 Open Access
Open Access
REVIEW
Multi-Omics Approach in the Transcriptional Regulation of Coumarin Biosynthetic Pathway
College of Life and Health, West Anhui University, Lu’an, China
* Corresponding Authors: Cheng Song. Email: ,
(This article belongs to the Special Issue: Plant Hormone Signaling and Environmental Cue Integration: Mechanisms and Regulatory Networks)
Phyton-International Journal of Experimental Botany 2026, 95(3), 7 https://doi.org/10.32604/phyton.2026.076582
Received 23 November 2025; Accepted 25 February 2026; Issue published 31 March 2026
Abstract
Coumarin, a significant secondary metabolite derived from the phenylpropanoid pathway, exhibits a broad spectrum of biological activities and medicinal properties. Structurally, coumarins are categorized into simple and angular types based on variations in the core structure of the parent skeleton, with further classifications as furanocoumarins and pyranocoumarins depending on their carbon atom count. In nature, coumarins are predominantly found in plants within families such as Apiaceae, Asteraceae, and Thymelaeaceae. The structural diversity observed among coumarins in higher plants is largely driven by coevolutionary interactions between herbivorous insects and their host plants. With recent advancements in multi-omics technologies, research has increasingly focused on unraveling the transcriptional regulation of key genes in the coumarin biosynthetic pathway. This review consolidates recent findings from genomics, transcriptomics, proteomics, and metabolomics studies to identify, characterize, and regulate coumarin-related genes. These insights provide a foundational reference for further research efforts in gene discovery, molecular regulation, and synthetic biology applications in the context of coumarin biosynthesis.Keywords
Coumarin, a phenylpropanolide synthesized via the shikimic acid pathway from phenylalanine and tyrosine, is known for its pleasant fragrance and is widely used as a flavoring agent in foods and beverages [1]. Found extensively in both monocots and dicots, coumarins were first isolated from Dipteryx odorata and are particularly abundant in families such as Apiaceae, Rutaceae, Asteraceae, Fabaceae, Oleaceae, Moraceae, and Thymelaeaceae. In nature, coumarins typically occur in either free form or as glycosides. Pharmacologically, coumarins exhibit a broad range of effects, including anti-inflammatory [2], antiviral [3], antibacterial [4], and cardioprotective properties [5], as well as the ability to induce apoptosis in cancer cells [6]. In plants, coumarins play roles in regulating growth and development, influencing processes such as seed germination, modulation of root-associated microbial communities, and enhancing antioxidant activity [7,8,9]. The synthesis and metabolism of coumarins involve complex interactions between multiple structural genes, transcriptional regulators, and environmental factors. Recent advancements in multi-omics technologies have intensified research on the transcriptional regulatory mechanisms governing coumarin biosynthesis genes, often utilizing high-throughput sequencing and integrated multi-omics approaches. This review provides a comprehensive overview of the coumarin biosynthetic pathway in plants, highlights recent discoveries related to enzymes involved in coumarin synthesis, and serves as a valuable reference and theoretical foundation for future research in this field.
2 Origin and Biosynthetic Pathway of Coumarins
Since the 1980s, coumarin biosynthesis has garnered global research interest as a more environmentally friendly and accessible alternative to semi-synthetic or chemical synthesis methods [10]. This growing interest in biosynthetic coumarins underscores a growing awareness of their potential applications and benefits [11]. In the natural synthesis pathway, the conversion of simple coumarins into angular coumarins, furanocoumarins, and pyranocoumarins involves isopentenylation of the 6- or 8-position on the parent carbon atom, followed by modification of the 7-position hydroxyl group to form angular coumarins containing furan or pyran rings. These angular coumarins undergo further transformation into furanocoumarins and pyranocoumarins [12]. In plants, phenylalanine ammonia-lyase (PAL) converts L-phenylalanine into trans-cinnamic acid, whereas cinnamic acid 2-hydroxylase (C2H) and cinnamic acid 4-hydroxylase (C4H) undergo ortho-hydroxylation and para-hydroxylation reactions, respectively, converting to ortho-coumaric acid and para-coumaric acid, respectively, and the subsequent reactions eventually form coumarin compounds or other secondary metabolites. In the coumarin biosynthetic pathway, a para-hydroxylation reaction occurs before ortho-hydroxylation. The pathway begins with the O-hydroxylation of trans-cinnamic acid, which diverges from phenylpropanoid biosynthesis [13]. Trans-cinnamic acid undergoes O-hydroxylation catalyzed by C2H to produce O-coumaric acid. Subsequently, O-coumaric acid reacts with UDP-glucose through the action of 2-coumaric acid O-β-glucosyltransferase, forming trans-coumaric acid β-glucoside. This compound accumulates in the vacuole, where it undergoes cis-trans isomerization. Under ultraviolet radiation, O-coumaric acid β-glucoside participates in a non-enzymatic photochemical reaction and is converted to cis-coumaric acid glucoside. At this stage, coumarin is in a bound (non-free) state. The trans-cinnamic acid pathway can also generate other simple hydroxylated coumarins, such as umbelliferone. Under the catalysis of PAL, phenylalanine generates cinnamic acid, which is the starting precursor of coumarin compounds [13,14]. Cinnamic acid generates p-coumaric acid catalyzed by C4H, and p-coumaric acid generates 2′,4′-dihydroxycinnamoyl coenzyme A catalyzed by 4-coumarate coenzyme ligase (4CL) and coumaryl coenzyme A hydroxylase (C2′H), subsequently forming umbelliferone (7-hydroxycoumarin) [15,16]. Umbelliferone is a precursor of esculetin, which is converted by 6-hydroxylation. In addition, p-coumaric acid generates caffeic acid and caffeoyl-CoA in sequence catalyzed by coumaroyl-3-hydroxylase (C3H), finally producing esculetin. Caffeic acid generates 6-hydroxyferuloyl-CoA through caffeic acid-O-methoxytransferase (COMT), 4CL, and feruloyl-coenzymeA-6′-hydroxylase (F6′H), while 6-Hydroxyferuloyl-CoA generates scopoletin catalyzed by coumarin synthase (COSY). Esculetin can also generate scopoletin catalyzed by O-methoxy transferase (OMT) and fraxetin catalyzed by scopoletin-8-hydroxylase (S8H). Pyranocoumarins and furanocoumarins are key representatives of tetrahydropyrans and tetrahydrofurans, respectively. Multi-omics analysis of the biosynthetic enzymes in P. praeruptorum revealed two types of enzyme that are essential for the biosynthesis of the pyran and furan rings in Apiaceae plants. These included three distinct prenyltransferases, which were responsible for the prenylation of the simple coumarin skeleton 7 into linear or angular precursors, and two novel CYP450 cyclases (namely DC and OC) which were crucial for the cyclization of the linear/angular precursors into either tetrahydropyran or tetrahydrofuran scaffolds. Biochemical characterization of cyclases revealed that acid/base-assisted epoxide ring opening plays a role in the enzyme-catalyzed tetrahydropyran and tetrahydrofuran ring refactoring process. The catalytic process has been proven to be facilitated by the acidic amino acids Glu303 in PpDC and Asp301 in PpOC [17]. Recently, researchers used the genomes and transcriptomes of over 30 Apiaceae plants to conduct phylogenetic analyses and reconstruct the evolutionary history of complex coumarins (CCs). p-Coumaroyl-CoA 2′-hydroxylase (C2′H), C-prenyltransferase (C-PT) and cyclase evolved concurrently at distinct points in the evolution of the Apiaceae family through ectopic and tandem duplications. Neofunctionalization endowed these enzymes with novel roles that were necessary for CC biosynthesis. C-PTs are considered as the key determinant of CC structural diversity due to their ability to target different positions in umbelliferone [18] (Fig. 1).
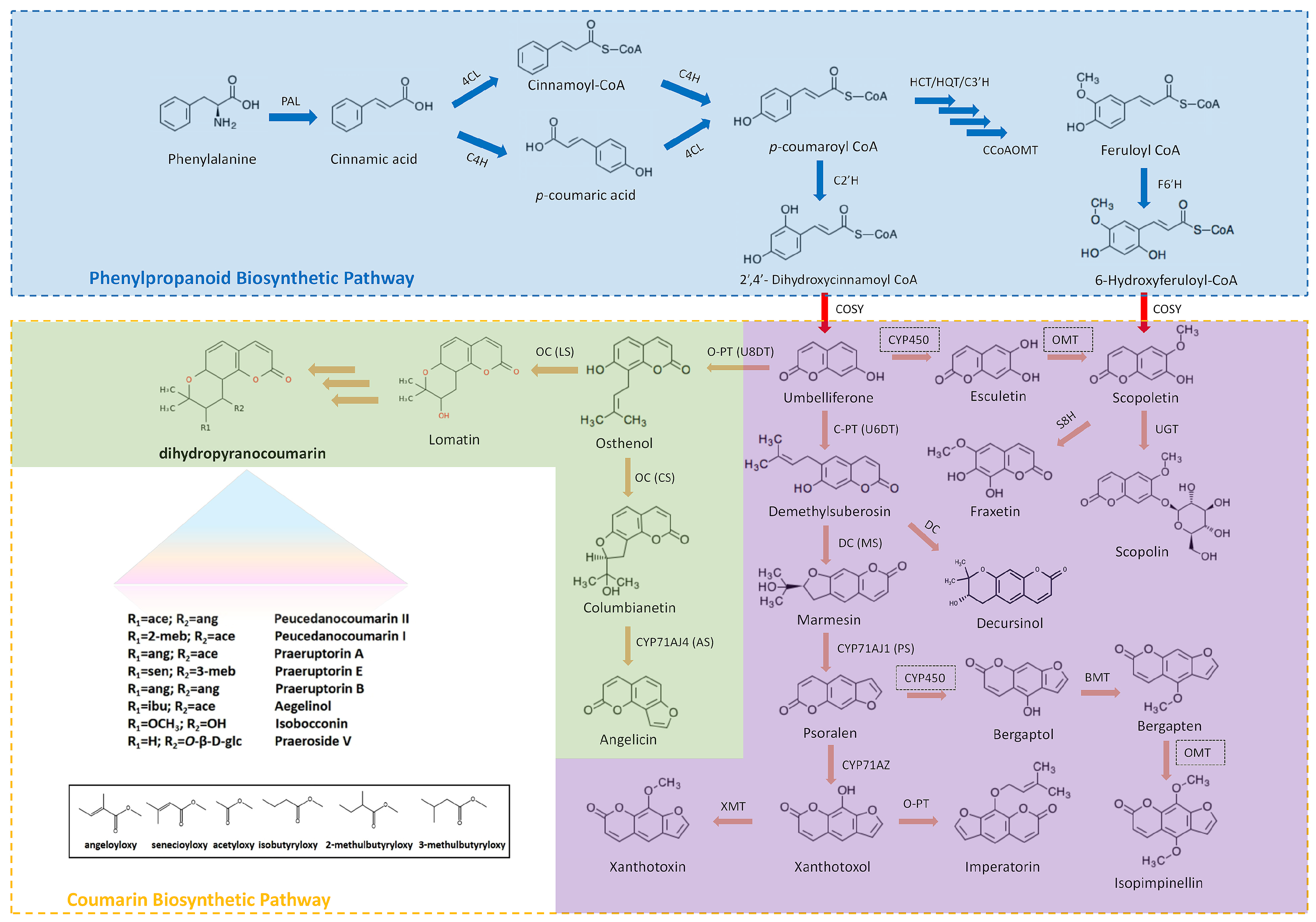
Figure 1: Biosynthesis of different types of coumarins and key catalytic enzymes.
The blue region indicates the phenylpropane biosynthesis pathway and the key enzymes. The purple region indicates the biosynthesis pathways for simple and linear coumarins. The green region indicates the biosynthesis pathways for complex and angular coumarins, including representative coumarin types in Peucedanum praeruptorum. PAL, phenylalanine ammonia lyase; C4H, cinnamate 4-hydroxylase; 4CL, 4-coumarate: coenzyme A ligase; HCT/HQT, hydroxycinnamoyl CoA shikimate/quinate hydroxycinnamoyl transferase; CCoAOMT, caffeoylCoA O-methyltransferase; C3′H, cinnamoyl ester 3′-hydroxylase; C2′H, p-coumaroyl CoA 2′-hydroxylase; F6′H, feruloyl-CoA 6′-hydroxylase; DC, demelthyisuberosin cyclase; OC, osthenol cyclase; COSY, coumarin synthase, U6PT, umbelliferone 6-prenyltransferase, U8PT, umbelliferone 8-prenyltransferase; MS, marmesin synthase; PS, psoralen synthase; CS, columbianetin synthase; AS, angelicin synthase; LS, lomatin synthase; XMT, xanthomol-O-methyltransferase; BMT, bergaptol-O-methyltransferase; S8H, scopoletin 8-hydroxylase.
3 Specific Functions and Catalytic Mechanisms of Key Enzymes in Coumarin Biosynthesis
Coumarins are primarily derived from the phenylpropanoid metabolic pathway and accumulate as secondary metabolites in various plant organs and tissues. Their differential accumulation is closely linked to the expression of key biosynthetic genes [19]. Advances in multi-omics approaches—including genomics, transcriptomics, proteomics, and synthetic biology—have enabled the identification of numerous genes involved in coumarin synthesis. Specific genes have been identified in plants such as Morus alba [20], Angelica dahurica [21], Cnidium monnieri [22], Pediculus alba [23], etc., furthering our understanding of coumarin biosynthesis across diverse species. Transcriptome sequencing analysis of five Trichosanthes meliloti samples showed significant differences in coumarin content, and the authors identified 213 unigenes belonging to the phenylalanine synthesis pathway based on the Kyoto Encyclopedia of Genes and Genomes (KEGG) classification of differentially expressed genes (DEGs). Seventeen unigenes were differentially expressed among different species, including genes related to the coumarin synthesis pathway, such as PAL, C4H, 4CL, C3H, HCT, CCoAOMT, COMT, and BGLU [23].
In plant kingdom, PAL genes are predominantly organized into gene families, with distinct subfamilies exhibiting varied expression patterns in response to environmental factors. PAL is also the first rate-limiting enzyme in the phenylpropanoid pathway and catalyzes the non-oxidative deamination of L-Phe to produce trans-cinnamic acid. PALs were first discovered in Hordeum vulgare and have been found in various plants [24]. PAL family genes have been identified in Arabidopsis, Populus, banana, rice, and walnut [25,26,27,28,29]. The overexpression of LjPAL1 delays the infection process and reduces the number of nodules formed Mesorhizobium loti infection in Lotus japonicus [30]. Expression levels of PAL genes were upregulated in wheat cultivars resistant to Fusarium graminearum [31]. Overexpression of AevPAL1 has been shown to significantly enhance resistance to the cereal cyst nematode in bread wheat [32]. PAL2 regulates secondary metabolism and confers manganese tolerance in Stylosanthes guianensis [33]. CsPAL9 and CsPAL7 were examined under heat stress for six hours at the transcriptome level, suggesting a potential role in cucumber’s tolerance to heat stress [34]. In Arabidopsis thaliana, AtPAL1, AtPAL2, and AtPAL4 are closely associated with lignin biosynthesis, with AtPAL1 and AtPAL2 also regulating total flavonoid levels [19]. A study on Peucedanum praeruptorum identified multiple genes linked to coumarin content, revealing a strong correlation between PAL gene expression and coumarin levels, suggesting that PAL expression influences the entire downstream of coumarin biosynthesis pathway [35]. Under salt stress, PAL expression increases within a few hours of exposure [36]. Beyond plants, the role of PAL has also been widely investigated in various microorganisms, which leverage phenylalanine metabolism to enter the glucose-to-caffeic acid pathway, synthesizing p-coumaric acid via the PAL enzyme and converting it to caffeic acid through a 3′-hydroxylation reaction [37]. Additionally, 4CL is a key enzyme in the phenylpropanoid pathway, with Class I 4CL involved in lignin synthesis and Class II 4CL in flavonoid synthesis [38]. Three paralogous genes, namely 4CL1, 4CL7, and 4CL10, have been isolated from P. praeruptorum. Pp4CL1 can regulate the p-coumaric acid and ferulic acid pathways and catalyze umbelliferone and scopoletin biosynthesis [39]. The 4CL and F6′H1 genes were found to be expressed in Saccharomyces cerevisiae, enabling the production of scopoletin and fraxetin from ferulic acid [40].
C4H belongs to the cytochrome P450 family and catalyzes the conversion of cinnamic acid to p-coumaric acid. RNA-seq was used to analyze the differences in the expression of phenylpropanoid and flavonoid biosynthesis genes between flowers of different colors and to screen three C4H genes, namely HaC4H1, HaC4H2, and HaC4H3. All three genes were successfully expressed in E. coli and catalyzed the conversion of cinnamic acid to p-coumaric acid. The function of the C4H gene was verified in P. praeruptorum and Angelica decursiva, confirming its ability to convert cinnamic acid to p-coumaric acid [41]. F6′H is a member of the monooxygenase family within the cytochrome P450 superfamily and catalyzes the o-hydroxylation of feruloyl-CoA to produce 6-hydroxyferuloyl-CoA. This enzyme was first discovered when studying scopoletin biosynthesis in A. thaliana, where AtF6′H1 mediates the o-hydroxylation of ferulic acid CoA to yield scopoletin and scopolamine, two of the most abundant coumarins in A. thaliana [42,43]. Subsequent research established that the F6′H enzyme specifically exhibits 6′-hydroxylase activity for feruloyl-CoA, which helped clarify the biosynthetic pathway of coumarin compounds [44]. BGLU is the ligase responsible for the synthesis of simple coumarins. It catalyzes the β-glucosidation of monosaccharides with coumarin nuclei, resulting in the formation of biologically active coumarin glycosides. These compounds play essential roles in critical physiological processes in plants, such as cell wall lignification and secondary metabolism [45,46]. In A. thaliana, AtBGLU21–23 not only control the hydrolysis of scopoletin produced in the roots but also contribute to the plant’s defense against pathogen attacks and abiotic stresses. Additionally, AtBGLU42 is involved in inducing resistance to bacterial diseases and promoting the secretion of linear coumarins in the rhizosphere under iron deficiency conditions [47]. O-methylation and oxygen methylation are two of the most important processes in coumarin biosynthesis; however, their underlying mechanisms remain poorly understood [48]. In Ammi majus, a specific bergaptol O-methyltransferase (BMT) was successfully cloned. Due to the strict substrate specificity of BMT, it has been predicted that at least one unknown enzyme is involved in the O-methylation of other hydroxylated coumarins, hinting at a potential pathway for the O-methoxylation of aescin and other hydroxylated coumarins [49]. Another enzyme, caffeic acid-O-methyltransferase (COMT-S) identified in P. praeruptorum participated in O-methylation of coumarin synthesis reactions was elucidated, shedding light on its catalytic mechanism [50].
COSY, a member of the BAHD-acyltransferase family, is responsible for the cis-trans isomerization and lactonization of umbelliferone [51]. Kim et al. elucidated the catalytic mechanism of COSY lactonization [52]. Vanholme et al. were the first to identify COSY in A. thaliana, demonstrating that it is a key enzyme in umbelliferone and scopolamine biosynthesis [51]. Prior to the discovery of COSY, from the conversion of o-hydroxycinnamoyl to coumarin was thought to occur spontaneously through photocatalysis. Study has shown that COSY plays an important role in coumarin synthesis in light-protected tissues such as roots, providing significant insights into the biosynthetic pathways of endogenous coumarins in plants. Recently, studies on the umbelliferone production pathway in engineered yeast showed that COSY plays a vital role in the conversion of 2′,4′-dihydroxycoumaroyl CoA to umbelliferone [53]. UDP glucosyltransferase (UGT), a member of the type I glycosyltransferase family, primarily facilitates the glycosylation of coumarin compounds, enhancing their hydrolysis and functional activity [54]. A novel glycosyltransferase isolated from Cistanche tubulosa, known as CtUGT1, catalyzes the glycosylation of hydroxycoumarins, effectively converting scopoletin to scopolamine [55]. In Melilotus officinalis, MaUGT68 and MaUGT186 not only participate in the glycosylation of coumarin compounds but also contribute to the synthesis of various stress response metabolites, enhancing the plant’s resistance to abiotic stress [56]. The UGT family generally share a common C-terminal domain and exhibit a degree of substrate promiscuity. The presence of a distinct substrate-binding domain endows UGTs with certain substrate specificity. For instance, UGT84A1 and UGT84A3 exhibit hydroxycinnamic acid glucosyltransferase activity [57]. AtUGT84A1 has a higher affinity for caffeic acid and p-coumaric acid, while AtUGT84A3 shows the strongest activity in binding ferulic acid [58].
4 Multi-Omics Technology for Identifying Active Ingredients in Medicinal Plants
Multi-omics technology integrates, and correlates two or more omics approaches to explore interactions among various molecules within biological systems. This comprehensive methodology includes genomics, epigenomics, transcriptomics, proteomics, metabolomics, and microbiome analyses. Together, these molecular dimensions influence and determine the phenotypes, traits, and behaviors of living organisms. In medicinal plants, multi-omics technologies can be used to detect cellular changes at the transcriptional, translational, and metabolic levels throughout their growth and development. Consequently, multi-omics joint analysis provides a comprehensive perspective on the dynamic changes that occur at the gene and protein levels during plant development. This approach enhances our understanding of mechanisms regulating metabolism in medicinal plants, facilitates the optimization of metabolite content, and supports the cultivation of superior varieties [59]. With China’s increasing investment and innovation in traditional Chinese medicine, particularly through major new drug initiatives, several high-level research platforms have been established. This has led to the emergence of numerous groundbreaking technologies. Notably, in 2015, journal Science published a special issue titled “The Art and Science of Traditional Chinese Medicine,” which introduced the concept of “Herbgenomics”. This field leverages existing multi-omics technologies to investigate genetic information and regulatory networks of herbs, aiming to elucidate their molecular mechanisms for preventing and treating human diseases. Herbgenomics effectively integrates traditional Chinese medicine with cutting-edge omics research, promoting the application of these technologies within the field. Its applications extend to medicinal plant model research, gene-assisted breeding, herbal synthetic biology, genetic resource protection and utilization, quality assessment and control of traditional Chinese medicine, and the development of traditional Chinese medicine [60]. In recent years, advancements in multi-omics technology have rapidly matured, significantly advancing the research of traditional Chinese medicine resources. The following section primarily outlines the research scope and application scenarios of genomics, proteomics, transcriptomics, and metabolomics in medicinal plants.
Genomics focuses on genes and chromosomes, emphasizing aspects such as gene structure, function, evolution, mapping, epigenomics, and genome editing [61]. It plays an essential role in elucidating genetic variations in plant genes, enhancing the efficiency of plant breeding, and serving as a crucial tool for plant genetic improvement. Genetic information significantly influences the types and yields of secondary metabolites in medicinal plants. The primary applications of genomics in the analysis of biosynthetic pathways include identification of putative biosynthetic gene clusters, expansion and contraction of gene families, and characterization of unique biosynthetic pathways that result from specialized evolutionary adaptations. By conducting comparative analyses of genome sequences from model plants, researchers can isolate sequences specific to target medicinal plants, thereby enhancing our understanding of their biosynthetic potential. When comparing species within the same genus, analyzing genomic data across different individuals and groups can enhance the efficiency of gene element screening and provide valuable references for functional research on new gene families. The transcriptome encompasses all RNA transcribed in a specific cell type, tissue, organ, or group of cells at a given developmental stage, including both mRNA and non-coding RNA [62,63,64]. The concept of the transcriptome was initially proposed to study gene expression in yeast [63]. In recent years, its application has expanded to life sciences, metabolic engineering, and medicinal plants. Early transcriptome sequencing technologies, such as GeneChip and Expressed Sequence Tags (EST), provided insights into gene expression levels in plants but were limited to species with complete whole-genome sequencing and annotated genes. With advancements in sequencing technology, next-generation sequencing has become the predominant method for transcriptome analysis. RNA-Seq can be utilized for species without complete genome maps, offering high throughput, accuracy, a wide detection range, and cost-effectiveness [65]. Recently, full-length transcriptome technology has emerged based on PacBio single-molecule real-time sequencing, allowing for more comprehensive transcript information and annotations, as well as the discovery of novel splice variants and genes. In medicinal plants, transcriptome data can be used to predict and analyze new transcripts, aiding in the identification of key regulatory genes associated with important plant traits or phenotypes.
Metabolomics primarily focuses on the metabolites produced by biological activities. Metabolites are the final products expressed by organisms, providing an intuitive representation of life processes through dynamic changes. Depending on the experimental purpose, researchers can choose from targeted metabolomics, non-targeted metabolomics, and broadly targeted metabolomics. When transcriptome and proteome analyses cannot fully explain complex regulatory activities, metabolite analysis can quantitatively reflect the physiological processes occurring in cells, tissues, and organs in real-time. In studying the transcriptional regulatory mechanism of coumarin biosynthetic genes, metabolomics can identify the metabolites and metabolic pathways related to coumarin synthesis by examining patterns of metabolite changes and predicting metabolic pathways. Ultra-high-performance liquid chromatography coupled with mass spectrometry (UHPLC-MS) has been employed to quantitatively detect secondary metabolites in A. thaliana. This analysis revealed significant changes in the content of simple coumarins (such as scopoletin, umbelliferone, and aescin) and their glycosides (including scopoletin, apodipine, and aescin), thereby enhancing our understanding of coumarin accumulation in A. thaliana [64]. Additionally, UPLC-MS technology has been utilized to analyze and identify the components of P. praeruptorum and A. decursiva, determining the main differential metabolites and conducting a quantitative analysis. This approach identified a total of 1085 major differential metabolites across 12 categories, with coumarins showing a significant difference, accounting for 10.1% of the total. Differential metabolites were predominantly enriched in the key metabolic pathways of phenolic acids, fatty acids, and other components [56]. Proteomics focuses on the proteome, studying the composition and regulation of proteins at the organismal or cellular level to gain a deeper understanding of various physiological and molecular mechanisms [65]. The proteome encompasses all proteins expressed by a genome, cell, or tissue, and varies with different tissues and environmental conditions. Researchers screen differentially expressed proteins by comparing proteomes from samples under varying conditions or treatments, integrating this information with transcriptome and metabolome data to address biological questions. This integration has become a popular research topic. Proteomics is widely used in medicinal plant research, often employed to analyze the metabolic pathways of bioactive compounds and study the pharmacological effects of active ingredients, thus promoting the effective utilization of medicinal plants [66,67]. For understanding the transcriptional regulation mechanism of coumarin biosynthesis genes, proteomics is frequently used to analyze post-translational modifications and interactions, identifying post-translational modification sites and predicting protein interactions. The comprehensive proteomic profiling of Artemisia annua, coupled with a quantitative, targeted sub-proteomic analysis of two chemotypes—one exhibiting high artemisinin content and the other low—yielded the identification of over 13,000 proteins via the genome sequence annotation database and 182 proteins through mass spectrometry. Mass spectrometry-based proteomics elucidated the trichome machinery of A. annua and its contribution to artemisinin synthesis [68]. Through a multi-omics analysis that combines metabolomics, transcriptomics, and ATAC-seq techniques, researchers identified several CYP450 enzymes, specifically CYP71AZ18 and CYP71AZ19/CYP83F95, which are responsible for the hydroxylation of psoralen. It was found that three CYP450 genes demonstrated strong signals of accessible chromatin regions, which illustrates the importance of chromatin accessibility in the biosynthesis of furocoumarins [69]. Monoterpene indole alkaloids (MIAs) are a family of natural products with anticancer activities. ScRNA-seq demonstrated the cell-type-specific expression of MIA biosynthetic pathways in the leaves and roots of C. roseus. Two reductases that facilitate the dehydration reaction to produce olivacine alkaloids from bisindole alkaloid precursors were identified when combined with single-cell metabolomics [70]. The genome sequencing and annotation of two Ocimum species reveal a heightened expression of genes involved in the phenylpropanoid and terpenoid biosynthetic pathways, as well as cytochrome P450s and some transcription factors. This discovery offers a novel approach to exploring the biosynthetic pathways of significant medicinal metabolites in relative species. Additionally, Ocimum tenuiflorum produces specialized metabolites with anticancer potential as a defense mechanism, including ursolic acid, oleanolic acid, apigenin, etc. Identifying the genes responsible for the biosynthesis of these anticancer metabolites may facilitate the development of targeted drugs [71].
5 Multi-Omics Analysis of Transcriptional Regulation of Coumarin Biosynthetic Genes
In recent years, an increasing number of studies have applied multi-omics technologies to investigate the transcriptional regulation mechanisms of coumarin biosynthesis genes in Apiaceae species [72]. The complete genome of Angelica dahurica was sequenced and assembled for the first time, resulting in the annotation of 114,310 genes and identification of key enzymes involved in coumarin biosynthesis [19]. Studies have shown that miRNA families have similar regulatory functions in different plants [73]. One study used high-throughput sequencing technology to analyze miRNAs that regulate coumarin biosynthesis genes in five white sweet clover samples with different coumarin contents. A total of 417 known and 76 unknown miRNAs were identified in the five samples. GO and KEGG were used to perform functional annotations on the predicted 4196 target genes, and two candidate HCT genes and three miRNAs were predicted to be involved in the coumarin biosynthetic pathway. Finally, the analysis indicated that genes such as PAL, C4H, HCT, and C3H are involved in coumarin biosynthesis [74]. Clarifying the synthesis, regulation, and physiological functions of metabolites is crucial. Advancements in transcriptome sequencing technology have provided robust support for elucidating biosynthesis and regulating plant metabolite abundance. Simple biological pathways for coumarin synthesis have been proposed in Cnidium monnieri and A. dahurica [75]. Additionally, the first chromosome-level genome of A. sinensis has been assembled [76]. Through joint analysis of the transcriptome and metabolome, 46 genes and 174 candidate transcription factors regulating simple coumarins in A. sinensis were identified, contributing to a more comprehensive understanding of the biosynthetic pathway and offering new insights into the molecular mechanisms and evolution of coumarin biosynthesis. Phylogenetic genomics revealed that PT gene, which is involved in the biosynthesis of furanocoumarins in Moraceae, Leguminosae, Rutaceae, and Apiaceae has undergone independent evolutionary processes, providing a valuable model for studying the evolutionary mechanisms of plant defense. Significant variations in the M. officinalis coumarin revealed multiple unigenes enriched in the phenylalanine pathway of coumarin biosynthesis, including PAL, C4H, 4CL, and C3H [77]. Two near-isogenic lines of M. officinalis were analyzed. First, the members of the UGT gene family in M. officinalis were identified at the whole-genome level, and their expression patterns were examined in conjunction with the transcriptomic data. During the functional analyses of the genes, we also explored the upstream transcriptional regulatory mechanisms, establishing a molecular basis for the regulation of scopoletin biosynthesis [78]. Furthermore, metabolomics and transcriptomics techniques were employed to analyze Arabidopsis mutants and overexpression lines, focusing on differences in gene structures related to primary metabolism and phenylpropanoid metabolism among the myb99, myb21-5, and myb24-5 mutants [79]. MYB99 is part of a regulatory triangle with MYB21 and MYB24, which are orthologs of the benzenoid emission genes I and II. Together with ODORANT1 (ODO1), these genes govern the production of scent in petunias. The identification of orthologous protein triads that produce similar secondary metabolites suggests the existence of analogous regulatory modules in other plants, which function to regulate different branches of the complex phenylpropanoid pathway.
The primary strategy for integrating proteomics and metabolomics analyses involves identifying significantly enriched pathways of differentially expressed proteins and metabolites using KEGG pathway annotation information. This allows for the selection of relevant pathways, followed by correlation analysis with transcriptomic data. Ultimately, key genes exhibiting significant differences at the transcriptional, protein, and metabolic levels were identified. Through the integration of transcriptome and proteome analyses, this study demonstrated that coumarin synthesis in mulberry is induced under salt stress, indicating that phenylpropanoid biosynthesis is involved in the plant’s response to salt stress. Additionally, genes and proteins related to plant hormone signal transduction and phenylpropanoid biosynthesis showed a positive correlation with the salt stress response [80]. In another study, the effects of ultraviolet light on the leaves of Clematis paniculata were analyzed to reveal its physiological mechanism at the protein level, leading to the discovery of the enzyme K18. Following UV-B induction, coumarin compounds appeared in the leaves, likely due to K18’s regulatory role. Subsequent studies could elucidate the biosynthetic mechanism of coumarin compounds in C. paniculata leaves resulting from UV induction at the protein level. The biosynthetic regulatory network of scopoletin in Melilotus alba, an important leguminous plant, has garnered significant attention. Using multi-omics analysis combined with in vitro protein interaction experiments, researchers found that scopoletin content correlated with the expression pattern of MabHLH11. MabHLH11 interacts with MaMYB4, and co-expression of these two genes promotes increased scopoletin accumulation [81]. This indicates that MabHLH11 positively regulates scopoletin accumulation, clarifying the molecular mechanism by which MaMYB4 and MabHLH11 interact to enhance scopoletin biosynthesis. The combination of transcriptomics and microbiome studies has been extensively employed to investigate the transcriptional regulation of coumarin biosynthesis. Host-endophyte interactions in secondary metabolite production represent a newly emerging field of research. Endophytes are often capable of producing defense compounds that mimic plant metabolites, with coumarins being a significant class due to their multifunctional biological properties. In cucumber, p-coumaric acid, a precursor to coumarin synthesis, is secreted as an autotoxin. The rhizosphere bacterial and fungal communities of cucumber seedlings treated with p-coumaric acid were analyzed using 16S rRNA gene sequencing and high-throughput sequencing to assess the feedback effects of the rhizosphere on seedling growth. The results indicated that p-coumaric acid reduced bacterial diversity while increasing fungal diversity and altering the composition of both bacterial and fungal communities [82]. Additionally, metagenomic sequencing was performed on endophytic bacteria and fungi in 35 samples of P. praeruptorum before and after bolting, identifying 2624 bacterial and 4281 fungal species, which significantly enriched the species and resources of endophytes associated with P. praeruptorum [83]. Studies exploring the diversity and differences among microorganisms and endophytes in P. praeruptorum provide deeper insights into the mechanism of plant-microorganism interactions and offer new avenues for investigating the coumarin synthesis pathway [84]. In our study, we have undertaken broad investigations into the biosynthetic pathways and transport mechanisms of coumarins in P. praeruptorum. We completed the first chromosome-level genome assembly using PacBio HiFiasm and Oxford Nanopore Technologies (ONT) sequencing, revealing the specific role of whole-genome duplication and tandem duplication events in the expansion of genes related to coumarin and terpenoid biosynthesis [23]. Heretofore, we also investigated the biological characteristics and bolting phenomenon of P. praeruptorum [85]. Through a series of physiological and biochemical studies, microscopic observations, and integrated transcriptomic and metabolomic analyses, we identified key gene families involved in coumarin synthesis and transport [86]. For instance, PT catalyzes the isoprenylation of coumarins, setting the stage for subsequent cyclization. ABCC and ABCG subfamilies in ABC transporter genes likely mediate the transmembrane transport of coumarins [87,88,89].
Certain secondary metabolites have been identified as signaling molecules in plant-microbe symbiosis, influencing the composition of host-associated microbiomes. In medicinal plants, endophyte-induced metabolic reprogramming is a common phenomenon. Coumarin, a secondary metabolite synthesized via the phenylpropanoid pathway, is prevalent in plant rhizospheres and plays a crucial role in the response of dicotyledonous plants to iron deficiency [54]. Under iron stress, coumarin is secreted to enhance the mobilization of stubborn iron sources, thereby meeting plants’ nutritional needs [90]. Subsequent studies have shown that coumarin can influence the community structure of rhizosphere microorganisms, although the specific mechanisms behind this effect remain unclear. A comparative transcriptome analysis has been conducted to model the role of coumarin in microbial contributions to alleviating iron deficiency in plants. This analysis identified numerous genes associated with iron deficiency responses, regulation, and transport, suggesting that plants secrete coumarins to enhance iron absorption by activating microbial-mediated iron mobilization processes rather than altering the microbial community structure or directly interacting with microorganisms [91]. In addition to affecting the composition of the rhizosphere microbiome, coumarins exhibit varying toxicities toward beneficial and pathogenic microorganisms [92]. In Arabidopsis mutants lacking the BGLU42 gene, the relative abundance of Proteobacteria increased and the relative abundance of Firmicutes decreased around their roots. While scopolamine inhibits the growth of soil-borne pathogens in roots, it does not affect rhizosphere bacteria. Our recent study indicated that endophytic fungi coordinate the early bolting and coumarin accumulation in P. praeruptorum. Penicillium restrictum demonstrated superior bolting retardation, enhancing plant resilience and prolonging the coumarin accumulation period [93]. Current metabolomics and microbiome studies have predominantly focused on model organisms, leaving gaps in our understanding of the regulatory mechanisms by which microbial communities influence secondary metabolites in host plants. Therefore, investigating the interactions between medicinal plants and their associated microbiomes is essential for elucidating the microbiome’s role in the synthesis of plant secondary metabolites.
The review highlights recent advancements in multi-omics technology related to key enzyme genes and transcriptional regulation within the coumarin biosynthetic pathway. It underscores the critical role of multi-omics in elucidating the mechanisms behind plant growth, development, and stress tolerance, thereby offering innovative strategies for studying coumarin biosynthesis regulation. While research has primarily focused on cloning and identifying individual genes in this pathway, the specific roles of newly discovered genes remain unclear, and the regulatory mechanisms of target genes—especially when involving multiple genes or transcription factors—are still not well defined. Moreover, genetic transformation systems for high-coumarin content medicinal plants like M. albus, P. praeruptorum, and A. dahurica are underdeveloped, limiting functional validation of these genes and delaying progress in breeding medicinal plant strains with optimized coumarin content. Future research should better integrate multi-omics approaches to thoroughly examine key enzyme genes involved and downstream pathways in coumarin biosynthesis. Following the cloning of relevant regulatory genes, functional verification should be conducted initially in model plants using transgenic technology before reintroduction into the original plant species for further validation. Additionally, advancements in multi-omics methodologies and data processing will be essential for deepening our understanding of the transcriptional regulatory mechanisms governing coumarin-related genes.
Acknowledgement:
Funding Statement: This work was supported by Traditional Chinese Medicine Inheritance and Innovation Research Project of Anhui Province (2025CCCX033), Demonstration Experiment Training Center of Anhui Provincial Department of Education (2022sysx033), and Quality Engineering Project of West Anhui University (wxxy2024011).
Availability of Data and Materials: Not applicable.
Ethics Approval: Not applicable.
Conflicts of Interest: The author declares no conflicts of interest.
References
1. Song J , Luo H , Xu Z , Zhang Y , Xin H , Zhu D , et al. Mining genes associated with furanocoumarin biosynthesis in an endangered medicinal plant, Glehnia littoralis. J Genet. 2020; 99( 1): 11. doi:10.1007/s12041-019-1170-6. [Google Scholar] [CrossRef]
2. Kamauchi H , Noji M , Kinoshita K , Takanami T , Koyama K . Coumarins with an unprecedented tetracyclic skeleton and coumarin dimers from chemically engineered extracts of a marine-derived fungus. Tetrahedron. 2018; 74( 23): 2846– 56. doi:10.1016/j.tet.2018.04.033. [Google Scholar] [CrossRef]
3. Hamdy AM , Khaddour Z , Al-Masoudi NA , Rahman Q , Hering-Junghans C , Villinger A , et al. Synthesis of arylated coumarins by Suzuki–Miyaura cross-coupling. Reactions and anti-HIV activity. Bioorganic Med Chem. 2016; 24( 21): 5115– 26. doi:10.1016/j.bmc.2016.08.029. [Google Scholar] [CrossRef]
4. Jia C , Zhang J , Yu L , Wang C , Yang Y , Rong X , et al. Antifungal activity of coumarin against candida albicans is related to apoptosis. Front Cell Infect Microbiol. 2019; 8: 445. doi:10.3389/fcimb.2018.00445. [Google Scholar] [CrossRef]
5. Wei B , Zhou J , Xu JJ , Cui J , Ping FF , Ling JJ , et al. Discovery of coumarin-derived imino sulfonates as a novel class of potential cardioprotective agents. Eur J Med Chem. 2019; 184: 111779. doi:10.1016/j.ejmech.2019.111779. [Google Scholar] [CrossRef]
6. Albratty M , El-Sharkawy KA , Alam S . Synthesis and antitumor activity of some novel thiophene, pyrimidine, coumarin, pyrazole and pyridine derivatives. Acta Pharm. 2017; 67( 1): 15– 33. doi:10.1515/acph-2017-0004. [Google Scholar] [CrossRef]
7. Chen BX , Peng YX , Gao JD , Zhang Q , Liu QJ , Fu H , et al. Coumarin-induced delay of rice seed germination is mediated by suppression of abscisic acid catabolism and reactive oxygen species production. Front Plant Sci. 2019; 10: 828. doi:10.3389/fpls.2019.00828. [Google Scholar] [CrossRef]
8. Stringlis IA , Yu K , Feussner K , de Jonge R , Van Bentum S , Van Verk MC , et al. MYB72-dependent coumarin exudation shapes root microbiome assembly to promote plant health. Proc Natl Acad Sci U S A. 2018; 115( 22): E5213– 22. doi:10.1073/pnas.1722335115. [Google Scholar] [CrossRef]
9. Stassen MJJ , Hsu SH , Pieterse CMJ , Stringlis IA . Coumarin communication along the microbiome–root–shoot axis. Trends Plant Sci. 2021; 26( 2): 169– 83. doi:10.1016/j.tplants.2020.09.008. [Google Scholar] [CrossRef]
10. Qian B , Váradi L , Trinchi A , Reichman SM , Bao L , Lan M , et al. The design and synthesis of fluorescent coumarin derivatives and their study for Cu2+ sensing with an application for aqueous soil extracts. Molecules. 2019; 24( 19): 3569. doi:10.3390/molecules24193569. [Google Scholar] [CrossRef]
11. Hara KY , Araki M , Okai N , Wakai S , Hasunuma T , Kondo A . Development of bio-based fine chemical production through synthetic bioengineering. Microb Cell Fact. 2014; 13( 1): 173. doi:10.1186/s12934-014-0173-5. [Google Scholar] [CrossRef]
12. Wang Y , Guan T , Yue X , Yang J , Zhao X , Chang A , et al. The biosynthetic pathway of coumarin and its genetic regulation in response to biotic and abiotic stresses. Front Plant Sci. 2025; 16: 1599591. doi:10.3389/fpls.2025.1599591. [Google Scholar] [CrossRef]
13. Zou Y , Teng Y , Li J , Yan Y . Recent advances in the biosynthesis of coumarin and its derivatives. Green Chem Eng. 2024; 5( 2): 150– 4. doi:10.1016/j.gce.2023.04.003. [Google Scholar] [CrossRef]
14. Poulton JE , McRee DE , Conn EE . Intracellular localization of two enzymes involved in coumarin biosynthesis in Melilotus alba. Plant Physiol. 1980; 65( 2): 171– 5. doi:10.1104/pp.65.2.171. [Google Scholar] [CrossRef]
15. Vialart G , Hehn A , Olry A , Ito K , Krieger C , Larbat R , et al. A 2-oxoglutarate-dependent dioxygenase from Ruta graveolens L. exhibits p-coumaroyl CoA 2′-hydroxylase activity (C2′H): A missing step in the synthesis of umbelliferone in plants. Plant J. 2012; 70( 3): 460– 70. doi:10.1111/j.1365-313X.2011.04879.x. [Google Scholar] [CrossRef]
16. Hijazin T , Radwan A , Abouzeid S , Dräger G , Selmar D . Uptake and modification of umbelliferone by various seedlings. Phytochemistry. 2019; 157: 194– 9. doi:10.1016/j.phytochem.2018.10.032. [Google Scholar] [CrossRef]
17. Zhao Y , He Y , Han L , Zhang L , Xia Y , Yin F , et al. Two types of coumarins-specific enzymes complete the last missing steps in pyran- and furanocoumarins biosynthesis. Acta Pharm Sin B. 2024; 14( 2): 869– 80. doi:10.1016/j.apsb.2023.10.016. [Google Scholar] [CrossRef]
18. Huang XC , Tang H , Wei X , He Y , Hu S , Wu JY , et al. The gradual establishment of complex coumarin biosynthetic pathway in Apiaceae. Nat Commun. 2024; 15( 1): 6864. doi:10.1038/s41467-024-51285-x. [Google Scholar] [CrossRef]
19. Zhao Y , Liu T , Luo J , Zhang Q , Xu S , Han C , et al. Integration of a decrescent transcriptome and metabolomics dataset of Peucedanum praeruptorum to investigate the CYP450 and MDR genes Involved in coumarins biosynthesis and transport. Front Plant Sci. 2015; 6: 996. doi:10.3389/fpls.2015.00996. [Google Scholar] [CrossRef]
20. Rostami Z , Fazeli A , Hojati Z . The isolation and expression analysis of cinnamate 4-hydroxylase and chalcone synthase genes of Scrophularia striata under different abiotic elicitors. Sci Rep. 2022; 12( 1): 8128. doi:10.1038/s41598-022-12361-8. [Google Scholar] [CrossRef]
21. Zhao L , Zhang S , Shan C , Shi Y , Wu H , Wu J , et al. De novo transcriptome assembly of Angelica dahurica and characterization of coumarin biosynthesis pathway genes. Gene. 2021; 791: 145713. doi:10.1016/j.gene.2021.145713. [Google Scholar] [CrossRef]
22. Shi Y , Zhang S , Peng D , Shan C , Zhao L , Wang B , et al. De novo transcriptome analysis of Cnidium monnieri (L.) Cuss and detection of genes related to coumarin biosynthesis. PeerJ. 2020; 8: e10157. doi:10.7717/peerj.10157. [Google Scholar] [CrossRef]
23. Song C , Zhang Y , Manzoor MA , Wei P , Yi S , Chu S , et al. A chromosome-scale genome of Peucedanum praeruptorum provide insights into Apioideae evolution and medicinal ingredient biosynthesis. Int J Biol Macromol. 2024; 255: 128218. doi:10.1016/j.ijbiomac.2023.128218. [Google Scholar] [CrossRef]
24. Uribe EG , Conn EE . The metabolism of aromatic compounds in higher plants. J Biol Chem. 1966; 241( 1): 92– 4. doi:10.1016/S0021-9258(18)96962-0. [Google Scholar] [CrossRef]
25. Huang J , Gu M , Lai Z , Fan B , Shi K , Zhou YH , et al. Functional analysis of the Arabidopsis PAL gene family in plant growth, development, and response to environmental stress. Plant Physiol. 2010; 153( 4): 1526– 38. doi:10.1104/pp.110.157370. [Google Scholar] [CrossRef]
26. Shi R , Shuford CM , Wang JP , Sun YH , Yang Z , Chen HC , et al. Regulation of phenylalanine ammonia-lyase (PAL) gene family in wood forming tissue of Populus trichocarpa. Planta. 2013; 238( 3): 487– 97. doi:10.1007/s00425-013-1905-1. [Google Scholar] [CrossRef]
27. Wang Z , Li JY , Jia CH , Li JP , Xu BY , Jin ZQ . Molecular cloning and expression of four phenylalanine ammonia lyase genes from banana interacting with Fusarium oxysporum. Biol Plant. 2016; 60( 3): 459– 68. doi:10.1007/s10535-016-0619-1. [Google Scholar] [CrossRef]
28. Zhan C , Li Y , Li H , Wang M , Gong S , Ma D , et al. Phylogenomic analysis of phenylalanine ammonia-lyase (PAL) multigene family and their differential expression analysis in wheat (Triticum aestivum L.) suggested their roles during different stress responses. Front Plant Sci. 2022; 13: 982457. doi:10.3389/fpls.2022.982457. [Google Scholar] [CrossRef]
29. Yan F , Li H , Zhao P . Genome-wide identification and transcriptional expression of the PAL Gene family in common walnut (Juglans Regia L.). Genes. 2019; 10( 1): 46. doi:10.3390/genes10010046. [Google Scholar] [CrossRef]
30. Chen Y , Li F , Tian L , Huang M , Deng R , Li X , et al. The phenylalanine ammonia lyase gene LjPAL1 is involved in plant defense responses to pathogens and plays diverse roles in lotus japonicus-rhizobium symbioses. Mol Plant Microbe Interact. 2017; 30( 9): 739– 53. doi:10.1094/MPMI-04-17-0080-R. [Google Scholar] [CrossRef]
31. Golkari S , Gilbert J , Ban T , Procunier JD . QTL-specific microarray gene expression analysis of wheat resistance to Fusarium head blight in Sumai-3 and two susceptible NILs. Genome. 2009; 52( 5): 409– 18. doi:10.1139/G09-018. [Google Scholar] [CrossRef]
32. Zhang H , Huang Q , Yi L , Song X , Li L , Deng G , et al. PAL-mediated SA biosynthesis pathway contributes to nematode resistance in wheat. Plant J. 2021; 107( 3): 698– 712. doi:10.1111/tpj.15316. [Google Scholar] [CrossRef]
33. Wang L , Li J , Liu L , Dong R , Liu G , Rao IM , et al. Phenylalanine ammonia-lyase 2 regulates secondary metabolism and confers manganese tolerance in Stylosanthes guianensis. Plant Physiol. 2024; 197( 1): kiaf005. doi:10.1093/plphys/kiaf005. [Google Scholar] [CrossRef]
34. Amjad M , Wang Y , Han S , Haider MZ , Sami A , Batool A , et al. Genome wide identification of phenylalanine ammonia-lyase (PAL) gene family in Cucumis sativus (cucumber) against abiotic stress. BMC Genom Data. 2024; 25( 1): 76. doi:10.1186/s12863-024-01259-1. [Google Scholar] [CrossRef]
35. Sui Z , Luo J , Yao R , Huang C , Zhao Y , Kong L . Functional characterization and correlation analysis of phenylalanine ammonia-lyase (PAL) in coumarin biosynthesis from Peucedanum praeruptorum Dunn. Phytochemistry. 2019; 158: 35– 45. doi:10.1016/j.phytochem.2018.11.006. [Google Scholar] [CrossRef]
36. Wei P , Li Y , Song C , Manzoor MA , Dai J , Yin Q , et al. Analysis of coumarin content and key enzyme genes expression involved in coumarin biosynthesis from Peucedanum praeruptorum Dunn at different stages. Acta Physiol Plant. 2023; 45( 12): 141. doi:10.1007/s11738-023-03619-3. [Google Scholar] [CrossRef]
37. Li G , Song C , Manzoor MA , Li D , Cao Y , Cai Y . Functional and kinetics of two efficient phenylalanine ammonia lyase from Pyrus bretschneideri. BMC Plant Biol. 2023; 23( 1): 612. doi:10.1186/s12870-023-04586-0. [Google Scholar] [CrossRef]
38. Gui J , Shen J , Li L . Functional characterization of evolutionarily divergent 4-Coumarate: Coenzyme A Ligases in rice. Plant Physiol. 2011; 157( 2): 574– 86. doi:10.1104/pp.111.178301. [Google Scholar] [CrossRef]
39. Liu T , Yao R , Zhao Y , Xu S , Huang C , Luo J , et al. Cloning, functional characterization and site-directed mutagenesis of 4-Coumarate: Coenzyme A Ligase (4CL) involved in coumarin biosynthesis in Peucedanum praeruptorum Dunn. Front Plant Sci. 2017; 8: 4. doi:10.3389/fpls.2017.00004. [Google Scholar] [CrossRef]
40. Zhao CH , Zhang RK , Qiao B , Li BZ , Yuan YJ . Engineering budding yeast for the production of coumarins from lignin. Biochem Eng J. 2020; 160: 107634. doi:10.1016/j.bej.2020.107634. [Google Scholar] [CrossRef]
41. Park CH , Park HW , Yeo HJ , Jung DH , Jeon KS , Kim TJ , et al. Combined transcriptome and metabolome analysis and evaluation of antioxidant and antibacterial activities in white, pink, and violet flowers of Angelica gigas. Ind Crops Prod. 2022; 188: 115605. doi:10.1016/j.indcrop.2022.115605. [Google Scholar] [CrossRef]
42. Kai K , Mizutani M , Kawamura N , Yamamoto R , Tamai M , Yamaguchi H , et al. Scopoletin is biosynthesized via ortho-hydroxylation of feruloyl CoA by a 2-oxoglutarate-dependent dioxygenase in Arabidopsis thaliana. Plant J. 2008; 55( 6): 989– 99. doi:10.1111/j.1365-313X.2008.03568.x. [Google Scholar] [CrossRef]
43. Döll S , Kuhlmann M , Rutten T , Mette MF , Scharfenberg S , Petridis A , et al. Accumulation of the coumarin scopolin under abiotic stress conditions is mediated by the Arabidopsis thaliana THO/TREX complex. Plant J. 2018; 93( 3): 431– 44. doi:10.1111/tpj.13797. [Google Scholar] [CrossRef]
44. Tao M , Liu S , Li Y , Liu A , Tian J , Liu Y , et al. Molecular characterization of a feruloyl-CoA 6′-hydroxylase involved in coumarin biosynthesis in Clematis terniflora DC. Plant Physiol Biochem. 2023; 196: 162– 70. doi:10.1016/j.plaphy.2023.01.046. [Google Scholar] [CrossRef]
45. Ahn YO , Shimizu B , Sakata K , Gantulga D , Zhou Z , Bevan DR , et al. Scopolin-hydrolyzing-glucosidases in roots of Arabidopsis. Plant Cell Physiol. 2010; 51( 1): 132– 43. doi:10.1093/pcp/pcp174. [Google Scholar] [CrossRef]
46. Schmidt S , Rainieri S , Witte S , Matern U , Martens S . Identification of a Saccharomyces cerevisiae glucosidase that hydrolyzes flavonoid glucosides. Appl Environ Microbiol. 2011; 77( 5): 1751– 7. doi:10.1128/AEM.01125-10. [Google Scholar] [CrossRef]
47. Tsai HH , Rodríguez-Celma J , Lan P , Wu YC , Vélez-Bermúdez IC , Schmidt W . Scopoletin 8-Hydroxylase-Mediated fraxetin production is crucial for iron mobilization. Plant Physiol. 2018; 177( 1): 194– 207. doi:10.1104/pp.18.00178. [Google Scholar] [CrossRef]
48. Zhao Y , Wang N , Zeng Z , Xu S , Huang C , Wang W , et al. Cloning, functional characterization, and catalytic mechanism of a Bergaptol O-Methyltransferase from Peucedanum praeruptorum Dunn. Front Plant Sci. 2016; 7: 722. doi:10.3389/fpls.2016.00722. [Google Scholar] [CrossRef]
49. Hehmann M , Lukačin R , Ekiert H , Matern U . Furanocoumarin biosynthesis in Ammi majus L.: Cloning of bergaptol O-methyltransferase. Eur J Biochem. 2004; 271( 5): 932– 40. doi:10.1111/j.1432-1033.2004.03995.x. [Google Scholar] [CrossRef]
50. Zhao Y , Wang N , Sui Z , Huang C , Zeng Z , Kong L . The molecular and structural basis of O-methylation reaction in coumarin biosynthesis in Peucedanum praeruptorum Dunn. Int J Mol Sci. 2019; 20( 7): 1533. doi:10.3390/ijms20071533. [Google Scholar] [CrossRef]
51. Vanholme R , Sundin L , Seetso KC , Kim H , Liu X , Li J , et al. COSY catalyses trans–cis isomerization and lactonization in the biosynthesis of coumarins. Nat Plants. 2019; 5( 10): 1066– 75. doi:10.1038/s41477-019-0510-0. [Google Scholar] [CrossRef]
52. Kim CY , Mitchell AJ , Kastner DW , Albright CE , Gutierrez MA , Glinkerman CM , et al. Emergence of a proton exchange-based isomerization and lactonization mechanism in the plant coumarin synthase COSY. Nat Commun. 2023; 14( 1): 597. doi:10.1038/s41467-023-36299-1. [Google Scholar] [CrossRef]
53. Wang P , Fan Z , Wei W , Yang C , Wang Y , Shen X , et al. Biosynthesis of the plant coumarin osthole by engineered Saccharomyces cerevisiae. ACS Synth Biol. 2023; 12( 8): 2455– 62. doi:10.1021/acssynbio.3c00321. [Google Scholar] [CrossRef]
54. Stringlis IA , De Jonge R , Pieterse CMJ . The age of coumarins in plant–microbe interactions. Plant Cell Physiol. 2019; 60( 7): 1405– 19. doi:10.1093/pcp/pcz076. [Google Scholar] [CrossRef]
55. Xu X , Yan Y , Huang W , Mo T , Wang X , Wang J , et al. Molecular cloning and biochemical characterization of a new coumarin glycosyltransferase CtUGT1 from Cistanche tubulosa. Fitoterapia. 2021; 153: 104995. doi:10.1016/j.fitote.2021.104995. [Google Scholar] [CrossRef]
56. Duan Z , Yan Q , Wu F , Wang Y , Wang S , Zong X , et al. Genome-wide analysis of the udp-glycosyltransferase family reveals its roles in coumarin biosynthesis and abiotic stress in Melilotus albus. Int J Mol Sci. 2021; 22( 19): 10826. doi:10.3390/ijms221910826. [Google Scholar] [CrossRef]
57. Yonekura-Sakakibara K . Functional genomics of family 1 glycosyltransferases in Arabidopsis. Plant Biotechnol. 2009; 26( 3): 267– 74. doi:10.5511/plantbiotechnology.26.267. [Google Scholar] [CrossRef]
58. Cheng C , Krishnakumar V , Chan AP , Thibaud-Nissen F , Schobel S , Town CD . Araport11: A complete reannotation of the Arabidopsis thaliana reference genome. Plant J. 2017; 89( 4): 789– 804. doi:10.1111/tpj.13415. [Google Scholar] [CrossRef]
59. Song C , Zhang Y , Chen R , Zhu F , Wei P , Pan H , et al. Label-free quantitative proteomics unravel the impacts of salt stress on Dendrobium huoshanense. Front Plant Sci. 2022; 13: 874579. doi:10.3389/fpls.2022.874579. [Google Scholar] [CrossRef]
60. Chakraborty P . Herbal genomics as tools for dissecting new metabolic pathways of unexplored medicinal plants and drug discovery. Biochimie Open. 2018; 6: 9– 16. doi:10.1016/j.biopen.2017.12.003. [Google Scholar] [CrossRef]
61. Marian AJ . Sequencing your genome: What does it mean? Methodist DeBakey Cardiovasc J. 2014; 10( 1): 3. doi:10.14797/mdcj-10-1-3. [Google Scholar] [CrossRef]
62. Wang Z , Gerstein M , Snyder M . RNA-Seq: A revolutionary tool for transcriptomics. Nat Rev Genet. 2009; 10( 1): 57– 63. doi:10.1038/nrg2484. [Google Scholar] [CrossRef]
63. Lockhart DJ , Dong H , Byrne MC , Follettie MT , Gallo MV , Chee MS , et al. Expression monitoring by hybridization to high-density oligonucleotide arrays. Nat Biotechnol. 1996; 14( 13): 1675– 80. doi:10.1038/nbt1296-1675. [Google Scholar] [CrossRef]
64. Perkowska I , Siwinska J , Olry A , Grosjean J , Hehn A , Bourgaud F , et al. Identification and quantification of coumarins by UHPLC-MS in Arabidopsis thaliana natural populations. Molecules. 2021; 26( 6): 1804. doi:10.3390/molecules26061804. [Google Scholar] [CrossRef]
65. Abril N , Gion JM , Kerner R , Müller-Starck G , Cerrillo RMN , Plomion C , et al. Proteomics research on forest trees, the most recalcitrant and orphan plant species. Phytochemistry. 2011; 72( 10): 1219– 42. doi:10.1016/j.phytochem.2011.01.005. [Google Scholar] [CrossRef]
66. Hashiguchi A , Tian J , Komatsu S . Proteomic contributions to medicinal plant research: From plant metabolism to pharmacological action. Proteomes. 2017; 5( 4): 35. doi:10.3390/proteomes5040035. [Google Scholar] [CrossRef]
67. Munakata R , Olry A , Takemura T , Tatsumi K , Ichino T , Villard C , et al. Parallel evolution of UbiA superfamily proteins into aromatic O-prenyltransferases in plants. Proc Natl Acad Sci U S A. 2021; 118( 17): e2022294118. doi:10.1073/pnas.2022294118. [Google Scholar] [CrossRef]
68. Chen M , Yan T , Ji L , Dong Y , Sidoli S , Yuan Z , et al. Comprehensive map of the Artemisia annua proteome and quantification of differential protein expression in chemotypes producing high versus low content of artemisinin. Proteomics. 2020; 20( 10): 1900310. doi:10.1002/pmic.201900310. [Google Scholar] [CrossRef]
69. Ji J , Han X , Zang L , Li Y , Lin L , Hu D , et al. Integrative multi-omics data provide insights into the biosynthesis of furanocoumarins and mechanisms regulating their accumulation in Angelica dahurica. Commun Biol. 2025; 8( 1): 649. doi:10.1038/s42003-025-08076-x. [Google Scholar] [CrossRef]
70. Li C , Wood JC , Vu AH , Hamilton JP , Rodriguez Lopez CE , Payne RME , et al. Single-cell multi-omics in the medicinal plant Catharanthus roseus. Nat Chem Biol. 2023; 19( 8): 1031– 41. doi:10.1038/s41589-023-01327-0. [Google Scholar] [CrossRef]
71. Upadhyay AK , Chacko AR , Gandhimathi A , Ghosh P , Harini K , Joseph AP , et al. Genome sequencing of herb Tulsi (Ocimum tenuiflorum) unravels key genes behind its strong medicinal properties. BMC Plant Biol. 2015; 15( 1): 212. doi:10.1186/s12870-015-0562-x. [Google Scholar] [CrossRef]
72. Bai M , Jiang S , Chu S , Yu Y , Shan D , Liu C , et al. The telomere-to-telomere (T2T) genome of Peucedanum praeruptorum Dunn provides insights into the genome evolution and coumarin biosynthesis. GigaScience. 2024; 13: giae025. doi:10.1093/gigascience/giae025. [Google Scholar] [CrossRef]
73. Axtell MJ , Snyder JA , Bartel DP . Common functions for diverse small RNAs of land plants. Plant Cell. 2007; 19( 6): 1750– 69. doi:10.1105/tpc.107.051706. [Google Scholar] [CrossRef]
74. Wu F , Luo K , Yan Z , Zhang D , Yan Q , Zhang Y , et al. Analysis of miRNAs and their target genes in five Melilotus albus NILs with different coumarin content. Sci Rep. 2018; 8( 1): 14138. doi:10.1038/s41598-018-32153-3. [Google Scholar] [CrossRef]
75. Wang S , Xie Y , Huo YW , Li Y , Abel PW , Jiang H , et al. Airway relaxation mechanisms and structural basis of osthole for improving lung function in asthma. Sci Signal. 2020; 13( 659): eaax0273. doi:10.1126/scisignal.aax0273. [Google Scholar] [CrossRef]
76. Han X , Li C , Sun S , Ji J , Nie B , Maker G , et al. The chromosome-level genome of female ginseng (Angelica sinensis) provides insights into molecular mechanisms and evolution of coumarin biosynthesis. Plant J. 2022; 112( 5): 1224– 37. doi:10.1111/tpj.16007. [Google Scholar] [CrossRef]
77. Wu F , Duan Z , Xu P , Yan Q , Meng M , Cao M , et al. Genome and systems biology of Melilotus albus provides insights into coumarins biosynthesis. Plant Biotechnol J. 2022; 20( 3): 592– 609. doi:10.1111/pbi.13742. [Google Scholar] [CrossRef]
78. Han L , Zhang L , He Y , Liao L , Li J , Xu S , et al. Three carbon-/oxygen-prenyltransferases responsible for furanocoumarin synthesis in Angelica dahurica. Ind Crops Prod. 2023; 200: 116814. doi:10.1016/j.indcrop.2023.116814. [Google Scholar] [CrossRef]
79. Battat M , Eitan A , Rogachev I , Hanhineva K , Fernie A , Tohge T , et al. A MYB triad controls primary and phenylpropanoid metabolites for pollen coat patterning. Plant Physiol. 2019; 180( 1): 87– 108. doi:10.1104/pp.19.00009. [Google Scholar] [CrossRef]
80. Yrjönen T , Eeva M , Kauppila TJ , Martiskainen O , Summanen J , Vuorela P , et al. Profiling of coumarins in Peucedanum palustre (L.) Moench populations growing in finland. Chem Biodivers. 2016; 13( 6): 700– 9. doi:10.1002/cbdv.201500198. [Google Scholar] [CrossRef]
81. Duan Z , Wang S , Zhang Z , Yan Q , Zhang C , Zhou P , et al. The MabHLH11 transcription factor interacting with MaMYB4 acts additively in increasing plant scopolin biosynthesis. Crop J. 2023; 11( 6): 1675– 85. doi:10.1016/j.cj.2023.06.011. [Google Scholar] [CrossRef]
82. Zhou X , Zhang J , Pan D , Ge X , Jin X , Chen S , et al. p-Coumaric can alter the composition of cucumber rhizosphere microbial communities and induce negative plant-microbial interactions. Biol Fertil Soils. 2018; 54( 3): 363– 72. doi:10.1007/s00374-018-1265-x. [Google Scholar] [CrossRef]
83. Wang Y , Liao R , Pan H , Wang X , Wan X , Han B , et al. Comparative metabolic profiling of the mycelium and fermentation broth of Penicillium restrictum from Peucedanum praeruptorum rhizosphere. Environ Microbiol Rep. 2024; 16( 3): e13286. doi:10.1111/1758-2229.13286. [Google Scholar] [CrossRef]
84. Liu L , Wang X , Chen S , Liu D , Song C , Yi S , et al. Fungal isolates influence the quality of Peucedanum praeruptorum Dunn. Front Plant Sci. 2022; 13: 1011001. doi:10.3389/fpls.2022.1011001. [Google Scholar] [CrossRef]
85. Song C , Zhang W , Manzoor MA , Sabir IA , Pan H , Zhang L , et al. Differential involvement of PEBP genes in early flowering of Peucedanum praeruptorum Dunn. Postharvest Biol Technol. 2024; 212: 112860. doi:10.1016/j.postharvbio.2024.112860. [Google Scholar] [CrossRef]
86. Song C , Zhang Y , Zhang Y , Yi S , Pan H , Liao R , et al. Genome sequencing-based transcriptomic analysis reveals novel genes in Peucedanum praeruptorum. BMC Genom Data. 2023; 24( 1): 53. doi:10.1186/s12863-023-01157-y. [Google Scholar] [CrossRef]
87. Song C , Li X , Jia B , Liu L , Ou J , Han B . De novo Transcriptome sequencing coupled with co-expression analysis reveal the transcriptional regulation of key genes involved in the formation of active ingredients in Peucedanum praeruptorum Dunn under bolting period. Front Genet. 2021; 12: 683037. doi:10.3389/fgene.2021.683037. [Google Scholar] [CrossRef]
88. Song C , Li X , Jia B , Liu L , Wei P , Manzoor MA , et al. Comparative transcriptomics unveil the crucial genes involved in coumarin biosynthesis in Peucedanum praeruptorum Dunn. Front Plant Sci. 2022; 13: 899819. doi:10.3389/fpls.2022.899819. [Google Scholar] [CrossRef]
89. Liao R , Yao J , Zhang Y , Liu Y , Pan H , Han B , et al. MYB transcription factors in Peucedanum Praeruptorum Dunn: The diverse roles of the R2R3-MYB subfamily in mediating coumarin biosynthesis. BMC Plant Biol. 2024; 24( 1): 1135. doi:10.1186/s12870-024-05864-1. [Google Scholar] [CrossRef]
90. Lefèvre F , Fourmeau J , Pottier M , Baijot A , Cornet T , Abadía J , et al. The Nicotiana tabacum ABC transporter NtPDR3 secretes O-methylated coumarins in response to iron deficiency. J Exp Bot. 2018; 69( 18): 4419– 31. doi:10.1093/jxb/ery221. [Google Scholar] [CrossRef]
91. Harbort CJ , Hashimoto M , Inoue H , Niu Y , Guan R , Rombolà AD , et al. Root-secreted coumarins and the microbiota interact to improve iron nutrition in Arabidopsis. Cell Host Microbe. 2020; 28( 6): 825– 37.e6. doi:10.1016/j.chom.2020.09.006. [Google Scholar] [CrossRef]
92. Voges MJEEE , Bai Y , Schulze-Lefert P , Sattely ES . Plant-derived coumarins shape the composition of an Arabidopsis synthetic root microbiome. Proc Natl Acad Sci U S A. 2019; 116( 25): 12558– 65. doi:10.1073/pnas.1820691116. [Google Scholar] [CrossRef]
93. Song C , Pan H , Wang Y , Liao R , Arif M , Wang H , et al. Penicillium restrictum reshapes root exudates and inhibits Peucedanum praeruptorum bolting through field inoculation and co-cultivation. Plant Pathol. 2025; 74( 7): 2035– 53. doi:10.1111/ppa.70007. [Google Scholar] [CrossRef]
Cite This Article
 Copyright © 2026 The Author(s). Published by Tech Science Press.
Copyright © 2026 The Author(s). Published by Tech Science Press.This work is licensed under a Creative Commons Attribution 4.0 International License , which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.


 Submit a Paper
Submit a Paper Propose a Special lssue
Propose a Special lssue View Full Text
View Full Text Download PDF
Download PDF Downloads
Downloads
 Citation Tools
Citation Tools