 Open Access
Open Access
ARTICLE
Detection and Characterization of an Isolate of Cucumber Mosaic Virus Infecting Catharanthus roseus Using Deep Sequencing
1 Department of Biology, Faculty of Science, University of Tabuk, Tabuk, Saudi Arabia
2 Plant Molecular Virology Lab, Department of Botany, School of Chemical and Life Sciences, Jamia Hamdard (A Deemed-to-Be University), New Delhi, India
3 Centre for Virology, School of Interdisciplinary Sciences & Technology, Jamia Hamdard (A Deemed-to-Be University), New Delhi, India
4 Department of Botany and Microbiology, College of Science, King Saud University, Riyadh, Saudi Arabia
5 Department of Studies and Basic Sciences, Applied College, University of Tabuk, Tabuk, Saudi Arabia
6 Department of Science and Basic Studies, Applied College, University of Tabuk, Tabuk, Saudi Arabia
7 Biodiversity Genomic Unit, Faculty of Science, University of Tabuk, Tabuk, Saudi Arabia
* Corresponding Authors: Zahid Hameed Siddiqui. Email: ; Md Salik Noorani. Email:
# These authors contributed equally to this work
(This article belongs to the Special Issue: Technological Advances for Sustainable Management and Biological Control of Plant Pests and Diseases)
Phyton-International Journal of Experimental Botany 2026, 95(4), 8 https://doi.org/10.32604/phyton.2026.076432
Received 20 November 2025; Accepted 11 March 2026; Issue published 28 April 2026
Abstract
Cucumber mosaic virus (CMV) is among the most widespread plant viruses, infecting over a thousand plant species, including Catharanthus roseus, a medicinal plant valued for producing the anticancer alkaloids vincristine and vinblastine. Despite its economic significance, genomic information on CMV infecting C. roseus in India has been lacking. In this study, we employed small RNA deep sequencing integrated with advanced bioinformatics to generate the first complete genome of CMV infecting C. roseus in India, followed by validation through RT-PCR and Sanger sequencing. The reconstructed tripartite CMV genome encodes replication, silencing suppressor, movement, and coat proteins, consistent with known genomic organization. Sequence identity and phylogenetic analyses revealed a close relationship with Iranian tobacco isolates, placing the Indian isolate (CR8-JH) within subgroup IB. Recombination analysis indicated inter-fragmental recombination events, while mutational profiling highlighted natural variation, particularly in genes associated with host interaction. Conserved UTR motifs were identified, suggesting functional roles in viral replication and genome stability. By applying next-generation sequencing for accurate viral detection and characterization, this study enhances diagnostic capacity and contributes to technology-driven, sustainable plant disease management.Graphic Abstract
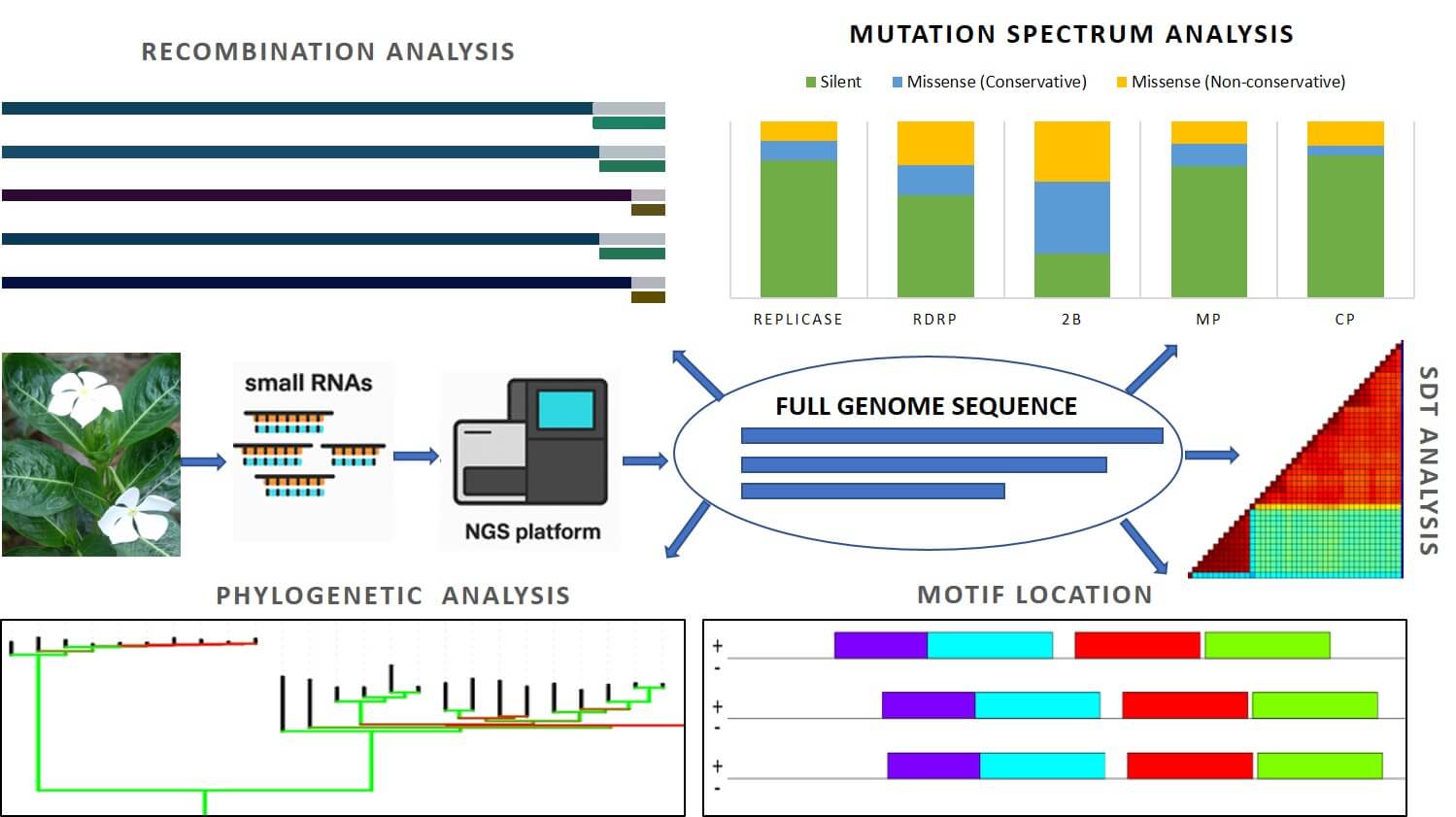
Keywords
Supplementary Material
Supplementary Material FileCite This Article
 Copyright © 2026 The Author(s). Published by Tech Science Press.
Copyright © 2026 The Author(s). Published by Tech Science Press.This work is licensed under a Creative Commons Attribution 4.0 International License , which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.


 Submit a Paper
Submit a Paper Propose a Special lssue
Propose a Special lssue View Full Text
View Full Text Download PDF
Download PDF Downloads
Downloads
 Citation Tools
Citation Tools