 Open Access
Open Access
ARTICLE
NeuroTriad-ViT: A Scalable and Interpretable Framework for Multi-Class Brain Tumor Classification via MRI and Knowledge Distillation
1 School of Computer Science and Engineering, Central South University, Changsha, China
2 Department of Computer Science, College of Computers and Information Technology, Taif University, Taif, Saudi Arabia
3 Electrical Engineering Department, University of Business and Technology, Jeddah, Saudi Arabia
4 Department of Electrical and Computer Engineering, North South University, Dhaka, Bangladesh
* Corresponding Authors: Zuping Zhang. Email: ; Ahmed Emara. Email:
Computers, Materials & Continua 2026, 87(3), 74 https://doi.org/10.32604/cmc.2026.076402
Received 20 November 2025; Accepted 28 January 2026; Issue published 09 April 2026
Abstract
The effective diagnosis and treatment planning require the correct classification of the cerebral neoplasia, such as glioma, meningioma, and pituitary tumors. The recent developments in the deep learning field have made a significant contribution to the field of image analysis in medicine; however, Vision Transformers (ViTs) have achieved good results but are computationally complex. This paper presents NeuroTriad-ViT, a proprietary large-scale Vision Transformer of 235 million parameters, which is represented as a high-performance teacher model to classify brain tumors. Knowledge distillation is applied in an attempt to transfer the representations that the teacher learned to lightweight student models, in which MobileNetV2 outperformed EfficientNet0-Lite. The models were conditioned on 24,455 MRI scans, which were combined from three publicly available datasets. The CNN-pretrained ViT-based hybrid architecture was the highest-accuracy heterogeneous hybrid model reported as teacher-nominated, with 96% accuracy, and the CNN -architecture-based ensemble models (using EfficientNet, VGG16, and DenseNet) had a maximum accuracy of 92%–95%. Comparatively, the NeuroTriad-ViT model had an accuracy of 98% and the distilled MobileNetV2 model had an accuracy of 99.32%, thus it can be seen as having better performance with lower computational cost. The interpretability of models was tested based on Grad-CAM and LIME, and the measures of insertion and deletion support the faithfulness of the explanations. Overall, the proposed framework enables efficient, interpretable, and scalable brain tumor diagnosis suitable for real-time clinical and mobile health deployment. The source code is publicly available at https://doi.org/10.5281/zenodo.17494928.Keywords
Accurately diagnosing and effectively managing brain tumors can frequently provide challenges. Globally, over 308,000 brain tumors are diagnosed every year, with an estimated 251,000 deaths due to the tumors annually [1,2]. Even though CNS tumors are not the main cause of cancer, they are disproportionately prevalent in the morbidity and mortality of younger populations, despite their minor percentage, which is less than 2% amongst all cancer types. As an example, the five-year relative survival of primary malignant brain tumors is only 36.6% [2]. It is thus of paramount importance that early and accurate distinction of high- and low-grade gliomas, meningiomas, and pituitary adenomas is made in terms of prognosis and treatment planning [3]. The heterogeneity of the tumors makes it difficult to make a correct diagnosis, and may frequently require more rapid and accurate imaging and analytical tools to do so [4].
Traditionally, the interpretation of neuro imaging studies has been the work of expert radiologists, and it is a process that depends upon years of experience and pattern recognition. Nevertheless, there are many difficulties in inter-rater variability, fatigue, and overlapping features of imaging. An example of such is gliomas, which have a heterogeneous appearance and can resemble other types of tumors, which require high levels of skill to identify them properly [2,4]. Artificial intelligence (AI), specifically deep learning, is one of the possible solutions, as it can deliver objective, quantitative, and accurate information in medical imaging. VGGNet, ResNet, DenseNet, and EfficientNet are convolutional neural networks that have shown expert-level accuracy in brain tumor classification through learning the hierarchical features of MRI images with high accuracy [5]. In spite of these improvements, CNNs cannot capture long-range dependencies and global structure signals important to tumors with complex morphology due to the local receptive fields.
Vision Transformers (ViTs) have changed the paradigm of medical imaging. In contrast to CNNs, ViTs process images as a sequence of patches and use self-attention to learn long-range spatial structure and global context by modeling the long-range structure and considering lots of information simultaneously in a single model [6]. This makes ViTs more efficient than traditional CNNs in various vision problems, such as brain tumor classification, where global structural patterns are needed to capture the classification viewpoint. ViTs are computationally expensive and memory-intensive, which is why they cannot be deployed in clinical practice, in particular, at the edge or on a mobile platform, even though their accuracy remains high [6]. Besides, they are complex, which makes them black-box models and an obstacle to clinical trust and interpretability.
To resolve the trade-off between the model with high capacity and the model with practical deployment, an innovative solution is needed. Knowledge distillation (KD) has become a powerful approach, which allows the learning representations of big teacher models to be transferred to small student models without any fundamental performance degradation in judgment [7]. Such a design can maintain the diagnostic capability of a ViT and allow quick, light, and deployable models that can be used in clinical environments in real-time.
To balance the high representational capacity of modern models with the pragmatic needs in clinical deployment, the present study presents NeuroTriad-ViT, a high-capacity Vision Transformer specifically designed for glioma, meningioma, and pituitary tumor classification. Knowledge distillation is used to extract the diagnostic capability of the large model (teacher) into a small MobileNetV2 (student), allowing for rapid and resource-efficient inference without compromising predictive fidelity. In addition to that, explainable artificial intelligence techniques, such as Gradient-weighted Class Activation Mapping (Grad-CAM), provide visual rationalizations of the outputs produced by a model, which can help build trust among clinicians and also aid in meeting the compliance requirements of the regulatory bodies [8].
The proposed framework provides a clinically deployable, accurate and interpretable solution that provides an appropriate trade-off between model performance and operational usability. By combining state-of-the-art Vision Transformers with knowledge distillation and explainability components, the system demonstrates potential in democratizing access to AI-assisted brain tumor diagnostics, especially those in resource-constrained environments where access to expert radiological know-how and high-performance computing resources is limited [1,5–8].
2.1 Advances in Brain Tumor Classification Using CNNs, Transfer Learning, Feature Fusion, and Ensemble Learning
Recent advances in deep learning, transfer learning, and hybrid models have significantly improved MRI-based brain tumor classification. Wong et al. (2025) [9] developed a data-augmented deep learning system achieving 99.24% accuracy, and Preetha et al. (2025) [10] showed that combining 3B Net with EfficientNet-B2 further enhances performance. Velpula et al. (2025) [11] used a collection of trained convolutional neural networks (CNNs) to obtain a 99.69% accuracy. Conversely, the RBEBT framework presented by Zhu et al. (2023) [12] combines ResNet 18 with randomized neural networks to optimize F-1 scores. By adding the ALCResNet module to the FusionNet architecture to refine their diagnostic grading, Abbas et al. (2025) [13] achieved a better diagnostic grading performance, whereas Ramakrishna et al. (2025) [14] used EfficientNet-B3 to better the performance in terms of accuracy and generalization. Lastly, Nazir et al. (2025) [15] performed an extensive survey on explainable artificial intelligence (XAI) in medical imaging, summarized the main techniques of salient XAI, data, and measurement of evaluation, and provided an overview of future research directions.
Hybrid architectures have also developed classification abilities. Tariq et al. (2025) [16] combined EfficientNet-V2 with Vision Transformers, achieving an accuracy of 96.02% and showing synergistic relations between CNN and transformer models. Shawly and Alsheikhy (2025) [17] put together the NDDRSATALFA structure, which is a combination of VGG-19, U-Net, and Dense Dynamic Residual Self-Attention Fusion module with a 99.32% accuracy level. Aloraini et al. (2023) [18] introduced TECNN, which is a convolutional neural network with a vision transformer, with an accuracy of 99.10% on Figshare and 96.75% on BraTS2018. Naim Islam et al. (2024) [19] adopted a CNN-LSTM hybrid model to learn a temporal feature, obtaining an accuracy up to 98.82%–99%. Haque et al. (2025) [20] created a deep-layered stack with EfficientNet-B0, MobileNetV2, GoogLeNet, and CapsuleNet, achieving a PR-AUC of 98.75% and alleviating the problems of class-imbalance.
Collectively, these studies demonstrate that CNNs, hybridization, attention mechanisms, and ensemble learning consistently achieve near-perfect accuracy, robustness, and clinical applicability in brain tumor classification.
2.2 Advances in Brain Tumor Classification with Vision Transformers
The recent literature in the classification of brain tumors has demonstrated the impressive potential of Vision Transformer (ViT) models, especially with hybrid and ensemble techniques. A ViT-CNN hybrid model presented by Chandraprabha et al. (2025) [21] demonstrated an accuracy of 99.64%, thus demonstrating that the combination of ViT and CNNs improves the extraction of features and the strength of classification. In like manner, Ahmed et al. (2024) [22] came up with a ViT-GRU hybrid that achieved an accuracy of 98.97% with Explainable AI (XAI) to enhance interpretability. The ViT-CB model (Tanone et al., 2025) [23] was the combination of Vision Transformers with PCA-based dimensionality reduction and CatBoost, with good sensitivity, specificity, and overall accuracy, and transparency can be provided by SHAP-based XAI. Ensemble approaches were also considered: Tummala et al. (2022) [24] utilized an ensemble of pre-trained and fine-tuned ViT models and obtained the highest accuracy of 98.70%. FTVT (ViT) models have been proven to be superior to traditional CNNs, while Reddy et al. (2024) [25] demonstrated that FTVT (ViT) models outperformed ResNet50 and MobileNetV2 with the same accuracy, confirming the superiority of ViT architectures over conventional CNNs. Islam et al. (2025) [26] further improved ViT models using XAI approaches, with an accuracy of 98.4%–99.3% and the significance of interpretability in clinical practice. Lastly, Asiri et al. (2024) [27] implemented a Swin Transformer-based method with an accuracy rate of 97%, which is better than traditional CNNs and regular ViT models, thus indicating that transformer variants can be used in the medical image analysis process. Taken together, these works indicate the power of ViT-based architectures, hybridization, ensemble learning, and explainable AI, with high accuracy and improved clinical reliability in brain tumor diagnosis.
2.3 Brain Tumor Classification via Knowledge Distillation: Recent Advances
Recent developments in medical imaging have highlighted the potential basis of change in the area of knowledge distillation (KD) and hybrid AI models to classify and segment brain tumors. Choi et al. (2023) [28] proposed a one-stage KD-based segmentation of brain tumors with limited modalities of MRI with Dice scores of 91.11% and 92.20% for the tumor core and the entire tumor, respectively, thus illustrating that efficient segmentation is feasible with fewer inputs. Continuing on the topic of lightweight architectures, Tan et al. (2024) [29] have created FM-LiteLearn that combines image fusion, model amplification, and KD, enhancing the classification performance by 9.4% and showing better generalization. After finishing the proposed system, Uddin et al. (2025) [30] achieved 97% and 99% accuracy when segmenting data, and using CLIP-ViT, respectively, 97% and 99% accuracy on segmented data, and 99% with CLIP-KDViT, respectively, thus confirming the clinical value of X-ray-based multi-disease diagnosis. Silpa and Satpathy (2024) [31] introduced Knowledge-Distilled ResNeXt as they had 95.3% accuracy and 96.7% F1-score, which suggests the strength of KD in medical image classification. To overcome the privacy issue, Gohari et al. (2024) [32] proposed FedBrain-Distill, a federated learning model that considers KD; the model had a higher performance in non-IID conditions and did not violate patient privacy. Khan et al. (2025) [33] applied KD to Ant Colony Optimization (ACO) and context-sensitive scaling of the temperature, and got very good accuracy on several datasets, thus highlighting the versatility of KD in various medical tasks. Finally, Liu et al. (2023) [34] designed a siamese self-distillation segmentation framework, which does not deteriorate under any missing modality setting, proving to be effective in the case of limited data. Taken together, these results support the efficacy of knowledge distillation (KD), hybrid approaches, and lightweight or federated models in improving the accuracy, efficiency, interpretability, and clinical applicability of AI-based medical image analysis.
This research report delineated the process employed for the classification of brain tumors, as specified in the methods portion of the manuscript. The preprocessing pipeline consisted of several normalization and augmentation steps that tend to enhance the standard of the images and provide dataset balance, thus enabling effective model training. The current research paper outlined the procedure that was used in the categorization of cerebral tumors, as indicated in the methodology section of the manuscript. Several normalization and augmentation processes made up the pretreatment pipeline, which tends to enhance image quality and offer dataset balance, allowing for efficient model training. Knowledge distillation was used to port knowledge on larger, more complex models to smaller models that are computationally efficient, thereby enhancing performance without scaling or efficiency degradation. Following a thorough examination of the models, their accuracy, computing efficiency, and reliability were assessed to assess the models’ capacity for prediction. All minor models do create predictions. Using methods like Gradient-based Local Interpretable Model-agnostic Explanations (LIME) with Class Activation Mapping (Grad-CAM)increased the interpretability. The decision-making process was clarified using these methods, as well as the transparency of model predictions shown in Fig. 1.
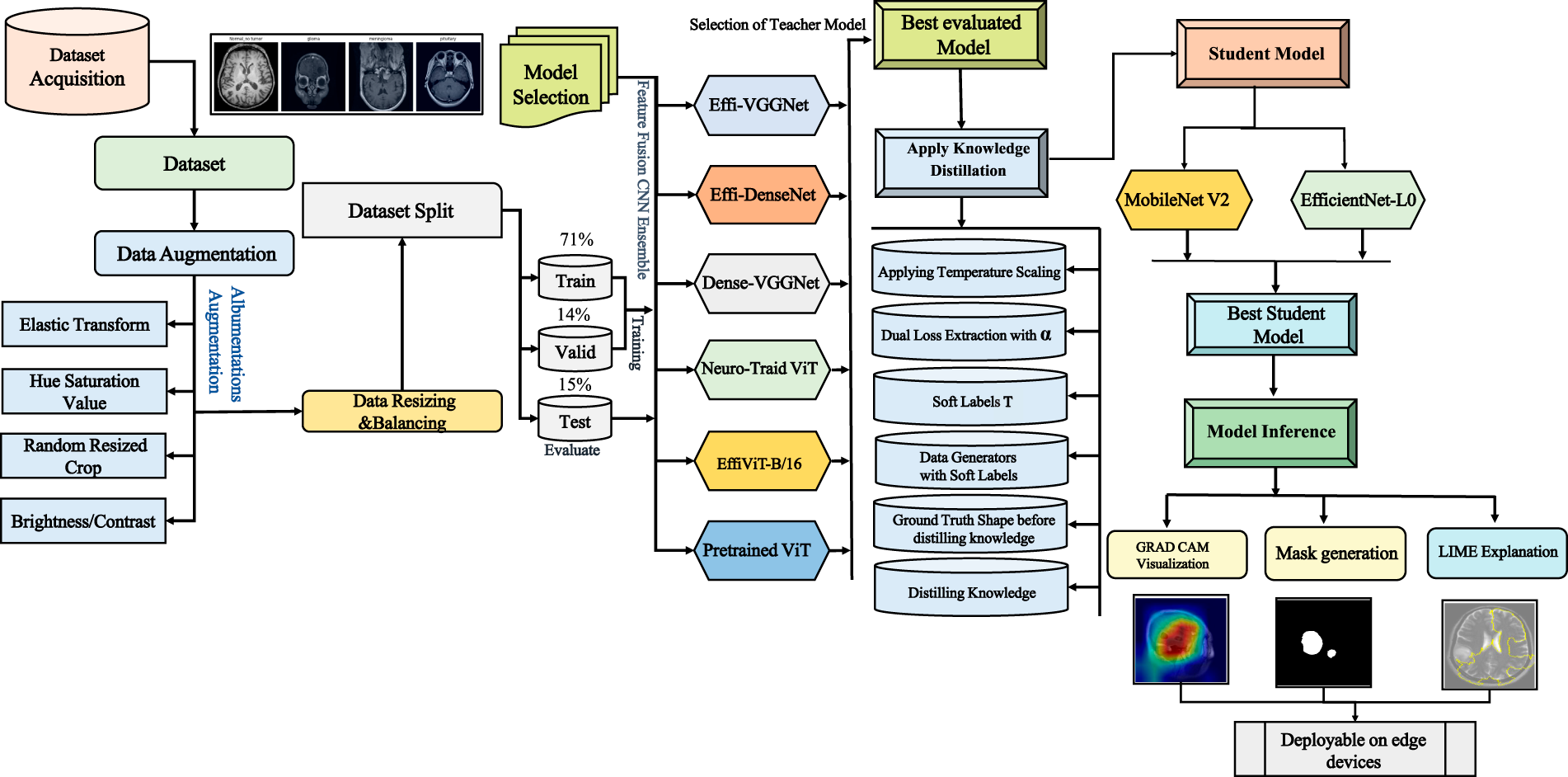
Figure 1: System design and workflow of NeuroTriad-ViT for brain tumor classification. The workflow schematic of the NeuroTriad-ViT illustrates that the system performs processing, classification, visualization, and training to identify an image and determine its optimal label.
3.1.1 Data Collection and Composition
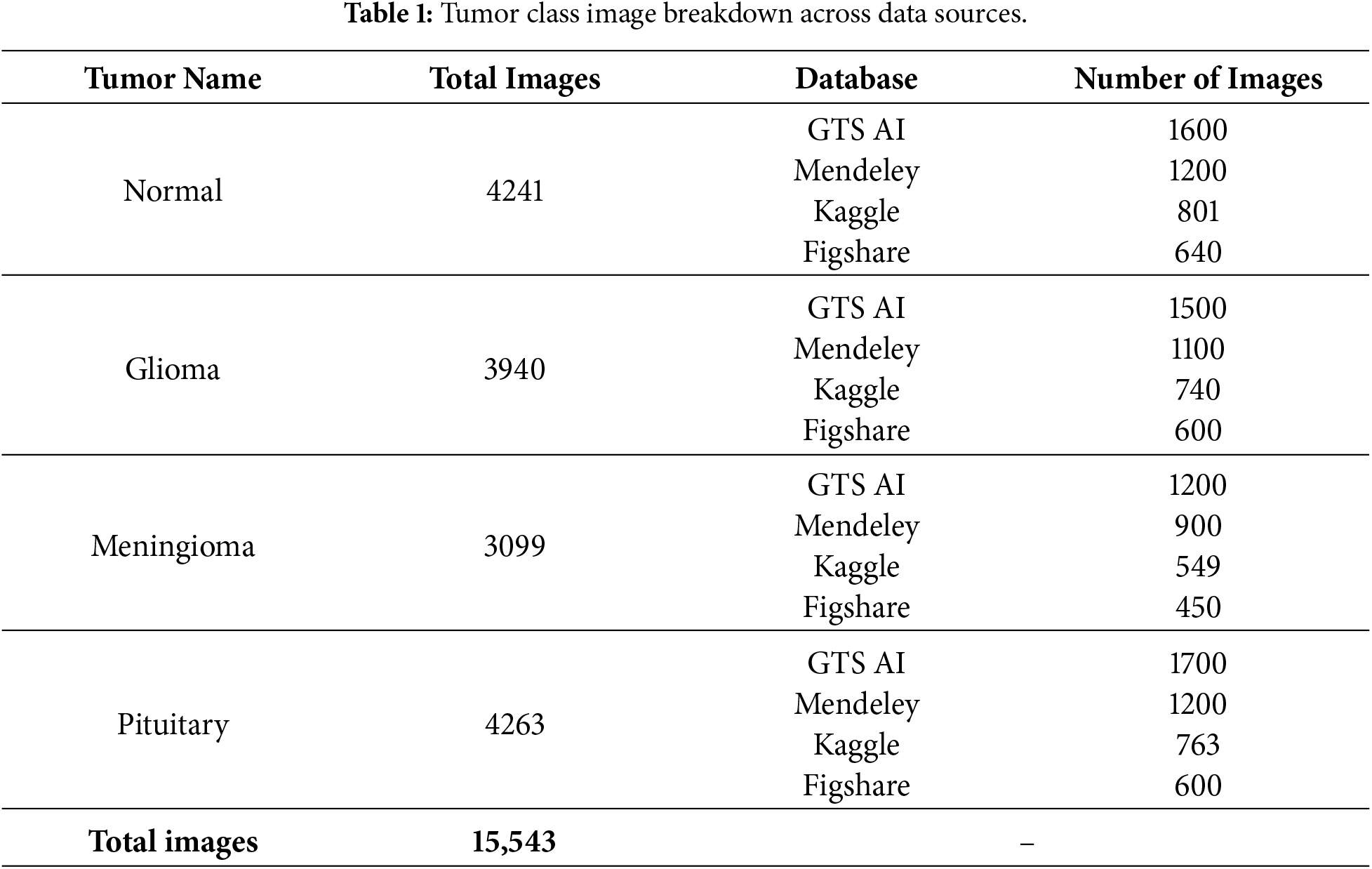
In this research, the dataset was generated by integrating and preprocessing several brain tumor classification datasets. All of them are T1-weighted MRI slices, widely accessible and well-known. Namely, the Mendeley dataset [35], the GTS AI dataset [36], Kaggle(BRISC dataset) [37], and the Figshare dataset [38] were utilized. The resultant dataset includes 15,543 magnetic high-resolution resonance imaging scans of four diagnostic classes: normal, glioma, meningioma, and pituitary tumors, a detailed description showing in the Table 1. Overall, the integration of the three datasets yielded a broad and balanced dataset for training and testing deep learning models to ensure high performance in brain tumor classification tasks.
3.1.2 Data Preprocessing and Downsampling
An extensive data preparation pipeline was implemented to optimize the brain tumor dataset for robust model training. The pixel intensities were normalized to grayscale intensities in the interval
Magnetic resonance images of Pituitary, Normal, Meningioma, and Glioma tumors varied in size from
Overall, this combined preprocessing and downsampling strategy significantly improved data quality, consistency, and interpretability, providing a robust, high-quality, and reproducible dataset for deep learning–based approaches
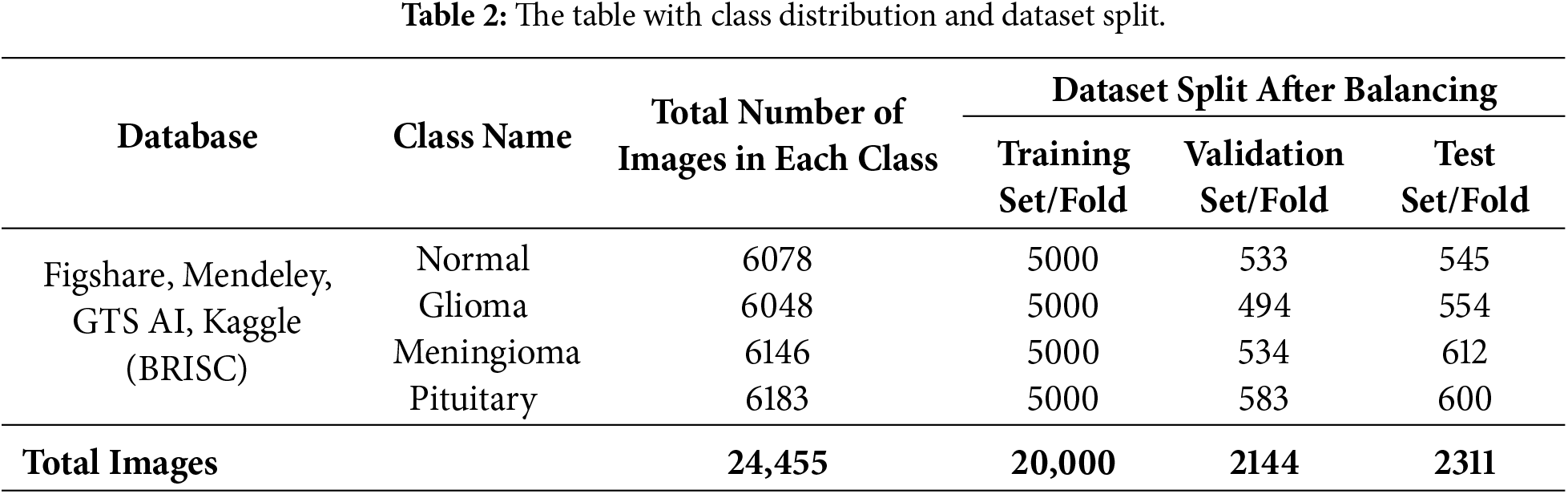
The image dataset of the brain tumor problem was carefully prepared to facilitate the classification. In particular, the class distribution of each tumor phenotype was stratified with respect to the distribution of images in order to maintain the balanced class distribution across all subsets. Table 2, showing a total of 24,555 images data were divided into 72% training, 14% validation, and 15% testing and. Following set best practices in medical imaging, this separation ensures that there is an adequate amount of data for model training and optimization while also maintaining a stable validation set and unbiased test set.
3.1.4 Training Data Augmentation and Balancing
The Albumentations pipeline library was employed to augment the training dataset in order to achieve variability and robustness in the model. Some of the transformations applied were:
• Random Crop (224
• Horizontal & Vertical Flips: Applied with probability
• Random 90° Rotations: Simulate multi-angle views with
• Elastic Transformations: Applied with parameters
• Brightness & Contrast Adjustments: Mimic varying imaging conditions in the range of
• Hue, Saturation & Value Modifications: Simulate lighting variations where Hue shift limited to
• Normalization: Align with CNN and Transformer preprocessing standards with
To obtain a balanced dataset, the underrepresented classes in the train set were increased to reach the target of 5000 images per disease, as shown in Table 3 and totaling of 24,555 images showing in the Table 2. These augmentations, as demonstrated in Fig. 2, minimized overfitting and also increased the diversity of classes using the Albumentations augmentation library pipeline. To achieve the balance, the training set was scaled up to 5000 images per class through random augmentation of the underexploited sample. Table 3, except that the CNNs, ensembles, and ViTs are all subjected to equal learning without favoring any samples [39].
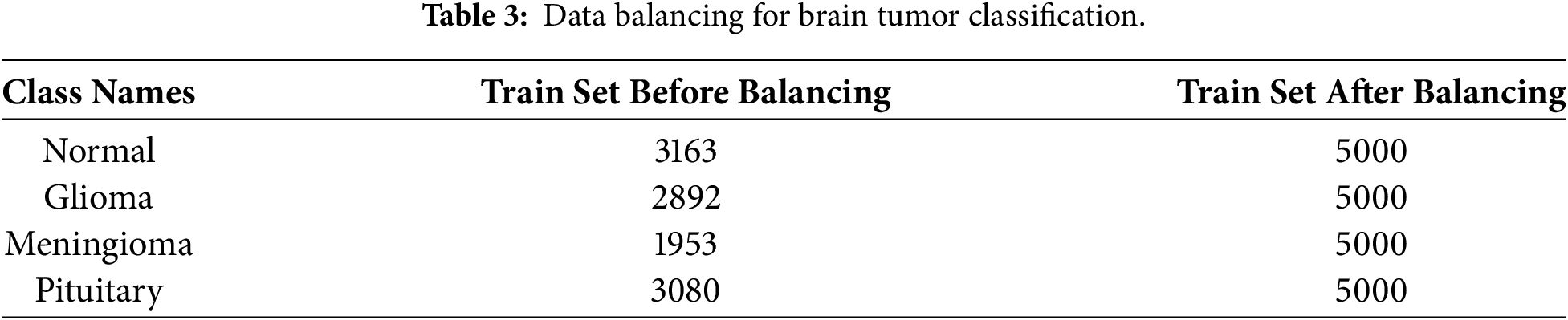
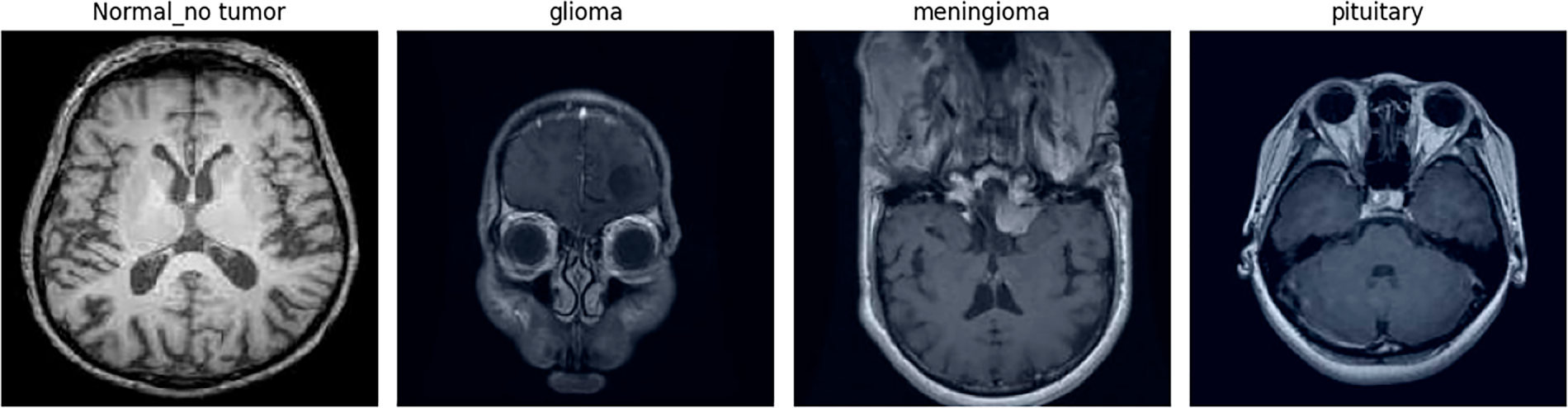
Figure 2: Random samples of each class after augmentation. Representative examples of augmented MRI images, showing variability in orientation, brightness, and scale across tumor categories.
3.2.1 Different CNN Architectures for Type I Heterogeneous Ensemble Models
In this study, several convolutional neural network frameworks were carefully selected as backbone models based on their complementary strengths in feature extraction and representation learning, forming the basis of a heterogeneous ensemble framework.VGG16, comprising 13 convolutional layers and 3 fully connected layers, employs
Moreover, by combining different CNN architectures, this study proposes three heterogeneous ensemble models:
• Effi-VGGNet (VGG16 + EfficientNetB2)
• Effi-DenseNet (DenseNet121 + EfficientNetB2)
• VGG-DenseNet (VGG16 + DenseNet121).
These ensembles leverage the complementary strengths of their constituent networks to enhance feature extraction, improve generalization, and achieve higher accuracy in brain tumor classification.
3.2.2 Type II Heterogeneous Ensemble: EffiViT-B/16
The suggested Type II ensemble provides a hybrid architecture of EffiViT-B/16. The base vision transformer (ViT-B/16) partitions the
The ensemble takes the final output of the two-member networks formulated in Eq. (1) and carries the summation of the final result to a completely connected classification head to apply a Softmax layer to produce the ultimate class probabilities:
Here,
3.2.3 Proposed Vision Transformer (NeuroTriad-ViT) Model Architecture
This study proposes a Model titled the NeuroTriad -ViT model, which is a version of a Vision Transformer that has been specifically adapted to classify brain tumors. In contrast to the traditional CNNs that can hardly capture long-range contextual dependencies, the NeuroTriad-ViT takes advantage of the “self-attention mechanisms that enable the model to capture long-range dependencies and complex patterns in space across the whole image. The GAP-based aggregation reduces dependence on a single token representation and improves robustness to spatially distributed tumor patterns. Architecture, as illustrated in Fig. 3, is mathematically formulated when it comes to multi-class tumor classification as indicated in the Eqs. (2) and (3).
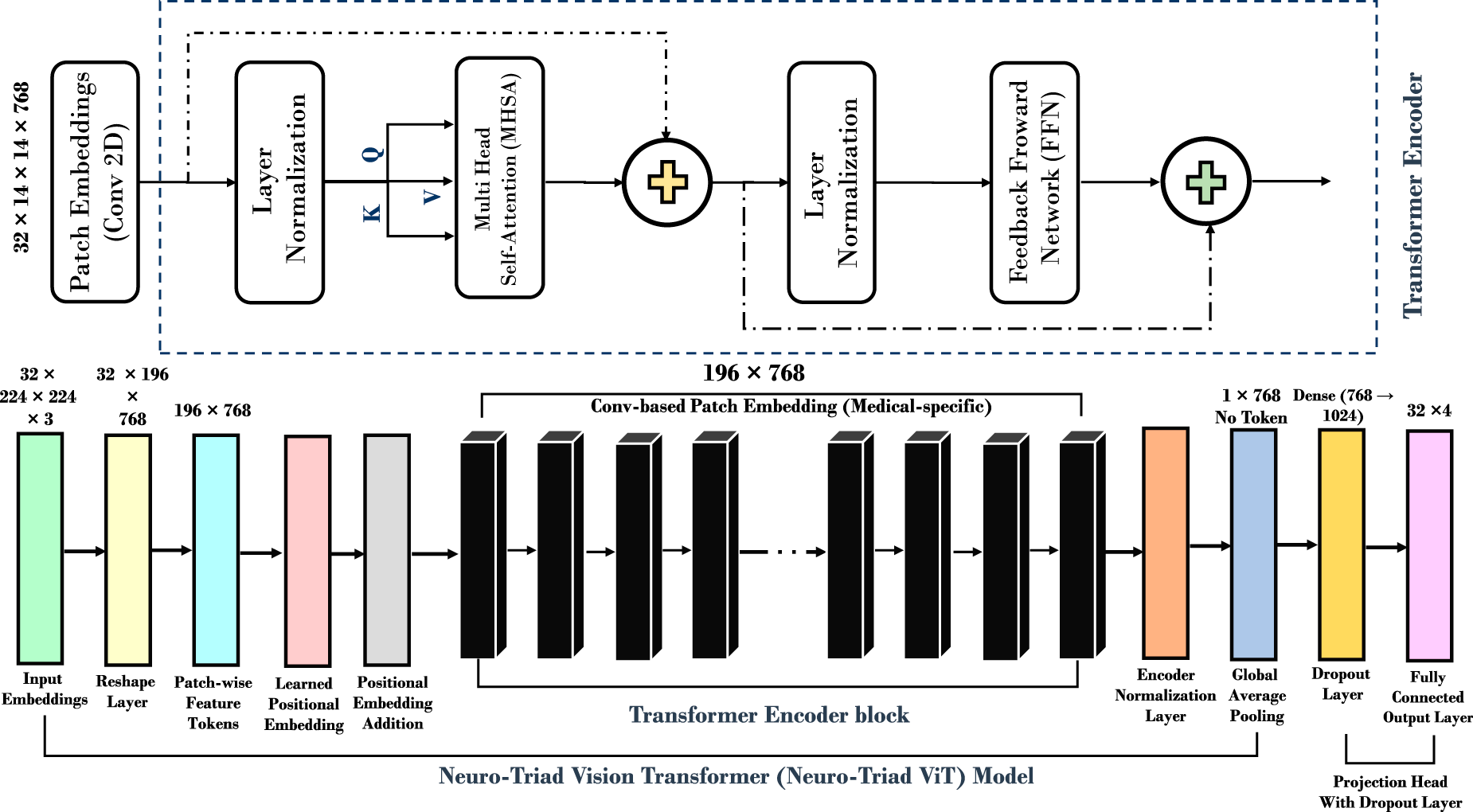
Figure 3: Neuro Triad-ViT: brain tumor classification model architecture. The complete architecture of the proposed NeuroTriad-ViT model illustrates convolution-based patch embedding, transformer encoder blocks, and GAP-based classification for multi-class brain tumor detection.
• Patch Embedding: Patch Embedding: Patch embedding is obtained by dividing the
• Positional Encoding: Positional encodings
• Transformer Layers: The model contains 12 transformer blocks, each comprising multi-head self-attention and feed-forward layers. The attention mechanism computes:
where
• Classification Head: Features from transformer blocks are aggregated using global average pooling (GAP), which replaces the standard class token in ViT-B/16 and allows for a more flexible and spatially aware feature summarization. This is subsequently passed through a fully connected layer of 1024 units with ReLU activation, followed by a dropout layer for regularization to improve learning and reduce overfitting. Two bias terms,
The model output
where
Overall, the proposed NeuroTriad-ViT enhances the standard ViT-B/16 by employing a GAP-based classification head that aggregates features across all patches instead of relying on a [CLS] token, improving sensitivity to distributed tumor patterns. The inclusion of two bias terms (
3.3 Dual-Supervision Knowledge Distillation with Temperature Scaling
In the present study, knowledge distillation (KD) is adopted as a core optimization strategy to reconcile the high diagnostic accuracy of computationally intensive teacher models with the deployment constraints of lightweight student networks in clinical environments [7]. Rather than introducing a novel distillation objective, the work emphasizes a principled, task-specific application of logit-based KD within a heterogeneous teacher–student framework for brain tumor classification. Unlike the traditional knowledge-distillation (KD) schemes, which rely on a single teacher with homogeneous knowledge, the current case evaluates a heterogeneous teacher ensemble that comprises six architectures: CNN-based hybrid models, CNN–Transformer hybrids with the canonical Vision Transformer (ViT-B/16), and a custom-designed Vision Transformer (NeuroTriad-ViT). Empirical validation identifies the diagnostically most reliable teacher, which is then used to reduce knowledge into lightweight student models. Two state-of-the-art student architectures, MobileNetV2 and EfficientNet-Lite0, are analyzed under the same distillation conditions, and the final student architecture is chosen according to the overall performance and efficiency measures.
The distillation protocol follows a two-supervision paradigm, with the student being supervised by both hard ground-truth labels and temperature-scaled soft probability distributions provided by the teacher. Although hard labels maintain diagnostic fidelity, inter-class similarities necessary for separating visually overlapping tumor subsets are contained in the soft labels. More importantly, the framework uses only logit-based distillation to avoid feature- or attention-level transfer, which is a deliberate choice that ensures compatibility between the architectures of heterogeneous teachers and maintains computational efficiency during student training.
Temperature Scaling and Soft Label Generation
To increase the expressiveness of the predictions of the teacher, a temperature parameter T is added to the logits of the teacher before the SoftMax normalization. Increased T also produces smoother probability distributions that reveal latent class relationships and uncertainty patterns learn by the teacher during training. Accordingly, such smooth outputs provide more informative supervisory information than that of one-hot labels, allowing the student model to acquire finer decision boundaries.
Eq. (4) shows that the probabilistic labels generated by the teacher network are computed as:
Dual-Loss Optimization Strategy
The optimization goal for the student is presented in the form of a weighted sum of categorical cross-entropy loss (Eq. (5)) and Kullback-Leibler (KL) divergence loss (Eq. (6)). The cross-entropy term penalizes deviation from the ground truth annotations, while the KL divergence term encourages the student’s softened model’s predictions to be similar to the predictions the teacher model generates. A balancing coefficient, with a multiplier denoted
The categorical cross-entropy loss function is defined as:
The KL divergence loss is given by:
The final distillation objective used to train the student network is expressed in Eq. (7):
3.3.1 Efficient Soft-Label Integration via Dynamic Data Generators
Rather than precomputing and storing soft labels, which is memory-intensive and inflexible, in this study, custom data generators are used, and these generators dynamically create the teacher predictions during training. This on-the-fly soft-label generation allows dual supervision to be easily supervised without any additional storage space, thereby saving a large amount of memory and making it simple to integrate the pipeline. Furthermore, this design enables the framework used for distillation to be reactivated/adapted to changing conditions without retraining/re-generating auxiliary dataset (s).
3.3.2 Lightweight Student Models for Knowledge Distillation
MobileNetV2 and EfficientNet-Lite0 are employed as lightweight student architectures to evaluate the effectiveness of knowledge distillation under deployment-oriented constraints. MobileNetV2 can maintain an effective tradeoff between the representational and computational cost through the use of incorporating the depthwise separable convolutions, inverted residual blocks, and linear bottleneck layers, making it a very viable choice in low-latency medical imaging inference. Additionally, EfficientNet-Lite0, another state-of-the-art lightweight architecture, tuned with the help of compound scaling to achieve even better accuracy efficiency trade-offs at a very low parameter cost. Its simplicity in streamlining design and a strong representation of features make it especially suitable in clinical environments that are cost-limited.
Altogether, although a traditional description of the distillation loss was used, the combination of various lightweight student models and metric-guided selection schemes strengthens the methodology and supports the ability of the presented framework to deliver medical-image classification efficiently.
3.4 Performance Evaluation Metrics
The models of the deep-learning performance was quantified using accuracy, precision, recall, F1-score, and ROC-AUC as evaluation criteria, and interpretability by means of Grad-CAM, LIME, Faithfulness (Insertion and Deletion), and inference analysis. Accuracy is a measure of correctness (ratio) overall that is calculated using the counts of true positives (TP), true negatives (TN), false positives (FP), and false negatives (FN). Precision, also known as the positive predictive value (PPV), refers to the percentage of the predicted positives that are the ones that are rightly identified. Recall, also known as sensitivity, is the fraction of actual positives correctly reported.
The F1-score is defined as the harmonic mean of Precision and Recall, with values approaching unity indicating strong model performance. Finally, ROC-AUC evaluates the model’s discriminative capability by measuring the area under the Receiver Operating Characteristic curve, where values near one signify excellent class separation.
3.5 Model Interpretability Using Grad-CAM and LIME Explanation
Model interpretability is achieved by employing Gradient-Weighted Class Activation Mapping (Grad-CAM) and Local Interpretable Model-Agnostic Explanations (LIME) to identify salient regions within the images that most strongly influence the model’s predictions. Grad-CAM identifies class-discriminative localization maps by weighting the final convolutional feature maps according to the gradients of the target class, as expressed in Eq. (8):
where
Complementarily, LIME explains model behavior by approximating the classifier locally using an interpretable surrogate model over perturbed superpixels. Region-wise attributions are obtained by solving the optimization problem in Eq. (9):
To quantitatively assess explanation reliability, tumor region masks are employed exclusively during evaluation to compute faithfulness scores based on insertion and deletion metrics. These masks serve as spatial references to verify whether regions highlighted by Grad-CAM and LIME are causally aligned with the model’s predictions and are not utilized during training or inference. Faithfulness is measured as the difference in prediction confidence when salient pixels are progressively added (insertion) or removed (deletion), as defined in Eq. (10):
where
This section compares the NeuroTriad-ViT teacher architecture and its knowledge distilled lightweight student models (e.g., MobileNetV2 and EfficientNet-Lite0) in the example of brain-tumor classification. The findings depict the great diagnostic accuracy of the Vision Transformer teacher and the performance improvements achieved with the help of knowledge distillation. A detailed analysis of model metrics, interpretability results, and drawbacks in comparison with modern hybrid CNN approaches demonstrates that the selected student model is an efficient and stable compromise in terms of accuracy, computational power, and readiness for deployment, and thus highlights the credibility and feasible applicability of the framework.
4.1 Hyperparameter Optimization and Training Protocols
Hyperparameter selection was critical to achieve robust convergence, reliable generalization, and reproducibility across all models. The ensemble CNNs (Effi-VGGNet, Effi-DenseNet, VGG-DenseNet) were trained with a learning rate of
The hybrid EffiViT-B/16 model combined CNN and Transformer components via penultimate-layer concatenation, using a similar learning rate, dropout probability of 0.5, and pretrained weights for both components. The ViT-B/16 model employed a lower learning rate of
The NeuroTriad-ViT teacher model was trained with a linear warm-up schedule from an initial learning rate of
For knowledge-distillation (KD) optimized student models, the MobileNetV2 student was trained with a temperature
4.2 The Emergence of a Mentor Model: Comparative Analysis and Teacher Selection for Transformational Medical AI
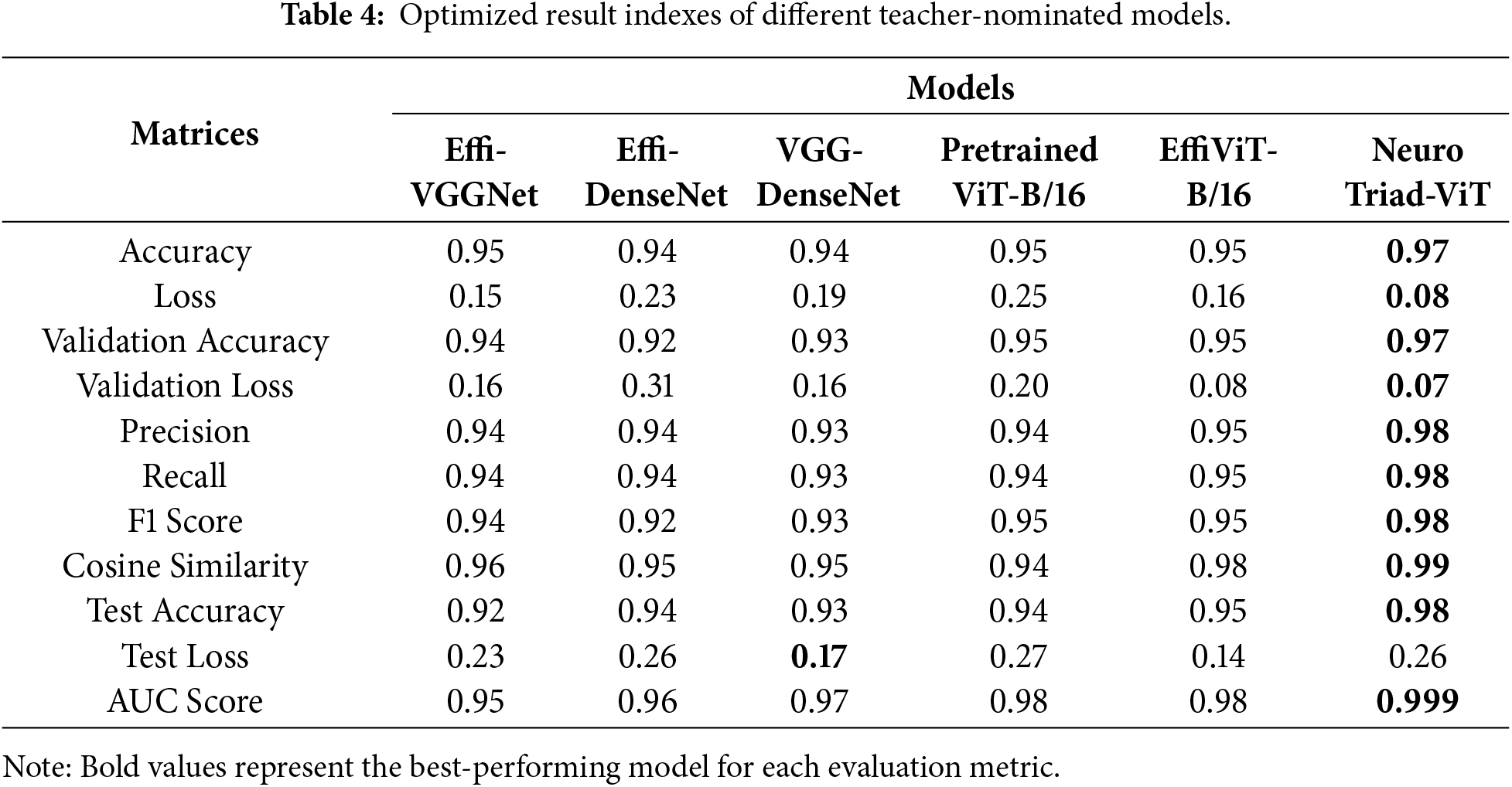
This sub-topic will make a comparative study of six of the evolved architectures that are employed to classify brain neoplasm in terms of the optimal trade-off between diagnostic performance and computational efficiency. Table 4, provides a summary of the state-of-the-art models hybrid CNNs and Vision Transformer (ViTs) and demonstrates that both the transformer-based and the hybrid designs achieve high quantitative results, whereas qualitative analyses reveal a significant improvement in feature representation and discriminative power.
Neuro Triad-ViT emerges as the overall leader, attaining training and test accuracies of 97% and 98%, respectively, shown in Fig. 4. Also, Fig. 5, demonstrates that it achieves the lowest cross-entropy losses and an AUC of 0.999; the ROC further demonstrates near-perfect class separability (high TPR, low FPR). Its strong calibration at challenging multi-class decision boundaries is corroborated by precision, recall, F1-Score = 0.98, and by a cosine similarity of 0.99 to the ground-truth label distributions—an attribute critical for high-stakes diagnostic systems.
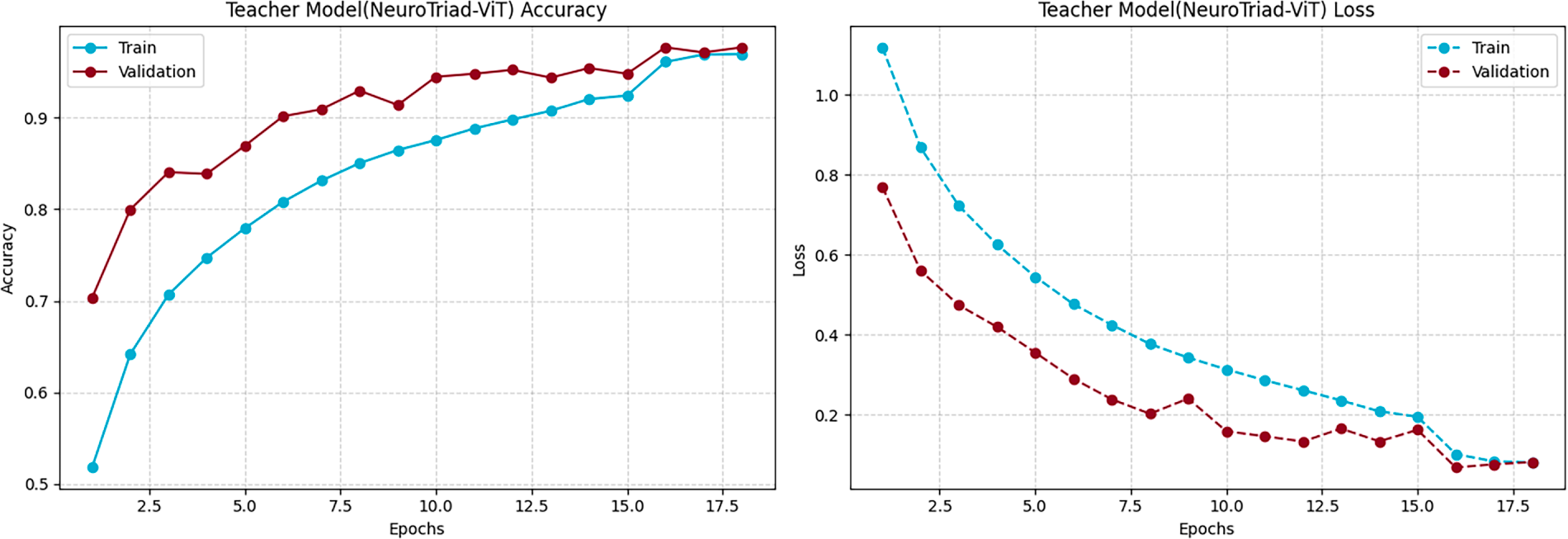
Figure 4: Training performances of the NeuroTriad-ViT model. Visualization of the training and validation accuracy and loss curves, demonstrating the model’s convergence behavior and learning stability throughout the training epochs.
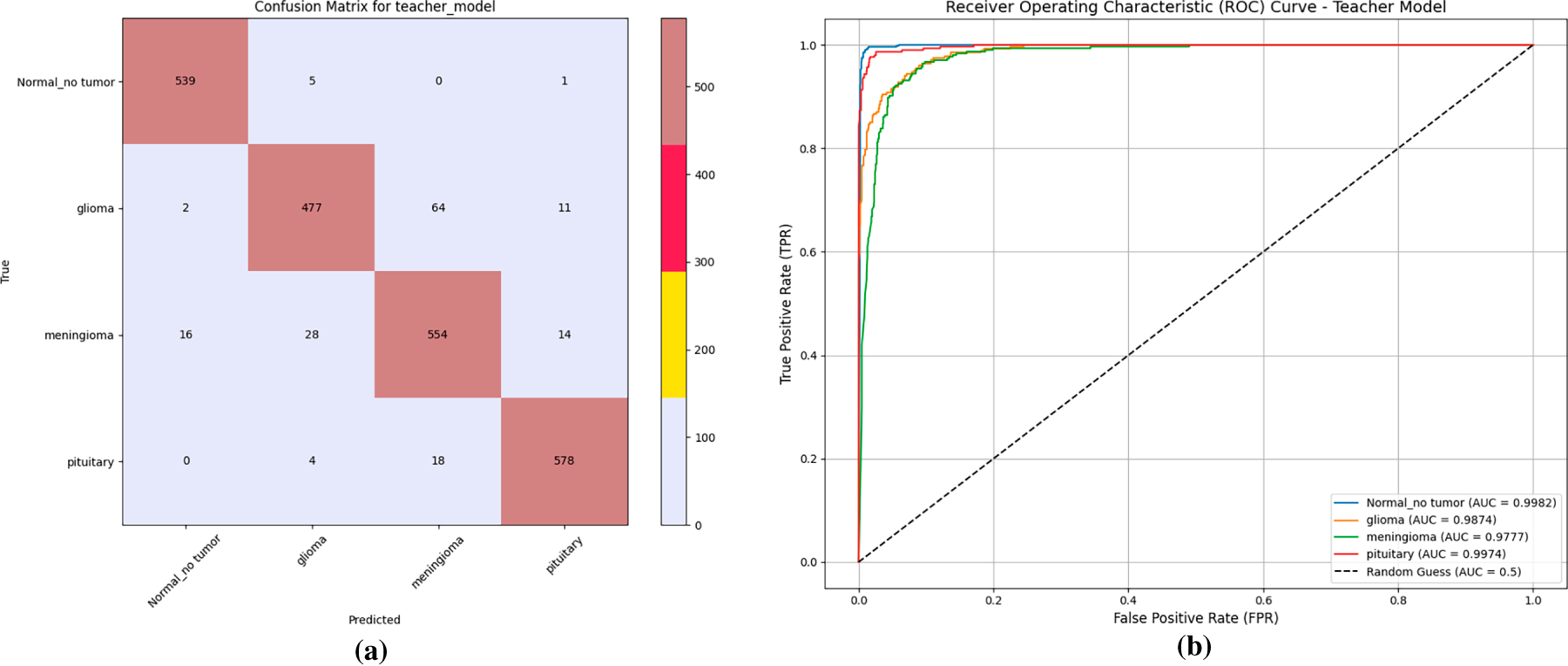
Figure 5: Performance evaluation of the teacher model. (a) Confusion matrix illustrating classification accuracy for each tumor category. (b) Receiver operating characteristic (ROC) curves showing class-wise area under the curve (AUC) performance of the model.
EffiViT-B/16 and the Pretrained ViT-B/16 also exhibit high expressiveness, each surpassing 95% accuracy and generalizing well. For EffiViT-B/16, the hybridization of convolutional inductive biases with ViT attention yields macro-F1 = 0.95 and AUC = 0.98, indicating effective synergy across the data manifold. Traditional hybrid baselines (Effi-VGGNet and VGG-DenseNet) are competitive but show limitations: weighted F1 scores of 0.94 and 0.93 alongside higher validation loss (0.16) suggest greater susceptibility to overfitting and weaker latent representations relative to transformer-augmented architectures. Effi-DenseNet performs lowest in most categories (validation accuracy = 92%; precision/recall/F1 = 0.93), indicating a performance plateau when stacking conventional CNN features without attention mechanisms. An overall performance comparison between all Teacher-nominated models, shown in a bar chart visualization of Fig. 6, provides clarification of performance patterns for each distinct candidate.
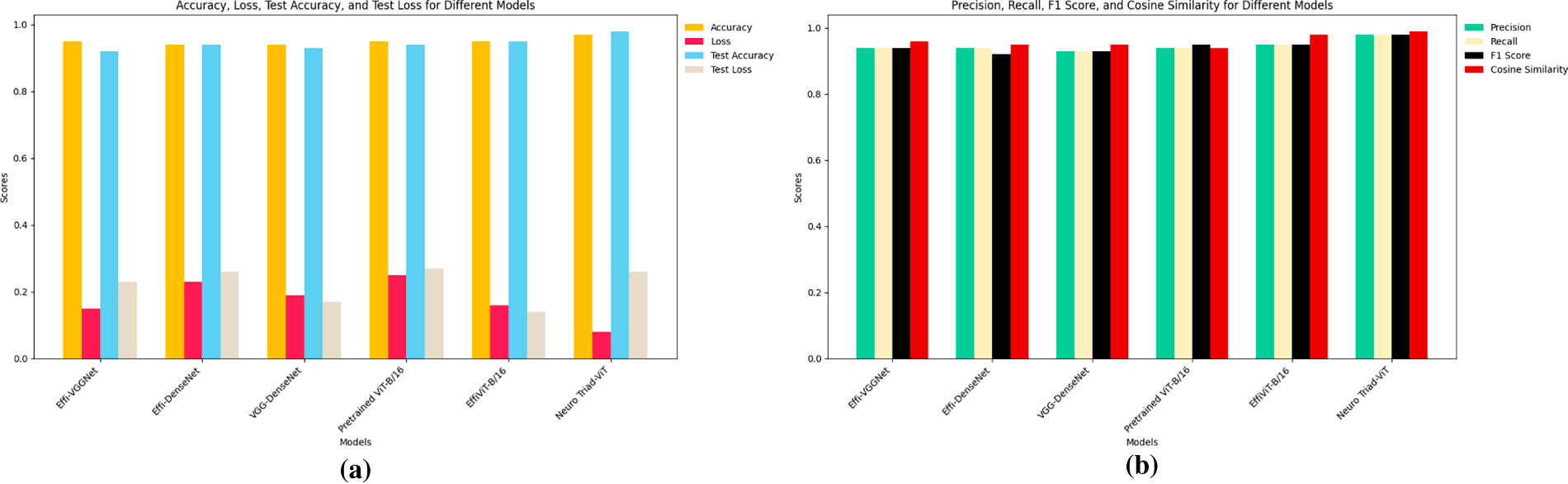
Figure 6: Performance evaluation of teacher-nominated models. (a) Accuracy, loss, test accuracy, and test loss comparison across different models. (b) Precision, recall, F1 score, and cosine similarity comparison across different models.
We go well beyond the standard benchmarking paradigms by developing a rigorous evaluation methodology, thereby certifying the most transferable and best-performing architecture, NeuroTriad-ViT, as the canonical teacher model in a knowledge-distillation framework. NeuroTriad-ViT achieves superior classification performance and effectively transfers knowledge to smaller student networks, establishing a benchmark for reproducibility, fine-tuning capability, and clinical applicability. Transformer-enriched architectures exhibit improved performance and richer feature representations, while their attention mechanisms ensure consistent generalization across validation and test sets, demonstrating robustness for clinical deployment. NeuroTriad-ViT shows high concordance with conventional diagnostic ratings, enhancing neuro-oncological care. As a teacher model, it provides a scalable foundation for student models to assimilate both feature-level and logit-level representations, enabling state-of-the-art performance in efficient and reliable brain tumor classification.
Overall, NeuroTriad-ViT opens new avenues in medical imaging AI and may serve as a precedent for future clinical AI systems, with subsequent research aimed at improving interpretability, computational efficiency, and real-world deployment.
4.3 Knowledge Distillation: Enhancing Performance and Efficiency of Brain Tumor Classification Models
Table 5 presents a comprehensive quantitative comparison between the NeuroTriad-ViT teacher model and its knowledge-distilled lightweight student counterparts, MobileNetV2 and EfficientNet-Lite0. The teacher achieves strong test accuracy (98.23%), a macro F1-score of 0.98, and an AUC of 0.99, demonstrating reliable decision boundaries, further supported by a cosine similarity score of 0.99. Following distillation, both student models improve substantially; however, MobileNetV2 consistently aligns most closely with the teacher across all metrics, attaining a test accuracy of 99.3% and a validation accuracy of 99.32%, with reduced validation and test losses (0.006 and 0.112).
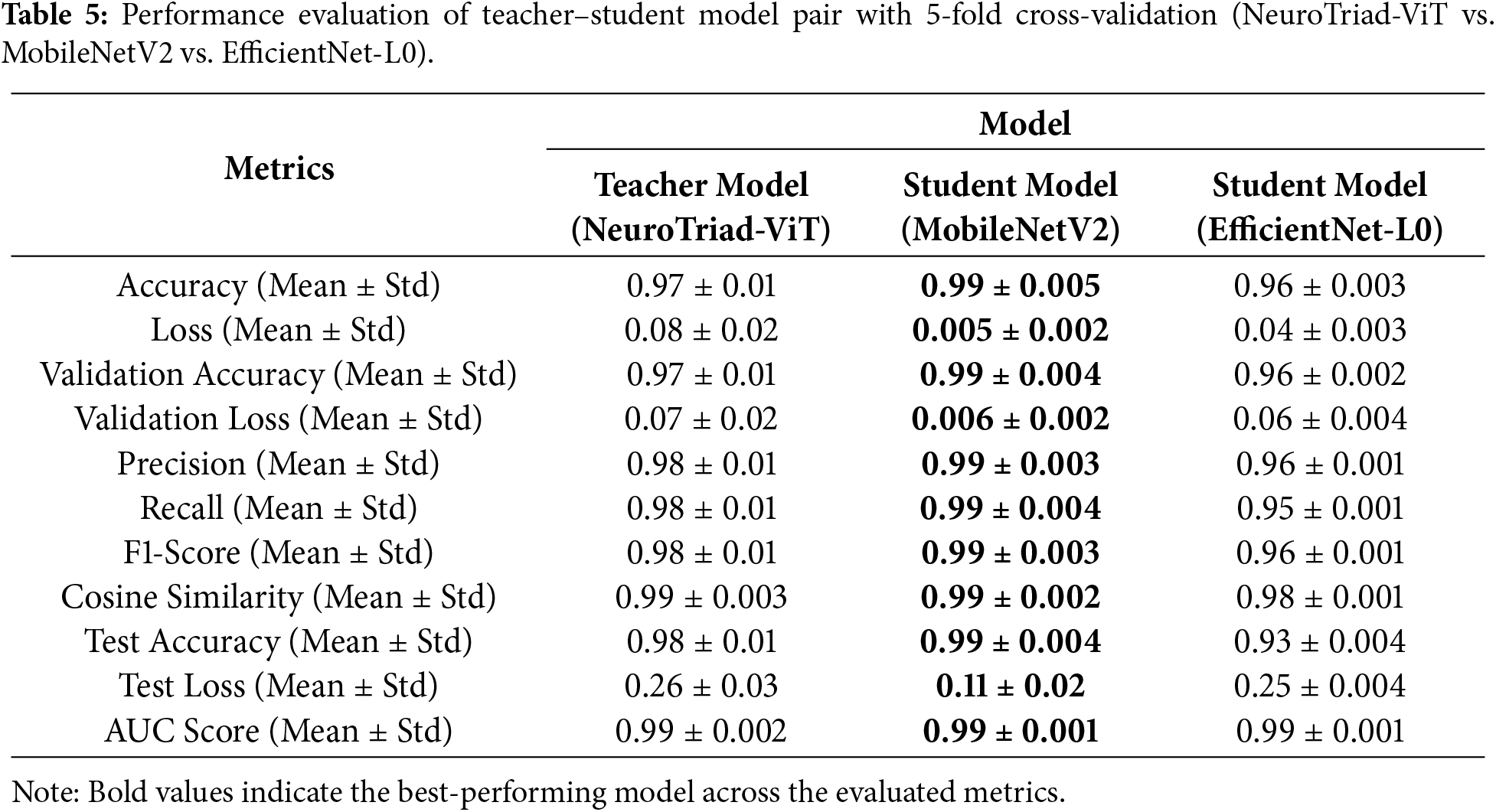
Table 5 provides a comprehensive quantitative analysis comparing the NeuroTriad-ViT teacher model with its knowledge-distilled lightweight student counterparts, MobileNetV2 and EfficientNet-Lite0. The teacher model achieves a test result of 98.23%, achieving a macro F1-score of 0.98 and an area under the receiver operating characteristic curve (AUC) of 0.99, demonstrating strong decision boundaries, further supported by a high cosine similarity of 0.99. Knowledge distillation significantly improves the performance of both student models. Among them, MobileNetV2 consistently aligns most closely with the teacher across all evaluation metrics, attaining a test accuracy of 99.12%, a validation accuracy of 99.32%, and validation and test losses of 0.006 and 0.112, respectively.
Although EfficientNet-Lite0 also benefits from distillation, it yields lower accuracy (96%), higher loss values (0.04), and reduced cosine similarity (0.98), indicating comparatively weaker preservation of the teacher’s decision geometry than MobileNetV2, as illustrated in Fig. 7. In contrast, precision, recall, and F1-score values all exceed 0.99, while the confusion matrices (Fig. 8) show fewer misclassifications across tumor classes, highlighting MobileNetV2’s superior internalization of the teacher’s inter-class relationships.
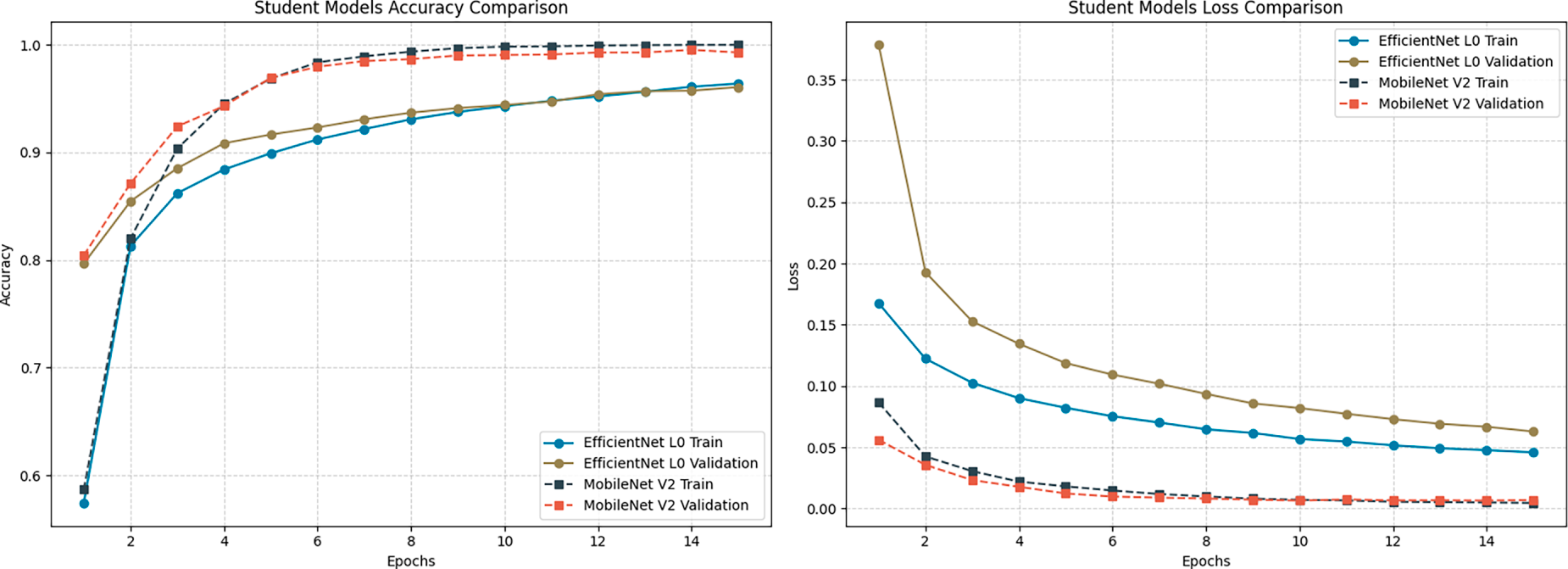
Figure 7: Accuracy and loss comparison of distilled Student models. Illustration of the training and validation accuracy and loss trajectories, showcasing the student model’s convergence characteristics and learning consistency over the training epochs.
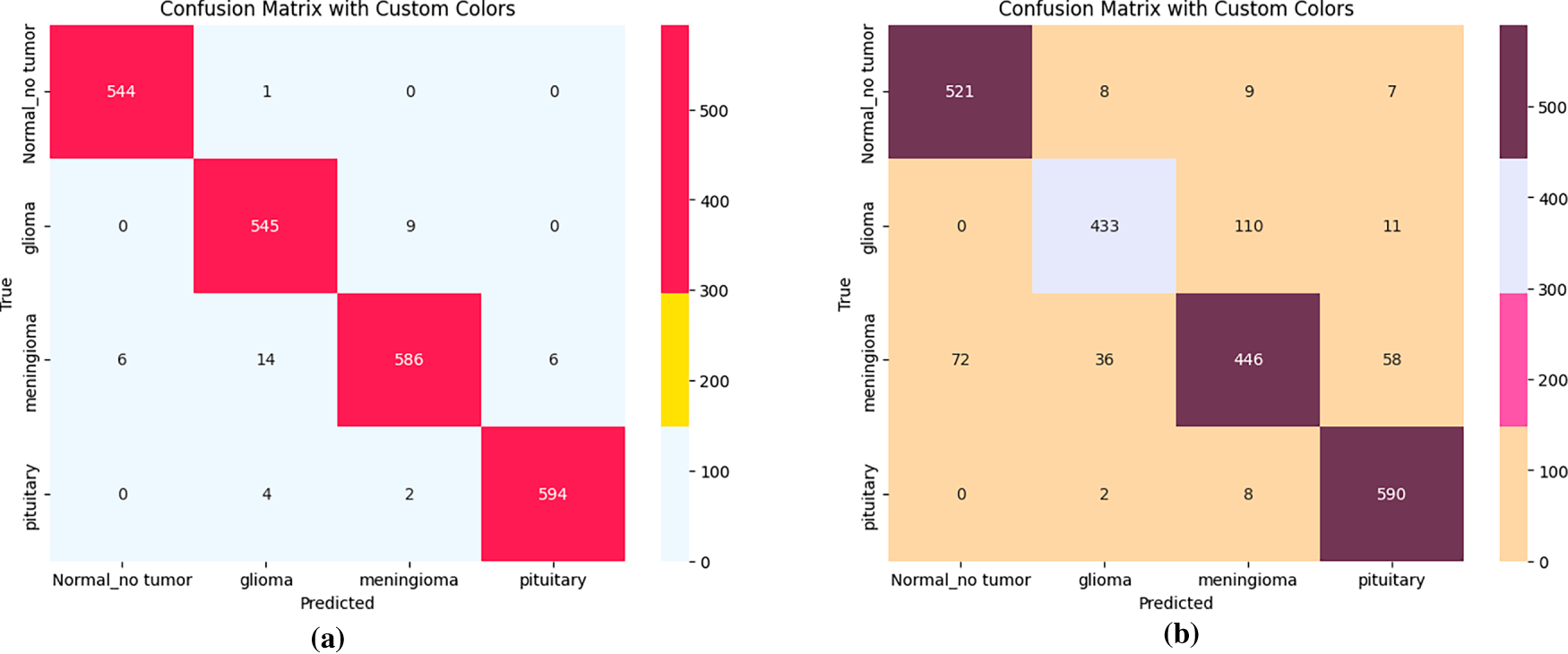
Figure 8: Confusion matrices of student models after knowledge distillation. (a) Confusion matrix for MobileNetV2 student model. (b) Confusion matrix for EfficientNet-L0 student model.
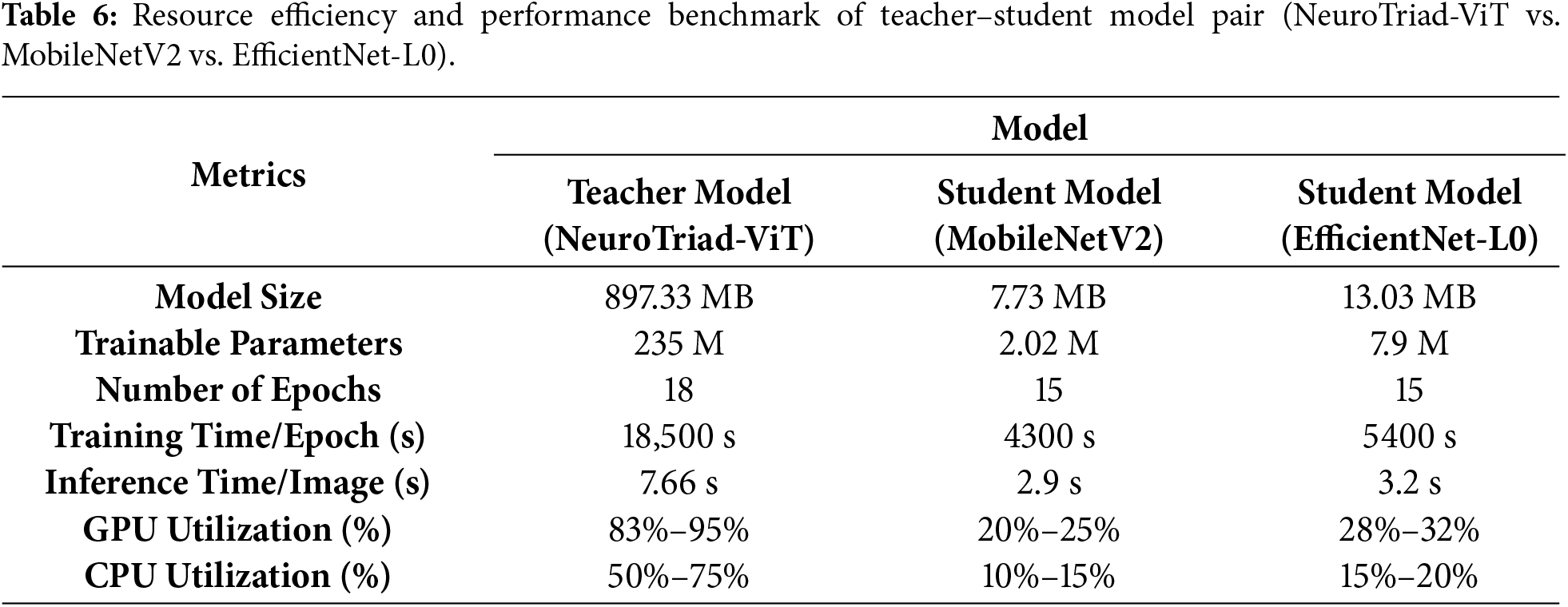
Beyond predictive performance, MobileNetV2 not only delivers higher accuracy and better alignment with the teacher’s decision boundaries but also demonstrates superior computational efficiency (Table 6). The MobileNetV2 the student model is substantially smaller than the teacher model, with a size of only 7.73 MB compared to 897.33 MB for NeuroTriad-ViT, and the number of trainable parameters is reduced by 235 million to just 2.02 million. In contrast, EfficientNet-Lite0, while still smaller than the teacher, occupies 13.02 MB with 7.9 million parameters, nearly four times more than MobileNetV2. This reduction in model dimensionality translates to faster training and inference and substantially lower hardware usage, with GPU and CPU consumption limited to 20%–25% and 10%–15%, respectively, of that required by MobileNetV2.
As the student model achieves extreme performance across evaluation metrics, the study further examines prediction confidence using reliability diagrams and Expected Calibration Error (ECE) to verify whether high accuracy corresponds to trustworthy probabilities. The teacher model is initially overconfident but is corrected via temperature scaling, reducing its ECE to 0.0696 without retraining. MobileNetV2 achieves an ECE of 0.0114, indicating near-perfect alignment between confidence and accuracy, as visually confirmed in the reliability diagram (Appendix A).
Overall, MobileNetV2 student model achieves the most effective knowledge transfer, retaining and in several metrics refining the teacher’s discriminative confidence while minimizing computational cost. Its superior performance across accuracy, loss, cosine similarity, and calibration metrics, combined with minimal parameters and resource demand, allows it to surpass EfficientNet-Lite0 and emerge as the optimal student model in the proposed framework, establishing it as the most clinically viable choice for deployment.
4.4 Model Interpretability with Grad-CAM, LIME, and Faithfulness Evaluation
The interpretability of the student model was analyzed using Grad-CAM and LIME visualizations, complemented by faithfulness scores (Insertion and Deletion metrics) computed over 10 representative images per class. These analyses were conducted after achieving state-of-the-art accuracy, thereby confirming that high predictive performance aligns with stable interpretability. The confidence scores of the sampled images remained consistently high (average 0.99), indicating strong predictive certainty of the model. Grad-CAM visualizations (Fig. 9) clearly highlight the spatial regions with the greatest impact on predictions across all four classes, including normal, glioma, pituitary, and meningioma, accurately localizing tumor regions and supporting the model’s structural focus. LIME results (Fig. 10b) further delineate superpixels contributing to model decisions, while class-specific masks (Fig. 10a) were generated solely for visualization in the faithfulness evaluation, not for training purposes.
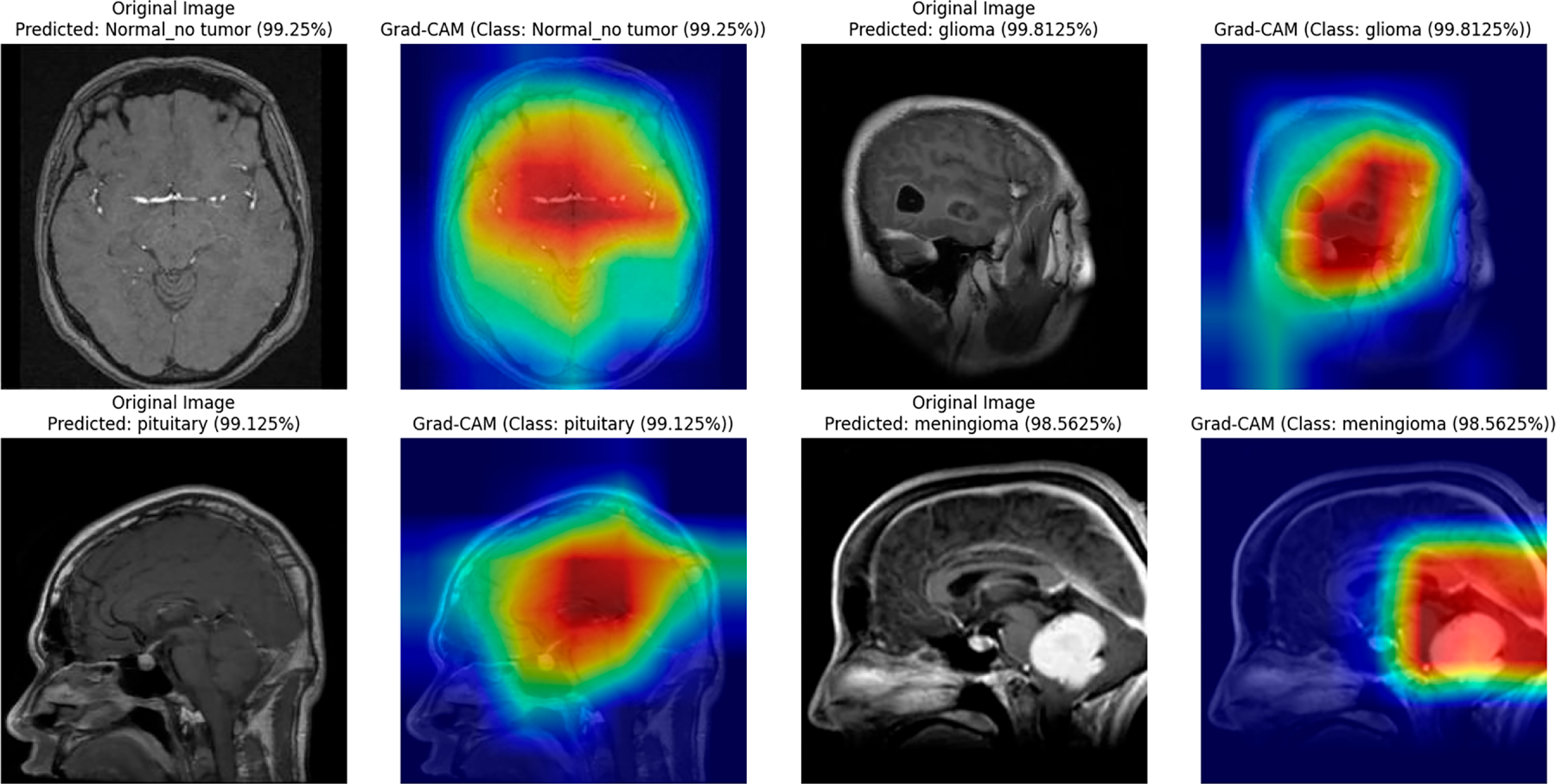
Figure 9: Grad-CAM visualizations for the student model across different brain tumor classes (glioma, pituitary, meningioma) and normal brain images.
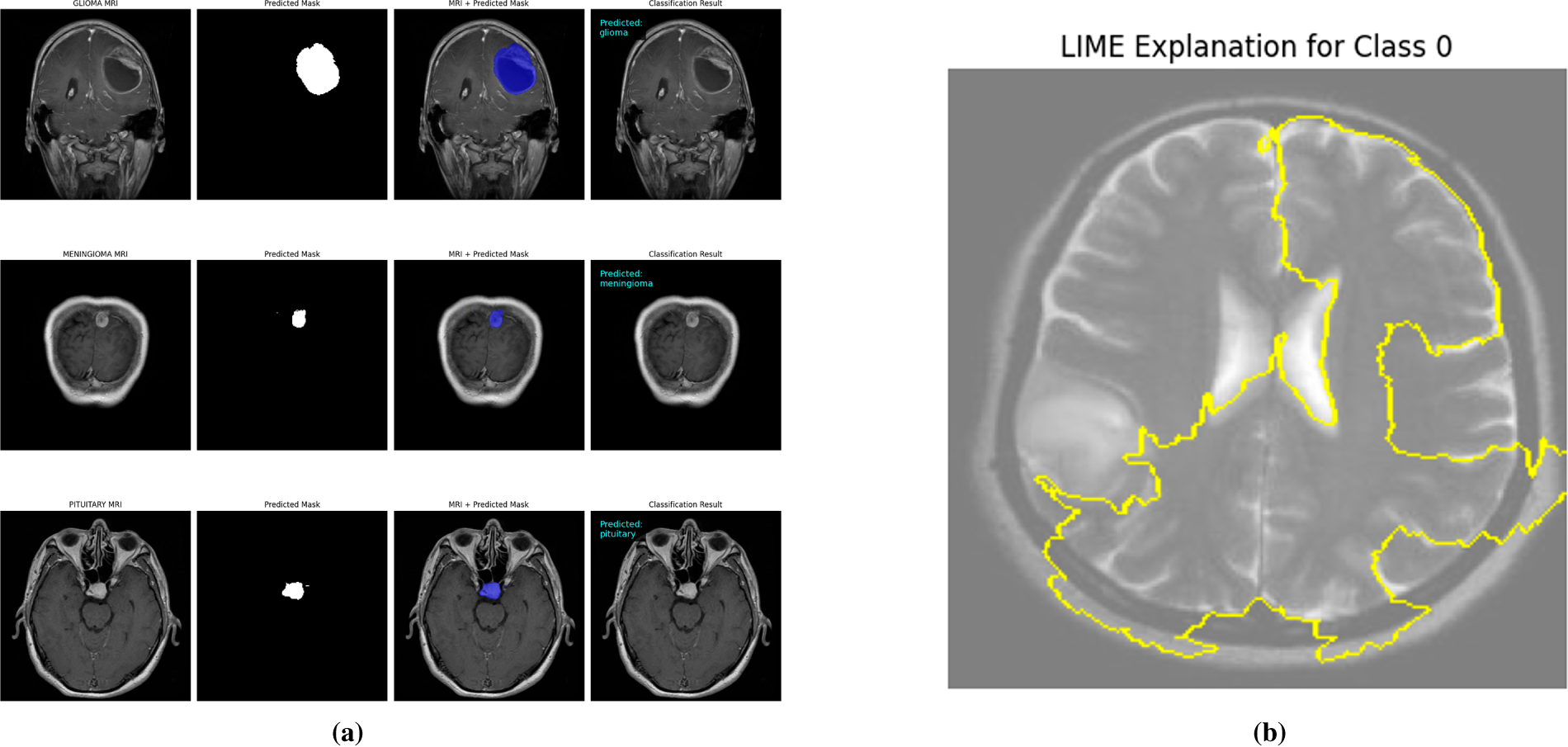
Figure 10: Visual explanations for student models after knowledge distillation. (a) Class-specific masks highlighting regions of interest for the MobileNetV2 student, illustrating model attention for each tumor type. (b) LIME-based explanations showing the predictive regions for the EfficientNet-L0 student, providing interpretable insights into model decisions.
Table 7 presents the Insertion and Deletion scores for both Grad-CAM and LIME. Notably, the normal class shows a low deletion score of 0.09 for Grad-CAM because removing non-informative regions in a normal brain has minimal effect on prediction, reflecting the absence of critical features to disrupt the model’s confident classification. Across all classes, the average insertion and deletion scores confirm that the highlighted regions consistently correspond to the model’s confident predictions (Insertion: 0.69 Grad-CAM, 0.73 LIME; Deletion: 0.45 Grad-CAM, 0.59 LIME).
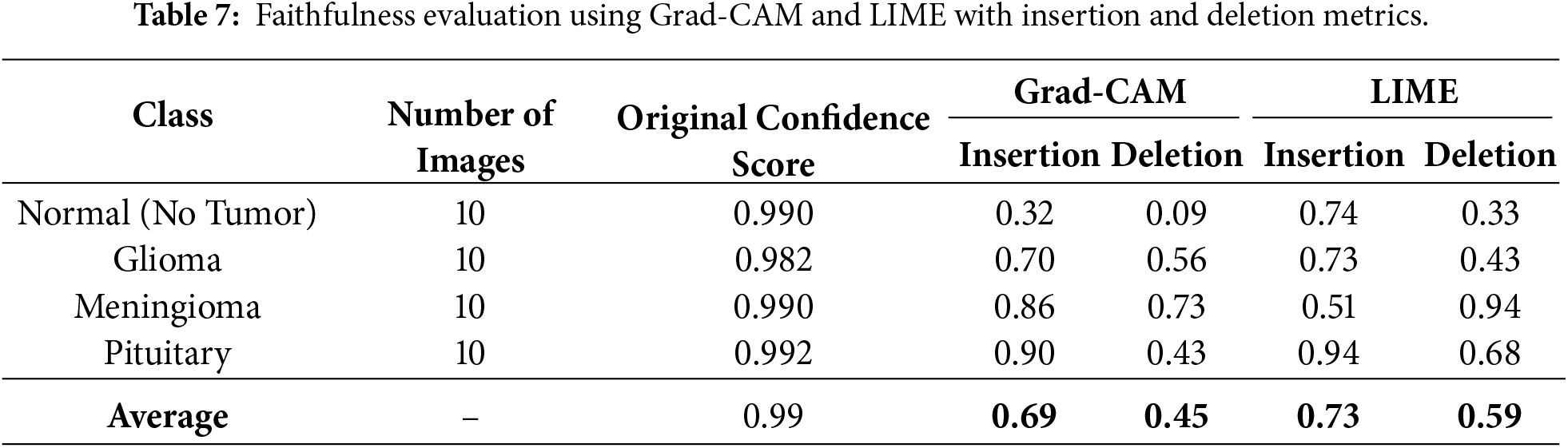
In summary, combining Grad-CAM, LIME, and mask-based faithfulness metrics provides a comprehensive interpretability framework, ensuring transparency, high diagnostic accuracy, and confidence in both teacher and student models for neurodiagnostic applications.
4.5 Comparison with Existing Studies
In order to demonstrate the importance of the proposed Neuro Triad-ViT framework and the dual-supervision knowledge distillation strategy, this section reviews its performance in comparison with previously reported performances of deep learning models for brain tumor classification is presented in Table 8.
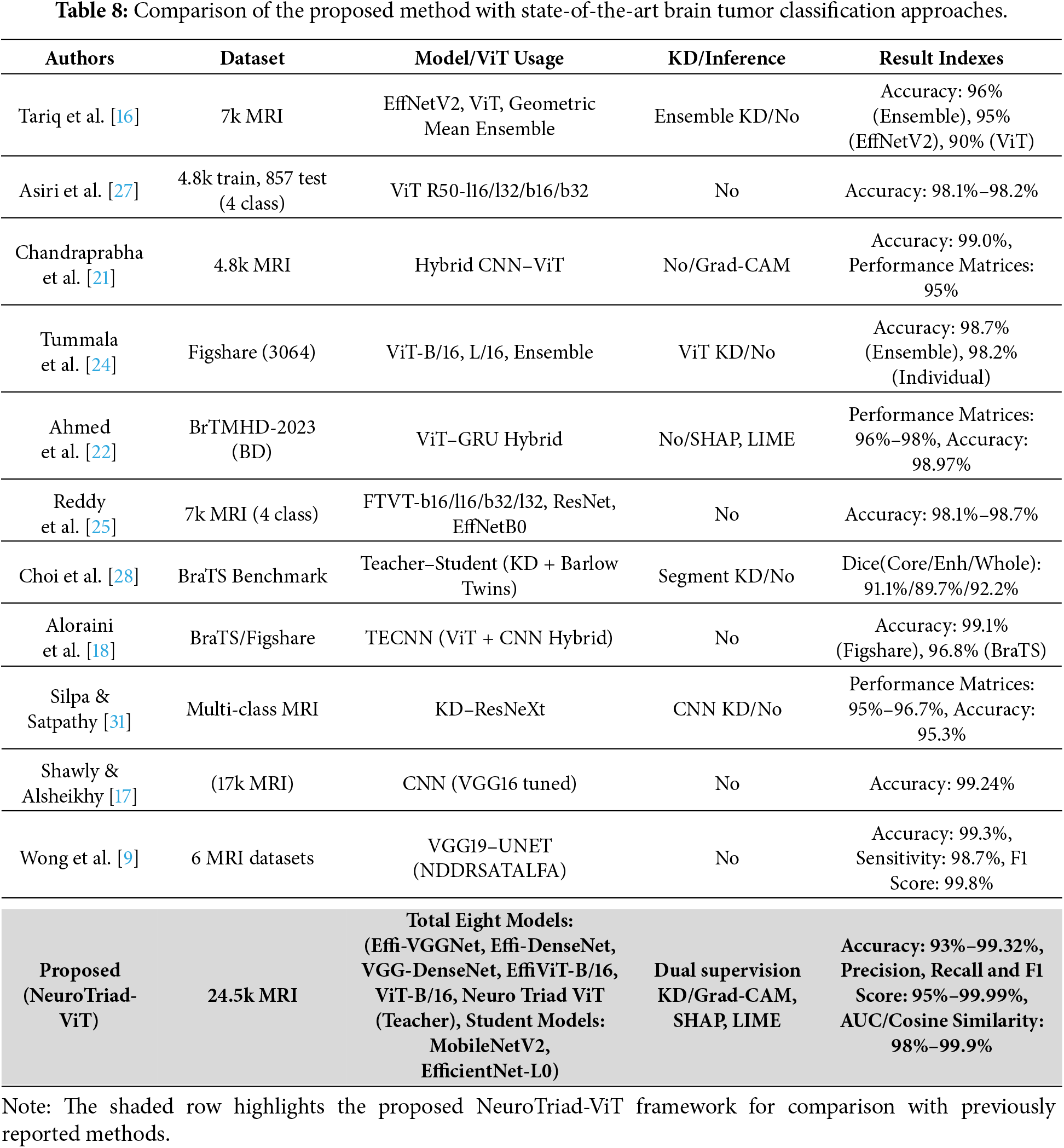
This comparative overview thus allows the assessment of the advancement included in this project from a systemic perspective by correlating the findings of prior studies with the most recent and effective strategies, such as CNN-based systems and ensemble models, which have been developed to construct transformer-level architectures. This, in turn, demonstrates the uniqueness of the proposed methodology. This research showed that, in conjunction with the use of the knowledge distillation method, Neuro Triad-ViT laid new state-of-the-art (SOTA) records in the current competitive landscape for medical artificial intelligence research.
In a broader context, the relative comparison of advanced deep learning approaches to brain tumor classification made in the current research has the potential to open up significant innovative opportunities for AI-driven medical diagnostics. Several deep learning models were explored, including recent convolutional networks, their hybrids with ViT, and transformer-based approaches. All of these models demonstrated benefits in terms of performance, interpretability, and computational efficiency. This study confirms that the NeuroTriad-viT model has better accuracy, robustness, and adaptability compared with conventional CNN-based and hybrid methods. Furthermore, the analysis implies that the integration of convolutional and vision transformer learning is very beneficial for medical image classification tasks.
Additionally, the study provided a framework for the Robust Brain Tumor detection and Classification (RBTC) experiment using an effective knowledge distillation scheme, which was designed to ensure knowledge transfer from a complex teacher model to a lightweight student model. This translation led to a significant reduction in computational needs with maintenance and improvement in some cases of accuracy and precision over a dataset of 24,555 brain tumor samples. In addition, decision transparency was ensured by color-coded visualization (Grad-CAM, LIME, and Mask-based Faithfulness).
Writing the last line, this study makes a bridge of Artificial Intelligence with Medical statement, combining the proposed NeuroTriad-ViT model with a knowledge-distillation-based RBTC framework provide a feasible pathway toward clinical implementation. This innovation may help bridge the gap between clinical practitioners and software developers, thereby accelerating the incorporation of artificial intelligence into radiological diagnostics. The proposed synergy aligns with the objectives of improving diagnostic accuracy, streamlining operational efficiency, and ultimately contributing to superior patient outcomes.
Despite the promising performance of the proposed architecture and its demonstrated robustness on a large-scale MRI dataset of 24,455 scans, the analysis is limited to a single curated dataset. Further validation on fully independent external datasets is required to assess generalizability across different acquisition protocols, scanner models, and clinical settings. Additionally, delays in institutional review board (IRB) approval restricted large-scale evaluation of rare tumor subtypes and highly heterogeneous demographic cohorts, which could introduce data bias. Addressing these limitations will improve the clinical reliability and applicability of the framework.
Further studies will be conducted using large-scale external validation across multiple institutions and geographic locations to assess robustness and generalization under heterogeneous imaging protocols and clinical settings. Future longitudinal research will evaluate real-world clinical performance and temporal stability. Additionally, efforts will be made to streamline clinical deployment pipelines, including integration into radiology workflows, scalability across hospital systems, and adherence to regulatory and ethical standards, thereby facilitating practical adoption in routine clinical practice.
In conclusion, the artificial intelligence aspect of medical diagnostics has been discussed. In particular, the paper proposed the model of NeuroTriad-ViT, which is used to classify brain tumors in an accurate and interpretable manner. The combination of convolutional neural networks with transformer-based architectures in the model brought it to the state-of-the-art accuracy, robustness, and clinical relevance. The utilization of knowledge distillation facilitated its deployment in the form of a lightweight student model (MobileNetV2) without considerable degradation in performance. By leveraging the interpretation modules of Grad-CAM and LIME, the current study paves the way for trustworthy, interpretable, and real-time AI-based diagnostic platforms in the health sector.
Acknowledgement: The authors express their sincere appreciation for the administrative assistance and technical resources provided by the supporting institutions.
Funding Statement: The research was funded by Taif University, Taif, Saudi Arabia (TU-DSPP-2024-210) and the National Natural Science Foundation of China (U23A20321, 62272487, 62272490).
Author Contributions: Conceptualization: Sultan Kahla, Zuping Zhang, Ahmed Emara. Methodology: Sultan Kahla, Majed Alsafyani, Mohammod Abdullah Bin Hossain. Validation: Sultan Kahla, Ahmed Emara, Majed Alsafyani. Investigation: Sultan Kahla, Zuping Zhang, Mohammod Abdullah Bin Hossain. Model design and implementation: Sultan Kahla, Abdulwahab Osman Sheikhdon. Supervision: Zuping Zhang, Ahmed Emara, Majed Alsafyani. Writing—original draft: Sultan Kahla, Zuping Zhang, Majed Alsafyani, Ahmed Emara, Mohammod Abdullah Bin Hossain, Abdulwahab Osman Sheikhdon. Writing—review & editing: Sultan Kahla, Zuping Zhang, Majed Alsafyani, Ahmed Emara, Mohammod Abdullah Bin Hossain, Abdulwahab Osman Sheikhdon. All authors reviewed and approved the final version of the manuscript.
Availability of Data and Materials: The datasets used to support the findings of this study are available from public data repositories on the classification of brain tumors in MRI. These include the Kaggle (BRISC 2025) dataset [37], Mendeley dataset [35], GTS AI dataset [36], and Figshare dataset [38].
Ethics Approval: This study was conducted in accordance with established ethical standards and guidelines for research involving human participants. The MRI datasets used for this study were collected from publicly available sources with the approval of prior institutional review board (IRB) and de-identified for the privacy of the patient. As no new human subjects were recruited and all data were anonymized, the need for additional ethical approval and informed consent was waived. The research follows the principles of the Declaration of Helsinki for ethical research on humans.
Conflicts of Interest: The authors declare no conflicts of interest.
Appendix A Reliability Diagram of Teacher and Student Models
This figure visualizes the calibration of predicted probabilities for both the teacher and MobileNetV2 student models. It illustrates the alignment between prediction confidence and actual accuracy, confirming that the student’s predictions are well-calibrated and reliable.
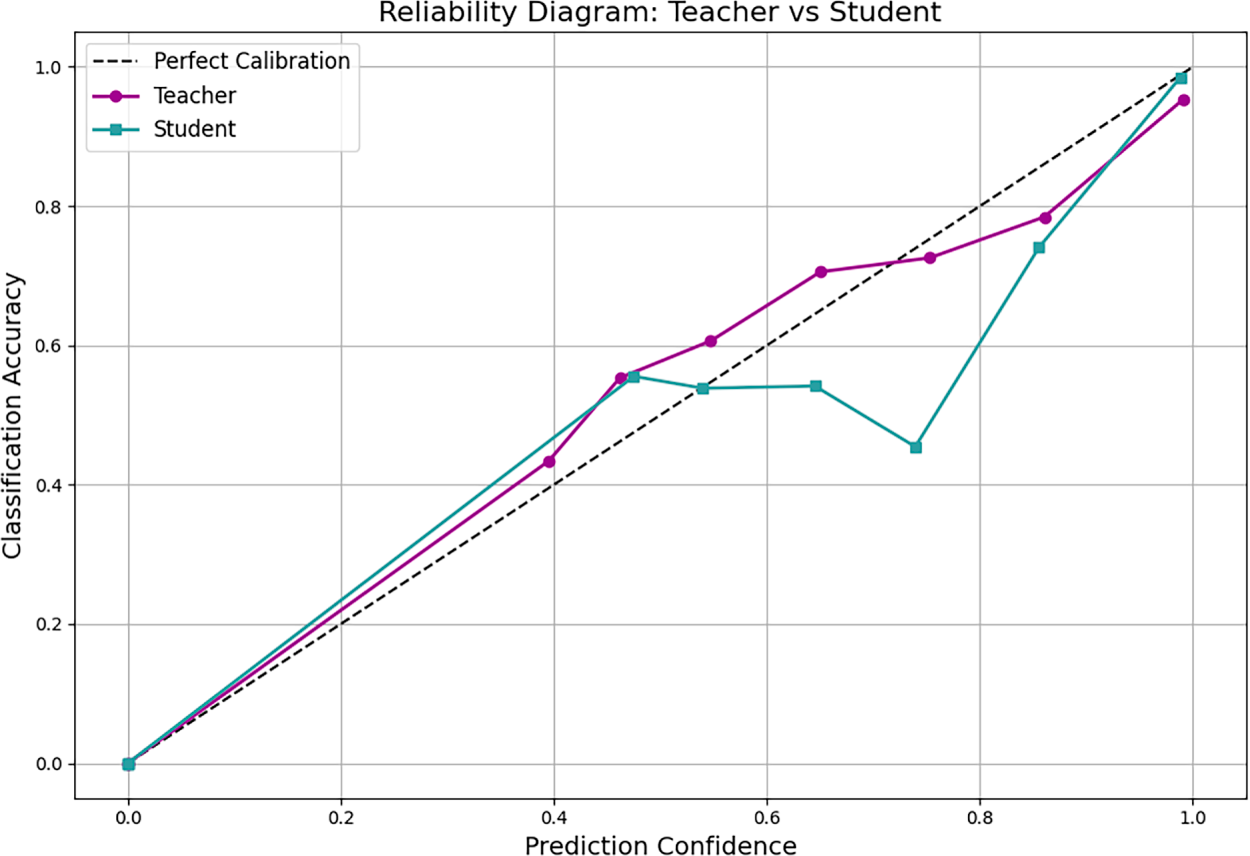
Figure A1: Reliability diagram showing the calibration of predicted probabilities for both the teacher and MobileNetV2 student models. The plot illustrates the alignment between prediction confidence and actual classification accuracy, confirming well-calibrated predictions.
References
1. Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2021;71(3):209–49. doi:10.3322/caac.21660. [Google Scholar] [PubMed] [CrossRef]
2. Tan AC, Ashley DM, López-Gyüera M, Tsoli M, Packer RJ. Brain tumors. Lancet. 2020;395(10235):1537–50. doi:10.1016/S0140-6736(20)30974-X. [Google Scholar] [CrossRef]
3. Louis DN, Perry A, Wesseling P, Brat DJ, Cree IA, Figarella-Branger D, et al. The 2021 WHO classification of tumors of the central nervous system: a summary. Neuro Oncol. 2021;23(8):1231–51. doi:10.1093/neuonc/noab106. [Google Scholar] [PubMed] [CrossRef]
4. Mabray MC, Barajas RF Jr, Villanueva-Meyer JE. Modern brain tumor imaging. Brain Tumor Res Treat. 2021;9(2):61–75. doi:10.14791/btrt.2015.3.1.8. [Google Scholar] [PubMed] [CrossRef]
5. Raza A, Ayub H, Khan JA, Ahmad I, Salama AS, Daradkeh YI, et al. A deep learning-based framework for brain tumor classification using transfer learning. Circuits Syst Signal Process. 2022;41(1):123–46. doi:10.1007/s00034-019-01246-3. [Google Scholar] [CrossRef]
6. Dosovitskiy A, Beyer L, Kolesnikov A, Weissenborn D, Zhai X, Unterthiner T, et al. An image is worth 16×16 words: transformers for image recognition at scale. arXiv:2010.11929. 2021. [Google Scholar]
7. Gou J, Yu B, Maybank SJ, Tao D. Knowledge distillation: a survey. Int J Comput Vis. 2021;129(6):1789–819. doi:10.1007/s11263-021-01453-Z. [Google Scholar] [CrossRef]
8. Selvaraju RR, Cogswell M, Das A, Vedantam R, Parikh D, Batra D. Grad-CAM: visual explanations from deep networks via gradient-based localization. In: Proceedings of the 2017 IEEE International Conference on Computer Vision (ICCV); 2017 October 22–29; Venice, Italy. Piscataway, NJ, USA: IEEE; 2017. p. 618–26. [Google Scholar]
9. Wong Y, Su ELM, Yeong CF, Holderbaum W, Yang C. Brain tumor classification using MRI images and deep learning techniques. PLoS One. 2025;20(5):e0322624. doi:10.1371/journal.pone.0322624. [Google Scholar] [PubMed] [CrossRef]
10. Preetha R, Priyadarsini MJP, Nisha JS. Hybrid 3B Net and EfficientNetB2 model for multi-class brain tumor classification. IEEE Access. 2025;13:63465–85. doi:10.1109/ACCESS.2025.3558411. [Google Scholar] [CrossRef]
11. Velpula V, Vadlamudi J, Janapati M, Kasaraneni P, Kumar Y, Challa PR, et al. Enhanced brain tumor classification using convolutional neural networks and ensemble voting classifier for improved diagnostic accuracy. Comput Electr Eng. 2025;123:110124. doi:10.1016/j.compeleceng.2025.110124. [Google Scholar] [CrossRef]
12. Zhu Z, Khan MA, Wang S, Zhang Y. RBEBT: a ResNet-based BA-ELM for brain tumor classification. Comput Mater Contin. 2023;74(1):101–11. doi:10.32604/cmc.2023.030790. [Google Scholar] [CrossRef]
13. Abbas R, Naijie G, Aldrees A, Umer M, Hakeem A, Alsubai S, et al. Advancing brain tumor segmentation and grading through integration of FusionNet and IBCO-based ALCResNet. Image Vis Comput. 2025;154(3):105432. doi:10.1016/j.imavis.2025.105432. [Google Scholar] [CrossRef]
14. Ramakrishna MT, Pothanaicker K, Selvaraj P, Khan SB, Venkatesan VK, Alzahrani S, et al. Leveraging EfficientNetB3 in a deep learning framework for high-accuracy MRI tumor classification. Comput Mater Contin. 2024;81(1):867–83. doi:10.32604/cmc.2024.053563. [Google Scholar] [CrossRef]
15. Nazir A, Raza D. Advancements in deep learning and explainable artificial intelligence for enhanced medical image analysis: a comprehensive survey and future directions. Eng Appl Artif Intell. 2025;158:111413. doi:10.1016/j.engappai.2025.111413. [Google Scholar] [CrossRef]
16. Tariq A, Iqbal M, Iqbal M, Ahmad A. Transforming brain tumor detection empowering multi-class classification with vision transformers and EfficientNetV2. IEEE Access. 2025;13:63857–76. doi:10.1109/access.2025.3555638. [Google Scholar] [CrossRef]
17. Shawly T, Alsheikhy AA. A novel dynamic residual self-attention transfer adaptive learning fusion approach for brain tumor diagnosis. Comput Mater Contin. 2025;82(3):4161–79. doi:10.32604/cmc.2025.061497. [Google Scholar] [CrossRef]
18. Aloraini M, Khan A, Aladhadh S, Habib S, Alsharekh M, Islam M. Combining the transformer and convolution for effective brain tumor classification using MRI images. Appl Sci. 2023;13(6):3680. doi:10.3390/app13063680. [Google Scholar] [CrossRef]
19. Naim Islam Md, Shafiul Azam Md, Samiul Islam Md, Hasan Kanchan M, Shahariar Parvez AHM, Monirul Islam Md. An improved deep learning-based hybrid model with ensemble techniques for brain tumor detection from MRI image. Inform Med Unlocked. 2024;47(1):101483. doi:10.1016/j.imu.2024.101483. [Google Scholar] [CrossRef]
20. Haque R, Khan M, Rahman H, Khan S, Siddiqui M, Limon Z, et al. Explainable deep stacking ensemble model for accurate and transparent brain tumor diagnosis. Comput Biol Med. 2025;191(6):110166. doi:10.1016/j.compbiomed.2025.110166. [Google Scholar] [PubMed] [CrossRef]
21. Chandraprabha K, Ganesan L, Baskaran K, Russo C, Bianconi A. A novel approach for the detection of brain tumor and its classification via end-to-end vision transformer—CNN architecture. Front Oncol. 2025;15:1508451. doi:10.3389/fonc.2025.1508451. [Google Scholar] [PubMed] [CrossRef]
22. Ahmed M, Hossain M, Islam M, Ali M, Nafi A, Ahmed M, et al. Brain tumor detection and classification in MRI using hybrid ViT and GRU model with explainable AI in Southern Bangladesh. Sci Rep. 2024;14(1):22797. doi:10.1038/s41598-024-71893-3. [Google Scholar] [PubMed] [CrossRef]
23. Tanone R, Li L, Saifullah S. ViT-CB: integrating hybrid vision transformer and CatBoost to enhanced brain tumor detection with SHAP. Biomed Signal Process Control. 2025;100:107027. doi:10.1016/j.bspc.2024.107027. [Google Scholar] [CrossRef]
24. Tummala S, Kadry S, Bukhari S, Rauf H. Classification of brain tumor from magnetic resonance imaging using vision transformers ensembling. Curr Oncol. 2022;29(10):7498–511. doi:10.3390/curroncol29100590. [Google Scholar] [PubMed] [CrossRef]
25. Reddy C, Reddy P, Janapati H, Assiri B, Shuaib M, Alam S, et al. A fine-tuned vision transformer based enhanced multi-class brain tumor classification using MRI scan imagery. Front Oncol. 2024;14:1400341. doi:10.3389/fonc.2024.1400341. [Google Scholar] [PubMed] [CrossRef]
26. Islam MA, Mridha M, Safran M, Alfarhood S, Mohsin Kabir M. Revolutionizing brain tumor detection using explainable AI in MRI images. NMR Biomed. 2025;38(3):e70001. doi:10.1002/nbm.70001. [Google Scholar] [PubMed] [CrossRef]
27. Asiri AA, Shaf A, Ali T, Pasha MA, Aamir M, Irfan M, et al. Advancing brain tumor classification through fine-tuned vision transformers: a comparative study of pre-trained models. Sensors. 2023;23(18):7913. doi:10.3390/s23187913. [Google Scholar] [PubMed] [CrossRef]
28. Choi Y, Al-Masni M, Jung K, Yoo R, Lee S, Kim D. A single stage knowledge distillation network for brain tumor segmentation on limited MR image modalities. Comput Methods Programs Biomed. 2023;240(5):107644. doi:10.1016/j.cmpb.2023.107644. [Google Scholar] [PubMed] [CrossRef]
29. Tan S, Cai Y, Zhao Y, Hu J, Chen Y. FM-LiteLearn: a lightweight brain tumor classification framework integrating image fusion and multi-teacher distillation strategies. In: Xie X, Styles I, Powathil G, Ceccarelli M, editors. Artificial Intelligence in Healthcare (AIiH 2024). Vol. 14976. Cham, Switzerland: Springer; 2024. p. 95–107. [Google Scholar]
30. Uddin R, Bin Hossain MA, Hossain MN, Chowdhury TK, Siddique S. A diagnostic pipeline for multi-class classification of chest X-ray images using knowledge distillation and semi-supervised segmentation. In: 2025 IEEE Region 10 Symposium (TENSYMP); 2025 Jul 7–9; Christchurch, New Zealand. Piscataway, NJ, USA: IEEE; 2025. p. 1–8. [Google Scholar]
31. Silpa P, Satpathy S. Advancing brain tumour detection and classification: knowledge distilled ResNeXt model for multi-class MRI analysis. Int J Comput Exp Sci Eng. 2024;10(4):1610–23. doi:10.22399/ijcesen.730. [Google Scholar] [CrossRef]
32. Gohari R, Aliahmadipour L, Valipour E. FedBrain-Distill: communication-efficient federated brain tumor classification using ensemble knowledge distillation on non-IID data. In: 2024 14th International Conference on Computer and Knowledge Engineering (ICCKE); 2024 Oct 29–30; Mashhad, Iran. Piscataway, NJ, USA: IEEE; 2024. p. 49–54. [Google Scholar]
33. Khan S, Asim M, Vollmer S, Dengel A. Robust & precise knowledge distillation-based novel context-aware predictor for disease detection in brain and gastrointestinal. arXiv:2505.06381. 2025. [Google Scholar]
34. Liu J, Jiao G, Wu Y. Siamese self-distillation for brain tumor segmentation with missing modalities. SSRN Electron J. 2023. doi:10.2139/ssrn.4333683. [Google Scholar] [CrossRef]
35. Rahman MM. Brain cancer—MRI dataset. Mendeley Data. 2024. doi:10.17632/mk56jw9rns.1. [Google Scholar] [CrossRef]
36. GTS AI. Brain tumor detection with advanced MRI dataset. GTS AI. 2025 [cited 2026 Jan 12]. Available from: https://gts.ai/dataset-download/brain-tumor-dataset-mri-scans/. [Google Scholar]
37. Kaggle. BRISC 2025: brain MRI dataset for tumor classification. Database: Kaggle. 2025 [cited 2026 Jan 12]. Available from: https://www.kaggle.com/datasets/briscdataset/brisc2025. [Google Scholar]
38. Figshare. Brain tumor dataset. Figshare. 2025. doi:10.6084/m9.figshare.28597496. [Google Scholar] [CrossRef]
39. Buslaev A, Parinov A, Khvedchenya E, Iglovikov V, Kalinin A. Albumentations: fast and flexible image augmentations. arXiv:1809.06839. 2018. [Google Scholar]
40. Simonyan K, Zisserman A. Very deep convolutional networks for large-scale image recognition. arXiv:1409.1556. 2014. [Google Scholar]
41. Tan M, Le QV. EfficientNet: rethinking model scaling for convolutional neural networks. arXiv:1905.11946. 2020. [Google Scholar]
42. Huang G, Liu Z, van der Maaten L, Weinberger KQ. Densely connected convolutional networks. arXiv:1608.06993. 2018. [Google Scholar]
43. Vaswani A, Shazeer N, Parmar N, Uszkoreit J, Jones L, Gomez A, et al. Attention is all you need. arXiv:1706.03762. 2017. [Google Scholar]
44. Ribeiro MT, Singh S, Guestrin C. “Why should I trust you?”: explaining the predictions of any classifier. In: Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining. New York, NY, USA: ACM; 2016. p. 1135–44. [Google Scholar]
Cite This Article
 Copyright © 2026 The Author(s). Published by Tech Science Press.
Copyright © 2026 The Author(s). Published by Tech Science Press.This work is licensed under a Creative Commons Attribution 4.0 International License , which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.


 Submit a Paper
Submit a Paper Propose a Special lssue
Propose a Special lssue View Full Text
View Full Text Download PDF
Download PDF Downloads
Downloads
 Citation Tools
Citation Tools