 Open Access
Open Access
REVIEW
A Survey of Convolutional Neural Network in Breast Cancer
School of Computing and Mathematical Sciences, University of Leicester, Leicester, LE1 7RH, UK
* Corresponding Author: Yu-Dong Zhang. Email:
(This article belongs to the Special Issue: Computer Modeling of Artificial Intelligence and Medical Imaging)
Computer Modeling in Engineering & Sciences 2023, 136(3), 2127-2172. https://doi.org/10.32604/cmes.2023.025484
Received 15 July 2022; Accepted 28 October 2022; Issue published 09 March 2023
Abstract
Problems: For people all over the world, cancer is one of the most feared diseases. Cancer is one of the major obstacles to improving life expectancy in countries around the world and one of the biggest causes of death before the age of 70 in 112 countries. Among all kinds of cancers, breast cancer is the most common cancer for women. The data showed that female breast cancer had become one of the most common cancers. Aims: A large number of clinical trials have proved that if breast cancer is diagnosed at an early stage, it could give patients more treatment options and improve the treatment effect and survival ability. Based on this situation, there are many diagnostic methods for breast cancer, such as computer-aided diagnosis (CAD). Methods: We complete a comprehensive review of the diagnosis of breast cancer based on the convolutional neural network (CNN) after reviewing a sea of recent papers. Firstly, we introduce several different imaging modalities. The structure of CNN is given in the second part. After that, we introduce some public breast cancer data sets. Then, we divide the diagnosis of breast cancer into three different tasks: 1. classification; 2. detection; 3. segmentation. Conclusion:Although this diagnosis with CNN has achieved great success, there are still some limitations. (i) There are too few good data sets. A good public breast cancer dataset needs to involve many aspects, such as professional medical knowledge, privacy issues, financial issues, dataset size, and so on. (ii) When the data set is too large, the CNN-based model needs a sea of computation and time to complete the diagnosis. (iii) It is easy to cause overfitting when using small data sets.Graphic Abstract
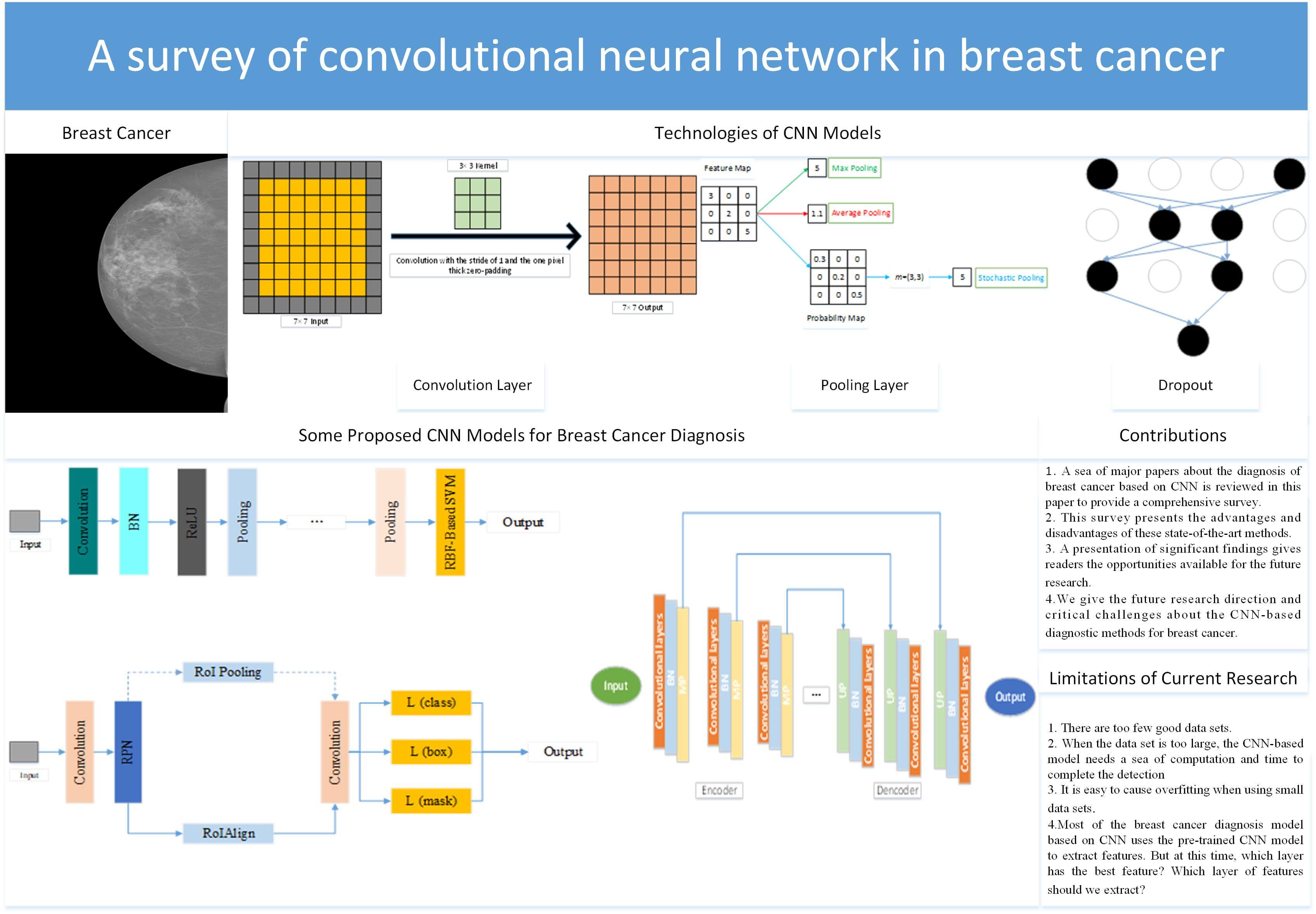
Keywords
Cite This Article
 Copyright © 2023 The Author(s). Published by Tech Science Press.
Copyright © 2023 The Author(s). Published by Tech Science Press.This work is licensed under a Creative Commons Attribution 4.0 International License , which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.


 Submit a Paper
Submit a Paper Propose a Special lssue
Propose a Special lssue View Full Text
View Full Text Download PDF
Download PDF Downloads
Downloads
 Citation Tools
Citation Tools