 Open Access
Open Access
ARTICLE
Pharmacotherapeutics and molecular docking studies of alpha-synuclein modulators as promising therapeutics for Parkinson’s disease
1 Department of Medical Elementology and Toxicology, Jamia Hamdard, New Delhi, 110062, India
2 Centre for Interdisciplinary Research in Basic Sciences, Jamia Millia Islamia, New Delhi, 110025, India
3 Department of Pharmaceutical Sciences, Gurukul Kangri University, Uttarakhand, 249404, India
4 Department of Pharmacology HIMSR, Jamia Hamdard, New Delhi, 110062, India
* Corresponding Author: RAZI AHMAD. Email:
BIOCELL 2022, 46(12), 2681-2694. https://doi.org/10.32604/biocell.2022.021224
Received 02 January 2022; Accepted 11 April 2022; Issue published 10 August 2022
Abstract
Parkinson’s disease (PD) is an age-related neurodegenerative ailment that affects dopamine-producing neurons in a specific area of the brain called the substantia nigra of the ventral midbrain. It is clinically characterized by movement disorder and marked with unusual synaptic protein alpha-synuclein accumulation in the brain. To date, only a few Food and Drug Administration (FDA) approved drugs are available on the market for the treatment of PD. Nonetheless, these drugs show parasympathomimetic related adverse events and remarkably higher toxicity; hence, it is important to find more efficacious molecules to treat PD. In our study, We chosen 22 natural compounds as inhibitors that potentially block the alpha-synuclein clump—the pathological hallmark of PD—and provide new avenues for its treatment. Most of these molecules exhibited good pharmacokinetic behaviors, making them decisively favorable drug candidates to cure PD. Molecular docking studies were performed to investigate the binding interactions between natural compounds and alpha-synuclein as anti-Parkinson drug targets. Among the examined compounds, curcumin and piperine emerged as promising phytochemicals with the highest binding affinity, key residual stable bindings and showed a good inhibitory features. Thus, the present study indicates that curcumin and piperine hold the potential to be developed as treatment options against PD. Experimental validations are needed for insights into their mechanism of action and potential clinical application.Keywords
Supplementary Material
Supplementary Material FileParkinson’s disease (PD) is characterized by a decrease in the number of neurons in the substantia nigra and the formation of Lewy bodies (Zhu and Gong, 2020). PD is the second most common neurodegenerative disease, followed by Alzheimer’s disease (M’Angale and Staveley, 2017). PD is accompanied by motor deficit due to selective loss or degeneration of dopaminergic neurons encompasses striatum, substantia nigra, nigrostriatal pathway (Khan et al., 2010). Presently this disorder affects ~1% of the population over 65-year of age (Huang et al., 2012). The accumulation of alpha-synuclein is the pathological hallmark of PD, encoded by the SNCA gene and plaque formation in substantia nigra, also called Lewy bodies (Alecu and Bennett, 2019). Numerous clinical symptoms associated with PD are motor deficits, abnormal movement or bradykinesia, Akinesia, rigidity, resting tremor, and abnormality in gait (Jayaraj et al., 2013). Other symptoms include anxiety, psychosis, sleep disturbances, depression, and memory impairment (Gupta et al., 2018). Oxidative stress plays a key role in the pathogenesis of PD (Wei et al., 2018); it is caused by the overproduction of free radicals, which produces ROS that creates an imbalance between reactive oxygen species and the antioxidant defense system, which can cause damage to DNA, proteins, and lipids (Ghaffari et al., 2018). PD can be categorized into genetically linked and sporadic forms; 90% of the patients suffering from PD carry a sporadic/idiopathic form of the disease, while the remaining are associated with the genetic form. PD linked to genetic form has mutations in genes like LRRK2, Parkin, Pink1, DJ-1, and alpha-synuclein (Lopert and Patel, 2016). Although the root cause is still unknown, several risk factors, such as gender, age, and family history, play a crucial role in the development of PD (Roshan et al., 2016).
Moreover, the deficiency of dopamine has been found to be strongly associated with inflammatory response and characterized by reactive microglia with large cell bodies and short process (Cataldi et al., 2016). Various other aspects that increase the occurrence of ‘PD are excessive use of herbicides, milk products, and being overweight (Nicholatos et al., 2018). Neuroinflammation is thought to be the main factor in the pathogenesis of PD. The activation of microglia and aggregation of cytokine is found in postmortem PD brains and experimental PD models (Xu et al., 2017; Campolo et al., 2017). The pathogenesis of PD is still not fully explained, but several important intracellular processes are linked, such as mitochondrial dysfunction, endoplasmic reticulum stress, aggregation of misfolded protein, and imbalance between apoptosis and autophagy (Liu et al., 2018). The main factors which cause neuronal death in PD are oxidative stress and mitochondrial dysfunction (Zhang et al., 2010). Genetic evidence shows that a high expression of alpha-synuclein is associated with the pathological process of PD, although its physiological function is poorly known. The soluble oligomer type or fibrillar state of alpha-synuclein can be neurotoxic (Finkelstein et al., 2017).
Alpha-Synuclein is a small protein containing 140 amino acid residues encoded by the SNCA gene with a transcript length of 3041 bp (Siddiqui et al., 2016). The early onset of PD is associated with a missense mutation, and genetic studies have confirmed its involvement in both types of PD (familial and sporadic) (Stefanis, 2012). While the protein is primarily localized in the central nervous system (Oikawa et al., 2016), it is tremendously expressed in the peripheral nervous system and presynaptic terminals of the brain that connect the synaptic vesicle apparatus. The expression of alpha-synuclein is elevated within the synapses of nigral dopamine neurons (Longhena et al., 2017). Plant-derived natural compounds can prevent or inhibit the process of various neurological disorders such as PD and other neurological dysfunction. Naturally occurring phytochemicals such as flavonoids and polyphenols have been shown to possess several pharmacological properties which play an important role in preventing neurodegeneration disorders (Alam et al., 2018).
In this study, various naturally available molecules were selected based on the available literature, and their pharmacological properties, such as antioxidants, anti-inflammatory, antitumor, etc., were examined. The potency of these molecules was compared with another natural molecule based on the molecular docking score. The result revealed these natural molecules as potent inhibitors against PD.
A schematic workflow of this study is represented in Fig. 1.
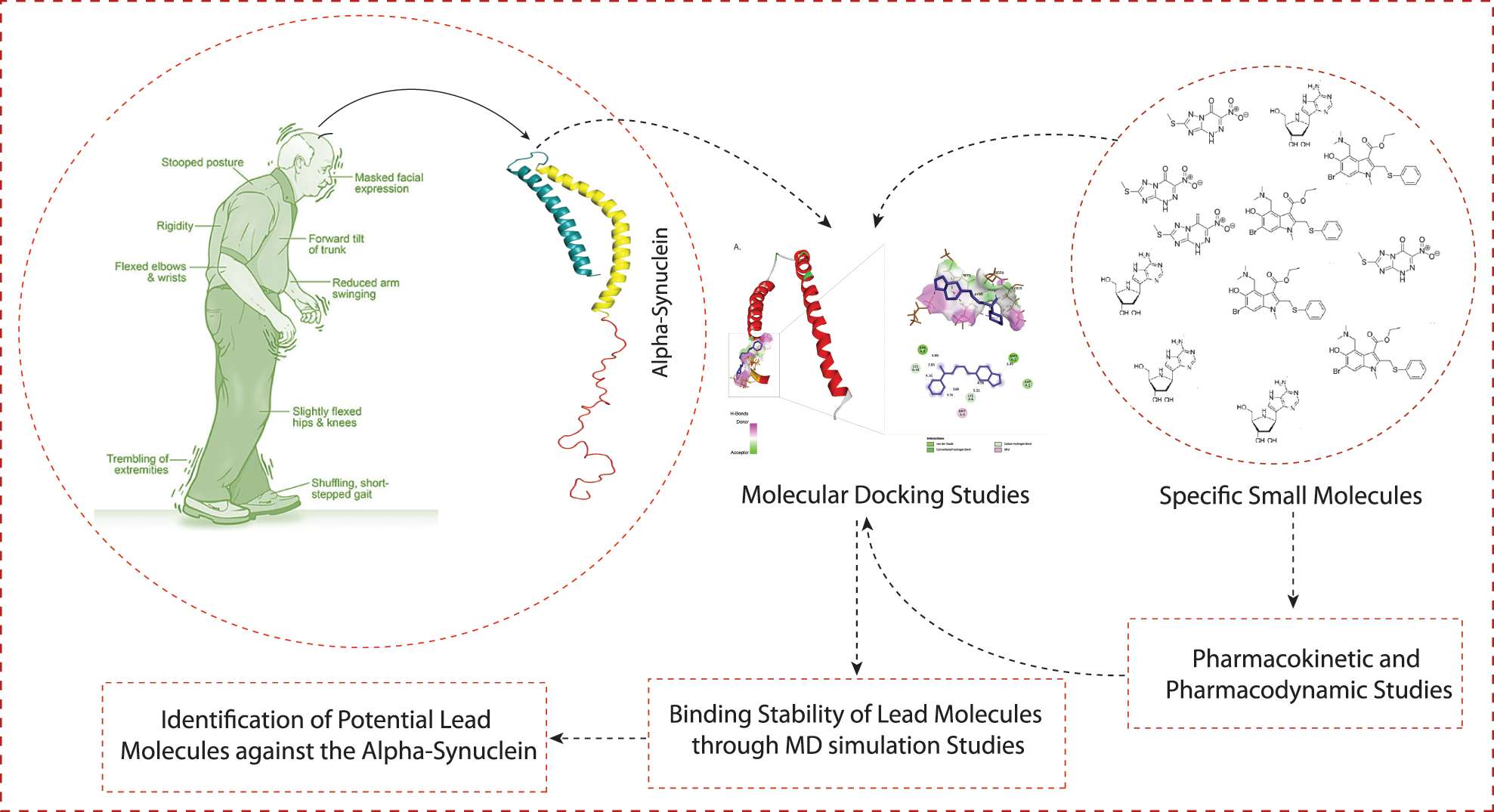
Figure 1: The schematic representation of workflow and methodology used in this study.
Selection of natural compounds
We selected natural compounds/molecules based on their symptomatic inhibition activity and good pharmacological evidence against various diseases. Selected molecules are potential drug candidates against various oxidative stress-based symptomatic diseases. The chemical structure of all these molecules or compounds was drawn using ChemBioDraw Ultra 12.0 (CambrigdeSoft), as shown in Fig. 2.
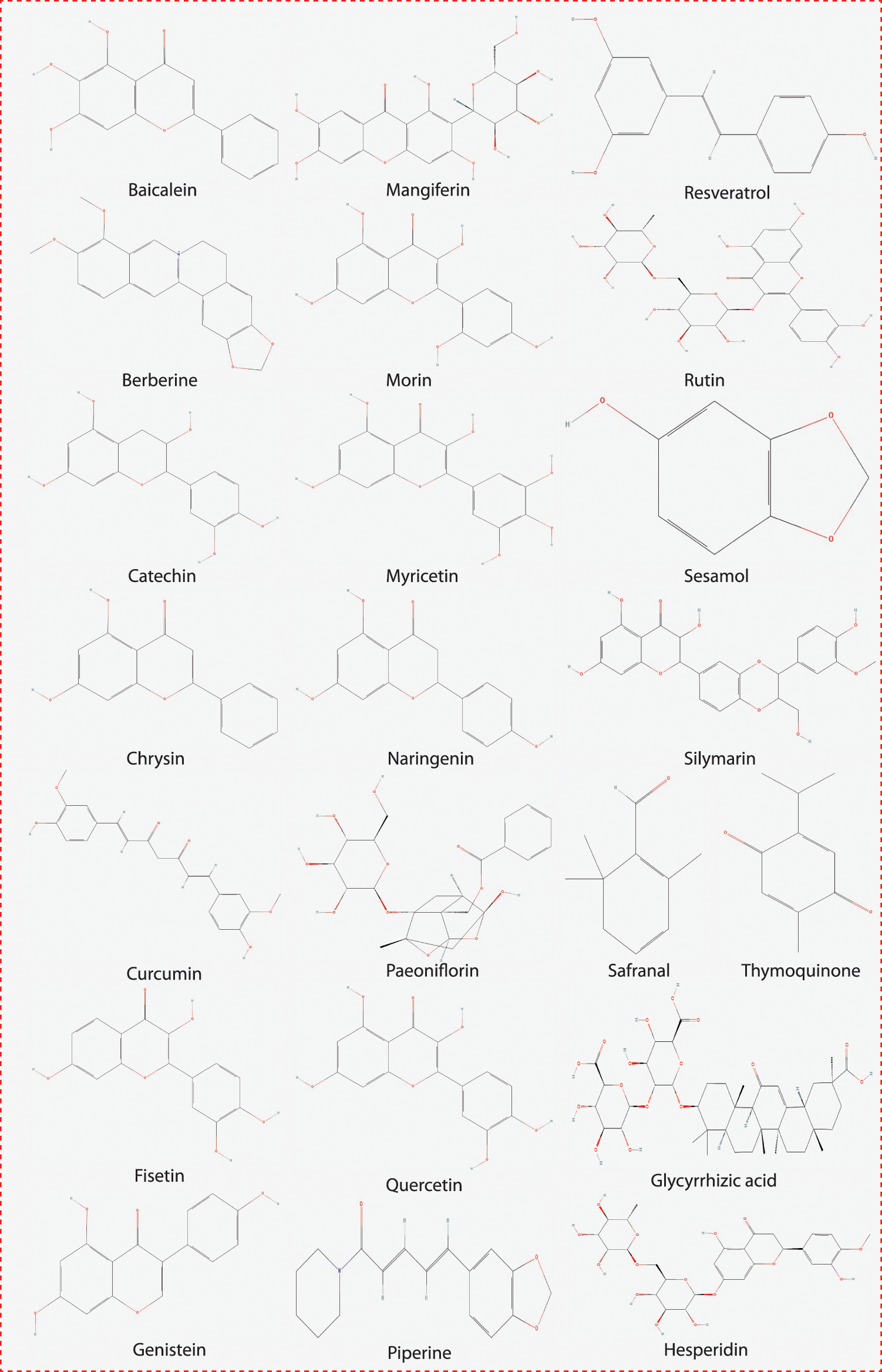
Figure 2: The chemical structure of the compounds examined in this study.
Calculation of basic pharmacokinetic parameters
To be considered a novel drug, a compound or molecule must be passed through multiple filters. In this way, most molecules that fail in the preclinical trial are eliminated from the queue of drug molecules (Wolohan and Clark, 2003). Pharmacokinetic properties, including absorption, distribution, metabolism, excretion, and toxicity, play a crucial role in drug discovery and design (Moroy et al., 2012). Physicochemical parameters such as MW, LogP, number of hydrogen bond donors (HBD), number of hydrogen bond acceptors (HBA), calculated topological polar surface area (TPSA), number of rotatable bonds (nrtB), and violations of Lipinski’s rule of five (n-Violation) was calculated using the in silico tool Molinspiration (Stierand and Rarey, 2010).
Prediction of the toxicity compounds
The prediction of toxicity of compounds is integral to drug design and development. In silico toxicity predictions are easy and rapid and can also minimize the number of animal usage. So, in this study, LD50 was estimated for all the selected natural bioactive markers. The LD50 is the lethal median dose required for killing 50% of a test population. It is the measure for evaluating the virulence of the medication or a poison. A lower LD50 dose is indicative of increased toxicity of the substance. Herein, an online tool, ProTox, was used to calculate these LD50 values (Drwal et al., 2014). Its carcinogenic, mutagenic, skin irritancy properties were determined using Discovery studio 2.5 (Accelrys Software Inc., San Diego, CA, USA) (Anon, 2009).
For this study, the amino acid sequence of the PD-associated target, alpha-synuclein (PDB ID: 1XQ8), was downloaded from the protein data bank (http://www.rcsb.org/pdb/home/home.do). For the crystal structure of targets, the crystallographic water molecules were removed, the missing hydrogen atoms were added, and the energy level of targets was minimized by the Swiss PDB viewer tool (Johansson et al., 2012).
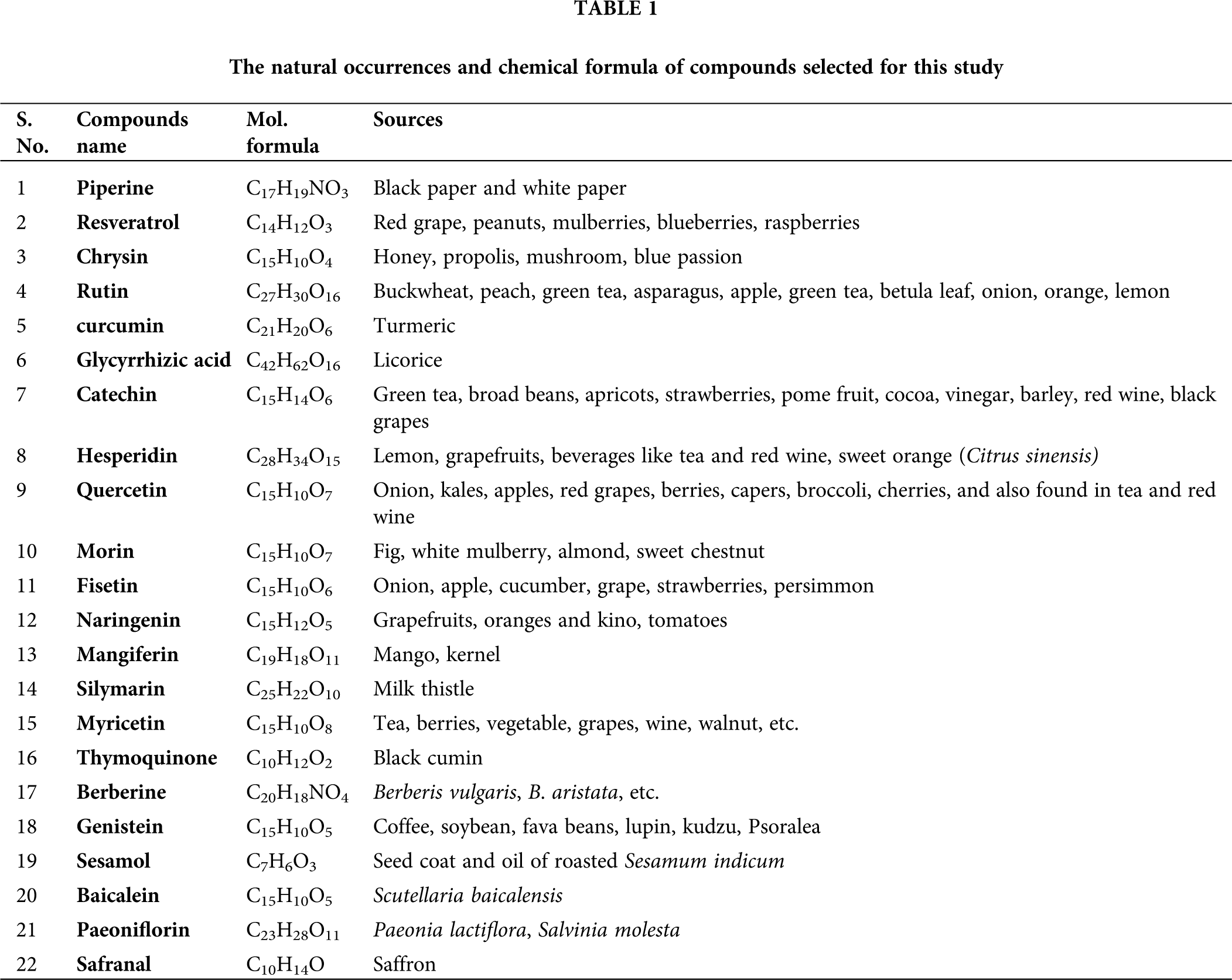
We selected 22 compounds based on their potential drug-like properties based on earlier reports. The structure of the selected 22 compounds was downloaded from the PubChem database to their 3D form, and ChemBio3DUltra was used to optimize the geometry of the compounds (PerkinElmer Informatics, Waltham, MA, USA). The list of compounds and their sources are listed in Table 1.
Molecular docking and molecular dynamics study
Docking studies generate important information regarding the orientation of the different compounds within the binding pocket of the target protein. In this study, 22 target compounds and four previously reported inhibitors (metaproterenol, clenbuterol, salbutamol, and gallic acid) were used as control (reference) drugs for their inhibition activity against the alpha-synuclein aggregation (Suppl. Fig. S1) were used as control (reference) drugs. All these compounds were docked individually to the alpha-synuclein with a grid center (grid coordination) and grid boxes of certain sizes for each receptor. The flexibility of the compounds was maintained when they interacted with the target protein under rigid conditions. Kollman charges and polar hydrogen atoms were included in the surviving structure. The grid size was set at
Further, MD simulations were performed using the GROMACS version 5.1 software using the standard GROMOS
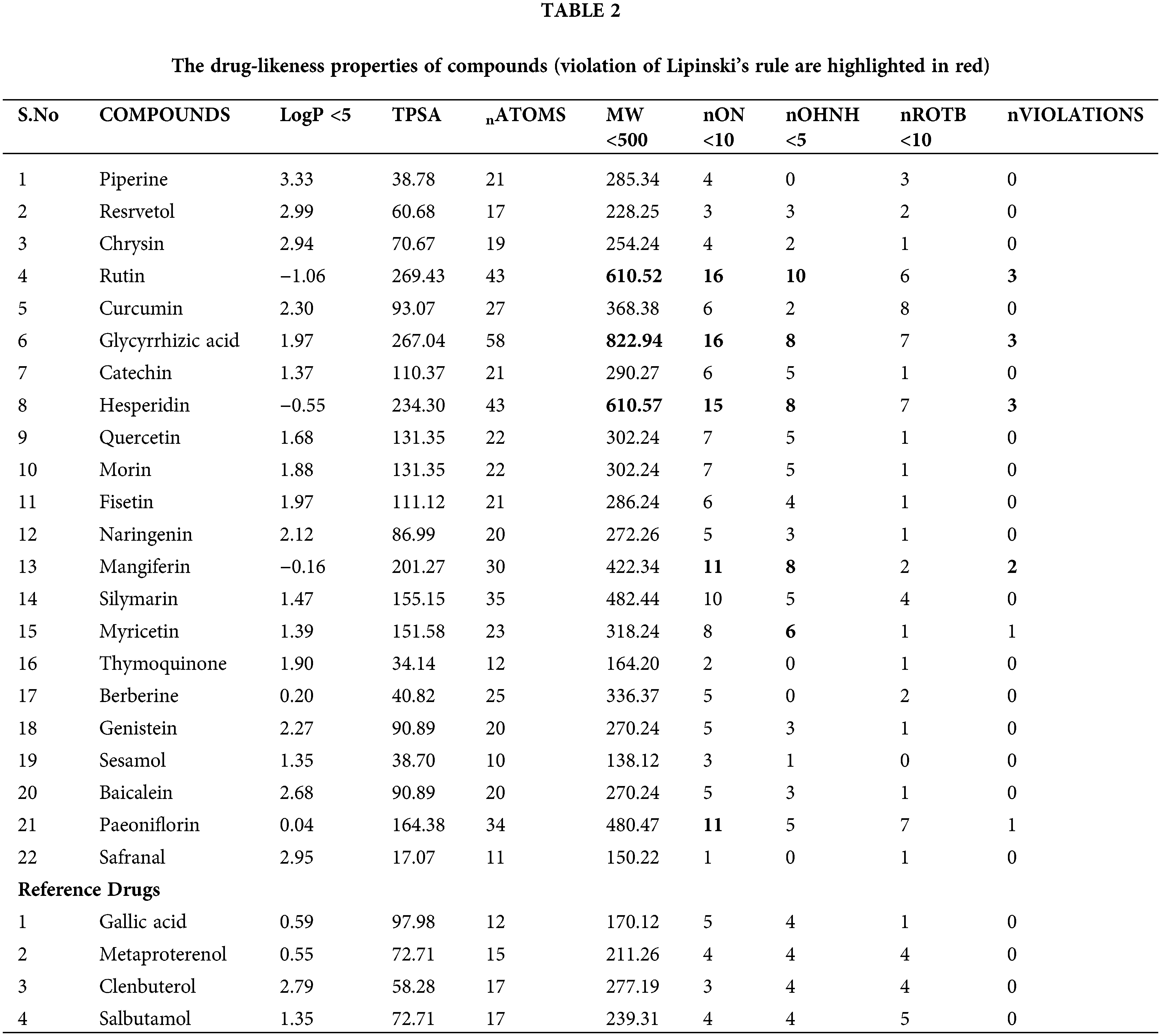
The bioactive markers must possess certain properties and pass basic filters like Lipinski’s rule of five (RO5) to be considered suitable drug candidates. Evaluation results of drug-likeness properties among selected compounds showed that the highest binding affinity of lead molecules followed Lipinski’s RO5, and it has a high potential to be used as a drug. “Lipinski rule of five” is a well-known factor in producing a drug moiety; proven available data from the analysis of the world drug index and helps enhance the drug properties, which possesses all the characteristics to be administered as an orally active drug molecule. This rule is formulated and followed by most orally administered drugs. Four different criteria were used to decide if a molecule is a drug such as (a) having a molecular weight of ≤500 kda, (b) LogP (logarithm of partition coefficient) ≤5, (c) to have ≤5 hydrogen bond donor sites, and (d) hydrogen bond acceptor sites should be ≤10. Molecules that violate the aforesaid rules are believed to have bioavailability-related problems. So given that in our case, most of the compounds passed the RO5 except rutin, glycyrrhizic acid, hesperidin, and mangiferin, which can thus have issues with bioavailability when taken orally.
TPSA measurement was used to determine the bioavailability of all-natural bio-active compounds in the study; it can be performed using Veber’s rule. For good oral bioavailability, TPSA measurement says that the molecule must have a rotatable bond, which is ≤10, and the TPSA values
Toxicity is another important aspect of the drug that needs to be accessed before considering any molecules as a potential drug. We used the online software PROTOX to predict the LD50 value of compounds. Expect a few, the bioactive moiety of all compounds in the current study were in non-toxic zone, with LD50 of more than 1000 mg/kg mark, except for piperine (330 mg/kg), quercetin (159 mg/kg), fisetin (159 mg/kg), mangiferin (2 mg/kg), myricetin (159 mg/kg), berberine (200 mg/kg), and sesamol (580 mg/kg). Besides, carcinogenicity, immunotoxicity, mutagenicity, and cytotoxicity were also predicted, and most of the compounds did not show any of the above toxicities, although a few compounds exhibited potentially severe toxicity problems; a detailed analysis of the results is listed in Table 3. The toxicity of the chemical compound was determined in the form of toxicity endpoints such as mutagenicity, carcinogenicity, some other endpoints, etc.
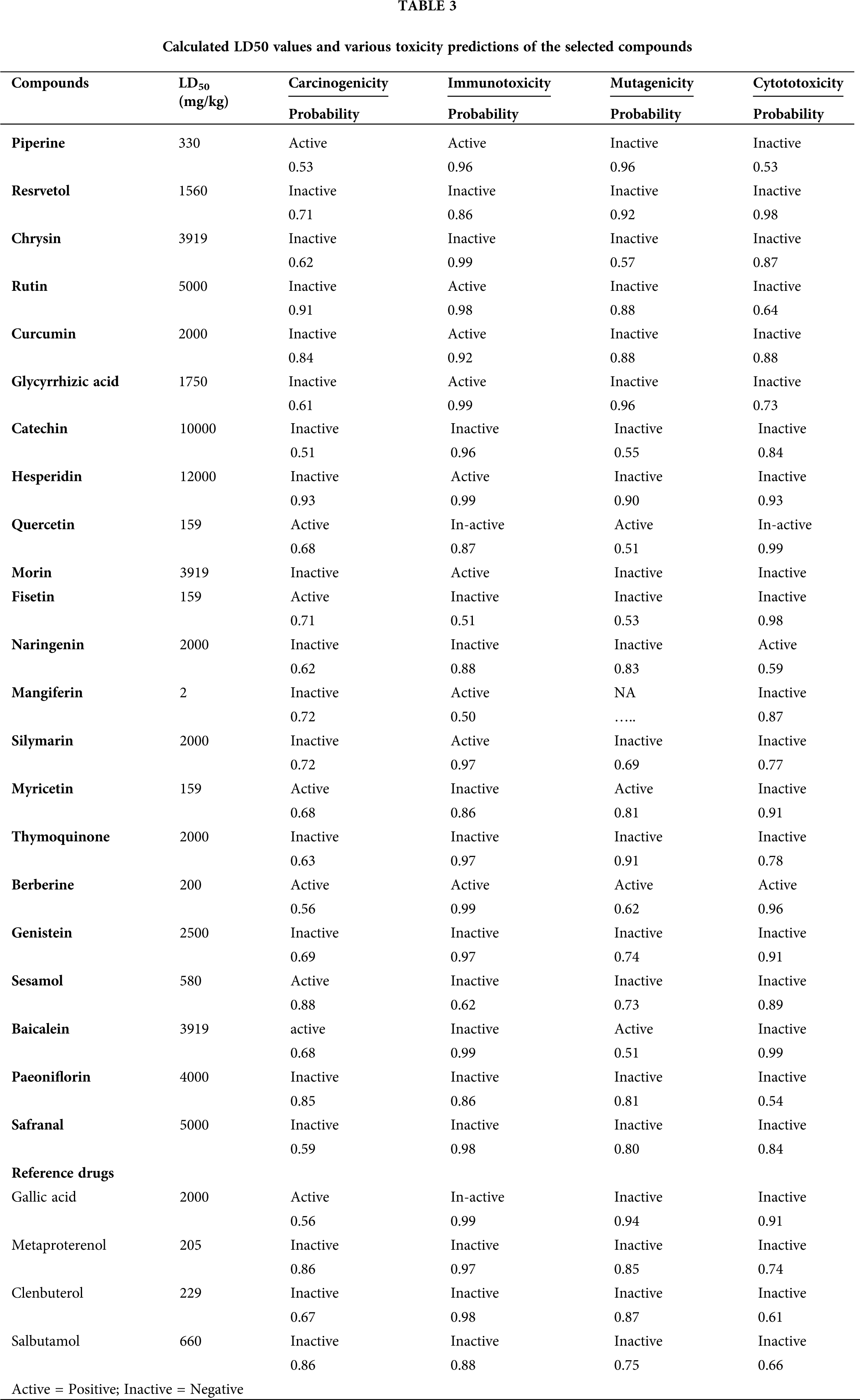
Further, moving up, toxicity may also measure quantitatively (LD50_lethal dose); qualitatively (active or inactive) both as well for special cell types followed by assays. For those compounds assumed as toxicity targets having very little binding possibility, a compound may be active for several toxicity endpoints, resulting in severe toxic effects. Hence, this assay serves as a crucial preclinical trial and is economical and time-saving.
To make sure the binding between the potential bioactive moiety of compounds (Table 3) and PD target (1X08), we performed molecular docking analysis (the score with the inhibition constant is mentioned in Table 4) and two compounds, namely curcumin and piperine show the highest score, bind with a good binding affinity (−9.10 kcal/mol and −8.6 kcal/mol, respectively) and form H-bonds with the residues, e.g., SER 9, MET 1, ASP 2. Piperine with 1X08 forms H-bonds with residues LYS 97, PHE 94, and LYS 96 (Fig. 3). Additionally, the molecular docking outcomes near the ligand surrounding amino acid residue (within 4 Å) show polar interaction over a small range that stabilized the complex formation (delocalization of charges with the help of ligands).
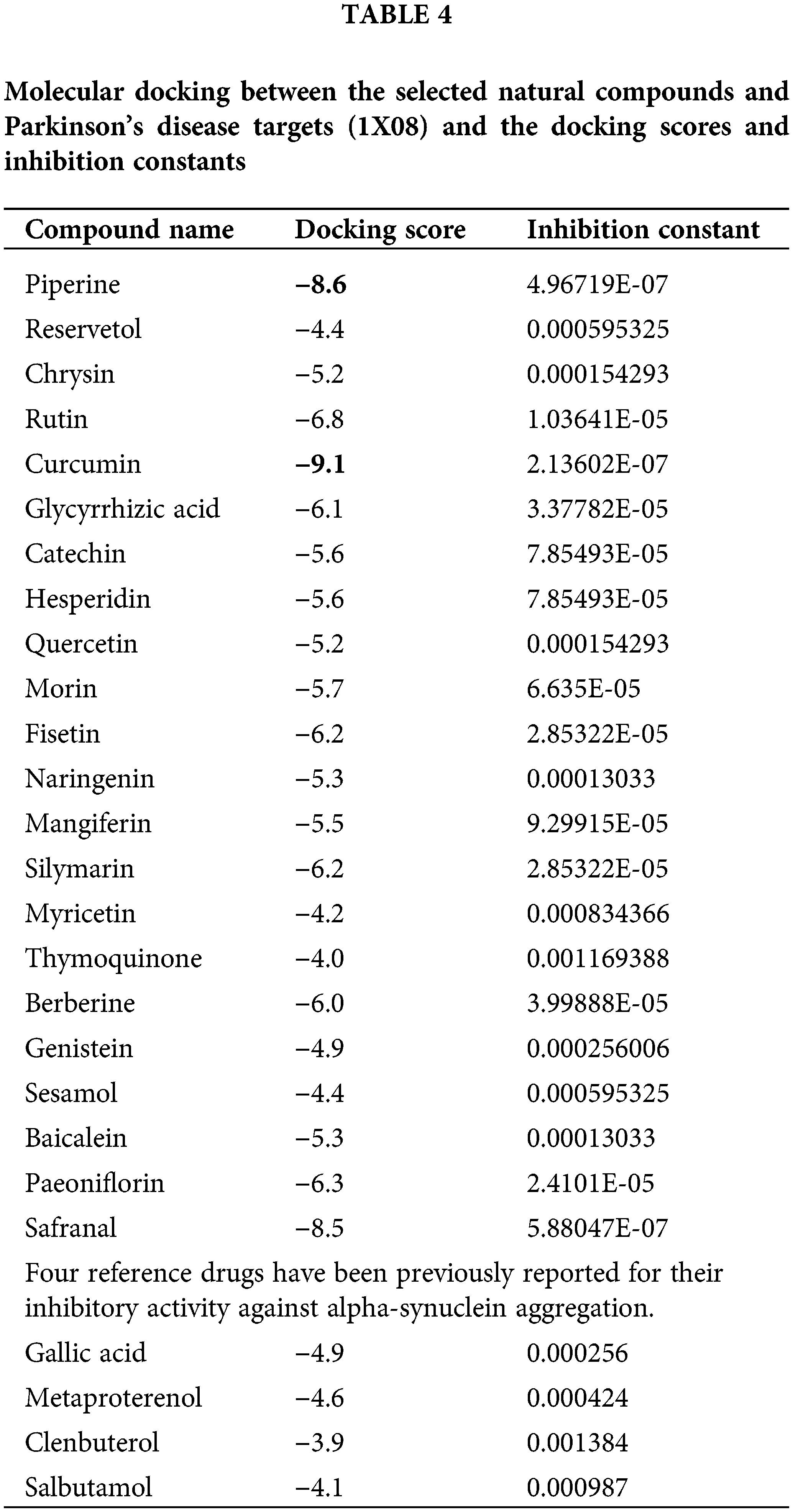
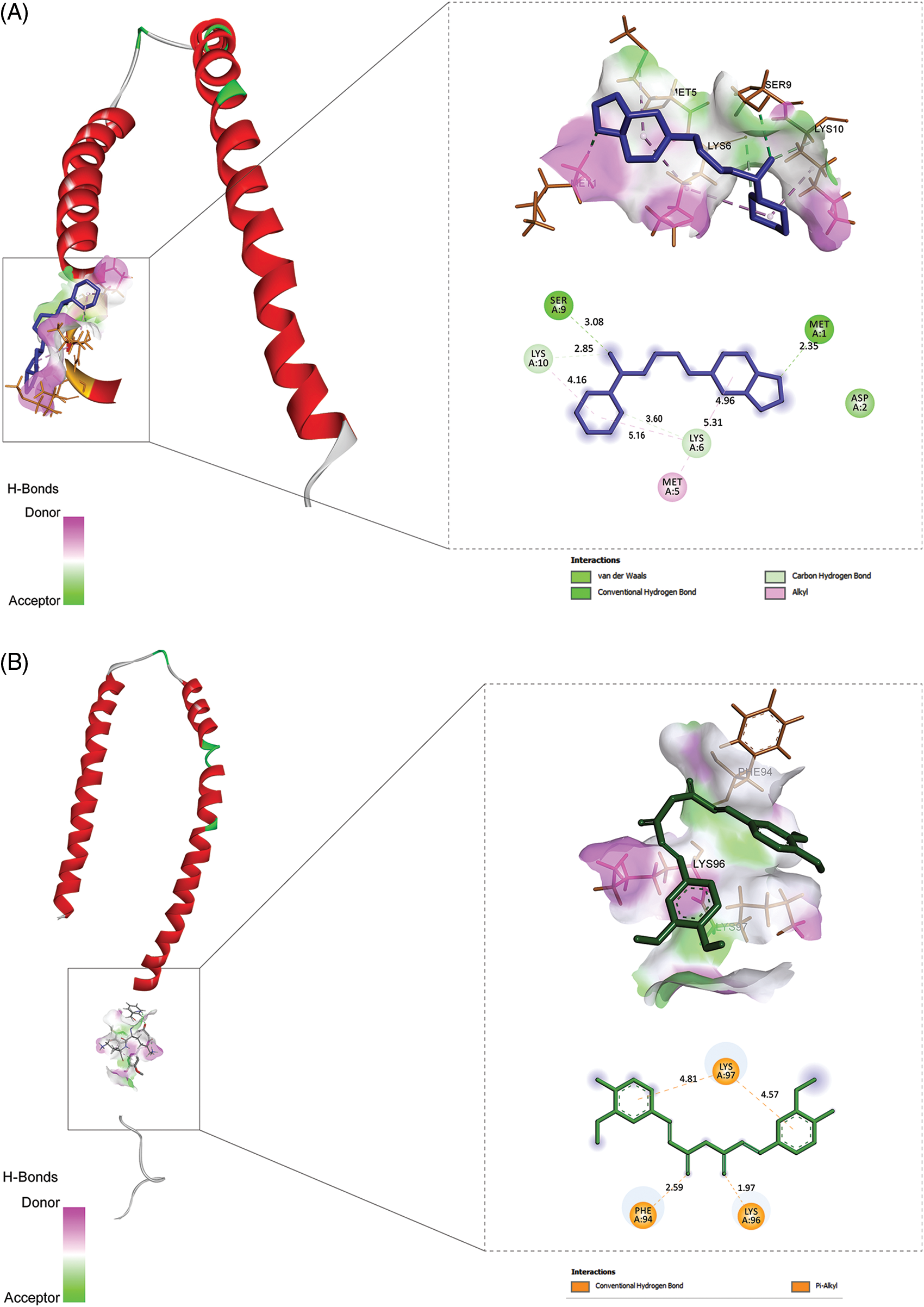
Figure 3: The results of molecular docking studies: (A) Piperine and 1XQ8 (B) Curcumin and 1XQ8.
Simulation results of molecular docking
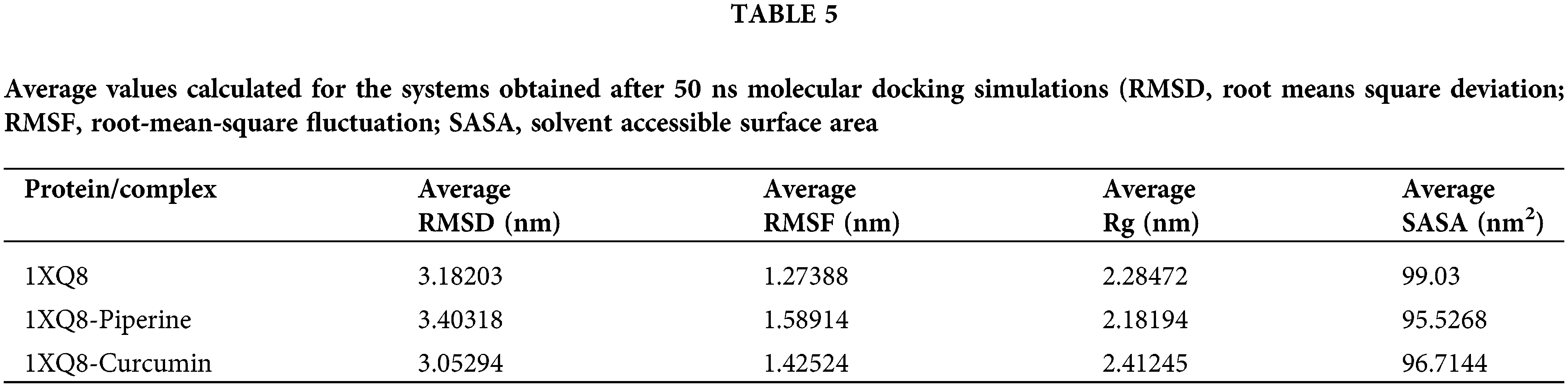
MD simulations of all atoms were performed for 50 ns to understand the mechanism of interaction of curcumin and piperine with 1X08. Several analyses were made using trajectories generated by simulations. The average root means square deviation (RMSD), the root-mean-square fluctuation (RMSF), solvent accessible surface area (SASA), and radius of gyration (Rg) were estimated, and the results are mentioned in Table 5.
Structural deviations and compactness
To evaluate conformational changes of alpha-synuclein upon ligand binding, RMSD was determined to estimate the stability of the system under-exploited solvent conditions. The RMSD plot suggested that the binding of curcumin and piperine stabilized 1X08 and led to fewer structural deviations from its native conformation. In Fig. 4A, curcumin and 1X08 complexes showed distinct deviation until 25 ns from the starting point. However, the complex appeared to stabilize until the end of the simulation. Although piperine followed the same pattern of deviation in this study, both the curcumin and piperine were bound perfectly into the cavity of 1X08 protein and stabilized the system; this stability is necessary to stop the action of proteins after the binding of curcumin and piperine.
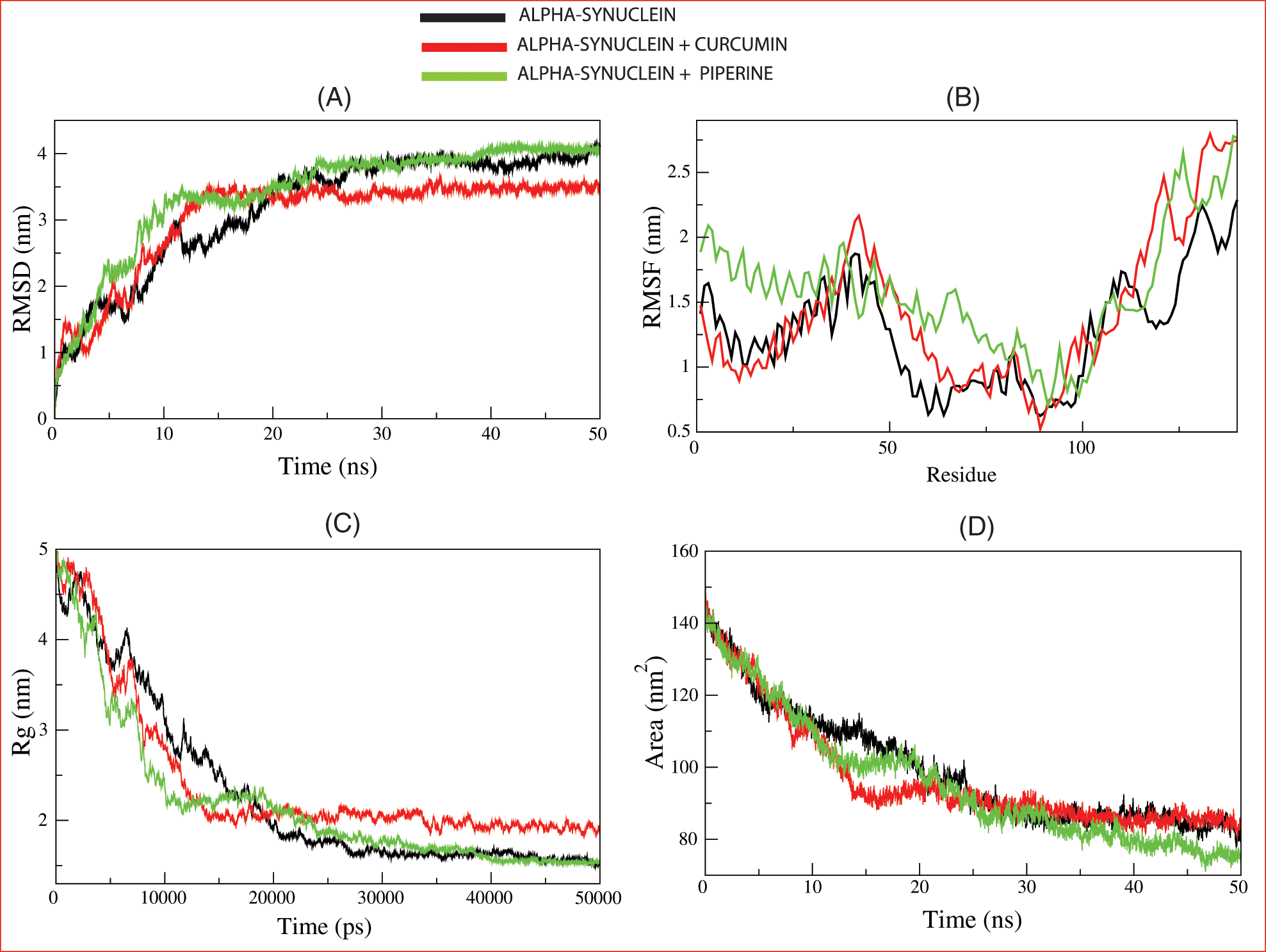
Figure 4: The molecular dynamic simulation of the lead compounds with the Parkinson disease target: (A) root means square deviation (RMSD), (B) root-mean-square fluctuation (RMSF), (C) Radius of gyration (Rg), and (D) solvent accessible surface area (SASA).
To calculate the average fluctuation of all residues during the simulation, the RMSF of the binding of IX08 upon ligands was plotted as a function of residue number, as shown in Fig. 4B. RMSF plot showed the presence of several residual fluctuations in different regions of the IX08 structure. These residual fluctuations were found to increase upon binding of curcumin and piperine in the region spanning in N-terminal to C-terminal and affect the dynamics behavior of residues. RMSF values fluctuated highly in IX08-curcumin, starting from residue 48, and in IX08-piperine, after residues 65. These findings indicate that the IX08 complexes have smaller RMSF values than the native IX08 structure. Thus, the observed drifting was accompanied by increased atomic fluctuations in the complex structure. The major fluctuations suggested the profound importance of these residues in the ligand-binding process.
Rg is linked to the three-dimensional volume of a protein and is applied to assess the stability of the protein in a biological system. A protein is supposed to have a higher Rg due to less tight packing. Here in our study, the Rg plot suggested more tight packing of 1X08 upon the binding of curcumin and piperine Fig. 4C. Minimal structural deviation can be seen in the Rg plot, and no conformational shift was observed after the binding of both drugs. The Rg plot showed a tight packing of 1X08 proteins from the initial to the endpoint of simulation after the binding of drugs, suggesting that both the drugs were bound tightly with the proteins and may stop the action of proteins.
Solvent accessible surface area
SASA is a protein’s surface area interacting with its solvent and indicates the folding-unfolding state and stability of the protein. The average SASA values for IX08-curcumin and IX08-Piperine were monitored during 50 ns MD simulations. The complexes presented here showed a lowered value of SASA with time, while the native IX08 structure showed a slightly larger SASA value Fig. 4D.
PD is the second-fastest-growing neurodegenerative disorder worldwide after Alzheimer’s disease and highly affects the elderly (Cornejo et al., 2021). The alpha-synuclein oligomer and fibrils are crucial for PD pathogenesis. A previous study has shown that the toxicity is caused by oligomers and dopaminergic cell death in the rat model of PD (Cascella et al., 2021). Hence an effective therapeutic approach could be by inhibiting the oligomerization of toxic oligomers into fibrils. Many recent studies have suggested the reduction of toxic oligomers via the modulation of natural compounds. In this perspective, curcumin has been shown to be effective against various diseases such as cancer, diabetes, cardiovascular, and other neurological disorders and might be a promising therapeutic agent. Curcumin possesses antioxidant and anti-inflammatory properties and detoxifies the effects of reactive oxygen species (Jha et al., 2016). It is a natural compound mainly found in the root of the Curcuma longa plant and is the main component of turmeric spices. Curcumin is biologically most active and abundant among the three curcuminoids (Spinelli et al., 2015). Research has proven that this compound possesses antioxidant, anti-inflammatory, cardioprotective, neuroprotective (Wang et al., 2020), anti-apoptotic (El Nebrisi et al., 2020), and chemopreventive (Scapagnini et al., 2011) properties. It also exhibits free radical scavenging activity, and such properties could combat nitrative and oxidative stress in various neurodegenerative disorders like PD and Alzheimer’s disease. Studies have strongly favored the clinical manifestation of curcumin in a different model of PD (Hu et al., 2015).
Besides, in one study, piperine attenuated the memory performance and enhanced myelin repair in Lys phosphatidylcholine-induced animal model of a rat via anti-inflammatory, antioxidants, and neuroprotective effects. Therefore, treatment with piperine increased the level of antioxidant capacity. Evidence-based experimental findings have proven that piperine is a promising therapeutic target for myelin repair in multiple sclerosis and improves memory performance (Roshanbakhsh et al., 2020). It is also effective against intracerebroventricular cognitive impairment induced by streptozotocin injections. Piperine could improve memory impairment and decrease amyloid-beta deposition through a neuroprotective antioxidant mechanism (Wang et al., 2019) and exhibit various pharmacological properties such as antidepressant, cognitive analgesic, anti-inflammatory, cytoprotective, and antioxidant effects. Compared with chemical entities, piperine has many beneficial properties, such as easily available natural plant material, low cost, well known, and safe to use (Ashour et al., 2016).
The impairment of the antioxidant defense mechanism is caused by enhanced oxidative stress in the brain and peripheral tissues in Alzheimer’s disease, amyotrophic lateral sclerosis, and PD. To date, available treatments for neurodegenerative diseases are not appropriately effective and have side effects. The standard gold treatment of PD using levodopa has copious side effects (Cornejo et al., 2021). Most of the drugs have failed to inhibit or limit the progress of Perkinson’s disease (Niedzielska et al., 2016). There is a global need for a new therapeutic alternative treatment for neurodegenerative disease and PD. Curcumin has been reported to possibly cross the blood-brain barrier and lead to several improvements in the pathological process of neurodegenerative disease (Salehi et al., 2020).
Molecular docking, homology modeling, and MD simulation are proven tools for the identification of potent inhibitors. Its approaches are frequently used to understand ligands-receptor interaction in computational drug designing. Apart from the in vitro and in vivo studies, these computational techniques strongly support manufacturing or designing novel drugs in the pharmaceutical area by deciphering the mechanism of drug-receptor interaction (Srivastava et al., 2010; Khan et al., 2021). In this study, curcumin and piperine were found to be comparatively more suitable ligands. The result suggested that the stability of the proteins increased after docking, and both the ligands were present for a longer period in the pocket of the protein (Table 4). Herein, curcumin and piperine were observed to bind to the active site of the alpha-synuclein protein and inhibit the self-association of targeted protein to act as drug candidates for PD. Earlier studies suggest diverse pharmacological properties of curcumin and piperine (Yu et al., 2017). Alpha-synuclein is the most characteristic protein of PD; therefore, it has important applications in therapeutics and clinical diagnosis (Dehay et al., 2015).
The present in silico study gives a better insight into the structure and function of alpha-synuclein. Several plant-based compounds are combined with cognitive protective compounds or molecules to improve neurological disorders in PD patients. Our current study expands the understanding of natural molecules or compounds, showing their potential in treating PD. Most importantly, we found two compounds (curcumin and piperine) as variable multi-target-directed compounds with antioxidant and neuroprotective activities. So, these molecules may be prospective therapy for PD and other neurobehavioral diseases. Further studies are crucially important to explore the specific targets in various brain regions and the mode of action, including the signaling pathway, blood-brain barrier (BBB), and mechanism of synergetic effect of the antioxidant agent on the target. This pharmaco-chemical approach can open new avenues to design effective, potential herbal moieties for the treatment of PD with negligible side effects.
Acknowledgement: The authors thank the Department of Science and Technology (DST), Government of India, and Jamia Hamdard for providing financial assistance to Rahat Ali during this experimental study.
Availability of Data and Materials: All data generated or analyzed during this study are included in this published article (and its supplementary information files).
Supplementary Material: The supplementary table is available online at DOI: https://10.32604/biocell.2022.021224.
Authors’ Contributions: RA and SKR conceived the study design and analyzed the data. R Ali curated data and performed statistical analyses. AA curated data and prepared figures. R Ali and AA performed molecular docking studies. All the authors read, edited, and approved the final manuscript.
Funding Statement: This work did not receive any specific grant from funding agencies in the public, commercial, or not-for-profit sectors.
Conflicts of Interest: The authors declare that they have no conflicts of interest to report regarding the present study.
References
Alam A, Tamkeen N, Imam N, Farooqui A, Ahmed MM et al. (2018). Pharmacokinetic and molecular docking studies of plant-derived natural compounds to exploring potential anti-Alzheimer Activity, 2018. In: Choudhary DK, Kumar M, Prasad R, Kumar V (eds.In Silico Approach for Sustainable Agriculture, pp. 217–238. Singapore: Springer Singapore. [Google Scholar]
Alecu I, Bennett SAL (2019). Dysregulated lipid metabolism and its role in α-synucleinopathy in Parkinson’s disease. Frontiers in Neuroscience 13: 1–22. DOI 10.3389/fnins.2019.00328. [Google Scholar] [CrossRef]
Anon (2009). Discovery Studio Modeling Environment, Release 2.5. San Diego, CA: Accelrys Software Inc. [Google Scholar]
Anon BIOVIA, Dassault Systèmes, BIOVIA Workbook, Release (2020). BIOVIA pipeline pilot, release 2020. San Diego: Dassault Systèmes. [Google Scholar]
Ashour EA, Majumdar S, Alsheteli A, Alshehri S, Alsulays B, Feng X, Gryczke A, Kolter K, Langley N, Repka MA (2016). Hot melt extrusion as an approach to improve solubility, permeability and oral absorption of a psychoactive natural product, piperine. Journal of Pharmacy and Pharmacology 68: 989–998. DOI 10.1111/jphp.12579. [Google Scholar] [CrossRef]
Bjelkmar P, Larsson P, Cuendet MA, Hess B, Lindahl E (2010). Implementation of the CHARMM force field in GROMACS: Analysis of protein stability effects from correction maps, virtual interaction sites, and water models. Journal of Chemical Theory and Computation 6: 459–466. DOI 10.1021/ct900549r. [Google Scholar] [CrossRef]
Campolo M, Casili G, Biundo F, Crupi R, Cordaro M, Cuzzocrea S, Esposito E (2017). The neuroprotective effect of dimethyl fumarate in an MPTP-mouse model of Parkinson’s disease: Involvement of reactive oxygen species/nuclear factor-κB/nuclear transcription factor related to NF-E2. Antioxidants and Redox Signaling 27: 453–471. DOI 10.1089/ars.2016.6800. [Google Scholar] [CrossRef]
Cascella R, Chen SW, Bigi A, Camino JD, Xu CK et al. (2021). The release’ of toxic oligomers from α-synuclein fibrils induces dysfunction in neuronal cells. Nature Communications 12: 1–16. DOI 10.1038/s41467-021-21937-3. [Google Scholar] [CrossRef]
Cataldi S, Codini M, Hunot S, Légeron FP, Ferri I et al. (2016). e-Cadherin in 1-methyl-4-phenyl-1,2,3,6-tetrahydropyridine-induced Parkinson disease. Mediators of Inflammation 2016: 1–7. DOI 10.1155/2016/3937057. [Google Scholar] [CrossRef]
Cheatham TEI, Miller JL, Fox T, Darden TA, Kollman PA (1995). Molecular dynamics simulations on solvated biomolecular systems: The particle mesh Ewald method leads to stable trajectories of DNA, RNA, and proteins. Journal of the American Chemical Society 117: 4193–4194. DOI 10.1021/ja00119a045. [Google Scholar] [CrossRef]
Cornejo A, Caballero J, Simirgiotis M, Torres V, Sánchez L et al. (2021). Dammarane triterpenes targeting α-synuclein: Biological activity and evaluation of binding sites by molecular docking. Journal of Enzyme Inhibition and Medicinal Chemistry 36: 154–162. DOI 10.1080/14756366.2020.1851216. [Google Scholar] [CrossRef]
Dehay B, Bourdenx M, Gorry P, Przedborski S, Vila M et al. (2015). Targeting α-synuclein for treatment of Parkinson’s disease: Mechanistic and therapeutic considerations. The Lancet Neurology 14: 855–866. DOI 10.1016/S1474-4422(15)00006-X. [Google Scholar] [CrossRef]
Drwal MN, Banerjee P, Dunkel M, Wettig MR, Preissner R (2014). ProTox: A web server for the in silico prediction of rodent oral toxicity. Nucleic Acids Research 42: W53–W58. DOI 10.1093/nar/gku401. [Google Scholar] [CrossRef]
Finkelstein DI, Billings JL, Adlard PA, Ayton S, Sedjahtera A et al. (2017). The novel compound PBT434 prevents iron mediated neurodegeneration and alpha-synuclein toxicity in multiple models of Parkinson’s disease. Acta Neuropathologica Communications 5: 53. DOI 10.1186/s40478-017-0456-2. [Google Scholar] [CrossRef]
Ghaffari F, Hajizadeh Moghaddam A, Zare M (2018). Research paper: Neuroprotective effect of quercetin nanocrystal in a 6-hydroxydopamine model of Parkinson disease: Biochemical and behavioral evidence. Basic and Clinical Neuroscience Journal 9: 317–324. DOI 10.32598/bcn.9.5.317. [Google Scholar] [CrossRef]
Gupta M, Kant K, Sharma R, Kumar A (2018). Evaluation of in silico anti-Parkinson potential of β-asarone. Central Nervous System Agents in Medicinal Chemistry 18: 128–135. DOI 10.2174/1871524918666180416153742. [Google Scholar] [CrossRef]
Hu S, Maiti P, Ma Q, Zuo X, Jones MR, Cole GM, Frautschy SA (2015). Clinical development of curcumin in neurodegenerative disease. Expert Review of Neurotherapeutics 15: 629–637. DOI 10.1586/14737175.2015.1044981. [Google Scholar] [CrossRef]
Huang HC, Chang P, Dai XL, Jiang ZF (2012). Protective effects of curcumin on amyloid-β-induced neuronal oxidative damage. Neurochemical Research 37: 1584–1597. DOI 10.1007/s11064-012-0754-9. [Google Scholar] [CrossRef]
Jayaraj RL, Ranjani V, Manigandan K, Elangovan N (2013). In silico docking studies to identify potent inhibitors of alpha-synuclein aggregation in Parkinson disease. Asian Journal of Pharmaceutical and Clinical Research 6: 127–131. [Google Scholar]
Jha NN, Ghosh D, Das S, Anoop A, Jacob RS, Singh PK, Ayyagari N, Namboothiri INN, Maji SK (2016). Effect of curcumin analogs on α-synuclein aggregation and cytotoxicity. Scientific Reports 6: 28511. DOI 10.1038/srep28511. [Google Scholar] [CrossRef]
Johansson MU, Zoete V, Michielin O, Guex N (2012). Defining and searching for structural motifs using DeepView/Swiss-PdbViewer. BMC Bioinformatics 13: 173. DOI 10.1186/1471-2105-13-173. [Google Scholar] [CrossRef]
Khan MM, Ahmad A, Ishrat T, Khan MB, Hoda MN et al. (2010). Resveratrol attenuates 6-hydroxydopamine-induced oxidative damage and dopamine depletion in rat model of Parkinson’s disease. Brain Research 1328: 139–151. DOI 10.1016/j.brainres.2010.02.031. [Google Scholar] [CrossRef]
Khan N, Bhat R, Patel AK, Ray P (2021). Discovery of small molecule inhibitors of chikungunya virus proteins (nsP2 and E1) using in silico approaches. Journal of Biomolecular Structure and Dynamics 39: 1373–1385. DOI 10.1080/07391102.2020.1731602. [Google Scholar] [CrossRef]
Liu J, Liu W, Lu Y, Tian H, Duan C, Lu L, Gao G, Wu X, Wang X, Yang H (2018). Piperlongumine restores the balance of autophagy and apoptosis by increasing BCL2 phosphorylation in rotenone-induced Parkinson disease models. Autophagy 14: 845–861. DOI 10.1080/15548627.2017.1390636. [Google Scholar] [CrossRef]
Longhena F, Faustini G, Missale C, Pizzi M, Spano P, Bellucci A (2017). The contribution of α-synuclein spreading to Parkinson’s disease synaptopathy. Neural Plasticity 2017: 1–15. DOI 10.1155/2017/5012129. [Google Scholar] [CrossRef]
Lopert P, Patel M (2016). Mitochondrial mechanisms of redox cycling agents implicated in Parkinson’s disease. Journal of Neural Transmission 123: 113–123. DOI 10.1007/s00702-015-1386-4. [Google Scholar] [CrossRef]
M’Angale PG, Staveley BE (2017). A loss of Pdxk model of Parkinson disease in Drosophila can be suppressed by Buffy. BMC Research Notes 10: 205. DOI 10.1186/s13104-017-2526-8. [Google Scholar] [CrossRef]
Moroy G, Martiny VY, Vayer P, Villoutreix BO, Miteva MA (2012). Toward in silico structure-based ADMET prediction in drug discovery. Drug Discovery Today 17: 44–55. DOI 10.1016/j.drudis.2011.10.023. [Google Scholar] [CrossRef]
El Nebrisi E, Javed H, Ojha SK, Oz M, Shehab S (2020). Neuroprotective effect of curcumin on the nigrostriatal pathway in a 6-hydroxydopmine-induced rat model of Parkinson’s disease is mediated by α7-nicotinic receptors. International Journal of Molecular Sciences 21: 7329. DOI 10.3390/ijms21197329. [Google Scholar] [CrossRef]
Nicholatos JW, Francisco AB, Bender CA, Yeh T, Lugay FJ, Salazar JE, Glorioso C, Libert S (2018). Nicotine promotes neuron survival and partially protects from Parkinson’s disease by suppressing SIRT6. Acta Neuropathologica Communications 6: 120. DOI 10.1186/s40478-018-0625-y. [Google Scholar] [CrossRef]
Niedzielska E, Smaga I, Gawlik M, Moniczewski A, Stankowicz P, Pera J, Filip M (2016). Oxidative stress in neurodegenerative diseases. Molecular Neurobiology 53: 4094–4125. DOI 10.1007/s12035-015-9337-5. [Google Scholar] [CrossRef]
Oikawa T, Nonaka T, Terada M, Tamaoka A, Hisanaga S, Hasegawa M (2016). α-Synuclein fibrils exhibit gain of toxic function, promoting Tau aggregation and inhibiting microtubule assembly. Journal of Biological Chemistry 291: 15046–15056. DOI 10.1074/jbc.M116.736355. [Google Scholar] [CrossRef]
Roshan MHK, Tambo A, Pace NP (2016). Potential role of caffeine in the treatment of Parkinson’s disease. The Open Neurology Journal 10: 42–58. DOI 10.2174/1874205X01610010042. [Google Scholar] [CrossRef]
Roshanbakhsh H, Elahdadi Salmani M, Dehghan S, Nazari A, Javan M, Pourabdolhossein F (2020). Piperine ameliorated memory impairment and myelin damage in lysolecethin induced hippocampal demyelination. Life Sciences 253: 117671. DOI 10.1016/j.lfs.2020.117671. [Google Scholar] [CrossRef]
Salehi B, Calina D, Docea A, Koirala N, Aryal S et al. (2020). Curcumin’s nanomedicine formulations for therapeutic application in neurological diseases. Journal of Clinical Medicine 9: 430. DOI 10.3390/jcm9020430. [Google Scholar] [CrossRef]
Scapagnini G, Sonya V, Nader AG, Calogero C, Zella D, Fabio G (2011). Modulation of Nrf2/ARE pathway by food polyphenols: A nutritional neuroprotective strategy for cognitive and neurodegenerative disorders. Molecular Neurobiology 44: 192–201. DOI 10.1007/s12035-011-8181-5. [Google Scholar] [CrossRef]
Siddiqui IJ, Pervaiz N, Abbasi AA (2016). The Parkinson disease gene SNCA: Evolutionary and structural insights with pathological implication. Scientific Reports 6: 24475. DOI 10.1038/srep24475. [Google Scholar] [CrossRef]
Spinelli KJ, Osterberg VR, Meshul CK, Soumyanath A, Unni VK (2015). Curcumin treatment improves motor behavior in α-synuclein transgenic mice. PLoS One 10: e0128510. DOI 10.1371/journal.pone.0128510. [Google Scholar] [CrossRef]
Van Der Spoel D, Lindahl E, Hess B, Groenhof G, Mark AE, Berendsen HJC (2005). GROMACS: Fast, flexible, and free. Journal of Computational Chemistry 26: 1701–1718. DOI 10.1002/jcc.20291. [Google Scholar] [CrossRef]
Srivastava V, Gupta SP, Siddiqi MI, Mishra BN (2010). Molecular docking studies on quinazoline antifolate derivatives as human thymidylate synthase inhibitors. Bioinformation 4: 357–365. DOI 10.6026/97320630004357. [Google Scholar] [CrossRef]
Stefanis L (2012). a-Synuclein in Parkinson’s disease. Cold Spring Harbor Perspectives in Medicine 2: a009399. DOI 10.1101/cshperspect.a009399 [Google Scholar]
Stierand K, Rarey M (2010). PoseView--molecular interaction patterns at a glance. Journal of Cheminformatics 2: P50. DOI 10.1186/1758-2946-2-S1-P50. [Google Scholar] [CrossRef]
Trott O, Olson AJ (2009). AutoDock Vina: Improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. Journal of Computational Chemistry 32: 455–461. DOI 10.1002/jcc.21334. [Google Scholar] [CrossRef]
Veber DF, Johnson SR, Cheng HY, Smith BR, Ward KW, Kopple KD (2002). Molecular properties that influence the oral bioavailability of drug candidates. Journal of Medicinal Chemistry 45: 2615–2623. DOI 10.1021/jm020017n. [Google Scholar] [CrossRef]
Wang C, Cai Z, Wang W, Wei M, Kou D, Li T, Yang Z, Guo H, Le W, Li S (2019). Piperine attenuates cognitive impairment in an experimental mouse model of sporadic Alzheimer’s disease. The Journal of Nutritional Biochemistry 70: 147–155. DOI 10.1016/j.jnutbio.2019.05.009. [Google Scholar] [CrossRef]
Wang WH, Shen CY, Chien YC, Chang WS, Tsai CW, Lin YH, Hwang JJ (2020). Validation of enhancing effects of curcumin on radiotherapy with F98/FGT glioblastoma-bearing rat model. International Journal of Molecular Sciences 21: 4385. DOI 10.3390/ijms21124385. [Google Scholar] [CrossRef]
Wei Z, Li X, Li X, Liu Q, Cheng Y (2018). Oxidative stress in Parkinson’s disease: A systematic review and meta-analysis. Frontiers in Molecular Neuroscience 11: 1–7. DOI 10.3389/fnmol.2018.00236. [Google Scholar] [CrossRef]
Wolohan PRN, Clark RD (2003). Predicting drug pharmacokinetic properties using molecular interaction fields and SIMCA. Journal of Computer-Aided Molecular Design 17: 65–76. DOI 10.1023/A:1024582008908. [Google Scholar] [CrossRef]
Xu Q, Langley M, Kanthasamy AG, Reddy MB (2017). Epigallocatechin gallate has a neurorescue effect in a mouse model of Parkinson disease. The Journal of Nutrition 147: 1926–1931. DOI 10.3945/jn.117.255034. [Google Scholar] [CrossRef]
Yu S, Yan H, Zhang L, Shan M, Chen P, Ding A, Li S (2017). A review on the phytochemistry, pharmacology, and pharmacokinetics of amentoflavone, a naturally-occurring biflavonoid. Molecules 22: 299. DOI 10.3390/molecules22020299. [Google Scholar] [CrossRef]
Zhang Z, Cao X, Xiong N, Wang H, Huang J, Sun S, Wang T (2010). Morin exerts neuroprotective actions in Parkinson disease models in vitro and in vivo. Acta Pharmacologica Sinica 31: 900–906. DOI 10.1038/aps.2010.77. [Google Scholar] [CrossRef]
Zhu M, Gong D (2020). A mouse model of 1-methyl-4-phenyl-1,2,3,6-tetrahydropyridine (MPTP)-induced Parkinson disease shows that 2-aminoquinoline targets JNK phosphorylation. Medical Science Monitor 26: 1–10. DOI 10.12659/MSM.920989. [Google Scholar] [CrossRef]
Zoete V, Cuendet MA, Grosdidier A, Michielin O (2011). SwissParam: A fast force field generation tool for small organic molecules. Journal of Computational Chemistry 32: 2359–2368. DOI 10.1002/jcc.21816. [Google Scholar] [CrossRef]
Supplementary Materials
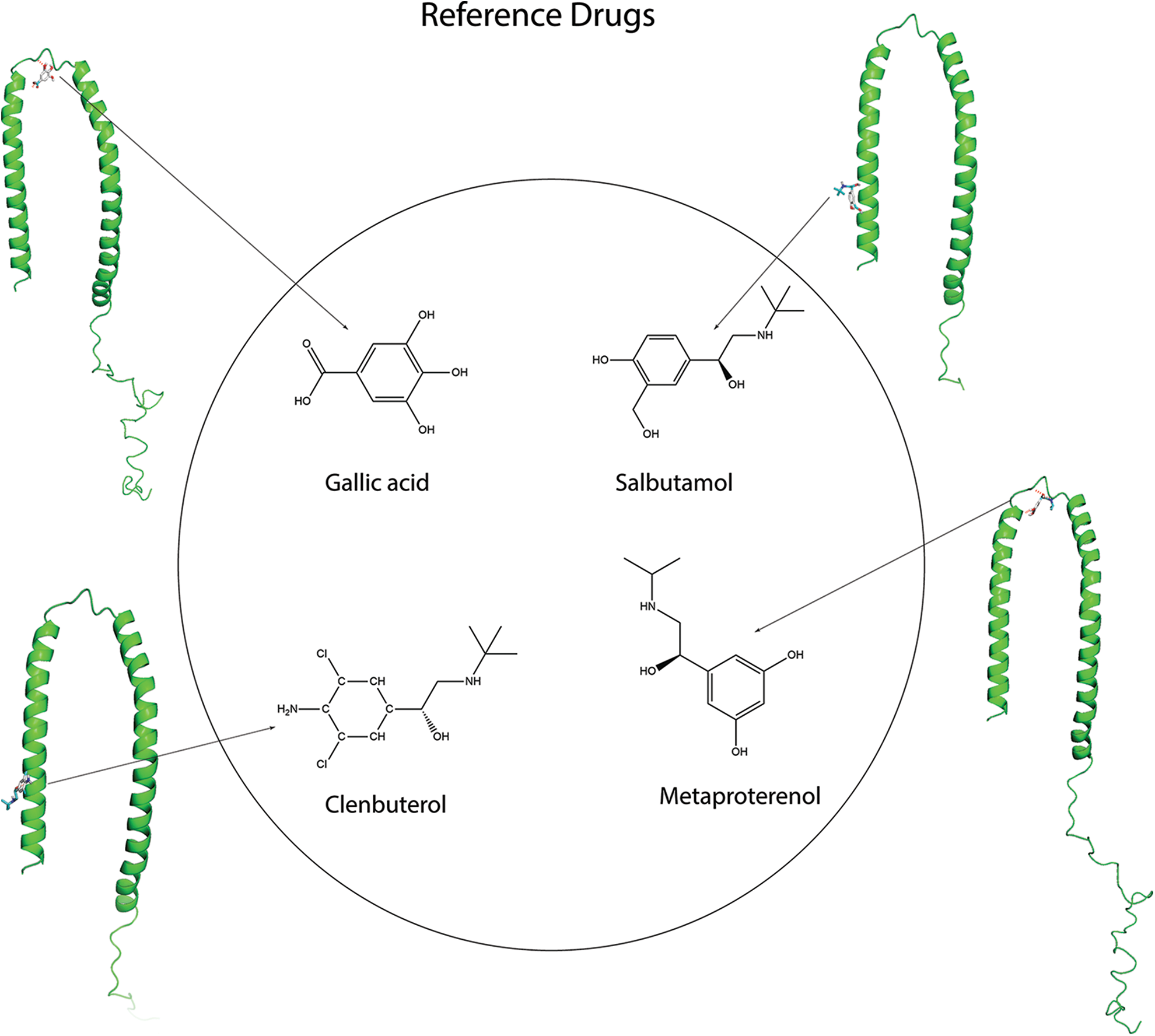
Figure S1: Four reference drugs (as control) used for their inhibition activity against the alpha-synuclein aggregation. So, these four compounds were docked individually to the alpha-synuclein and compared the binding affinity with our target compounds.
Table S1. The results of ADME of the 22 examined compounds
Cite This Article
 Copyright © 2022 The Author(s). Published by Tech Science Press.
Copyright © 2022 The Author(s). Published by Tech Science Press.This work is licensed under a Creative Commons Attribution 4.0 International License , which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.


 Submit a Paper
Submit a Paper Propose a Special lssue
Propose a Special lssue View Full Text
View Full Text Download PDF
Download PDF Downloads
Downloads
 Citation Tools
Citation Tools