 Open Access
Open Access
ARTICLE
Differential Activation of Defense Enzymes in Clonal Lines of Agave americana Derived from Chemical Mutagenesis in Response to Fusarium oxysporum Infection
Tecnológico Nacional de México, División de Estudios de Posgrado e Investigación, Instituto Tecnológico de Tuxtla Gutiérrez, Carretera Panamericana Km 1080, Tuxtla Gutiérrez, Chiapas, México
* Corresponding Authors: Joaquín Adolfo Montes-Molina. Email: ; Carlos Alberto Lecona-Guzmán. Email:
(This article belongs to the Special Issue: Advances in Ornamental Plants: Micropropagation, Plant Biotechnology, Chromosome Doubling, Mutagenesis, Plant Breeding, Environmental Stress Tolerance, and Postharvest Physiology)
Phyton-International Journal of Experimental Botany 2026, 95(3), 14 https://doi.org/10.32604/phyton.2026.076451
Received 21 November 2025; Accepted 18 February 2026; Issue published 31 March 2026
Abstract
Agave americana L. is potentially a source of functional and nutritive compounds. However, its yield has been compromised by vascular wilt, which is associated with the presence of the Fusarium oxysporum. In response to this phytosanitary problem, the implementation of efficient methods to mitigate the damage. Biotechnological techniques offer a viable alternative to improve and increase the production of species of interest via genetic improvement. By use of mutagenic chemical agents, these techniques have been consolidated as a powerful tool to induce genetic variability and select genotypes with greater tolerance to pathogens. In this study, we evaluated the activity of three pathogenesis-related (PR) enzymes, namely β-1,3-glucanase (PR2), chitinase (PR3), and peroxidase (PR9), that were present in second generation (F2) plants belonging to two resistant clonal lines and to one A. americana line that was susceptible to F. oxysporum. This was used to generate information that can be applied to biotechnology strategies for genetic preservation of the species and agro-industrial use. Plants that were six months old were selected, which were susceptible (Control) and resistant (Clone 1, Clone 4). These were inoculated with F. oxysporum and kept under controlled greenhouse conditions. Enzyme activity was assessed at 0 and 72 hpi and then 7, 15, and 30 dpi. The results showed a significant decrease in β-1,3-glucanase enzyme activity in susceptible plants (control) and in clone 4, as compared to clone 1 plants. Regarding chitinase activity, a progressive increase was observed in control plants and clone 1 over time, while a steadier regulation was detected in clone 4. Lastly, peroxidase activity decreased over time in infected control plants, while dynamic variations were observed in both clonal lines, suggesting a regulated response. These findings indicate that enzyme activity plays a key role in defense against pathogens and that each clonal line of A. americana could activate different defense mechanisms against infection by F. oxysporum.Keywords
Supplementary Material
Supplementary Material FileAgave americana L. belongs to the Agavaceae family. The agave genus exhibits remarkable biomass productivity and resilience under drought conditions, making it particularly suitable for cultivation in arid and semi-arid environments due to its crassulacean acid metabolism (CAM) [1].
As it is found within the agave genus, it serves as a reference species for investigating physiological and molecular responses to biotic and abiotic stress conditions. Additionally, its economic relevance is underscored by its use in the production of natural fibers, bio-inputs, and culturally significant traditional beverages [2].
Despite its agronomic and ecological importance, the natural distribution of A. americana in Mexico has experienced a notable decline. The reduction is attributed to factors such as diminished crop diversification, progressive soil degradation, and increased incidence of phyto pathological disorders, all of which constrain genetic recombination and hinder the evolutionary potential of local [3]. One of the main threats to this crop is vascular wilt, caused by Fusarium oxysporum, which affects its productive and ecological viability [4].
In response to pathogenic invasion, plants initiate a cascade of biochemical defense strategies, many of which involve the overexpression of pathogenesis-related proteins (PR). The main causes of PR toxicity are their hydrolytic, proteinases inhibitory, and membrane permeabilizing properties. Mainly, the proteins involved are PR2 and PR3, which are classified as glucanases and chitinases, respectively, as well as peroxidases (PR9) [5].
The implementation of methodology that focuses on obtaining pest and disease-resistant genotypes via mutagenic techniques, such as the use of ethyl methanesulfonate to induce hereditable changes in genetic structure, has been efficiently employed as a tool for genetic plant improvement [6]. This approach has proved to be an environmentally sustainable strategy, as it is based on early gene expression pertaining to plant defense [7]. This mechanism allows plants to respond more quickly to stress, thus reducing the use of agrochemical products like insecticides, bactericides, and fungicides [8]. Moreover, Wagh et al. [9] point out that the economic and agronomic consequences of pests and diseases are severe and are further exacerbated by the intensive use of pesticides, which promotes the emergence of pathogens that are resistant to chemical compounds.
Recently, Reyes-Zambrano et al. [10] reported obtaining somaclonal variants resistant to F. oxysporum at the in vitro level, which resulted from the induction of mutations with ethyl methanesulfonate (EMS).
The study of enzymes related to plant defense allows establishing functional markers for the selection of resistant genotypes [11]. Therefore, the objective of this study was to quantify the enzymatic activity of β-1,3-glucanase, chitinases, and peroxidases in second generation (F2) A. americana plants that were susceptible and resistant to F. oxysporum.
The plant material was obtained from the germplasm bank of Agave americana L. at the plant tissue culture laboratory of the Technological National of Mexico, Tuxtla Gutiérrez Campus. Mother plants (first generation) resistant to F. oxysporum were obtained in vitro by chemical mutagenesis, Reyes-Zambrano et al. [10]. The second generation of resistant plants was obtained from shoots of the mother plants under in vitro conditions, two resistant clonal lines (clone 1 and clone 4) as well as one susceptible line (control) were selected, both approximately six months old, were acclimatized in greenhouse conditions, placed in pots with a mixture of peat moss and agrolite as substrate in a (1:1) proportion.
Inoculation was carried out following the methodology described by Reyes-Zambrano et al. [12]. The inoculum consisted of a suspension of F. oxysporum conidia (2 × 108 conidia mL−1) corresponding to the ITTG_Foxy_C6 isolate, previously obtained from the basal tissue of an A. americana stem, which was registered in GenBank under access number MT791313. Before inoculation, plant roots were superficially wounded with a scalpel, and the inoculum was then applied directly to the roots. Treated plants were placed in a growth chamber at 28°C and a relative humidity between 60 and 90%. After inoculation, samples were collected at 0, 72 h post-inoculation (hpi), and at 7, 15, and 30 days post-inoculation (dpi) and stored at −4°C until ready to be used for enzyme activity determination.
2.1.1 Extraction of Defense-Related Enzymes (PR)
For the extraction of PR proteins (glucanases and chitinases), 1 g of fresh plant material was weighed and then macerated in liquid nitrogen, after which it was homogenized with phosphate buffer in a 1:1 (w/v) ratio [13]. For the extraction of peroxidases, 100 mM sodium phosphate buffer (pH 7) supplemented with 0.1 mM EDTA and 1% (w/v) polyvinylpyrrolidone (PVP) was used in a 1:1 (w/v) ratio. Samples were then centrifuged at 10,000 rpm for 20 min at 4°C, whereby the supernatant was used for enzymatic determinations. The soluble protein content in the supernatant contained within the crude enzyme extract was determined by the Bradford method [14].
2.1.2 β-1,3-Glucanase (PR2) Activity
β-1,3-glucanase activity was quantified by the modified method reported by Fink et al. [15]. The reaction mixture contained 150 μL of enzyme extract, 150 μL of Laminarin (2 mg mL−1), and 150 μL of Somogyi’s reagent [16]. After incubation with boiling water for 40 min, 150 μL of Nelson’s reagent [17] and 1.5 mL of distilled water were added. Subsequently, the absorbance was measured at 610 nm. The specific activity was expressed as mg of glucose mg−1 protein min−1, using a glucose standard curve for determinations.
2.1.3 Chitinase Activity (PR3)
Chitinase activity was quantified following the protocol established by Boller et al. [18], with minor adjustments to optimize reaction conditions. The assay mixture contained 0.2 mL of colloidal chitin and 0.4 mL of enzyme extract, which were homogenized with a vortex. Subsequently, 100 μL of 0.8 M sodium tetraborate buffer was added, and the reaction mixture was incubated in a water bath at 37°C for 60 min. After incubation, the samples were then centrifuged at 4000 rpm for 5 min. Once the time had elapsed, 0.4 mL of the supernatant was collected and subjected to boiling for 3 min. Afterwards, the samples were allowed to cool, and 1 mL of p-dimethylaminobenzaldehyde (10%) was added to each sample. The mixture was vortexed briefly and incubated again at 30°C for 20 min. Absorbance readings were carried out at 585 nm using a spectrophotometer. A standard curve was made with N-acetylglucosamine. Chitinase activity was expressed as μmol min−1 mg−1 of protein, according to the method described by Reissig et al. [19].
2.1.4 Peroxidase Activity (PR9)
The enzymatic activity of peroxidase was assessed following the protocol described by Hammerschmidt et al. [20]. The assay consisted of 3 mL of a freshly prepared mixture containing 0.25% (v/v) guaiacol, 10 mM sodium phosphate buffer (pH 6), and hydrogen peroxide as the substrate. The reaction was initiated by adding 100 μL of enzyme extract, and the increase in absorbance was measured at 470 nm/min using a spectrophotometer. Enzymatic activity was calculated based on the rate of absorbance and expressed as U mg of protein−1.
The experimental design was completely randomized. The values that are shown correspond to the mean value of experiments performed in triplicate. To assess statistically significant differences between treatments, data was subjected to a one-way analysis of variance (ANOVA), followed by Tukey’s multiple comparison test, considering a significance level of p ≤ 0.05. Tukey’s test was also applied to obtain the Honestly Significant Difference (HSD).
3.1 Defense-Related Enzyme (PR) Activity
The 30-day interaction between plantlets of A. americana and F. oxysporum induced the activation of enzymatic responses as a component of plant defense mechanisms, which occurred at different times after inoculation.
3.2 Specific β-1,3-Glucanases Activity
The findings show that the control and clone 1 treatments without inoculation presented basal levels of β-1,3-glucanase enzymatic activity, which increased slightly over time, likely happening as a response to phenological development or environmental stimuli (Fig. 1 and S1(Supplementary Materials)). In Clone 1 plants that were inoculated with F. oxysporum, an early induction of enzymatic activity was observed, where the highest levels occurred at 72 hpi and 7 dpi. This suggests that plants from clone 1 have a more efficient defense system, possibly associated with the rapid activation of PR2 genes that are involved in the degradation of components within the cell wall of fungi. Lastly, an increase in enzymatic activity was observed over time in control treatment inoculated with F. oxysporum. However, this was lower than what was observed for the case of inoculated Clone 1.
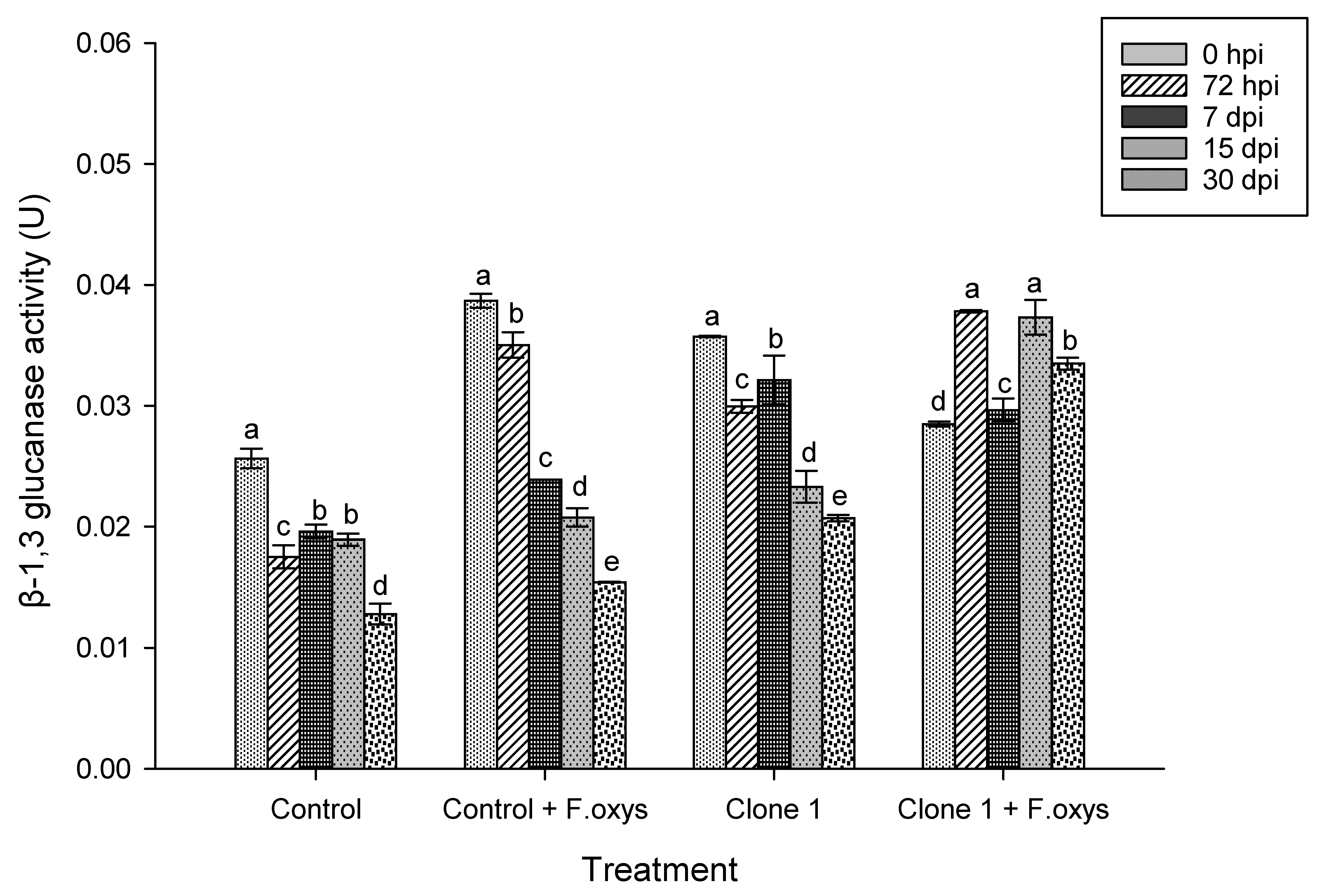
Figure 1: Enzymatic activity observed for β-1,3 glucanases during the advance of wilt disease in A. americana clone 1 plants. Values represent the mean ± standard error (SE). Group Means with different letters are significantly different (Tukey, p ≤ 0.05). HSD control = 0.006, HSD control + F. oxysporum = 0.006, Clone 1 = 0.00, Clone 1 + F. oxysporum = 0.00. hpi: hours post-inoculation; dpi: days post-inoculation.
Regarding the activity in non-inoculated clone 4 plants, a slight increase was observed at 72 hpi and 7 dpi (Fig. 2 and S1), possibly as a response to phenological development or environmental stimuli. On the other hand, the highest enzymatic activity of β-1,3-glucanases in inoculated control plants was observed at 0 hpi, where a gradual decrease over time was noticed. A similar behavior was observed in clone 4 plants, which had a higher enzymatic activity at 0 and 72 hpi, and, very importantly, a statistically significant difference regarding the enzymatic activity at 30 dpi was found.
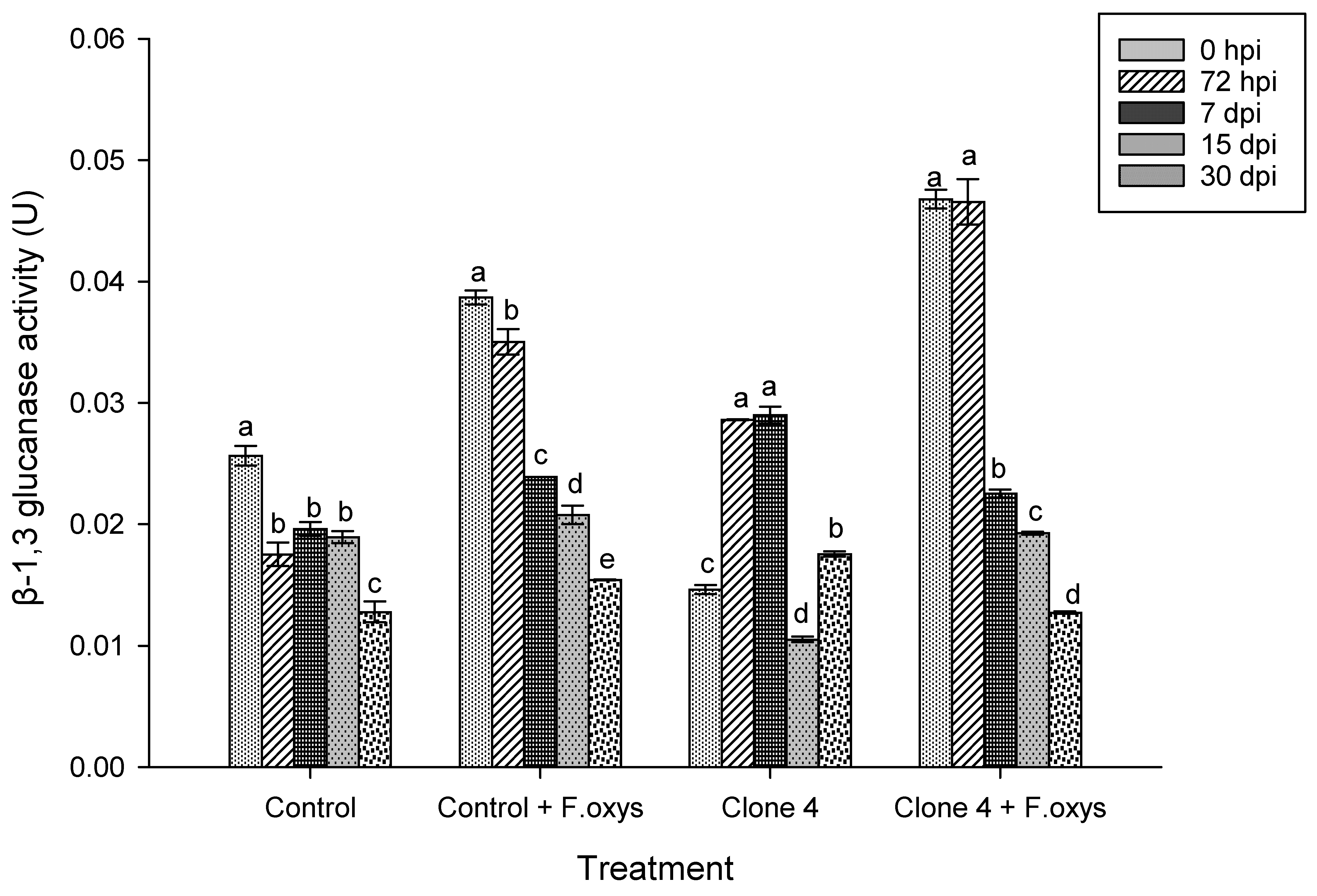
Figure 2: Enzymatic activity observed for β-1,3 glucanases during the advance of wilt disease in A. americana clone 4 plants. Values represent mean ± standard error (SE). Group means with different letters are significantly different (Tukey, p ≤ 0.05). HSD control = 0.006, HSD control + F. oxysporum = 0.006, Clone 4 = 0.006, Clone 4 + F. oxysporum = 0.006. hpi: hours post-inoculation; dpi: days post-inoculation.
3.3 Specific Chitinase Activity
The activity of chitinase enzyme in A. americana control plants inoculated with F. oxysporum showed an increase in activity after 7 dpi, although the highest activity was seen at 30 dpi. This can be seen in Fig. 3 and S1. A statistically significant difference was found after comparisons with all other assessed times were performed. The same behavior could be seen in clone 1 plants, even though the concentration of this enzyme was lower than for the infected control.
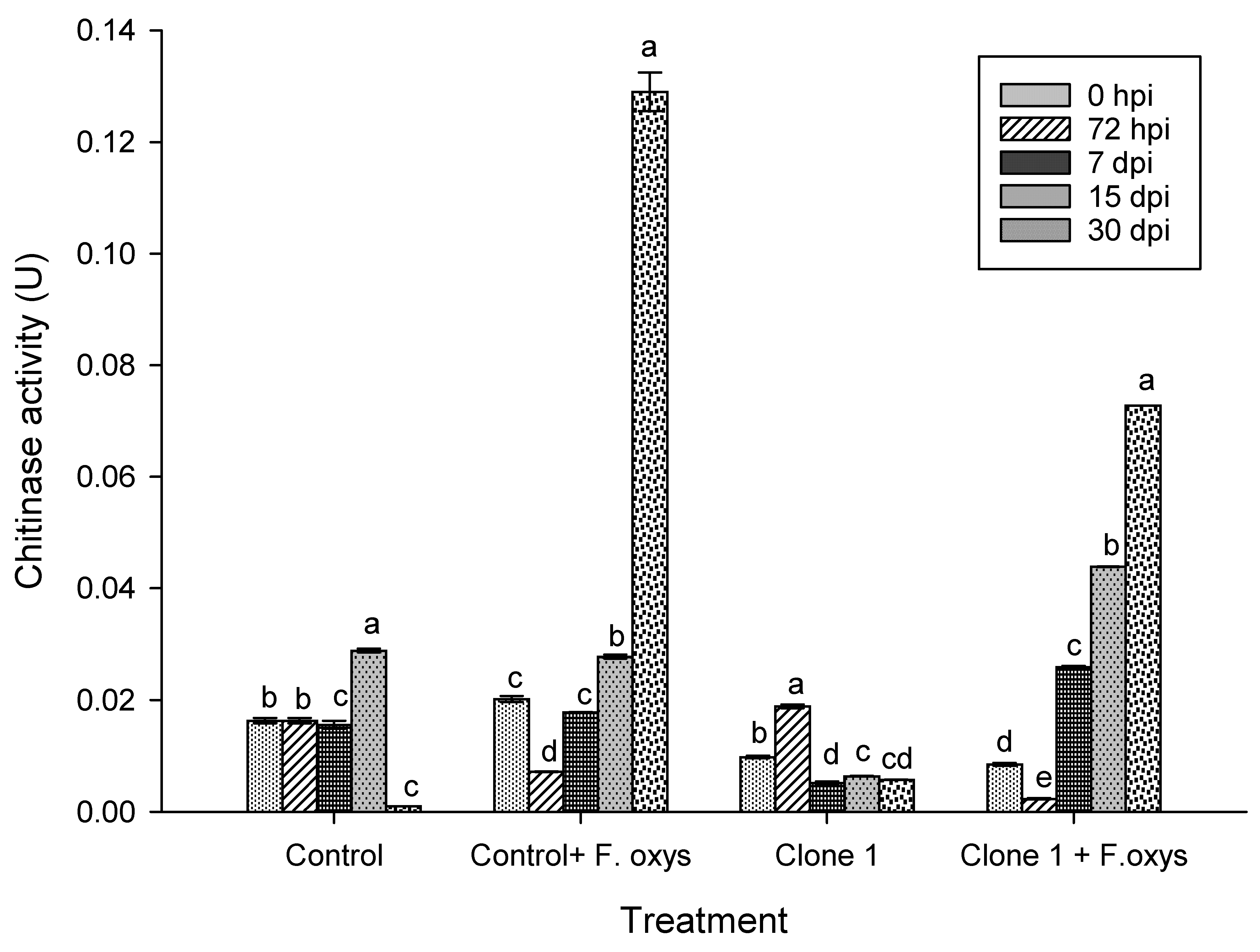
Figure 3: Enzymatic activity of chitinases during the advance of wilt disease in A. americana clone 1 plants. Values represent mean ± standard error (SE). Group means with different letters are significantly different (Tukey, p ≤ 0.05). HSD control = 0.006, HSD control + F. oxysporum = 0.006, Clone 1 = 0.006, Clone 1 + F. oxysporum = 0.00. hpi: hours post-inoculation; dpi: days post-inoculation.
In treatments lacking the pathogen, the enzymatic activity of chitinase was low (Fig. 4 and S1), where a slight increase was observed at 15 dpi in control plants. Additionally, in the case of clone 4 control plants, an increase in activity occurred at 7 and 15 dpi, while in infected clone 4 plants, the highest activity was observed later on at 15 dpi and 30 dpi. This activity then decreased significantly, suggesting a regulated and efficient defense strategy.
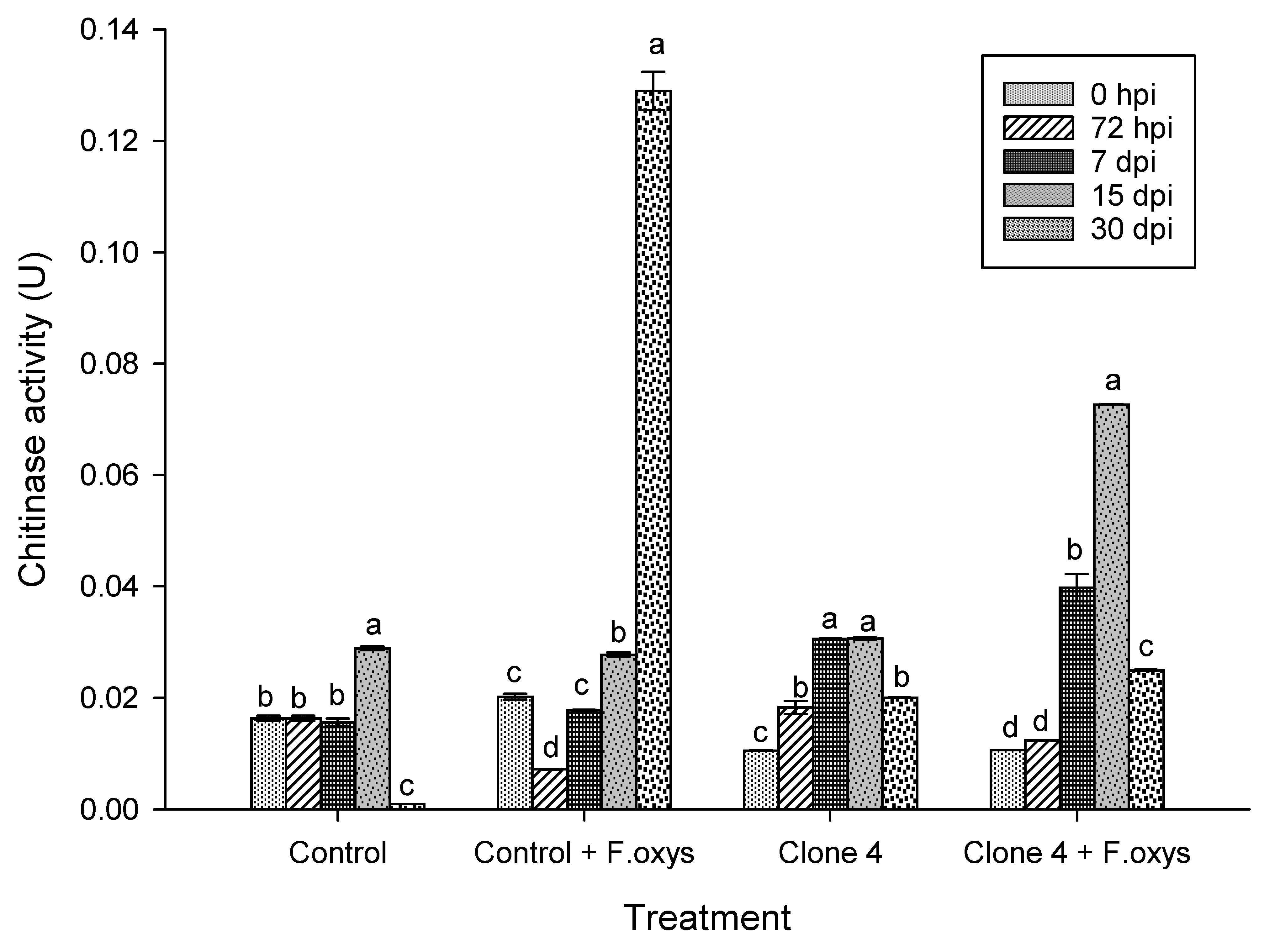
Figure 4: Enzymatic activity of chitinases during the advance of wilt disease in A. americana clone 4 plants. Values represent mean ± standard error (SE). Group means with different letters are significantly different (Tukey, p ≤ 0.05). HSD control = 0.006, HSD control + F. oxysporum = 0.006, Clone 4 = 0.006, Clone 4 + F. oxysporum = 0.006. hpi: hours post-inoculation; dpi: days post-inoculation.
3.4 Specific Peroxidases Activity
Results regarding enzymatic activity of peroxidases showed a statistically significant difference between treatments and time (Fig. 5 and S1). The uninoculated control showed basal activity, whereas clone 1 plants inoculated with F. oxysporum showed a progressive increase in enzymatic activity with a maximum peak at 7 dpi, which was higher than the rest of the groups that were assessed. In inoculated control plants, an increase was observed from 0 hpi to 7 dpi. However, these plants did not present a statistically significant difference and this activity decreased drastically after 15 dpi.
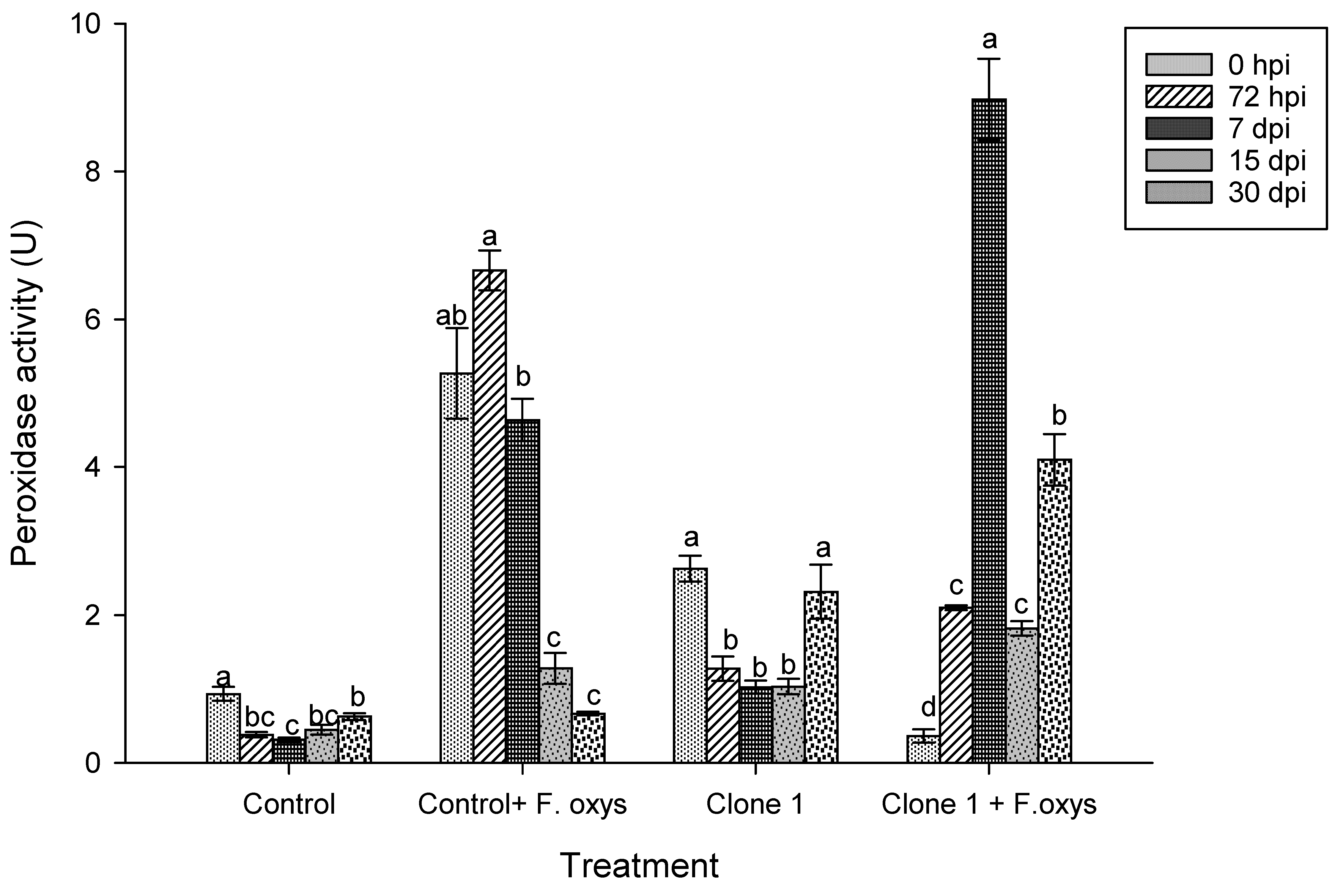
Figure 5: Enzymatic activity of peroxidases during the advance of wilt disease in A. americana clone 1 plants. Values represent mean ± standard error (SE). Group means with different letters are significantly different (Tukey, p ≤ 0.05). HSD control = 0.27, HSD control + F. oxysporum = 1.57, Clone 1 = 0.95, Clone 1 + F. oxysporum = 1.38. hpi: hours post-inoculation; dpi: days post-inoculation.
Regarding the activity of the peroxidase enzyme in non-inoculated clone 4 plants (Fig. 6 and S1), a basal activity was observed, while in inoculated plants a significant increase was observed at 15 dpi, suggesting an activation of the antioxidant defense system in response to the pathogen.
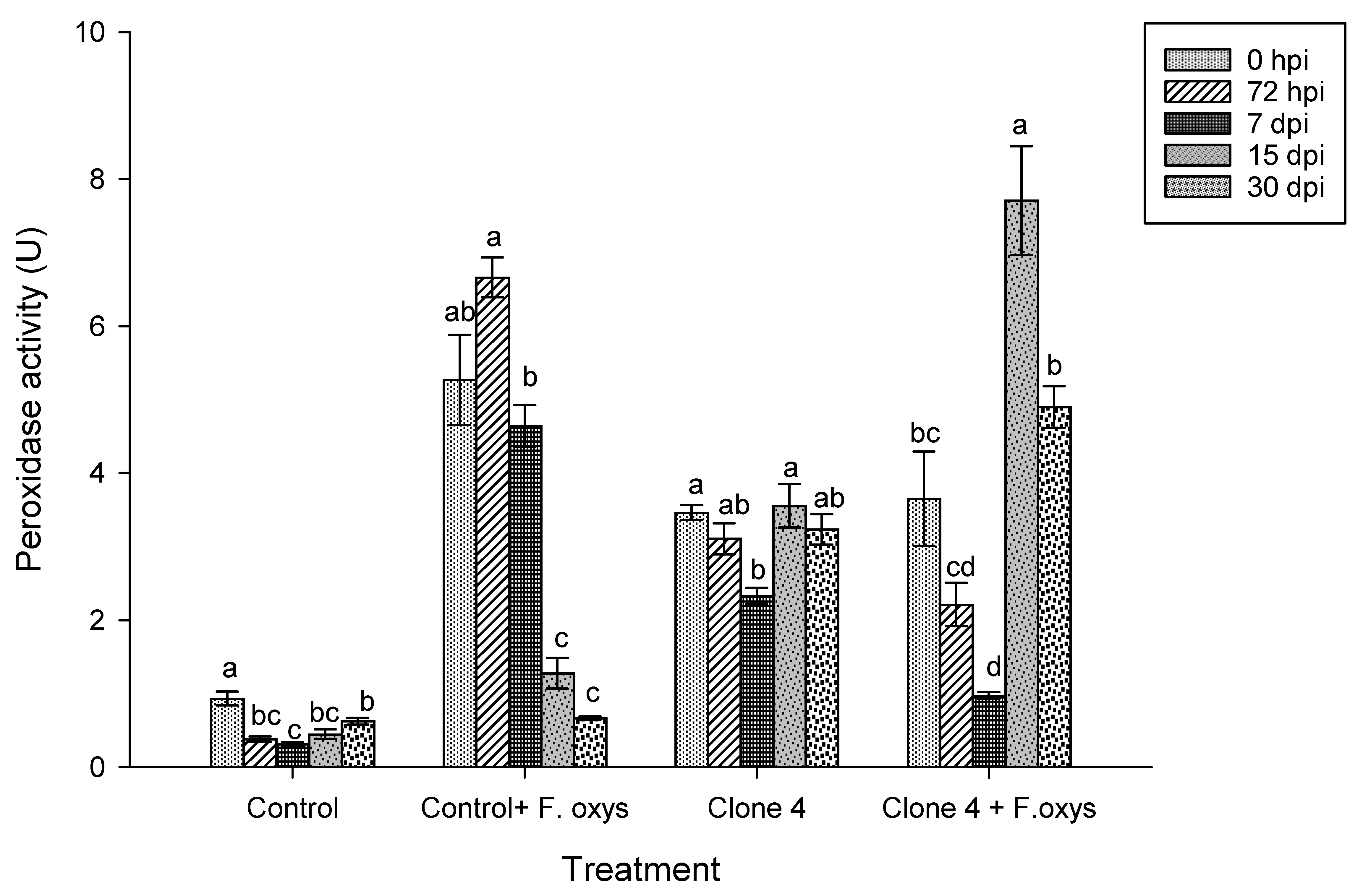
Figure 6: Enzymatic activity of peroxidases during the progression of wilt disease in A. americana clone 4 plants. Values represent mean ± standard error (SE). Group means with different letters are significantly different (Tukey, p ≤ 0.05). HSD control = 0.27, HSD control + F. oxysporum = 1.57, Clone 4 = 0.91, Clone 4 + F. oxysporum = 1.49. hpi: hours post-inoculation; dpi: days post-inoculation.
Plants are exposed to various pathogens in their natural habitat, and these interactions trigger multiple defensive responses involving both constitutive and inducible barriers [21]. These defense mechanisms, called “Systemic Resistance” are activated only in the presence of the pathogen, where inhibitors are released as a means of defense, such as accumulation of phytoalexins, and production of enzymes related to pathogenesis (PR) and secondary metabolites including, salicylic acid, jasmonic acid, among others [5]. PR proteins have an important function in the defense against pathogens containing β-1,3-glucan and chitin.
The obtained results show the activity of enzymes involved in the defense mechanism of A. americana plants against infection by F. oxysporum. As part of the results, a modulated response was observed in plants from treatments with ethyl methanesulfonate. These results tentatively suggest that the second generation of these clonal lines, which result from genetic improvement when subjected to chemical agents, could contribute to the differential activation of defense mechanisms in A. americana. This is then reflected in the regulation of the activity of enzymes such as glucanases (PR2), chitinases (PR3), and peroxidases (PR9).
In regard to enzyme β-1,3-glucanase activity, it has been described that this enzyme inhibits mycelial growth and the formation of sporangia. It acts by hydrolyzing the cell walls of fungal pathogens, usually in combination with another enzyme, such as chitinases and peroxidases, which belong to the group of PR2 proteins [5,22].
Reyes-Zambrano et al. [12] reported the activity of β-1,3-glucanase in A. americana plants inoculated with F. oxysporum, whereby a slight increase in its activity at 15 dpi was observed, compared to the non-inoculated ones. In our study, β-1,3-glucanase activity correlated directly with time. This suggests that the F2 plants from clone 4 that were obtained with EMS showed regulated enzymatic activity, thus counteracting the presence of F. oxysporum and preventing the appearance of wilt disease symptoms.
López-Velázquez et al. [23] carried out a study to research ways to control rust in coffee (Coffea arabica) by using chitosan. They observed that β-1,3-glucanase activity decreased during the first 12 h after infection with Hemileia vastatrix and the response continued up until 24 h, suggesting that the pathogen managed to suppress enzymatic activity and infect the plant tissue. Recently, López-Velázquez et al. [24] reported that β-1,3-glucanase activity increased significantly in pepper plantlets (Capsicum annuum) infected with Phytophthora capsici and treated with inulin solutions from dahlia tubers (Dahlia sp.), occurring as early as 2 dpi when compared with the control. Activity decreased significantly at 7 dpi before increasing again at 9 dpi. Similarly, when infected plantlets were treated with inulin, an increase in activity was observed, starting at day 0 dpi and continuing throughout days 1, 2, 3, and 5 dpi. Thus, the use of inulin originating from dahlia is capable of increasing activity of β-1,3-glucanases in a systemic and early manner in pepper plants (Capsicum annuum) inoculated with P. capsici. Rizal et al. [25] reported an increase in the expression of β-1,3-glucanases enzyme activity in resistant lentil (Lens culinaris Medikus) genotypes, as compared to susceptible ones, after inoculation with Stemphylium botryosum Wallr.
On the other hand, it has been described that the synthesis of chitinases in plants is involved in protection against both biotic and abiotic factors. Plant chitinase have the ability to hydrolyze pathogen’s cell walls and trigger a defense response, restricting fungal growth by degrading chitin or by stimulating acquired systemic resistance, which results from various defense mechanisms such as the oxidative burst, the hypersensitivity response, salicylic acid generation, and lignin deposition, leading to increased production of other PR proteins [26]. The development of modified plants via techniques such as chemical mutagenesis favors yields by improving their characteristics, whereby the synthesis of a greater amount of chitinase is the main purpose behind alteration of plants, as it allows many of them to counteract the pathogens [27].
In this sense, Reyes Zambrano et al. [12], observed an increase in enzymatic activity of chitinases in A. americana plants that had been infected with F. oxysporum while still lacking mutagenic treatments (control plants) after 15 dpi, as compared to uninfected plants. At 30 dpi, this activity decreased significantly in infected plants, as compared to uninfected ones.
In our study, an increase in chitinase activity was observed as of 7 dpi in infected control plants, as well as in clone 1, which increased over time. However, the activity of this enzyme decreased significantly at 30 dpi for clone 4, thus suggesting inhibition of the pathogen.
Studies conducted by Liu et al. [28] in orchids (Bletilla striata) infected with Coleosporium bletiae demonstrated a decrease in chitinase activity in resistant orchids at 2 dpi, followed by a sharp recovery. In contrast, susceptible plants exhibited a gradual decline during 0–4 dpi followed by a delayed increase. These findings indicate that resistant material displayed greater sensitivity and enzymatic induction. Camacho-Luna et al. [29] evaluated the interaction between Trichoderma asperellum and Alternaria leek in onion plants (Allium cepa), finding significant differences in the activity of chitinases and glucanases, in regard to control plants.
Regarding peroxidase enzyme activity, it has been described that peroxidase is a key enzyme in the antioxidative defense of plants, which is responsible for breaking down hydrogen peroxide (H2O2), which catalyzes the polymerization of lignin monomers to reinforce the cell wall, thus reducing pathogen entry and modulating redox changes in distant tissues. In turn, this contributes towards establishing Systemic Acquired Resistance [30]. Gaibor-Vaca et al. [22] described a constant increase of peroxidase after 2 dpi in stems and leaves from chili plants (Capsicum annuum) that were resistant to P. capsici. As for the agave genus, Chávez-Sánchez et al. [31] reported an increase in peroxidases at 72 hpi in potentially resistant Agave tequilana blue var. plantlets that were inoculated with F. solani.
In addition, López-Velázquez et al. [23] reported that the use of chitosan for rust control in coffee plants (Coffea arabica) favors the activity of peroxidase enzymes starting at 12 hpi. It has been shown that the implementation of genetic improvement is an efficient method to obtain resistant genotypes that are suitable for crop health management, due to its cost-benefit ratio and low environmental impact [32]. Such strategies represent an alternative that would allow minimizing the use of chemical agents while safeguarding food security [33,34].
The use of resistant Agave varieties represents an effective strategy for the control of phytosanitary diseases, such as F. oxysporum-induced wilt in A. americana. Our findings show that each of the assessed A. americana clonal lines activates different defense mechanisms against infection by F. oxysporum. This suggests that the combined enzymatic activity occurring to β-1,3-glucanase, chitinase, and peroxidase interactions plays a key role in the defense response against pathogens. Furthermore, persisting resistance in second-generation (F2) plants strongly suggests that genetic improvement via chemical mutagenesis with ethyl methanesulfonate is a promising strategy for inducing resistance in this crop.
The results of this study indicate great feasibility in applying this approach in genetic improvement programs for other crops of agricultural interest, which would, in turn, contribute to more sustainable production systems with less dependence on fungicides. However, it is necessary to evaluate persistence in subsequent generations and under field conditions. Furthermore, transcriptomic and molecular analyses should be carried out in parallel to correlate enzymatic activity with the gene expression of pathogenesis-related proteins.
Acknowledgement:
Funding Statement: The authors received no specific funding for this study.
Author Contributions: The authors confirm contribution to the paper as follows: Conceptualization, Carlos Alberto Lecona-Guzmán, Joaquín Adolfo Montes-Molina; methodology, Carlos Alberto Lecona-Guzmán, Sugey Vásquez-Hernández, Joaquín Adolfo Montes-Molina; validation, Victor Manuel Ruiz-Valdiviezo, Federico Antonio Gutierrez-Miceli, Nancy Ruiz-Lau; formal analysis, Sugey Vásquez-Hernández, Joaquín Adolfo Montes-Molina, Carlos Alberto Lecona-Guzmán; writing—original draft preparation, Sugey Vásquez-Hernández, Joaquín Adolfo Montes-Molina, Carlos Alberto Lecona-Guzmán; supervision, Victor Manuel Ruiz-Valdiviezo, Federico Antonio Gutierrez-Miceli, Nancy Ruiz-Lau. All authors reviewed and approved the final version of the manuscript.
Availability of Data and Materials: The authors confirm that the data supporting the findings of this study are available within the article or its Supplementary Materials.
Ethics Approval: Not applicable.
Conflicts of Interest: The authors declare no conflicts of interest.
Supplementary Materials: The supplementary material is available online at https://www.techscience.com/doi/10.32604/phyton.2026.076451/s1.
References
1. Ávila-Reyes SV , Jiménez-Aparicio AR , Melgar-Lalanne G , Fajardo-Espinoza FS , Hernández-Sánchez H . Mezcal: A review of chemistry, processing, and potential health benefits. Foods. 2025; 14( 8): 1408. doi:10.3390/foods14081408. [Google Scholar] [CrossRef]
2. Lecona-Guzmán CA , Gutiérrez-Miceli FA , Reyes-Zambrano SJ , Vásquez-Hernández S , Joya-Dávila JG . American century plant (Agave americana L.) breeding. In: Breeding and biotechnology of leaf, fruit, and seed fiber crops. Berlin/Heidelberg, Germany: Springer; 2025. p. 109– 53. doi:10.1007/978-3-032-00407-9_3. [Google Scholar] [CrossRef]
3. Ruiz Mondragón KY , Klimova A , Aguirre-Planter E , Valiente-Banuet A , Lira R , Sanchez-de la Vega G , et al. Differences in the genomic diversity, structure, and inbreeding patterns in wild and managed populations of Agave potatorum Zucc. used in the production of Tobalá mezcal in Southern Mexico. PLoS One. 2023; 18( 11): e0294534. doi:10.1371/journal.pone.0294534. [Google Scholar] [CrossRef]
4. Quiñones-Aguilar EE , Montoya-Martínez AC , Rincón-Enríquez G , López-Pérez L . Inoculación de bulbilos de Agave tequilana con hongos micorrízicos arbusculares: Efecto en el crecimiento y biocontrol contra Fusarium oxysporum. Agric Sci Technol. 2022; 24( 1): 3043. (In Spainish). doi:10.21930/rcta.vol24_num1_art:3043. [Google Scholar] [CrossRef]
5. Rai AK , Sahoo LP , Sahoo M , Seth T , Patra C . Induced resistance mechanism in plant and its importance in agriculture. Int J Plant Soil Sci. 2024; 36( 5): 1– 22. doi:10.9734/ijpss/2024/v36i54497. [Google Scholar] [CrossRef]
6. Chen L , Duan L , Sun M , Yang Z , Li H , Hu K , et al. Current trends and insights on EMS mutagenesis application to studies on plant abiotic stress tolerance and development. Front Plant Sci. 2023; 13: 1052569. doi:10.3389/fpls.2022.1052569. [Google Scholar] [CrossRef]
7. Wang Z , Bart RS . Using targeted genome methylation for crop improvement. J Exp Bot. 2025; 76( 9): 2394– 404. doi:10.1093/jxb/eraf131. [Google Scholar] [CrossRef]
8. Flors V , Kyndt T , Mauch-Mani B , Pozo MJ , Ryu CM , Ton J . Enabling sustainable crop protection with induced resistance in plants. Front Sci. 2024; 2: 1407410. doi:10.3389/fsci.2024.1407410. [Google Scholar] [CrossRef]
9. Wagh SG , Patil AM , Patil GB , Bhor SA , Pawar KR , Shinde H . Programmable plant immunity: Synthetic biology for climate-resilient agriculture. SynBio. 2026; 4( 1): 1. doi:10.3390/synbio4010001. [Google Scholar] [CrossRef]
10. Reyes-Zambrano S , Alberto Lecona-Guzmán C , Antonio Gutirez-Miceli F , Manuel Ruiz-Valdiviezo V , Anayansi Vargas-Diaz A . Selection and analysis of polymorphisms in somaclonal variants of Agave americana resistant to Fusarium oxysporum via an ethyl methanesulphonate treatment. Phyton. 2021; 90( 6): 1727– 39. doi:10.32604/phyton.2021.016171. [Google Scholar] [CrossRef]
11. Le Cordon Bleu U , Menéndez E , Navarro J , Le Cordon Bleu U , López J , Le Cordon Bleu U , et al. Mecanismos de defensa en plantas. Proteínas relacionadas con la patogenicidad. Le Cordon Bleu Univ Res J. 2020; 7( 2): 98– 109. (In Spainish). doi:10.36955/riulcb.2020v7n2.010. [Google Scholar] [CrossRef]
12. Reyes-Zambrano SJ , Lecona-Guzmán CA , Gutiérrez-Miceli FA , Santana-Buzzy N , Islas-Flores I , Tzec-Simá M , et al. Scanning electron microscopy and enzymatic analysis in Agave americana during Fusarium oxysporum infection. Rev Mex De Fitopatología Mex J Phytopathol. 2020; 38( 3): 408– 19. doi:10.18781/r.mex.fit.2005-3. [Google Scholar] [CrossRef]
13. Santos-Espinoza AM , González-Mendoza D , Ruiz-Valdiviezo VM , Luján-Hidalgo MC , Jonapa-Hernández F , Valdez-Salas B , et al. Changes in the physiological and biochemical state of peanut plants (Arachis hypogaea L.) induced by exposure to green metallic nanoparticles. Int J Phytoremediat. 2021; 23( 7): 747– 54. doi:10.1080/15226514.2020.1856037. [Google Scholar] [CrossRef]
14. Bradford MM . A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem. 1976; 72: 248– 54. doi:10.1016/0003-2697(76)90527-3. [Google Scholar] [CrossRef]
15. Fink W , Liefland M , Mendgen K . Chitinases and β-1, 3-glucanases in the apoplastic compartment of oat leaves (Avena sativa L.). Plant Physiol. 1988; 88( 2): 270– 5. doi:10.1104/pp.88.2.270. [Google Scholar] [CrossRef]
16. Somogyi M . A new reagent for the determination of sugars. J Biol Chem. 1945; 160( 1): 61– 8. doi:10.1016/S0021-9258(18)43097-9. [Google Scholar] [CrossRef]
17. Nelson N . A photometric adaptation of the somogyi method for the determination of glucose. J Biol Chem. 1944; 153( 2): 375– 80. doi:10.1016/S0021-9258(18)71980-7. [Google Scholar] [CrossRef]
18. Boller T , Gehri A , Mauch F , Vögeli U . Chitinase from Phaseolus vulgaris leaves. In: Biomass part B: Lignin, pectin, and chitin. Amsterdam, The Netherlands: Elsevier; 1988. p. 479– 84. doi:10.1016/0076-6879(88)61062-7. [Google Scholar] [CrossRef]
19. Reissig JL , Storminger JL , Leloir LF . A modified colorimetric method for the estimation of N-acetylamino sugars. J Biol Chem. 1955; 217( 2): 959– 66. doi:10.1016/S0021-9258(18)65959-9. [Google Scholar] [CrossRef]
20. Hammerschmidt R , Nuckles EM , Kuć J . Association of enhanced peroxidase activity with induced systemic resistance of cucumber to Colletotrichum lagenarium. Physiol Plant Pathol. 1982; 20( 1): 73– 82. doi:10.1016/0048-4059(82)90025-X. [Google Scholar] [CrossRef]
21. Kumar P , Pandey S , Pati PK . Interaction between pathogenesis-related (PR) proteins and phytohormone signaling pathways in conferring disease tolerance in plants. Physiol Plant. 2025; 177( 2): e70174. doi:10.1111/ppl.70174. [Google Scholar] [CrossRef]
22. Gaibor-Vaca DG , García-Bazurto GL , Garcés-Fiallos FR . Mecanismos de defensa en plantas de Capsicum contra Phytophthora capsici. Bionatura. 2022; 7( 2): 1– 11. doi:10.21931/rb/2022.07.02.25. [Google Scholar] [CrossRef]
23. López-Velázquez JC , García-Morales S , Espinosa-Andrews H , Montero-Cortés MI , Qui-Zapata JA . Control de la roya del café utilizando moléculas de quitosanos de diferentes orígenes. In: XLIII Encuentro Nacional de la AMIDIQ. Puerto Vallarta, México: AMDIQ; 2022. p. 23– 6. [Google Scholar]
24. López-Velázquez JC , García-Morales S , Qui-Zapata JA , García-Carvajal ZY , Navarro-López DE , García-Varela R . Inducción de respuesta de defensa por inulina de tubérculos de dalia (Dahlia sp.) en Capsicum annuum. Mex J Phytopathol. 2024; 42: 1. (In Spainish). doi:10.18781/r.mex.fit.2305-2. [Google Scholar] [CrossRef]
25. Rizal S , Saha P , Mondal P , Mondol MSA , Datta J , Ahmadi T , et al. Physio-Biochemical changes in lentil genotypes under biotic stress induced by Stemphylium botryosum Wallr. BMC Plant Biol. 2025; 25( 1): 514. doi:10.1186/s12870-025-06508-8. [Google Scholar] [CrossRef]
26. Khan RS , Iqbal A , Bibi A , Khalil I , Ul Islam Z , Jan F , et al. Plant chitinases: Types, structural classification, antifungal potential and transgenic expression in plants for enhanced disease resistance. Plant Cell Tissue Organ Cult PCTOC. 2024; 156( 3): 75. doi:10.1007/s11240-024-02696-7. [Google Scholar] [CrossRef]
27. Lucas-Bautista JA , Bautista-Baños S , Ventura-Aguilar RI , Hernández-López M . Quitinasas en plantas y posible uso Como biomarcadores para el diseño de biosensores en la detección de hongos fitopatógenos. Remexca. 2022; 13( 4): 701– 13. doi:10.29312/remexca.v13i4.2717. [Google Scholar] [CrossRef]
28. Liu Q , Lu X , Wu Q , Lu Z , Qin R , Huang K , et al. Transcriptomics combined with physiological analysis provides insights into the mechanism of resistance to Coleosporium bletiae in Bletilla striata. Front Plant Sci. 2025; 16: 1604512. doi:10.3389/fpls.2025.1604512. [Google Scholar] [CrossRef]
29. Camacho-Luna V , Flores-Moctezuma HE , Rodríguez-Monroy M , Montes-Belmont R , Sepúlveda-Jiménez G . Inducción de la respuesta de defensa de plantas de cebolla en la interacción con Trichoderma asperellum y Alternaria porri. Remexca. 2021; 12( 4): 685– 98. doi:10.29312/remexca.v12i4.2683. [Google Scholar] [CrossRef]
30. Pandey R , Digambar WD , Padmaja O , Sampath V , Ashok PE , Yadav A , et al. Role of plant defence enzymes in combating fungal infections: A review. J Adv Biol Biotechnol. 2025; 28( 9): 96– 110. doi:10.9734/jabb/2025/v28i92863. [Google Scholar] [CrossRef]
31. Chávez-Sánchez C , Mancilla-Margalli NA , Montero-Cortés MI , Gutiérrez-Miceli FA , Briceño-Félix GA , Simpson Williamson JK , et al. Asexually propagated Agave tequilana var. azul exhibits variation in genetic markers and defence responses to Fusarium solani. AoB Plants. 2022; 14( 3): plac027. doi:10.1093/aobpla/plac027. [Google Scholar] [CrossRef]
32. Mariz BL , Caixeta ET , de Resende MDV , de Oliveira ACB , de Almeida DP , Alves DR . Exploring the genetic potential for multi-resistance to rust and other coffee phytopathogens in breeding programs. Plants. 2025; 14( 3): 391. doi:10.3390/plants14030391. [Google Scholar] [CrossRef]
33. Danso Ofori A , Zheng T , Titriku JK , Appiah C , Xiang X , Kandhro AG , et al. The role of genetic resistance in rice disease management. Int J Mol Sci. 2025; 26( 3): 956. doi:10.3390/ijms26030956. [Google Scholar] [CrossRef]
34. Limenie AD , Alehegn M . Genetic engineering for cereal crop yield improvement and disease resistant breeding. Sci World J. 2025; 2025( 1): 6743917. doi:10.1155/tswj/6743917. [Google Scholar] [CrossRef]
Cite This Article
 Copyright © 2026 The Author(s). Published by Tech Science Press.
Copyright © 2026 The Author(s). Published by Tech Science Press.This work is licensed under a Creative Commons Attribution 4.0 International License , which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.


 Submit a Paper
Submit a Paper Propose a Special lssue
Propose a Special lssue View Full Text
View Full Text Download PDF
Download PDF Downloads
Downloads
 Citation Tools
Citation Tools