the T2 stage (Fig. 8C,D). This indicated a lower degree of membrane damage in OE lines than in WT plants under salt stress. This further revealed that membrane damage reduced and plant salt tolerance could be enhanced in the OE lines.
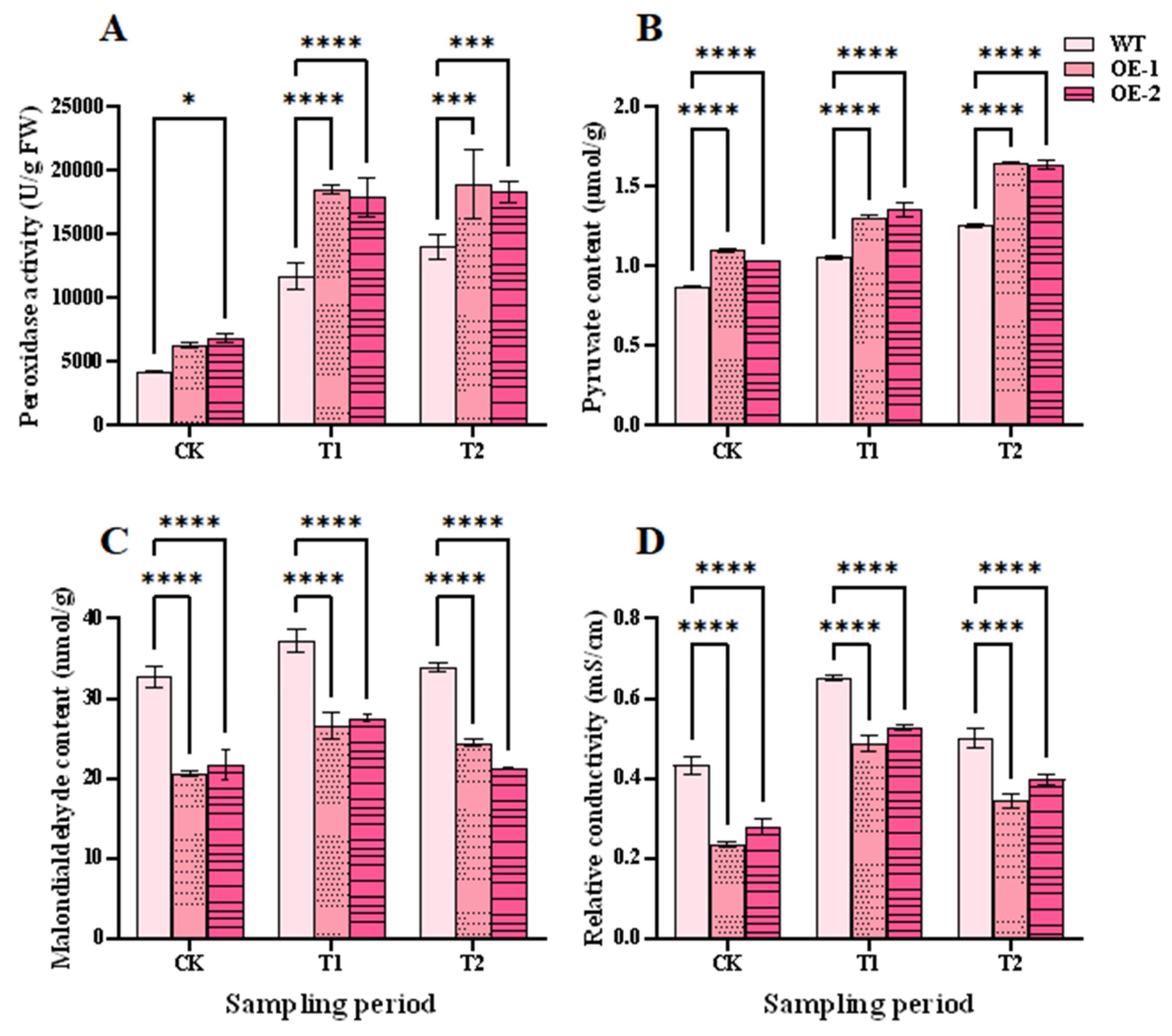
Figure 8: Effects of salt stress and recovery on antioxidant capacity in wheat. (A) Peroxidase activity. (B) Pyruvate content. (C) Malondialdehyde content. (D) Relative conductivity. CK, T1 and T2 represent the control, 24 h after salt stress, and 1 h after recovery, respectively. Data are presented as means ± SDs (n = 3). The significance between the WT plants and OE lines was analyzed: *represents p < 0.05; ***represents p < 0.001; and ****represents p < 0.0001.
3.8 TtHVA22 OE Effect on Photosynthetic Pigment Contents
Photosynthetic pigments can absorb, transfer, and convert light energy, and produce carbohydrate to maintain plant growth and metabolism [38]. Under normal conditions, contents of the three photosynthetic pigments of the OE lines were higher than in WT plants (Fig. 9). At the T1 and T2 stages, contents of chlorophyll a and b in both the OE lines and the WT plants decreased (Fig. 9A,B) but were significantly higher in OE lines than WT plants. Therefore, higher chlorophyll contents could be maintained, and more organic materials might be synthesized to maintain plant growth and metabolism in the OE lines under salt stress.
During the CK, T1, and T2 periods, the carotenoid content of both OE lines displayed a trend of initial increase and then decrease (Fig. 9C). Under salt-stress conditions, although the carotenoid content of all wheat plants increased, content of the OE-1 and OE-2 lines was 1.30 and 1.36 mg/g FW, respectively, remarkably higher than that of WT plants at only 1.07 mg/g FW. Therefore, photosynthetic pigments were more stable in the OE lines than WT plants. This contributed to plant photosynthesis and growth under salt stress.
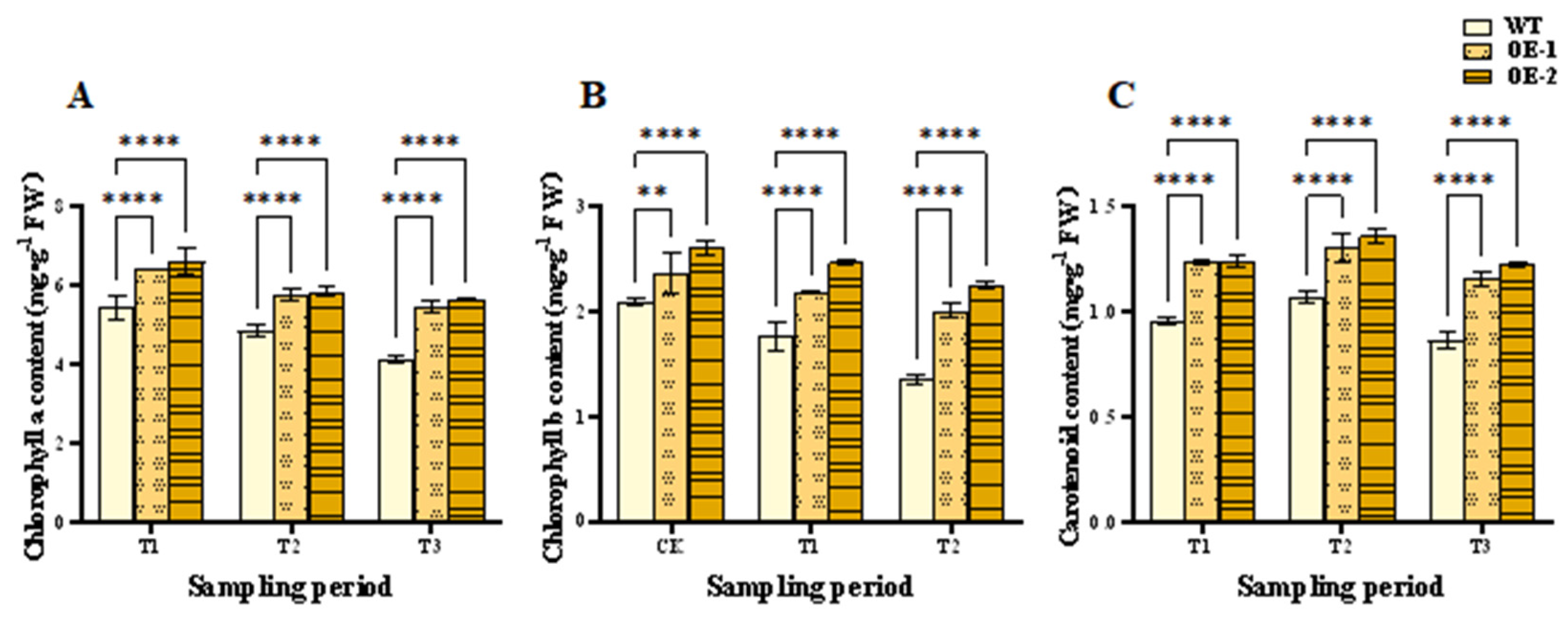
Figure 9: Effects of salt stress and recovery on wheat photosynthetic pigment contents. (A) Chlorophyll a content. (B) Chlorophyll b content. (C) Carotenoid content. CK, T1 and T2 represent the control, 24 h after salt stress, and 1 h after recovery, respectively. Data are presented as means ± SDs (n = 3). The significance between the WT plants and OE lines was analyzed: **represents p < 0.01; and ****represents p < 0.0001.
3.9 Correlations between Physiological and Biochemical Indicators
Proline content did not correlate with soluble sugar, malondialdehyde, and chlorophyll a contents, nor relative conductivity (Table S7). All the other parameters correlated significantly with each other. The levels of ABA, soluble proteins, pyruvate, chlorophyll b, carotenoid, and peroxidase activity were significantly positively correlated with each other. These results demonstrated that these substances might work together to form a defense network for protecting wheat under salt stress.
3.10 TtHVA22 OE Effects on Agronomic Traits
In 2024 and 2025, plant height was significantly lower for the OE lines than WT plants. Stem diameter, flag leaf length/width, tiller number, spike length, spikelet number, 1000-grain weight, and grain yield per plant were remarkably higher for the OE lines than WT plants in the two years (Fig. 10A-I). The results indicated better agronomic traits for the OE lines. The shorter plant height and thicker stems of the OE lines improved lodging resistance. The large flag leaves of the OE lines could promote photosynthetic capacity compared to the WT plants, thereby enhancing wheat grain growth.
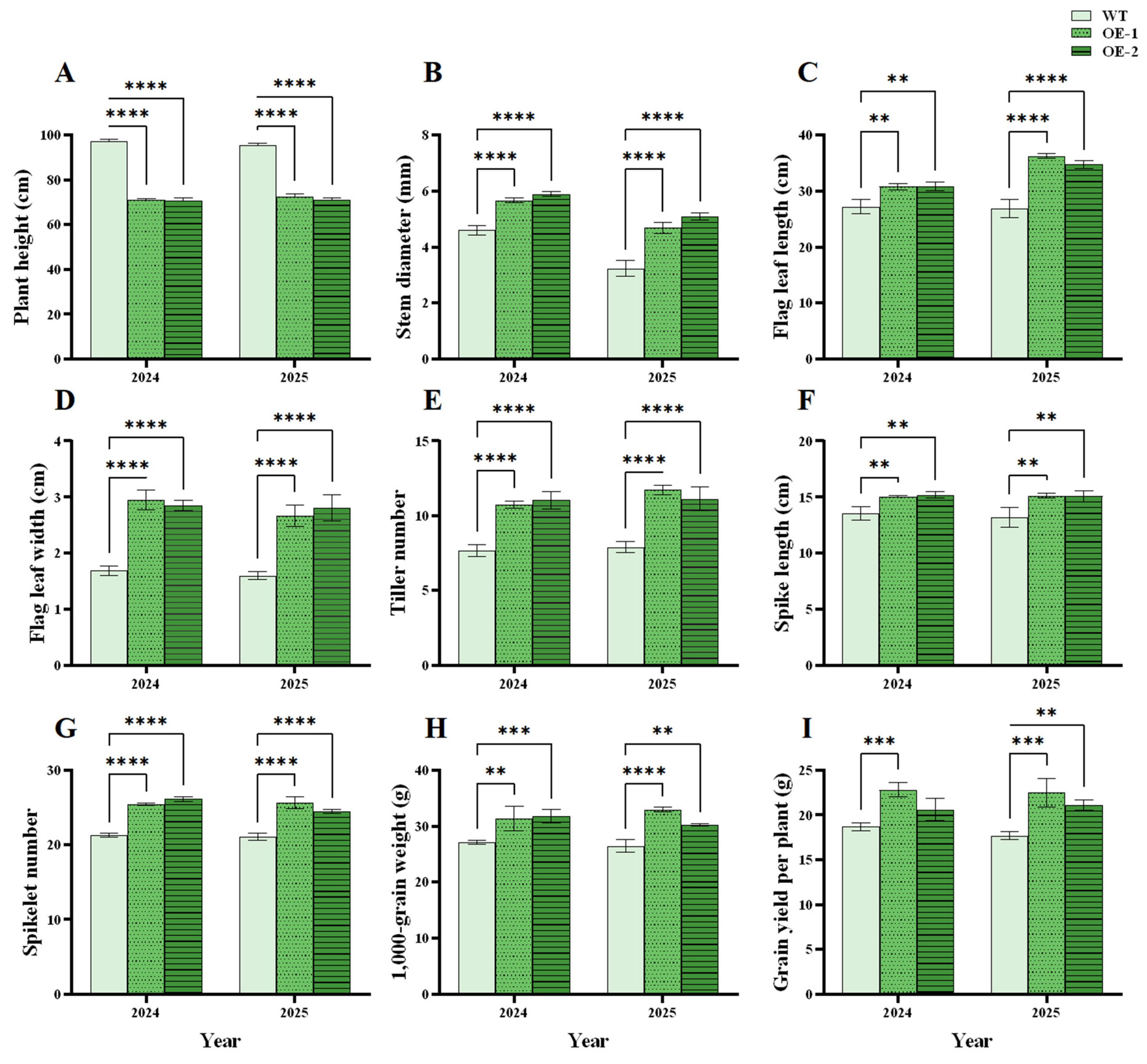
Figure 10: Effects of TtHVA22 overexpression on wheat agronomic traits during the reproductive growth period. (A) Plant height. (B) Stem diameter. (C) Flag leaf length. (D) Flag leaf width. (E) Tiller number. (F) Spike length. (G) Spikelet number. (H) The 1000-grain weight. (I) Grain yield per plant. Data are presented as means ± SDs (n = 3). The significance between the WT plants and OE lines was analyzed: **represents p < 0.01; ***represents p < 0.001; and **** represents p < 0.0001.
Grain yield per plant was significantly positively correlated with stem diameter, flag leaf area, tiller number, spike length, spikelet number, and 1000-grain weight (p < 0.05) in both 2024 and 2025, while plant height was negatively correlated with grain yield per plant (2024: r = −0.763, p = 0.0167; 2025: r = −0.856, p = 0.0032) (Table S8). Thus, these correlated traits might play key roles in enhancing grain yield.
4.1 TtHVA22 Cloning and Expression Pattern
Protein HVA22 and its homologs have been found in various organisms, including plants [18,21], animals [41,42], and fungi [22], indicating their significance in cellular physiological processes. In this study, TtHVA22 cloned from salt-tolerant Tritipyrum “Y1805” encoded a hydrophilic protein with 156 amino acids and contained transmembrane structures. The phylogenetic tree showed that TtHVA22 and homologs from common wheat and Th. elongatum were grouped in the same branch (Fig. 2H), suggesting that TtHVA22 was evolutionarily and functionally similar to common wheat and Th. elongatum.
In Arabidopsis thaliana, AtHVA22-E OE can enhance plant salt tolerance, while knockout of HVA22 makes plants more sensitive to adverse stress [19]. The GhHVA22E1D gene in cotton plays a positive role in plant response to salt and drought stresses; GhHVA22E1D OE can increase cotton salt and drought tolerance, while silencing GhHVA22E1D reduces plant salt and drought tolerance [20]. Similar results were obtained in our experiment. The TtHVA22 expression level in roots increased under salt stress (Fig. 4). Plant roots were directly and severely influenced by salt stress, which led to a remarkably higher expression of TtHVA22 in roots compared to WT plants. Therefore, TtHVA22 expression was highly upregulated and sensitive in roots under salt stress.
4.2 Wheat Salt Tolerance Was Enhanced in the OE Lines
Salt stress promotes ABA accumulation by activating the key genes (such as TaNCED) in the ABA synthesis pathway [43]. In a saline environment, ABA can alleviate plant damage by regulating ion transport, stomatal aperture, osmolyte synthesis, and the antioxidant system [44,45]. Here, ABA contents of OE-1 and OE-2 lines were 1.39- and 1.46-fold that of WT plants at the T1 stage, respectively (Fig. 7A). ABA content could increase quickly under salt stress, which contributed to enhancing plant salt tolerance in the OE lines.
Under salt stress, wheat accumulates osmolytes (including soluble sugars, proline, and soluble proteins) to reduce cellular osmotic potential, thereby promoting water uptake from the external environment and maintaining cell turgor pressure and normal physiological functions. In this experiment, the OE lines had 1.34- to 1.54-fold soluble sugar and 1.30- to 1.40-fold soluble protein levels compared to WT plants under salt stress (Fig. 7B,C), possibly enhancing their osmotic adjustment capacity, maintaining cellular water balance, and reducing cellular damage by salt stress. Additionally, proline can protect membrane systems and enzyme activities, scavenge ROS, and reduce plant malondialdehyde content [46]. Proline interacts with the ABA signaling pathway to modulate stress-related gene expression [47]. In this work, proline contents were significantly higher in the OE-1 and OE-2 lines than WT plants at the T1 stage (Fig. 7D). Overall, osmolyte levels increased, cell osmotic potential decreased, and cell water loss was alleviated in the OE lines under salt stress.
Under environmental stress, plants usually accumulate excessive ROS, which triggers lipid peroxidation reactions, generating toxic substances, and damaging the integrity and fluidity of membrane structures, leading to ion leakage and impaired cellular function [48,49]. The ROS can cause protein function loss or even degradation through amino acid residue modification [50]. Relative conductivity is positively correlated with cell membrane damage [39,41]. The oxidative damage caused by ROS is one core mechanism of salt-stress injury in wheat, and salt-tolerant wheat varieties usually effectively control ROS level by enhancing antioxidant enzyme activity or accumulating non-enzyme antioxidants to alleviate the harm. Tomato plants with silenced SIHVA22 had higher levels of hydrogen peroxide and malondialdehyde, and significantly reduced antioxidant enzyme activities after drought treatment, indicating that SIHVA22 plays an important role in drought resistance [21]. Here, the peroxidase activity was still higher in the OE-1 and OE-2 lines than WT plants at the T2 stage (Fig. 8A). The relative conductivity and malondialdehyde content were significantly higher for WT plants than the OE lines (Fig. 8C,D). The increase in peroxidase activity could remove harmful ROS in the OE lines, and enhance wheat salt tolerance under salt stress.
Under salt stress, chlorophyll is not only a core functional substance for photosynthesis, but also a sensitive indicator of stress damage [51]. Salt stress hinders photosynthesis by inducing osmotic stress and ion toxicity in crops, and significantly reduces chlorophyll content in wheat leaves [52,53]. Here, chlorophyll contents were remarkably lower in WT plants than OE lines under salt stress and recovery conditions (Fig. 9A,B). Carotenoid levels are positively correlated with salt tolerance, as carotenoids quench singlet oxygen and scavenge free radicals, thereby reducing photooxidative damage [54]. Additionally, carotenoids serve as precursors for ABA synthesis, and their accumulation may enhance salt tolerance through the ABA signaling pathway [55]. In this work, carotenoid contents were remarkably higher in the OE lines than WT plants at the T1 stage (Fig. 9C). The stability of photosynthetic pigment content and structure directly reflected stability of the photosynthetic apparatus, and improved salt tolerance depended on protecting photosynthetic pigments through multiple mechanisms such as ion balance, antioxidant defense, and hormone regulation [56]. The photosynthetic pigments were more stable in the OE lines than WT plants, which may contribute to plant photosynthesis and growth under salt stress.
The recovery period of 1 h was screened by the preliminary tests. However, the recovery period was too short to reflect physiological recovery. We will extend the recovery period (e.g., 24 h) in the future experiments.
Agronomic traits are closely related to wheat yield, quality, stress resistance, and adaptability [57,58]. Dwarf wheat plants are less likely to experience stem breakage or lodging [59]. The flag leaves, as the core of wheat “source” organ, directly determine the size and fullness of the “sink” (grains), maintain a certain photosynthetic capacity under adverse conditions, and reduce stress impact on grain development, which is a key link among photosynthesis, stress resistance, and yield formation [60,61]. In the spring of 2024 and of 2025, severe lodging occurred in the WT plants in the field, while the OE lines showed strong lodging resistance due to their short and thick stems. In addition, the length and width of flag leaves, spike length, tiller number, spikelet number, 1000-grain weight, and grain yield per plant were remarkably greater in OE lines than WT plants (Fig. 10). The results revealed that not only wheat salt tolerance but also growth and yield were improved in the OE lines.
Based on previous transcriptome data of Tritipyrum “Y1805”, the salt-tolerant gene TtHVA22 was screened and cloned. Growth and physiological indicators proved that salt-stress damage could be alleviated in the OE lines. Under salt stress, TtHVA22 was upregulated in roots. It could alleviate salt-stress damage by enhancing the levels of ABA and osmolytes and impacting the antioxidant capacity and photosynthetic pigment stability. Additionally, wheat lodging resistance and yield were improved in the OE lines. Therefore, Tritipyrum TtHVA22 was a precious gene for salt-tolerant wheat breeding.
Acknowledgement:
Funding Statement: This work was supported by the National Natural Science Foundation of China (32160442 and 32560458). Funding has no influence on study design, interpretation of data, writing, and publishing.
Author Contributions: Study conception and design: Tingting Yuan, Suqin Zhang; data collection: Mutong Li, Zhishun Yu, Jiangyun Huang; analysis and interpretation of results: Tingting Yuan; draft manuscript preparation: Tingting Yuan, Qingqin Zhang, Suqin Zhang, Guangdong Geng. All authors reviewed and approved the final version of the manuscript.
Availability of Data and Materials: All data generated or analyzed during this study are provided in this published article and its supplementary data files.
Ethics Approval: Not applicable.
Conflicts of Interest: The authors declare no conflicts of interest.
Supplementary Materials: The supplementary material is available online at https://www.techscience.com/doi/10.32604/phyton.2026.075984/s1. Table S1: The primers used in the experiment; Table S2: The reaction mixture for quantitative real-time PCR; Table S3: The primary endogenous genes in the recipient wheat showing high homology (>95%) to the TtHVA22 sequence; Table S4: TtHVA22 Primer-BLAST results; Table S5: The relative expression level (log2 fold change) of Tel5E01G132800 in two wheat materials; Table S6: TtHVA22 copy number in the two overexpression lines; Table S7: Correlations among wheat physiological and biochemical parameters under salt stress; Table S8: Correlation between grain yield per plant and other traits in the TtHVA22 overexpression lines in 2024 and 2025.
References
1. FAO (Food and Agriculture Organization of the United Nations). Global status of salt-affected soils. Rome, Italy: FAO; 2024. doi:10.4060/cd3044en. [Google Scholar] [CrossRef]
2. Zhu W , Gu S , Jiang R , Zhang X , Hatano R . Saline–alkali soil reclamation contributes to soil health improvement in China. Agriculture. 2024; 14( 8): 1210. doi:10.3390/agriculture14081210. [Google Scholar] [CrossRef]
3. Huang XQ , Wang LX , Xu MX , Röder MS . Microsatellite mapping of the powdery mildew resistance gene Pm5e in common wheat (Triticum aestivum L.). Theor Appl Genet. 2003; 106( 5): 858– 65. doi:10.1007/s00122-002-1146-3. [Google Scholar] [CrossRef]
4. Liu X , Xie X , Zheng C , Wei L , Li X , Jin Y , et al. RNAi-mediated suppression of the abscisic acid catabolism gene OsABA8ox1 increases abscisic acid content and tolerance to saline–alkaline stress in rice (Oryza sativa L.). Crop J. 2022; 10( 2): 354– 67. doi:10.1016/j.cj.2021.06.011. [Google Scholar] [CrossRef]
5. Muhammad Aslam M , Waseem M , Jakada BH , Okal EJ , Lei Z , Ahmad Saqib HS , et al. Mechanisms of abscisic acid-mediated drought stress responses in plants. Int J Mol Sci. 2022; 23( 3): 1084. doi:10.3390/ijms23031084. [Google Scholar] [CrossRef]
6. Liu X , Wei J , Li S , Li J , Cao H , Huang D , et al. MdHY5 positively regulates cold tolerance in apple by integrating the auxin and abscisic acid pathways. New Phytol. 2025; 246( 5): 2155– 73. doi:10.1111/nph.20333. [Google Scholar] [CrossRef]
7. Toh S , Imamura A , Watanabe A , Nakabayashi K , Okamoto M , Jikumaru Y , et al. High temperature-induced abscisic acid biosynthesis and its role in the inhibition of gibberellin action in Arabidopsis seeds. Plant Physiol. 2008; 146( 3): 1368– 85. doi:10.1104/pp.107.113738. [Google Scholar] [CrossRef]
8. Sah SK , Reddy KR , Li J . Abscisic acid and abiotic stress tolerance in crop plants. Front Plant Sci. 2016; 7: 571. doi:10.3389/fpls.2016.00571. [Google Scholar] [CrossRef]
9. Bharath P , Gahir S , Raghavendra AS . Abscisic acid-induced stomatal closure: an important component of plant defense against abiotic and biotic stress. Front Plant Sci. 2021; 12: 615114. doi:10.3389/fpls.2021.615114. [Google Scholar] [CrossRef]
10. Leung J , Valon C , Moreau B , Boeglin M , Lefoulon C , Joshi-Saha A , et al. The ABC of abscisic acid action in plant drought stress responses. Biol Aujourdhui. 2012; 206( 4): 301– 12. doi:10.1051/jbio/2012029. [Google Scholar] [CrossRef]
11. Lin R , Zou T , Mei Q , Wang Z , Zhang M , Jian S . Genome-wide analysis of the late embryogenesis abundant (LEA) and abscisic acid-, stress-, and ripening-induced (ASR) gene superfamily from Canavalia rosea and their roles in salinity/alkaline and drought tolerance. Int J Mol Sci. 2021; 22( 9): 4554. doi:10.3390/ijms22094554. [Google Scholar] [CrossRef]
12. Parveen A , Ahmar S , Kamran M , Malik Z , Ali A , Riaz M , et al. Abscisic acid signaling reduced transpiration flow, regulated Na+ ion homeostasis and antioxidant enzyme activities to induce salinity tolerance in wheat (Triticum aestivum L.) seedlings. Environ Technol Innov. 2021; 24: 101808. doi:10.1016/j.eti.2021.101808. [Google Scholar] [CrossRef]
13. Shen Q , Uknes SJ , Ho TH . Hormone response complex in a novel abscisic acid and cycloheximide-inducible barley gene. J Biol Chem. 1993; 268( 31): 23652– 60. doi:10.1016/S0021-9258(19)49512-4. [Google Scholar] [CrossRef]
14. Shen Q , Zhang P , Ho TH . Modular nature of abscisic acid (ABA) response complexes: composite promoter units that are necessary and sufficient for ABA induction of gene expression in barley. Plant Cell. 1996; 8( 7): 1107– 19. doi:10.2307/3870355. [Google Scholar] [CrossRef]
15. Brands A , Ho TD . Function of a plant stress-induced gene, HVA22. Synthetic enhancement screen with its yeast homolog reveals its role in vesicular traffic. Plant Physiol. 2002; 130( 3): 1121– 31. doi:10.1104/pp.007716. [Google Scholar] [CrossRef]
16. Straub PF , Shen Q , Ho TD . Structure and promoter analysis of an ABA- and stress-regulated barley gene, HVA1. Plant Mol Biol. 1994; 26( 2): 617– 30. doi:10.1007/BF00013748. [Google Scholar] [CrossRef]
17. Shen Q , Chen CN , Brands A , Pan SM , Ho TH . The stress- and abscisic acid-induced barley gene HVA22 developmental regulation and homologues in diverse organisms. Plant Mol Biol. 2001; 45( 3): 327– 40. doi:10.1023/a:1006460231978. [Google Scholar] [CrossRef]
18. Liang Y , Meng F , Zhao X , He X , Liu J . OsHLP1 is an endoplasmic-reticulum-phagy receptor in rice plants. Cell Rep. 2023; 42( 12): 113480. doi:10.1016/j.celrep.2023.113480. [Google Scholar] [CrossRef]
19. Guo WJ , Ho TH . An abscisic acid-induced protein, HVA22, inhibits gibberellin-mediated programmed cell death in cereal aleurone cells. Plant Physiol. 2008; 147( 4): 1710– 22. doi:10.1104/pp.108.120238. [Google Scholar] [CrossRef]
20. Zhang H , Yuan Y , Xing H , Xin M , Saeed M , Wu Q , et al. Genome-wide identification and expression analysis of the HVA22 gene family in cotton and functional analysis of GhHVA22E1D in drought and salt tolerance. Front Plant Sci. 2023; 14: 1139526. doi:10.3389/fpls.2023.1139526. [Google Scholar] [CrossRef]
21. Wai AH , Waseem M , Cho LH , Kim ST , Lee DJ , Kim CK , et al. Comprehensive genome-wide analysis and expression pattern profiling of the SlHVA22 gene family unravels their likely involvement in the abiotic stress adaptation of tomato. Int J Mol Sci. 2022; 23( 20): 12222. doi:10.3390/ijms232012222. [Google Scholar] [CrossRef]
22. Ye W , Hossain R , Pröbsting M , Ali AAM , Han L , Miao Y , et al. Knock-out of BnHva22c reduces the susceptibility of Brassica napus to infection with the fungal pathogen Verticillium longisporum. Crop J. 2024; 12( 2): 503– 14. doi:10.1016/j.cj.2024.02.012. [Google Scholar] [CrossRef]
23. Yang Z , Mu Y , Wang Y , He F , Shi L , Fang Z , et al. Characterization of a novel TtLEA2 gene from tritipyrum and its transformation in wheat to enhance salt tolerance. Front Plant Sci. 2022; 13: 830848. doi:10.3389/fpls.2022.830848. [Google Scholar] [CrossRef]
24. Peng Z , Wang Y , Geng G , Yang R , Yang Z , Yang C , et al. Comparative analysis of physiological, enzymatic, and transcriptomic responses revealed mechanisms of salt tolerance and recovery in tritipyrum. Front Plant Sci. 2022; 12: 800081. doi:10.3389/fpls.2021.800081. [Google Scholar] [CrossRef]
25. Li M , Yang X , Yu Z , Yuan T , Wang W , Zhang S , et al. Tritipyrum Aux/IAA13L increases chlorophyll content and yield in wheat. Phyton. 2025; 94( 10): 3175– 88. doi:10.32604/phyton.2025.070731. [Google Scholar] [CrossRef]
26. Weng H , Pan A , Yang L , Zhang C , Liu Z , Zhang D . Estimating number of transgene copies in transgenic rapeseed by real-time PCR assay withHMG I/Y as an endogenous reference gene. Plant Mol Biol Rep. 2004; 22( 3): 289– 300. doi:10.1007/BF02773139. [Google Scholar] [CrossRef]
27. Mu Y , Shi L , Tian H , Tian H , Zhang J , Zhao F , et al. Characterization and transformation of TtMYB1 transcription factor from Tritipyrum to improve salt tolerance in wheat. BMC Genom. 2024; 25( 1): 163. doi:10.1186/s12864-024-10051-5. [Google Scholar] [CrossRef]
28. Yang S , Yu Z , Yuan T , Li M , Zhang C , Zhang Q , et al. Aldose reductase has a function in salt tolerance of wheat. Gene. 2025; 944: 149295. doi:10.1016/j.gene.2025.149295. [Google Scholar] [CrossRef]
29. Zhang J , Yang S , Li M , Yuan T , Yu Z , Song D , et al. A novel phospholipase a(1) gene from tritipyrum improves wheat early ripening and salt tolerance. J Agric Food Chem. 2025; 73( 28): 17713– 29. doi:10.1021/acs.jafc.4c12792. [Google Scholar] [CrossRef]
30. Schmittgen TD , Livak KJ . Analyzing real-time PCR data by the comparative CT method. Nat Protoc. 2008; 3( 6): 1101– 8. doi:10.1038/nprot.2008.73. [Google Scholar] [CrossRef]
31. Porra RJ , Thompson WA , Kriedemann PE . Determination of accurate extinction coefficients and simultaneous equations for assaying chlorophylls a and b extracted with four different solvents: verification of the concentration of chlorophyll standards by atomic absorption spectroscopy. Biochim Biophys Acta BBA Bioenerg. 1989; 975( 3): 384– 94. doi:10.1016/S0005-2728(89)80347-0. [Google Scholar] [CrossRef]
32. Song H , Huang Y , Gu B . QTL-Seq identifies quantitative trait loci of relative electrical conductivity associated with heat tolerance in bottle gourd (Lagenaria siceraria). PLoS One. 2020; 15( 11): e0227663. doi:10.1371/journal.pone.0227663. [Google Scholar] [CrossRef]
33. Yu J , Cang J , Lu Q , Fan B , Xu Q , Li W , et al. ABA enhanced cold tolerance of wheat ‘dn1’ via increasing ROS scavenging system. Plant Signal Behav. 2020; 15( 8): 1780403. doi:10.1080/15592324.2020.1780403. [Google Scholar] [CrossRef]
34. Munns R , Tester M . Mechanisms of salinity tolerance. Annu Rev Plant Biol. 2008; 59: 651– 81. doi:10.1146/annurev.arplant.59.032607.092911. [Google Scholar] [CrossRef]
35. Ashraf M , Harris PJC . Potential biochemical indicators of salinity tolerance in plants. Plant Sci. 2004; 166( 1): 3– 16. doi:10.1016/j.plantsci.2003.10.024. [Google Scholar] [CrossRef]
36. Kaur G , Asthir B . Proline: a key player in plant abiotic stress tolerance. Biol Plant. 2015; 59( 4): 609– 19. doi:10.1007/s10535-015-0549-3. [Google Scholar] [CrossRef]
37. Parida AK , Das AB . Salt tolerance and salinity effects on plants: a review. Ecotoxicol Environ Saf. 2005; 60( 3): 324– 49. doi:10.1016/j.ecoenv.2004.06.010. [Google Scholar] [CrossRef]
38. Nelson DL , Cox MM . Lehninger Principles of Biochemistry. New York, NY, USA: Recording for the Blind & Dyslexic; 2017. [Google Scholar]
39. Dionisio-Sese ML , Tobita S . Antioxidant responses of rice seedlings to salinity stress. Plant Sci. 1998; 135( 1): 1– 9. doi:10.1016/S0168-9452(98)00025-9. [Google Scholar] [CrossRef]
40. Mittler R . Oxidative stress, antioxidants and stress tolerance. Trends Plant Sci. 2002; 7( 9): 405– 10. doi:10.1016/s1360-1385(02)02312-9. [Google Scholar] [CrossRef]
41. Hu J , Shibata Y , Zhu PP , Voss C , Rismanchi N , Prinz WA , et al. A class of dynamin-like GTPases involved in the generation of the tubular ER network. Cell. 2009; 138( 3): 549– 61. doi:10.1016/j.cell.2009.05.025. [Google Scholar] [CrossRef]
42. Voeltz GK , Prinz WA , Shibata Y , Rist JM , Rapoport TA . A class of membrane proteins shaping the tubular endoplasmic reticulum. Cell. 2006; 124( 3): 573– 86. doi:10.1016/j.cell.2005.11.047. [Google Scholar] [CrossRef]
43. Yang R , Yang Z , Xing M , Jing Y , Zhang Y , Zhang K , et al. TaBZR1 enhances wheat salt tolerance via promoting ABA biosynthesis and ROS scavenging. J Genet Genom. 2023; 50( 11): 861– 71. doi:10.1016/j.jgg.2023.09.006. [Google Scholar] [CrossRef]
44. Saradadevi R , Palta JA , Siddique KHM . ABA-mediated stomatal response in regulating water use during the development of terminal drought in wheat. Front Plant Sci. 2017; 8: 1251. doi:10.3389/fpls.2017.01251. [Google Scholar] [CrossRef]
45. Zou Z , Khan A , Khan A , Tao Z , Zhang S , Long Q , et al. Activation of ABA signaling pathway and up-regulation of salt-responsive genes confer salt stress tolerance of wheat (Triticum aestivum L.) seedlings. Agronomy. 2024; 14( 9): 2095. doi:10.3390/agronomy14092095. [Google Scholar] [CrossRef]
46. Al Hinai MS , Ullah A , Al-Rajhi RS , Farooq M . Proline accumulation, ion homeostasis and antioxidant defence system alleviate salt stress and protect carbon assimilation in bread wheat genotypes of Omani origin. Environ Exp Bot. 2022; 193: 104687. doi:10.1016/j.envexpbot.2021.104687. [Google Scholar] [CrossRef]
47. Verslues PE , Sharma S . Proline metabolism and its implications for plant-environment interaction. Arab Book. 2010; 8: e0140. doi:10.1199/tab.0140. [Google Scholar] [CrossRef]
48. Nadarajah KK . ROS homeostasis in abiotic stress tolerance in plants. Int J Mol Sci. 2020; 21( 15): 5208. doi:10.3390/ijms21155208. [Google Scholar] [CrossRef]
49. Puniran-Hartley N , Hartley J , Shabala L , Shabala S . Salinity-induced accumulation of organic osmolytes in barley and wheat leaves correlates with increased oxidative stress tolerance: in planta evidence for cross-tolerance. Plant Physiol Biochem. 2014; 83: 32– 9. doi:10.1016/j.plaphy.2014.07.005. [Google Scholar] [CrossRef]
50. Mittler R , Zandalinas SI , Fichman Y , Van Breusegem F . Reactive oxygen species signalling in plant stress responses. Nat Rev Mol Cell Biol. 2022; 23( 10): 663– 79. doi:10.1038/s41580-022-00499-2. [Google Scholar] [CrossRef]
51. Turan S , Tripathy BC . Salt-stress induced modulation of chlorophyll biosynthesis during de-etiolation of rice seedlings. Physiol Plant. 2015; 153( 3): 477– 91. doi:10.1111/ppl.12250. [Google Scholar] [CrossRef]
52. Nassar RMA , Kamel HA , Ghoniem AE , Alarcón JJ , Sekara A , Ulrichs C , et al. Physiological and anatomical mechanisms in wheat to cope with salt stress induced by seawater. Plants. 2020; 9( 2): 237. doi:10.3390/plants9020237. [Google Scholar] [CrossRef]
53. van Zelm E , Zhang Y , Testerink C . Salt tolerance mechanisms of plants. Annu Rev Plant Biol. 2020; 71: 403– 33. doi:10.1146/annurev-arplant-050718-100005. [Google Scholar] [CrossRef]
54. Bekkering C , Yu S , Kuo CC , Tian L . Distinct growth patterns in seedling and tillering wheat plants suggests a developmentally restricted role of HYD2 in salt-stress response. Plant Cell Rep. 2024; 43( 5): 119. doi:10.1007/s00299-024-03206-x. [Google Scholar] [CrossRef]
55. Huang Y , Zhou J , Li Y , Quan R , Wang J , Huang R , et al. Salt stress promotes abscisic acid accumulation to affect cell proliferation and expansion of primary roots in rice. Int J Mol Sci. 2021; 22( 19): 10892. doi:10.3390/ijms221910892. [Google Scholar] [CrossRef]
56. Vives-Peris V , López-Climent MF , Moliner-Sabater M , Gómez-Cadenas A , Pérez-Clemente RM . Morphological, physiological, and molecular scion traits are determinant for salt-stress tolerance of grafted Citrus plants. Front Plant Sci. 2023; 14: 1145625. doi:10.3389/fpls.2023.1145625. [Google Scholar] [CrossRef]
57. Gharib MAAH , Qabil N , Salem AH , Ali MMA , Awaad HA , Mansour E . Characterization of wheat landraces and commercial cultivars based on morpho-phenological and agronomic traits. Cereal Res Commun. 2021; 49( 1): 149– 59. doi:10.1007/s42976-020-00077-2. [Google Scholar] [CrossRef]
58. Yang H , Chen R , Chen Y , Li H , Wei T , Xie W , et al. Agronomic and physiological traits associated with genetic improvement of phosphorus use efficiency of wheat grown in a purple lithomorphic soil. Crop J. 2022; 10( 4): 1151– 64. doi:10.1016/j.cj.2021.11.010. [Google Scholar] [CrossRef]
59. Van De Velde K , Thomas SG , Heyse F , Kaspar R , Van Der Straeten D , Rohde A . N-terminal truncated RHT-1 proteins generated by translational reinitiation cause semi-dwarfing of wheat Green Revolution alleles. Mol Plant. 2021; 14( 4): 679– 87. doi:10.1016/j.molp.2021.01.002. [Google Scholar] [CrossRef]
60. Sattar A , Sher A , Ijaz M , Ul-Allah S , Rizwan MS , Hussain M , et al. Terminal drought and heat stress alter physiological and biochemical attributes in flag leaf of bread wheat. PLoS One. 2020; 15( 5): e0232974. doi:10.1371/journal.pone.0232974. [Google Scholar] [CrossRef]
61. Yang D , Liu Y , Cheng H , Chang L , Chen J , Chai S , et al. Genetic dissection of flag leaf morphology in wheat (Triticum aestivum L.) under diverse water regimes. BMC Genet. 2016; 17( 1): 94. doi:10.1186/s12863-016-0399-9. [Google Scholar] [CrossRef]
Cite This Article
 Copyright © 2026 The Author(s). Published by Tech Science Press.
Copyright © 2026 The Author(s). Published by Tech Science Press.This work is licensed under a Creative Commons Attribution 4.0 International License , which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.


 Submit a Paper
Submit a Paper Propose a Special lssue
Propose a Special lssue View Full Text
View Full Text Download PDF
Download PDF Downloads
Downloads
 Citation Tools
Citation Tools