 Open Access
Open Access
ARTICLE
Multi-Scale Transcriptomic Sequencing Data Analysis Reveals LINC00467 is Associated with Malignant Progression in Breast Cancer: An In Silico and In Vitro Study
1 Department of Urology, Xiangya Hospital, Central South University, Changsha, China
2 NHC Key Laboratory of Human Stem Cell and Reproductive Engineering, Institute of Reproductive and Stem Cell Engineering, Central South University, Changsha, China
3 Department of Plastic Surgery, The Third Xiangya Hospital, Central South University, Changsha, China
4 Department of Anesthesiology, The Second Xiangya Hospital, Central South University, Changsha, China
5 Guangxi Key Laboratory of Reproductive Health and Birth Defect Prevention, Maternal and Child Health Hospital of Guangxi Zhuang Autonomous Region, Nanning, China
6 Genetic and Metabolic Central Laboratory, Birth Defects Prevention and Control Institute of Guangxi Zhuang Autonomous Region, Nanning, China
7 National Institute of Drug Clinical Trial, Xiangya Hospital, Central South University, Changsha, China
8 China National Clinical Research Center for Geriatric Disorders, Xiangya Hospital, Central South University, Changsha, China
* Corresponding Authors: Zailong Qin. Email: ; Jie Guo. Email:
; Junbin Yuan. Email:
# These authors contributed equally to this work
(This article belongs to the Special Issue: Breast Cancer Biomarkers and Drug Targets Discoveries Towards a More Personalized Treatment Setting)
Oncology Research 2026, 34(4), 26 https://doi.org/10.32604/or.2026.067601
Received 07 May 2025; Accepted 26 December 2025; Issue published 23 March 2026
Abstract
Objective: Long non-coding RNAs have been found to play a pivotal role in breast cancer, yet the majority of these lncRNAs remain to be thoroughly investigated. This study aimed to explore the role of differentially expressed long non-coding RNAs (lncRNAs) in breast cancer stemness and drug sensitivity. Methods: Database mining was performed to evaluate the expression of LINC00467 in different types of breast cancer and its association with clinical features. The function of LINC00467 was examined through colony formation assays, quantitative reverse transcription PCR (qRT-PCR), and western blotting following LINC00467 silencing in breast cancer cell lines. Results: LINC00467 was significantly upregulated in various breast cancer subtypes with spatial specificity. Silencing LINC00467 reduced clonogenic capacity and downregulated the stemness-associated factor LIN28B as well as phosphorylated RAC-alpha serine/threonine-protein kinase (p-AKT). The transcription factors specificity protein 1 (SP1) and E2F transcription factor 1 (E2F1) were predicted to bind to the LINC00467 promoter. Furthermore, breast cancer samples with high LINC00467 expression displayed reduced sensitivity to AKT inhibitors, and high LINC00467 expression was negatively correlated with the therapeutic response to programmed cell death 1 (PD-1) antibodies. Conclusion: Our findings suggest that spatially expressed LINC00467 may promote breast cancer stemness by regulating AKT signaling and could serve as a potential new therapeutic target and indicator of drug sensitivity in breast cancer.Keywords
Breast cancer is one of the most common malignant tumors in women, and its incidence continues to rise worldwide [1]. According to global cancer statistics, breast cancer accounts for 25% of all cancer cases and 15% of cancer-related deaths [2]. Current treatment strategies typically involve a combination of hormonal therapy, surgery, chemotherapy, and radiotherapy. However, prognosis remains poor due to multidrug resistance, recurrence, and metastasis. Consequently, identifying specific biomarkers and therapeutic targets is critical for the early diagnosis and effective treatment of breast cancer [3].
Long non-coding RNAs (lncRNAs) are a class of RNA molecules longer than 200 nucleotides that lack protein-coding potential and are generally poorly conserved [4]. They regulate diverse biological processes through mechanisms such as anchoring transcription factors, modulating alternative splicing, activating microRNAs, and recruiting chromatin-modifying enzymes [5]. In recent years, lncRNAs have received increasing attention because of their crucial roles in various human diseases, including breast cancer [6]. Several studies have found that lncRNAs are aberrantly expressed in a variety of cancers, including breast cancer [7]. Such dysregulated lncRNAs may serve as valuable biomarkers for cancer screening and diagnosis [8,9]. Moreover, their altered expression can influence tumor progression and chemotherapy resistance [10,11]. For example, LINC00467, a testis-specific lncRNA, has been shown to reduce viability and promote apoptosis when silenced in adult neuroblastoma cells [12,13]. Consistent with these findings, our previous study demonstrated that elevated LINC00467 expression was associated with poorer overall survival in breast cancer patients. Furthermore, we revealed that LINC00467 promotes breast cancer progression by directly interacting with miR-138-5p and LIN28B [14]. However, the utilization of spatial transcriptomics and single-cell transcriptome data remains relatively low, with a vast amount of spatially specific gene expression and function yet to be elucidated. The role of lncRNAs in the spatial heterogeneity of breast cancer is almost entirely unexplored. In this study, we integrated spatial transcriptomics, single-cell RNA sequencing, and bulk RNA sequencing data for analysis and identified a spatially highly variable lncRNA—LINC00467—with important functions in breast cancer.
The pronounced heterogeneity and stemness of breast cancer cells contribute to their low sensitivity to therapeutic drugs [15]. In recent years, spatial transcriptomics and single-cell transcriptomics have become important approaches for elucidating the pathogenesis of breast cancer [16]. However, their application remains limited, and the expression and function of many spatially specific genes are still poorly characterized. In particular, the role of lncRNAs in the spatial heterogeneity of breast cancer remains largely unexplored.
In this study, we integrated spatial transcriptomic, single-cell sequencing, and bulk RNA sequencing data to identify LINC00467, a spatially highly variable lncRNA with potential functional significance in breast cancer. However, its relationship with breast cancer stemness and drug sensitivity has not been fully clarified. Here, we conducted cell experiments, molecular biology experiments and a series of bioinformatics analysis to explore the role of LINC00467 in breast cancer prognosis and drug treatment response.
Spatial transcriptomic and single-cell sequencing data from breast cancer patients were downloaded from STOmicsDB (https://db.cngb.org/stomics/), the spatial resolution of the spatial transcriptome chip is 55 μm. Venny 2.1 online tool (https://bioinfogp.cnb.csic.es/tools/venny/index.html) was used to draw the venn diagram. Clustering analysis was performed in R (v4.2.1) using the ‘Seurat’ package (v4.2.3). Specifically, the FindClusters function was applied to the spatial transcriptomic data, and each cluster was classified and labeled as either “Tumor” or “Non-Tumor” based on corresponding pathological images. The VlnPlot function was then used to visualize the expression of target lncRNAs across clusters. Spatially specific genes were identified using SpatialDE (https://github.com/Teichlab/SpatialDE) with default parameters. Genes with the lowest q-values were intersected with lncRNAs annotated in the Ensembl database (https://www.ensembl.org/index.html?redirect=no) [17], yielding 10 spatially specific lncRNAs. These 10 candidates were further intersected with lncRNAs significantly differentially expressed in the TCGA breast cancer (BRCA) cohort available in GEPIA2 (|log2FC| > 1, p < 0.05), and the final target molecules were obtained. Using the TCGA BRCA cohort data from the BEST online database (https://rookieutopia.hiplot.com.cn/app_direct/BEST/), samples were stratified into high- and low-expression groups based on the median expression level of LINC00467. Gene Set Enrichment Analysis (GSEA) and mapping of the top 20 high-frequency mutated genes were then performed. Overall survival (OS) curves were generated using the Kaplan-Meier Plotter online tool ( http://kmplot.com/analysis/index.php?p=service) [18]. Expression levels of LINC00467 across the four breast cancer subtypes, as well as corresponding image mapping, were analyzed using GEPIA2 (http://gepia2.cancer-pku.cn/#index) [19], based on the TCGA BRCA cohort data. The correlation between LINC00467 expression and ER, KI67, PR positive and treatment methods in breast cancer patients was analyzed using the dataset GSE21653 and GSE9893 [20,21]. The single-cell RNA sequencing dataset GSE212461 was used to analyze the expression pattern of LINC00467 in different breast cancer cell subclusters [22]. Finally, correlations between LINC00467 and ALDH2, ALDH6A1, CD44, LIN28B, SP1, and E2F1 were assessed using GEPIA2. Associations between LINC00467 and tumor-associated signaling pathways in breast cancer were analyzed through GSCA (https://guolab.wchscu.cn/GSCA/#/) based on TCGA BRCA cohort data [23]. The competitive endogenous RNA (ceRNA) regulatory network analysis of LINC00467 was performed using the RNAInter online database (http://www.rnainter.org/) [24]. AnimalTFDB (https://guolab.wchscu.cn/AnimalTFDB4/#/) [25], RegNetwork (http://www.regnetworkweb.org/search.jsp) [26], and PROMO (http://alggen.lsi.upc.es/cgi-bin/promo_v3/promo/promoinit.cgi?dirDB=TF_8.3) [27] online tools were used to predict the transcription factors of LINC00467, and finally take the intersection of the prediction results of the three databases. We used the chromatin immunoprecipitation seq (chip-seq) data of SP1 and E2F1 of MCF-7 and MDA-MB-231 cells in the UCSC database [28] to analyze the binding of these two transcription factors in the promoter region of LINC00467.
2.2 Cell Culture and Transfection
All cell lines (MCF-7 and MDA-MB-231) used in this study were purchased from Wuhan Procell Company, authenticated by short tandem repeat (STR) profiling, and confirmed to be free of mycoplasma contamination. Cells were cultured in Dulbecco’s Modified Eagle’s Medium (Thermo Fisher Scientific, cat#11960044, Waltham, MA, USA) containing 10% fetal bovine serum (Thermo Fisher Scientific, cat#A5670801, Waltham, MA, USA), 100 U/mL penicillin, and 100 µg/mL streptomycin (Thermo Fisher Scientific, cat#15140122, Waltham, MA, USA). Cultures were maintained at 37°C in a humidified atmosphere with 5% CO2. Plasmid transfection was carried out using the Lipofectamine 3000 Transfection Kit (Thermo Fisher Scientific, cat#L3000015, Waltham, MA, USA) following the manufacturer’s instructions. Small interfering RNA (siRNA) sequences targeting LINC00467 were synthesized by RiboBio (Guangzhou, China, cat#siB000000). The sequence used was: LINC00470-siRNA: CTGAGTTGCAGAAACAAAT.
2.3 Quantitative Real-Time PCR
MCF-7 and MDA-MB-231 cells were washed three times with pre-cooled phosphate-buffered saline (1× PBS, pH = 7.4), and total RNA was extracted using TRIzol reagent (2 mL TRIzol per 60 mm dish; Thermo Fisher Scientific, cat#10296028CN, Waltham, MA, USA). A 1–2 μL aliquot was used to assess RNA concentration and quality at wavelengths of 260 and 280 nm with a spectrophotometer (Thermo Fisher Scientific, Waltham, MA), and the remaining RNA was stored at –80°C. The expression of LINC00467 and LIN28B was quantified by real-time fluorescence quantitative polymerase chain reaction (qPCR) using the LightCycler 480 system (Roche, Basel, Switzerland), with β-actin serving as the internal control. Primers were designed and synthesized by Shanghai Biotechnology Co., Ltd., with the following sequences: LINC00467-F: 5′-TCGTCTTCAGGAAGCCAGAC-3′; LINC00467-R: 5′-TGGAAATCAAAAGGGTCAGC-3′; LIN28B-F: 5′-CACGAGTTTGGAGCTGAGGG-3′; LIN28B-R: 5′-AGGTAGACTTTGCAACCGGG-3′; β-actin-F: 5′-TCACCAACTGGGACGACATG-3′; β-actin-R: 5′-GTCACCGGAGTCCATCACGAT-3′.
Digested MCF-7 and MDA-MB-231 cells were resuspended in culture medium, and cell numbers were determined using a hemocytometer (Hausser Scientific, Horsham, PA, USA) under the 1 mm × 1 mm × 0.1 mm grid counting mode. Cells were seeded at a density of 1000 cells per well for colony formation assays. After 10 days of incubation, colonies were washed twice with 1× PBS (pH = 7.4), stained with 1% crystal violet for 15 min, and photographed for colony counting.
Protein blotting was performed as previously described [29]. Briefly, total protein was extracted from MCF-7 cells using RIPA lysis buffer (Beyotime, cat# P0038, Beijing, China), and protein concentrations were determined with the BCA Protein Assay Kit (Thermo Fisher Scientific, cat#A65453, Waltham, MA, USA). Equal amounts (50 μg) of protein were separated on 10% SDS-PAGE gels and transferred onto polyvinylidene difluoride (PVDF) membranes. Membranes were blocked with non-fat powdered milk (Beyotime, cat# P0216-300g, Beijing, China) at room temperature for 1 h and then incubated overnight at 4°C with the following primary antibodies: anti-AKT (1:1000, Cell Signaling Technology, cat#9272, Danvers, MA, USA), anti-p-AKT (1:1000, Cell Signaling Technology, cat#4060, Danvers, MA, USA), and monoclonal mouse anti-GAPDH (1:3000, ProMab Biotechnologies, cat#20035, Changsha, China). After washing, membranes were incubated with the appropriate secondary antibody (1:2000, Cell Signaling Technology, cat#7076, Danvers, MA, USA) for 1 h at room temperature. Protein bands were visualized using an enhanced chemiluminescence (ECL) detection kit (Millipore, cat#WBKLS0500, Darmstadt, Germany). Each experiment was repeated three times.
Statistical analyses were performed using GraphPad Prism 9 software (GraphPad Software, San Diego, CA, USA). Data are presented as mean ± SEM. Comparisons between two groups were conducted using Student’s t-test or Wilcoxon test. Correlation analysis was performed using Spearman correlation analysis. Survival analysis was performed using the log-rank test. And p < 0.05 was considered statistically significant.
3.1 The Expression of LINC00467 in Breast Cancer and Its Correlation with the Clinicopathological Characteristics of Patients with Breast Cancer
To identify spatially highly variable lncRNAs associated with breast cancer, we first downloaded the top 500 candidates from the STOmicsDB spatial transcriptome database and intersected them with lncRNAs annotated in the Ensembl database [30]. Differentially expressed lncRNAs in breast cancer were further filtered using the GEPIA2 database [19]. This analysis yielded two breast cancer–associated spatially highly variable lncRNAs, LINC00467 and LINC00342 (Fig. 1A). Downscaling clustering of the STOmicsDB spatial transcriptome data using the ‘Seurat’ package in R identified 13 subgroups (Fig. 1B,C). Pathologist-annotated Ensembl markers were then used to characterize each subgroup (Fig. 1D,E). We observed marked heterogeneity in LINC00467 expression across subgroups: it was nearly absent in non-tumor tissues but strongly expressed in tumor tissues (Fig. 1F). Analysis of an additional slice from the same patient confirmed the consistent expression pattern and spatial localization of LINC00467 in breast cancer (Fig. 1G,H). However, LINC00342 showed a lower positive rate and weaker expression in breast cancer samples compared with LINC00467, and it was also detected in non-tumor regions (highlighted by red circles) (Fig. A1). Based on these findings, LINC00467 was selected for subsequent analyses.
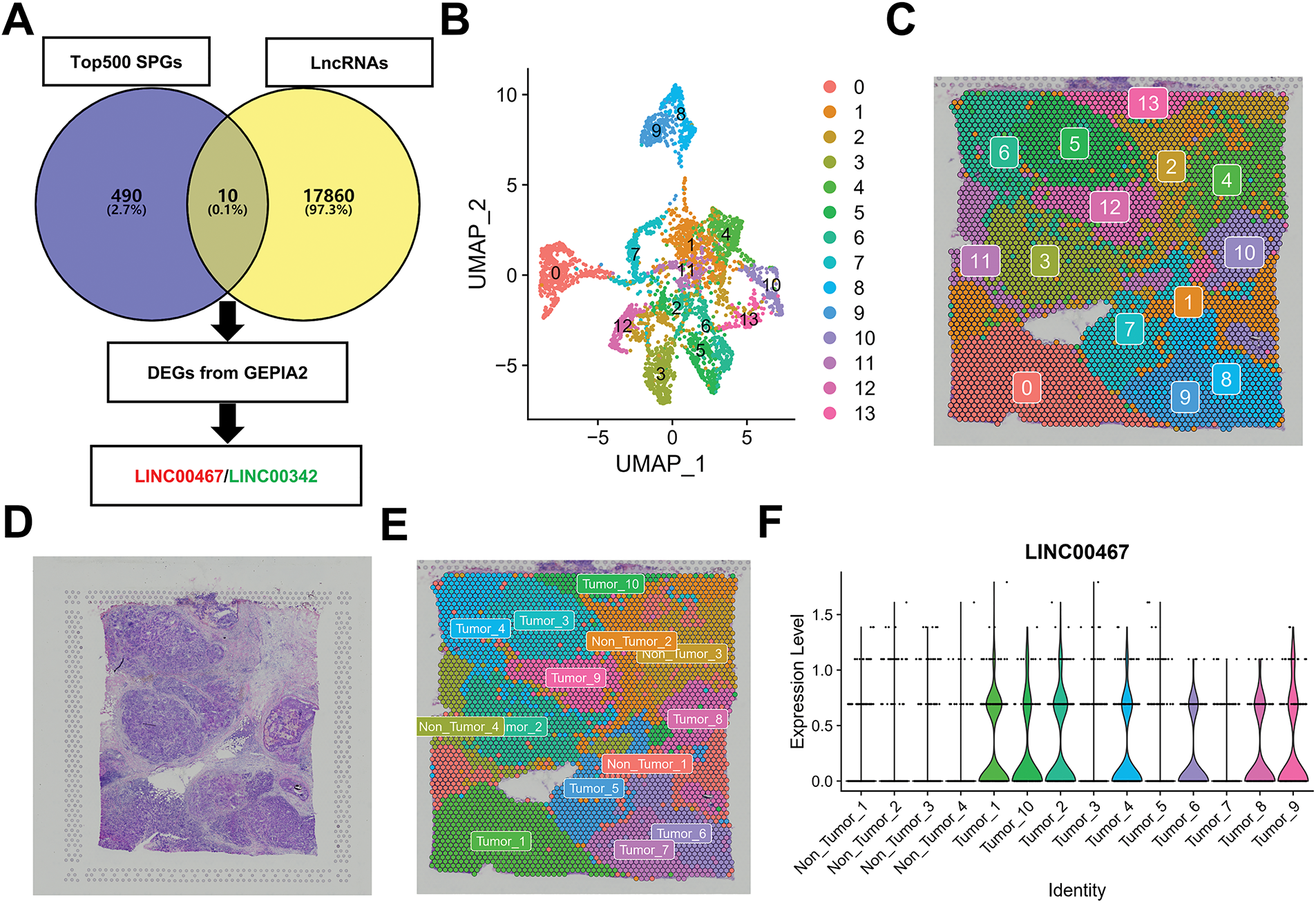
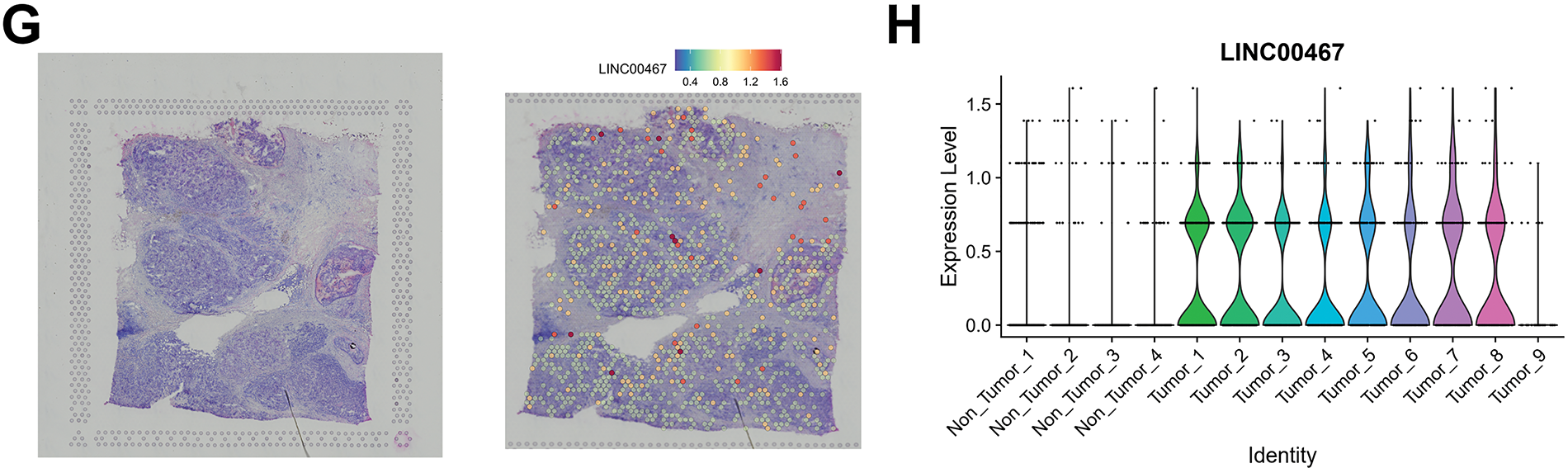
Figure 1: Screening of spatially hypervariable lncRNAs. (A) Workflow for identifying breast cancer spatially hypervariable lncRNAs: the top 500 spatially variable genes from STOmicsDB were intersected with lncRNAs from the Ensembl database, and further filtered using GEPIA2, yielding LINC00467 and LINC00342. (B,C) Downscaling clustering analysis of spatial transcriptome data using the ‘Seurat’ package identified 13 subgroups. (D) Hematoxylin and eosin (H&E) staining of the corresponding patient section. (E) Pathologist’s annotation of each subgroup. (F) Expression of LINC00467 across subgroups (Section 1), showing minimal expression in non-tumor tissues and high expression in tumor tissues. (G) Analysis of an additional breast cancer section from STOmicsDB confirmed consistent expression and spatial localization of LINC00467. (H) The expression level of LINC00467 in each subgroup of Section 2.
We used the GEPIA2 database to analyze the correlation between LINC00467 expression and the clinicopathological characteristics of breast cancer patients. LINC00467 was significantly upregulated in Luminal A and Luminal B subtypes and showed a slight, non-significant increase in Basal-like and HER2 subtypes (Fig. 2A). Its expression was not associated with the age of breast cancer adenocarcinoma patients (Fig. 2B). However, LINC00467 expression was positively correlated with estrogen receptor (ER) and progesterone receptor (PR) expression but negatively correlated with the proliferation marker Ki67 (Fig. 2C). Furthermore, LINC00467 expression varied according to treatment regimen, being significantly higher in patients treated with X-ray compared with those treated with tamoxifen (Fig. 2D).
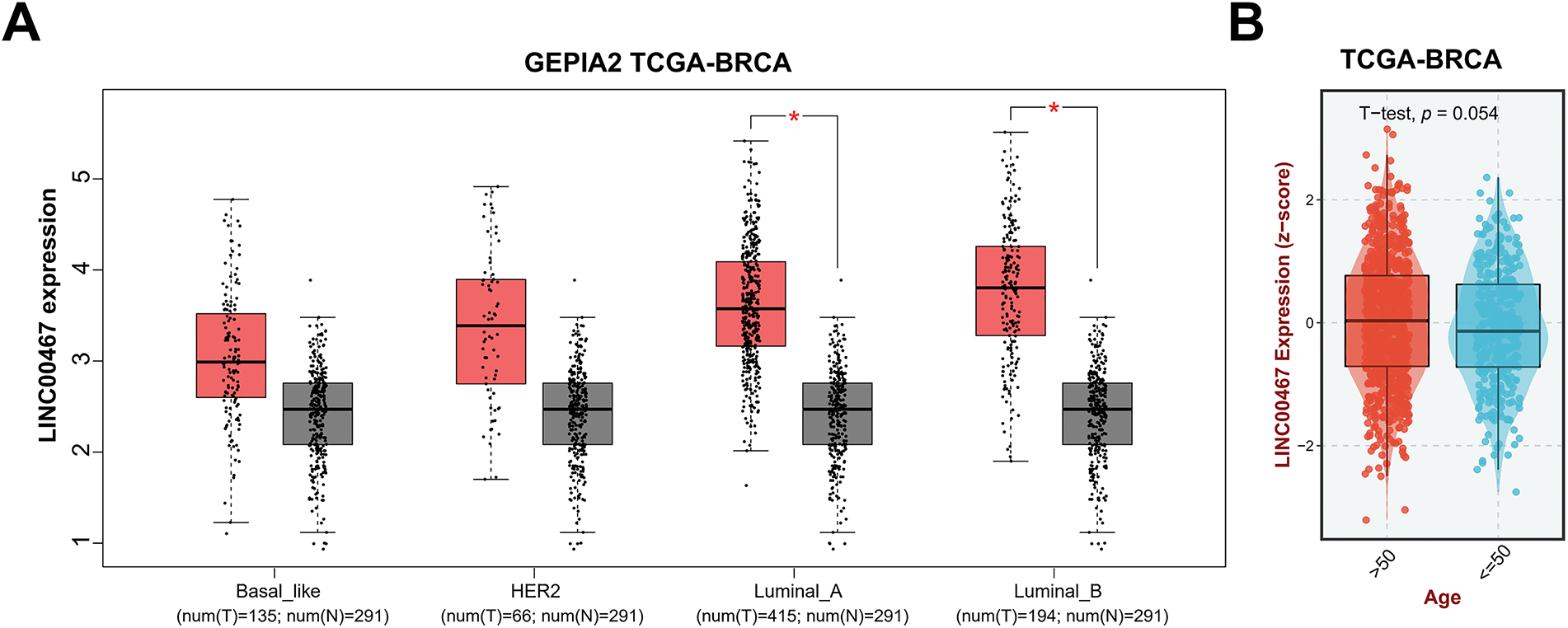
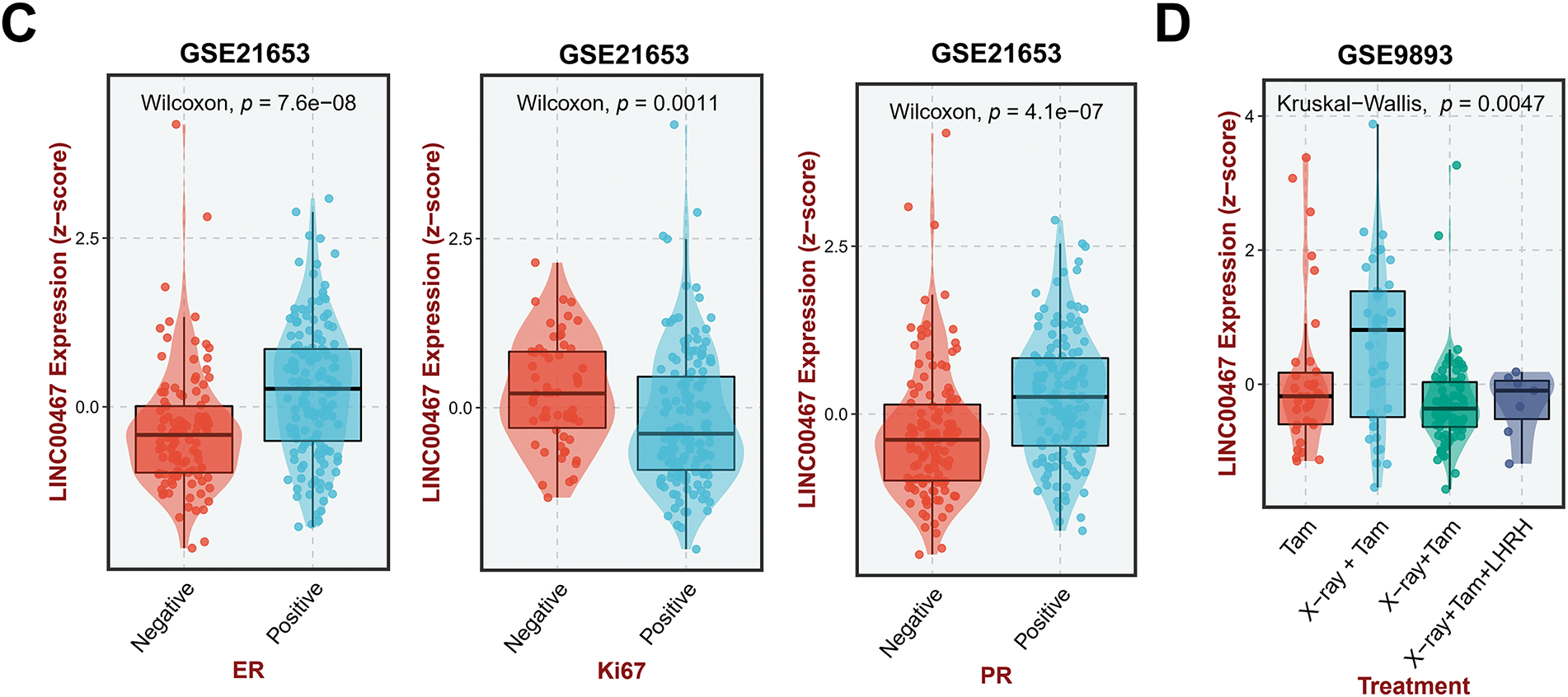
Figure 2: Correlation of LINC00467 expression with clinicopathological characteristics of breast cancer patients. (A) LINC00467 expression was significantly upregulated in Luminal A and Luminal B subtypes, with a slight but non-significant increase in Basal-like and HER2 subtypes. (B) LINC00467 expression showed no correlation with patient age. (C) LINC00467 expression was higher in estrogen receptor (ER)-positive and progesterone receptor (PR)-positive breast cancer patients but lower in Ki67-positive patients. (D) LINC00467 expression varied across treatment regimens and was significantly higher in patients treated with X-ray compared with those treated with luteinizing hormone-releasing hormone (LHRH) or tamoxifen (Tam). *p < 0.05.
We further integrated single-cell sequencing and the TCGA data to analyze the correlation between LINC00467 and tumor cell stemness in breast cancer patients. Tumor cells from the GSE212461 dataset were downscaled into four subgroups (Fig. 3A,B). LINC00467 showed low expression in subgroup 2 and high expression in subgroup 0. Proliferation and stemness analysis revealed that the proliferation marker MKI67 was highly expressed in subgroup 2, whereas breast cancer stemness-related markers were highly expressed in subgroup 0 (Fig. 3C–E). To further substantiate their relationship, we examined the spatial co-localization of LINC00467 with stem cell markers (e.g., CD44). The data demonstrated a positive correlation between LINC00467 and the stem cell marker CD44 at the gene expression level (Fig. 3F). The positive correlation between LINC00467 and stemness-related markers (ALDH2, ALDH6A1, and CD44) was further confirmed using TCGA BRCA data (Fig. 3G). Collectively, these findings suggest that high LINC00467 expression represents a poor prognostic factor in breast cancer and may contribute to maintaining tumor cell stemness.
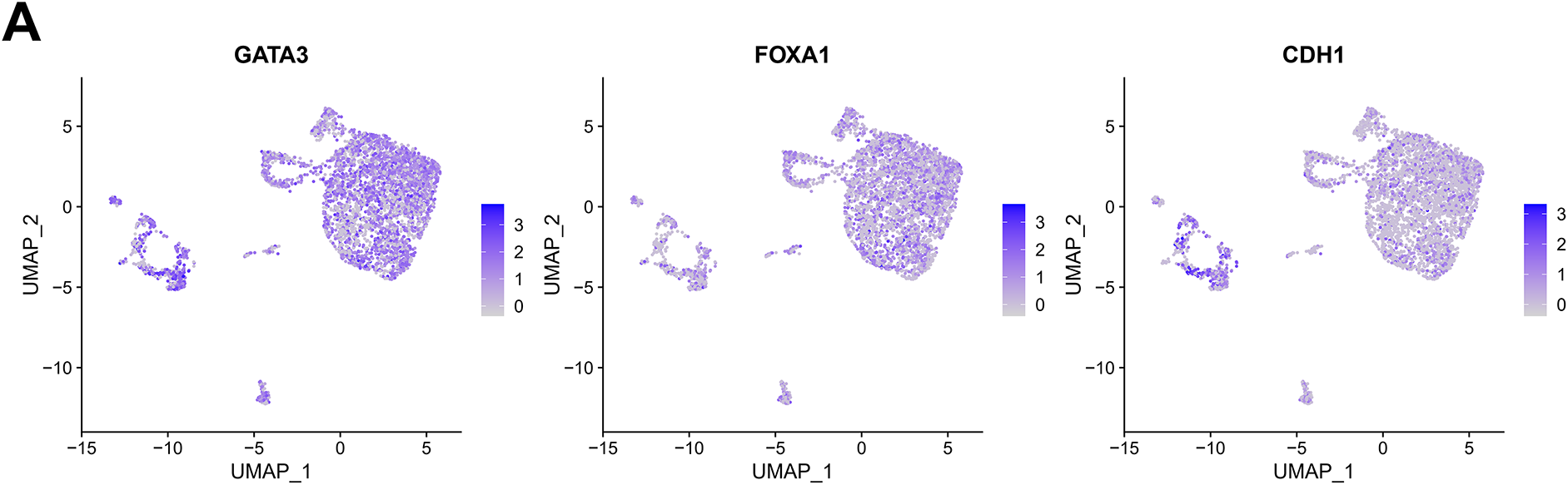
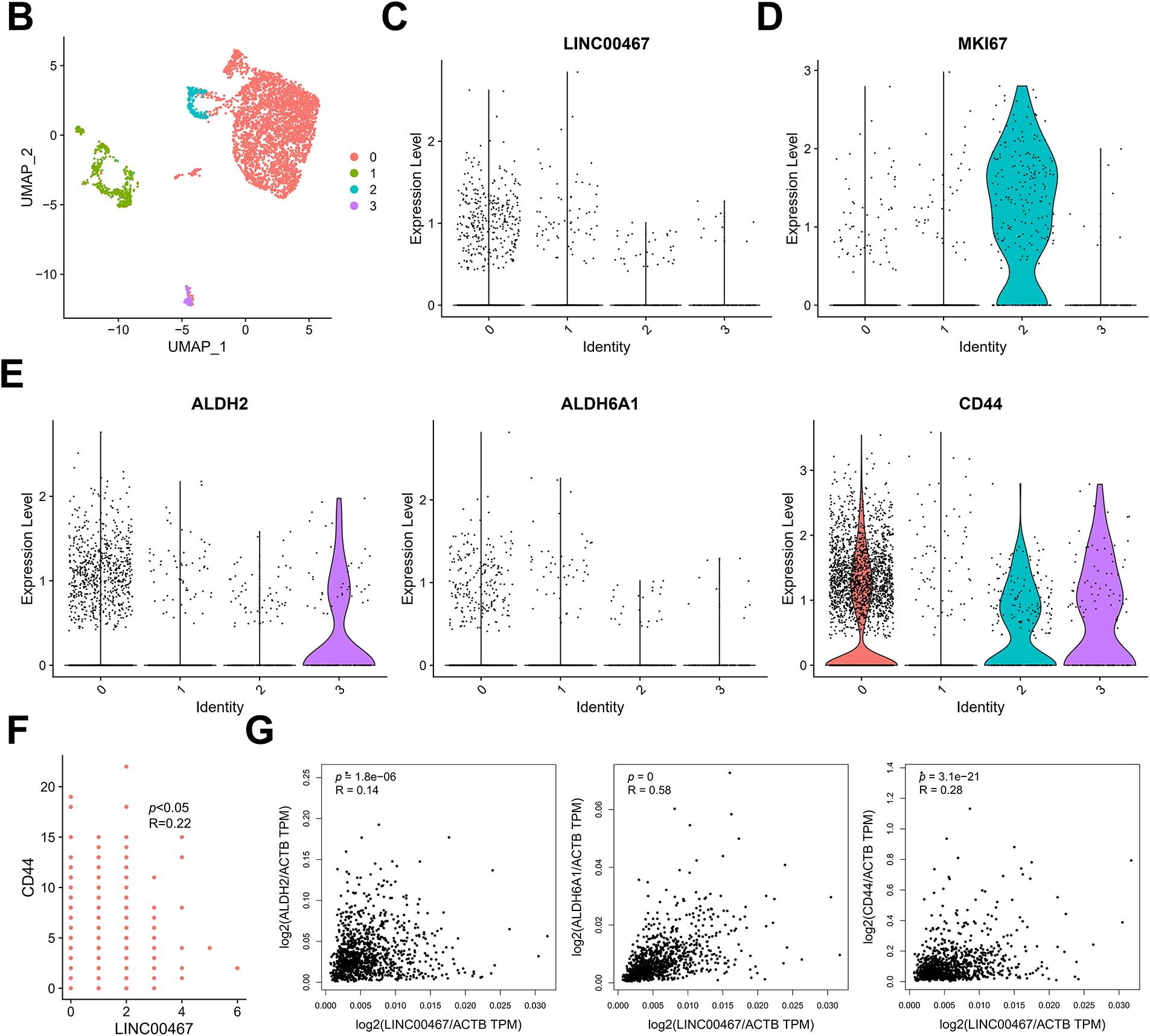
Figure 3: LINC00467 is associated with stemness in breast cancer. (A) Marker gene display of breast cancer cells in the GSE212461 dataset. (B) Tumor cells in the GSE212461 dataset were downscaled into 4 subpopulations. (C) LINC00467 showed higher expression in subpopulation 0. (D) Proliferation marker MKI67 was highly expressed in subpopulation 2. (E) The breast cancer stemness-associated marker was highly expressed in subpopulation 0. (F) Spatial transcriptomics analysis identified a significant positive correlation between LINC00467 and the stem cell marker CD44 at the gene expression level. (G) Validation of the positive correlation between LINC00467 and stemness-associated markers (ALDH2, ALDH6A1, and CD44) using TCGA BRCA data.3.2 Silencing of LINC00467 significantly inhibited clone formation in breast cancer cells as well as cancer cell stemness.
3.2 Silencing of LINC00467 Significantly Inhibited Clone Formation in Breast Cancer Cells as Well as Cancer Cell Stemness
To further investigate the function of LINC00467 in breast cancer, we designed siRNA specifically targeting LINC00467. Following transfection in both MCF-7 and MDA-MB-231 cells, LINC00467 expression was significantly downregulated (Fig. 4A,B). Colony formation assays revealed that silencing LINC00467 markedly reduced the clonogenic capacity of both MCF-7 and MDA-MB-231 cells (Fig. 4C,D). In an effort to elucidate the functional partners of LINC00467, we conducted computational screening using the RNAInter database (http://www.rnainter.org/). The analysis revealed that LINC00467 may interact with several let-7 family miRNAs, including hsa-let-7b-5p, hsa-let-7e-5p, hsa-let-7g-5p, and hsa-let-7i-5p [24]. Given that let-7 family miRNAs have been well-established as negative regulators targeting LIN28B [31], a key factor in modulating cellular stemness, we propose that LINC00467 may function as a molecular sponge for let-7 miRNAs, thereby alleviating their inhibitory effect on LIN28B. (Fig. A2). Given that the multipotency factor LIN28B is known to promote stemness in breast cancer cells, we next analyzed data from the TCGA BRCA cohort and identified a significant positive correlation between LINC00467 and LIN28B expression (Fig. 4E). Consistently, knockdown of LINC00467 significantly suppressed LIN28B expression (Fig. 4F), suggesting that LINC00467 may regulate breast cancer stemness, at least in part, through LIN28B.
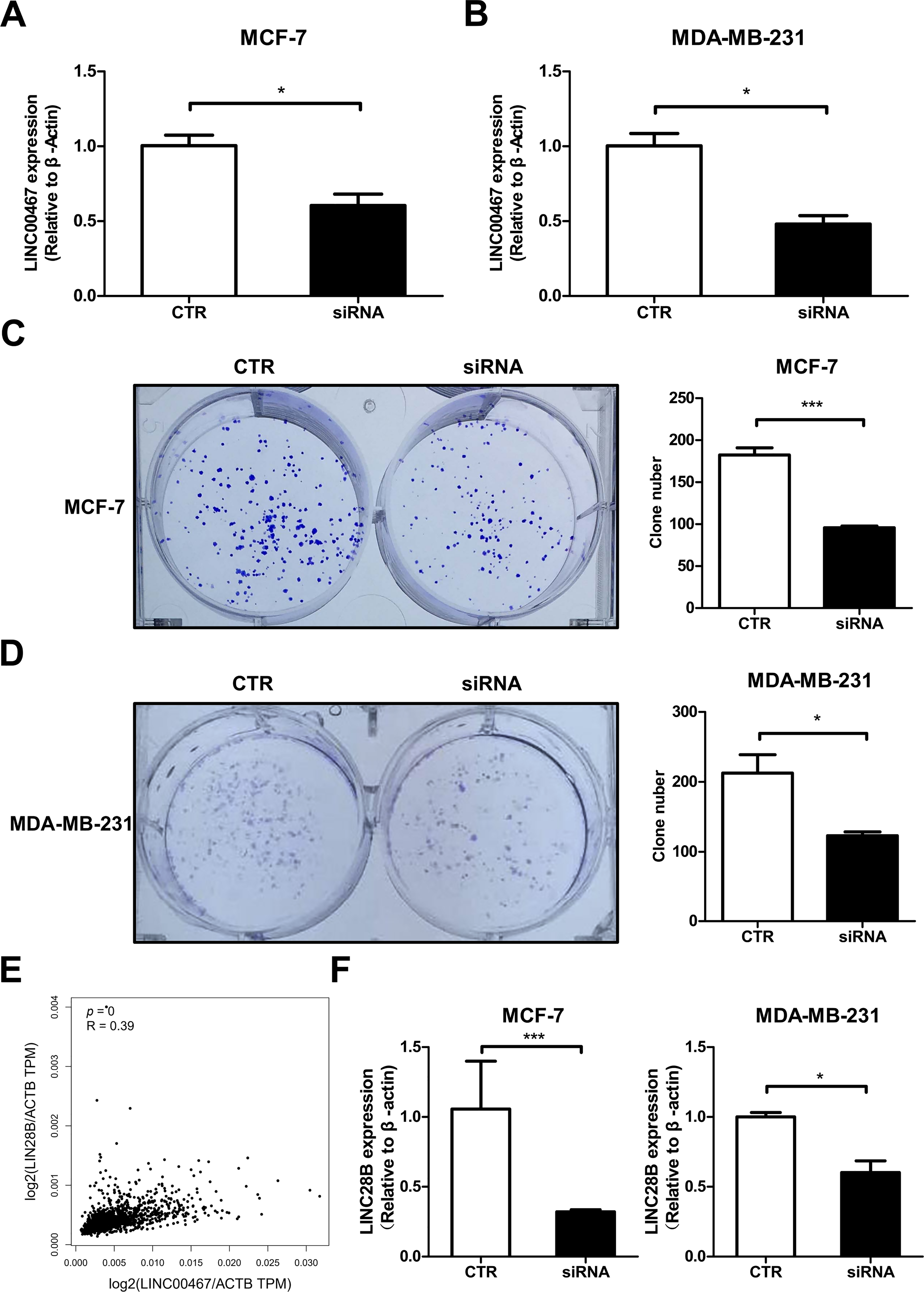
Figure 4: LINC00467 correlates with clonogenic ability and stemness genes in breast cancer. (A,B) qRT-PCR detection of LINC00467 expression in MCF-7 and MDA-MB-231 cells after transfection with LINC00467 siRNA, CTR, control.(C,D) Silencing of LINC00467 significantly inhibits breast cancer cell clonogenic ability. (E) TCGA BRCA cohort analysis showing a significant positive correlation between LINC00467 and LIN28B expression. (F) qRT-PCR analysis of LIN28B expression after silencing LINC00467. Knockdown of LINC00467 significantly suppressed LIN28B expression. *p < 0.05, ***p < 0.001.
3.3 LINC00467 Alters AKT Signaling and Correlates with Drug Treatment Sensitivity
To further explore the downstream signaling pathways regulated by LINC00467, we analyzed its association with tumor-related pathways using the TCGA BRCA cohort data. LINC00467 expression showed a significant positive correlation with the PI3K/AKT pathway (Fig. 5A). Consistently, silencing LINC00467 markedly reduced both total AKT and phosphorylated AKT protein levels (Fig. 5B). Moreover, breast cancer cells with high LINC00467 expression exhibited increased sensitivity to AKT inhibitors compared with those with low LINC00467 expression (Fig. 5C,D). We further examined the mutational landscape and its relationship with immunotherapeutic response in patients stratified by LINC00467 expression. Using TCGA BRCA cohort data from the BEST online database, samples were divided into high- and low-expression groups based on the median LINC00467 expression level. The top 20 most frequently mutated genes revealed that TP53 mutations were less common in the high-expression group, whereas PIK3CA mutations were more prevalent (Fig. 6A). In addition, LINC00467 expression was significantly higher in patients who were non-responsive (NR) to anti-PD-1 immunotherapy compared with responders, suggesting that LINC00467 may serve as a predictive marker of immunotherapy efficacy (Fig. 6B,C). It is noteworthy that higher expression of LINC00467 correlates with poorer prognosis in patients receiving anti-PD-1 therapy, whereas it is associated with better prognosis in those treated with anti-PD-L1 therapy. (Fig. 6D–F). We performed a correlation analysis using the publicly available BRCA dataset from the GEPIA2 database. The results demonstrated a significant positive correlation between the expression of LINC00467 and CD274 (PD-L1) (Fig. A3).
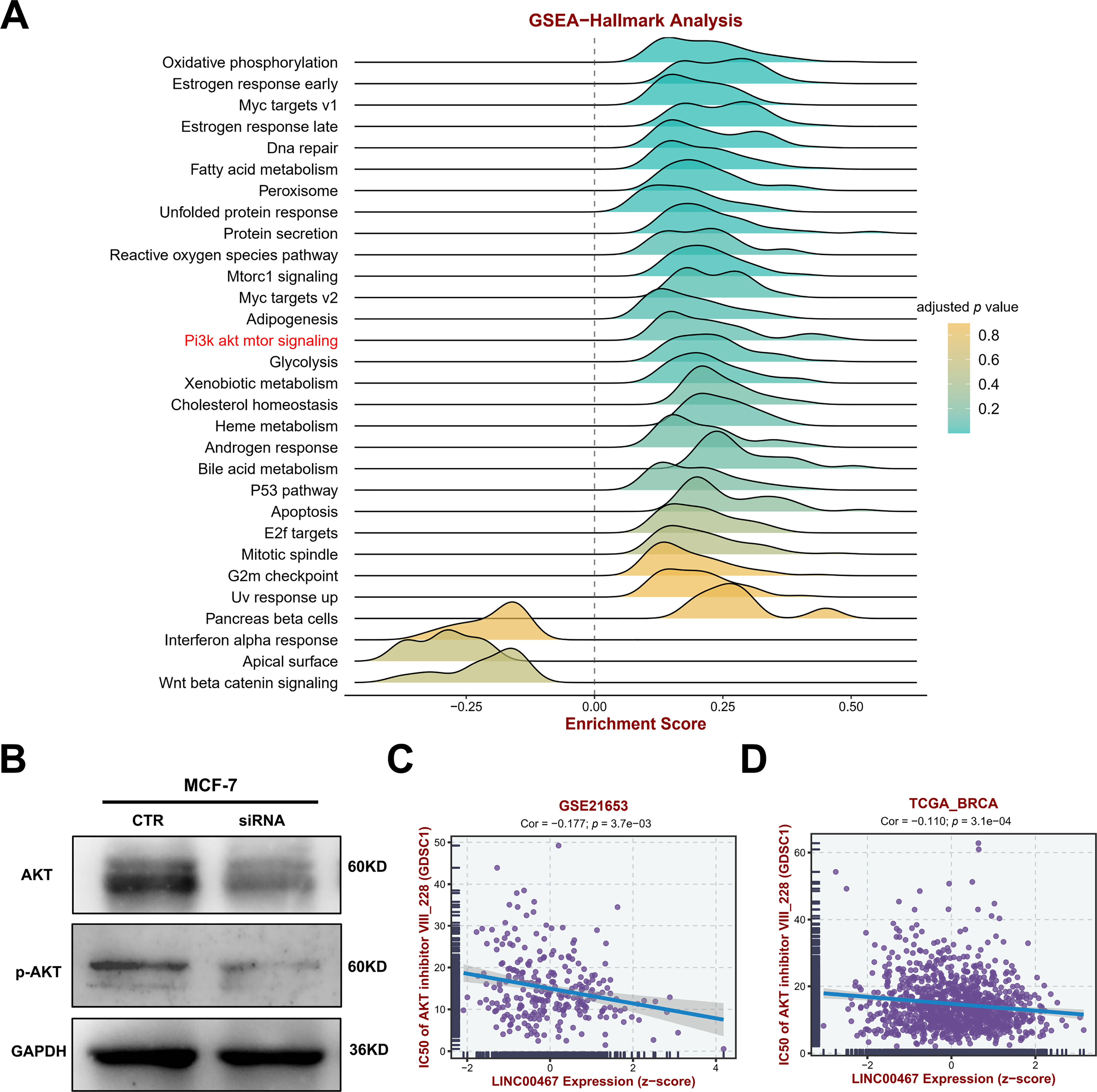
Figure 5: LINC00467 regulates AKT signaling and correlates with AKT-related drug sensitivity. (A) GSEA enrichment analysis using TCGA BRCA cohort data from the BEST online database, dividing samples into two groups of high and low LINC00467 expression based on median expression, showing that LINC00467 was significantly associated with AKT signaling. (B) Silencing of LINC00467 significantly inhibited AKT expression. (C,D) Analysis of the relationship between LINC00467 expression and AKT-related drug sensitivity using GSE21653 and TCGA BRCA data from the BEST online tool. LINC00467 expression was significantly negatively correlated with AKT-related drug sensitivity.
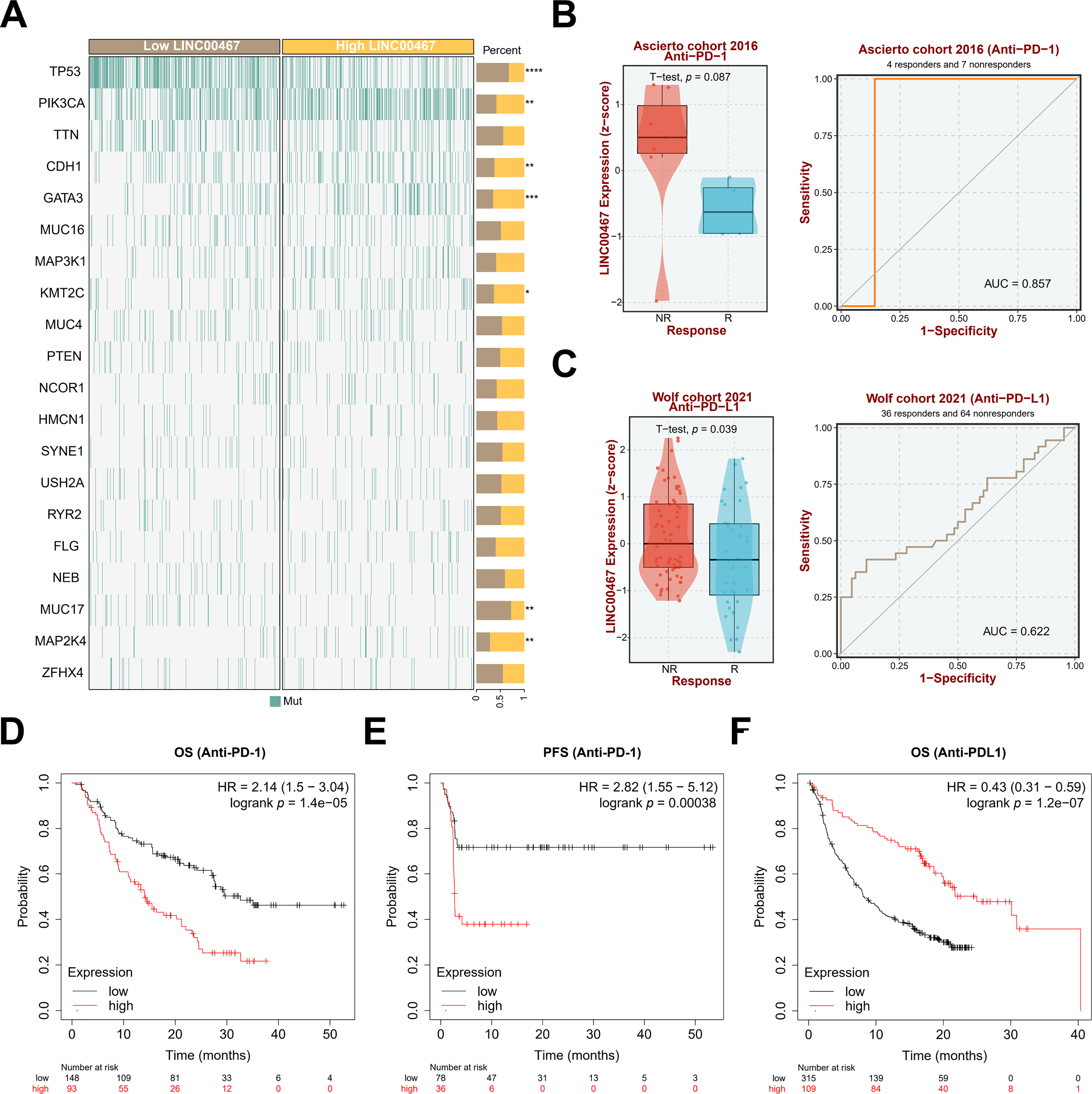
Figure 6: Mutation profiles in high- and low-LINC00467 expression groups and their relationship with immunotherapy response in tumor patients. (A) TCGA BRCA cohort data from the BEST online database were stratified into high- and low-LINC00467 expression groups to generate the top 20 most frequently mutated genes. TP53 showed a lower mutation frequency in the high-LINC00467 group, whereas PIK3CA and CDH1 mutations were more frequent. (B) LINC00467 expression was significantly higher in the non-responsive (NR) group compared with the responsive (R) group of patients receiving anti-PD-1 immunotherapy. (C) Elevated LINC00467 expression was associated with worse prognosis among patients treated with anti-PD-1 immunotherapy. (D) The correlation between the expression level of LINC00467 and the overall survival of patients treated with anti-PD-1. (E) The correlation between the expression level of LINC00467 and the progression-free survival of patients treated with anti-PD-1. (F) The correlation between the expression level of LINC00467 and the overall survival of patients treated with anti-PDL-1. *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001.
3.4 SP1 and E2F1 May be the Regulatory Transcription Factors of LINC00467
To explore the upstream transcriptional regulation of LINC00467, three transcription factor prediction tools—AnimalTFDB (https://guolab.wchscu.cn/AnimalTFDB4/#/), RegNetwork (http://www.regnetworkweb.org/search.jsp), and PROMO (http://alggen.lsi.upc.es/cgi-bin/promo_v3/promo/promoinit.cgi?dirDB=TF_8.3)—were used. The intersection of predictions from these databases identified two potential transcription factors, SP1 and E2F1 (Fig. 7A). LINC00467 expression was positively correlated with both factors (Fig. 7B). Analysis of publicly available ChIP-seq data revealed significant peaks for E2F1 and SP1 in the promoter region of LINC00467 (Fig. 7C), suggesting that these transcription factors may directly regulate its expression. E2F1 expression was significantly upregulated in Luminal A, Luminal B, Basal-like, and HER2 breast cancer subtypes (Fig. A4A). Stratification of patients into high- and low-E2F1 groups based on median expression showed that higher E2F1 expression was associated with poorer overall survival (Fig. A4B). In contrast, SP1 expression was significantly elevated in Luminal A and Luminal B breast cancer tissues compared with normal tissues but showed no significant difference in Basal-like or HER2 subtypes (Fig. A4C). Notably, higher SP1 expression was associated with better survival (Fig. A4D).
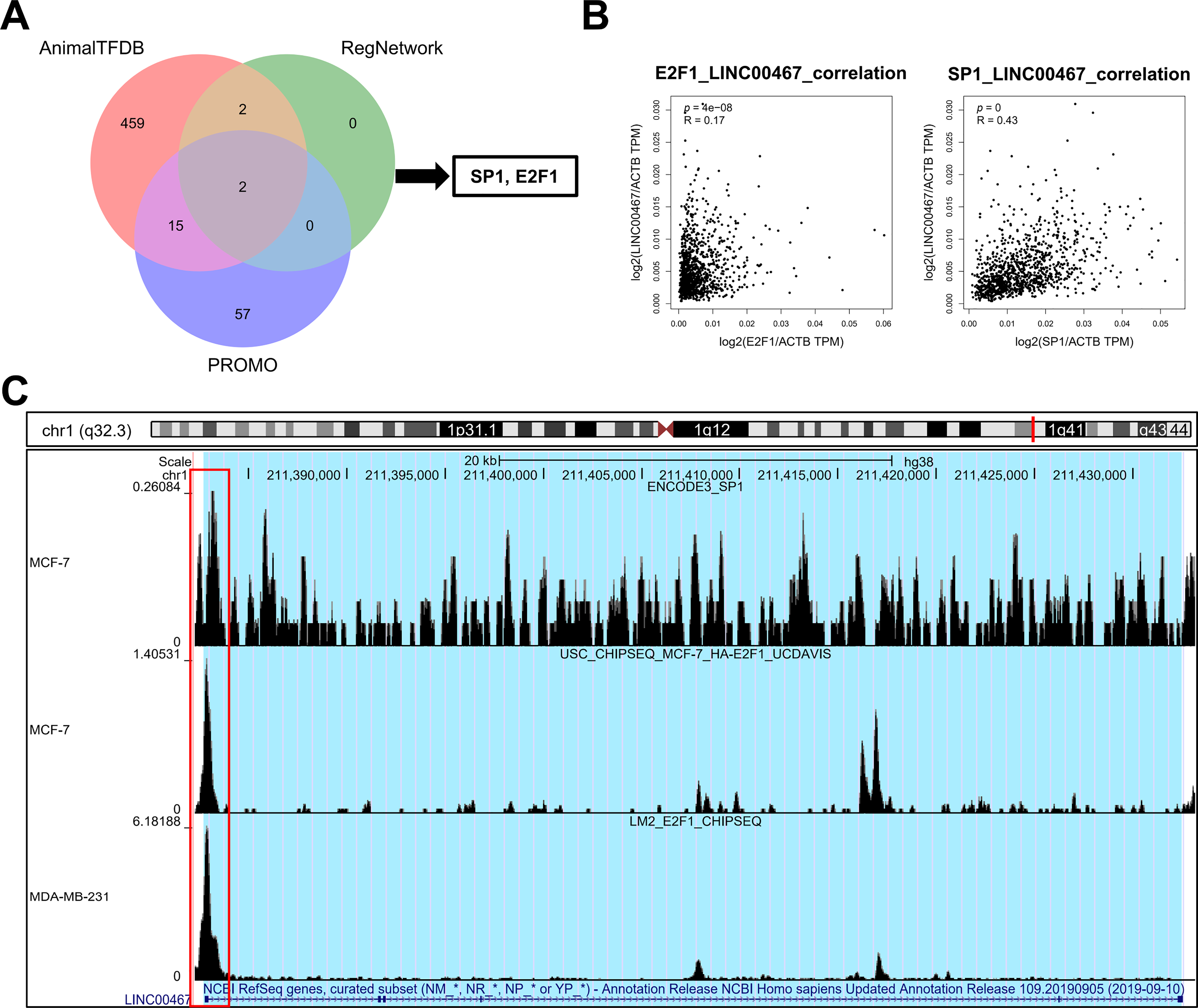
Figure 7: Prediction and validation of transcription factors regulating LINC00467. (A) Transcription factors of LINC00467 were predicted using three databases (AnimalTFDB, RegNetwork, and PROMO), with E2F1 and SP1 identified as common candidates. (B) Correlation analysis showed that LINC00467 expression was positively associated with E2F1 and SP1 expression. (C) Chip-seq data revealed significant binding peaks for E2F1 and SP1 near the LINC00467 promoter region, suggesting direct transcriptional regulation.
The intra-tumoral heterogeneity (ITH) theory differs from the molecular typing theory of breast cancer. ITH refers to the coexistence of multiple subtypes within the same tumor (spatial heterogeneity) and the dynamic interconversion between phenotypes over time (temporal heterogeneity) [32]. Single-cell RNA sequencing (scRNA-seq) enables detailed characterization of cell states and population heterogeneity, thereby providing insights into the mechanisms underlying breast cancer progression and metastasis, and offering a foundation for individualized therapy [33]. Using scRNA-seq, researchers have characterized tumor heterogeneity across different molecular subtypes and identified specific cell populations linked to poor prognosis or therapeutic response [34]. Single-cell profiling of the breast cancer tumor microenvironment has revealed immune cell subpopulations that may be potential targets for immunotherapy [35]. In addition, scRNA-seq can be applied to investigate intercellular signaling, regulatory single-cell states, and immune cell distribution in breast cancer [36], as well as to assess the relationship between immune infiltration and treatment outcomes [37]. Despite these advances, several challenges remain. These include elucidating the role of ITH in limiting the efficacy of targeted therapies, devising strategies to identify drug-resistant subpopulations that contribute to poor long-term outcomes, and developing rational combination therapies that simultaneously target multiple signaling pathways [38]. Methodologically, single-cell suspensions prepared through enzymatic digestion disrupt the spatial context of cells within tissues. By contrast, spatial transcriptomics preserves spatial localization while capturing transcriptomic information, thereby providing more accurate insights into in situ gene expression, cellular functions, microenvironmental interactions, lineage tracing, and disease pathology [39,40].
In this study, we integrated spatial transcriptomics, single-cell sequencing, and bulk sequencing analyses to demonstrate that LINC00467, which displays spatial expression characteristics, is associated with breast cancer stemness, immunotherapy response, and drug sensitivity. Functional experiments further showed that silencing LINC00467 inhibited the clonogenic capacity of breast cancer cells, potentially through regulation of the AKT pathway. AKT, also known as protein kinase B, is a central regulator of proliferation, metastasis, and invasion in breast cancer and is linked to multiple oncogenic markers and metastatic cascades [41]. Dysregulation of the PI3K/AKT signaling pathway occurs in up to 70% of breast cancers and is consistently associated with poor prognosis, underscoring AKT as an attractive therapeutic target [42]. LncRNAs with tissue-specific expression profiles may represent tissue specificity at different stages of breast cancer and thus may serve as biomarkers or therapeutic targets for breast cancer diagnosis and prognosis [43]. Several lncRNAs associated with breast cancer stemness have been identified, and once validated in prospective studies, they could enhance prediction accuracy and guide individualized treatment [44]. Notably, LIN28 is an RNA-binding protein first identified in C. elegans and plays an important role in its development; in mammals, LIN28 has two structural and functional homologs, LIN28A and LIN28B, and is highly expressed in various tumors, promoting cellular reprogramming and thus maintaining stem cell self-renewal and pluripotency [45–47]. Analysis of the TCGA BRCA cohort data revealed a significant positive correlation between LINC00467 and LIN28B, while silencing LINC00467 significantly inhibited LIN28B expression, suggesting that LINC00467 may affect the stemness of breast cancer through LIN28B.
Due to the lack of direct empirical evidence. Computational screening via the RNAInter database revealed that LINC00467 potentially interacts with multiple let-7 family miRNAs, including hsa-let-7b-5p, hsa-let-7e-5p, hsa-let-7g-5p, and hsa-let-7i-5p [24]. This is highly relevant as the let-7 family members are established negative regulators of LIN28B, a key driver of cellular stemness [31]. Thus, our bioinformatic finding suggests a potential mechanism where LINC00467 may function as a molecular sponge for let-7 miRNAs, thereby attenuating their suppressive effect on LIN28B. The precise molecular mechanism underlying the regulation between LINC00467 and the AKT pathway has yet to be experimentally validated. The direct interaction between LINC00467 and the tumor suppressor miR-138-5p was confirmed by luciferase reporter and biotin RNA pull-down assays, demonstrating that LINC00467 acts as a miRNA sponge to repress miR-138-5p expression. RIP assays further revealed that LINC00467 directly binds to lin-28 homolog B (LIN28B), a key oncogene in breast cancer, leading to an increase in LIN28B protein levels [48]. However, the precise mechanism by which this interaction enhances LIN28B protein expression requires further investigation. The potential association between LINC00467 and resistance to anti-PD-1 therapy remains an area of observational research. Conducting mechanistic explorations using co-culture models or multiplex immunohistochemistry techniques is of paramount importance. Supporting its clinical relevance, one study analyzing LINC00467 expression in 113 normal breast tissues and 1091 breast cancer specimens reported significant overexpression in tumor tissues compared to normal controls [49]. Consistent with this, qRT-PCR analysis of 70 collected breast cancer samples and matched adjacent tissues demonstrated remarkable upregulation of LINC00467 in cancerous samples. Further analysis indicated that LINC00467 expression was positively correlated with tumor stage (p = 0.0164) and lymph node metastasis (p = 0.0248) in these patients [49]. Addressing these aspects will be a primary focus of subsequent research to fully delineate the functional and clinical significance of LINC00467 in breast cancer.
Several limitations should be noted in this study. Spatial transcriptomic analysis was performed on only a single patient sample, limiting the generalizability of LINC00467 expression patterns across breast cancer contexts. The observed inverse correlation between LINC00467 expression and AKT inhibitor sensitivity relies solely on computational analyses of public datasets and requires functional validation using patient-derived organoids or isogenic cell lines. The precise mechanism through which LINC00467 regulates AKT signaling remains unclear. Furthermore, this study lacks Western blot analysis of stemness markers (e.g., ALDH, CD44, CD24) in knockout vs. control cells and requires further validation via rescue experiments and RNA-protein interaction studies. The proposed role of LINC00467 in cancer stemness also needs confirmation via stemness marker detection or in vivo xenograft models. Furthermore, its association with anti-PD-1 therapy resistance remains speculative and requires validation through co-culture systems or multiplex immunohistochemistry. Addressing these aspects will be a focus of subsequent analyses to fully elucidate the functional and clinical significance of LINC00467 in breast cancer.
Our study indicated that elevated LINC00467 expression is linked to cancer stemness traits in breast cancer. Functionally, LINC00467 knockdown was shown to impair clonogenicity and attenuate AKT signaling pathway activation. Taken together, LINC00467 represents a potential predictor of drug response in breast cancer.
Acknowledgement: Not applicable.
Funding Statement: This work was supported by grants from the Natural Science Foundation of Hunan Province (2023JJ30879, 2026JJ80914).
Author Contributions: The authors confirm contribution to the paper as follows: Conceptualization, Junbin Yuan, Zailong Qin and Jie Guo; investigation, Hui Zha, Chao Li and Jia Chen; formal analysis, Hao Bo and Jie Guo; writing—original draft preparation, Hui Zha, Chao Li and Jia Chen; writing—review and editing, Hui Zha, Zhaolan Hu. All authors reviewed and approved the final version of the manuscript.
Availability of Data and Materials: The data that support the findings of this study are available from the Corresponding Author, Junbin Yuan, upon reasonable request.
Ethics Approval: Not applicable.
Conflicts of Interest: The authors declare no conflicts of interest
Appendix A
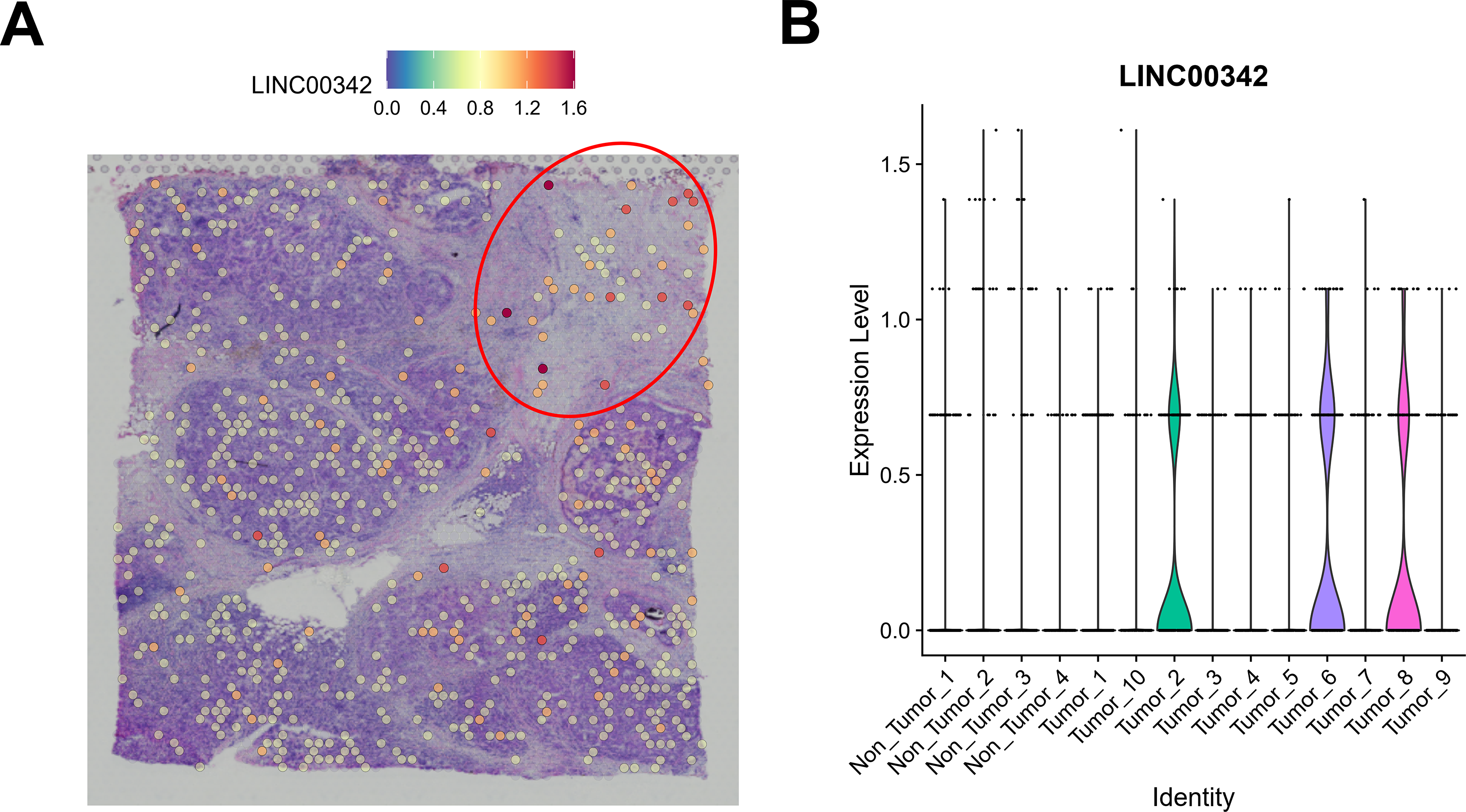
Figure A1: Spatial expression characteristics of LINC00342. (A) Spatial expression of LINC00342 in breast cancer samples, showing lower expression positivity compared with LINC00467. Red circles indicate that LINC00342 is also expressed in non-tumor regions. (B) Violin plot showing the distribution of LINC00342 expression across the different subgroups.
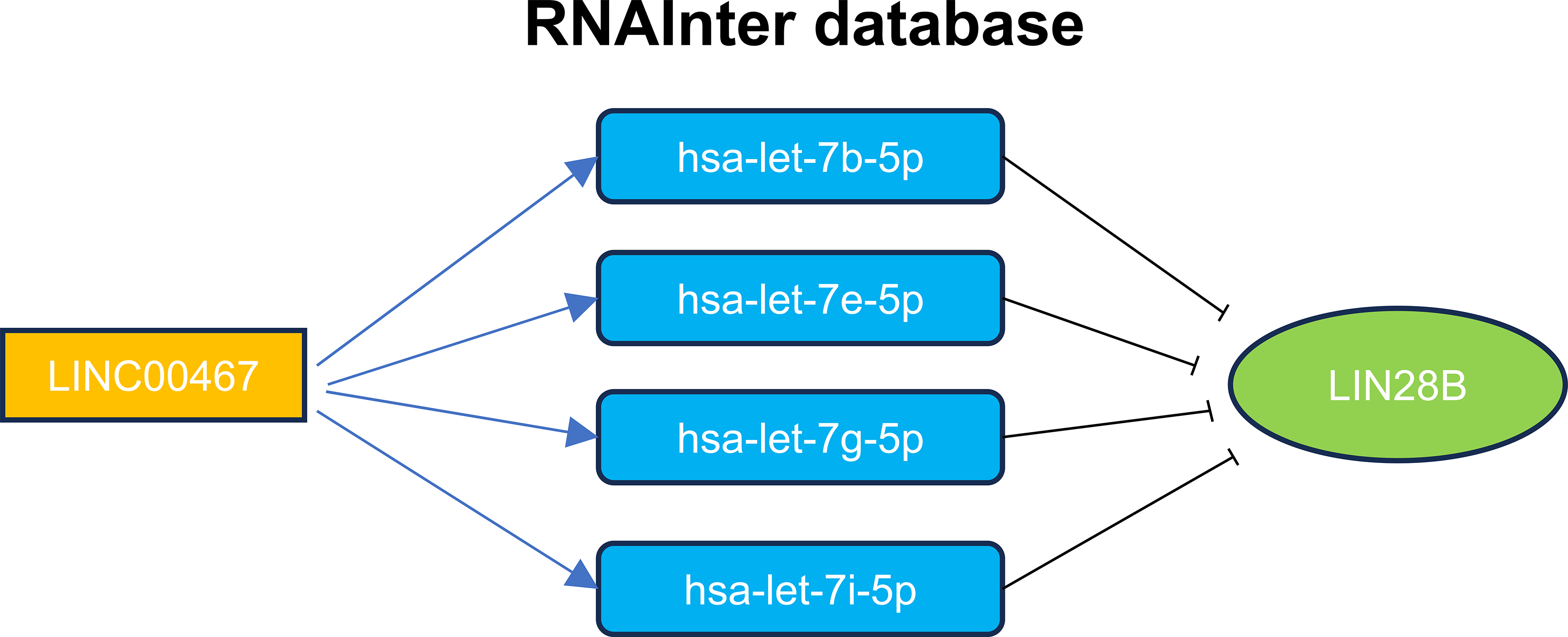
Figure A2: This model proposes that LINC00467 functions as a molecular sponge, sequestering let-7 family miRNAs and thereby potentially upregulating LIN28B to promote cellular stemness.
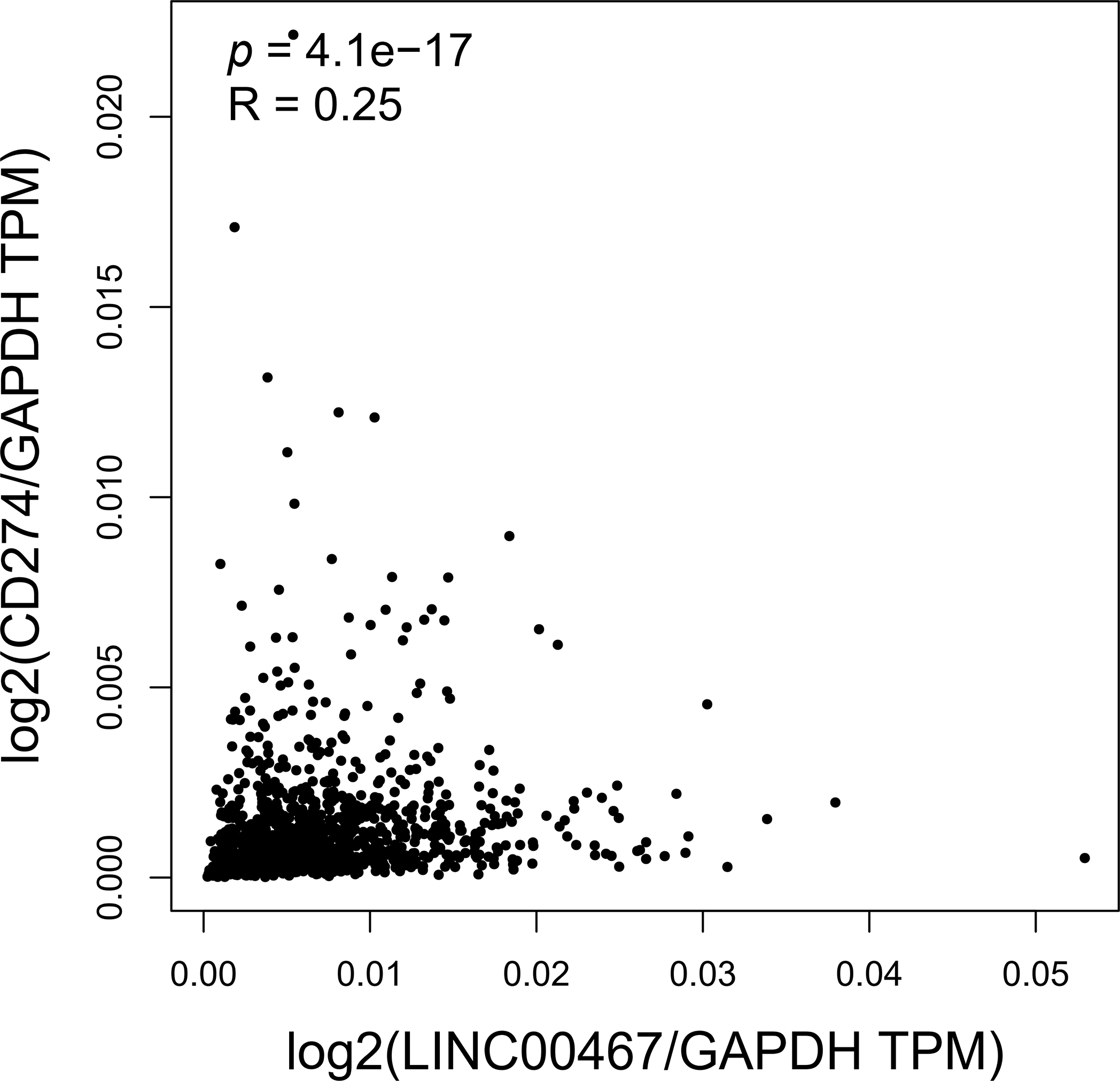
Figure A3: Analysis of the BRCA dataset from the GEPIA2 database reveals a significant positive correlation between LINC00467 and CD274 (PD-L1) expression.
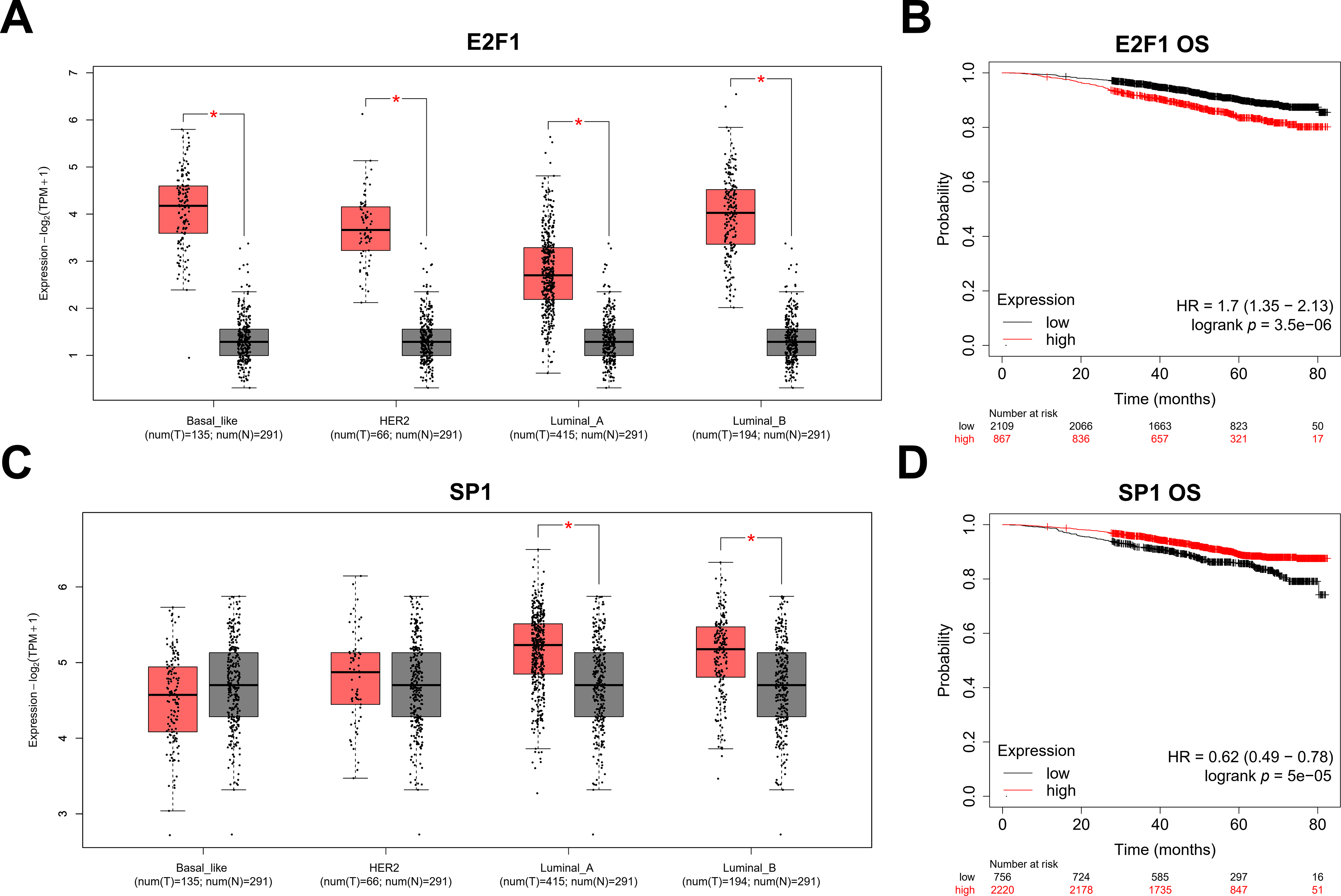
Figure A4: Expression of transcription factors E2F1 and SP1 in breast cancer and their correlation with prognosis. (A) E2F1 expression was significantly upregulated in Luminal A, Luminal B, Basal-like, and HER2 subtypes. (B) Kaplan–Meier survival analysis based on TCGA BRCA data, stratified by median E2F1 expression, showed that high E2F1 expression was associated with reduced overall survival. (C) SP1 expression was significantly upregulated in Luminal A and Luminal B breast cancer tissues compared to normal tissues. In contrast, there was no significant difference in expression in Basal-like and HER2 subtypes. (D) The samples were divided into two groups based on the median level of SP1 expression, and in the group with high SP1 expression, the survival rate was higher. *p < 0.05.
References
1. Sarhangi N, Hajjari S, Heydari SF, Ganjizadeh M, Rouhollah F, Hasanzad M. Breast cancer in the era of precision medicine. Mol Biol Rep. 2022;49(10):10023–37. doi:10.1007/s11033-022-07571-2. [Google Scholar] [PubMed] [CrossRef]
2. Kawiak A. Molecular research and treatment of breast cancer. Int J Mol Sci. 2022;23(17):9617. doi:10.3390/ijms23179617. [Google Scholar] [PubMed] [CrossRef]
3. Zhang L, Chen W, Liu S, Chen C. Targeting breast cancer stem cells. Int J Biol Sci. 2023;19(2):552–70. doi:10.7150/ijbs.76187. [Google Scholar] [PubMed] [CrossRef]
4. Giuliani B, Tordonato C, Nicassio F. Mechanisms of long non-coding RNA in breast cancer. Int J Mol Sci. 2023;24(5):4538. doi:10.3390/ijms24054538. [Google Scholar] [PubMed] [CrossRef]
5. Liu F, Tian T, Zhang Z, Xie S, Yang J, Zhu L, et al. Long non-coding RNA SNHG6 couples cholesterol sensing with mTORC1 activation in hepatocellular carcinoma. Nat Metab. 2022;4(8):1022–40. doi:10.1038/s42255-022-00616-7. [Google Scholar] [PubMed] [CrossRef]
6. Luo Y, Tang W, Xiang S, Feng J, Zu X. Non-coding RNAs in breast cancer: implications for programmed cell death. Cancer Lett. 2022;550:215929. doi:10.1016/j.canlet.2022.215929. [Google Scholar] [PubMed] [CrossRef]
7. Tang Y, Tian W, Zheng S, Zou Y, Xie J, Zhang J, et al. Dissection of FOXO1-induced LYPLAL1-DT impeding triple-negative breast cancer progression via mediating hnRNPK/β-catenin complex. Research. 2023;6:0289. doi:10.34133/research.0289. [Google Scholar] [PubMed] [CrossRef]
8. Ghasemian A, Omear HA, Mansoori Y, Mansouri P, Deng X, Darbeheshti F, et al. Long non-coding RNAs and JAK/STAT signaling pathway regulation in colorectal cancer development. Front Genet. 2023;14:1297093. doi:10.3389/fgene.2023.1297093. [Google Scholar] [PubMed] [CrossRef]
9. Liu Y, Liu Y, Ye S, Feng H, Ma L. A new ferroptosis-related signature model including messenger RNAs and long non-coding RNAs predicts the prognosis of gastric cancer patients. J Transl Int Med. 2023;11(2):145–55. doi:10.2478/jtim-2023-0089. [Google Scholar] [PubMed] [CrossRef]
10. Xiong XD, Ren X, Cai MY, Yang JW, Liu X, Yang JM. Long non-coding RNAs: an emerging powerhouse in the battle between life and death of tumor cells. Drug Resist Updat. 2016;26:28–42. doi:10.1016/j.drup.2016.04.001. [Google Scholar] [PubMed] [CrossRef]
11. Yu Z, Tang H, Chen S, Xie Y, Shi L, Xia S, et al. Exosomal LOC85009 inhibits docetaxel resistance in lung adenocarcinoma through regulating ATG5-induced autophagy. Drug Resist Updat. 2023;67:100915. doi:10.1016/j.drup.2022.100915. [Google Scholar] [PubMed] [CrossRef]
12. Chen X, Luo Q, Xiao Y, Zhu J, Zhang Y, Ding J, et al. LINC00467: an oncogenic long noncoding RNA. Cancer Cell Int. 2022;22(1):303. doi:10.1186/s12935-022-02733-5. [Google Scholar] [PubMed] [CrossRef]
13. Gao Y, Li J, Ma M, Fu W, Ma L, Sui Y, et al. Prognostic prediction of m6A and ferroptosis-associated lncRNAs in liver hepatocellular carcinoma. J Transl Int Med. 2024;12(5):526–9. doi:10.1515/jtim-2024-0023. [Google Scholar] [PubMed] [CrossRef]
14. Bo H, Zhang W, Zhong X, Chen J, Liu Y, Cheong KL, et al. LINC00467, driven by copy number amplification and DNA demethylation, is associated with oxidative lipid metabolism and immune infiltration in breast cancer. Oxid Med Cell Longev. 2021;2021:4586319. doi:10.1155/2021/4586319. [Google Scholar] [PubMed] [CrossRef]
15. Yang F, Xiao Y, Ding JH, Jin X, Ma D, Li DQ, et al. Ferroptosis heterogeneity in triple-negative breast cancer reveals an innovative immunotherapy combination strategy. Cell Metab. 2023;35(1):84–100.e8. doi:10.1016/j.cmet.2022.09.021. [Google Scholar] [PubMed] [CrossRef]
16. Liang Y, Zhang H, Song X, Yang Q. Metastatic heterogeneity of breast cancer: molecular mechanism and potential therapeutic targets. Semin Cancer Biol. 2020;60:14–27. doi:10.1016/j.semcancer.2019.08.012. [Google Scholar] [PubMed] [CrossRef]
17. Martin FJ, Amode MR, Aneja A, Austine-Orimoloye O, Azov AG, Barnes I, et al. Ensembl 2023. Nucleic Acids Res. 2023;51(D1):D933–41. doi:10.1093/nar/gkac958. [Google Scholar] [PubMed] [CrossRef]
18. Győrffy B. Survival analysis across the entire transcriptome identifies biomarkers with the highest prognostic power in breast cancer. Comput Struct Biotechnol J. 2021;19:4101–9. doi:10.1016/j.csbj.2021.07.014. [Google Scholar] [PubMed] [CrossRef]
19. Tang Z, Kang B, Li C, Chen T, Zhang Z. GEPIA2: an enhanced web server for large-scale expression profiling and interactive analysis. Nucleic Acids Res. 2019;47(W1):W556–60. doi:10.1093/nar/gkz430. [Google Scholar] [PubMed] [CrossRef]
20. Sabatier R, Finetti P, Cervera N, Lambaudie E, Esterni B, Mamessier E, et al. A gene expression signature identifies two prognostic subgroups of basal breast cancer. Breast Cancer Res Treat. 2011;126(2):407–20. doi:10.1007/s10549-010-0897-9. [Google Scholar] [PubMed] [CrossRef]
21. Chanrion M, Negre V, Fontaine H, Salvetat N, Bibeau F, Mac Grogan G, et al. A gene expression signature that can predict the recurrence of tamoxifen-treated primary breast cancer. Clin Cancer Res. 2008;14(6):1744–52. doi:10.1158/1078-0432.CCR-07-1833. [Google Scholar] [PubMed] [CrossRef]
22. Foster DS, Januszyk M, Delitto D, Yost KE, Griffin M, Guo J, et al. Multiomic analysis reveals conservation of cancer-associated fibroblast phenotypes across species and tissue of origin. Cancer Cell. 2022;40(11):1392–406.e7. doi:10.1016/j.ccell.2022.09.015. [Google Scholar] [PubMed] [CrossRef]
23. Liu CJ, Hu FF, Xia MX, Han L, Zhang Q, Guo AY. GSCALite: a web server for gene set cancer analysis. Bioinformatics. 2018;34(21):3771–2. doi:10.1093/bioinformatics/bty411. [Google Scholar] [PubMed] [CrossRef]
24. Kang J, Tang Q, He J, Li L, Yang N, Yu S, et al. RNAInter v4.0: RNA interactome repository with redefined confidence scoring system and improved accessibility. Nucleic Acids Res. 2022;50(D1):D326–32. doi:10.1093/nar/gkab997. [Google Scholar] [PubMed] [CrossRef]
25. Shen WK, Chen SY, Gan ZQ, Zhang YZ, Yue T, Chen MM, et al. AnimalTFDB 4.0: a comprehensive animal transcription factor database updated with variation and expression annotations. Nucleic Acids Res. 2023;51(D1):D39–45. doi:10.1093/nar/gkac907. [Google Scholar] [PubMed] [CrossRef]
26. Liu ZP, Wu C, Miao H, Wu H. RegNetwork: an integrated database of transcriptional and post-transcriptional regulatory networks in human and mouse. Database. 2015;2015:bav095. doi:10.1093/database/bav095. [Google Scholar] [PubMed] [CrossRef]
27. Messeguer X, Escudero R, Farré D, Núñez O, Martínez J, Albà MM. PROMO: detection of known transcription regulatory elements using species-tailored searches. Bioinformatics. 2002;18(2):333–4. doi:10.1093/bioinformatics/18.2.333. [Google Scholar] [PubMed] [CrossRef]
28. Perez G, Barber GP, Benet-Pages A, Casper J, Clawson H, Diekhans M, et al. The UCSC genome browser database: 2025 update. Nucleic Acids Res. 2025;53(D1):D1243–9. doi:10.1093/nar/gkae974. [Google Scholar] [PubMed] [CrossRef]
29. Luo Y, Wang X, Niu W, Zhou Y, Li M, Ma J, et al. BRD7 stabilizes P53 via dephosphorylation of MDM2 to inhibit tumor growth in breast cancer harboring wild-type P53. J Cancer. 2022;13(5):1436–48. doi:10.7150/jca.67447. [Google Scholar] [PubMed] [CrossRef]
30. Xu Z, Wang W, Yang T, Chen J, Huang Y, Gould J, et al. STOmicsDB: a database of spatial transcriptomic data. BioRxiv. 2022. doi:10.1101/2022.03.11.481421. [Google Scholar] [CrossRef]
31. Viswanathan SR, Daley GQ, Gregory RI. Selective blockade of microRNA processing by Lin28. Science. 2008;320(5872):97–100. doi:10.1126/science.1154040. [Google Scholar] [PubMed] [CrossRef]
32. Baek M, Chang JT, Echeverria GV. Methodological advancements for investigating intra-tumoral heterogeneity in breast cancer at the bench and bedside. J Mammary Gland Biol Neoplasia. 2020;25(4):289–304. doi:10.1007/s10911-020-09470-3. [Google Scholar] [PubMed] [CrossRef]
33. Peng J, Sun BF, Chen CY, Zhou JY, Chen YS, Chen H, et al. Single-cell RNA-seq highlights intra-tumoral heterogeneity and malignant progression in pancreatic ductal adenocarcinoma. Cell Res. 2019;29(9):725–38. doi:10.1038/s41422-019-0195-y. [Google Scholar] [PubMed] [CrossRef]
34. Yan X, Xie Y, Yang F, Hua Y, Zeng T, Sun C, et al. Comprehensive description of the current breast cancer microenvironment advancements via single-cell analysis. J Exp Clin Cancer Res. 2021;40(1):142. doi:10.1186/s13046-021-01949-z. [Google Scholar] [PubMed] [CrossRef]
35. Puram SV, Tirosh I, Parikh AS, Patel AP, Yizhak K, Gillespie S, et al. Single-cell transcriptomic analysis of primary and metastatic tumor ecosystems in head and neck cancer. Cell. 2017;171(7):1611–24.e24. doi:10.1016/j.cell.2017.10.044. [Google Scholar] [PubMed] [CrossRef]
36. Ho DW, Tsui YM, Chan LK, Sze KM, Zhang X, Cheu JW, et al. Single-cell RNA sequencing shows the immunosuppressive landscape and tumor heterogeneity of HBV-associated hepatocellular carcinoma. Nat Commun. 2021;12(1):3684. doi:10.1038/s41467-021-24010-1. [Google Scholar] [PubMed] [CrossRef]
37. Li R, Ferdinand JR, Loudon KW, Bowyer GS, Laidlaw S, Muyas F, et al. Mapping single-cell transcriptomes in the intra-tumoral and associated territories of kidney cancer. Cancer Cell. 2022;40(12):1583–99.e10. doi:10.1016/j.ccell.2022.11.001. [Google Scholar] [PubMed] [CrossRef]
38. Zou Y, Ye F, Kong Y, Hu X, Deng X, Xie J, et al. The single-cell landscape of intratumoral heterogeneity and the immunosuppressive microenvironment in liver and brain metastases of breast cancer. Adv Sci. 2023;10(5):2203699. doi:10.1002/advs.202203699. [Google Scholar] [PubMed] [CrossRef]
39. Lv J, Shi Q, Han Y, Li W, Liu H, Zhang J, et al. Spatial transcriptomics reveals gene expression characteristics in invasive micropapillary carcinoma of the breast. Cell Death Dis. 2021;12(12):1095. doi:10.1038/s41419-021-04380-6. [Google Scholar] [PubMed] [CrossRef]
40. Mandair D, Reis-Filho JS, Ashworth A. Biological insights and novel biomarker discovery through deep learning approaches in breast cancer histopathology. npj Breast Cancer. 2023;9(1):21. doi:10.1038/s41523-023-00518-1. [Google Scholar] [PubMed] [CrossRef]
41. Vasan N, Cantley LC. At a crossroads: how to translate the roles of PI3K in oncogenic and metabolic signalling into improvements in cancer therapy. Nat Rev Clin Oncol. 2022;19(7):471–85. doi:10.1038/s41571-022-00633-1. [Google Scholar] [PubMed] [CrossRef]
42. Eustace AJ, Lee MJ, Colley G, Roban J, Downing T, Buchanan PJ. Aberrant calcium signalling downstream of mutations in TP53 and the PI3K/AKT pathway genes promotes disease progression and therapy resistance in triple negative breast cancer. Cancer Drug Resist. 2022;5(3):560–76. doi:10.20517/cdr.2022.41. [Google Scholar] [PubMed] [CrossRef]
43. Sideris N, Dama P, Bayraktar S, Stiff T, Castellano L. LncRNAs in breast cancer: a link to future approaches. Cancer Gene Ther. 2022;29(12):1866–77. doi:10.1038/s41417-022-00487-w. [Google Scholar] [PubMed] [CrossRef]
44. Arriaga-Canon C, Contreras-Espinosa L, Aguilar-Villanueva S, Bargalló-Rocha E, García-Gordillo JA, Cabrera-Galeana P, et al. The clinical utility of lncRNAs and their application as molecular biomarkers in breast cancer. Int J Mol Sci. 2023;24(8):7426. doi:10.3390/ijms24087426. [Google Scholar] [PubMed] [CrossRef]
45. Zou H, Luo J, Guo Y, Liu Y, Wang Y, Deng L, et al. RNA-binding protein complex LIN28/MSI2 enhances cancer stem cell-like properties by modulating Hippo-YAP1 signaling and independently of Let-7. Oncogene. 2022;41(11):1657–72. doi:10.1038/s41388-022-02198-w. [Google Scholar] [PubMed] [CrossRef]
46. Sa-Nguanraksa D, Pongthong W, Samarnthai N, Mitpakdi K, Chuangsuwanich T, Limjindaporn T, et al. Expression of androgen receptor and its regulatory molecule Lin28 in non-luminal subtype breast cancer. Mol Clin Oncol. 2020;12(6):511–8. doi:10.3892/mco.2020.2029. [Google Scholar] [PubMed] [CrossRef]
47. Han R, Yang H, Ling C, Lu L. Neurospora crassa is a potential source of anti-cancer agents against breast cancer. Breast Cancer. 2022;29(6):1032–41. doi:10.1007/s12282-022-01383-9. [Google Scholar] [PubMed] [CrossRef]
48. Changizian M, Nourisanami F, Hajpoor V, Parvaresh M, Bahri Z, Motovali-Bashi M. LINC00467: a key oncogenic long non-coding RNA. Clin Chim Acta. 2022;536:112–25. doi:10.1016/j.cca.2022.09.013. [Google Scholar] [PubMed] [CrossRef]
49. Zhang Y, Sun Y, Ding L, Shi W, Ding K, Zhu Y. Long non-coding RNA LINC00467 correlates to poor prognosis and aggressiveness of breast cancer. Front Oncol. 2021;11:643394. doi:10.3389/fonc.2021.643394. [Google Scholar] [PubMed] [CrossRef]
Cite This Article
 Copyright © 2026 The Author(s). Published by Tech Science Press.
Copyright © 2026 The Author(s). Published by Tech Science Press.This work is licensed under a Creative Commons Attribution 4.0 International License , which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.


 Submit a Paper
Submit a Paper Propose a Special lssue
Propose a Special lssue View Full Text
View Full Text Download PDF
Download PDF Downloads
Downloads
 Citation Tools
Citation Tools