 Open Access
Open Access
ARTICLE
Metochalcone induces senescence-associated secretory phenotype via JAK2/STAT3 pathway in breast cancer
1 Department of Pharmacology, West China School of Pharmacy, Sichuan University, Chengdu, China
2 State Key Laboratory of Southwestern Chinese Medicine Resources, Chengdu University of Traditional Chinese Medicine, Chengdu, China
3 Key Laboratory of Drug-Targeting and Drug Delivery System of the Education Ministry, Sichuan Engineering Laboratory for Plant-Sourced Drug and Sichuan Research Center for Drug Precision Industrial Technology, Sichuan University, Chengdu, China
4 Chengdu No. 1 Pharmaceutical Co., Ltd., Pengzhou, China
5 Chengdu Push Bio-Technology Co., Ltd., Chengdu, China
* Corresponding Authors: CHENG PENG. Email: ; FU PENG. Email:
(This article belongs to the Special Issue: Application of Multi-omics Analysis in Cancer Immunotherapy)
Oncology Research 2024, 32(5), 943-953. https://doi.org/10.32604/or.2023.044775
Received 08 August 2023; Accepted 24 November 2023; Issue published 23 April 2024
Abstract
Breast and lung cancers are the leading causes of mortality and most frequently diagnosed cancers in women and men, respectively, worldwide. Although the antitumor activity of chalcones has been extensively studied, the molecular mechanisms of isoliquiritigenin analog 2', 4', 4-trihydroxychalcone (metochalcone; TEC) against carcinomas remain less well understood. In this study, we found that TEC inhibited cell proliferation of breast cancer BT549 cells and lung cancer A549 cells in a concentration-dependent manner. TEC induced cell cycle arrest in the S-phase, cell migration inhibition in vitro, and reduced tumor growth in vivo. Moreover, transcriptomic analysis revealed that TEC modulated the activity of the JAK2/STAT3 and P53 pathways. TEC triggered the senescence-associated secretory phenotype (SASP) by repressing the JAK2/STAT3 axis. The mechanism of metochalcone against breast cancer depended on the induction of SASP via deactivation of the JAK2/STAT3 pathway, highlighting the potential of chalcone in senescence-inducing therapy against carcinomas.Graphic Abstract
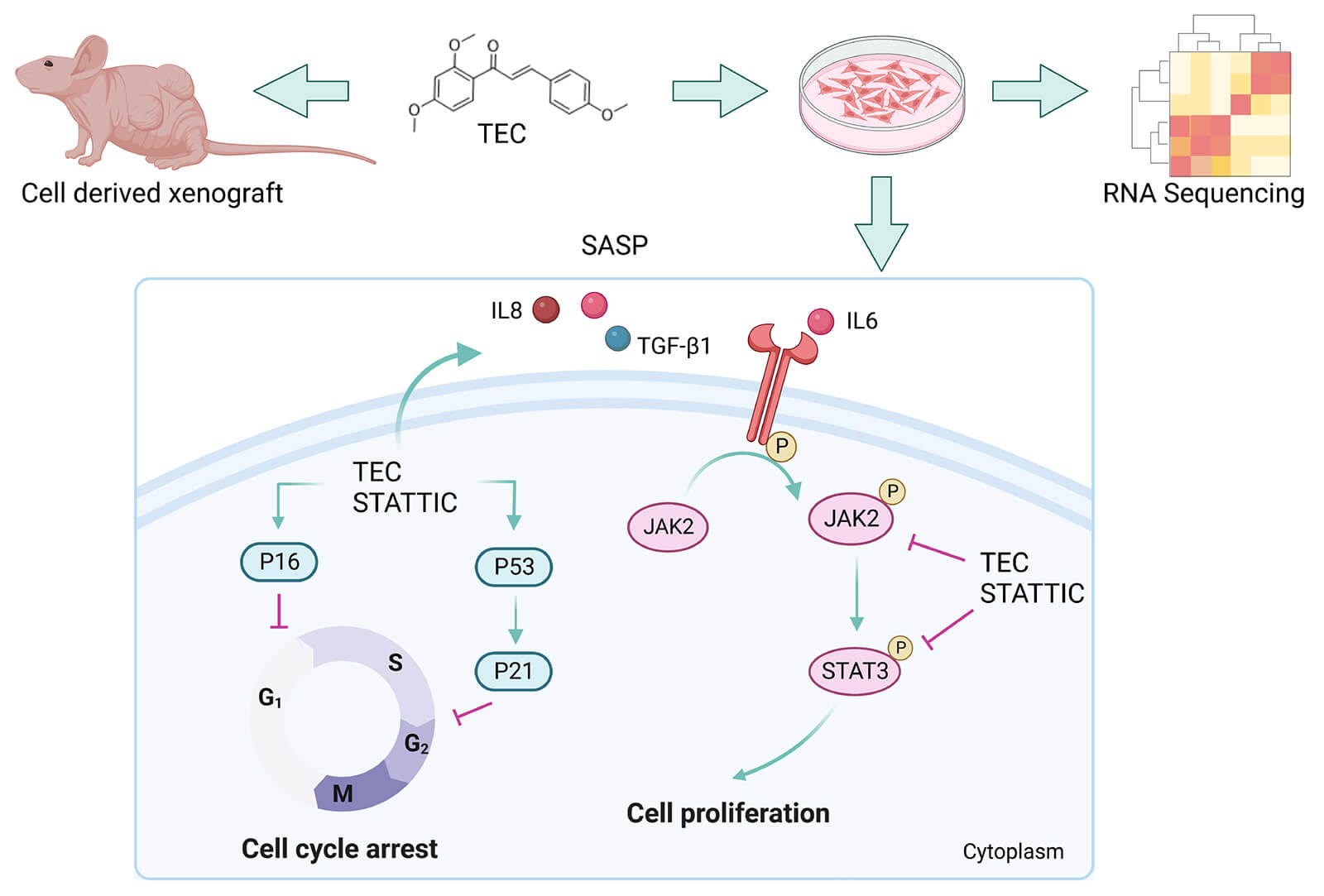
Keywords
Cite This Article
 Copyright © 2024 The Author(s). Published by Tech Science Press.
Copyright © 2024 The Author(s). Published by Tech Science Press.This work is licensed under a Creative Commons Attribution 4.0 International License , which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.


 Submit a Paper
Submit a Paper Propose a Special lssue
Propose a Special lssue View Full Text
View Full Text Download PDF
Download PDF Downloads
Downloads
 Citation Tools
Citation Tools