 Open Access
Open Access
ARTICLE
Co-expression of CCR7 and H3K9me3 identifies aggressive B-cell lymphoma with bone marrow infiltration and poor prognosis
1 Department of Hematology, The Second Affiliated Hospital of Anhui Medical University, Hefei, China
2 Department of Pathology, The First Affiliated Hospital of Anhui Medical University, Hefei, China
3 Department of Pathogen Biology, School of Basic Medicine, Tongji Medical College and State Key Laboratory for Diagnosis and Treatment of Severe Zoonostic Infectious Disease, Huazhong University of Science and Technology, Wuhan, China
* Corresponding Authors: Yang Wan. Email: ; Jiyu Wang. Email:
; Qianshan Tao. Email:
,
These authors contributed equally to this work
European Cytokine Network 2026, 37(1), 25-39. https://doi.org/10.32604/ecn.2026.077875
Received 18 December 2025; Accepted 05 March 2026; Issue published 13 April 2026
Abstract
Objectives: B-cell lymphoma exhibits significant clinical heterogeneity, necessitating improved biomarkers for risk stratification. C-C chemokine receptor 7 (CCR7) and trimethylation of histone H3 lysine 9 (H3K9me3) are implicated in cellular senescence and tumor invasion. While the clinical significance of their co-expression in lymphomagenesis remains unclear. This study aims to define the expression profiles of CCR7 and H3K9me3 in B-cell lymphoma, explore their correlation with aggressive clinical indicators, and evaluate their combined prognostic value. Methods: The expression of CCR7 and H3K9me3 in tumor tissues from B-cell lymphoma patients was analyzed by immunohistochemical (IHC) double-staining. The mechanistic association between the two was verified by co-immunoprecipitation assays and Western blot (WB) experiments detecting changes in cellular H3K9me3 levels following CCR7 ligand stimulation. The association between co-expression and patient clinical parameters, tumor burden, and progression-free survival (PFS) was evaluated through correlation analysis, Kaplan-Meier survival curves, and Cox regression analysis. Results: H3K9me3 expression was predominantly nuclear, whereas CCR7 was expressed on the cell membrane. Both markers were significantly upregulated in aggressive lymphomas and positively correlated with LDH, β2-microglobulin, and neutrophil percentage. An interaction between CCR7 and H3K9me3 could be demonstrated in that CCR7 ligand stimulation resulted in an upregulation of H3K9me3 expression. Enhanced H3K9me3 expression was associated with bone marrow infiltration. High expression of CCR7 was associated with poorer progression-free survival (PFS), whereas high expression of both CCR7 and H3K9me3 identified patients with the worst prognosis. Univariate and multivariate Cox regression analysis indicated that the combined expression was a potential prognostic biomarker for B-cell lymphoma. Conclusion: Co-elevated CCR7 and H3K9me3 expression defines a high-risk B-cell lymphoma subgroup with high tumor burden, bone marrow infiltration, and poor prognosis, highlighting their potential as biomarkers for risk stratification and candidate therapeutic targeting.Keywords
Supplementary Material
Supplementary Material FileB-cell lymphoma is a group of heterogeneous hematological malignancies, and diffuse large B-cell lymphoma (DLBCL) is the most common type [1–3]. The World Health Organization (WHO) classifies B-cell lymphoma at multiple levels based on histomorphology, immunophenotype, molecular features, cell of origin, and genetic abnormalities. Yet distinct B-cell lymphoma subtypes exhibit marked heterogeneity [4,5]. Therefore, identifying biomarkers that can complement existing classifications, enable more precise prognostic stratification, and guide therapeutic decision-making remains an urgent challenge in this field [6].
C-C chemokine receptor 7 (CCR7), a lymphocyte-specific G protein-coupled receptor [7], which plays a central role in directing lymphocyte migration and homing to secondary lymphoid organs, guiding cells into the T-cell zones of lymph nodes through interactions with its ligands C-C motif chemokine ligand 19 (CCL19) and C-C motif chemokine ligand 21 (CCL21) [8–12]. In various malignancies, including hematologic cancers such as chronic lymphocytic leukemia (CLL), high CCR7 expression correlates with increased aggressiveness, lymph node metastasis, and poor prognosis [13]. Our previous studies link CCR7 to cellular senescence, suggesting that its dysregulation may enhance stem-like properties and chemoresistance in lymphoma cells, ultimately associating with unfavorable outcomes [10,11,14]. This senescence-related mechanism might facilitate tumor cell evasion from growth arrest while promoting invasive potential [14].
Aberrant epigenetic regulation, particularly histone modification, is another hallmark of cancer that influences chromatin structure and gene expression [15,16]. Notably, trimethylation of lysine 9 on histone H3 (H3K9me3), a classical marker of transcriptionally silent heterochromatin, is frequently dysregulated in tumors [17]. Altered H3K9 methylation can drive malignant progression by silencing tumor suppressor genes or activating oncogenes [18]. In DLBCL, while the prognostic role of marks like Enhancer of Zeste Homolog 2 (EZH2)-mediated trimethylation of lysine 27 on histone H3 (H3K27me3) has been explored [19], the clinical relevance of H3K9me3 remains less clear. Importantly, H3K9me3 itself has been implicated in promoting cellular senescence [20], suggesting a potential functional convergence with CCR7 in modulating lymphoma biology.
We hypothesized that the co-expression pattern of CCR7 and H3K9me3 might identify a distinct, high-risk subgroup of B-cell lymphoma characterized by bone marrow infiltration and inferior prognosis [5,21,22]. This study aimed to delineate the expression profiles of CCR7 and H3K9me3 in B-cell lymphoma, investigate their associations with clinical indicators of aggressive disease, and evaluate their combined prognostic value. Our findings are expected to provide novel insights for risk stratification and highlight potential candidate biomarkers and therapeutic targets for high-risk B-cell lymphoma.
From May 2020 to September 2025, a total of 26 B-cell lymphoma patients were enrolled in this study at the Second Affiliated Hospital of Anhui Medical University. The diagnosis and staging of all patients with B-cell lymphoma were performed in accordance with the criteria of the Lugano International Conference on Malignant Lymphomas. The specific diagnostic results for each patient are presented in Table S1 (Supplementary Material). This study was approved by the Institutional Review Board (IRB) Institutional of the Second Hospital of Anhui Medical University (No. 20190034). All methods were performed in accordance with relevant guidelines and regulations. All patients enrolled in the study have signed informed consent. All participants provided informed consent in line with the Declaration of Helsinki.
2.2 Diagnosis of Bone Marrow Infiltration
Bone marrow infiltration was determined by morphological assessment of bone marrow aspirate and/or trephine biopsy slides. All cases were reviewed independently by two experienced hematopathologists, who were blinded to each other’s findings and the patients’ clinical information. Any discrepancies were resolved by consensus review to reach a final diagnosis.
The human DLBCL cell lines SU-DHL-4 were obtained from the Shanghai Cell Bank of the Chinese Academy of Sciences. Cells were cultured in RPMI-1640 medium (Hyclone, USA, SH30809.01) supplemented with 10% fetal bovine serum (Lonsera, Uruguay, S711-001S) and maintained at 37°C in a humidified atmosphere containing 5% CO2. Mycoplasma detection was performed routinely for all cell lines, and no contamination was observed. Cell line identity was verified by short tandem repeat (STR) analysis.
2.4 Immunohistochemical Double-Staining
Paraffin-embedded lymph node tissues were sectioned at 3 μm. After deparaffinization and rehydration, antigen retrieval was performed in citrate buffer, and endogenous peroxidase activity was blocked with 0.3% hydrogen peroxide solution. For H3K9me3 staining (HRP-DAB detection system), sections were incubated with the primary anti-histone H3 (trimethyl K9) antibody (ChIP Grade, 1:400 dilution). After rinsing, the HRP-conjugated secondary antibody was applied for incubation. Sections were then rinsed with phosphate-buffered saline (PBS) to remove excess solution, and 100 μL of freshly prepared DAB chromogen solution (1:20 dilution) was added dropwise; color development was monitored microscopically for 5–8 min. Non-specific binding sites were subsequently blocked with protein blocking reagent to eliminate cross-reactivity between the two detection systems. For CCR7 staining (AP-Fast Red detection system), sections were incubated with the primary CCR7 rabbit monoclonal antibody, followed by rinsing and incubation with AP-conjugated secondary antibody. Ready-to-use Fast Red chromogen solution (1:20 dilution) was then used for staining for 1–3 min, and the reaction was terminated by rinsing with distilled water. Hematoxylin staining solution was added dropwise for counterstaining for 1–2 min, followed by washing with purified water to return to blue. Aqueous mounting medium was added dropwise to fully cover the tissue, which was dried at 50–60°C; neutral balsam was then applied dropwise for permanent mounting prior to microscopic observation. All antibodies and reagents used in this study are listed in Table S2 (Supplementary Material).
Appropriate negative controls were included for IHC staining. For each primary antibody, an isotype-matched control antibody was applied in parallel at the same concentration to exclude non-specific binding. In addition, omission of the primary antibody was performed as an additional negative control to confirm the specificity of staining. For double-staining experiments, single-stain controls were performed in parallel for each marker to confirm the absence of signal bleed-through and chromogen interference. Each chromogen was tested individually under the same imaging conditions to ensure no cross-signal between channels.
2.5 Staining Evaluation and Scoring
CCR7 positive staining was observed in the cell membrane, and positive staining for H3K9me3 was observed in the cell nucleus. The average count of 5 random high-power fields (HPF) (×400) for each slice was obtained by an Olympus BX51TF microscope (Olympus, Japan). Areas with obvious necrosis, hemorrhage, tissue folding, or non-neoplastic tissue infiltration were excluded. ensuring coverage of different spatial regions of the tumor to minimize sampling bias caused by intratumoral heterogeneity. All IHC slides were independently scored by two hematological pathologists with more than 10 years of clinical experience, who were blinded to patients’ clinical information (age, gender, disease subtype, clinical stage, treatment outcome, and follow-up data) and experimental grouping. A calibration session was conducted using 5 randomly selected slides before formal scoring to unify the interpretation criteria for staining intensity and positive cell proportion. After scoring, the Kappa value was calculated to assess inter-observer consistency. The immunohistochemistry scoring adopts a semi-quantitative interpretation method, where the expression levels of two antigens are scored separately. Quantification was based on the proportion of positive cells and staining intensity, in accordance with general IHC standards. Staining intensity was graded as follows: for DAB staining (H3K9me3), 0 (colorless), 1 (light brown), 2 (brownish yellow), 3 (dark brown); for Fast Red staining (CCR7), 0 (colorless), 1 (light red), 2 (bright red), 3 (dark red). For staining range, the percentage of positively stained cells relative to the total cell number was recorded under 5 randomly selected high-power fields (400×), with scores assigned as 0 (0%), 1 (0–25%), 2 (25–50%), 3 (50–75%), and 4 (75–100%). The expression level of a single antigen was calculated as the product of the staining range score and the staining intensity score. Score 1 denoted H3K9me3 expression, Score 2 denoted CCR7 expression, and Score 3 denoted the co-expression of H3K9me3 and CCR7.
2.6 Co-Immunoprecipitation (Co-IP)
To verify the interaction between CCR7 and H3K9me3, CoIP assay was performed from two opposite perspectives using specific antibodies against CCR7 and H3K9me3, respectively. SU-DHL-4 cells were harvested and rinsed with ice-cold PBS. Total protein was extracted using radioimmunoprecipitation assay (RIPA) lysis buffer supplemented with phenylmethanesulfonyl fluoride (PMSF) at a 1:100 dilution. The cell lysates were clarified by centrifugation at 12,000× g for 10 min at 4°C, and the supernatants were collected. For the immunoprecipitation assay, two groups of lysates were set up: one group was incubated with Protein A+G Magnetic Beads (BeyoMag™, China) in the presence of specific antibody against CCR7, and the other group was incubated with the same magnetic beads and specific antibody against H3K9me3, both at 4°C for 6 h with gentle rotation. Non-specific IgG was used as a negative control in both groups. Following incubation, the immunocomplexes were magnetically separated and washed three times with RIPA buffer to remove non-specific binding. The bound proteins were eluted by boiling in SDS-PAGE sample loading buffer at 100°C for 10 min. Finally, the immunoprecipitates and whole cell lysates (Input) were subjected to WB analysis to detect the interacting protein (H3K9me3 for CCR7 immunoprecipitation and CCR7 for H3K9me3 immunoprecipitation).
2.7 CCR7 Ligand Stimulation Assay
To detect the expression level of H3K9me3 in SU-DHL-4 cells after CCR7 stimulation by its ligand, CCL21, WB assay was performed. SU-DHL-4 cells were seeded and cultured to an appropriate confluence, then starved in serum-free medium for 12 h. The cells were divided into three groups: the experimental group treated with CCL21 at the appropriate concentration, the first control group (untreated cells), and the second control group treated with diluent without CCL21. After incubation for 24 h, total protein was extracted using RIPA lysis buffer supplemented with 1:100 diluted PMSF. The protein samples were subjected to SDS-PAGE electrophoresis, transferred to PVDF membrane, and detected by WB using specific antibody against H3K9me3. GAPDH was used as an internal reference.
Statistical analyses were conducted using SPSS V.29.0.2 software (IBM Corp., Armonk, NY, USA) and GraphPad Prism V.10.4.1 (GraphPad Software, La Jolla, CA, USA). The normality of quantitative data was assessed using the Shapiro–Wilk test. All quantitative data are presented as the Mean ± Standard Deviation (SD). The chi-square test was applied for categorical variables, while the two-sided Student’s t-test was used for quantitative variables with a normal distribution. The relationship between BMI and clinical parameters was assessed using either the Spearman or Pearson correlation test, depending on data distribution characteristics. Progression-free survival (PFS) was defined as the time from diagnosis until disease progression, relapse, death from any cause, or the last follow-up. Survival estimates were generated using the Kaplan–Meier method, with differences evaluated by the stratified log-rank test. Univariate hazard ratios (HR) with 95% confidence intervals (CI) were calculated using the Cox proportional hazards regression model. A p value < 0.05 was considered statistically significant. All experiments were performed in triplicate (n = 3 independent replications).
3.1 Characteristics of B-Cell Lymphoma Patients in Our Cohort
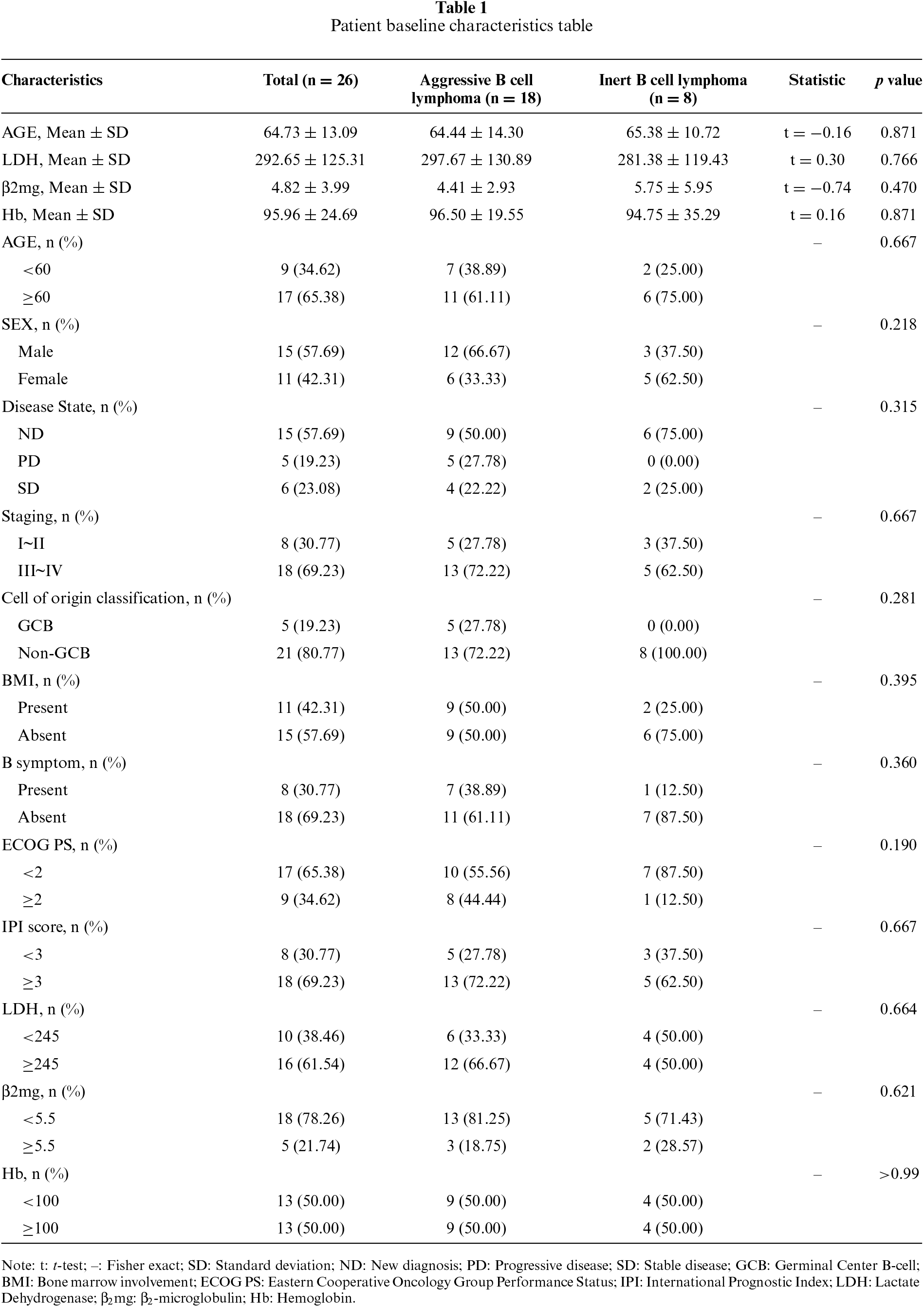
This study included 26 patients with B-cell lymphoma, with a median age of 64.73 years (range: 37–86 years). Among them, 8 patients (30.77%) had indolent lymphoma, and 18 patients (69.23%) had aggressive lymphoma. 11 patients (42.31%) had bone marrow involvement. 8 patients (30.77%) were classified as stage I–II, and 18 patients (69.23%) as stage III–IV. According to the IPI scoring system, 8 patients (30.77%) were classified as low-risk, and 18 patients (69.23%) as high-risk. Additional clinical characteristics are summarized in table 1.
3.2 High Expression of H3K9me3 and CCR7 in Aggressive B-Cell Lymphoma Tissues
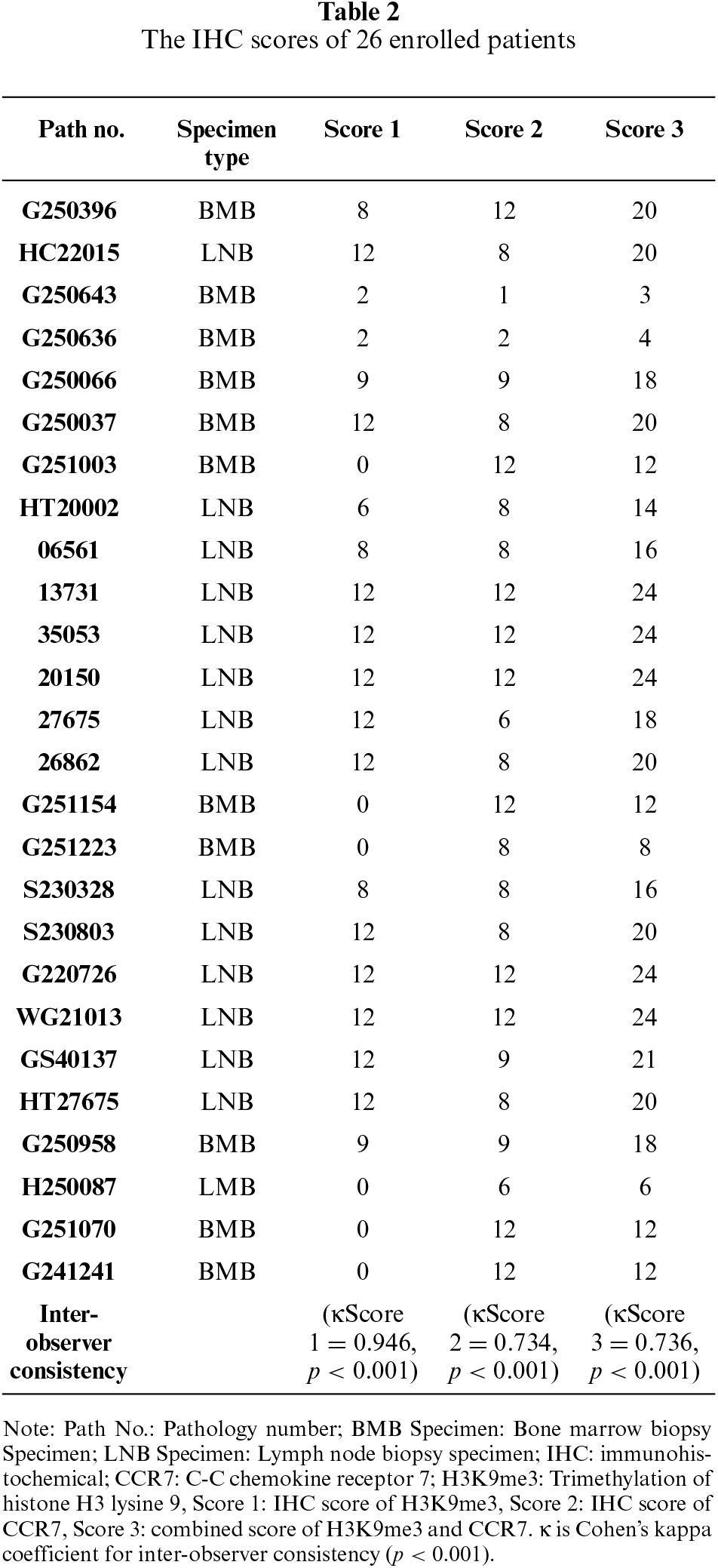
To investigate the expression patterns of H3K9me3 and CCR7 in indolent vs aggressive lymphomas, we collected bone marrow biopsy and lymph node specimens from B cell lymphoma patients and performed IHC double staining. The results showed that H3K9me3 was predominantly localized in the nucleus, whereas CCR7 was mainly expressed on the cell membrane (figures 1A–D and 2A–D). Semi-quantitative IHC analysis further demonstrated that H3K9me3 expression was markedly upregulated in the bone marrow of aggressive lymphomas, whereas it was nearly undetectable in indolent lymphoma tissues (figure 1C,D). Semi-quantitative immunohistochemical analysis was performed by two independent senior pathologists, who were blinded to the clinical characteristics of the patients. Any discrepancies in scoring were resolved through consensus discussion to yield a final unified score for each case. Inter-observer agreement was assessed using Cohen’s kappa coefficient, which demonstrated excellent consistency for H3K9me3 (κScore 1 = 0.946, p < 0.001) and substantial consistency for CCR7 (κScore 2 = 0.734, p < 0.001) as well as the combined score (κScore 3 = 0.736, p < 0.001) (table 2). Statistical comparison using the Mann-Whitney U test confirmed a significant difference between the two groups (p = 0.0030) (table 2). CCR7, in contrast, was expressed in both groups, but there was no significant difference (figure 1C,D). Similar results were also observed in the lymph node specimens (figure 2C,D).
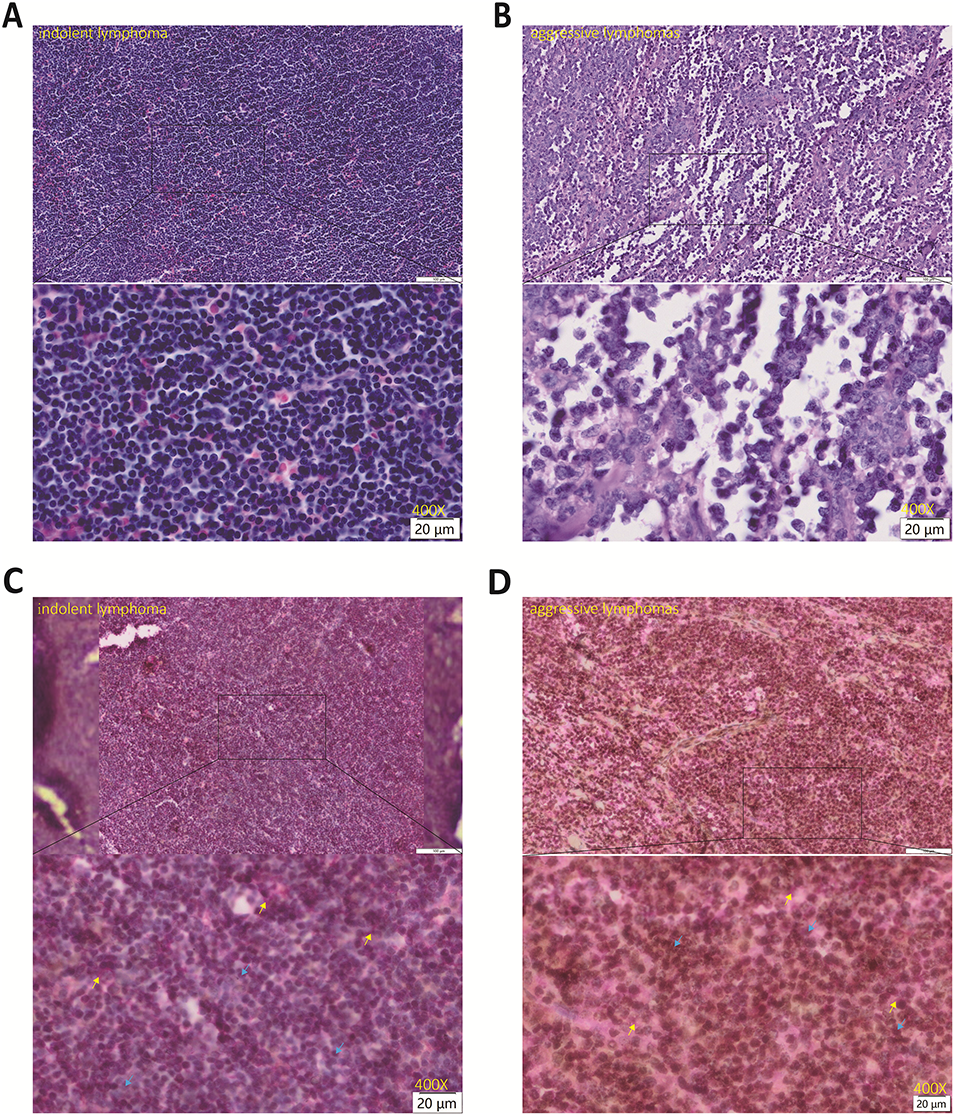
Figure 1: Hematoxylin and Eosin (HE) staining and IHC double staining results of H3K9me3 and CCR7 in lymph node of patients with lymphoma (Original magnification: ×400; scale bar = 20 μm). H3K9me3 (brown staining) was predominantly localized in the cell nucleus, while CCR7 (red staining) was mainly expressed on the cell membrane. (A) HE staining of indolent lymphoma tissues; (B) HE staining of aggressive lymphoma tissues; (C) IHC double staining of H3K9me3 and CCR7 in indolent lymphoma tissues; (D) IHC double staining of H3K9me3 and CCR7 in aggressive lymphoma tissues. H3K9me3 was indicated by blue arrows and CCR7 was indicated by yellow arrows. Positive signals were determined according to the instructions of the staining kit. HE: Hematoxylin and Eosin; IHC: immunohistochemical; CCR7: C-C chemokine receptor 7; H3K9me3: Trimethylation of histone H3 lysine 9.
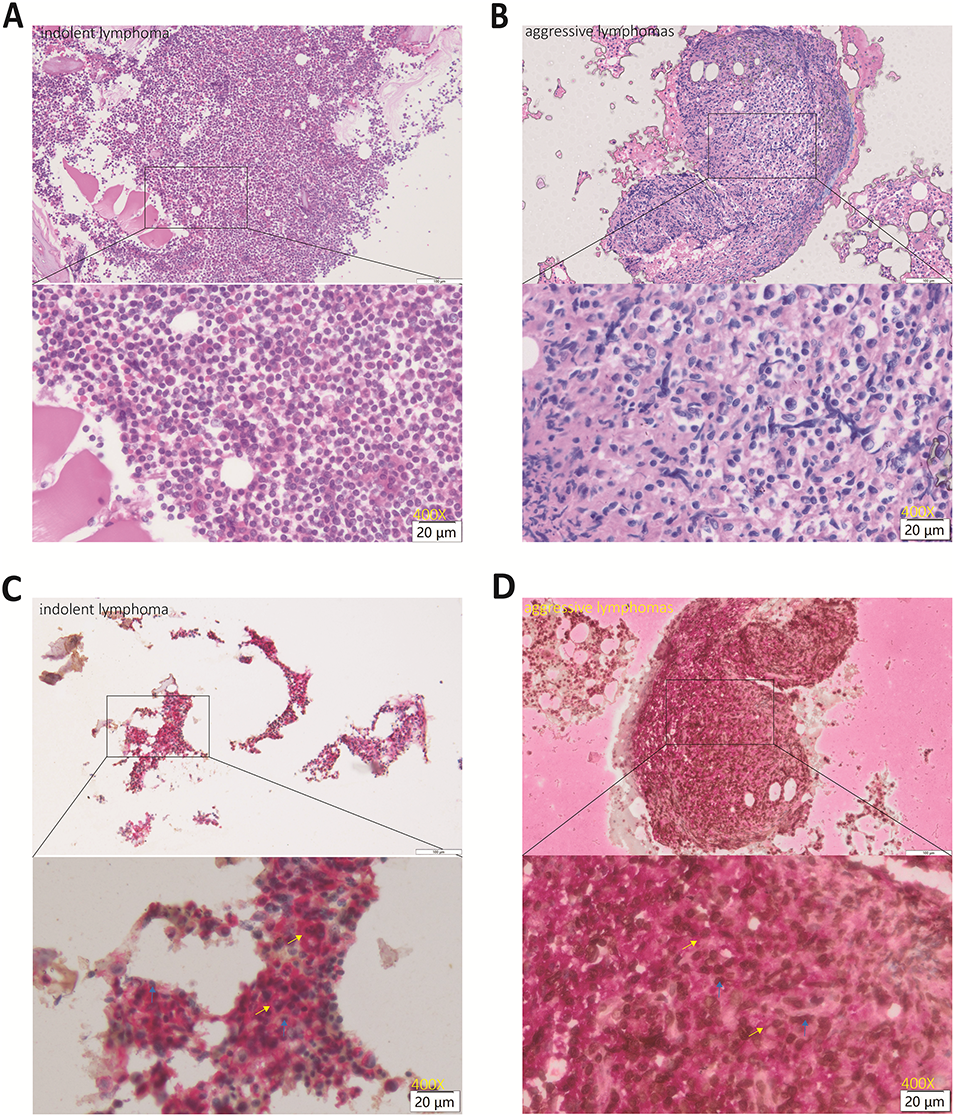
Figure 2: HE staining and IHC double staining results of H3K9me3 and CCR7 in bone marrow tissues of patients with lymphoma (Original magnification: ×400; scale bar = 20 μm). H3K9me3 (brown staining) was predominantly localized in the cell nucleus, while CCR7 (red staining) was mainly expressed on the cell membrane. (A) HE staining of indolent lymphoma tissues; (B) HE staining of aggressive lymphoma tissues; (C) IHC double staining of H3K9me3 and CCR7 in indolent lymphoma tissues; (D) IHC double staining of H3K9me3 and CCR7 in aggressive lymphoma tissues. H3K9me3 was indicated by blue arrows, and CCR7 was indicated by yellow arrows. Positive signals were determined according to the instructions of the staining kit. HE: Hematoxylin and Eosin; IHC: immunohistochemical; CCR7: C-C chemokine receptor 7; H3K9me3: Trimethylation of histone H3 lysine 9.
3.3 CCR7 and H3K9me3 Have a Direct Protein Interaction and CCR7 Ligand Stimulation Upregulates H3K9me3 Expression
To explore the molecular association between CCR7 and H3K9me3, co-immunoprecipitation (Co-IP) assays were performed using the human diffuse large B-cell lymphoma (DLBCL) SU-DHL-4 cell line, and the interaction between the two molecules was verified by Western blot (WB) technology. The results showed that when the cell lysate was immunoprecipitated with a specific anti-CCR7 antibody, the target band of H3K9me3 was clearly detected in the immunoprecipitate; conversely, CCR7 was also successfully enriched when immunoprecipitation was performed with a specific anti-H3K9me3 antibody (figure 3A). No corresponding target bands were detected in the non-specific immunoglobulin G (IgG) negative control group, while clear bands of CCR7 and H3K9me3 proteins were observed in the Input loading control group, confirming the reliability of the experimental results. These findings indicated that CCR7 and H3K9me3 have a direct protein-protein interaction in DLBCL cells.
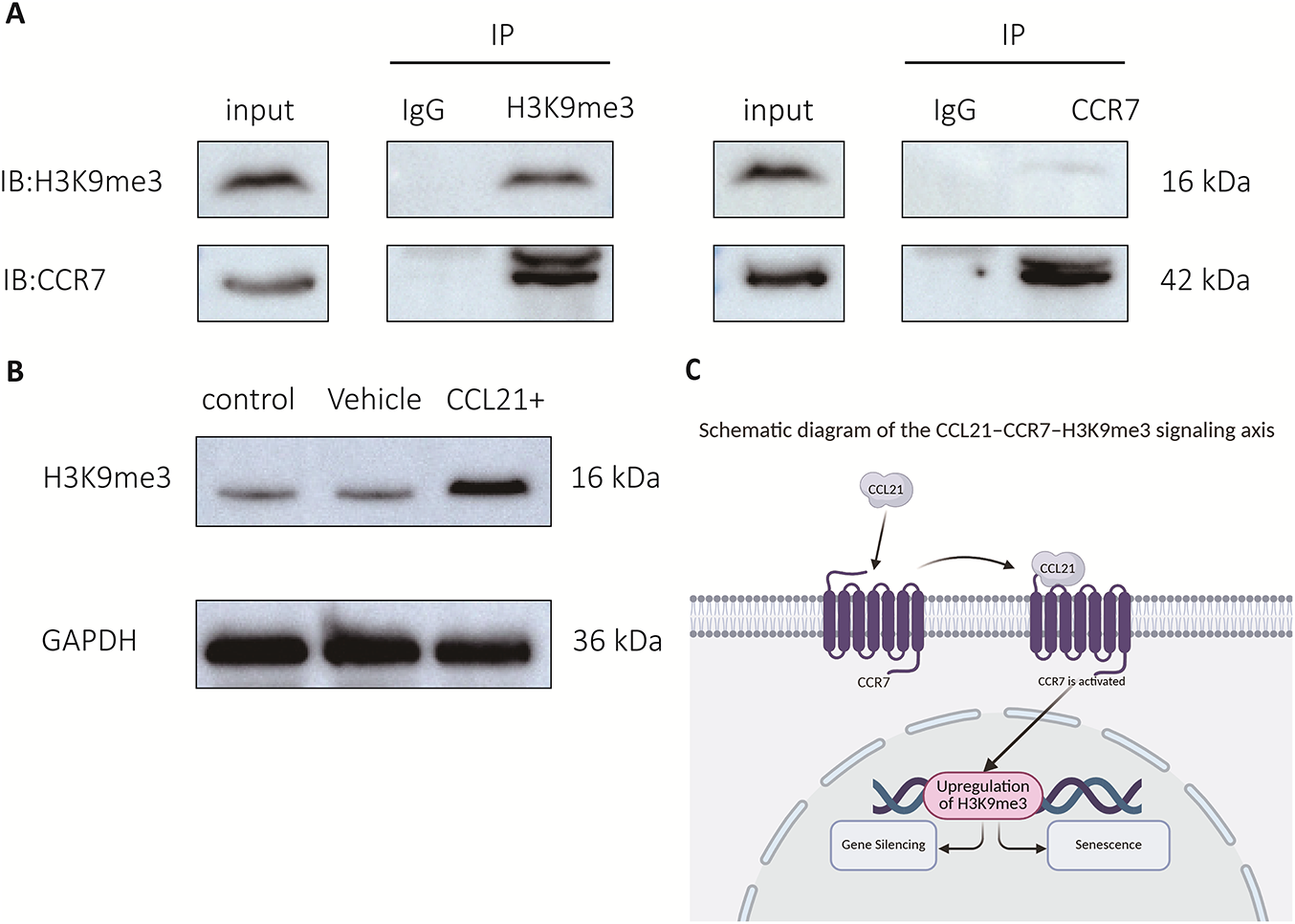
Figure 3: Verification results of the interaction between CCR7 and H3K9me3 by Co-IP (WB) and Changes in H3K9me3 expression level in SU-DHL-4 cells after CCL21 stimulation (WB). (A) Input, total cellular protein loading control; IP-CCR7, immunoprecipitation group with anti-CCR7 antibody; IP-H3K9me3, immunoprecipitation group with anti-H3K9me3 antibody; IgG, non-specific immunoglobulin G negative control. (B) Control, untreated control group; Vehicle, blank control group with diluent only; CCL21, CCR7 ligand stimulation group; GAPDH, internal reference protein. (C) Extracellular CCL21 ligand binds to and activates its receptor CCR7 on the cell membrane. Activated CCR7 signaling promotes the upregulation of H3K9me3 in the nucleus, which further induces gene silencing and ultimately leads to cellular senescence. CCR7: C-C chemokine receptor 7; H3K9me3: Trimethylation of histone H3 lysine 9; Co-IP: Co-Immunoprecipitation; CCL21: C-C motif chemokine ligand 21; IgG: Immunoglobulin G; GAPDH: Glyceraldehyde-3-phosphate dehydrogenase; IB: Immunoblotting; IP: Immunoprecipitation.
To further clarify the functional regulatory relationship between CCR7 and H3K9me3, WB technology was used to detect the changes in the expression level of H3K9me3 in SU-DHL-4 cells after stimulation with CCL21, the ligand of CCR7. The results showed that the protein expression level of H3K9me3 in the CCL21 stimulation group was significantly upregulated compared with the two control groups (figure 3B). Glyceraldehyde-3-phosphate dehydrogenase (GAPDH) was used as the internal reference protein, and its band intensity was uniform in all groups, excluding the influence of protein loading differences on the experimental results. This result indicated that the CCR7 signaling pathway can be activated by its ligand and positively regulates the expression of H3K9me3, suggesting a functional synergy between CCR7 and H3K9me3 in DLBCL cells. Based on these findings, we proposed a CCL21-CCR7-H3K9me3 regulatory axis (figure 3C), which provides a cellular and molecular experimental basis for their co-expression characteristics in clinically aggressive B-cell lymphoma tissues.
3.4 The Expression Levels of H3K9me3 and CCR7 Are Positively Correlated with Tumor Burden
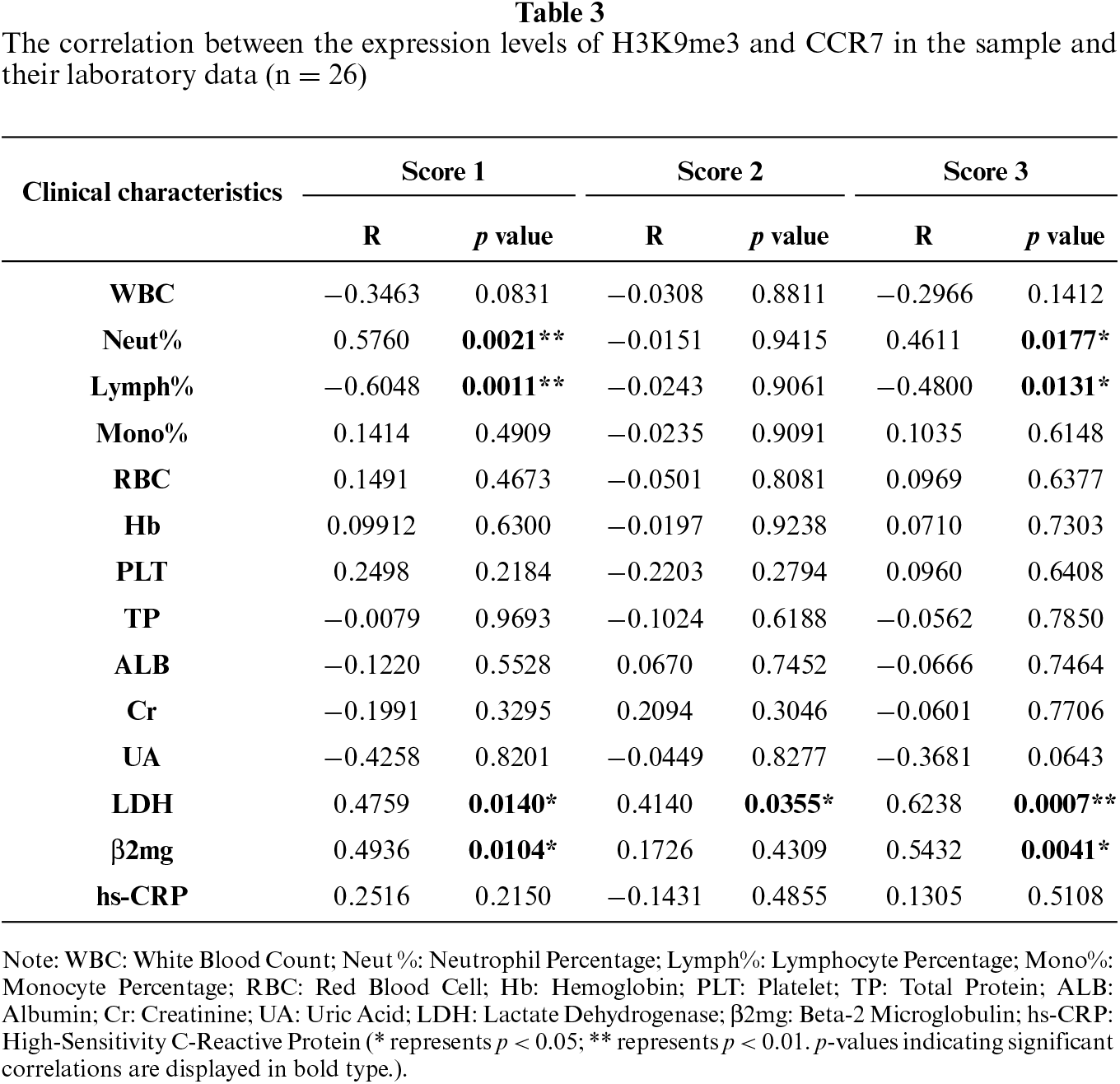
We further analyzed the correlations between the expression levels of H3K9me3 and CCR7 in pathological tissues and the clinical data of patients. The results showed that H3K9me3 expression was positively correlated with serum LDH (R = 0.4759, p = 0.0140), β2-microglobulin (R = 0.4936, p = 0.0104), and neutrophil percentage (R = 0.5760, p = 0.0021), but negatively correlated with lymphocyte percentage (R = −0.6048, p = 0.0011) (figure 4A–D; table 3). CCR7 expression was positively correlated with LDH (R = 0.4140, p = 0.0355) (figure 4E; table 3). The combined score of the two markers was positively correlated with LDH (R = 0.6238, p = 0.0007), β2-microglobulin (R = 0.5432, p = 0.0041), and neutrophil percentage (R = 0.4611, p = 0.0177) (figure 4F–H; table 3).
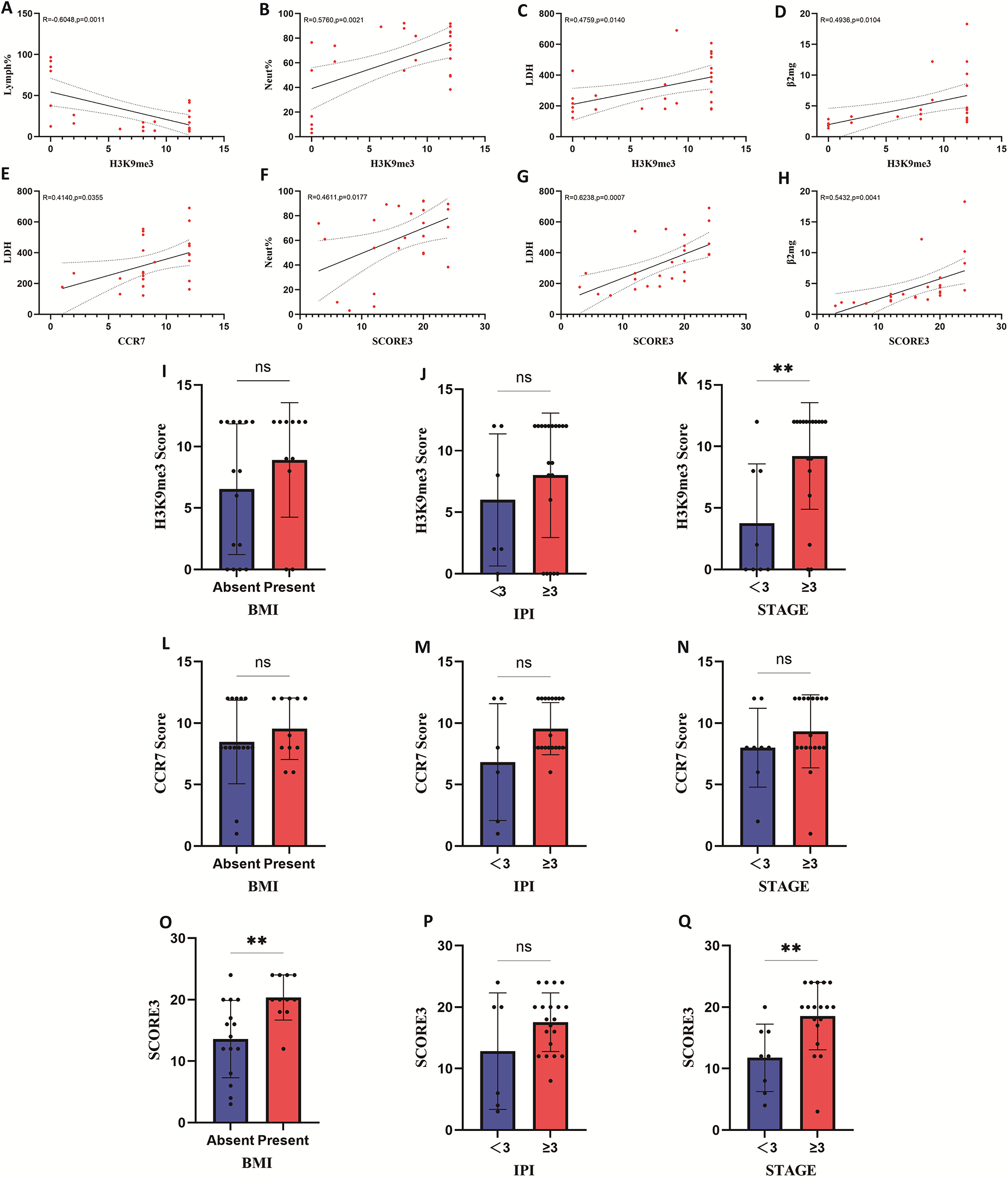
Figure 4: Correlation of H3K9me3, CCR7 and their combined expression with clinical parameters and group differences in clinical stratification. (A–D) Correlation between H3K9me3 expression level and lymphocyte percentage, neutrophil percentage, lactate dehydrogenase (LDH), and β-2 Microglobulin (β-2mg). (E) Correlation between CCR7 expression level and LDH. (F–H) Correlation between the co-expression level of H3K9me3 and CCR7 and neutrophil percentage, LDH, and β-2mg. (I–K) Correlation between H3K9me3 expression in patients and different bone marrow infiltration (BMI) statuses, different IPI scores, and different disease stages. (L–N) Correlation between CCR7 expression in patients and different BMI statuses, different IPI scores, and different disease stages. (O–Q) Correlation between the co-expression of H3K9me3 and CCR7 in patients and different BMI statuses, different IPI scores, and different disease stages. CCR7: C-C chemokine receptor 7; H3K9me3: Trimethylation of histone H3 lysine 9; LDH: Lactate Dehydrogenase; β-2mg: β2-microglobulin; BMI: Bone marrow involvement; IPI: International Prognostic Index; STAGE: clinical stage. ** represents p < 0.01; ns: not significant.
Additionally, we analyzed the association between H3K9me3/CCR7 expression and clinical pathological characteristics (BMI, IPI score, clinical stage). H3K9me3 expression had no significant correlation with BMI or IPI score (figure 4I,J, p > 0.05), but was significantly associated with clinical stage (figure 4K, p < 0.001). In contrast, CCR7 expression showed no significant correlation with BMI, IPI score, or clinical stage (figure 4L–N, all p > 0.05). Notably, the combined score was significantly associated with BMI (figure 4O, p < 0.001), while no significant correlation was found between the combined score and IPI score (figure 4P, p > 0.05). The combined score was also significantly associated with clinical stage (figure 4Q, p < 0.001).
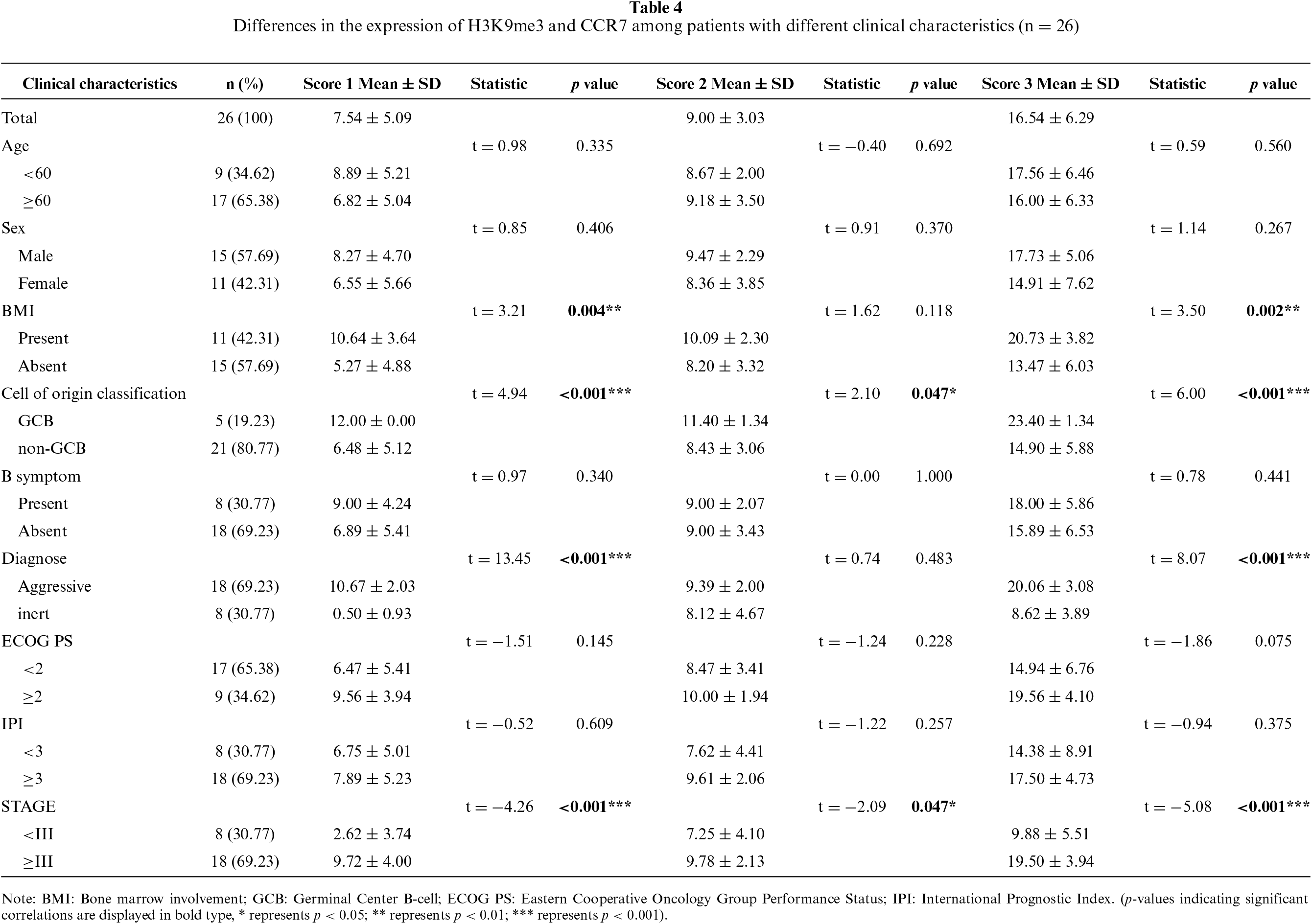
Table 4 investigates the between-group differences in the expression levels of H3K9me3 (Score 1), CCR7 (Score 2), and their combined score (Score 3) across different clinical stratifications in 26 patients with B-cell lymphoma, with no significant differences observed in Score 1, Score 2 and Score 3 relative to patients’ age, gender, B symptoms, Eastern Cooperative Oncology Group performance status (ECOG PS) and International Prognostic Index (IPI) score (all p > 0.05). In contrast, bone marrow infiltration (BMI) was associated with markedly elevated Score 1 (p = 0.004**) and Score 3 (p = 0.002**), while Score 2 showed no statistical difference in this regard (p = 0.118). Aggressive B-cell lymphoma presented with dramatically higher Score 1 and Score 3 compared with indolent subtypes (p < 0.001*** for both), and Score 2 expression did not differ between the two disease types (p = 0.483). Advanced clinical stage (≥III) was correlated with significantly increased Score 1 (p < 0.001***), Score 2 (p = 0.047*) and Score 3 (p < 0.001***), and Germinal Center B-cell (GCB) subtype also exhibited significantly higher expression of the three scores than non-GCB subtype (p < 0.001*** for Score 1 and Score 3, p = 0.047* for Score 2). Based on these findings, we conclude that H3K9me3 and CCR7 may be associated with tumor burden and clinical progression in patients with B-cell lymphoma.
3.5 High Expression of H3K9me3 and CCR7 Is Associated with Poor Prognosis in Patients with B-Cell Lymphoma
The above findings suggest that elevated expression of H3K9me3 and CCR7 may be linked to disease onset and clinical tumor burden. To further explore their relationship with patient survival outcomes, we divided patients into high- and low-expression groups based on the median expression levels and generated Kaplan–Meier survival curves for analysis. The median follow-up time in this study was 27 months (range: 3–82 months). During follow-up, 9 patients (34.6%) experienced disease progression, while no deaths were reported. Kaplan-Meier analysis indicated that patients with high H3K9me3 expression had significantly poorer survival outcomes compared with those in the low-expression group (HR = 2.070, 95%CI: 0.877–71.610, p = 0.033) (figure 5A). Consistently, high CCR7 expression was also linked to markedly inferior survival prognosis in patients (HR = 8.611, 95%CI: 1.051–70.570, p = 0.016) (figure 5B). We additionally developed a composite Score 3 to comprehensively assess the prognostic value of H3K9me3 and CCR7, and stratified patients into high- and low-Score 3 subgroups for survival analysis. The results showed that patients in the high-Score 3 group had significantly worse survival outcomes than those in the low-Score 3 group (HR = 5.003, 95%CI: 0.977–25.630, p = 0.034) (figure 5C). These findings indicate that dual high expression of H3K9me3 and CCR7 may serve as an important prognostic factor in B-cell lymphoma (figure 5).
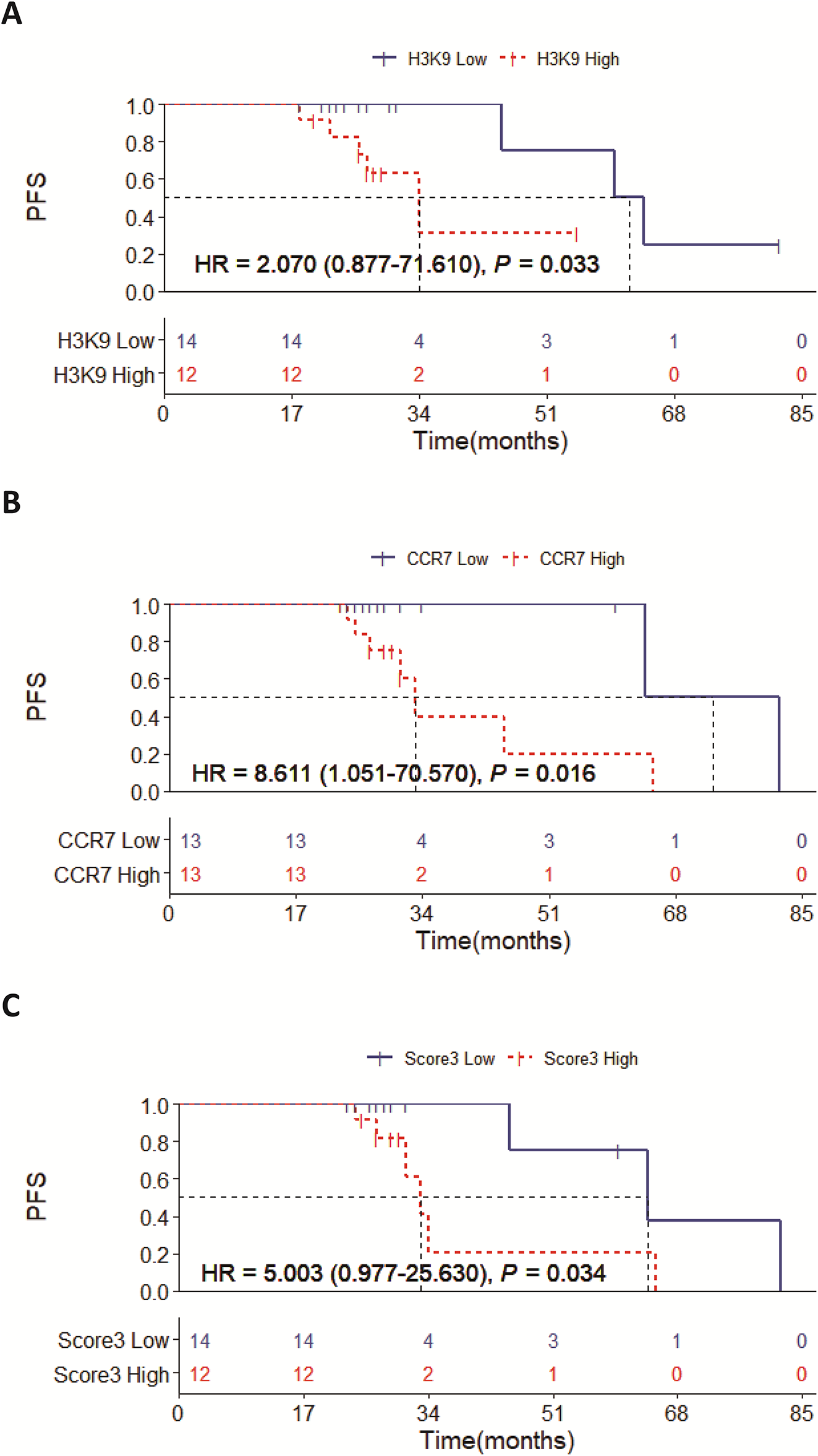
Figure 5: Kaplan-Meier survival curves for PFS. (A) Progression-Free Survival (PFS) of patients with different H3K9me3 expression levels. (B) PFS of patients with different CCR7 expression levels. (C) PFS of patients with different combined expression levels of H3K9me3 and CCR7. PFS: Progression-Free Survival; CCR7: C-C chemokine receptor 7; H3K9me3: Trimethylation of histone H3 lysine 9.
3.6 Combined H3K9me3/CCR7 Score as a Potential Prognostic Biomarker
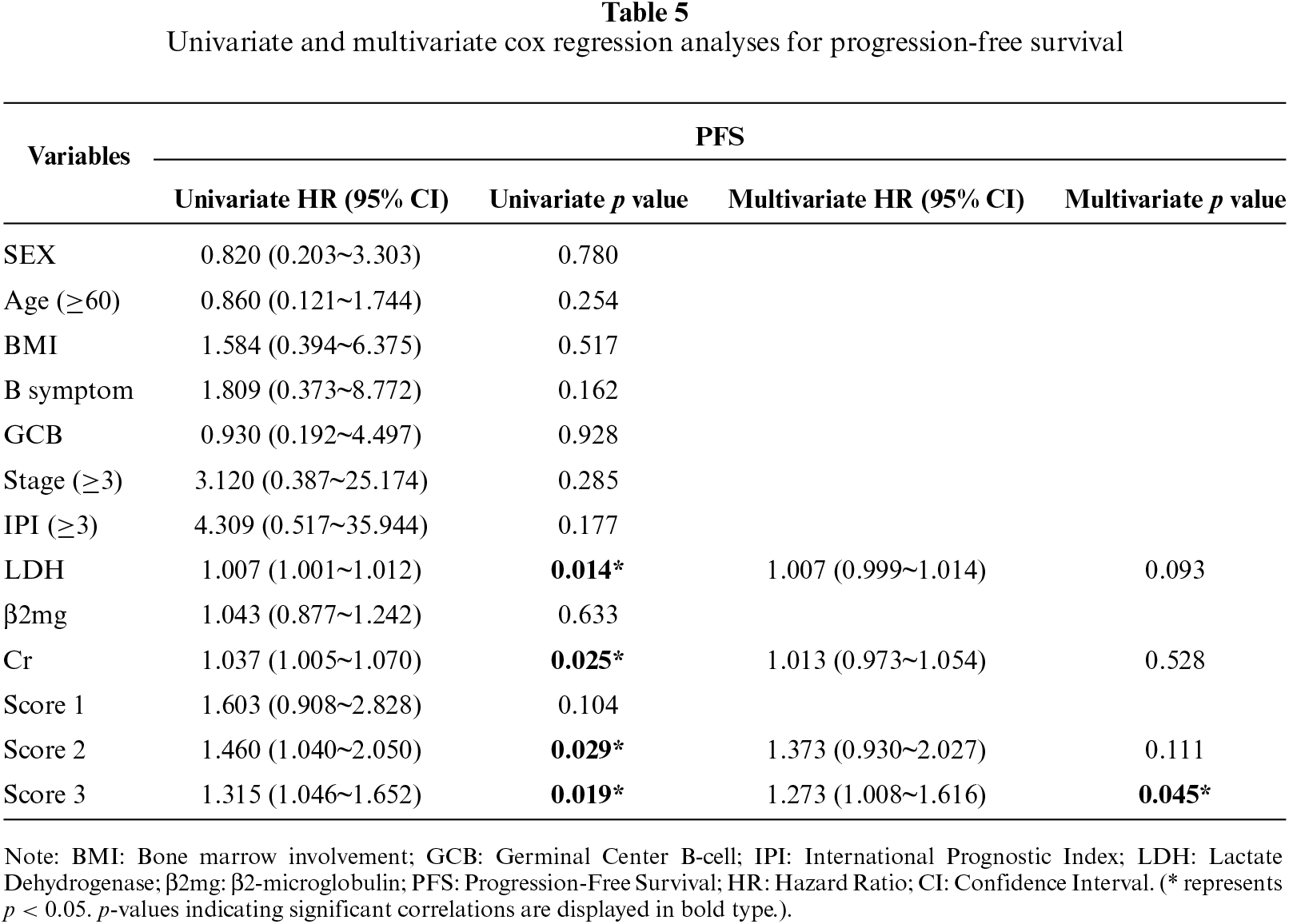
To further clarify the relationship between the expression levels of H3K9me3 and CCR7 and clinical outcomes, we performed univariate and multivariate Cox proportional hazards regression analyses to evaluate the prognostic value of relevant indicators for progression-free survival (PFS) in 26 patients with B-cell lymphoma (table 5). Univariate analysis revealed that lactate dehydrogenase (LDH) (HR = 1.007, 95%CI: 1.001~1.012, p = 0.014), serum creatinine (Cr) (HR = 1.037, 95%CI: 1.005~1.070, p = 0.025), CCR7 IHC score (Score 2) (HR = 1.460, 95%CI: 1.040~2.050, p = 0.029) and the combined H3K9me3/CCR7 IHC score (Score 3) (HR = 1.315, 95%CI: 1.046~1.652, p = 0.019) were significantly correlated with PFS in the enrolled patients. When these statistically significant variables from the univariate analysis were further included in the multivariate Cox regression model, the combined H3K9me3/CCR7 score (Score 3) may serve as a potential prognostic biomarker for unfavorable PFS in B-cell lymphoma patients (HR = 1.273, 95%CI: 1.008~1.616, p = 0.045). Collectively, these findings suggest that the combined expression score of H3K9me3 and CCR7 may act as a potential prognostic biomarker for predicting unfavorable PFS in patients with B-cell lymphoma, which could represent a potential novel prognostic biomarker for this patient population, warranting further validation in larger cohorts.
Bone marrow infiltration by lymphoma cells is a known adverse prognostic factor in B-cell lymphomas [23]. Studies have shown that even minimal bone marrow involvement can significantly affect patient outcomes [24,25]. In this study, through analysis of the clinicopathological characteristics of 26 patients with B-cell lymphoma, we preliminarily hypothesized that the expression patterns and clinical significance of the histone methylation mark H3K9me3 and the chemokine receptor CCR7 in aggressive lymphomas. We note that H3K9me3 was predominantly localized in the nucleus and CCR7 was mainly expressed on the cell membrane [26,27], which are the intrinsic subcellular characteristics of histone modification markers and G protein-coupled chemokine receptors, respectively. This observation is not a mere reflection of the expected antigen distribution, but a scientific validation with specific research implications in the pathological context of B-cell lymphoma. By applying IHC double-staining technology, we for the first time verified the co-expression of the two markers with typical intrinsic localization in the same tumor tissue/cell microenvironment of B-cell lymphoma, which excluded the experimental bias caused by separate detection of single markers and provided a reliable morphological basis for the semi-quantitative scoring and correlation analysis of their expression levels in clinical samples. More importantly, the clear confirmation of their cross-subcellular structure localization (cell membrane vs. nucleus) provided a key clue for our subsequent exploration of the molecular regulatory relationship between the two markers, and further guided the in vitro experiments to verify that the membrane-localized CCR7 can regulate the expression of nuclear-localized H3K9me3 through ligand-activated signaling pathways. This cross-subcellular regulatory interaction is a novel finding of our study, and the accurate verification of subcellular localization is the prerequisite for the discovery of this molecular mechanism. Our key finding is that H3K9me3 and CCR7 exhibit coordinated high expression in the bone marrow of aggressive B-cell lymphomas. Their combined high expression is strongly associated with bone marrow infiltration and high tumor burden, and serves as a potential prognostic biomarker influencing progression-free survival (PFS). This discovery not only suggests that the chemokine axis and epigenetic remodeling may be closely associated with lymphoma progression but also provides clinical evidence supporting the concept that “senescence-associated stemness” contributes to unfavorable outcomes [28].
H3K9me3 (trimethylation of lysine 9 on histone H3) is a classic marker of heterochromatin formation and gene silencing that participates in the assembly of senescence-associated heterochromatin foci (SAHF) [29,30], and its expression pattern and functional role exhibit heterogeneity across different tumor types [31–34]. In hematologic malignancies, H3K9me3 has been identified as a poor prognostic marker in AML patients with high expression correlated with unfavorable survival [35], while its clinical significance and regulatory mechanisms in B-cell lymphoma, especially in relation to bone marrow infiltration, remain largely uncharacterized. In the present study, we demonstrated that H3K9me3 is highly expressed in the bone marrow tissues of aggressive B-cell lymphoma and correlates with increased tumor burden, suggesting that H3K9me3 may promote lymphoma cell migration and bone marrow colonization through epigenetic regulation of invasion-related genes. This epigenetic regulatory role of H3K9me3 in driving tumor cell infiltration is consistent with the general functional role reported in other tumor types [36], and our in vitro experiments further confirmed that H3K9me3 expression is positively regulated by CCR7 signaling in DLBCL cells, providing a direct molecular link between this epigenetic marker and chemokine-mediated bone marrow infiltration. Specifically, H3K9me3-mediated gene silencing may repress the expression of molecules that limit cell migration and infiltration (e.g., adhesion molecules, migration inhibitors), thereby enhancing the ability of lymphoma cells to invade and adapt to the bone marrow microenvironment [36–38]. These findings elucidate the biological basis for the association between H3K9me3 and bone marrow involvement in aggressive B-cell lymphoma, and support the potential of H3K9me3 as an epigenetic biomarker for evaluating the risk of bone marrow infiltration in B-cell lymphoma patients.
Recent studies have shown that CCR7 expression is significantly upregulated during chemotherapy-induced senescence of B-cell lymphoma cells. Wang et al. found in a doxorubicin-induced B-cell lymphoma senescence model that a subset of memory B cells was strongly CCR7-positive, and these cells were significantly increased in senescent cells compared with normally proliferating cells. Further experiments demonstrated that the CCR7 ligand CCL21 promotes lymphoma cell migration and invasion and enhances stem-like phenotypes, suggesting that CCR7 drives the conversion of senescent cells toward a tumor stem-like state [14]. The consolidation of cellular senescence depends on heterochromatin formation, in which trimethylation of lysine 9 on histone H3 (H3K9me3) plays a key role [39]. The Suv39h1 enzyme, mediated by the Rb/E2F axis, deposits H3K9me3 at E2F target genes to form condensed heterochromatin structures, thereby stabilizing G1-phase arrest [40]. Schleich et al. observed in a transgenic mouse lymphoma model that lymphomas with a high “SUVARness” gene expression signature dominated by H3K9me3 exhibited a stronger senescence response to chemotherapy, and high H3K9me3 levels were significantly associated with favorable outcomes in DLBCL patients [40]. In patient cohorts, those with high H3K9me3 expression in tumor tissues showed markedly better long-term survival than the low-expression group, indicating that H3K9me3 serves not only as an epigenetic marker of senescence execution but also as a prognostic indicator [41].
Interestingly, in the aforementioned B-cell lymphoma senescence model, CCR7+ cells co-expressed CD44 and H3K9me3, suggesting that cells expressing CCR7 also exhibit prominent H3K9me3 chromatin features. This finding implies that CCR7 signaling may interact with H3K9me3-mediated heterochromatin homeostasis: CCR7-driven pathways (such as the ARHGAP18/IKBα pathway) could potentially interfere with the consolidation of senescence by H3K9me3, allowing a subset of cells to escape heterochromatin-repressed senescence and regain proliferative capacity. Therefore, a functional link between CCR7 and H3K9me3 can be envisioned: lymphoma cells with high CCR7 expression may acquire stem-like traits by modulating histone demethylase activity, maintaining certain senescence markers while gaining proliferative potential, ultimately promoting relapse and therapy resistance.
From a clinical perspective, CCR7 and H3K9me3 together constitute valuable assessment indicators. Studies have shown that B-cell lymphoma patients with high CCR7 expression are more prone to bone marrow infiltration and have shorter progression-free survival (PFS), with the poorest prognosis observed in patients co-expressing high levels of CCR7 and CXCR4 [42]. On the other hand, high H3K9me3 expression indicates an intact senescence program, favorable treatment response, and is significantly associated with improved prognosis [40]. In this study, high H3K9me3 expression was associated with poor PFS in B-cell lymphoma patients, which is inconsistent with the results of Schleich et al. [40]. This discrepancy may be due to the different study cohorts: Schleich et al. focused on DLBCL patients receiving standard chemoimmunotherapy, while our study included various subtypes of B-cell lymphoma with different treatment regimens, and the sample size was relatively small with limited progression events. In addition, the combined detection of CCR7 and H3K9me3 in our study showed a stronger prognostic value than single-marker detection, which may explain the different prognostic implications of H3K9me3 alone. These findings provide a theoretical basis for individualized monitoring and therapeutic strategies for B-cell lymphoma patients.
Although this study identified the clinical significance and molecular association of CCR7/H3K9me3 co-expression in aggressive B-cell lymphoma, it has certain limitations. This single-center retrospective study with a small sample size and short follow-up only used PFS as the prognostic endpoint, which restricts the generalizability of the results. Due to the limited sample size and only 9 progression events in this cohort, the multivariate Cox regression model was underpowered to fully adjust for established clinical risk factors, including IPI score and disease stage. Besides, semi-quantitative IHC scoring has subjective bias, despite double-blind independent assessment by two senior hematopathologists and robust interobserver consistency, excluding infiltrating CCR7+ T cells via pathological morphology and spatial localization has inherent methodological limitations and cellular experiments with a single cell line failed to clarify the key binding domains and downstream signaling pathways of CCR7 and H3K9me3. The lack of external cohort verification, subgroup analyses and in vivo animal experiments also weakens the evidence for clinical translation, and future studies will address these deficiencies by expanding samples, conducting multicenter investigations and perfecting experimental and clinical validation.
In summary, this study elucidates the aberrant expression patterns of H3K9me3 and CCR7 in aggressive B-cell lymphomas and demonstrates the significant value of their combined assessment in evaluating tumor burden and predicting the risk of bone marrow involvement. The co-elevated expression of H3K9me3 and CCR7 may indicate a malignant clonal subpopulation characterized by high migratory capacity and senescence-associated stem-like traits. Epigenetic interventions or chemokine blockade targeting this subpopulation could provide potential novel therapeutic strategies to improve the prognosis of high-risk lymphoma patients. Although the sample size in this study was relatively small, the statistically significant differences observed suggest the reliability of these findings, warranting further validation in larger cohorts.
In conclusion, this study demonstrated that H3K9me3 and CCR7 are co-elevated in bone marrow tissues of aggressive B-cell lymphomas and are significantly associated with high tumor burden. Their combined high expression serves as a potential prognostic biomarker for progression-free survival. Extracellular CCL21 ligand binds to and activates its receptor CCR7 on the cell membrane. Activated CCR7 signaling promotes the upregulation of H3K9me3 in the nucleus, which further induces gene silencing and ultimately leads to cellular senescence.
Acknowledgement: Not applicable.
Funding Statement: This work was supported by grants from the National Natural Science Foundation of China (No. 82500257) and the Anhui Province Key Project for Clinical Medical Research Translation (202427b10020040).
Author Contributions: The authors confirm contribution to the paper as follows: study conception and design: Jiyu Wang, Qianshan Tao, Yang Wan; Data collection: Jiawen Chen, Keke Huang, Yajie Zhang, Dandan Chen, Furun An, Jinlan Li; analysis and interpretation of results: Jiyu Wang, Jiawen Chen, Zelin Liu, Yanjie Ruan, Ying Pan; draft manuscript preparation: Jiyu Wang, Jiawen Chen, Zelin Liu. All authors reviewed and approved the final version of the manuscript.
Availability of Data and Materials: The data that support the findings of this study are available from the corresponding author, Jiyu Wang, upon reasonable request and with permission from Second Affiliated Hospital of Anhui Medical University.
Ethics Approval: This study was approved by the Institutional Review Board (IRB) Institutional of the Second Hospital of Anhui Medical University (No. 20190034). All methods were performed in accordance with relevant guidelines and regulations. All patients enrolled in the study have signed informed consent. All participants provided informed consent in line with the Declaration of Helsinki.
Conflicts of Interest: The authors declare no conflicts of interest.
Supplementary Materials: The supplementary material is available online at https://www.techscience.com/doi/10.32604/ecn.2026.077875/s1.
Abbreviations
| ALB | Albumin |
| AP | Alkaline phosphatase |
| ARHGAP18 | Rho GTPase-activating protein 18 |
| BMB | Bone marrow biopsy specimen |
| BMI | Bone marrow involvement |
| CLL | Chronic lymphocytic leukemia |
| CCL19 | C-C motif chemokine ligand 19 |
| CCL21 | C-C motif chemokine ligand 21 |
| CCR7 | C-C chemokine receptor 7 |
| CI | Confidence interval |
| Co-IP | Co-Immunoprecipitation |
| Cr | Creatinine |
| CXCR4 | C-X-C chemokine receptor type 4 |
| DAB | 3,3′-Diaminobenzidine |
| DLBCL | Diffuse large B-cell lymphoma |
| ECOG PS | Eastern Cooperative Oncology Group Performance Status |
| ERK1/2 | Extracellular regulated protein kinases 1/2 |
| EZH2 | Enhancer of Zeste Homolog 2 |
| GAPDH | Glyceraldehyde-3-phosphate dehydrogenase |
| GCB | Germinal Center B-cell |
| Hb | Hemoglobin |
| H3K9me3 | Trimethylation of histone H3 lysine 9 |
| H3K27me3 | Trimethylation of histone H3 lysine 27 |
| HE | Hematoxylin and Eosin Staining |
| HIF-1 | Hypoxia-inducible factor 1 alpha |
| HPF | High-power fields |
| HR | Hazard ratio |
| HRP | Horseradish peroxidase |
| hs-CRP | High-Sensitivity C-Reactive Protein |
| IKBα | Inhibitor of nuclear factor kappa-B alpha |
| IP | Immunoprecipitation |
| IPI | International Prognostic Index |
| IRB | Institutional Review Board |
| KDM4B | Lysine demethylase 4B |
| LDH | Lactate Dehydrogenase |
| LNB | Lymph node biopsy specimen |
| Lymph% | Lymphocyte Percentage |
| lnc-Dpf3 | Long non-coding RNA Dpf3 |
| Mono% | Monocyte Percentage |
| ND | New diagnosis |
| Neut% | Neutrophil Percentage |
| PBS | Phosphate-buffered saline |
| PD | Progressive disease |
| PFS | Progression-free survival |
| PLT | Platelet |
| PMSF | Phenylmethanesulfonyl fluoride |
| PVDF | Polyvinylidene fluoride |
| RBC | Red Blood Cell |
| RIPA | Radioimmunoprecipitation assay |
| SAHF | Senescence-associated heterochromatin foci |
| Scr | Serum creatinine |
| SD | Stable disease/Standard Deviation |
| SETDB1 | SET domain bifurcated 1 |
| SETDB2 | SET domain bifurcated 2 |
| SDS-PAGE | Sodium dodecyl sulfate-polyacrylamide gel electrophoresis |
| SUV39H1 | Suppressor of variegation 3-9 homolog 1 |
| TP | Total Protein |
| UA | Uric Acid |
| WB | Western blot |
| WBC | White Blood Count |
| WHO | World Health Organization |
| YEATS2 | YEATS domain containing 2 |
| β2mg | β2-microglobulin |
References
1. Miao Y, Medeiros LJ, Xu-Monette ZY, Li J, Young KH. Dysregulation of cell survival in diffuse large B cell lymphoma: mechanisms and therapeutic targets. Front Oncol 2019;9:107. doi:10.3389/fonc.2019.00107. [Google Scholar] [PubMed] [CrossRef]
2. Sakhawat U, Shehadah A, Mazhar U et al. Cecal involvement of diffuse large B-cell lymphoma: a rare extra-nodal presentation. J Med Cases 2025;16(12):475–86. doi:10.14740/jmc5192. [Google Scholar] [PubMed] [CrossRef]
3. Puneeth HK, Kumar SR, Ananthaneni A, Jakkula A. Physical torment: a predisposition for diffuse B-cell lymphoma. J Oral Maxillofac Pathol 2019;23(4):17. doi:10.4103/jomfp.jomfp_7_18. [Google Scholar] [PubMed] [CrossRef]
4. Harrington F, Greenslade M, Talaulikar D, Corboy G. Genomic characterisation of diffuse large B-cell lymphoma. Pathology 2021;53(3):367–76. doi:10.1016/j.pathol.2020.12.003. [Google Scholar] [PubMed] [CrossRef]
5. Yang W, Tang B, Xu D, Yang W. Lenalidomide regulates the CCL21/CCR7/ERK1/2 axis to inhibit migration and proliferation in diffuse large B-cell lymphoma. Oncol Res 2025;33(1):199–212. doi:10.32604/or.2024.050036. [Google Scholar] [PubMed] [CrossRef]
6. Atallah-Yunes SA, Khurana A, Maurer M. Challenges identifying DLBCL patients with poor outcomes to upfront chemoimmunotherapy and its impact on frontline clinical trials. Leuk Lymphoma 2024;65(4):430–39. doi:10.1080/10428194.2023.2298705. [Google Scholar] [PubMed] [CrossRef]
7. Birkenbach M, Josefsen K, Yalamanchili R, Lenoir G, Kieff E. Epstein-Barr virus-induced genes: first lymphocyte-specific G protein-coupled peptide receptors. J Virol 1993;67(4):2209–20. doi:10.1128/jvi.67.4.2209-2220.1993. [Google Scholar] [PubMed] [CrossRef]
8. Legler DF, Uetz-von Allmen E, Hauser MA. CCR7: roles in cancer cell dissemination, migration and metastasis formation. Int J Biochem Cell Biol 2014;54:78–82. doi:10.1016/j.biocel.2014.07.002. [Google Scholar] [PubMed] [CrossRef]
9. Liu J, Zhang X, Chen K et al. CCR7 chemokine receptor-inducible lnc-Dpf3 restrains dendritic cell migration by inhibiting HIF-1α-mediated glycolysis. Immunity 2019;50(3):600–15.e15. doi:10.1016/j.immuni.2019.01.021. [Google Scholar] [PubMed] [CrossRef]
10. Zhang J, Qiu X. A Mendelian randomization study of the gut microbiota and risk of knee osteoarthritis and the mediating role of immune cells. Sci Rep 2025;15(1):28455. doi:10.1038/s41598-025-14007-x. [Google Scholar] [PubMed] [CrossRef]
11. Lin SY, Yao BY, Hu CJ, Chen HW. Induction of robust immune responses by CpG-ODN-loaded hollow polymeric nanoparticles for antiviral and vaccine applications in chickens. Int J Nanomed 2020;15:3303–18. doi:10.2147/ijn.s241492. [Google Scholar] [PubMed] [CrossRef]
12. Salem A, Alotaibi M, Mroueh R, Basheer HA, Afarinkia K. CCR7 as a therapeutic target in Cancer. Biochim Biophys Acta BBA Rev Cancer 2021;1875(1):188499. doi:10.1016/j.bbcan.2020.188499. [Google Scholar] [PubMed] [CrossRef]
13. Bill CA, Allen CM, Vines CM. C-C chemokine receptor 7 in cancer. Cells 2022;11(4):656. doi:10.3390/cells11040656. [Google Scholar] [PubMed] [CrossRef]
14. Wang J, Tao Q, Huang K et al. Chemotherapy-induced cellular senescence promotes stemness of aggressive B-cell non-Hodgkin’s lymphoma via CCR7/ARHGAP18/IKBα signaling activation. J Immunother Cancer 2025;13(1):e009356. doi:10.1136/jitc-2024-009356. [Google Scholar] [PubMed] [CrossRef]
15. Grubb T, Koochaki P, Matar S et al. The SLC1A1/EAAT3 dicarboxylic amino acid transporter is an epigenetically dysregulated nutrient carrier that sustains oncogenic metabolic programs. Nat Commun 2025;16(1):10012. doi:10.1038/s41467-025-64983-x. [Google Scholar] [PubMed] [CrossRef]
16. Ji K, Chen G, Wang Y, Li Y, Chen J, Feng M. YEATS2: a novel cancer epigenetic reader and potential therapeutic target. Cancer Cell Int 2025;25(1):162. doi:10.1186/s12935-025-03797-9. [Google Scholar] [PubMed] [CrossRef]
17. Padeken J, Methot SP, Gasser SM. Establishment of H3K9-methylated heterochromatin and its functions in tissue differentiation and maintenance. Nat Rev Mol Cell Biol 2022;23(9):623–40. doi:10.1038/s41580-022-00483-w. [Google Scholar] [PubMed] [CrossRef]
18. Saha N, Muntean AG. Insight into the multi-faceted role of the SUV family of H3K9 methyltransferases in carcinogenesis and cancer progression. Biochim Biophys Acta BBA Rev Cancer 2021;1875(1):188498. doi:10.1016/j.bbcan.2020.188498. [Google Scholar] [PubMed] [CrossRef]
19. Oh EJ, Yang WI, Cheong JW, Choi SE, Yoon SO. Diffuse large B-cell lymphoma with histone H3 trimethylation at lysine 27: another poor prognostic phenotype independent of c-Myc/Bcl2 coexpression. Hum Pathol 2014;45(10):2043–50. doi:10.1016/j.humpath.2014.07.002. [Google Scholar] [PubMed] [CrossRef]
20. Park JW, Bae YS. Dephosphorylation of p53 Ser 392 enhances trimethylation of histone H3 Lys 9 via SUV39h1 stabilization in CK2 downregulation-mediated senescence. Mol Cells 2019;42(11):773–82. doi:10.14348/molcells.2019.0018. [Google Scholar] [PubMed] [CrossRef]
21. Zhang Y, Qian S, Wen Q et al. SUV39H1 is a prognosis and immune microenvironment-related biomarker in diffuse large B-cell lymphoma. Clin Transl Oncol 2023;25(8):2438–50. doi:10.1007/s12094-023-03128-2. [Google Scholar] [PubMed] [CrossRef]
22. Chen X, Soma L, Murphy C, Tretiakova M, Naresh KN, Fromm JR. Utility of CCR7 to differentiate classic Hodgkin lymphoma and other B-cell lymphomas by flow cytometry and immunohistochemistry. Am J Clin Pathol 2025;163(2):266–76. doi:10.1093/ajcp/aqae119. [Google Scholar] [PubMed] [CrossRef]
23. Liu Z, Li Q, Liu R et al. Development and validation of novel models based on clonality immunoglobulin gene rearrangement for evaluation of bone marrow involvement and prognostic prediction in patients with diffuse large B-cell Lymphoma: a multicenter retrospective study. Front Immunol 2025;16:1547056. doi:10.3389/fimmu.2025.1547056. [Google Scholar] [PubMed] [CrossRef]
24. Cho YA, Yang WI, Song JW, Min YH, Yoon SO. The prognostic significance of monoclonal immunoglobulin gene rearrangement in conjunction with histologic B-cell aggregates in the bone marrow of patients with diffuse large B-cell lymphoma. Cancer Med 2016;5(6):1066–73. doi:10.1002/cam4.679. [Google Scholar] [PubMed] [CrossRef]
25. Adams HJA, Nievelstein RAJ, Kwee TC. Opportunities and limitations of bone marrow biopsy and bone marrow FDG-PET in lymphoma. Blood Rev 2015;29(6):417–25. doi:10.1016/j.blre.2015.06.003. [Google Scholar] [PubMed] [CrossRef]
26. Shi M, Chen D, Yang D, Liu XY. CCL21-CCR7 promotes the lymph node metastasis of esophageal squamous cell carcinoma by up-regulating MUC1. J Exp Clin Cancer Res 2015;34(1):149. doi:10.1186/s13046-015-0268-9. [Google Scholar] [PubMed] [CrossRef]
27. Lien MY, Tsai HC, Chang AC et al. Chemokine CCL4 induces vascular endothelial growth factor C expression and lymphangiogenesis by miR-195-3p in oral squamous cell carcinoma. Front Immunol 2018;9:412. doi:10.3389/fimmu.2018.00412. [Google Scholar] [PubMed] [CrossRef]
28. Alduaij W, Collinge BJ, Ben-Neriah S et al. Molecular determinants of clinical outcomes in a real-world diffuse large B-cell lymphoma population. Blood 2023;141(20):2493–507. doi:10.1182/blood.2022018248. [Google Scholar] [PubMed] [CrossRef]
29. Mendelsohn AR, Larrick JW. Stem cell depletion by global disorganization of the H3K9me3 epigenetic marker in aging. Rejuvenation Res 2015;18(4):371–5. doi:10.1089/rej.2015.1742. [Google Scholar] [PubMed] [CrossRef]
30. Zhang J, Donahue G, Gilbert MB, Lapidot T, Nicetto D, Zaret KS. Distinct H3K9me3 heterochromatin maintenance dynamics govern different gene programmes and repeats in pluripotent cells. Nat Cell Biol 2024;26(12):2115–28. doi:10.1038/s41556-024-01547-z. [Google Scholar] [PubMed] [CrossRef]
31. Akano I, Hebert JM, Tiedemann RL et al. The SWI/SNF-related protein SMARCA3 is a histone H3K23 ubiquitin ligase that regulates H3K9me3 in cancer. Mol Cell 2025;85(15):2885–99.e8. doi:10.1016/j.molcel.2025.06.020. [Google Scholar] [PubMed] [CrossRef]
32. Hu S, Wang X, Wang T et al. Differential enrichment of H3K9me3 in intrahepatic cholangiocarcinoma. BMC Med Genom 2022;15(1):185. doi:10.1186/s12920-022-01338-1. [Google Scholar] [PubMed] [CrossRef]
33. Zhou M, Li Y, Lin S et al. H3K9me3, H3K36me3, and H4K20me3 expression correlates with patient outcome in esophageal squamous cell carcinoma as epigenetic markers. Dig Dis Sci 2019;64(8):2147–57. doi:10.1007/s10620-019-05529-2. [Google Scholar] [PubMed] [CrossRef]
34. Zhou M, Yan JQ, Chen QX et al. Association of H3K9me3 with breast cancer prognosis by estrogen receptor status. Clin Epigenet 2022;14(1):135. doi:10.1186/s13148-022-01363-y. [Google Scholar] [PubMed] [CrossRef]
35. Monaghan L, Massett ME, Bunschoten RP et al. The emerging role of H3K9me3 as a potential therapeutic target in acute myeloid leukemia. Front Oncol 2019;9:705. doi:10.3389/fonc.2019.00705. [Google Scholar] [PubMed] [CrossRef]
36. Lu C, Yang D, Sabbatini ME et al. Contrasting roles of H3K4me3 and H3K9me3 in regulation of apoptosis and gemcitabine resistance in human pancreatic cancer cells. BMC Cancer 2018;18(1):149. doi:10.1186/s12885-018-4061-y. [Google Scholar] [PubMed] [CrossRef]
37. Becker JS, Nicetto D, Zaret KS. H3K9me3-dependent heterochromatin: barrier to cell fate changes. Trends Genet 2016;32(1):29–41. doi:10.1016/j.tig.2015.11.001. [Google Scholar] [PubMed] [CrossRef]
38. Torrano J, Al Emran A, Hammerlindl H, Schaider H. Emerging roles of H3K9me3, SETDB1 and SETDB2 in therapy-induced cellular reprogramming. Clin Epigenet 2019;11(1):43. doi:10.1186/s13148-019-0644-y. [Google Scholar] [PubMed] [CrossRef]
39. Deng P, Yuan Q, Cheng Y et al. Loss of KDM4B exacerbates bone-fat imbalance and mesenchymal stromal cell exhaustion in skeletal aging. Cell Stem Cell 2021;28(6):1057–73.e7. doi:10.1016/j.stem.2021.01.010. [Google Scholar] [PubMed] [CrossRef]
40. Schleich K, Kase J, Dörr JR et al. H3K9me3-mediated epigenetic regulation of senescence in mice predicts outcome of lymphoma patients. Nat Commun 2020;11(1):3651. doi:10.1038/s41467-020-17467-z. [Google Scholar] [PubMed] [CrossRef]
41. Tanaka M, Harada H, Kimura H. The role of H3K9me3 in oral squamous cell carcinoma. Biochem Biophys Res Commun 2023;640:56–63. doi:10.1016/j.bbrc.2022.11.102. [Google Scholar] [PubMed] [CrossRef]
42. Du H, Zhang L, Li G et al. CXCR4 and CCR7 expression in primary nodal diffuse large B-cell lymphoma—a clinical and immunohistochemical study. Am J Med Sci 2019;357(4):302–10. doi:10.1016/j.amjms.2019.01.008. [Google Scholar] [PubMed] [CrossRef]
Cite This Article
 Copyright © 2026 The Author(s). Published by Tech Science Press.
Copyright © 2026 The Author(s). Published by Tech Science Press.This work is licensed under a Creative Commons Attribution 4.0 International License , which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.


 Submit a Paper
Submit a Paper Propose a Special lssue
Propose a Special lssue View Full Text
View Full Text Download PDF
Download PDF Downloads
Downloads
 Citation Tools
Citation Tools