 Open Access
Open Access
ARTICLE
Dioscin Regulates Mitochondrial Autophagy and Cell Cycle to Promote Pulpal Stem Cell Differentiation and Mineralization
1 Department of Stomatology, Naval Medical University, Shanghai, China
2 Department of Stomatology, Wusong Central Hospital, Shanghai, China
3 Department of Stomatology, The First Affiliated Hospital of Naval Medical University, Shanghai, China
* Corresponding Author: Qiang Zhu. Email:
BIOCELL 2026, 50(5), 10 https://doi.org/10.32604/biocell.2026.076758
Received 26 November 2025; Accepted 12 February 2026; Issue published 13 May 2026
Abstract
Background: Human dental pulp stem cells (hDPSCs) are promising for dental tissue regeneration. Dioscin (Dio), a natural compound, has various biological activities, but its effects on hDPSCs are unclear. This study aims to systematically elucidate the effects of Dio on promoting the osteogenic differentiation of hDPSCs and the underlying molecular mechanisms. Methods: Characterized hDPSCs were treated with Dio. Cell viability, proliferation, osteogenic differentiation (alkaline phosphatase (ALP) activity, Alizarin Red S (ARS)), and migration (Transwell) were assessed. Mitophagy (fluorescence, Western blot for PTEN-induced kinase 1 (PINK1), parkin RBR E3 ubiquitin-protein ligase (PRKN), microtubule-associated protein 1 light chain 3-II/I (LC3II/I), sequestosome 1 (p62)) and cell cycle (flow cytometry, cyclin D1 (Cyclin-D1), cellular myelocytomatosis oncogene (c-Myc)) were evaluated. The phosphatidylinositol 3-kinase (PI3K) inhibitor LY294002 and runt-related transcription factor 2 (Runx2) overexpression were used to investigate the PI3K/protein kinase B (AKT)/Runx2 pathway. Results: hDPSCs displayed mesenchymal stem cell characteristics. Dio treatment enhanced hDPSC viability, significantly increased ALP activity and ARS staining intensity, and promoted cell migration. It also increased mitophagy (increased colocalization of mitochondria and lysosomes, upregulated protein expression of PINK1 and PRKN, an increased LC3II/I ratio) and promoted cell cycle progression (increased S-phase cells, Cyclin-D1, c-Myc). Dio activated the PI3K/AKT pathway, upregulating Runx2. LY294002 reversed Dio’s effects, while Runx2 overexpression enhanced them. Conclusion: Dio is associated with enhanced hDPSC proliferation, osteogenesis, migration, mitophagy, and cell cycle progression, partly through activation of the PI3K/AKT pathway and upregulation of Runx2. These findings support the potential application of Dio in dental pulp regeneration and bone tissue engineering.Graphic Abstract
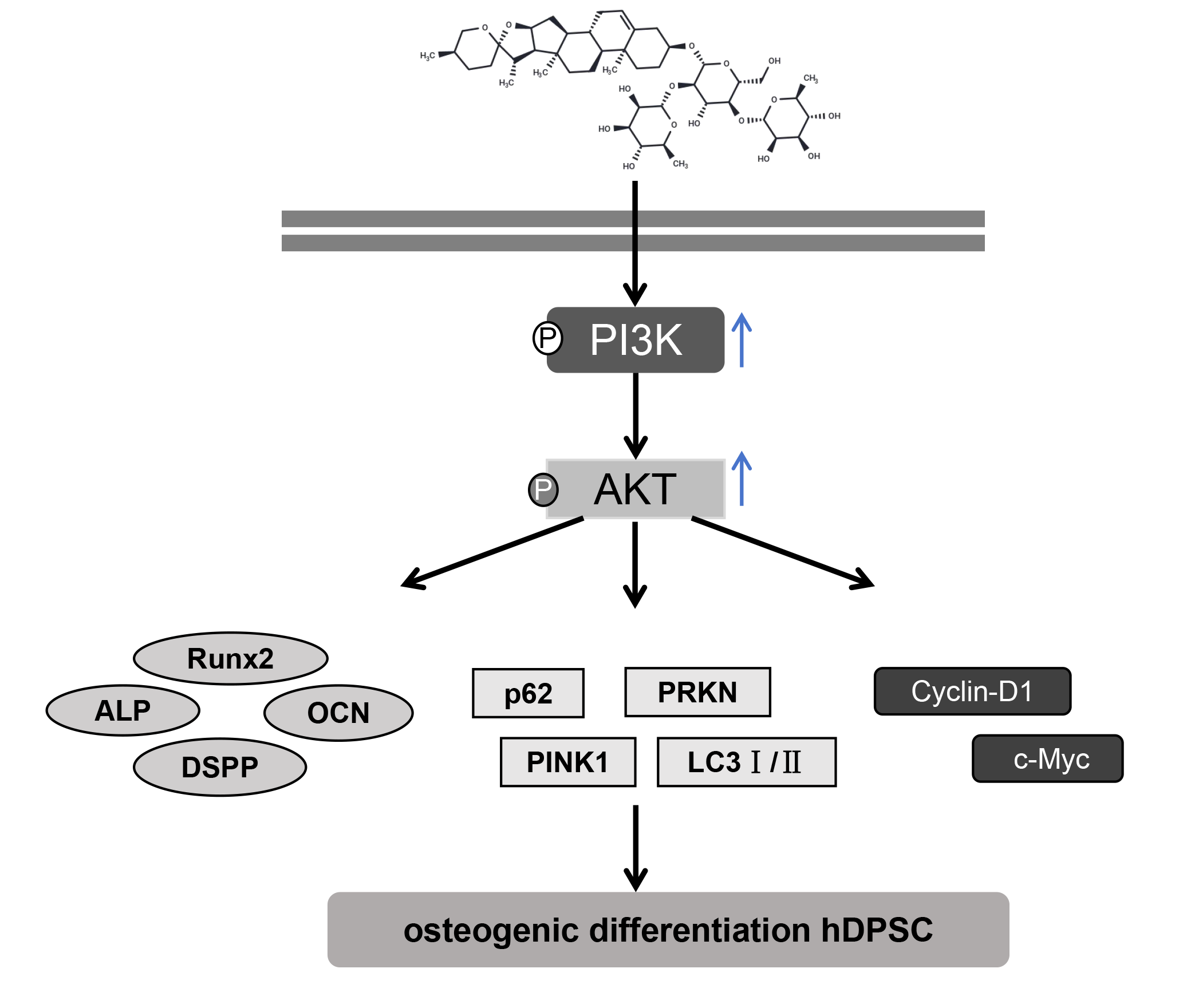
Keywords
Supplementary Material
Supplementary Material FileDental pulp stem cells (DPSCs) are a type of mesenchymal stem cell residing in dental pulp tissue, possessing self-renewal capacity and multipotent differentiation potential [1]. They are considered ideal seed cells for pulp tissue repair, dentin regeneration, and stem cell-based tissue engineering therapies [2]. One of the core biological functions of DPSCs is their ability to differentiate into odontoblast-like cells and secrete extracellular matrix (ECM), ultimately forming mineralized dentin structures [3]. The Phosphatidylinositol 3-kinase/Protein Kinase B (PI3K/AKT) signaling pathway has been extensively studied due to its pivotal role in regulating cell survival, proliferation, metabolism, and differentiation, making it a key focus in research on DPSC differentiation among numerous related pathways [4,5,6]. The activation of the PI3K/AKT pathway has been demonstrated to significantly enhance the differentiation potential of mesenchymal stem cells (MSCs) toward osteoblastic and odontoblastic lineages [7,8]. A key downstream effector of this pathway is runt-related transcription factor 2 (Runx2) [9]. Runx2, which serves as an indispensable molecular switch, governs the differentiation and mineralization of DPSCs; its knockout impairs odontoblast differentiation in early dentinogenesis [10]. However, how the PI3K/AKT pathway regulates the activity and expression of Runx2 during DPSC differentiation and mineralization, and the interplay between this signaling axis and other key cellular events, remains an area requiring further investigation.
The determinants of cellular differentiation extend beyond the PI3K/AKT pathway to include the cell’s energy metabolism and proliferation cycle. Research indicates that moderate mitophagy is not merely a clearance mechanism but also participates in regulating cell differentiation by modulating reactive oxygen species (ROS) levels, thereby indirectly influencing the expression of differentiation-related genes [11,12]. Concurrently, the fate decisions of stem cells are closely associated with the progression of their cell cycle. Stem cells typically need to exit the cell cycle (entering G0/G1 arrest) to initiate differentiation programs [13]. Evidence suggests that changes in the expression of cell cycle proteins, such as cyclins and cyclin-dependent kinases, can affect stem cell proliferation and differentiation [14,15]. How changes in the PI3K/AKT pathway, a major pro-proliferative signal, coordinate mitophagy, the cell cycle, and the initiation of differentiation programs, particularly in DPSCs, still warrant further investigation.
In recent years, active compounds extracted from natural plants have received widespread attention due to their excellent biocompatibility and fewer side effects. Dioscin (Dio), a steroidal saponin isolated from plants of the Dioscorea genus, possesses antioxidant, anti-inflammatory, and lipid-lowering effects [16]. In the context of periodontitis, Dio has been found to elevate the levels of key osteogenic markers in human periodontal ligament stem cells (hPDLSCs) and potentiate their ability to undergo osteogenic differentiation [17]. In bone-related studies, it has been reported that Dio can promote the osteogenic differentiation of rat bone marrow mesenchymal stem cells (BMSCs) by activating the AMPK signaling pathway [18]. Nevertheless, the potential influence of Dio on the differentiation and mineralization of hDPSCs, along with its underlying molecular mechanisms, is yet to be elucidated and demands thorough investigation. The present study was designed to systematically investigate the novel function of Dio in stimulating the odonto/osteogenic differentiation of hDPSCs. Our findings are expected to provide fresh theoretical perspectives on the regulatory network governing DPSC differentiation and to establish an experimental basis for future therapeutic strategies utilizing Dio to regenerate the dentin-pulp complex.
2.1 Cell Culture and Treatment
hDPSCs (PT-5025) were purchased from LONZA (Walkersville, MD, USA). The cells were cultured in DMEM medium (Gibco, 11965092, Grand Island, NY, USA) supplemented with 10% FBS (Gibco, A5670701), 50 U/mL penicillin, and 50 μg/mL streptomycin (ABclonal, BR00080, Wuhan, China). Cells were cultured in an incubator at 37°C with 5% CO2, and the culture medium was replaced every 3 days. When cell confluence reached 80–90%, they were passaged, and cells at passages 3–8 were used for subsequent experiments. All cells were authenticated by the supplier using short tandem repeat (STR) profiling and were routinely tested for mycoplasma contamination during the experimental period using a PCR-based mycoplasma detection kit (Beyotime, C0301, Shanghai, China); all results were negative.
Upon plating at 3 × 104 cells/well in 6-well plates, hDPSCs were cultured in osteogenic or chondrogenic induction media for a 21-day course. Differentiation was monitored and characterized via specific cell staining methods at predetermined intervals during the induction. The osteogenic induction medium consisted of DMEM supplemented with 10% FBS, 50 U/mL penicillin, 50 μg/mL streptomycin, 0.1 mM dexamethasone (Sigma, 265005, St. Louis, MO, USA), 10 mM β-glycerophosphate (MCE, HY-126304, Monmouth Junction, NJ, USA), and 50 μg/mL ascorbic acid (Sigma, 1043003). The chondrogenic induction medium was composed of DMEM supplemented with 5% FBS, 0.1 mM dexamethasone, 10 ng/mL TGF-β3 (MCE, HY-P7120), 50 μg/mL ascorbic acid, 1% sodium pyruvate (Aladdin, 113-24-6, Shanghai, China), and 50 mg/mL insulin transferrin selenium (ITS) premix (Corning, CLS354350, Corning, NY, USA).
To investigate the function of Runx2, hDPSCs overexpressing Runx2 (OE-Runx2 group) were constructed using Lipofectamine 2000 transfection reagent (Invitrogen, 11668019, Carlsbad, CA, USA), with a corresponding empty vector control group (OE-NC group). For plasmid transfection, 4 μL of Lipofectamine 2000 and 4 μg of the pcDNA-RUNX2 construct (RiboBio, Guangzhou, China) were added to each well and transfected into the cell line. The pcDNA-Runx2 plasmid was constructed as follows: total RNA was extracted from cells and reverse-transcribed into cDNA. The target fragment was amplified by PCR using the primers (forward: 5′-GCTAGCATGGACTACAAAGACGATGACGACAAGGTGATGCGTATTCCCGTAGATCCGAG-3′; reverse: 5′-CTCGAGATATGGTCGCCAAACAGATTCATCC-3′). The PCR product was then digested with NheI-XhoI and cloned into the pcDNA3.1 vector to generate the pcDNA-Runx2 plasmid.
To investigate the impact of Dio on DPSC differentiation and mineralization, hDPSCs were exposed to a concentration gradient of Dio (MCE, HY-N0124) ranging (0, 0.5, 1, 2, 4, 8, 16 μM) for a period of 7 days. Based on cell viability assays, Dio concentrations of 1, 2, and 4 μM were selected for subsequent experiments. Investigation into Dio’s regulatory role in the PI3K/AKT/Runx2 cascade, additional groups were established: Dio+LY294002, Dio+LY294002+OE-NC, and Dio+LY294002+OE-Runx2. In the Dio+LY294002 group, cells were co-cultured with 10 μM LY294002 (a PI3K inhibitor; MCE, HY-10108) and Dio for 7 days. In the Dio+LY294002+OE-NC and Dio+LY294002+OE-Runx2 groups, after being transfected with the empty vector or Runx2 overexpression vector, cells were then exposed to a combined treatment of LY294002 and Dio. In addition, to further evaluate the effects of Dio on autophagic flux and mitophagy in hDPSCs, Bafilomycin A1 (Baf A1, a late-stage autophagy inhibitor, HY-100558, MCE) was used to interfere with autophagic flux. First, the cytotoxicity of Baf A1 to hDPSCs was assessed using the CCK-8 assay. Cells were seeded into 96-well plates and treated with Baf A1 at final concentrations of 0, 5, 10, 20, 40, 80, and 160 nM for 24 h. Based on the Western blot results, 20 nM was determined as the Baf A1 concentration for subsequent experiments. Four experimental groups were then established: control group, 2 μM Dio group, 20 nM Baf A1 group, and 2 μM Dio + 20 nM Baf A1 co-treatment group. Baf A1 was added to the culture medium at a final concentration of 20 nM during the last 4–6 h of Dio treatment to block the fusion of autophagosomes with lysosomes.
After the 8-day induction period, cells (initially seeded at 100 cells/well in 6-well plates) were fixed with 10% formalin (Aladdin, 50-00-0) for 30 min and stained with 0.1% toluidine blue (Aladdin, 6586-04-5) for 30 min. The plates were then washed with PBS (pH 7.2, 0.01 M) and ready for microscopic (CKX53, Olympus, Tokyo, Japan) evaluation.
Cells were seeded into 24-well plates at a density of 5 × 104 cells per well and then switched to adipogenic induction medium (Solarbio, D3514, Beijing, China) for induction for 21 days. Cells were fixed with 4% paraformaldehyde (PFA) for 30 min and stained with Oil Red O staining solution (Beyotime, C0157) for 30 min. After washing with PBS, images were captured under a microscope (Eclipse Ti-S, Nikon, Tokyo, Japan).
2.4 Alizarin Red S (ARS) Staining
Upon plating in 24-well plates (5 × 104 cells/well), cells underwent a 21-day osteogenic induction. After fixation with 95% absolute ethanol (Aladdin, 64-17-5), stained with ARS staining solution (30 min; Beyotime, C0148). After staining, 10% acetic acid solution was added to dissolve the calcium deposits, and the optical density (OD) was measured at 490 nm using a microplate reader (Bio-Rad, Benchmark™ Plus, Hercules, CA, USA). Quantitative analysis was performed using SPSS 26.0 (IBM Corp., Armonk, NY, USA).
Cells (2 × 105) were harvested in a 15 mL tube and incubated in chondrogenic induction medium for 21 days. When cell pellets formed, the tubes were gently agitated to suspend the spheroids in the medium. Following induction, with medium changes every 2–3 days, the cell pellets were fixed (4% PFA, 30 min), paraffin-embedded, and sectioned at 5 μm. The resulting sections were stained with Alcian blue (Aladdin, 33864-99-2) for 1 h prior to observation.
When the cells reached approximately 80% confluence, they were digested with 0.25% trypsin for 3 min, and the enzymatic reaction was terminated by adding an equal volume of complete medium containing 10% FBS. The cell suspension was then collected and centrifuged. After resuspension, the cell density was adjusted to 1 × 107 cells/mL and incubated with the following antibodies (1:20 dilution) at 4°C for 30 min: CD34 (571801), CD45 (772353), CD44 (749713), CD73 (772799), and CD90 (774385), purchased from BD Biosciences (San Jose, CA, USA). Samples were analyzed for fluorescence using a flow cytometer (FACSCanto II, BD Biosciences), and the data were subsequently processed with FlowJo software (version 7.6.1).
For cell cycle analysis, collected cells were fixed in 70% ethanol (−20°C, 4 h). After ethanol fixation, cells were resuspended in cold PBS (pH 7.2, 0.01 M) and stained with PI stain solution (Beyotime, C1052) in the dark (37°C, 30 min). Samples were then analyzed on a flow cytometer, and the resulting data were processed with FlowJo software’s cell cycle module for phase quantification (G1, S, G2).
To determine cell viability, hDPSCs were plated in 96-well plates at 5 × 103 cells/well and exposed to a gradient of Dio concentrations (0, 0.5, 1, 2, 4, 8, 16 μM) for 7 days. At designated time points (days 1, 3, 7), 10 μL of CCK-8 reagent (ABclonal, RM02823) was added to each well. The plates were incubated for 3 h, and the OD value was measured at 450 nm.
2.8 Alkaline Phosphatase (ALP) Activity Assay
To evaluate hDPSC differentiation, ALP activity was determined using a specific kit (Beyotime, P0321). Cells were plated in 6-well plates and treated according to the protocols in Section 2.1. Cells were washed with PBS (pH 7.2, 0.01 M) and then placed on ice. Subsequently, 100 μL of lysis buffer (Beyotime, P0013J) was added, followed by centrifugation at 12,000 rpm for 15 min at 4°C using an Eppendorf 5417R refrigerated centrifuge (Eppendorf, Hamburg, Germany). The supernatant was collected for analysis. A standard curve was generated using standard samples. Following a 10-min reaction initiated by the addition of a chromogenic substrate, the process was terminated with 100 μL of stop solution. The absorbance was then measured at 405 nm using a microplate reader (Benchmark Plus, Bio-Rad, Hercules, CA, USA), and ALP activity for each sample was determined from the standard curve.
Cell migration was evaluated using Transwell chambers (8 μm pore, Corning). Briefly, 200 μL of serum-free medium containing hDPSCs was added to the upper chamber, while the lower chamber was filled with DMEM supplemented with 10% FBS as a chemoattractant. After 24 h, non-migrated cells on the upper surface were removed by swabbing. The migrated cells on the lower membrane were fixed with 4% PFA for 15 min, stained with 0.1% crystal violet (Sigma, V5265) for 20 min, and counted in five random microscopic fields. Results are expressed as fold-change relative to the control group (set to 1).
2.10 Colocalization of Mitochondria and Lysosomes
A dual-fluorescence labeling approach was employed to co-stain mitochondria and lysosomes, enabling the assessment of Dio’s effect on mitophagy. After performing specific treatments on the cells as described in Section 2.1, hDPSCs in each group were sequentially incubated with 75 nM LysoTracker Red (Beyotime, Y060275) and 200 nM MitoTracker Green (Beyotime, C1048) for 30 min each. The cells were then stained with DAPI (Beyotime, C1006) for 5 min and fluorescence microscopy (Olympus, CKX53, Tokyo, Japan). The level of colocalization was assessed by the Pearson’s correlation coefficient (PCC) in a semi-quantitative analysis. In this assay, green fluorescence indicated mitochondria, red fluorescence indicated lysosomes, and the orange signal resulting from their colocalization suggested the potential occurrence of mitophagy.
2.11 Transmission Electron Microscopy (TEM)
After treatment with Dio, the cells were digested and centrifuged at 1000 rpm for 10 min to collect the cell pellets. The pellets were first fixed overnight at 4°C with 2.5% glutaraldehyde fixative (Solarbio, P1126), and then washed twice with 0.1 M sodium dimethylarsenate buffer (Sigma, 1.06277) for 10 min each. Subsequently, the samples were fixed with 1% osmium acid at 4°C for 1.5 h and washed twice with distilled water for 10 min each. After fixation, the samples were dehydrated in a graded ethanol series (30%, 50%, 70%, and 90%) followed by acetone (90% and 100%), with each step lasting 10–15 min. The dehydrated samples were then infiltrated with Epon812 resin (Electron Microscopy Sciences, 14120, Hatfield, PA, USA) diluted with acetone, and incubated sequentially in 33%, 66%, and 100% (w/v) resin for approximately 3 h at each concentration. After resin infiltration, the samples were polymerized in an oven at 37°C for 12 h, 45°C for 12 h, and finally at 60°C for 48 h to obtain embedding blocks. Ultrathin sections of approximately 70 nm were cut from the cured blocks, stained with lead citrate (Macklin, L885990, Shanghai, China), and examined and photographed using a transmission electron microscope (H-7650, Hitachi, Tokyo, Japan).
Total RNA was extracted from cells using TRIzol reagent (15596026, Invitrogen) according to the manufacturer’s instructions. RNA (1 μg) was reverse transcribed into cDNA using a reverse transcription kit (R323-01, Vazyme, Nanjing, China). The reverse transcription reaction system consisted of 4 μL 4× gDNA wiper Mix, 1 μg total RNA, and RNase-free water added to a final volume of 16 μL, followed by incubation at 42°C for 2 min. Then, 4 μL 5× HiScript III qRT SuperMix was added, and the mixture was incubated at 37°C for 15 min and heated at 85°C for 5 s. qPCR amplification was performed on a real-time PCR instrument (CFX Opus, Bio-Rad) using SYBR Green PCR Master Mix (CN830, Takara, Kyoto, Japan). The qPCR reaction mixture (per well) contained 10 μL SYBR Premix Ex Taq (2×), 0.4 μL primer (10 μM), 0.4 μL ROX Reference Dye (50×), 2 μL cDNA template, and ddH2O. The PCR cycling conditions were as follows: initial denaturation at 95°C for 30 s, followed by 40 cycles of denaturation at 95°C for 5 s and annealing/extension at 60°C for 34 s. Each sample was run in triplicate, and a melting curve analysis was performed to verify primer specificity. GAPDH was used as the internal control, and relative gene expression was calculated using the 2−ΔΔCt method. The primer sequences were as follows: Runx2, forward 5′-GGAGTGGACGAGGCAAGAGTTT-3′ and reverse 5′-AGCTTCTGTCTGTGCCTTCTGG-3′; GAPDH, forward 5′-CTCTGCTCCTCCTGTTCGACA-3′ and reverse 5′-ACGACCAAATCCGTTGACTC-3′.
After protein extraction from treated hDPSCs with RIPA buffer (Solarbio, R0010) and concentration determination by BCA kit (PC0040, Solarbio), equal protein aliquots were resolved by SDS-PAGE and electroblotted onto PVDF membranes. The membranes were blocked with blocking buffer (Beyotime, P0023) for 1 h prior to immunoblotting. They were then incubated overnight at 4°C with primary antibodies against ALP (1:500, Abbexa, abx271785, Cambridge, UK), Osteocalcin (OCN, 1:500, GeneTex, GTX55255, Irvine, CA, USA), Dentin Sialophosphoprotein (DSPP, 1:1000, Proteintech, 55396-1-AP, Wuhan, China), Runx2 (1:200, Proteintech, 20700-1-AP), Sequestosome 1 (p62, 1:10000, Proteintech, 18420-1-AP), PTEN-induced putative kinase 1 (PINK1, 1:2000, Proteintech, 23274-1-AP), Parkin RBR E3 ubiquitin-protein ligase (PRKN, 1:2000, Proteintech, 14060-1-AP), Microtubule-associated protein 1A/1B-light chain 3 (LC3I/II, 1:3000, Proteintech, 14600-1-AP), Cyclin-D1 (1:5000, Proteintech, 60186-1-Ig), c-Myc (1:2000, Proteintech, 10828-1-AP), p-PI3K (1:1000, CST, #4249, Danvers, MA, USA), PI3K (1:200, Proteintech, 20584-1-AP), AKT (1:5000, Proteintech, 10176-2-AP), p-AKT (1:2000, CST, #4060), and β-actin (1:5000, Proteintech, 20536-1-AP). The following day, after being washed with TBST, the membranes were incubated with an HRP-labeled secondary antibody (1:5000, #7074, CST) for 1 h. Protein bands were visualized using an ECL kit (Beyotime, P0018). β-actin was used as the internal control, and the grayscale values of the protein bands were quantified using ImageJ software (version 1.53c).
All experimental data are presented as the mean ± standard deviation (Mean ± SD). Cell experiments were independently repeated six times, with five replicate wells set up in each independent experiment, and the average value was used for statistical analysis. Before performing parametric tests, data were tested for normality (Shapiro–Wilk test) and homogeneity of variance. Statistical analyses were conducted using GraphPad Prism software. Comparisons between two groups were performed using Student’s t-test, while comparisons among multiple groups were conducted using one-way analysis of variance (ANOVA) followed by Bonferroni post hoc test. Effect sizes for between-group differences were expressed as fold changes, defined as the ratio of the mean value in the treatment group to that in the control group. A p value of less than 0.05 was considered statistically significant.
3.1 Characterization of hDPSCs
The characteristics of hDPSCs were investigated through flow cytometry and cell staining. Immunophenotyping by flow cytometry identified hDPSCs as positive for the MSC markers (CD44, CD73, CD90) and negative for the hematopoietic markers (CD34, CD45) (Fig. 1A). Furthermore, toluidine blue staining indicated that hDPSCs were capable of forming dense cell colonies by day 8 (Fig. 1B), suggesting a strong proliferative capacity. Following 21 days of adipogenic induction, Oil Red O staining demonstrated successful lipid droplet formation, confirming adipogenic differentiation capability (Fig. 1C). Concurrently, ARS staining confirmed that the osteogenic induction of hDPSCs for 21 days resulted in the formation of mineralized nodules (Fig. 1D), demonstrating a capacity for osteogenic differentiation. Following 21 days of chondrogenic induction, Alcian Blue staining demonstrated the secretion of acidic mucopolysaccharides by chondrocytes (Fig. 1E). Collectively, these findings verify that the hDPSCs possess the typical characteristics and multi-lineage differentiation potential.

Figure 1: Characterization of hDPSCs. (A) Flow cytometric analysis of surface antigen expression on hDPSCs (CD34, CD45, CD44, CD73, CD90). (B) Toluidine blue staining for the assessment of colony-forming ability. Scale bar: 100 μm. (C) Oil Red O staining following 21-day adipogenic differentiation induction. Scale bar: 100 μm. (D) ARS staining following 21-day osteogenic differentiation induction. Scale bar: 100 μm. (E) Alcian blue staining following 21-day chondrogenic differentiation induction. Scale bar: 100 μm. n = 6.
3.2 Dio Promotes the Viability, Osteogenesis, and Migration of DPSCs
A CCK-8 assay was used to evaluate the effect of different concentrations of Dio (chemical structure shown in Fig. 2A) on the viability of hDPSCs. On days 1, 3, and 7, treatment with 8 μM and 16 μM Dio inhibited cell viability. At the 3- and 7-day time points, Dio significantly promoted hDPSC viability in a dose-dependent manner (i.e., 1, 2, and 4 μM) (Fig. 2B). Based on these results, concentrations of 1, 2, and 4 μM Dio were chosen for subsequent experiments.
In cellular osteogenic differentiation, ALP functions as a key early marker during osteogenesis [19]. The ALP activity assay results showed that Dio treatment significantly upregulated the ALP activity of hDPSCs on day 8 post-osteogenic induction (2 μM Dio: 3.21-fold, p < 0.001) (Fig. 2C). Mineralized nodules, which are deposits formed from the calcification of the ECM secreted by osteoblasts, serve as direct evidence of successful in vitro osteogenic differentiation [20]. Moreover, Dio treatment increased the formation of mineralized nodules on day 21 of osteogenic induction (ARS staining, 2 μM Dio: 5.23-fold, p < 0.001; Fig. 2D,E), indicating that Dio can enhance the mineralization capacity of hDPSCs. OCN is primarily synthesized and secreted by mature osteoblasts and is a specific marker for the terminal stage of osteoblast differentiation [21]; DSPP is a specific and highly expressed marker of odontoblasts [22]; and Runx2 is the earliest and most crucial osteoblast-specific transcription factor [23]. The protein levels of ALP, OCN, DSPP, and Runx2 were upregulated by Dio treatment at the early (day 3) and middle (day 7) phases of osteogenic induction (Fig. 2F–J). Regarding cell migration, Dio treatment significantly enhanced the migratory ability of hDPSCs (2 μM Dio: 1.88-fold, p < 0.001; Fig. 2K,L). In summary, Dio, at appropriate concentrations, effectively promotes the viability, osteogenic differentiation, and migration of hDPSCs.

Figure 2: Dio promotes the viability, osteogenesis, and migration of DPSCs. (A) Chemical structure of Dio. (B) CCK-8 across a range of Dio concentrations and time points. (C) ALP activity measurement on day 8. (D,E) Mineralization assessment by ARS staining and quantification on day 21. Scale bar: 100 μm. (F–J) Protein expression analysis of osteogenic markers via Western blot on days 3 and 7. (K,L) Cell migration evaluation by Transwell assay. Scale bar: 100 μm. The 0 μM Dio group was treated with DMSO and served as the vehicle control. n = 6.
3.3 Dio Promotes Mitophagy and Cell Cycle Progression in DPSCs
After 24 h of Dio treatment, the co-localization of mitochondria (green fluorescence) and lysosomes (red fluorescence) in hDPSCs was significantly increased (Fig. 3A,B), suggesting that Dio promotes the occurrence of mitophagy. p62 is often used as a marker for mitophagy [24]. Upon mitochondrial damage, PINK1 accumulates on the outer membrane, where it serves to activate PRKN. This PINK1-PRKN complex subsequently orchestrates a selective autophagic response, which is crucial for maintaining a healthy mitochondrial network and proper cellular function [25,26]. LC3-II, produced by the lipidation of LC3-I with phosphatidylethanolamine, plays an indispensable role in autophagy by enabling the formation of autophagosomes [27]. Western Blot results further verified that Dio elevated the protein levels of PINK1 (2 μM Dio: 2.79-fold, p < 0.001), PRKN (2 μM Dio: 1.64-fold, p < 0.001), and the LC3-II/I ratio (2 μM Dio: 1.21-fold, p < 0.001), whilst concurrently reducing p62 expression (2 μM Dio: 1.69-fold, p < 0.001) (Fig. 3C–G). TEM results showed that the number of mitochondria engulfed by lysosomes increased with the dose of Dio (Fig. 3H). In addition, MitoTracker Green staining revealed that mitochondrial fluorescence intensity gradually decreased as the concentration of Dio increased; in the Dio-treated groups (especially at 2 μM), mitochondrial fluorescence intensity markedly declined within 24 h and remained at a relatively low level for at least 48 h (Fig. 3I).
Regarding the cell cycle, Dio treatment significantly altered the distribution of hDPSCs, increasing the proportion of cells in the S phase and decreasing the proportion in the G1 phase (Control: G1: 48%, G2: 3.73%; 4 μM Dio: G1: 34.4%, G2: 8.5%) (Fig. 3J,K). This suggests that Dio may promote the proliferative activity of hDPSCs, driving more cells from the quiescent phase (G1) into the DNA synthesis phase (S), thereby accelerating cell cycle progression. c-Myc acts as a pivotal transcription factor that stimulates Cyclin D1 expression, either directly or indirectly, to drive cell cycle progression from G1 to S phase and consequently promote proliferation [28]. Dio treatment promoted cell cycle progression in hDPSCs, as further evidenced by Western Blot results showing upregulated expression of the key cell cycle proteins Cyclin-D1 (2 μM Dio: 1.44-fold, p < 0.001) and c-Myc (2 μM Dio: 3.07-fold, p < 0.001) (Fig. 3L–N). Collectively, these results demonstrate that Dio promotes mitophagy and cell cycle progression in hDPSCs in vitro.

Figure 3: Dio promotes mitophagy and the cell cycle in DPSCs. (A,B) Mitochondrial and lysosomal colocalization assay. Scale bar: 50 μm. (C–G) Protein expression analysis of p62, PINK1, PRKN, and LC3-II/I. (H) Transmission electron microscopy (TEM) images showed that the number of mitochondria engulfed by lysosomes gradually increased with the concentration of Dio. Red arrows indicate mitochondria surrounded by lysosomes. (I) MitoTracker Green staining was used to analyze changes in total mitochondrial mass. (J,K) Cell cycle profiling by flow cytometry. (L–N) Protein expression analysis of Cyclin D1 and c-Myc. The 0 μM Dio group was treated with DMSO and served as the vehicle control. n = 6.
3.4 Dio Activates the PI3K/AKT Pathway to Promote Runx2 Expression
Following 7-day Dio treatment, hDPSCs exhibited increases in p-PI3K (2 μM Dio: 2.05-fold, p < 0.001) and p-AKT (2 μM Dio: 1.77-fold, p < 0.001), but not total PI3K/AKT (Fig. 4A–C), indicating PI3K/AKT pathway activation. A significant upregulation of Runx2 was also observed (2 μM Dio: 1.52-fold, p < 0.001; Fig. 4D). To verify the role of the PI3K/AKT signaling pathway in this process, cells were treated with the PI3K inhibitor LY294002. Compared to the group treated with Dio alone, the expression levels of p-PI3K, p-AKT, and Runx2 were all significantly decreased in the group treated with Dio in combination with 10 μM LY294002 (p < 0.001, Fig. 4E–H). This demonstrates that LY294002 can effectively inhibit the Dio-induced activation of the PI3K/AKT pathway and the upregulation of Runx2. These findings collectively confirm that Dio promotes Runx2 expression through PI3K/AKT pathway activation.

Figure 4: Dio activates the PI3K/AKT pathway to promote Runx2 expression. (A–D) Protein level analysis of the PI3K/AKT pathway and Runx2 after Dio treatment (1, 2, 4 μM; 7 days). (E–H) Protein level analysis following pharmacological inhibition of PI3K with LY294002. The 0 μM Dio group was treated with DMSO and served as the vehicle control.
3.5 Dio Promotes the Proliferation, Osteogenesis, and Migration of DPSCs by Activating Runx2 via the PI3K/AKT Pathway
To elucidate the roles of the PI3K/AKT signaling pathway and Runx2 in Dio-induced promotion of hDPSC proliferation, osteogenesis, and migration, PI3K inhibitor LY294002 treatment and Runx2 overexpression experiments were added in this study. Western blot and qPCR analyses confirmed the effectiveness of the Runx2 overexpression vector (OE-Runx2), which successfully upregulated Runx2 expression in hDPSCs (Fig. 5A,B). CCK-8 assay results showed that on days 3 and 7, Dio significantly promoted the proliferation of hDPSCs, whereas the addition of LY294002 partially reversed this pro-proliferative effect (Fig. 5C). Notably, in the presence of LY294002, OE-Runx2 was able to rescue the pro-proliferative effect of Dio, suggesting a key role for Runx2. Dio treatment promoted osteogenic differentiation by upregulating the protein expression of ALP, OCN, DSPP, and Runx2. These effects were significantly inhibited by LY294002 but were rescued by Runx2 overexpression (Fig. 5D–H). During the osteogenic induction process, LY294002 attenuated the enhancing effect of Dio on ALP activity on day 8 (Fig. 5I), and on day 21, it diminished the promoting effect of Dio on mineralized nodule formation (Fig. 5J,K). Runx2 overexpression was able to partially reverse these inhibitory effects. Furthermore, Transwell migration assay results indicated that the ability of Dio to promote hDPSC migration was inhibited by LY294002 and could be restored by Runx2 overexpression (Fig. 5L,M). Together, these results demonstrate that Dio promotes the proliferation, osteogenic differentiation, and migration of hDPSCs, in association with activation of Runx2 via the PI3K/AKT signaling pathway.

Figure 5: Dio promotes DPSC proliferation, osteogenesis, and migration by activating Runx2 via the PI3K/AKT pathway. (A,B) Verification of Runx2 overexpression efficiency. (C) Assessment of cell proliferation using the CCK-8 assay at days 1, 3, and 7. (D–H) Western blot analysis of osteogenesis-related proteins (ALP, OCN, DSPP, RUNX2). (I) ALP activity assay on day 8 of osteogenic induction. (J,K) ARS staining and quantification after 21 days of osteogenic induction. Scale bar: 100 μm. (L,M) Transwell migration assay. Scale bar: 100 μm. n = 6.
3.6 Dio Promotes Mitophagy and Cell Cycle by Activating Runx2 via the PI3K/AKT Pathway
We further focused on the mechanism by which Dio enhances mitophagy and cell cycle progression, specifically its action of activating Runx2 by PI3K/AKT pathway activation. After 24 h of Dio treatment, the co-localization of mitochondria and lysosomes in hDPSCs was significantly increased, indicating that mitophagy was promoted (Fig. 6A,B). When the PI3K inhibitor LY294002 was added, this co-localization was significantly inhibited, whereas Runx2 overexpression could partially rescue the Dio-induced mitophagy. Western Blot results further confirmed that Dio upregulated PINK1, LC3-II/I, and PRKN expression, and downregulated the expression of p62 (Fig. 6C–G). These effects were inhibited by LY294002 but were rescued by Runx2 overexpression. The ability of Dio to promote cell cycle progression in hDPSCs was inhibited by LY294002 and could be restored by Runx2 overexpression (Fig. 6H,I). Western Blot results also indicated that Dio upregulated c-Myc and Cyclin-D1 expression, and these effects were similarly inhibited by LY294002 and could be rescued by Runx2 overexpression (Fig. 6J–L). In summary, Dio acts through the PI3K/AKT pathway to activate Runx2, and this activation is associated with enhanced mitophagy and accelerated cell-cycle progression in hDPSCs.

Figure 6: Dio promotes mitophagy and the cell cycle by activating Runx2 via the PI3K/AKT pathway. (A,B) Mitochondria-lysosome colocalization assay. Scale bar: 50 μm. (C–G) Mitophagy marker analysis (p62, PINK1, PRKN, LC3-II/I). (H,I) Cell cycle profiling. (J–L) Cell cycle protein analysis (Cyclin D1, c-Myc). n = 6.
The hDPSCs in this study were confirmed to be MSCs, as evidenced by their expression of typical MSC markers (CD44, CD73, CD90) and absence of hematopoietic markers (CD34, CD45). This immunophenotype is consistent with that of MSCs, and the cells possess in vitro differentiation potential [29,30]. Cell staining results confirmed that hDPSCs could differentiate into adipocytes, osteoblasts, and chondrocytes under specific induction conditions. Furthermore, toluidine blue staining verified that hDPSCs have a strong colony-forming ability; stem cells with high colony-forming capacity typically possess greater expansion potential. This comprehensive characterization provided a reliable cell model for an in-depth investigation into the effects of Dio on the biological behavior of hDPSCs. Cui et al., using single-cell RNA sequencing, revealed changes in the cellular composition of hDPSCs during in vitro culture and identified a subpopulation with higher proliferative capacity and multi-lineage differentiation potential [31]. This research highlights the heterogeneity of hDPSCs and the potential phenotypic changes that may occur after in vitro expansion, but it also indicates that specific subpopulations can maintain their original stem cell characteristics, offering more precise guidance for the clinical application of hDPSCs. Through in-depth research on hDPSC subpopulations, cells with greater therapeutic potential can be screened, thereby improving the efficiency and specificity of regenerative therapies. Li et al. emphasized that hDPSCs and other dental-derived MSCs possess extensive differentiation capacity, similar to bone marrow MSCs, giving rise to odontogenic, osteogenic, chondrogenic, adipogenic, and neurogenic lineages [32]. This broad potential makes them promising candidates for regenerating dental tissues as well as for repairing bone, cartilage, and nerve defects.
The results of this study clearly indicate that Dio, at appropriate concentrations (1, 2, and 4 μM), promotes dose-dependent hDPSCs viability. In contrast, higher concentrations (8 and 16 μM) of Dio inhibited cell viability, suggesting that the biological effects of Dio are concentration-dependent. In future clinical applications, precise dose control of Dio will be crucial to avoid potential cytotoxicity and maximize its therapeutic effects. In terms of osteogenic differentiation, Dio treatment upregulated the ALP activity of hDPSCs; as an important early marker of osteogenesis, ALP activity is closely related to the mineralization capacity of the ECM. Subsequently, on day 21 of osteogenic induction, Dio treatment significantly increased the formation of mineralized nodules, which serve as direct evidence of successful in vitro osteogenic differentiation and reflect the cell’s ability to secrete and calcify the ECM. Together, these results indicate that Dio enhances osteogenic differentiation and mineralization in hDPSCs, a key capability for bone and dental pulp regeneration, as mineralization is indispensable for functional bone tissue formation [33]. Furthermore, Dio treatment resulted in a dose-dependent increase in the expression of osteogenic proteins across early and late stages, such as ALP, OCN, DSPP, and Runx2. The upregulation of DSPP, a specific marker for odontoblasts, suggests that Dio may also have a promoting effect on dentin regeneration, which is of great significance for the repair of the pulp-dentin complex. The elevated expression of these markers suggests that Dio significantly promotes the osteogenic differentiation of hDPSCs by targeting multiple stages, spanning from early induction to terminal mineralization [34]. Transwell migration assays demonstrated that Dio significantly enhanced the migration of hDPSCs. Cell migration is a vital part of stem cell homing, tissue repair, and regeneration, especially after dental pulp injury, where hDPSCs need to migrate from surrounding tissues to the site of injury for repair [35]. The ability of Dio to promote hDPSC migration in vitro suggests a potential role in cellular processes related to dental pulp repair, but its effects on actual tissue regeneration and healing in vivo require further study. This is crucial for in vivo dental pulp regeneration, as effective cell homing is a prerequisite for successful regeneration. Additionally, Naringenin has also been found to stimulate the osteo/odontogenic differentiation and migration of hDPSCs [36], which further supports the potential of natural products in dental pulp regeneration.
This study further investigated the effects of Dio on mitophagy and the cell cycle in hDPSCs. Following Dio treatment, the colocalization of mitochondria and lysosomes in hDPSCs was markedly increased, indicating that Dio promotes the occurrence of mitophagy. Consistently, the expression of mitophagy-related proteins PINK1, PRKN, and LC3-II/I was upregulated, while p62 expression was downregulated. PINK1 and PRKN are key initiating factors of mitophagy [26], and an increased LC3-II/I ratio together with p62 degradation reflects autophagosome formation and activation of autophagic flux [37]. To further verify that Dio-induced autophagy represents genuine activation of autophagic flux rather than impaired degradation, the late-stage autophagy inhibitor Baf A1 was introduced for combined assessment. The results showed that when Dio was co-administered with Baf A1, the accumulation levels of LC3-II and p62 were significantly higher than those in the Baf A1 alone group (Fig. S1). This further accumulation of autophagy-related proteins under conditions of terminal blockade strongly demonstrates that Dio enhances the initiation and formation of autophagosomes, rather than inhibiting their degradation, thereby exhibiting a distinct feature of “enhanced autophagic flux.” This mechanism suggests that Dio can, to some extent, counteract the detrimental effects of Baf A1 or other pathological factors on hDPSCs by promoting an intact and functional mitophagic flux. Therefore, Dio facilitates the timely removal and quality control of damaged mitochondria by activating PINK1/PRKN-mediated mitophagy. Such optimization of the intracellular environment not only maintains the health status of hDPSCs under stress conditions but also provides essential metabolic energy support and homeostatic maintenance for subsequent cell proliferation, cell-cycle progression, and the exertion of osteogenic differentiation potential. With respect to the cell cycle, the G1 phase is the period of cell growth and preparation for DNA synthesis, whereas the S phase is the stage of DNA synthesis [38]. An increased proportion of cells in S phase is generally associated with enhanced proliferative activity, thereby providing a sufficient number of cells to support differentiation [39]. Flow cytometry results showed that Dio treatment significantly increased the proportion of hDPSCs in the S phase and decreased the proportion of cells in the G1 phase, suggesting that Dio promoted the transition of cells from G1 to S phase and accelerated cell cycle progression. This finding is consistent with recent reports on hDPSC differentiation, which indicate that BACH1, a factor that drives cell cycle progression, is upregulated during physiological differentiation, whereas its inhibition leads to G0/G1 arrest and impaired differentiation [40]. These results imply that, at early stages, an appropriate level of cell cycle activity is an integral part of the normal differentiation program. This pro-proliferative effect was further supported by the Dio-induced upregulation of Cyclin-D1 and c-Myc—a key regulator of the G1/S transition and its upstream transcription factor, respectively [41,42]. These results collectively indicate that Dio can promote mitochondrial autophagy and cell cycle progression of hDPSCs in vitro, thereby providing a more favorable intracellular environment for their proliferation and differentiation, suggesting that Dio may have potential application value in cellular processes related to pulp regeneration.
Healthy mitochondrial function is crucial for stem cells to maintain their stemness and differentiation capacity. Studies have shown that Dio protects the kidneys from 3-Chloro-1, 2-propanediol (3-MCPD)-induced injury by regulating autophagy and mitochondrial dynamics [43]. In bone metabolic research, mitophagy has emerged as a key regulator involved in osteogenic differentiation and mineral deposition. Autophagy is a necessary process for the in vitro mineralization of osteoblasts; it promotes extracellular mineralization by facilitating the secretion of apatite crystals from autophagic vacuoles in osteoblasts [44]. The reported role of the importin subunit beta1-Activating transcription factor 4 (KPNB1-ATF4) axis in promoting odontoblastic differentiation via mitophagy induction [45] lends theoretical support to our findings that Dio-induced mitophagy is linked to enhanced osteogenesis in hDPSCs, highlighting mitophagy’s importance in bone formation. In contrast, disruptions in mitochondrial quality control, such as dysfunction and elevated ROS, can suppress stem cell functionality [46]. Therefore, by promoting mitophagy to clear damaged mitochondria, Dio helps maintain mitochondrial health in hDPSCs, thereby supporting their differentiation and mineralization processes, which are crucial for dental pulp regeneration. Mitophagy and the cell cycle are tightly coupled biological processes. Studies demonstrate that impaired autophagy triggers cell cycle arrest, while enhanced autophagic activity supports cell cycle progression [47]. The precise regulation of the cell cycle ensures that cells can undergo sufficient expansion before differentiation, providing an adequate cell reserve for tissue repair while avoiding the risk of tumorigenesis from excessive proliferation [48]. The balance between proliferation and differentiation is crucial for successful stem cell-based tissue regeneration. Dio effectively promotes this equilibrium by increasing the S-phase cell population to expand the cell pool, while simultaneously enhancing the differentiation and mineralization capacity of hDPSCs to meet the requirements of osteogenesis.
The PI3K/AKT pathway is a vital intracellular signaling cascade that regulates diverse processes, including cell proliferation, differentiation, migration, and survival [49]. Our investigation into the molecular mechanism of Dio revealed that it activates this pathway in hDPSCs by enhancing the phosphorylation of PI3K and AKT, while concurrently upregulating the expression of the key transcription factor Runx2. The key evidence comes from the PI3K inhibitor LY294002, which suppressed Dio-induced phosphorylation of PI3K/AKT and Runx2 expression, thereby inhibiting Dio’s pro-osteogenic and pro-migratory effects. The partial rescue by Runx2 overexpression confirms its downstream role. This mechanistic insight is consistent with prior reports: the IIS/PI3K/AKT/mTOR pathway regulates Runx2 and osteogenesis in hDPSCs [50]; Dio-activated PI3K/AKT promotes bone formation [51,52]; and this pathway is established as a mediator of osteo/odontogenic differentiation [53,54]. Thus, our results are well-corroborated, identifying the PI3K/AKT/Runx2 axis as central to Dio’s action on hDPSCs.
The present study was mainly based on in vitro experiments and still lacks systematic in vivo evidence. In the future, it will be necessary to further validate the regenerative effects and safety of Dio in relevant animal models of pulp injury and bone defects, so as to evaluate its true therapeutic potential in pulp repair and bone regeneration. Meanwhile, in vivo optimization of the dosing concentration, route of administration, and dosing frequency of Dio is warranted to achieve the best efficacy while ensuring safety and minimizing potential side effects. At the mechanistic level, this study primarily focused on the promotion of osteogenic/odontogenic differentiation by Dio via the PI3K/AKT/Runx2 pathway and preliminarily suggested its involvement in mitophagy and cell cycle regulation. However, it remains unclear how this pathway precisely regulates the expression of mitophagy- and cell cycle-related genes, and whether critical intermediate molecules or transcription factors are involved. Moreover, the observed changes in autophagy- and cell cycle-related proteins in this study are phenomena associated with Runx2 activity and PI3K/AKT pathway activation, rather than validated direct transcriptional targets. Future studies should employ ChIP-qPCR/ChIP-seq and promoter luciferase reporter assays to further determine whether there is direct binding and transcriptional activation between Runx2 and the promoters of these genes. In addition, gene-editing strategies (such as CRISPR/Cas9-mediated knockout or overexpression of key genes), together with interventions using specific pathway inhibitors/agonists, combined with high-throughput approaches including proteomics, transcriptomics, and metabolomics, could be used to construct a global molecular network of Dio action and functionally verify key nodes, thereby clarifying causal relationships and signaling cascades. At present, quantitative evidence for mitochondrial degradation is still mainly based on MitoTracker Green staining, TEM, and mitochondria–lysosome colocalization. Subsequent work will further strengthen this line of evidence by assessing mtDNA copy number, mitochondrial protein levels (e.g., TOM20, VDAC1), and mtROS production.
In addition, previous studies have suggested that Dio can regulate cell proliferation and differentiation by inhibiting the Wnt/β-catenin pathway [55]. In the present study, we focused only on the PI3K/AKT/Runx2 axis and did not systematically investigate the involvement of other signaling pathways; therefore, the possibility that Dio exerts integrated effects through the coordination of multiple pathways or interactions with various molecules cannot be excluded. In the future, it will be necessary to further elucidate the crosstalk between the PI3K/AKT/Runx2 pathway and Wnt/β-catenin, among others, from the perspective of multi-pathway and multi-target interaction networks, so as to gain a more comprehensive understanding of the mechanisms of Dio. This will also provide a theoretical basis for the combined application of Dio with other drugs or bioactive factors, as well as for precise interventions targeting specific signaling axes.
In summary, this study indicates that Dio activates the PI3K/AKT pathway and upregulates Runx2, which in turn contributes to enhanced proliferation, osteogenic differentiation, migration, mitophagy, and cell cycle progression of hDPSCs. Our findings not only support the considerable promise of Dio in dental pulp regeneration and bone tissue engineering but also suggest its multifaceted molecular mechanisms. This work provides an experimental and theoretical basis for the potential application of Dio, a natural bioactive compound, in promoting the differentiation and mineralization of DPSCs.
Acknowledgement:
Funding Statement: The authors received no specific funding for this study.
Author Contributions: Zhiye Zhou: Conceived and designed the research, conducted experiments, and analyzed data. Drafted and revised the manuscript critically for important intellectual content. Jianan Chen: Contributed to the acquisition, analysis, and interpretation of data. Provided substantial intellectual input during the drafting and revision of the manuscript. Qiang Zhu: Participated in the conception and design of the study. Played a key role in data interpretation and manuscript preparation. All authors reviewed and approved the final version of the manuscript.
Availability of Data and Materials: The data that support the findings of this study are available from the corresponding author [Qiang Zhu], upon reasonable request.
Ethics Approval: Not applicable.
Conflicts of Interest: The authors declare no conflicts of interest.
Supplementary Materials: The supplementary material is available online at https://www.techscience.com/doi/10.32604/biocell.2026.076758/s1.
Abbreviations
| Abbreviations | Full name |
| DPSCs | Dental Pulp Stem Cells |
| PI3K/AKT | Phosphoinositide 3-kinase/Protein Kinase B |
| Runx2 | Runt-related transcription factor 2 |
| Dio | Dioscin |
| hPDLSCs | Human Periodontal Ligament Stem Cells |
| ARS | Alizarin Red S |
| ALP | Alkaline Phosphatase |
| PCC | Pearson’s correlation coefficient |
| OCN | Osteocalcin |
| DSPP | Dentin Sialophosphoprotein |
| p62 | Sequestosome 1 |
| PINK1 | PTEN-induced putative kinase 1 |
| PRKN | Parkin RBR E3 ubiquitin-protein ligase |
| LC3II/I | Microtubule-associated protein 1A/1B-light chain 3 |
| MSCs | Mesenchymal Stem Cells |
| ECM | Extracellular Matrix |
| 3-MCPD | 3-Chloro-1, 2-propanediol |
| KPNB1-ATF4 | Importin subunit beta1-Activating transcription factor 4 |
| ROS | Reactive Oxygen Species |
References
1. Cao LL , Zhang YJ , Wang JW , Tian F , Wang CF . Studies on microRNA regulation of multidirectional differentiation of dental pulp stem cells: a narrative review. Eur Rev Med Pharmacol Sci. 2022; 26( 6): 1816– 24. doi:10.26355/eurrev_202203_28325. [Google Scholar] [CrossRef]
2. Ahmed GM , Abouauf EA , AbuBakr N , Fouad AM , Dörfer CE , Fawzy El-Sayed KM . Cell-based transplantation versus cell homing approaches for pulp-dentin complex regeneration. Stem Cells Int. 2021; 2021: 8483668. doi:10.1155/2021/8483668. [Google Scholar] [CrossRef]
3. Nowwarote N , Petit S , Ferre FC , Dingli F , Laigle V , Loew D , et al. Extracellular matrix derived from dental pulp stem cells promotes mineralization. Front Bioeng Biotechnol. 2022; 9: 740712. doi:10.3389/fbioe.2021.740712. [Google Scholar] [CrossRef]
4. Xie H , Lin Y , Fang F . Glycogen synthase kinase-3β inhibitor promotes the migration and osteogenic differentiation of rat dental pulp stem cells via the β-catenin/PI3K/Akt signaling pathway. J Dent Sci. 2022; 17( 2): 802– 10. doi:10.1016/j.jds.2021.09.035. [Google Scholar] [CrossRef]
5. Dong Q , Zhang H , Zhang Q , Fei X , Ruan J , He L . Dimethyloxalylglycine regulates osteogenesis of dental pulp stem cells through PI3K/AKT signaling pathways. Tissue Cell. 2025; 96: 103012. doi:10.1016/j.tice.2025.103012. [Google Scholar] [CrossRef]
6. Xu S , Xie X , Li C , Liu Z , Zuo D . Micromolar sodium fluoride promotes osteo/odontogenic differentiation in dental pulp stem cells by inhibiting PI3K/AKT pathway. Arch Oral Biol. 2021; 131: 105265. doi:10.1016/j.archoralbio.2021.105265. [Google Scholar] [CrossRef]
7. Gao F , Xia SL , Wang XH , Zhou XX , Wang J . Cornuside I promoted osteogenic differentiation of bone mesenchymal stem cells through PI3K/Akt signaling pathway. J Orthop Surg Res. 2021; 16( 1): 397. doi:10.1186/s13018-021-02508-0. [Google Scholar] [CrossRef]
8. Kajiura K , Umemura N , Ohkoshi E , Ohta T , Kondoh N , Kawano S . Shikonin induces odontoblastic differentiation of dental pulp stem cells via AKT-mTOR signaling in the presence of CD44. Connect Tissue Res. 2021; 62( 6): 689– 97. doi:10.1080/03008207.2020.1865937. [Google Scholar] [CrossRef]
9. Xu Y , Chen S , Huang L , Han W , Shao Y , Chen M , et al. Epimedin C alleviates glucocorticoid-induced suppression of osteogenic differentiation by modulating PI3K/AKT/RUNX2 signaling pathway. Front Pharmacol. 2022; 13: 894832. doi:10.3389/fphar.2022.894832. [Google Scholar] [CrossRef]
10. Pan H , Yang Y , Xu H , Jin A , Huang X , Gao X , et al. The odontoblastic differentiation of dental mesenchymal stem cells: molecular regulation mechanism and related genetic syndromes. Front Cell Dev Biol. 2023; 11: 1174579. doi:10.3389/fcell.2023.1174579. [Google Scholar] [CrossRef]
11. Yazdankhah M , Ghosh S , Shang P , Stepicheva N , Hose S , Liu H , et al. BNIP3L-mediated mitophagy is required for mitochondrial remodeling during the differentiation of optic nerve oligodendrocytes. Autophagy. 2021; 17( 10): 3140– 59. doi:10.1080/15548627.2020.1871204. [Google Scholar] [CrossRef]
12. Gu Y , Zhou Q , Sheng S , Yang H , Huang D , Zhang Q , et al. Bone remodeling stimulated by Wnt-mediated mitophagy regulated extracellular vesicles in subchondral bone contributes to osteoarthritis development. Theranostics. 2025; 15( 18): 10007– 27. doi:10.7150/thno.111724. [Google Scholar] [CrossRef]
13. Lee W , Eo SR , Choi JH , Kim YM , Nam MH , Seo YK . The osteogenic differentiation of human dental pulp stem cells through G0/G1 arrest and the p-ERK/runx-2 pathway by sonic vibration. Int J Mol Sci. 2021; 22( 18): 10167. doi:10.3390/ijms221810167. [Google Scholar] [CrossRef]
14. Wijnen R , Pecoraro C , Carbone D , Fiuji H , Avan A , Peters GJ , et al. Cyclin dependent kinase-1 (CDK-1) inhibition as a novel therapeutic strategy against pancreatic ductal adenocarcinoma (PDAC). Cancers. 2021; 13( 17): 4389. doi:10.3390/cancers13174389. [Google Scholar] [CrossRef]
15. Muhr J , Hagey DW . The cell cycle and differentiation as integrated processes: cyclins and CDKs reciprocally regulate Sox and Notch to balance stem cell maintenance. Bioessays. 2021; 43( 7): e2000285. doi:10.1002/bies.202000285. [Google Scholar] [CrossRef]
16. Wang M , Zhang Y , Ni S , Sun M , Wu Q , Wu X , et al. The anti-cancer activity of Dioscin: an update and future perspective. Med Oncol. 2025; 42( 3): 63. doi:10.1007/s12032-024-02572-6. [Google Scholar] [CrossRef]
17. Cong S , Peng Q , Cao L , Yi Q , Liu Y , Li L , et al. Diosgenin prevents periodontitis by inhibiting inflammation and promoting osteogenic differentiation. Oral Dis. 2024; 30( 4): 2497– 510. doi:10.1111/odi.14708. [Google Scholar] [CrossRef]
18. You M , Jing J , Tian D , Qian J , Yu G . Dioscin stimulates differentiation of mesenchymal stem cells towards hypertrophic chondrocytes in vitro and endochondral ossification in vivo. Am J Transl Res. 2016; 8( 9): 3930– 8. [Google Scholar]
19. Zhou P , Shi JM , Song JE , Han Y , Li HJ , Song YM , et al. Establishing a deeper understanding of the osteogenic differentiation of monolayer cultured human pluripotent stem cells using novel and detailed analyses. Stem Cell Res Ther. 2021; 12( 1): 41. doi:10.1186/s13287-020-02085-9. [Google Scholar] [CrossRef]
20. Wan HY , Shin RLY , Chen JCH , Assunção M , Wang D , Nilsson SK , et al. Dextran sulfate-amplified extracellular matrix deposition promotes osteogenic differentiation of mesenchymal stem cells. Acta Biomater. 2022; 140: 163– 77. doi:10.1016/j.actbio.2021.11.049. [Google Scholar] [CrossRef]
21. da Silva Sasso GR , Florencio-Silva R , de Pizzol-Júnior JP , Gil CD , de Jesus Simões M , Sasso-Cerri E , et al. Additional insights into the role of osteocalcin in osteoblast differentiation and in the early steps of developing alveolar process of rat molars. J Histochem Cytochem. 2023; 71( 12): 689– 708. doi:10.1369/00221554231211630. [Google Scholar] [CrossRef]
22. Gao Y , Lin Y , Li Y , Zeng W , Chen Z . Interplay of RUNX2 and KLF4 in initial commitment of odontoblast differentiation. J Cell Biochem. 2024; 125( 7): e30577. doi:10.1002/jcb.30577. [Google Scholar] [CrossRef]
23. Hojo H , Ohba S . Runt-related transcription factors and gene regulatory mechanisms in skeletal development and diseases. Curr Osteoporos Rep. 2023; 21( 5): 485– 92. doi:10.1007/s11914-023-00808-4. [Google Scholar] [CrossRef]
24. Nakayama T , Suzuki K , Mitsutake N . Calorie restriction in radiation-exposed mice affects the expression of autophagy-related protein p62. BMC Cancer. 2025; 25( 1): 1388. doi:10.1186/s12885-025-14771-z. [Google Scholar] [CrossRef]
25. Nguyen TT , Wei S , Nguyen TH , Jo Y , Zhang Y , Park W , et al. Mitochondria-associated programmed cell death as a therapeutic target for age-related disease. Exp Mol Med. 2023; 55( 8): 1595– 619. doi:10.1038/s12276-023-01046-5. [Google Scholar] [CrossRef]
26. Han R , Liu Y , Li S , Li XJ , Yang W . PINK1-PRKN mediated mitophagy: differences between in vitro and in vivo models. Autophagy. 2023; 19( 5): 1396– 405. doi:10.1080/15548627.2022.2139080. [Google Scholar] [CrossRef]
27. Chen T , Tu S , Ding L , Jin M , Chen H , Zhou H . The role of autophagy in viral infections. J Biomed Sci. 2023; 30( 1): 5. doi:10.1186/s12929-023-00899-2. [Google Scholar] [CrossRef]
28. Kubickova A , De Sanctis JB , Hajduch M . Isoform-directed control of c-myc functions: understanding the balance from proliferation to growth arrest. Int J Mol Sci. 2023; 24( 24): 17524. doi:10.3390/ijms242417524. [Google Scholar] [CrossRef]
29. Jovic D , Yu Y , Wang D , Wang K , Li H , Xu F , et al. A brief overview of global trends in MSC-based cell therapy. Stem Cell Rev Rep. 2022; 18( 5): 1525– 45. doi:10.1007/s12015-022-10369-1. [Google Scholar] [CrossRef]
30. Peng Y , Jiang H , Zuo HD . Factors affecting osteogenesis and chondrogenic differentiation of mesenchymal stem cells in osteoarthritis. World J Stem Cells. 2023; 15( 6): 548– 60. doi:10.4252/wjsc.v15.i6.548. [Google Scholar] [CrossRef]
31. Cui Y , Ji W , Gao Y , Xiao Y , Liu H , Chen Z . Single-cell characterization of monolayer cultured human dental pulp stem cells with enhanced differentiation capacity. Int J Oral Sci. 2021; 13( 1): 44. doi:10.1038/s41368-021-00140-6. [Google Scholar] [CrossRef]
32. Li B , Ouchi T , Cao Y , Zhao Z , Men Y . Dental-derived mesenchymal stem cells: state of the art. Front Cell Dev Biol. 2021; 9: 654559. doi:10.3389/fcell.2021.654559. [Google Scholar] [CrossRef]
33. Wawrzyniak A , Balawender K . Structural and metabolic changes in bone. Animals. 2022; 12( 15): 1946. doi:10.3390/ani12151946. [Google Scholar] [CrossRef]
34. Vallecillo-Rivas M , Fernández-Romero E , Pérez-Segura M , Toledano R , Amar-Zetouni A , Toledano M , et al. Efficacy of topical application of corticosteroids in the remineralization of dental pulp tissue. A systematic review of the literature. J Dent. 2024; 150: 105333. doi:10.1016/j.jdent.2024.105333. [Google Scholar] [CrossRef]
35. Lampiasi N . The migration and the fate of dental pulp stem cells. Biology. 2023; 12( 5): 742. doi:10.3390/biology12050742. [Google Scholar] [CrossRef]
36. Kim Y , Park HJ , Kim MK , Kim YI , Kim HJ , Bae SK , et al. Naringenin stimulates osteogenic/odontogenic differentiation and migration of human dental pulp stem cells. J Dent Sci. 2023; 18( 2): 577– 85. doi:10.1016/j.jds.2022.08.029. [Google Scholar] [CrossRef]
37. Pei W , Huang X , Ni B , Zhang R , Niu G , You H . Selective STAT3 inhibitor alantolactone ameliorates osteoarthritis via regulating chondrocyte autophagy and cartilage homeostasis. Front Pharmacol. 2021; 12: 730312. doi:10.3389/fphar.2021.730312. [Google Scholar] [CrossRef]
38. Wang RC , Wang Z . Synchronization of cultured cells to G1, S, G2, and M phases by double thymidine block. In: Cell-cycle synchronization. New York, NY, USA: Springer; 2022. p. 61– 71. doi:10.1007/978-1-0716-2736-5_5. [Google Scholar] [CrossRef]
39. Milletti G , Colicchia V , Cecconi F . Cyclers’ kinases in cell division: from molecules to cancer therapy. Cell Death Differ. 2023; 30( 9): 2035– 52. doi:10.1038/s41418-023-01196-z. [Google Scholar] [CrossRef]
40. Liu C , Yu J , Liu B , Liu M , Song G , Zhu L , et al. BACH1 regulates the proliferation and odontoblastic differentiation of human dental pulp stem cells. BMC Oral Health. 2022; 22( 1): 536. doi:10.1186/s12903-022-02588-2. [Google Scholar] [CrossRef]
41. Wang Z . Regulation of cell cycle progression by growth factor-induced cell signaling. Cells. 2021; 10( 12): 3327. doi:10.3390/cells10123327. [Google Scholar] [CrossRef]
42. Zhu J , Zheng Z , Yin Z , Ding L , Li C , Wang X , et al. miR-146b overexpression promotes bladder cancer cell growth via the SMAD4/C-MYC/Cyclin D1 axis. Front Oncol. 2025; 15: 1565638. doi:10.3389/fonc.2025.1565638. [Google Scholar] [CrossRef]
43. Zhong Y , Sun D , Yao Y , Liu Q , Guo T , Wang X , et al. Autophagy and mitochondrial dynamics contribute to the protective effect of diosgenin against 3-MCPD induced kidney injury. Chem Biol Interact. 2022; 355: 109850. doi:10.1016/j.cbi.2022.109850. [Google Scholar] [CrossRef]
44. Behera J , Ison J , Tyagi A , Mbalaviele G , Tyagi N . Mechanisms of autophagy and mitophagy in skeletal development, diseases and therapeutics. Life Sci. 2022; 301: 120595. doi:10.1016/j.lfs.2022.120595. [Google Scholar] [CrossRef]
45. Prateeksha P , Naidu P , Das M , Barthels D , Das H . KLF2 regulates neural differentiation of dental pulp-derived stem cells by modulating autophagy and mitophagy. Stem Cell Rev Rep. 2023; 19( 8): 2886– 900. doi:10.1007/s12015-023-10607-0. [Google Scholar] [CrossRef]
46. Lin Q , Chen J , Gu L , Dan X , Zhang C , Yang Y . New insights into mitophagy and stem cells. Stem Cell Res Ther. 2021; 12( 1): 452. doi:10.1186/s13287-021-02520-5. [Google Scholar] [CrossRef]
47. Ziegler DV , Huber K , Fajas L . The intricate interplay between cell cycle regulators and autophagy in cancer. Cancers. 2021; 14( 1): 153. doi:10.3390/cancers14010153. [Google Scholar] [CrossRef]
48. Matthews HK , Bertoli C , de Bruin RAM . Cell cycle control in cancer. Nat Rev Mol Cell Biol. 2022; 23( 1): 74– 88. doi:10.1038/s41580-021-00404-3. [Google Scholar] [CrossRef]
49. Yu JSL , Cui W . Proliferation, survival and metabolism: the role of PI3K/AKT/mTOR signalling in pluripotency and cell fate determination. Development. 2016; 143( 17): 3050– 60. doi:10.1242/dev.137075. [Google Scholar] [CrossRef]
50. E L , Shan Y , Luo Y , Feng L , Dai Y , Gao M , et al. Insulin promotes the bone formation capability of human dental pulp stem cells through attenuating the IIS/PI3K/AKT/mTOR pathway axis. Stem Cell Res Ther. 2024; 15( 1): 227. doi:10.1186/s13287-024-03843-9. [Google Scholar] [CrossRef]
51. Wu S , Zhao F , Zhao J , Li H , Chen J , Xia Y , et al. Dioscin improves postmenopausal osteoporosis through inducing bone formation and inhibiting apoptosis in ovariectomized rats. Biosci Trends. 2019; 13( 5): 394– 401. doi:10.5582/bst.2019.01186. [Google Scholar] [CrossRef]
52. Zhuo Y , Li M , Jiang Q , Ke H , Liang Q , Zeng LF , et al. Evolving roles of natural terpenoids from traditional Chinese medicine in the treatment of osteoporosis. Front Endocrinol. 2022; 13: 901545. doi:10.3389/fendo.2022.901545. [Google Scholar] [CrossRef]
53. Nasir NJN , Arifin N , Ahmad Amin Noordin KB , Yusop N . PI3K/AKT signaling mediate collagen type 1-induced osteogenic differentiation of dental pulp stem cells via focal adhesion mechanism. J Appl Oral Sci. 2025; 33: e20250173. doi:10.1590/1678-7757-2025-0173. [Google Scholar] [CrossRef]
54. Gao S , Chen B , Zhu Z , Du C , Zou J , Yang Y , et al. PI3K-Akt signaling regulates BMP2-induced osteogenic differentiation of mesenchymal stem cells (MSCs): a transcriptomic landscape analysis. Stem Cell Res. 2023; 66: 103010. doi:10.1016/j.scr.2022.103010. [Google Scholar] [CrossRef]
55. Ge Y , Ding S , Feng J , Du J , Gu Z . Diosgenin inhibits Wnt/β-catenin pathway to regulate the proliferation and differentiation of MG-63 cells. Cytotechnology. 2021; 73( 2): 169– 78. doi:10.1007/s10616-021-00454-7. [Google Scholar] [CrossRef]
Cite This Article
 Copyright © 2026 The Author(s). Published by Tech Science Press.
Copyright © 2026 The Author(s). Published by Tech Science Press.This work is licensed under a Creative Commons Attribution 4.0 International License , which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.


 Submit a Paper
Submit a Paper Propose a Special lssue
Propose a Special lssue View Full Text
View Full Text Download PDF
Download PDF Downloads
Downloads
 Citation Tools
Citation Tools