 Open Access
Open Access
REVIEW
Halotolerant PGPR at the Soil–Plant–Microbiome Interface: Microbial Strategies to Enhance Crop Adaptation to Salinity
1 College of Agriculture and Bioengineering, Taizhou Vocational College of Science & Technology, Taizhou, China
2 Institute of Computing Technology, Chinese Academy of Sciences, Beijing, China
* Corresponding Authors: Shuqin Gao. Email: ; Congcong Zheng. Email:
Phyton-International Journal of Experimental Botany 2026, 95(3), 3 https://doi.org/10.32604/phyton.2026.076579
Received 23 November 2025; Accepted 05 February 2026; Issue published 31 March 2026
Abstract
Global climate change has intensified drought and soil salinization, posing serious threats to crop productivity and ecosystem stability. Traditional physical and chemical reclamation methods are often expensive, energy-intensive, and unsustainable. In contrast, halotolerant plant growth-promoting rhizobacteria (HT-PGPR) have emerged as a promising, eco-friendly strategy to address extreme climate change-induced land salinization. HT-PGPR enhance plant tolerance by regulating osmotic balance, ion homeostasis, antioxidant defense, and phytohormone signaling. Current evidence for these effects is largely based on greenhouse pot and microcosm studies, while their validation in field experiments remain limited. In addition, HT-PGPR can improve nutrient availability through nitrogen fixation, phosphorus and potassium solubilization, and siderophore production. This review synthesizes recent advances in the physiological and molecular mechanisms underlying HT-PGPR-mediated stress alleviation, evaluates their performance in well-controlled greenhouse experiments and field applications, and outlines key criteria for strain selection. However, the translation of controlled-environment benefits into consistent field performance remains challenging, due to factors including inoculated microbial survival under fluctuating edaphic conditions, competitive interactions with native soil microbiota, and spatiotemporal variability in climate. Furthermore, we explore emerging innovative approaches such as AI-assisted design of synthetic microbial community for the remediation of saline-alkaline soils. Integrating HT-PGPR into sustainable management practices holds substantial potential for improving crop adaptation and productivity in saline environments under a changing climate.Keywords
Soil salinization poses a serious global threat to agricultural sustainability. Recent studies report that salinization affects more than 1 billion hectares of land worldwide, with approximately 76 million hectares attributed to human activities [1,2]. According to the Food and Agriculture Organization (FAO), approximately 20% of irrigated land globally is currently affected by salinity, with salt-affected soils posing a growing threat to global agricultural productivity [3]. The total area of salt-affected land continues to expand globally, driven by poor irrigation practices and climate change, based on FAO global assessments and peer-reviewed analyses [2,4,5]. In China, saline-alkali soils cover more than 90 million hectares, primarily distributed across the northeastern, northwestern, northern, and coastal regions [6]. With the increasing severity of extreme climate events, soil salinization is expected to intensify, underscoring the urgent need for cost-effective, environmentally friendly, and sustainable reclamation strategies.
1.2 Limitations of Conventional Reclamation Methods
Conventional approaches for saline soil reclamation, including physical leaching and chemical amendments such as gypsum, can temporarily lower soil salinity but suffer from several major limitations [7]. First, their high water demand make them impractical in arid and semi-arid regions where freshwater resources are already scarce [4]. In addition, repeated application of chemical amendments may disturb soil nutrient balance and health, and over time induce soil compaction, structural degradation, and a decline in microbial activity [8]. These methods are also economically unsustainable due to the substantial input costs associated with large-scale implementation [9]. Moreover, conventional practices mainly offer short-term relief without addressing the underlying drivers of salinization, often resulting in salt re-accumulation after reclamation [10]. Critically, such approaches neglect the restoration of soil ecological integrity, failing to recover native biodiversity, functional microbial communities, and essential ecosystem processes [11].
In contrast, biological approaches, particularly those based on halotolerant plant growth-promoting rhizobacteria (HT-PGPR), offer a more sustainable and cost-effective alternative. HT-PGPR improve soil health and plant productivity by modulating microbial community composition and function in the soil [12,13]. Therefore, future reclamation efforts should increasingly emphasize microbial and biological restoration techniques to achieve both ecosystem recovery and sustainable agricultural development.
1.3 Microbial Approaches for Saline Soil Rehabilitation: The Role of HT-PGPR
Microorganisms, especially HT-PGPR, have demonstrated substantial potential in mitigating salt stress and enhancing plant performance in saline environments. HT-PGPR promote plant growth through multiple, well-documented mechanisms, including the synthesis of osmoprotectants, 1-aminocyclopropane-1-carboxylate (ACC) deaminase activity and phytohormone production [7]. As a biological approach, microbial inoculation offers several advantages, such as low cost, high efficiency, and strong environmental compatibility, making it highly aligned with the principles of sustainable agriculture [9]. Beyond direct plant physiological modulation, HT-PGPR can restructure rhizosphere microbial communities, stimulate root system development, and enhance overall plant adaptability to saline stress [14]. These microorganisms also contribute to soil nutrient availability through biological nitrogen fixation, phosphate and potassium solubilization, and siderophore production, thereby supporting nutrient acquisition under salinized conditions [7]. Insights into the metabolic pathways and molecular mechanisms underpinning HT-PGPR function under salt stress provide a mechanistic foundation for developing new bio-mediated reclamation technologies [10]. Research on plant-microbe interactions further elucidate how HT-PGPR influence root morphology, ion homeostasis, and stress-related physiological processes to improve plant salt tolerance [14]. The application of such microbial inoculants not only but also enhances crop yield and quality, delivering significant agronomic and ecological benefits [9].
In summary, microorganisms represent a powerful and ecologically sound tool for mitigating salt stress and rehabilitating saline soils. However, the potential of microbial-based approaches remains far from fully realized, and thus future understanding of the synergistic interactions between HT-PGPR and other beneficial microbes, and advancing the development of high-efficiency synthetic microbial consortia for soil restoration and sustainable agricultural production are necessary [7]. A deeper understanding of microbial metabolic and molecular mechanisms under salt stress will further accelerate the design of next-generation bioremediation strategies. Accordingly, this review summarizes the physiological, biochemical, and microbial mechanisms through which HT-PGPR enhance plant salt tolerance, evaluates their practical applications in saline soil restoration, and highlights critical research gaps that must be addressed to advance their use in sustainable agriculture.
2 Mechanisms of Plant Salt Tolerance Mediated by HT-PGPR
2.1 Physiological and Biochemical Mechanisms
HT-PGPR enhance plant salt tolerance by regulating osmolyte biosynthesis pathways in host plants. Proline, one of the most prominent osmoprotectants, shows a context-dependent regulatory pattern shaped by specific plant-microbe interactions. Inoculation with ACC deaminase-producing strains such as Bacillus megaterium UPMR2 or Enterobacter sp. UPMR18 upregulates proline biosynthetic genes and enhances nutrient uptake, resulting in elevated proline accumulation in salt-stressed okra seedlings [15]. In contrast, inoculation of tomato with Bacillus megaterium CJLC2 under 200 mM NaCl stress reduced leaf proline content by 21-33% compared to stressed controls in a greenhouse pot experiment [16].
Other osmolytes, such as betaines and trehalose, also play vital roles in osmotic regulation (Fig. 1). Co-inoculation with Pseudomonas pseudoalcaligenes and Bacillus pumilus markedly increases betaine-type quaternary ammonium compounds in rice leaves, helping stabilize photosystem II proteins and maintain oxygen evolution in a greenhouse experiment under 43 mM NaCl conditions [15,17]. Trehalose functions as a molecular stabilizer that protects cellular proteins by replacing water molecules and preventing denaturation under osmotic and thermal stress [18]. Notably, many HT-PGPR can synthesize osmolytes more rapidly and efficiently than their plant hosts. For example, strain Bacillus zhangzhouensis increases soluble sugar concentrations in maize leaves by 109%, a response consistent with the presence of six proline-related biosynthetic genes identified in its genome [19]. Together, these compatible solutes form a “molecular shield” that preserves cellular osmotic balance, maintains macromolecular stability, and supports enzymatic function under saline conditions. It is important to recognize, however, that osmolyte-mediated responses vary substantially among different plant-microbe combinations, reflecting both microbial genetic diversity and species-specific plant salt tolerance strategies.
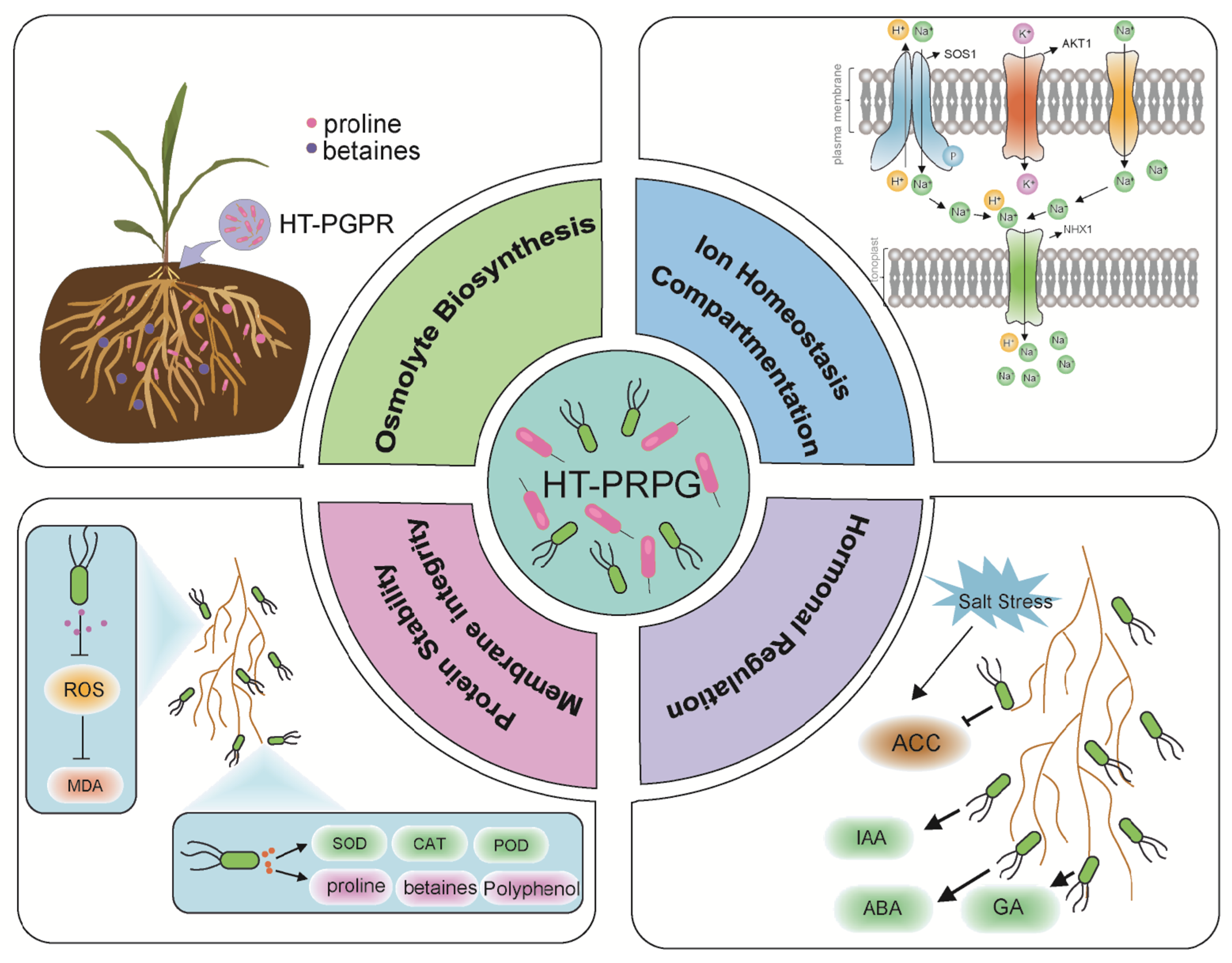
Figure 1: Biochemical and physiological responses of plant to salt stress mediated by halotolerant plant growth-promoting bacteria (HT-PGPB). HT-PGPB enhance plant tolerance by promoting osmolyte accumulation, maintaining ion homeostasis through selective uptake and exclusion, stabilizing cellular membranes and proteins, and modulating key phytohormone signaling pathways.
2.1.2 Ion Homeostasis and Compartmentation
Laboratory and greenhouse studies indicate that that HT-PGPR can contribute to the reestablishment of ion homeostasis in plants, often associated with the modulation of host plant genes encoding Na+/H+ antiporters such as SOS1 and NHX1 (Fig. 1) [5,20]. At the transcriptional level, microbes influence these transport processes. For example, at 200 mM NaCl level, inoculation of Kocuria rhizophila upregulates maize ZmHKT1 and ZmNHX1/2/3 gene expression in a greenhouse pot experiment, thereby enhancing K+ uptake and Na+ exclusion [21]. Volatile organic compounds (VOCs) emitted by Bacillus amyloliquefaciens FZB42 induce NHX1 and HKT1 expression in Arabidopsis, resulting in reduced Na+ accumulation [22]. Collectively, microbial treatments have been shown to be associated with the upregulation of ion transporter genes (e.g., HKT1, NHX1) in model systems, which correlates with improved ion balance [5,21,22]. It is important to note that such upregulation may entail metabolic costs for the plant, as energy and resources are diverted to transporter protein synthesis and ion pumping. Moreover, exopolysaccharides (EPS) secreted by PGPR can bind Na+ ions in the rhizosphere, acting as an “ion trap” to restrict Na+ translocation into roots [23].
These mechanisms maintain a low cytosolic Na+/K+ ratio, preserving enzyme function and membrane stability under salinity stress. Species-specific adaptations are evident: halophytes like Salicornia bigelovii enhance vacuolar Na+ sequestration with increasing salinity, as demonstrated by comparing plants grown in a greenhouse under 200 mM NaCl versus 5 mM NaCl conditions [24], whereas glycophytes rely more heavily on HKT1-mediated Na+ recirculation and exclusion from shoots [25]. Understanding the specificity of microbial regulation in plant ion transport networks will be essential for designing precision bioinoculant strategies.
2.1.3 Membrane Integrity and Protein Stability
HT-PGPR also protect plant cells from osmotic and oxidative damage by stabilizing cellular membranes and proteins. Salt stress disrupts reactive oxygen species (ROS) homeostasis, leading to lipid peroxidation and malondialdehyde (MDA) accumulation [15]. Inoculation with Pantoea dispersa or Enterococcus faecium significantly reduced MDA levels in mung bean leaves subjected to salt stress (50 and 100 mM NaCl) in a greenhouse trail, while meta-analyses show PGPR treatments reduce MDA accumulation by 23–24% across tolerant and sensitive plants [26]. The protection arises from coordinated activation of enzymatic (SOD, POD, CAT) and non-enzymatic (proline, betaine, polyphenols) antioxidant systems. For example, Dietzia natronolimnaea STR1, under saline stress conditions (e.g., 100 mM NaCl in hydroponic culture or 150 mM NaCl in soil pot experiments), enhanced the activities and transcript levels of SOD, POD, and CAT in wheat leaves [27]. Similarly, Bacillus cereus Pb25 promoted proline accumulation in mung bean, whereas Pseudomonas pseudoalcaligenes produced substantial quantities of glycine-betaine in rice seedlings [17,28].
Beyond antioxidant defenses, microbial inoculation also modulates aquaporins and ion transporter expression to maintain cellular homeostasis. In a potted experiment under 100 mM NaCl salt stress, Bacillus spp. upregulates aquaporin genes LePIP1, LeAQP2, and LeTRAMP in tomato leaves, enhancing water transport while enhancing LeNHX1 and tomPRO2, which contributed to reduced ion leakage and improved salt tolerance [16]. Genome analysis of Bacillus velezensis and Bacillus zhangzhouensis strains reveal catalase and Na+ efflux genes consistent with these physiological effects [19]. Collectively, these responses constitute a microbial-induced “molecular chaperone” network maintaining redox balance, membrane fluidity, and protein conformation, thereby enhancing plant resilience under salinity stress.
HT-PGPR modulate phytohormone signaling networks by integrating auxin, ethylene, cytokinin, and gibberellin pathways to coordinate plant responses to salt stress. Salinity typically suppresses auxin biosynthesis; however, Pseudomonas and Bacillus species can synthesize indole-3-acetic acid (IAA) via the tryptophan-dependent pathway, enhancing root surface area and promoting lateral root formation in soybean [29]. Similarly, strain Bacillus zhangzhouensis synthesized IAA, promoting maize root growth and raising the K+/Na+ ratio to 1.98 under salt stress [16]. Notably, the effect of microbial IAA is often dose-dependent; for example, optimal root promotion in Arabidopsis was observed at specific IAA concentrations produced by Pseudomonas sp., while higher concentrations could be inhibitory [29].
Microbes containing ACC deaminase, such as Pseudomonas putida, degrade the ethylene precursor ACC and reduce ethylene accumulation by 30-50% [18]. Bacillus megaterium CJLC2 decreases ethylene emission in tomato by 32% [14]. In addition, cytokinins secreted by Bacillus amyloliquefaciens B1619 delay leaf senescence and cooperate with gibberellins to regulate stomatal conductance and photosynthetic activity [30]. Hormonal crosstalk is highly dose- and context-dependent, e.g., low IAA promotes root hair development, whereas excessive IAA inhibits primary root elongation [31]. Deciphering the spatial and temporal architecture of microbial hormone regulation will be essential for moving from correlative observations to a mechanistic framework capable of guiding targeted manipulation of plant stress-response networks.
2.2 Molecular Pathways: Antioxidant and Signal Transduction Networks
HT-PGPR enhance plant salt tolerance by coordinating ROS homeostasis and activating multi-tiered signaling cascades. Salt stress disrupts redox equilibrium, leading to the accumulation of superoxide (O2−) and hydrogen peroxide (H2O2). In wheat, inoculation with Dietzia natronolimnaea STR1 markedly elevated the activities of antioxidant enzymes, including SOD, POD, CAT, APX, GPX, and GR [15]. In parallel, microbial VOCs trigger MAPK cascades that activate transcription factors DREB2A/B and AREB/ABF, which in turn regulate downstream antioxidant genes via antioxidant response elements and dehydration-responsive elements [18]. H2O2 serves as a “double-edged signal”: at low levels it activates Ca2+ and CDPK pathways, whereas at high levels it triggers programmed cell death. Bacillus velezensis, which carried five catalase-encoding genes, efficiently decomposes H2O2 into H2O and O2, thereby reducing maize leaf MDA content to 83% of control [19]. Additional microbial metabolites such as acetylcholine and γ-aminobutyric acid analogs further regulate stomatal movement, decreasing ROS production while maintaining photosynthetic efficiency [6].
Together, these processes form a finely tuned “redox-sensing network”, where NADPH oxidase (Rboh) amplifies ROS signaling and the ascorbate-glutathione cycle serves as a redox buffer. Strain-specific regulatory pathways further modulate plant ion homeostasis. For example, Bacillus velezensis induces the expression of plant H+-PPase, thereby enhancing the vacuolar proton gradient that facilitates Na+ sequestration [19], while Arthrobacter protophormiae synthesizes dimethylsulfoniopropionate (DMSP) to activate jasmonic acid signaling [18]. Decoding receptor-level, cross-kingdom signaling mechanisms between microbes and host plants remains a major frontier for designing precision microbial inoculants.
2.3 Nutrient Utilization and Symbiosis
2.3.1 Major Elements Availability
HT-PGPR enhance plant nutrient acquisition under salt stress via biological nitrogen fixation, phosphate solubilization, and potassium mobilization. It’s reported that in saline-alkali soils, up to 80% of soil phosphorus exists in insoluble mineral forms due to high pH; HT-PGPR release organic acids (e.g., oxalic, citric) to solubilize these forms, thereby improving plant P uptake. Pseudomonas inoculation increased fresh weight in cabbage seedlings, and Bacillus enhanced tomato P absorption by 48% [32].
Brevibacterium frigoritolerans, Bacillus velezensis, and Bacilluszhangzhouensis strains produced halos on insoluble phosphate and silicate media, increasing maize K+ content by 34% under 150 mM NaCl [19]. Nitrogen-fixing bacteria such as Rhizobium and Mesorhizobium establish symbiotic nodules with legumes, while Azospirillum lipoferum improved N accumulation and maintained nitrogenase activity under low oxygen microenvironments [15,33]. These nutrient-mobilizing processes often act synergistically; for instance, K-solubilizing bacteria also enhance P release, while N-fixing bacteria facilitate K mineralization through rhizosphere acidification. Moreover, PGPR inoculation increased the K+/Na+ ratio in tomato roots from 0.78 to 1.28 by regulating HKT1 and NHX1 expression [19]. Together, these mechanisms optimize nutrient availability and ion balance, reinforcing plant resilience in saline environments.
2.3.2 Siderophore Production and Plant Interaction
HT-PGPR produce siderophores that chelate Fe3+, enhancing plant iron acquisition under saline conditions [34]. Strains Bacillus velezensis and Bacillus zhangzhouensis synthesize catecholate-type siderophores encoded by nonribosomal peptide synthetase (NRPS) and fur-regulated gene clusters [19].
Siderophores not only suppress wheat sheath blight by 68% but also increase iron bioavailability. Arthrobacter protophormiae inoculation increased Fe content in tomato by 2.3-fold and upregulated FER and FIT expression [6,35]. Bacillus species typically produce bacillibactin-type siderophores, whose biosynthesis correlates positively with salinity levels, while Pseudomonas tends to generate pyochelin-type molecules [19]. Fe3+-siderophore complexes enter plant cells via YSL transporters, activating transcription factors FIT1 and bHLH038, which regulate IRT1 and FRO2 for Fe uptake [36]. Thus, siderophore production integrates microbial iron scavenging with host Fe homeostasis, establishing a cooperative defense-nutrition interface under salt stress.
2.3.3 Crop-Specific Applications
HT-PGPR exhibit crop-specific adaptations in enhancing salt tolerance through distinct physiological and molecular pathways. In rice (Oryza sativa), co-inoculation with Pseudomonas pseudoalcaligenes and Bacillus pumilus elevated betaine-type osmolytes and proline accumulation by 2.8-fold under 43 mM salt stress. In wheat (Triticum aestivum), Azospirillum lipoferum increased N accumulation by 37% while modulating HKT1 and AKT1 expression involved in ion homeostasis [17]. In maize (Zea mays), multiple Bacillus strains improved growth and antioxidant capacity. Bacillus tropicalis increased plant height by 45%, Fictibacillus arsenicus enhanced CAT and SOD activities, reducing MDA by 31%, and Bacillus velezensis improved both proline content and membrane stability [16]. In legumes, Rhizobium-Medicago sativa symbiosis maintained high leghaemoglobin and nitrogenase activity, while Mesorhizobium increased nodule numbers by 2.3-fold. Bacillus subtilis-Acacia interaction boosted nitrate and nitrite reductase activity by 42% [15,37].
These findings highlight that monocots (e.g., rice, wheat) primarily rely on ion regulation, legumes on symbiotic nitrogen fixation, and C4 crops like maize can achieve salt tolerance through integrated antioxidant and osmotic mechanisms [38]. Future work should focus on developing plant-microbe compatibility models to optimize HT-PGPR applications across saline agroecosystems.
3 Major Groups of Halotolerant Microorganisms
Among halotolerant microorganisms, PGPR represent the core functional group, primarily including the genera Bacillus, Pseudomonas, and Halomonas. Representative Bacillus species, B. subtilis, significantly increased 38.4% and 56.7% in glycine betaine and proline of Acacia gerrardii Benth under 200 mM NaCl stress in a greenhouse experiment [39]. Halomonas species exhibited remarkable adaptability by switching between halophilic and non-halophilic growth modes, accumulating compatible solutes such as ectoine and hydroxyectoine to maintain osmotic balance [40]. For example, H. desiderata and strain STR8 increased proline accumulation in maize leaves by 61% under 500 mM NaCl stress [41].
These bacterial taxa display functional complementarity. Bacillus species are efficient phosphate and potassium solubilizers, Pseudomonas species excel in siderophore-mediated pathogen suppression, while Halomonas specializes in the biosynthesis of organic osmolytes [42]. At the molecular level, bacterial halotolerance is linked to conserved salt-stress operons such as proU (proline transport) and bet (glycine betaine biosynthesis) [43]. Future comparative genomics studies are needed to elucidate the evolutionary trajectories of salt adaptation across genera, providing a theoretical basis for the design of multi-strain, synergistic “microbial consortia” for saline agriculture.
3.2 Fungi (Arbuscular Mycorrhizal Fungi and Endophytic Fungi)
Halotolerant fungi, primarily arbuscular mycorrhizal fungi (AMF) and endophytic fungi, can enhance plant salt tolerance through symbiotic interactions and extensive hyphal networks. AMF species such as Funneliformis mosseae, Rhizophagus intraradices, and Claroideoglomus etunicatum facilitate nutrient and water transport, improving Na+ efflux and ionic homeostasis. For instance, AMF inoculation increased Na+ extrusion efficiency by 47% and raised the K+/Na+ ratio to 2.8 in rice roots [44]. This effect is closely associated with the upregulation of aquaporin genes (PIP2;1 and TIP4;1), enabling alfalfa leaves to maintain 78% relative water content under 200 mM NaCl stress [45]. The endophytic fungus Serendipita indica mitigates salt-induced oxidative stress by activating the host antioxidant system, enhancing SOD and POD activities by 1.9-fold and 2.3-fold, respectively, while reducing MDA content by 41% in wheat leaves [46]. Synergistic AMF-PGPR interactions further enhance tolerance. Co-inoculation of F. mosseae and Methylobacterium oryzae CBMB20 increased maize root colonization by 68%, reduced proline accumulation, and elevated glycine betaine levels, suggesting a shift from a damage-repair to a pre-adaptive osmotic regulation strategy [47]. At the molecular level, AMF secrete effector-like proteins that modulate host gene expression. For example, the SiSSP1 effector interacts with the rice transcription factor OsTFIIBγ to activate OsNHX1, facilitating vacuolar Na+ sequestration [45]. These findings underscore fungi as biointerfaces coupling ionic balance, water transport, and host gene regulation under salinity stress.
3.3 Archaea (Extremely Halophilic Archaea)
Extremely Halophilic archaea, mainly belonging to the Euryarchaeota and Thaumarchaeota phyla, maintain cellular homeostasis under saturated saline conditions through unique cellular adaptations. They express high-affinity K+ transporters (e.g., KdpATPase) and produce acidic, halostable proteins (such as ferredoxins and ribosomal proteins) to maintain enzymatic function. For instance, Halobacterium retains enzymatic stability even at 4.5 mM NaCl [48]. Their membranes, composed of diether lipids, form a molecular shield with superior hydrolytic resistance compared to bacterial ester membranes, enabling Methanococcus species to maintain membrane integrity under 15% salinity fluctuation [48].
Metabolic complementarity between archaea and bacteria has also been observed in many previous studies. For example, members of the Halanaerobiales convert bacterial metabolites into osmoprotectants; when co-cultured with Halomonas, glycerol degradation efficiency increases 3.2-fold [49]. Inoculation with the halophilic archaeon Halolamina pelagica CDK2 significantly enhances plant growth under saline conditions, increasing grain yield by 30-50% and improving total protein and chlorophyll content, while reducing antioxidant enzyme activity [50].
However, archaea represent only a small fraction of halotolerant microorganism studies, and direct evidence of plant-archaea interactions remains limited [51]. Addressing the challenges of archaeal cultivation and integrating 16S sequencing with haloproteomics will be essential for identifying novel compatible solutes (e.g., ectoine derivatives, glycine betaine analogs) with potential applications in developing salt-tolerant crops. Such research could ultimately bridge fundamental gaps between prokaryotic and eukaryotic strategies of salt adaptation.
Collectively, these microbial groups exhibit distinct yet functionally convergent strategies to enhance plant performance under salinity. A comparative overview of representative taxa, mechanistic traits, and plant physiological outcomes is presented in Table 1.
Table 1: Representative halotolerant microorganisms, their salt-tolerance mechanisms, and associated plant responses.
| Functional Category | Representative Strains/Genera | Functional Description | References |
|---|---|---|---|
| Hormonal Regulation and Signal Transduction | Enterobacter cloacae, Dietzia natronolimnaea, Pseudomonas PS01 | Modify ethylene and auxin levels; induce expression of TaMYB and TaWRKY genes; regulate stress-responsive genes; activate LOX2 and jasmonic acid (JA) signaling pathways | Yaish et al., 2015 [52]; Bharti et al., 2016 [27]; Chu et al 2019 [53] |
| Antioxidant Defense and Oxidative Stress Mitigation | Arthrobacter sp., Bacillus sp., Bacillus pumilus AM11, Exiguobacterium sp. AM25, Bacillus fortis SSB21 | Increase proline content and total biomass; enhance glutathione, catalase, and peroxidase activities; decrease ROS accumulation | Sziderics et al., 2007 [54]; Ali et al., 2017 [55]; Yasin et al., 2018 [56] |
| Ion Homeostasis and Osmotic Regulation | Bacillus subtilis, Variovorax paradoxus, Planococcus rifetoensis, Pseudomonas migulae 8R6, Pseudomonas fluorescens YSs6, Kocuria rhizophila Y1 | Reduce Na+ accumulation and ROS production; maintain ionic and osmotic balance through ACC deaminase activity that lowers host ACC levels | Ali et al., 2014 [57]; Hashem et al., 2016 [37]; Zhou et al., 2017 [58]; Li et al., 2020 [21] |
| Photosynthesis and Nutrient Metabolism | Halomonas desiderata STR8, Bacillus pumilus AM11, Bacillus cereus Pb25, Exiguobacterium sp. AM25 | Enhance photosynthesis and chlorophyll content; solubilize phosphate and improve available soil P; facilitate nutrient absorption | Bharti et al., 2014 [59]; Islam et al., 2016 [28]; Ali et al., 2017 [55] |
| Biofilm Formation and Rhizosphere Colonization | Planococcus rifietoensis, Pseudomonas sp. P34-L, Halomonas sp. Exo1 | Produce biofilms and extracellular polysaccharides; colonize the rhizosphere, improve root architecture and soil phosphorus availability; promote plant growth under stress | Qurashi and Sabri, 2012 [60]; Mukherjee et al., 2019 [23] |
| Induced Systemic Tolerance | Bacillus amyloliquefaciens FZB42, Bacillus subtilis strain GB03 | Induce systemic salt tolerance via transcriptional regulation; emit VOCs that reduce Na+ accumulation, increase chlorophyll content, and maintain leaf osmotic potential and membrane integrity | Han et al., 2017 [61]; Liu et al., 2017 [62] |
| Symbiotic Nitrogen Fixation and Growth Promotion | Burkholderia cepacia, Bradyrhizobium (RA-5), Kocuria rhizophila Y1 | Fix atmospheric nitrogen, improving nodule formation efficiency in legumes under salt stress; stimulate seed germination under saline conditions | Bano et al., 2015 [63]; Li et al., 2020 [21] |
4 Applications of HT-PGPR in Saline Soil Remediation
4.1 Enhancing Crop Salt Stress Tolerance
4.1.1 Field Performance of Wheat (Dietzia), Soybean (Pseudomonas), and Tomato (Bacillus)
HT-PGPR markedly enhance crop salt tolerance and field yield through multi-mechanistic cooperation. Under saline conditions, these microbes optimize plant metabolism by synthesizing osmolytes (e.g., proline, betaine), regulating ion transporter genes (e.g., HKT family), and activating antioxidant and systemic defense pathways. In wheat, Dietzia inoculation upregulated TaMYB/TaWRKY transcription factors, elevated SOD and CAT activities, increased proline accumulation by 47%, and lowered MDA by 20%, leading to a 23% yield increase under salt stress [27]. For instance, inoculation of wheat with Dietzia natronolimnaea STR1 in a field trial at a single site resulted in a 23% yield increase under saline conditions [64]; multi-site validation is needed to confirm this effect across diverse environments.
In soybean, Pseudomonas strains optimized root architecture by synthesizing IAA while reducing ethylene accumulation through ACC deaminase activity, increasing root surface area by 51%, P uptake by 37%, and biomass gain by 42% in saline fields [7,65]. Similarly, in a greenhouse trial with tomato, inoculation with a Bacillus subtilis consortium under 200 mM NaCl resulted in a significant 13.9% increase in fresh biomass weight per plant (p < 0.05) compared to the non-inoculated control [66]. Collectively, these studies demonstrate that HT-PGPR act as biological “stress modulators”, engineering robust plant-soil feedback loops that translate molecular benefits into tangible yield gains in saline agroecosystems.
4.1.2 Cross-Crop Applicability
HT-PGPR exhibit broad cross-crop applicability by integrating conserved stress-mitigation pathways with species-specific adaptive responses. Their core mechanisms, osmolyte synthesis, EPS secretion, antioxidant activation, and SOS pathway regulation, provide a universal foundation for mitigating osmotic stress and ion toxicity in diverse crops [7]. Wheat inoculated with Dietzia exhibited proline accumulation via TaMYB/TaWRKY activation [27]; soybean-Pseudomonas interactions decreased ethylene levels and improved P mobilization [65]; and Bacillus-treated tomato plants formed EPS-mediated biofilms that restricted Na+ translocation and improved fruit quality [67].
At the same time, functional specialization enables strain-host matching: Rhizobium enhances N fixation in legumes, while Kushneria promotes growth of halophytes like Suaeda salsa through phosphate solubilization [68,69]. Field and pot studies have reported that HT-PGPR inoculation, often combined with organic amendments, can reduce soil electrical conductivity (EC) (e.g., a field study with wheat showed a 9.4% reduction after one growing season [70]; a greenhouse study with maize reported a 31% decrease in soil EC under the phosphogypsum and bacteria co-inoculation [71]. Thus, the cross-crop versatility of HT-PGPR reflects a modular and scalable functional architecture, supporting their deployment as universal microbial tools in saline agriculture.
4.2 Improving Soil Quality and Health
4.2.1 Soil EC and pH Regulation
Beyond enhancing plant resilience, HT-PGPR play vital roles in restoring soil health, particularly in regulating soil EC and pH. By secreting EPS and organic acids, these microbes decrease soluble Na+ concentrations through cation bridging, hydrogen bonding, and anion adsorption, contributing to 18–35% reductions in EC [7]. Low-molecular-weight acids (e.g., oxalic, citric acids) neutralize alkaline compounds, reducing soil pH from 9.4 to 8.6 [70]. During P solubilization, Pseudomonas moraviensis releases H+ ions that lower soil pH and increase P availability, while Bacillus cereus combined with phosphogypsum improves organic matter and dehydrogenase activity [7]. Microbial biofilms act as “natural ion exchangers”, sequestering Na+ and promoting soil aggregation, porosity, and infiltration [21]. Together, these microbially mediated processes drive a progressive shift toward chemically balanced and biologically active soils, laying the foundation for long-term saline-alkaline land improvement.
4.2.2 Increasing Soil Organic Matter and Nutrient Cycling (N/P/K)
HT-PGPR enhance soil organic matter (SOM) and nutrient cycling through synergistic biochemical pathways. EPS serves as a carbon-rich pool that promotes soil aggregation. For instance, soil microorganisms secreting EPS are important for improving soil aggregates, which are key parameters for sustainable soils. Inoculation of bacteria with high EPS production can significantly improve the stability of soil aggregates, enhance soil fertility and improve soil quality [72,73,74]. Nitrogen-fixing Pseudomonas aeruginosa, combined with organic amendments, significantly increased soil N levels compared with uninoculated controls, while in the P cycle, strains such as Oceobacillus picturae and Bacillus licheniformis MH48 increase available P by 18–35% [71]. Potassium mobilization occurs via ion exchange and mineral weathering, raising leaf K content by 14% under low-salt and 8% under high-salt conditions [15]. Enhanced soil enzyme activities (dehydrogenase, phosphatase) foster organic matter mineralization and nutrient turnover, generating a positive microbe-soil-nutrient feedback loop [5]. These improvements demonstrate that HT-PGPR serve as ecological catalysts, accelerating soil nutrient renewal and rebuilding the biogeochemical functionality of saline-degraded systems.
4.2.3 Enhancing Soil Water Retention and Microbial Diversity
HT-PGPR significantly improve soil water retention and microbial diversity through biophysical and ecological mechanisms. EPS produced by strains such as Alcaligenes latus and Pseudomonas aeruginosa CMG1421 functions as natural hydrogels that enhance aggregate stability and porosity [7,60]. This “bio-sponge effect” increased soil infiltration by 35% and root water potential by 28% [15]. EPS-soil complexes create nutrient-rich microhabitats that support diverse microbial communities, increasing microbial biomass carbon by 67% [7]. Planococcus rifietoensis RT4 and Halomonas variabilis HT1 significantly enhanced soil aggregation [75]. Enhanced microbial diversity was reflected in higher actinomycete and diazotroph populations, increased dehydrogenase activity, and improved carbon turnover [69]. Notably, biofilm-associated quorum sensing enhances microbial recruitment and stabilizes a root–microbiome mutualistic network, resulting in a 1.8–2.3-unit increase in the Shannon diversity index under salt stress [70]. These coordinated improvements in water dynamics and microbial ecology contribute to building resilient soil microecosystems capable of withstanding combined salinity and drought pressures.
4.3 Development of Microbial Bioformulations
4.3.1 Selection Criteria for Halotolerant Strains
Screening of HT-PGPR strains should adopt a multi-parameter evaluation system based on key physiological indicators. EPS yield directly correlates with salt-stress mitigation efficiency; high-yielding strains (e.g., Halomonas sp. EX01) can produce EPS accounting for 40-90% of dry biomass under salt stress, forming dense ion-binding matrices that restrict Na+ migration toward plant roots [76]. Conversely, an EPS-deficient mutant of Pantoea alhagi NX-11 lost antioxidant activity, reinforcing the positive linkage between EPS abundance and oxidative defense [66]. ACC deaminase activity must be verified both genetically and enzymatically. High-performance strains such as Pseudomonas sp. UW4 degrade plant ACC into α-ketobutyrate and ammonia, reducing ethylene accumulation by 35-50% under salt stress [67]. The activity threshold should exceed 4.8 μmol α-KB mg−1h−1 [65], with gene confirmation using acdSf3/acdSr3 primers to ensure acdS conservation [66]. Biofilm formation, typically quantified via crystal violet staining, represents another essential trait. Biofilm-forming strains such as Enterobacter cloacae establish ion-filtering microcolonies within rhizosphere pores, enhancing macroaggregate stability by up to 60% [77]. Collectively, EPS provides the structural scaffold, ACC deaminase maintains the rhizosphere physiological balance, and biofilms strengthen ecological persistence, together forming a “physical-physiological-ecological” triple-protection mechanism [7]. These criteria have guided the industrial development of more than 20 core strains (e.g., Bacillus amyloliquefaciens, Pseudomonas fluorescens), establishing a robust scientific foundation for selecting HT-PGPR-based bioformulations in saline soil remediation.
4.3.2 Design Principles of Synthetic Microbial Consortia
The rational design of synthetic microbial consortia (SynComs) represents a major step toward improving productivity and remediation efficiency in saline-alkaline soils. SynComs combine strains with complementary halotolerance and growth-promoting functions to form synergistic and resilient communities. Three fundamental principles should be used to guide design: (i) inter-strain interactions, understanding symbiotic, antagonistic, or competitive relationships enables optimized combinations that maximize cooperation and minimize interference [69]; (ii) environmental adaptability, selecting highly salt-tolerant and stress-adaptive strains ensures survival and colonization under fluctuating saline conditions [71]; and (iii) stability and persistence, optimizing formulation and encapsulation technologies (e.g., microcapsules) helps shield microbes from environmental fluctuations and extend field efficacy [69,70]. Together, these principles underpin the construction of stable, high-performance SynComs that sustainably enhance soil productivity and accelerate saline–alkaline land restoration.
Although these rationally designed SynComs, constructed based on inter-strain synergy, environmental adaptability, and stability principles, may effectively enhance plant salt tolerance and improve soil quality through various mechanisms, their successful field application still faces numerous limitations and challenges. The main bottlenecks include: poor survival and colonization efficiency of introduced strains in complex soil environments (especially under salt stress); intense spatial and nutritional competition with native microbial communities; high heterogeneity of field soil physicochemical properties (such as salinity, pH, and texture); and immature inoculant formulation and delivery technologies, which make it difficult to ensure long-term viability and efficacy of the bacterial cells. These factors lead to inconsistent and unstable growth-promoting and salt-tolerance effects in field applications, which often differ significantly from laboratory results. Future research and application need to focus on developing more environmentally adaptable inoculant formulations (e.g., using nanomaterials, adding protective agents), utilizing microbial community synergies, and conducting targeted design and testing based on specific soil conditions, in order to bridge the gap between experimental research and practical application.
4.3.3 Role and Prospects of Artificial Intelligence (AI) in Designing Halotolerant Microbial Consortia
The construction of halotolerant PGPR consortia has shifted from empirical screening toward AI-assisted rational design. Artificial intelligence integrates multi-omics datasets, predicts microbial interactions, and optimizes SynCom composition, substantially improving design efficiency, functional predictability, and ecological stability. Machine learning (ML) and deep learning (DL) can integrate metagenomic, metabolomic, and transcriptomic data to identify key functional strains associated with salt tolerance, including genes for compatible solute synthesis (ectoine, betaine), ion homeostasis (Na+/H+ antiporters), and auxin or quorum-sensing signal production. Feature selection algorithms (e.g., random forest, SVM) can prioritize strains most responsive to salt stress while minimizing redundancy, predicting compatibility and long-term stability within SynComs.
AI-based ecological modeling, using network analysis and Lotka-Volterra dynamics, can anticipate cooperative or competitive interactions, avoiding inhibitory combinations. Dynamic simulations also optimize inoculation order and strain ratios to maintain SynCom stability. In practical applications, AI integrates environmental variables (salinity, pH, moisture) to design location-specific SynComs. Real-time monitoring (fluorescence imaging, qPCR) coupled with DBTL (Design-Build-Test-Learn) cycles enables continuous optimization. Platforms such as BacterAI iteratively refine SynCom performance using automated experimental feedback [78,79]. Moreover, DL-based phenotyping (e.g., CNN analysis of plant images) enables early prediction of plant salt-tolerance responses, facilitating proactive management. While AI-assisted design shows promise for microbial strain selection and SynCom optimization, its application in rhizosphere systems remains nascent and is supported by only limited, proof-of-concept studies. Current models are often trained on greenhouse-derived datasets, making them vulnerable to data bias and limited field transferability [80]. Consequently, AI/ML predictions require rigorous mechanistic validation and field-scale testing before reliable application.
4.4 To What Extent Does the Salinity Gradient Determine Microbial Remediation Potential?
The efficacy of SynComs in ameliorating salt stress is likely not uniform across soil salinity gradients. Theoretical and empirical evidence suggests that their beneficial impacts may be more pronounced in lightly to moderately saline soils, while potentially diminishing in severely saline-alkaline conditions [43]. In light saline soils, the primary constraints on plant growth are often osmotic stress and initial ionic imbalance. Here, PGPR/SynComs can effectively deploy their core mechanisms, such as the production of osmoprotectants (e.g., proline, glycine betaine) [66], ACC deaminase-driven ethylene reduction [71], and moderate enhancement of nutrient solubilization and phytohormone modulation [77]. These interventions can significantly improve seedling establishment, root growth, and overall plant vigor, providing a crucial buffer against the initial salinity shock [64].
However, as salinity intensifies to moderate and severe levels, the extreme osmotic pressure, specific ion toxicity (Na+, Cl−), and deteriorated soil physicochemical properties (e.g., high pH, poor structure) exceed the physiological tolerance of many beneficial strains [43,60]. Severe stress reduces microbial survival and colonization, while intense ion toxicity and extreme pH damage microbial enzymes and impair key plant-growth-promoting functions [81]. Although some halotolerant strains exhibit remarkable resilience [9,42], their ability to meaningfully influence plant physiology under such extreme abiotic stress may be substantially curtailed [70].
In summary, while bio-inoculants represent a promising strategy for saline soil remediation, their application might be most effective as a proactive or early-intervention measure in lightly to moderately affected soils, whereas their role in severely degraded lands may be limited unless integrated with other amelioration practices (e.g., organic amendments, physical leaching) [7,82].
5 Conclusion and Future Perspectives
HT-PGPR provide a sustainable biological approach for enhancing productivity and ecological restoration in saline soils. By regulating osmotic adjustment, ion homeostasis, and plant hormonal-signaling pathways, HT-PGPR improve plant salt tolerance while reshaping rhizosphere functions. However, field-scale translation remains challenging due to limited adaptability, inconsistent stability, and competition with native microbiota. To overcome these constraints, future research should focus on several key aspects.
(1) Screening efficient and resilient HT-PGPR strains. Advanced approaches such as high-throughput sequencing, genome mining, and precision editing can facilitate the discovery of highly adaptable and functionally consistent strains, followed by validation across diverse soil and climatic conditions.
(2) Developing novel HT-PGPR formulations and inoculation technologies. Integrating modern biotechnology with material science may enable innovative encapsulation or carrier technologies that improve microbial survival, rhizosphere colonization, and long-term persistence in saline environments.
(3) Enhancing AI-assisted construction of SynComs. Building specialized datasets for halotolerant microbes and applying artificial intelligence algorithms can facilitate tailored SynComs optimized for specific salinity-alkalinity contexts.
Overall, HT-PGPR represent an eco-friendly and promising solution for saline soil remediation and sustainable agriculture. With the integration of microbial ecology, systems biology, and intelligent design technologies, their applications are expected to evolve from experimental validation toward large-scale, data-driven, and intelligent agricultural practices.
Acknowledgement:
Funding Statement: This work was supported by the Chinese Academy of Sciences, the Visiting Research & Practice Program (FG2025311) of Department of Education of Zhejiang Province, Science and Technology Plan Project of the Science and Technology Bureau of Taizhou (25nya06) and the New seedling Talents program, High-Level Talent Research Initiation Fund (322503002) and Provincial-Level Research Incubation Program (26PYK15) of Taizhou Vocational College of Science & Technology.
Author Contributions: Hualiang Zhang, Shuqin Gao and Congcong Zheng: Conceptualization, Data curation, Funding acquisition, Writing—original draft, Writing—review & editing. Honglong Zhao, Tianru Qu, Hao Jiang and Yucheng Zhang: Conceptualization, Writing—original draft, Writing—review & editing. All authors reviewed and approved the final version of the manuscript.
Availability of Data and Materials: Not applicable.
Ethics Approval: Not applicable.
Conflicts of Interest: The authors declare no conflicts of interest.
References
1. Wicke B , Smeets E , Dornburg V , Vashev B , Gaiser T , Turkenburg W , et al. The global technical and economic potential of bioenergy from salt-affected soils. Energy Environ Sci. 2011; 4( 8): 2669– 81. doi:10.1039/C1EE01029H. [Google Scholar] [CrossRef]
2. Hossain MS . Present scenario of global salt affected soils, its management and importance of salinity research. Int Res J Biol Sci. 2019; 1( 1): 1– 3. [Google Scholar]
3. Paul D , Lade H . Plant-growth-promoting rhizobacteria to improve crop growth in saline soils: A review. Agron Sustain Dev. 2014; 34( 4): 737– 52. doi:10.1007/s13593-014-0233-6. [Google Scholar] [CrossRef]
4. Semida WM , Abd El-Mageed TA , Abdalla RM , Hemida KA , Howladar SM , Leilah AAA , et al. Sequential antioxidants foliar application can alleviate negative consequences of salinity stress in Vicia faba L. Plants. 2021; 10( 5): 914. doi:10.3390/plants10050914. [Google Scholar] [CrossRef]
5. AbuQamar SF , El-Saadony MT , Saad AM , Desoky EM , Elrys AS , El-Mageed TAA , et al. Halotolerant plant growth-promoting rhizobacteria improve soil fertility and plant salinity tolerance for sustainable agriculture—A review. Plant Stress. 2024; 12: 100482. doi:10.1016/j.stress.2024.100482. [Google Scholar] [CrossRef]
6. Pan Y , Liu W , Meng J , Song T , Xie J . Reserch progress on enhancing plant growth under salt stress by salt-tolerant growth-promoting bacteria. Chin J Bioprocess Eng. 2024; 22( 2): 182– 8. (In Chinese). [Google Scholar]
7. Kabir AH , Baki MZI , Ahmed B , Mostofa MG . Current, faltering, and future strategies for advancing microbiome-assisted sustainable agriculture and environmental resilience. New Crops. 2024; 1: 100013. doi:10.1016/j.ncrops.2024.100013. [Google Scholar] [CrossRef]
8. Qadir M , Quillérou E , Nangia V , Murtaza G , Singh M , Thomas RJ , et al. Economics of salt-induced land degradation and restoration. Nat Resour Forum. 2014; 38( 4): 282– 95. doi:10.1111/1477-8947.12054. [Google Scholar] [CrossRef]
9. Shrivastava P , Kumar R . Soil salinity: A serious environmental issue and plant growth promoting bacteria as one of the tools for its alleviation. Saudi J Biol Sci. 2015; 22( 2): 123– 31. doi:10.1016/j.sjbs.2014.12.001. [Google Scholar] [CrossRef]
10. Lu Y , Fricke W . Salt stress—Regulation of root water uptake in a whole-plant and diurnal context. Int J Mol Sci. 2023; 24( 9): 8070. doi:10.3390/ijms24098070. [Google Scholar] [CrossRef]
11. Seleiman MF , Talha Aslam M , Ahmed Alhammad B , Umair Hassan M , Maqbool R , Umer Chattha M , et al. Salinity stress in wheat: Effects, mechanisms and management strategies. Phyton. 2022; 91( 4): 667– 94. doi:10.32604/phyton.2022.017365. [Google Scholar] [CrossRef]
12. Yuan J , Cao H , Qin W , Yang S , Zhang D , Zhu L , et al. Genomic and modern biotechnological strategies for enhancing salt tolerance in crops. New Crops. 2025; 2: 100057. doi:10.1016/j.ncrops.2024.100057. [Google Scholar] [CrossRef]
13. He J , Chen Y , Zhang M , Qiu Y , Zhou H , Li M . Current perspectives on improving soybean performance on saline-alkaline lands. New Crops. 2026; 3: 100079. doi:10.1016/j.ncrops.2025.100079. [Google Scholar] [CrossRef]
14. Seleiman MF , Ahmad A , Alhammad BA , Tola E . Exogenous application of zinc oxide nanoparticles improved antioxidants, photosynthetic, and yield traits in salt-stressed maize. Agronomy. 2023; 13( 10): 2645. doi:10.3390/agronomy13102645. [Google Scholar] [CrossRef]
15. Habib SH , Kausar H , Saud HM . Plant growth-promoting rhizobacteria enhance salinity stress tolerance in okra through ROS-scavenging enzymes. BioMed Res Int. 2016; 2016: 6284547. doi:10.1155/2016/6284547. [Google Scholar] [CrossRef]
16. Luo H , Wu H-J , Xie Y-L , Gao X-W . Effects of Bacillus megaterium CJLC2 on the growth and the salt-tolerance related physiological and biochemical characters of tomato under salt stress. J Plant Prot. 2013; 40( 5): 431– 6. (In Chinese). [Google Scholar]
17. Jha Y , Subramanian RB , Patel S . Combination of endophytic and rhizospheric plant growth promoting rhizobacteria in Oryza sativa shows higher accumulation of osmoprotectant against saline stress. Acta Physiol Plant. 2011; 33( 3): 797– 802. doi:10.1007/s11738-010-0604-9. [Google Scholar] [CrossRef]
18. Ma J , Xiang Z , Zhang D , Liao L , Wang C , Peng M . Research progress of plant growth-promoting bacteria in salt tolerance of plants. Heilongjiang Agricutural Sci. 2024;( 1): 108– 15. (In Chinese). [Google Scholar]
19. Wang ZL . Construction and genome sequencing of salt tolerant and growth-promoting synthetic community in maize rhizosphere [ master’s thesis]. Taian, China: Shandong Agricultural University; 2022. (In Chinese). [Google Scholar]
20. Latif A , Ahmad R , Ahmed J , Mueen H , Ali Khan S , Bibi G , et al. Novel halotolerant PGPR strains alleviate salt stress by enhancing antioxidant activities and expression of selected genes leading to improved growth of Solanum lycopersicum. Sci Hortic. 2024; 338: 113625. doi:10.1016/j.scienta.2024.113625. [Google Scholar] [CrossRef]
21. Li X , Sun P , Zhang Y , Jin C , Guan C . A novel PGPR strain Kocuria rhizophila Y1 enhances salt stress tolerance in maize by regulating phytohormone levels, nutrient acquisition, redox potential, ion homeostasis, photosynthetic capacity and stress-responsive genes expression. Environ Exp Bot. 2020; 174: 104023. doi:10.1016/j.envexpbot.2020.104023. [Google Scholar] [CrossRef]
22. Shi LN , Lu LX , Ye JR , Shi HM . The endophytic strain ZS-3 enhances salt tolerance in Arabidopsis thaliana by regulating photosynthesis, osmotic stress, and ion homeostasis and inducing systemic tolerance. Front Plant Sci. 2022; 13: 820837. doi:10.3389/fpls.2022.820837. [Google Scholar] [CrossRef]
23. Mukherjee P , Mitra A , Roy M . Halomonas rhizobacteria of Avicennia marina of Indian sundarbans promote rice growth under saline and heavy metal stresses through exopolysaccharide production. Front Microbiol. 2019; 10: 1207. doi:10.3389/fmicb.2019.01207. [Google Scholar] [CrossRef]
24. Parks GE . Increased vacuolar Na+/H+ exchange activity in Salicornia bigelovii Torr. in response to NaCl. J Exp Bot. 2002; 53( 371): 1055– 65. doi:10.1093/jexbot/53.371.1055. [Google Scholar] [CrossRef]
25. Berthomieu P , Conéjéro G , Nublat A , Brackenbury WJ , Lambert C , Savio C , et al. Functional analysis of AtHKT1 in Arabidopsis shows that Na+ recirculation by the phloem is crucial for salt tolerance. EMBO J. 2003; 22( 9): 2004– 14. doi:10.1093/emboj/cdg207. [Google Scholar] [CrossRef]
26. Panwar M , Tewari R , Nayyar H . Native halo-tolerant plant growth promoting rhizobacteria Enterococcus and Pantoea sp. improve seed yield of Mungbean (Vigna radiata L.) under soil salinity by reducing sodium uptake and stress injury. Physiol Mol Biol Plants. 2016; 22( 4): 445– 59. doi:10.1007/s12298-016-0376-9. [Google Scholar] [CrossRef]
27. Bharti N , Pandey SS , Barnawal D , Patel VK , Kalra A . Plant growth promoting rhizobacteria Dietzia natronolimnaea modulates the expression of stress responsive genes providing protection of wheat from salinity stress. Sci Rep. 2016; 6: 34768. doi:10.1038/srep34768. [Google Scholar] [CrossRef]
28. Islam F , Yasmeen T , Arif MS , Ali S , Ali B , Hameed S , et al. Plant growth promoting bacteria confer salt tolerance in Vigna radiata by up-regulating antioxidant defense and biological soil fertility. Plant Growth Regul. 2016; 80( 1): 23– 36. doi:10.1007/s10725-015-0142-y. [Google Scholar] [CrossRef]
29. Mal S , Panchal S . Drought and salt stress mitigation in crop plants using stress-tolerant auxin-producing endophytic bacteria: A futuristic approach towards sustainable agriculture. Front Plant Sci. 2024; 15: 1422504. doi:10.3389/fpls.2024.1422504. [Google Scholar] [CrossRef]
30. Yang XY , Chen ZY , Jiang PP , Zhang RS , Liu YZ , Liu YF . Growth-promotion effect of Bacillus amyloliquefaciens B1619 on tomato plant. Chin J Biol Control. 2016; 32( 03): 349– 56. (In Chinese). [Google Scholar]
31. Mantelin S . Plant growth-promoting bacteria and nitrate availability: Impacts on root development and nitrate uptake. J Exp Bot. 2003; 55( 394): 27– 34. doi:10.1093/jxb/erh010. [Google Scholar] [CrossRef]
32. Costa-Gutierrez SB , Lami MJ , Santo MCC , Zenoff AM , Vincent PA , Molina-Henares MA , et al. Plant growth promotion by Pseudomonas putida KT2440 under saline stress: Role of eptA. Appl Microbiol Biotechnol. 2020; 104( 10): 4577– 92. doi:10.1007/s00253-020-10516-z. [Google Scholar] [CrossRef]
33. López-Gómez M , Palma F , Lluch C . Strategies of salt tolerance in the rhizobia-legume symbiosis. In: Beneficial plant-microbial interactions. Boca Raton, FL, USA: CRC Press; 2013. p. 99– 121. doi:10.1201/b15251-6. [Google Scholar] [CrossRef]
34. Crowley DE , Wang YC , Reid CPP , Szaniszlo PJ . Mechanisms of iron acquisition from siderophores by microorganisms and plants. Plant Soil. 1991; 130( 1): 179– 98. doi:10.1007/BF00011873. [Google Scholar] [CrossRef]
35. Peng HY , Shao YD , Li H , Li PG , Ding BL , Yang TJ , et al. Study on the mechanism of increased iron availability by rhizosphere iron-solubilizing bacteria in combination with AM fungi in calcareous soil. Acta Pedol Sin. 2024; 61( 04): 1111– 22. (In Chinese). [Google Scholar]
36. Barnawal D , Bharti N , Pandey SS , Pandey A , Chanotiya CS , Kalra A . Plant growth-promoting rhizobacteria enhance wheat salt and drought stress tolerance by altering endogenous phytohormone levels and TaCTR1/TaDREB2 expression. Physiol Plant. 2017; 161( 4): 502– 14. doi:10.1111/ppl.12614. [Google Scholar] [CrossRef]
37. Hashem A , Abd_Allah EF , Alqarawi AA , Al-Huqail AA , Shah MA . Induction of osmoregulation and modulation of salt stress in Acacia gerrardii Benth. by arbuscular mycorrhizal fungi and Bacillus subtilis (BERA 71). BioMed Res Int. 2016; 2016: 6294098. doi:10.1155/2016/6294098. [Google Scholar] [CrossRef]
38. Roy SJ , Negrão S , Tester M . Salt resistant crop plants. Curr Opin Biotechnol. 2014; 26: 115– 24. doi:10.1016/j.copbio.2013.12.004. [Google Scholar] [CrossRef]
39. Behera BK , Das P , Maharana J , Paria P , Mandal SN , Meena DK , et al. Draft genome sequence of the extremely halophilic bacterium Halomonas salina strain CIFRI1, isolated from the east coast of India. Genome Announc. 2015; 3: e01321-14. doi:10.1128/genomea.01321-14. [Google Scholar] [CrossRef]
40. Hernández-Canseco J , Bautista-Cruz A , Sánchez-Mendoza S , Aquino-Bolaños T , Sánchez-Medina PS . Plant growth-promoting halobacteria and their ability to protect crops from abiotic stress: An eco-friendly alternative for saline soils. Agronomy. 2022; 12( 4): 804. doi:10.3390/agronomy12040804. [Google Scholar] [CrossRef]
41. Bharti N , Barnawal D , Maji D , Kalra A . Halotolerant PGPRs prevent major shifts in indigenous microbial community structure under salinity stress. Microb Ecol. 2015; 70( 1): 196– 208. doi:10.1007/s00248-014-0557-4. [Google Scholar] [CrossRef]
42. Li HP , Ma HB , Zhang JL . Halo-tolerant plant growth-promoting bacteria-mediated plant salt resistance and microbiome-based solutions for sustainable agriculture in saline soils. FEMS Microbiol Ecol. 2025; 101( 5): fiaf037. doi:10.1093/femsec/fiaf037. [Google Scholar] [CrossRef]
43. Kumar Arora N , Fatima T , Mishra J , Mishra I , Verma S , Verma R , et al. Halo-tolerant plant growth promoting rhizobacteria for improving productivity and remediation of saline soils. J Adv Res. 2020; 26: 69– 82. doi:10.1016/j.jare.2020.07.003. [Google Scholar] [CrossRef]
44. Porcel R , Aroca R , Azcon R , Ruiz-Lozano JM . Regulation of cation transporter genes by the arbuscular mycorrhizal symbiosis in rice plants subjected to salinity suggests improved salt tolerance due to reduced Na+ root-to-shoot distribution. Mycorrhiza. 2016; 26( 7): 673– 84. doi:10.1007/s00572-016-0704-5. [Google Scholar] [CrossRef]
45. Augé RM , Toler HD , Saxton AM . Arbuscular mycorrhizal symbiosis and osmotic adjustment in response to NaCl stress: A meta-analysis. Front Plant Sci. 2014; 5: 562. doi:10.3389/fpls.2014.00562. [Google Scholar] [CrossRef]
46. Vimal SR , Patel VK , Singh JS . Plant growth promoting Curtobacterium albidum strain SRV4: An agriculturally important microbe to alleviate salinity stress in paddy plants. Ecol Indic. 2019; 105: 553– 62. doi:10.1016/j.ecolind.2018.05.014. [Google Scholar] [CrossRef]
47. Lee Y , Krishnamoorthy R , Selvakumar G , Kim K , Sa T . Alleviation of salt stress in maize plant by co-inoculation of arbuscular mycorrhizal fungi and Methylobacterium oryzae CBMB20. J Korean Soc Appl Biol Chem. 2015; 58( 4): 533– 40. doi:10.1007/s13765-015-0072-4. [Google Scholar] [CrossRef]
48. Ventosa A , Nieto JJ , Oren A . Biology of moderately halophilic aerobic bacteria. Microbiol Mol Biol Rev. 1998; 62( 2): 504– 44. doi:10.1128/mmbr.62.2.504-544.1998. [Google Scholar] [CrossRef]
49. Meinzer M , Ahmad N , Nielsen BL . Halophilic plant-associated bacteria with plant-growth-promoting potential. Microorganisms. 2023; 11( 12): 2910. doi:10.3390/microorganisms11122910. [Google Scholar] [CrossRef]
50. Naitam MG , Ramakrishnan B , Grover M , Kaushik R . Rhizosphere-dwelling halophilic archaea: a potential candidate for alleviating salinity-associated stress in agriculture. Front Microbiol. 2023; 14: 1212349. doi:10.3389/fmicb.2023.1212349. [Google Scholar] [CrossRef]
51. Liu C , Baffoe DK , Zhan Y , Zhang M , Li Y , Zhang G . Halophile, an essential platform for bioproduction. J Microbiol Meth. 2019; 166: 105704. doi:10.1016/j.mimet.2019.105704. [Google Scholar] [CrossRef]
52. Yaish MW , Antony I , Glick BR . Isolation and characterization of endophyticplant growth-promoting bacteria from date palm tree (Phoenix dactylifera L.) andtheir potential role in salinity tolerance. Antonie Van Leeuwenhoek. 2015; 107: 1519– 32. doi:10.1007/s10482-015-0445-z [Google Scholar] [CrossRef]
53. Chu TN , Tran BTH , Van Bui L , Hoang MTT . Plant growth-promoting rhizobacterium Pseudomonas PS01 induces salt tolerance in Arabidopsis thaliana. BMC Res Notes. 2019; 12( 1): 11. doi:10.1186/s13104-019-4046-1. [Google Scholar] [CrossRef]
54. Sziderics AH , Rasche F , Trognitz F , Sessitsch A , Wilhelm E . Bacterialendophytes contribute to abiotic stress adaptation in pepper plants (Capsicum annuum L.). Can J Microbiol. 2007; 53( 11): 1195– 202. doi:10.1139/W07-082. [Google Scholar] [CrossRef]
55. Ali A , Shahzad R , Khan AL , Halo BA , Al-Yahyai R , Al-Harrasi A , et al. Endophytic bacterial diversity of Avicennia marina helps to confer resistance against salinity stress in Solanum lycopersicum. J Plant Interact. 2017; 12( 1): 312– 22. doi:10.1080/17429145.2017.1362051 [Google Scholar] [CrossRef]
56. Yasin NA , Akram W , Khan WU , Ahmad SR , Ahmad A , Ali A , et al. Halotolerant plant-growth promoting rhizobacteria modulate gene expression and osmolyte production to improve salinity tolerance and growth in Capsicum annum L. Environ Sci Pollut Res. 2018; 25( 23): 23236– 50. doi:10.1007/s11356-018-2381-8. [Google Scholar] [CrossRef]
57. Ali S , Charles TC , Glick BR . Amelioration of high salinity stress damage byplant growth-promoting bacterial endophytes that contain ACC deaminase. Plant Physiol Biochem. 2014; 80: 160– 7. doi:10.1016/j.plaphy.2014.04.003. [Google Scholar] [CrossRef]
58. Zhou N , Zhao S , Tian C . Effect of halotolerant rhizobacteria isolated fromhalophytes on the growth of sugar beet (Beta vulgaris L.) under salt stress. FEMS Microbiol Lett. 2017; 364: 11. doi:10.1093/femsle/fnx091. [Google Scholar] [CrossRef]
59. Bharti N; Barnawal D; Awasthi A; Yadav A; Kalra A . Plant growth promoting rhizobacteria alleviatesalinity induced negative effects on growth, oil content and physiological status in Mentha arvensis. Acta Physiol Plant. 2014; 36: 45– 60. doi:10.1007/s11738-013-1385-8. [Google Scholar] [CrossRef]
60. Qurashi AW , Sabri AN . Bacterial exopolysaccharide and biofilm formation stimulate chickpea growth and soil aggregation under salt stress. Braz J Microbiol. 2012; 43( 3): 1183– 91. doi:10.1590/s1517-83822012000300046. [Google Scholar] [CrossRef]
61. Han QQ , Wu YN , Gao HJ , Xu R , Paré PW , Shi H , et al. Improved salt tolerance of medicinal plant Codonopsis pilosula by Bacillus amyloliquefaciens GB03. Acta Physiol Plant. 2017; 39( 1): 35. doi:10.1007/s11738-016-2325-1. [Google Scholar] [CrossRef]
62. Liu S , Hao H , Lu X , Zhao X , Wang Y , Zhang YB , et al. Transcriptome profiling of genes involved in induced systemic salt tolerance conferred by Bacillus amyloliquefaciens FZB42 in Arabidopsis thaliana. Sci Rep. 2017; 7( 1): 10795. doi:10.1038/s41598-017-11308-8. [Google Scholar] [CrossRef]
63. Bano DA , Singh R , Waza SA , Singh N . Effect of cowpea Bradyrhizobium (RA-5) and Burkholderia cepacia (RRE-5) on growth parameters of pigeonpea under saltstress conditions. J Pure Appl Microbiol. 2015; 9: 2539– 46. [Google Scholar]
64. Nadeem SM , Ahmad Zahir Z , Naveed M , Nawaz S . Mitigation of salinity-induced negative impact on the growth and yield of wheat by plant growth-promoting rhizobacteria in naturally saline conditions. Ann Microbiol. 2013; 63( 1): 225– 32. doi:10.1007/s13213-012-0465-0. [Google Scholar] [CrossRef]
65. Saravanakumar D , Samiyappan R . ACC deaminase from Pseudomonas fluorescens mediated saline resistance in groundnut (Arachis hypogea) plants. J Appl Microbiol. 2007; 102( 5): 1283– 92. doi:10.1111/j.1365-2672.2006.03179.x. [Google Scholar] [CrossRef]
66. Li Z , Chang S , Ye S , Chen M , Lin L , Li Y , et al. Differentiation of 1-aminocyclopropane-1-carboxylate (ACC) deaminase from its homologs is the key for identifying bacteria containing ACC deaminase. FEMS Microbiol Ecol. 2015; 91( 10): fiv112. doi:10.1093/femsec/fiv112. [Google Scholar] [CrossRef]
67. Mahmood S , Daur I , Al-Solaimani SG , Ahmad S , Madkour MH , Yasir M , et al. Plant growth promoting rhizobacteria and silicon synergistically enhance salinity tolerance of mung bean. Front Plant Sci. 2016; 7: 876. doi:10.3389/fpls.2016.00876. [Google Scholar] [CrossRef]
68. Roychowdhury R , Qaiser TF , Mukherjee P , Roy M . Isolation and characterization of a Pseudomonas aeruginosa strain PGP for plant growth promotion. Proc Natl Acad Sci Ind Sect B Biol Sci. 2019; 89( 1): 353– 60. doi:10.1007/s40011-017-0946-9. [Google Scholar] [CrossRef]
69. Chen MX , Wei Z , Tian L , Tan Y , Huang JD , Dai L . Design and application of synthetic microbial communities. Chin Sci Bull. 2021; 66( 03): 273– 83. (In Chinese). doi:10.1360/TB-2020-0487. [Google Scholar] [CrossRef]
70. Glick BR . Plant growth-promoting bacteria: Mechanisms and applications. Scientifica. 2012; 2012: 963401. doi:10.6064/2012/963401. [Google Scholar] [CrossRef]
71. Munns R , Tester M . Mechanisms of salinity tolerance. Annu Rev Plant Biol. 2008; 59: 651– 81. doi:10.1146/annurev.arplant.59.032607.092911. [Google Scholar] [CrossRef]
72. Sandhya V , Ali SZ . The production of exopolysaccharide by Pseudomonas putida GAP-P45 under various abiotic stress conditions and its role in soil aggregation. Microbiology. 2015; 84( 4): 512– 9. doi:10.1134/S0026261715040153. [Google Scholar] [CrossRef]
73. Zhang M , Wu Y , Qu C , Huang Q , Cai P . Microbial extracellular polymeric substances (EPS) in soil: From interfacial behaviour to ecological multifunctionality. Geo Bio Interfaces. 2024; 1: e4. doi:10.1180/gbi.2024.4. [Google Scholar] [CrossRef]
74. Egamberdieva D , Jabborova D , Berg G . Synergistic interactions between Bradyrhizobium japonicum and the endophyte Stenotrophomonas rhizophila and their effects on growth, and nodulation of soybean under salt stress. Plant Soil. 2016; 405( 1): 35– 45. doi:10.1007/s11104-015-2661-8. [Google Scholar] [CrossRef]
75. Zhang XQ , Chen H , Qiao YQ , Cao CF , Du SZ , Li W , et al. Effects of different long-term fertilization models on culturable microbial quantity and enzyme Activities of lime concretion black soil. Ecol Environ Sci. 2016; 25( 08): 1283– 90. (In Chinese). [Google Scholar]
76. Ashraf M , Hasnain S , Berge O , Mahmood T . Inoculating wheat seedlings with exopolysaccharide-producing bacteria restricts sodium uptake and stimulates plant growth under salt stress. Biol Fertil Soils. 2004; 40( 3): 157– 62. doi:10.1007/s00374-004-0766-y. [Google Scholar] [CrossRef]
77. Halder A , Banerjee J , Bhattacharyya P , Pramanik K , Debnath A . Isolation of lentil-specific salt tolerant nitrogen fixing bacteria from Murshidabad district of West Bengal. J Crop Weed. 2016; 12( 3): 14– 9. [Google Scholar]
78. Rubrice R , Gueneau V , Briandet R , Cornuejols A , Guigue V . A machine learning framework for the prediction and analysis of bacterial antagonism in biofilms using morphological descriptors. Artif Intell Life Sci. 2025; 8: 100137. doi:10.1016/j.ailsci.2025.100137. [Google Scholar] [CrossRef]
79. Dama AC , Kim KS , Leyva DM , Lunkes AP , Schmid NS , Jijakli K , et al. BacterAI maps microbial metabolism without prior knowledge. Nat Microbiol. 2023; 8( 6): 1018– 25. doi:10.1038/s41564-023-01376-0. [Google Scholar] [CrossRef]
80. Zhang Z , Chen G , Hussain W , Pan Y , Yang Z , Liu Y , et al. Machine learning and network analysis with focus on the biofilm in Staphylococcus aureus. Comput Struct Biotechnol J. 2024; 23: 4148– 60. doi:10.1016/j.csbj.2024.11.011. [Google Scholar] [CrossRef]
81. Wang W , Sun X , Zheng F , Zhang Z , Wang Z , Qu L , et al. Salt–alkali-resistant phosphate-solubilizing bacterium: Kushneria sp. YCWA18 improves soil available phosphorus and promotes the growth of Suaeda salsa. J Plant Growth Regul. 2024; 43( 1): 272– 82. doi:10.1007/s00344-023-11083-z. [Google Scholar] [CrossRef]
82. Jiang H , Li S , Wang T , Chi X , Qi P , Chen G . Interaction between halotolerant phosphate-solubilizing bacteria (Providencia rettgeri strain TPM23) and rock phosphate improves soil biochemical properties and peanut growth in saline soil. Front Microbiol. 2021; 12: 777351. doi:10.3389/fmicb.2021.777351. [Google Scholar] [CrossRef]
Cite This Article
 Copyright © 2026 The Author(s). Published by Tech Science Press.
Copyright © 2026 The Author(s). Published by Tech Science Press.This work is licensed under a Creative Commons Attribution 4.0 International License , which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.


 Submit a Paper
Submit a Paper Propose a Special lssue
Propose a Special lssue View Full Text
View Full Text Download PDF
Download PDF Downloads
Downloads
 Citation Tools
Citation Tools