 Open Access
Open Access
REVIEW
Holocentric Chromosomes in Plants: Historical Overview, Developments and Challenges
1 Institute of Natural Sciences, Federal University of Alfenas, Alfenas, Brazil
2 Department of Biology, Federal University of Lavras, Lavras, Brazil
* Corresponding Author: Ana Luisa Arantes Chaves. Email:
(This article belongs to the Special Issue: Karyotypic Shifts: Shaping Plant Diversity and Driving Evolutionary Processes)
Phyton-International Journal of Experimental Botany 2026, 95(3), 2 https://doi.org/10.32604/phyton.2026.077185
Received 04 December 2025; Accepted 04 February 2026; Issue published 31 March 2026
Abstract
Holocentric chromosomes, characterized by kinetochore activity distributed along almost their entire length, have attracted growing interest in cytogenetics and evolutionary biology, particularly because of their adaptive significance. This review provides a historical overview of research on holocentricity, from the earliest microscopic observations to recent advances made possible by modern molecular and cytogenetic techniques. We discuss the main hypotheses concerning the origin and evolution of holocentric chromosomes, emphasizing their multiple independent origins across diverse lineages, as well as the potential selective pressures underlying their maintenance and diversification. In addition, we examine the principal methods used for their identification, with particular emphasis on the immunolocalization of CENH3, a centromere-specific variant of histone H3, and discuss its applications, limitations, and recent methodological advances, including the development of universal markers. A comprehensive understanding of the structural and functional diversity of holocentric chromosomes is essential for elucidating the mechanisms of karyotype evolution across taxa and for shedding light on the processes that shaped early terrestrial life.Keywords
Cytogenetics is the branch of biology that investigates chromosomes, which are defined as DNA molecules associated with histone and non-histone proteins and organized at a high level of condensation [1]. The term chromosome was introduced in 1888 by Heinrich Wilhelm Gottfried Waldeyer and derives from the Greek khrôma (“color”) and sôma (“body”), reflecting what early cytologists could observe under light microscopy: intensely stained structures visible during cell division [2]. Cytotaxonomy, in turn, focuses on the classification of organisms through comparative chromosomal analyses and is closely linked to phylogenetic systematics [3]. Key cytogenetic features include chromosome number, morphology, and behavior, the latter primarily examined during meiosis [1,4]. Together, these data provide valuable insights into evolutionary relationships among species and can reinforce taxonomic delimitations based on morphological criteria [2].
Chromosomes can be morphologically characterized by the presence of a primary constriction corresponding to a centromeric unit; such chromosomes are referred to as monocentric [5]. Monocentric chromosomes are commonly classified as metacentric, submetacentric, acrocentric, or telocentric based on measurements of the arm ratio [6] or the centromeric index, which is calculated as the ratio between the length of the short arm and the total chromosome length of the haploid set [7].
On the other hand, holocentric chromosomes exhibit centromeric activity distributed along almost their entire length (Fig. 1) [8,9,10]. They have been reported in protists, invertebrates, and plants, with particularly high representation in the families Cyperaceae, Convolvulaceae, Droseraceae, and Juncaceae [11,12]. In addition, metapolycentric chromosomes have been identified in some species of Pisum L. and Lathyrus L. (Fabaceae). These chromosomes are characterized by elongated primary constrictions and are considered to represent an intermediate state between monocentric and holocentric chromosomal organizations [5].
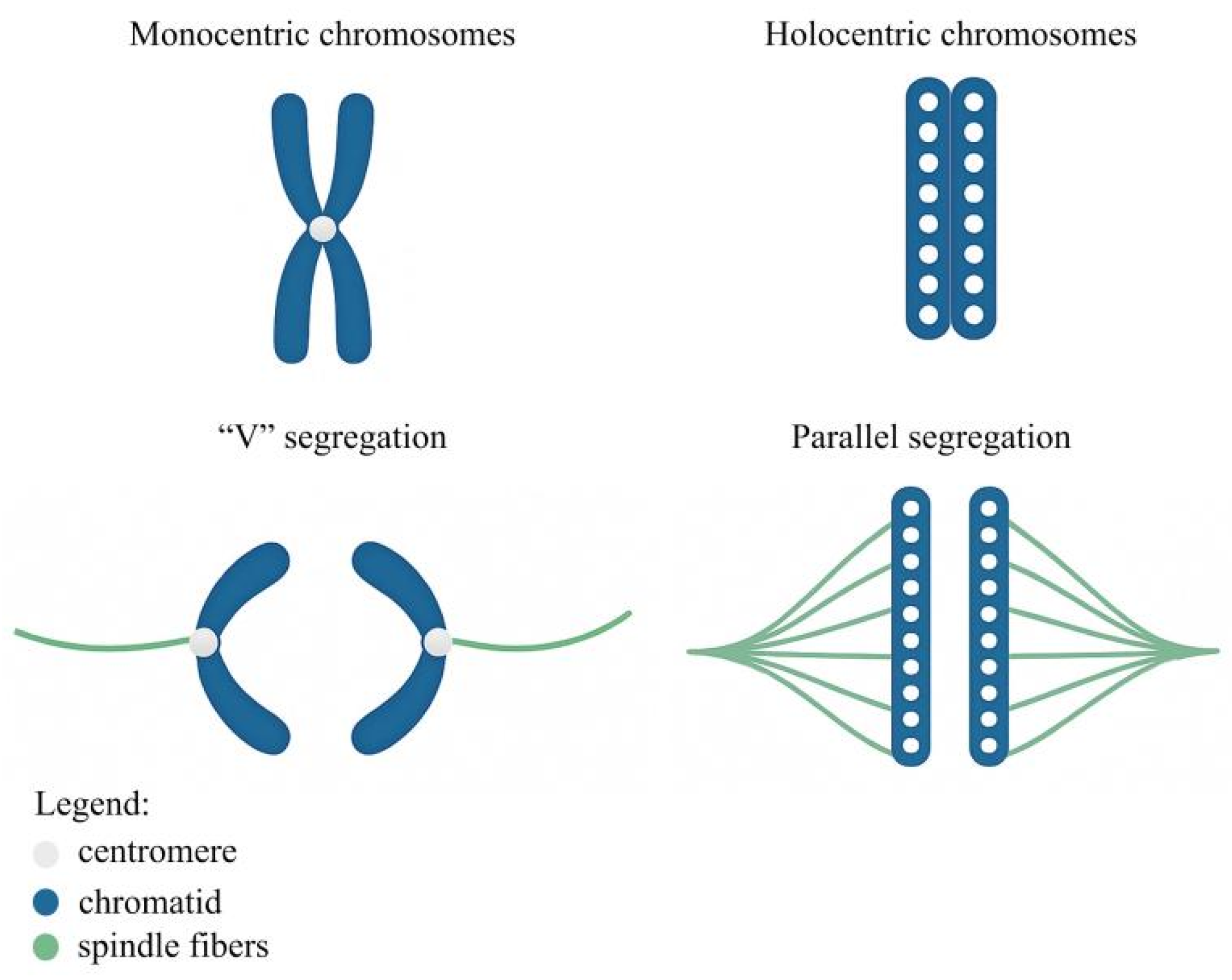
Figure 1: Main structural differences and anaphase behaviors between monocentric and holocentric chromosomes.
In general, centromeres are composed primarily of satellite DNA and/or transposable elements and are therefore highly heterochromatic, a feature that contributes to chromatid cohesion [10,13]. As a result, they are typically located in transcriptionally silent regions, although they may also harbor functionally active genes [14,15]. Accordingly, centromeric regions are characterized by pronounced sequence diversity, which may vary among chromosomes within the same individual [16,17,18], reflecting their rapid evolutionary dynamics [10,19].
From a structural standpoint, centromeres are characterized by the presence of a specialized variant of histone H3, known as CENH3 in plants [20,21]. This variant is essential for kinetochore assembly, a multiprotein complex responsible for microtubule attachment during cell division [22,23]. In animals, several chaperone proteins are involved in the deposition and recognition of CENH3 (CENP-A); however, the corresponding mechanisms in plants remain poorly understood [24]. Consequently, centromere activity does not depend on a specific nucleotide sequence but is instead determined by epigenetic regulation [25,26], illustrating the paradox between the high diversity of centromeric DNA sequences and the strong conservation of centromere function [27,28].
This article provides a historical overview of the main hypotheses concerning the origin and diversification of holocentric chromosomes, highlighting key advances and ongoing debates in the field. In addition, we review the principal techniques used to study these chromosomes, emphasizing their contributions to elucidating structural, functional, and evolutionary mechanisms. By doing so, we seek to establish a consolidated framework to support future investigations and to broaden the understanding of chromosomal diversity across different taxonomic groups.
2 Historical Overview, Origin and Evolution
Owing to the ability of chromosomal fragments to be stably inherited, holocentric species often exhibit natural variation in chromosome number [29,30,31,32]. The mechanisms underlying these numerical changes have been described using the terms agmatoploidy (chromosome fission) and simploidy (chromosome fusion) [33,34,35,36]. The concept of agmatoploidy was first proposed by Malheiros-Gardé and Gardé [37] to explain karyotypic variation in Luzula DC (Juncaceae). Based on the heritability of chromosomal fragments, the authors argued that variation in chromosome number in this genus did not result from polyploidization but rather from chromosome fission. Subsequently, the term simploidy was introduced by Luceño and Guerra [33] to describe cases in which a reduction in chromosome number is associated with an increase in chromosome size.
Chromosome fission is thought to occur at hotspots known as fragile sites, which are regions particularly prone to breakage following replication inhibition [30,38,39,40] and are also present in monocentric organisms [41]. For these fragments to be stabilized as independent chromosomes, the acquisition of telomeric sequences at broken or dysfunctional ends is required [42]. Telomeres prevent the formation of sticky chromosomes and, consequently, chromosome fusion [43,44]. In this context, the concepts of agmatoploidy and simploidy are indistinguishable from ascending and descending dysploidy, as all involve chromosome fissions and fusions without substantial DNA loss [44,45,46,47]. Accordingly, Guerra [48] argued that, because no distinct mechanisms have been proposed to explain these processes in holocentric chromosomes, there is no justification for adopting separate terminology.
The stable inheritance of chromosomal fragments has led to the hypothesis that holocentricity may promote higher rates of chromosome number variation [49,50]. However, a comparative study of monocentric and holocentric insects demonstrated that chromosome number evolution proceeds at statistically similar rates in both groups [51]. In Cryptangieae (Cyperaceae), for instance, a high frequency of chromosomal abnormalities has been observed during meiosis, particularly the presence of unaligned chromosomes [52]. Despite this, species in the group exhibit high pollen viability and no intraspecific variation in chromosome number (i.e., no distinct cytotypes). This pattern may be associated with the formation of pseudomonads, a process in which three of the four nuclei produced during male meiosis undergo apoptosis. This mechanism, which is recurrent in Cyperaceae but rare among other angiosperms, likely contributes to the elimination of cells carrying chromosomal anomalies [52].
In addition to structural chromosomal alterations, polyploidy and genome restructuring processes are also widespread among holocentric species [30,35,53,54,55]. In monocentric taxa, an increase in chromosome number is often associated with higher ploidy levels [56], although some species do not exhibit a corresponding increase in DNA content [57,58,59]. Under holocentric conditions, however, this relationship is rarely evident, as chromosome fissions and fusions can increase the basic chromosome number without changes in ploidy level [8].
In this context, chromosome counts should be complemented by nuclear DNA quantification to reliably assess the occurrence of polyploidy in holocentric species [8]. Additionally, the analysis of multivalent pairing during diakinesis is essential for identifying polyploids [60,61]. Finally, reconstructing the ancestral state of the basic chromosome number represents an effective strategy for mapping the mechanisms underlying changes in chromosome complements over the evolutionary history of a group, thereby allowing the distinction between polyploidy and dysploidy events [52,62,63,64].
Comparative phylogenetic analyses indicate that holocentricity is a homoplastic trait, having evolved independently in multiple lineages [65]. Because it occurs in relatively few groups, most authors regard holocentricity as a derived condition. Nonetheless, its evolutionary origin remains debated. Current estimates suggest that holocentricity has arisen at least 19 times among eukaryotes: six times in plants and 13 times in animals, including nematodes, velvet worms, insects, centipedes, and arachnids [65].
Holocentric chromosomes have been proposed as plesiomorphic, based on the hypothesis that intense UV radiation may have exerted strong deleterious effects during the early expansion of terrestrial life [66]. Under such conditions, organisms bearing holocentric chromosomes may have gained an adaptive advantage by tolerating chromosomal fragmentation. In contrast, the localized centromere of monocentric chromosomes represents a more specialized structure for chromosome segregation and is therefore regarded as a derived condition [67]. In this case, the confinement of centromeric activity to a single region, typically positioned away from chiasmata, may confer an adaptive advantage by concentrating kinetochore activity and promoting accurate chromosome segregation during meiosis [4,68]. By comparison, holocentric chromosomes generally exhibit a reduced number of chiasmata (usually one or two per chromosome) and the presence of three or more chiasmata may impair proper homologous chromosome separation [49,69,70,71]. Consequently, while monocentricity may support higher rates of gene recombination along chromosomes, holocentricity may compensate for this limitation through increased variation in chromosome number [49].
Holocentricity has been proposed to originate from an expansion of kinetochore activity mediated by transposable elements [65]. Similarly, another hypothesis suggests that centromeres evolved from telomeres, with the transposition of retroelements and the accumulation of subtelomeric repeats giving rise to the earliest centromeric regions [72,73]. In addition, Nagaki et al. [14] proposed that a 90° shift in the orientation of kinetochore assembly, extending along the chromosome axis toward the telomeric regions, may have contributed to the emergence of holocentric chromosomes.
Other hypotheses emphasize selective pressures associated with centromere structure. Malik and Henikoff [74] suggested that holocentric chromosomes may have evolved as a strategy to avoid the so-called “centromere drive”, a process in which larger centromeres, or those enriched in repetitive DNA, gain a transmission advantage during meiosis [75,76]. Because centromeric activity in holocentric chromosomes is distributed along the entire chromosome length, competition among centromeres would be minimized, thereby reducing the accumulation of potentially deleterious repetitive sequences. Subsequently, Bureš and Zedek [77] proposed the concept of “holokinetic drive” to explain the evolution of holocentric chromosomes, suggesting that larger chromosomes may be preferentially transmitted during meiosis, with consequences for genome size and complexity over evolutionary time.
Holocentric chromosomes are regarded as an apomorphic feature in Droseraceae [78] and as a synapomorphy for most clades of Juncaceae and Cyperaceae [11,79]. For a long time, the principal representatives of the Cyperid clade (Thurniaceae, Juncaceae, and Cyperaceae) were thought to be exclusively holocentric [80]. However, the recent discovery that Prionium serratum (L.f.) Drège (Thurniaceae) possesses monocentric chromosomes indicates that holocentricity arose independently at least twice after the divergence of Juncaceae and Cyperaceae [80]. In addition, monocentric chromosomes have also been reported in species from different sections of Juncus L. (Juncaceae) [79]. Accordingly, Guerra et al. [79] emphasized that the assumption of holocentricity should not be generalized to other genera of Juncaceae or to groups historically considered holocentric without robust cytogenetic evidence.
Holocentric chromosomes are thought to be widespread in Cyperaceae [8]. This family exhibits the greatest variation in chromosome number among angiosperms, ranging from 2n = 4 to 2n = 226, largely as a result of chromosome fissions and fusions [2,30,81]. Multiple modes of chromosomal evolution occur within the family, although dysploidy predominates [82,83]. Polyploidy appears to play a more limited role and has been reported mainly in Rhynchospora Vahl, Mapania Aubl., Fimbristylis Vahl, and Eleocharis R.Br. [82]. It has further been proposed that increased species diversification rates in Cyperaceae may be associated with karyotypic changes in combination with shifts in photosynthetic pathways [82]. Nevertheless, despite the large number of species in the family, chromosome counts are available for only a small fraction, with most data concentrated in Carex L. [81,84].
Evolutionary transitions between monocentricity and holocentricity can occur even among species within the same genus, as exemplified by Drosera L. (Droseraceae), a group of carnivorous plants [78]. In the parasitic genus Cuscuta L. (Convolvulaceae), holocentricity is restricted to the subgenus Cuscuta, whereas the subgenera Grammica and Monogynella retain monocentric chromosomes [85]. The transition to holocentricity in this lineage was accompanied by pronounced changes in kinetochore organization, including gene loss and the disruption of centromeric protein localization, such as CENH3 and CENP-C [86]. In addition, degeneration of the mitotic spindle assembly checkpoint has been reported, indicating a profound reorganization of chromosome segregation mechanisms [86]. This restructuring is further reflected in epigenetic modifications, including the loss of H2A histone phosphorylation and the elimination of centromeric retrotransposons, as well as a reduction in the basic chromosome number (x = 7) relative to monocentric species (x = 15 or 16) [85].
Similarly, in animal groups such as Lepidoptera, holocentricity is associated with major evolutionary processes. In Bombyx mori L., for example, high rates of chromosomal rearrangements have been documented [11]. In Erebia aethiops Esper, chromosomal fusions have led to a reduced chromosome number, along with inversions on the Z chromosome, suggesting an important role for holocentricity in speciation through chromosomal rearrangements [87]. More broadly, the presence of holocentric chromosomes is linked to evolutionary adaptations such as increased tolerance to chromosomal breakage, which may enhance survival under stressful environmental conditions and promote rapid karyotype evolution [66]. This property likely confers a significant adaptive advantage in contexts characterized by frequent exposure to clastogenic agents. Consequently, the evolution of holocentric chromosomes involves multiple genomic mechanisms and cellular adaptations, reflecting a complex convergent process that has occurred repeatedly throughout eukaryotic evolutionary history.
3 Methods for the Study of Holocentric Chromosomes
Classical (conventional) cytogenetic approaches can be used to assess V-shaped or parallel chromosome segregation, the latter being typical of holocentric chromosomes; however, such patterns are more readily observed in species with larger chromosomes. In the absence of an evident primary constriction, sister chromatids in holocentric chromosomes are oriented and segregate parallel to the metaphase plate during both mitosis and meiosis [8,88,89,90]. In contrast, monocentric chromosomes exhibit V-shaped segregation owing to the presence of a single, localized centromeric unit [8].
Meiosis in holocentric species also displays distinctive features and can be categorized into two main types. In the telokinetic type, homologous chromosomes form a configuration known as a “box”, which ensures their movement as a single unit during meiosis I [8,91,92,93]. In the holokinetic type, the reductional division occurs during meiosis II, which is why this process is also referred to as “inverted meiosis” [8,9,90,94]. These meiotic behaviors can be observed using light microscopy [52,90], but they are more accurately characterized using molecular cytogenetic techniques [9].
3.2 Chromosomal Break Induction Experiments
One of the earliest experimental approaches used to detect holocentric chromosomes involved the induction of chromosomal breaks by ionizing radiation, followed by analysis using flow cytometry. In holocentric species, the CENH3 protein is distributed along the entire chromosome, allowing fragments generated by breakage to segregate normally during both mitosis and meiosis without forming micronuclei [11,42,94]. Micronuclei are extranuclear structures that arise from chromosomal breakage or segregation errors and typically result in the loss of genetic material [95].
Based on this principle, ionizing radiation, including X-rays and gamma rays, has been applied to various plant tissues as an experimental strategy to assess the presence of holocentricity. The key evidence considered is the occurrence of chromosomal fragmentation without substantial DNA loss, measured in picograms, which suggests that the resulting fragments remain functional [96,97].
Currently, the primary technique for identifying holocentric chromosomes is the immunolocalization of CENH3. In monocentric chromosomes, CENH3 is confined to the primary constriction, whereas in holocentric chromosomes it is distributed along nearly the entire chromosome length [98]. These contrasting patterns can be confirmed by immunolocalization using specific antibodies [71,98]. However, as with centromeric DNA sequences, CENH3 exhibits considerable variability in amino acid composition and may be species-specific, which can limit its applicability [27,99,100]. Recently, Oliveira et al. [101] introduced new universal markers for the detection of functional centromeres, which are expected to facilitate their use in non-model plant systems.
Because of the high cost associated with the development and production of species-specific CENH3 antibodies, tubulin labeling has emerged as a viable alternative [100,101,102]. Tubulin, the main structural component of mitotic spindle fibers, can be visualized to directly assess spindle–chromosome interactions during cell division. This approach enables the indirect inference of centromere distribution and behavior, particularly in holocentric chromosomes, where spindle fibers attach along almost the entire chromosome length.
4 Conclusions and Future Perspectives
Despite methodological and conceptual advances in recent decades, research on holocentric chromosomes continues to face significant challenges, particularly regarding the accurate identification of this condition in non-model organisms. The limited availability of specific molecular tools, interspecific variation in centromeric proteins, and the scarcity of cytogenetic data for many taxa hinder the systematic characterization of holocentricity across extant biodiversity.
Addressing these challenges requires clearly defined criteria for selecting suitable model organisms. Species with small, stable chromosome numbers are particularly valuable. Ideal models should also combine well-characterized genomes supported by robust genomic resources (e.g., high-quality genome assemblies and linkage maps) and practical traits that facilitate experimental handling. These traits include straightforward cultivation, rapid and reliable propagation, and consistent flowering. Additionally, organisms amenable to controlled experimental manipulation, such as induced chromosome fragmentation, can provide key insights into kinetochore organization, chromosome segregation, and karyotype evolution. Comparative approaches integrating plants, invertebrates, and nematodes may further elucidate the evolutionary and functional significance of holocentricity across different ecological and selective contexts.
Building on these considerations, plant lineages in which sex chromosomes could occur in a holocentric background represent a particularly promising avenue for research, as this scenario is rare and largely unexplored [103,104,105]. Although dioecious species exist in several holocentric plant families, there is currently no formally published cytogenetic evidence of heteromorphic sex chromosomes (i.e., differentiated XY or ZW systems) in any of them. This lack of documented sex chromosomes makes these groups especially valuable for investigating whether and how sex-linked regions might arise and evolve in a holocentric background. The potential interaction between holocentricity and putative sex-determining regions could have profound implications for meiotic behavior, genome stability, and karyotype evolution, representing an unexplored frontier in plant cytogenetics. Beyond contributing to our understanding of holocentric chromosome organization, this review emphasizes the importance of expanding cytogenetic and genomic studies to taxonomically diverse plant groups, particularly those combining dioecy with holocentric chromosomes.
Acknowledgement:
Funding Statement: The authors received no specific funding for this study.
Author Contributions: The authors confirm contribution to the paper as follows: conceptualization, Ana Luisa Arantes Chaves and Marco Tulio Mendes Ferreira; methodology, Ana Luisa Arantes Chaves; validation, Ana Luisa Arantes Chaves and Marco Tulio Mendes Ferreira; formal analysis, Ana Luisa Arantes Chaves; investigation, Ana Luisa Arantes Chaves; data curation, Ana Luisa Arantes Chaves; writing—original draft preparation, Ana Luisa Arantes Chaves; writing—review and editing, Marco Tulio Mendes Ferreira; visualization, Ana Luisa Arantes Chaves. All authors reviewed and approved the final version of the manuscript.
Availability of Data and Materials: Not applicable.
Ethics Approval: Not applicable.
Conflicts of Interest: The authors declare no conflicts of interest.
Glossary
| CENH3 | Centromeric Histone H3 variant. Centromere-specific variant of histone H3 that replaces canonical H3 in nucleosomes at functional centromeres. It plays an essential role in specifying centromere identity and assembling the kinetochore, thereby ensuring accurate chromosome segregation during cell division. Term A name traditionally used for plants. |
| CENP-A | Centromere Protein A. Centromere-specific histone H3 variant that replaces canonical H3 in nucleosomes at active centromeres. It is essential for establishing centromere identity and serves as the foundation for kinetochore assembly, ensuring faithful chromosome segregation during cell division. Term name traditionally used for animals, especially vertebrates. |
References
1. Singh RJ . Plant cytogenetics. Boca Raton, FL, USA: CRC Press; 2016. 528 p. [Google Scholar]
2. Vimala Y , Lavania S , Lavania UC . Chromosome change and karyotype differentiation–implications in speciation and plant systematics. Nucleus. 2021; 64( 1): 33– 54. doi:10.1007/s13237-020-00343-y. [Google Scholar] [CrossRef]
3. Guerra M . Cytotaxonomy: the end of childhood. Plant Biosyst Int J Deal Aspects Plant Biol. 2012; 146( 3): 703– 10. doi:10.1080/11263504.2012.717973. [Google Scholar] [CrossRef]
4. Zickler D , Kleckner N . Meiosis: dances between homologs. Annu Rev Genet. 2023; 57: 1– 63. doi:10.1146/annurev-genet-061323-044915. [Google Scholar] [CrossRef]
5. Neumann P , Navrátilová A , Schroeder-Reiter E , Koblížková A , Steinbauerová V , Chocholová E , et al. Stretching the rules: monocentric chromosomes with multiple centromere domains. PLoS Genet. 2012; 8( 6): e1002777. doi:10.1371/journal.pgen.1002777. [Google Scholar] [CrossRef]
6. Levan A , Fredga K , Sandberg AA . Nomenclature for centromeric position on chromosomes. Hereditas. 1964; 52( 2): 201– 20. doi:10.1111/j.1601-5223.1964.tb01953.x. [Google Scholar] [CrossRef]
7. Guerra M . Reviewing the chromosome nomenclature of Levan et al. Braz J Genet. 1986; 9: 741– 3. [Google Scholar]
8. Bureš P , Zedek F , Marková M . Holocentric chromosomes. In: Greilhuber J , Doležel J , Wendel JF , editors. Plant genome diversity volume 2. Vienna, Austria: Springer; 2012. p. 187– 208. [Google Scholar]
9. Heckmann S , Macas J , Kumke K , Fuchs J , Schubert V , Ma L , et al. The holocentric species Luzula elegans shows interplay between centromere and large-scale genome organization. Plant J. 2013; 73( 4): 555– 65. doi:10.1111/tpj.12054. [Google Scholar] [CrossRef]
10. Steckenborn S , Marques A . Centromere diversity and its evolutionary impacts on plant karyotypes and plant reproduction. New Phytol. 2025; 245( 5): 1879– 86. doi:10.1111/nph.20376. [Google Scholar] [CrossRef]
11. Melters DP , Paliulis LV , Korf IF , Chan SWL . Holocentric chromosomes: convergent evolution, meiotic adaptations, and genomic analysis. Chromosome Res. 2012; 20( 5): 579– 93. doi:10.1007/s10577-012-9292-1. [Google Scholar] [CrossRef]
12. Ibiapino A , García MA , Amorim B , Baez M , Costea M , Stefanović S , et al. The evolution of cytogenetic traits in Cuscuta (Convolvulaceae), the genus with the most diverse chromosomes in angiosperms. Front Plant Sci. 2022; 13: 842260. doi:10.3389/fpls.2022.842260. [Google Scholar] [CrossRef]
13. Bernard P , Maure JF , Partridge JF , Genier S , Javerzat JP , Allshire RC . Requirement of heterochromatin for cohesion at centromeres. Science. 2001; 294( 5551): 2539– 42. doi:10.1126/science.1064027. [Google Scholar] [CrossRef]
14. Nagaki K , Kashihara K , Murata M . Visualization of diffuse centromeres with centromere-specific histone H3 in the holocentric plant Luzula nivea. Plant Cell. 2005; 17( 7): 1886– 93. doi:10.1105/tpc.105.032961. [Google Scholar] [CrossRef]
15. Liu Q , Liu Y , Shi Q , Su H , Wang C , Birchler JA , et al. Emerging roles of centromeric RNAs in centromere formation and function. Genes Genom. 2021; 43( 3): 217– 26. doi:10.1007/s13258-021-01041-y. [Google Scholar] [CrossRef]
16. Bao W , Zhang W , Yang Q , Zhang Y , Han B , Gu M , et al. Diversity of centromeric repeats in two closely related wild rice species, Oryza officinalis and Oryza rhizomatis. Mol Genet Genom. 2006; 275( 5): 421– 30. doi:10.1007/s00438-006-0103-2. [Google Scholar] [CrossRef]
17. Lim KB , Yang TJ , Hwang YJ , Kim JS , Park JY , Kwon SJ , et al. Characterization of the centromere and peri-centromere retrotransposons in Brassica rapa and their distribution in related Brassica species. Plant J. 2007; 49( 2): 173– 83. doi:10.1111/j.1365-313X.2006.02952.x. [Google Scholar] [CrossRef]
18. Su H , Liu Y , Liu C , Shi Q , Huang Y , Han F . Centromere satellite repeats have undergone rapid changes in polyploid wheat subgenomes. Plant Cell. 2019; 31( 9): 2035– 51. doi:10.1105/tpc.19.00133. [Google Scholar] [CrossRef]
19. Talbert PB , Henikoff S . What makes a centromere? Exp Cell Res. 2020; 389( 2): 111895. doi:10.1016/j.yexcr.2020.111895. [Google Scholar] [CrossRef]
20. Talbert PB , Henikoff S . Phylogeny as the basis for naming histones. Trends Genet. 2013; 29( 9): 499– 500. doi:10.1016/j.tig.2013.06.009. [Google Scholar] [CrossRef]
21. Kursel LE , Malik HS . Centromeres. Curr Biol. 2016; 26( 12): R487– 90. doi:10.1016/j.cub.2016.05.031. [Google Scholar] [CrossRef]
22. Navarro AP , Cheeseman IM . Chromosome segregation: evolving a plastic chromosome–microtubule interface. Curr Biol. 2020; 30( 4): R174– 7. doi:10.1016/j.cub.2019.12.058. [Google Scholar] [CrossRef]
23. Hara M , Fukagawa T . Dynamics of kinetochore structure and its regulations during mitotic progression. Cell Mol Life Sci. 2020; 77( 15): 2981– 95. doi:10.1007/s00018-020-03472-4. [Google Scholar] [CrossRef]
24. Keçeli BN , Jin C , Van Damme D , Geelen D . Conservation of centromeric histone 3 interaction partners in plants. J Exp Bot. 2020; 71( 17): 5237– 46. doi:10.1093/jxb/eraa214. [Google Scholar] [CrossRef]
25. Sullivan BA , Blower MD , Karpen GH . Determining centromere identity: cyclical stories and forking paths. Nat Rev Genet. 2001; 2( 8): 584– 96. doi:10.1038/35084512. [Google Scholar] [CrossRef]
26. Wong CYY , Lee BCH , Yuen KWY . Epigenetic regulation of centromere function. Cell Mol Life Sci. 2020; 77( 15): 2899– 917. doi:10.1007/s00018-020-03460-8. [Google Scholar] [CrossRef]
27. Henikoff S , Ahmad K , Malik HS . The centromere paradox: stable inheritance with rapidly evolving DNA. Science. 2001; 293( 5532): 1098– 102. doi:10.1126/science.1062939. [Google Scholar] [CrossRef]
28. Ma R , Zhu B . Pericentromeric sequences, where a conservation paradox occurs. Trends Cell Biol. 2025; 35( 12): 1007– 15. doi:10.1016/j.tcb.2025.01.011. [Google Scholar] [CrossRef]
29. Kandul NP , Lukhtanov VA , Pierce NE . Karyotypic diversity and speciation in Agrodiaetus butterflies. Evolution. 2007; 61( 3): 546– 59. doi:10.1111/j.1558-5646.2007.00046.x. [Google Scholar] [CrossRef]
30. Roalson EH . A synopsis of chromosome number variation in the Cyperaceae. Bot Rev. 2008; 74( 2): 209– 393. doi:10.1007/s12229-008-9011-y. [Google Scholar] [CrossRef]
31. Escudero M , Hipp AL , Luceño M . Karyotype stability and predictors of chromosome number variation in sedges: a study in Carex section Spirostachyae (Cyperaceae). Mol Phylogenet Evol. 2010; 57( 1): 353– 63. doi:10.1016/j.ympev.2010.07.009. [Google Scholar] [CrossRef]
32. Dincă V , Lukhtanov VA , Talavera G , Vila R . Unexpected layers of cryptic diversity in wood white Leptidea butterflies. Nat Commun. 2011; 2: 324. doi:10.1038/ncomms1329. [Google Scholar] [CrossRef]
33. Luceño M , Guerra M . Numerical variations in species exhibiting holocentric chromosomes: a nomenclatural proposal. Caryologia. 1996; 49( 3–4): 301– 9. doi:10.1080/00087114.1996.10797374. [Google Scholar] [CrossRef]
34. Stoeva MP , Uzunova K , Popova ED , Stoyanova K . Patterns and levels of variation within section Phacocystis of genus Carex (Cyperaceae) in Bulgaria. Phytol Balc. 2005; 11( 1): 45– 62. [Google Scholar]
35. Hipp AL , Rothrock PE , Roalson EH . The evolution of chromosome arrangements in Carex (Cyperaceae). Bot Rev. 2009; 75( 1): 96– 109. doi:10.1007/s12229-008-9022-8. [Google Scholar] [CrossRef]
36. Záveská Drábková L . A survey of karyological phenomena in the Juncaceae with emphasis on chromosome number variation and evolution. Bot Rev. 2013; 79( 3): 401– 46. doi:10.1007/s12229-013-9127-6. [Google Scholar] [CrossRef]
37. Malheiros-Gardé N , Gardé A . Chromosome number in Luzula multiflora Lej. Genet Iber. 1950; 4: 91– 4. [Google Scholar]
38. Manicardi GC , Nardelli A , Mandrioli M . Fast chromosomal evolution and karyotype instability: recurrent chromosomal rearrangements in the peach potato aphid Myzus persicae (Hemiptera: Aphididae). Biol J Linn Soc. 2015; 116( 3): 519– 29. doi:10.1111/bij.12621. [Google Scholar] [CrossRef]
39. Monti V , Lombardo G , Loxdale HD , Manicardi GC , Mandrioli M . Continuous occurrence of intra-individual chromosome rearrangements in the peach potato aphid, Myzus persicae (Sulzer) (Hemiptera: Aphididae). Genetica. 2012; 140( 1–3): 93– 103. doi:10.1007/s10709-012-9661-x. [Google Scholar] [CrossRef]
40. Lipnerová I , Bures P , Horová L , Smarda P . Evolution of genome size in Carex (Cyperaceae) in relation to chromosome number and genomic base composition. Ann Bot. 2013; 111( 1): 79– 94. doi:10.1093/aob/mcs239. [Google Scholar] [CrossRef]
41. Aguilera A , García-Muse T . Causes of genome instability. Annu Rev Genet. 2013; 47: 1– 32. doi:10.1146/annurev-genet-111212-133232. [Google Scholar] [CrossRef]
42. Jankowska M , Fuchs J , Klocke E , Fojtová M , Polanská P , Fajkus J , et al. Holokinetic centromeres and efficient telomere healing enable rapid karyotype evolution. Chromosoma. 2015; 124( 4): 519– 28. doi:10.1007/s00412-015-0524-y. [Google Scholar] [CrossRef]
43. Ferreira MG , Miller KM , Cooper JP . Indecent exposure: when telomeres become uncapped. Mol Cell. 2004; 13( 1): 7– 18. doi:10.1016/s1097-2765(03)00531-8. [Google Scholar] [CrossRef]
44. Schubert I , Lysak MA . Interpretation of karyotype evolution should consider chromosome structural constraints. Trends Genet. 2011; 27( 6): 207– 16. doi:10.1016/j.tig.2011.03.004. [Google Scholar] [CrossRef]
45. Lysak MA . Live and let die: centromere loss during evolution of plant chromosomes. New Phytol. 2014; 203( 4): 1082– 9. doi:10.1111/nph.12885. [Google Scholar] [CrossRef]
46. Escudero M , Martín-Bravo S , Mayrose I , Fernández-Mazuecos M , Fiz-Palacios O , Hipp AL , et al. Karyotypic changes through dysploidy persist longer over evolutionary time than polyploid changes. PLoS One. 2014; 9( 1): e85266. doi:10.1371/journal.pone.0085266. [Google Scholar] [CrossRef]
47. Mandáková T , Lysak MA . Post-polyploid diploidization and diversification through dysploid changes. Curr Opin Plant Biol. 2018; 42: 55– 65. doi:10.1016/j.pbi.2018.03.001. [Google Scholar] [CrossRef]
48. Guerra M . Agmatoploidy and symploidy: a critical review. Genet Mol Biol. 2016; 39( 4): 492– 6. doi:10.1590/1678-4685-GMB-2016-0103. [Google Scholar] [CrossRef]
49. Escudero M , Hipp AL , Hansen TF , Voje KL , Luceño M . Selection and inertia in the evolution of holocentric chromosomes in sedges (Carex, Cyperaceae). New Phytol. 2012; 195( 1): 237– 47. doi:10.1111/j.1469-8137.2012.04137.x. [Google Scholar] [CrossRef]
50. Mandrioli M , Manicardi GC . Holocentric chromosomes. PLoS Genet. 2020; 16( 7): e1008918. doi:10.1371/journal.pgen.1008918. [Google Scholar] [CrossRef]
51. Ruckman SN , Jonika MM , Casola C , Blackmon H . Chromosome number evolves at equal rates in holocentric and monocentric clades. PLoS Genet. 2020; 16( 10): e1009076. doi:10.1371/journal.pgen.1009076. [Google Scholar] [CrossRef]
52. Chaves ALA , Ferreira MTM , Escudero M , Luceño M , Costa SM . Chromosomal evolution in Cryptangieae Benth. (Cyperaceae): evidence of holocentrism and pseudomonads. Protoplasma. 2024; 261( 3): 527– 41. doi:10.1007/s00709-023-01915-w. [Google Scholar] [CrossRef]
53. Vanzela ALL , Cuadrado A , Guerra M . Localization of 45S rDNA and telomeric sites on holocentric chromosomes of Rhynchospora tenuis Link (Cyperaceae). Genet Mol Biol. 2003; 26( 2): 199– 201. doi:10.1590/s1415-47572003000200014. [Google Scholar] [CrossRef]
54. Yano O , Hoshino T . Phylogenetic relationships and chromosomal evolution of Japanese Fimbristylis (Cyperaceae) using nrDNA ITS and ETS 1f sequence data. Acta Phytotax Geobot. 2007; 57( 3): 205– 17. doi:10.18942/apg.KJ00004622872. [Google Scholar] [CrossRef]
55. da Silva CRM , González-Elizondo MS , Laforga Vanzela AL . Chromosome reduction in Eleocharis maculosa (Cyperaceae). Cytogenet Genome Res. 2008; 122( 2): 175– 80. doi:10.1159/000163096. [Google Scholar] [CrossRef]
56. Frawley LE , Orr-Weaver TL . Polyploidy. Curr Biol. 2015; 25( 9): R353– 8. doi:10.1016/j.cub.2015.03.037. [Google Scholar] [CrossRef]
57. Leitch IJ , Bennett MD . Genome downsizing in polyploid plants. Biol J Linn Soc. 2004; 82( 4): 651– 63. doi:10.1111/j.1095-8312.2004.00349.x. [Google Scholar] [CrossRef]
58. Chaves ALA , Chiavegatto RB , Benites FRG , Techio VH . Comparative karyotype analysis among cytotypes of Cynodon dactylon (L.) Pers. (Poaceae). Mol Biol Rep. 2019; 46( 5): 4873– 81. doi:10.1007/s11033-019-04935-z. [Google Scholar] [CrossRef]
59. Chaves ALA , Carvalho PHM , Ferreira MTM , Benites FRG , Techio VH . Genomic constitution, allopolyploidy, and evolutionary proposal for Cynodon Rich. based on GISH. Protoplasma. 2022; 259( 4): 999– 1011. doi:10.1007/s00709-021-01716-z. [Google Scholar] [CrossRef]
60. Hofstatter PG , Thangavel G , Castellani M , Marques A . Meiosis progression and recombination in holocentric plants: what is known? Front Plant Sci. 2021; 12: 658296. doi:10.3389/fpls.2021.658296. [Google Scholar] [CrossRef]
61. Soares NR , Mollinari M , Oliveira GK , Pereira GS , Vieira MLC . Meiosis in polyploids and implications for genetic mapping: a review. Genes. 2021; 12( 10): 1517. doi:10.3390/genes12101517. [Google Scholar] [CrossRef]
62. Glick L , Mayrose I . ChromEvol: assessing the pattern of chromosome number evolution and the inference of polyploidy along a phylogeny. Mol Biol Evol. 2014; 31( 7): 1914– 22. doi:10.1093/molbev/msu122. [Google Scholar] [CrossRef]
63. Ribeiro T , Buddenhagen CE , Thomas WW , Souza G , Pedrosa-Harand A . Are holocentrics doomed to change? Limited chromosome number variation in Rhynchospora Vahl (Cyperaceae). Protoplasma. 2018; 255( 1): 263– 72. doi:10.1007/s00709-017-1154-4. [Google Scholar] [CrossRef]
64. Carta A , Bedini G , Peruzzi L . A deep dive into the ancestral chromosome number and genome size of flowering plants. New Phytol. 2020; 228( 3): 1097– 106. doi:10.1111/nph.16668. [Google Scholar] [CrossRef]
65. Escudero M , Márquez-Corro JI , Hipp AL . The phylogenetic origins and evolutionary history of holocentric chromosomes. Syst Bot. 2016; 41( 3): 580– 5. [Google Scholar]
66. Zedek F , Bureš P . Holocentric chromosomes: from tolerance to fragmentation to colonization of the land. Ann Bot. 2018; 121( 1): 9– 16. doi:10.1093/aob/mcx118. [Google Scholar] [CrossRef]
67. Sybenga J . Specialization in the behaviour of chromosomes on the meiotic spindle. Genetica. 1981; 57( 2): 143– 51. doi:10.1007/BF00131240. [Google Scholar] [CrossRef]
68. Egel R , Penny D . On the origin of meiosis in eukaryotic evolution: coevolution of meiosis and mitosis from feeble beginnings. In: Recombination and meiosis. Berlin/Heidelberg, Germany: Springer; 2007. p. 249– 88. [Google Scholar]
69. Nokkala S , Kuznetsova VG , Maryanska-Nadachowska A , Nokkala C . Holocentric chromosomes in meiosis. I. Restriction of the number of chiasmata in bivalents. Chromosome Res. 2004; 12( 7): 733– 9. doi:10.1023/B:CHRO.0000045797.74375.70. [Google Scholar] [CrossRef]
70. de Bigliardo GR , Gabriel Virla E , Caro S , Murillo Dasso S . Karyotype and male pre-reductional meiosis of the sharpshooter Tapajosa rubromarginata (Hemiptera: Cicadellidae). Rev Biol Trop. 2011; 59( 1): 309– 14. doi:10.15517/rbt.v59i1.3200. [Google Scholar] [CrossRef]
71. Cabral G , Marques A , Schubert V , Pedrosa-Harand A , Schlögelhofer P . Chiasmatic and achiasmatic inverted meiosis of plants with holocentric chromosomes. Nat Commun. 2014; 5: 5070. doi:10.1038/ncomms6070. [Google Scholar] [CrossRef]
72. Villasante A , Abad JP , Méndez-Lago M . Centromeres were derived from telomeres during the evolution of the eukaryotic chromosome. Proc Natl Acad Sci U S A. 2007; 104( 25): 10542– 7. doi:10.1073/pnas.0703808104. [Google Scholar] [CrossRef]
73. Villasante A , Méndez-Lago M , Abad JP , Montejo de Garcíni E . The birth of the centromere. Cell Cycle. 2007; 6( 23): 2872– 6. doi:10.4161/cc.6.23.5047. [Google Scholar] [CrossRef]
74. Malik HS , Henikoff S . Major evolutionary transitions in centromere complexity. Cell. 2009; 138( 6): 1067– 82. doi:10.1016/j.cell.2009.08.036. [Google Scholar] [CrossRef]
75. Kursel LE , Malik HS . The cellular mechanisms and consequences of centromere drive. Curr Opin Cell Biol. 2018; 52: 58– 65. doi:10.1016/j.ceb.2018.01.011. [Google Scholar] [CrossRef]
76. Dudka D , Lampson MA . Centromere drive: model systems and experimental progress. Chromosome Res. 2022; 30( 2–3): 187– 203. doi:10.1007/s10577-022-09696-3. [Google Scholar] [CrossRef]
77. Bureš P , Zedek F . Holokinetic drive: centromere drive in chromosomes without centromeres. Evolution. 2014; 68( 8): 2412– 20. doi:10.1111/evo.12437. [Google Scholar] [CrossRef]
78. Kolodin P , Cempírková H , Bureš P , Horová L , Veleba A , Francová J , et al. Holocentric chromosomes may be an apomorphy of Droseraceae. Plant Syst Evol. 2018; 304( 10): 1289– 96. doi:10.1007/s00606-018-1546-8. [Google Scholar] [CrossRef]
79. Guerra M , Ribeiro T , Felix LP . Monocentric chromosomes in Juncus (Juncaceae) and implications for the chromosome evolution of the family. Bot J Linn Soc. 2019; 191( 4): 475– 83. doi:10.1093/botlinnean/boz065. [Google Scholar] [CrossRef]
80. Baez M , Kuo YT , Dias Y , Souza T , Boudichevskaia A , Fuchs J , et al. Analysis of the small chromosomal Prionium serratum (Cyperid) demonstrates the importance of reliable methods to differentiate between mono- and holocentricity. Chromosoma. 2020; 129( 3–4): 285– 97. doi:10.1007/s00412-020-00745-6. [Google Scholar] [CrossRef]
81. Rice A , Glick L , Abadi S , Einhorn M , Kopelman NM , Salman-Minkov A , et al. The chromosome counts database (CCDB)—a community resource of plant chromosome numbers. New Phytol. 2015; 206( 1): 19– 26. doi:10.1111/nph.13191. [Google Scholar] [CrossRef]
82. Márquez-Corro JI , Martín-Bravo S , Spalink D , Luceño M , Escudero M . Inferring hypothesis-based transitions in clade-specific models of chromosome number evolution in sedges (Cyperaceae). Mol Phylogenet Evol. 2019; 135: 203– 9. doi:10.1016/j.ympev.2019.03.006. [Google Scholar] [CrossRef]
83. Márquez-Corro JI , Martín-Bravo S , Jiménez-Mejías P , Hipp AL , Spalink D , Naczi RFC , et al. Macroevolutionary insights into sedges (Carex: Cyperaceae): the effects of rapid chromosome number evolution on lineage diversification. J Syst Evol. 2021; 59( 4): 776– 90. doi:10.1111/jse.12730. [Google Scholar] [CrossRef]
84. Więcław H , Kalinka A , Koopman J . Chromosome numbers of Carex (Cyperaceae) and their taxonomic implications. PLoS One. 2020; 15( 2): e0228353. doi:10.1371/journal.pone.0228353. [Google Scholar] [CrossRef]
85. Neumann P , Oliveira L , Čížková J , Jang TS , Klemme S , Novák P , et al. Impact of parasitic lifestyle and different types of centromere organization on chromosome and genome evolution in the plant genus Cuscuta. New Phytol. 2021; 229( 4): 2365– 77. doi:10.1111/nph.17003. [Google Scholar] [CrossRef]
86. Neumann P , Oliveira L , Jang TS , Novák P , Koblížková A , Schubert V , et al. Disruption of the standard kinetochore in holocentric Cuscuta species. Proc Natl Acad Sci U S A. 2023; 120( 21): e2300877120. doi:10.1073/pnas.2300877120. [Google Scholar] [CrossRef]
87. Pazhenkova EA , Lukhtanov VA . Whole-genome analysis reveals the dynamic evolution of holocentric chromosomes in satyrine butterflies. Genes. 2023; 14( 2): 437. doi:10.3390/genes14020437. [Google Scholar] [CrossRef]
88. Dopchiz LP , Poggio L . Meiosis and pollen grain development in Isolepis cernua f. cernua (Cyperaceae). Caryologia. 1999; 52( 3–4): 197– 201. doi:10.1080/00087114.1998.10589173. [Google Scholar] [CrossRef]
89. Pérez R , Rufas JS , Suja JA , Page J , Panzera F . Meiosis in holocentric chromosomes: orientation and segregation of an autosome and sex chromosomes in Triatoma infestans (Heteroptera). Chromosome Res. 2000; 8( 1): 17– 25. doi:10.1023/a:1009266801160. [Google Scholar] [CrossRef]
90. Da Silva CRM , González-Elizondo MS , Vanzela ALL . Reduction of chromosome number in Eleocharis subarticulata (Cyperaceae) by multiple translocations. Bot J Linn Soc. 2005; 149( 4): 457– 64. doi:10.1111/j.1095-8339.2005.00449.x. [Google Scholar] [CrossRef]
91. Monen J , Maddox PS , Hyndman F , Oegema K , Desai A . Differential role of CENP-A in the segregation of holocentric C. elegans chromosomes during meiosis and mitosis. Nat Cell Biol. 2005; 7( 12): 1248– 55. doi:10.1038/ncb1331. [Google Scholar] [CrossRef]
92. Dumont J , Oegema K , Desai A . A kinetochore-independent mechanism drives anaphase chromosome separation during acentrosomal meiosis. Nat Cell Biol. 2010; 12( 9): 894– 901. doi:10.1038/ncb2093. [Google Scholar] [CrossRef]
93. Maximiano da Silva CR , Trevisan R , González-Elizondo MS , Ferreira JM , Vanzela ALL . Karyotypic diversification and its contribution to the taxonomy of Eleocharis (Cyperaceae) from Brazil. Aust J Bot. 2010; 58( 1): 49– 60. doi:10.1071/bt09185. [Google Scholar] [CrossRef]
94. Pazy B , Plitmann U . Holocentric chromosome behaviour inCuscuta (Cuscutaceae). Plant Syst Evol. 1994; 191( 1): 105– 9. doi:10.1007/BF00985345. [Google Scholar] [CrossRef]
95. Terradas M , Martín M , Tusell L , Genescà A . Genetic activities in micronuclei: is the DNA entrapped in micronuclei lost for the cell? Mutat Res. 2010; 705( 1): 60– 7. doi:10.1016/j.mrrev.2010.03.004. [Google Scholar] [CrossRef]
96. Sheikh SA , Kondo K , Hoshi Y . Study on diffused centromeric nature of Drosera chromosomes. Cytologia. 1995; 60( 1): 43– 7. doi:10.1508/cytologia.60.43. [Google Scholar] [CrossRef]
97. Vanzela ALL , Colaço W . Mitotic and meiotic behavior of γ irradiated holocentric chromosomes of Rhynchospora pubera (Cyperaceae). Acta Sci. 2002; 24( 2): 611– 4. doi:10.4025/actascibiolsci.v24i0.2364. [Google Scholar] [CrossRef]
98. Wanner G , Schroeder-Reiter E , Ma W , Houben A , Schubert V . The ultrastructure of mono- and holocentric plant centromeres: an immunological investigation by structured illumination microscopy and scanning electron microscopy. Chromosoma. 2015; 124( 4): 503– 17. doi:10.1007/s00412-015-0521-1. [Google Scholar] [CrossRef]
99. Drinnenberg IA , Henikoff S , Malik HS . Evolutionary turnover of kinetochore proteins: a ship of Theseus? Trends Cell Biol. 2016; 26( 7): 498– 510. doi:10.1016/j.tcb.2016.01.005. [Google Scholar] [CrossRef]
100. Oliveira L , Neumann P , Jang TS , Klemme S , Schubert V , Koblížková A , et al. Mitotic spindle attachment to the holocentric chromosomes of Cuscuta europaea does not correlate with the distribution of CENH3 chromatin. Front Plant Sci. 2020; 10: 1799. doi:10.3389/fpls.2019.01799. [Google Scholar] [CrossRef]
101. Oliveira L , Neumann P , Mata-Sucre Y , Kuo YT , Marques A , Schubert V , et al. KNL1 and NDC80 represent new universal markers for the detection of functional centromeres in plants. Chromosome Res. 2024; 32( 1): 3. doi:10.1007/s10577-024-09747-x. [Google Scholar] [CrossRef]
102. Marques A , Schubert V , Houben A , Pedrosa-Harand A . Restructuring of holocentric centromeres during meiosis in the plant Rhynchospora pubera. Genetics. 2016; 204( 2): 555– 68. doi:10.1534/genetics.116.191213. [Google Scholar] [CrossRef]
103. Negrutiu I , Vyskot B , Barbacar N , Georgiev S , Moneger F . Dioecious plants. A key to the early events of sex chromosome evolution. Plant Physiol. 2001; 127( 4): 1418– 24. [Google Scholar]
104. Käfer J , Marais GAB , Pannell JR . On the rarity of dioecy in flowering plants. Mol Ecol. 2017; 26( 5): 1225– 41. doi:10.1111/mec.14020. [Google Scholar] [CrossRef]
105. Charlesworth D , Harkess A . Why should we study plant sex chromosomes? Plant Cell. 2024; 36( 5): 1242– 56. doi:10.1093/plcell/koad278. [Google Scholar] [CrossRef]
Cite This Article
 Copyright © 2026 The Author(s). Published by Tech Science Press.
Copyright © 2026 The Author(s). Published by Tech Science Press.This work is licensed under a Creative Commons Attribution 4.0 International License , which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.


 Submit a Paper
Submit a Paper Propose a Special lssue
Propose a Special lssue View Full Text
View Full Text Download PDF
Download PDF Downloads
Downloads
 Citation Tools
Citation Tools