 Open Access
Open Access
REVIEW
Research Advances and Therapeutic Potential of Gut Microbiota in Metabolic Diseases
1 Health Sciences Center, Yangtze University, 434023, China
2 Department of Psychiatry and Psychology, Wuhan Veterans’ and Preferential Treatment Hospital, Wuhan, 430024, China
* Corresponding Authors: Wangui Yu. Email: ; Xiaochun Peng. Email:
# These authors contributed equally to this work and should be considered co-first authors
(This article belongs to the Special Issue: Cellular and Molecular Mechanisms of Gut Microbiota, Oxidative Stress, and Inflammation in Health and Disease)
BIOCELL 2026, 50(4), 6 https://doi.org/10.32604/biocell.2026.075338
Received 29 October 2025; Accepted 24 December 2025; Issue published 21 April 2026
Abstract
The gut microbiota plays a pivotal role in maintaining host metabolic homeostasis. Accumulating evidence has demonstrated that dysbiosis of the gut microbiota is closely associated with metabolic disorders, including obesity, type 2 diabetes mellitus (T2DM), and non-alcoholic fatty liver disease (NAFLD). These alterations affect energy harvest, bile acid and short-chain fatty acid metabolism, intestinal barrier integrity, and low-grade inflammation, thereby contributing to insulin resistance and ectopic fat accumulation. In this narrative review, we summarize current knowledge on microbiome-host interactions in metabolic diseases, with a focus on energy metabolism, immune regulation, and inflammatory pathways. We further discuss the therapeutic potential and limitations of microbiota-targeted interventions, such as probiotics, prebiotics, dietary modifications, and fecal microbiota transplantation (FMT), highlighting the heterogeneity of clinical responses and safety considerations. Finally, we outline emerging precision strategies and future research directions aimed at integrating multi-omics profiling and patient stratification to inform personalized prevention and treatment of metabolic diseases.Graphic Abstract
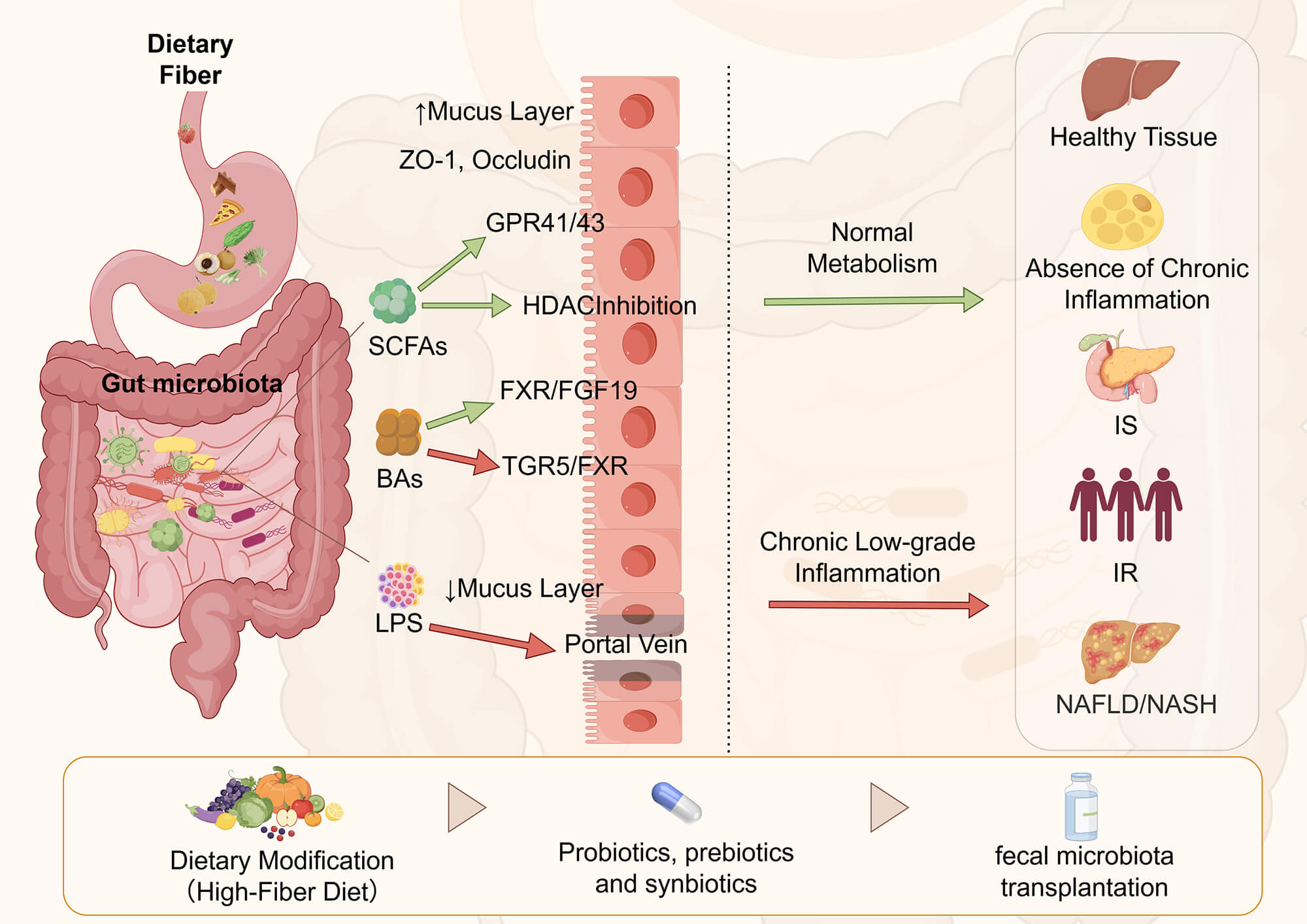
Keywords
Metabolic diseases have emerged as a critical global public health challenge. According to the latest data from the Global Burden of Disease Study 2019 [1], the prevalence of metabolic disorders—including obesity, T2DM, and NAFLD—has shown a persistent upward trajectory worldwide over the past two decades, with particularly pronounced growth in nations of higher socioeconomic development. This trend has imposed a substantial burden on healthcare systems, underscoring its urgency as a global priority. Epidemiological evidence links the burden of metabolic diseases to interconnected dysregulatory processes involving insulin resistance, glucose/lipid metabolism disturbances, and chronic low-grade inflammation [1].
In recent years, the gut microbiota has emerged as a focal point in understanding the pathogenesis of metabolic diseases, functioning as the human body’s “second genome”. Through various mechanisms, the gut microbiota regulates host metabolism, encompassing energy metabolism, short-chain fatty acid (SCFA) production, bile acid (BA) metabolism, and gut-brain axis signaling [2]. Although the precise molecular interactions between the gut microbiota and the host remain incompletely understood, existing evidence demonstrates that targeted interventions, such as probiotic/prebiotic supplementation, dietary modifications, and FMT, can yield clinically significant effects [3].
Most of the evidence discussed in this review comes from studies employing high-throughput sequencing and mass spectrometry methods [4]. 16S rRNA gene sequencing remains one of the most widely used techniques; it provides cost-effective taxonomic maps with resolution at the genus level, but offers limited insight into strain-level variation and functional capabilities [4,5]. Shotgun metagenomic sequencing overcomes some of these limitations by capturing total microbial DNA, enabling finer taxonomic resolution and functional profiling [5,6]. Metatranscriptomics studies active transcriptional components in the microbiome, revealing environment-dependent changes in microbial gene expression under dietary, pharmacological, or disease conditions [7]. Proteomics and metabolomics provide complementary information at the protein and metabolite levels; particularly, targeted and non-targeted metabolomics analysis of serum, urine, and fecal samples is critical for quantifying SCFAs, bile acids, and other microbial metabolites that mediate microbiota-host interactions [8]. Integrated analyses (e.g., the combination of metagenomics and metabolomics) provide a more comprehensive view of microbial functions than single modalities [9].
However, each method has inherent biases and limitations, including variations in DNA extraction, library preparation, sequencing platforms, bioinformatics workflows, and standardization strategies [4,5]. Batch effects, insufficient sample sizes, and inadequate dietary or medication controls may further confound the results, leading to inconsistent findings across studies [4,5]. Furthermore, most clinical microbiome studies remain cross-sectional, which limits the ability to establish causal relationships. These methodological considerations are critical for interpreting the strength of evidence linking gut microbiota changes to metabolic diseases, while also providing a foundation for designing robust studies that can distinguish between correlations and causation [9].
Here, we summarize and critically appraise recent advances on the role of the gut microbiota in obesity, T2DM, and NAFLD, with a particular focus on microbial metabolites (SCFAs, BA, and LPS) and microbiota-targeted therapies. Finally, we synthesize key knowledge gaps, outline future research directions, and discuss translational challenges that need to be addressed before microbiome-based interventions can be rationally integrated into the management of obesity, T2DM, and NAFLD.
2 Composition and Functions of the Gut Microbiota
The gut microbiota refers to the microbial community inhabiting the intestinal tract, while the ‘gut microbiome’ denotes its collective genome and functional potential [10]. We employ the concept of “dysbiosis” to describe deviations in composition and function compared to the healthy reference state. Early classical estimates suggested the human gut contained approximately 1014 bacterial cells, at least ten times more than human cells [11]. Recent systematic recalculations have revised this ratio to about 1:1, indicating roughly equal numbers of bacterial and human cells in the body [12].
Research indicates that the gut microbiota of healthy adults exhibits highly complex ecosystem characteristics, with its core composition dominated by Firmicutes (particularly butyrate-producing Clostridia strains) and Bacteroidetes (such as the metabolically regulatory B. thetaiotaomicron) [13]. Additionally, this ecosystem includes functionally important minor phyla such as Proteobacteria, Actinobacteria, Verrucomicrobia, and Fusobacteria [14]. Although these microbial groups are present in lower abundance, they play indispensable synergistic roles in maintaining gut microbial homeostasis and host health [12] (Fig. 1). The Firmicutes and Bacteroidetes phyla, as core gut microbiota, play indispensable regulatory roles in human health. Recent studies have revealed that gut Firmicutes encode numerous genes involved in dietary fiber fermentation and actively interact with the intestinal mucosa, playing a pivotal role in maintaining gut homeostasis [13]. The exceptional polysaccharide-degrading capacity of Bacteroides species plays a pivotal role in maintaining gut microbial ecology by generating essential metabolic substrates that support the growth of other commensal bacteria, thereby fostering a mutually beneficial symbiotic network within the intestinal microbiota [14].

Figure 1: In healthy individuals, the gut microbiota is predominantly composed of the phyla Firmicutes (e.g., Lactobacillus, Clostridium) and Bacteroidetes (e.g., Bacteroides, Prevotella). Actinobacteria (e.g., Bifidobacterium) and Verrucomicrobia (e.g., Akkermansia muciniphila) contribute to mucosal protection through metabolic regulation, while Proteobacteria typically exist at low abundance under homeostatic conditions. Aberrant expansion of Proteobacteria is strongly associated with microbial dysbiosis and pro-inflammatory states (Original illustration by Figdraw2.0, a scientific research platform for online drawing)
These SCFAs exhibit multiple physiological functions: butyrate serves as the primary energy source for colonocytes and maintains a hypoxic intestinal environment through PPARγ activation; acetate is metabolized in the liver via the portal vein and exerts anti-inflammatory effects through GPR43 receptor signaling; while propionate participates in hepatic gluconeogenesis. Their biosynthesis involves diverse metabolic pathways: acetate is produced through glycolysis or the Wood-Ljungdahl pathway; butyrate is derived from butyryl-CoA conversion, primarily by bacterial species such as Ruminococcaceae and Lachnospiraceae; and propionate can be synthesized via three distinct pathways—acrylate, succinate, and propanediol [15]. Notably, these SCFAs can interconvert (e.g., 24% of acetate is converted to butyrate) and collectively form an intricate metabolic network that plays pivotal roles in maintaining intestinal barrier function, regulating immune homeostasis, and modulating energy metabolism [15]. Bile acids play a central role in regulating energy metabolism, maintaining cholesterol homeostasis, and controlling the expression of bile acid transporters, making them a classic example of microbiota-host co-metabolites [16] Therefore, dysregulation of the gut microbiota leads to alterations in microbial-derived metabolites, which subsequently contribute to the development of various metabolic diseases [17]. While metabolic diseases are influenced by both genetic predisposition and environmental factors, their incidence among individuals sharing the same genetic background and energy intake exhibits a significant correlation with gut microbiota composition. It is now well-established through emerging research that dysregulation of the gut microbiota is substantially linked to a variety of metabolic disorders, among which obesity, T2DM, and NAFLD are particularly notable examples [2]. As summarized in Fig. 2, dysbiosis-driven alterations in SCFAs, bile acids, and LPS converge on barrier dysfunction and chronic low-grade inflammation, thereby promoting obesity, T2DM, and NAFLD/MASLD.

Figure 2: Dysbiosis alters the production of microbial metabolites such as SCFAs, bile acids, and LPS, thereby impairing gut barrier function, triggering low-grade inflammation, and contributing to obesity, T2DM, and NAFLD. Microbiota-targeted interventions—including dietary modulation, probiotics, prebiotics, and fecal microbiota transplantation (FMT)—aim to restore eubiosis and rebalance these metabolic and inflammatory pathways (Original illustration by Figdraw2.0, a scientific research platform for online drawing)
3 Gut Microbiota and Metabolic Diseases: Mechanistic Links
Intestinal microbiota dysbiosis is closely associated with the development of various metabolic disorders, including obesity, T2DM, and NAFLD, which share several underlying mechanisms. These mechanisms involve alterations in SCFAs production, disruption of the bile acid-FXR/TGR5-FGF15/19 axis, and increased intestinal permeability caused by low-grade inflammation driven by lipopolysaccharide (LPS) [18–20]. In this section, we first outline these common pathways and then focus on disease-specific microbial and metabolic features.
3.1 Shared Mechanisms across Obesity, T2DM, and NAFLD
3.1.1 SCFAs: Energy Metabolism, Barrier Function, and Inflammation
SCFAs are key metabolites produced by gut microbiota through the fermentation of dietary fiber, primarily consisting of acetate, propionate, and butyrate [19,21]. Butyric acid, as the main energy substrate of colonic cells, is closely related to the promotion of oxygen consumption and epithelial hypoxia, and the maintenance of an anaerobic environment, which is favorable for the growth of obligate anaerobic bacteria [22,23]. After entering the portal circulation, acetic acid and propionic acid participate in hepatic gluconeogenesis, lipogenesis, and cholesterol synthesis according to the specific environmental needs [18,22].
Beyond their role as substrates, SCFAs act as signaling molecules by activating GPCRs (GPR41/FFAR3, GPR43/FFAR2) and inhibiting histone deacetylases [24,25]. They stimulate the secretion of intestinal insulin-like growth factor (IGF) and satiety hormones (GLP-1, PYY), regulate sympathetic nerve output and energy expenditure, and promote the differentiation and function of regulatory T cells [24]. In addition, SCFAs, particularly butyrate, can, via receptor-mediated signaling and epigenetic regulation, be shown in experimental studies and in some clinical observations to be associated with the maintenance of tight-junction integrity, epithelial homeostasis, and anti-inflammatory responses, thereby contributing to improved intestinal barrier function [22,24,26]. As highlighted in the review by SIMão et al., SCFA-driven epigenetic and mitochondrial reprogramming plays a key role in metabolic disorders [27].
In obesity, T2DM, and NAFLD, microbiota dysbiosis is frequently associated with reduced abundance of key butyrate-producing bacteria (e.g., Fecalibacterium prausnitzii and Rohschia spp), as well as alterations in the SCFA profile [28–30]. However, human data remain heterogeneous, with SCFAs typically exhibiting dual effects of dose and environmental dependence—supporting metabolic health when produced in physiological amounts, but potentially increasing extra calories or causing ectopic lipid deposition when overproduced [21,31].
3.1.2 Bile Acids-FXR/TGR5-FGF15/19 Axis
Bile acids (BAs), a class of crucial signaling molecules, are synthesized by cholesterol in the liver and subsequently modified by gut microbiota [32]. Extensive research demonstrates that BAs function as signaling molecules by mediating the tight junctions between the liver and gut, thereby regulating glucose, lipid, and energy metabolism, as well as inflammatory responses [33–35]. BAs activate the nuclear receptor FXR and the membrane receptor TGR5, forming a complex signaling network that regulates glucose, lipid, and energy metabolism [36,37]. FXR in ileal cells induces fibroblast growth factor 15/19 (FGF15/19), which feeds back to the liver to inhibit bile acid synthesis and regulate gluconeogenesis and lipid metabolism [16,37]. TGR5 activation on intestinal endocrine cells and brown adipose tissue promotes GLP-1 secretion, thermogenesis, and energy expenditure [38,39].
Bacterial dysbiosis is frequently associated with the expansion of bile acid pools and increased hydrophobicity, along with alterations in the activation patterns of FXR/TGR5 [40]. Abnormal or excessive FXR signaling promotes hepatic adipogenesis and disrupts glucose homeostasis, whereas TGR5 signaling deficiency reduces GLP-1 secretion and thermogenesis [41,42]. These bile acid-mediated endocrine circuits are increasingly recognized as shared mediators of obesity, insulin resistance, and NAFLD, providing a mechanistic basis for the development of FXR/TGR5 modulators and bile acid isolators currently under investigation [40].
3.1.3 LPS and Metabolic Inflammation
LPS, a component of Gram-negative bacterial cell walls, exhibits pro-inflammatory properties. It has been identified as a key trigger for the onset and progression of mild systemic inflammation [43]. The disruption of SCFA production and BA signaling converges at the intestinal barrier, where reduced butyrate availability, mucin-degrading microbiota, and Western diets collectively compromise tight junctions and mucus layers, ultimately increasing paracellular permeability [20,44]. This enables LPS and other microbiota-associated molecular patterns (MAMPs) to migrate into the portal vein and systemic circulation, a phenomenon termed ‘metabolic endotoxemia’ [45,46]. One of the earliest reports on the association between intestinal barrier function, insulin resistance, and elevated circulating LPS levels originated from a study in obese mice [44]. Subsequent studies further confirmed that antibiotic therapy effectively reduces both intestinal and systemic LPS levels, accompanied by improved glucose tolerance, thereby establishing the gut microbiota as a key source of endotoxins [44]. Mechanistically, a high-fat diet (HFD) downregulates tight junction-associated gene expression, thereby increasing intestinal permeability [44,47]. As a protein complex between epithelial cells, tight junctions primarily regulate paracellular pathway permeability to prevent harmful substances from entering the body through this route [47]. Furthermore, studies have elucidated the molecular mechanism by which LPS induces increased tight junction permeability. This effect is demonstrated in both in vitro and in vivo experiments through enhanced expression and localization of Toll-like receptor 4 (TLR4) and CD14 on intestinal epithelial cell membranes [48].
LPS activates pattern recognition receptors (e.g., TLR4) on immune cells, endothelial cells, and parenchymal cells in adipose tissue, liver, and muscle, thereby activating the NF-κB and JNK signaling pathways and promoting the production of pro-inflammatory mediators such as TNF-α and IL-1β [49,50]. This chronic ‘superinflammation’ disrupts insulin receptor signaling, promotes lipid deposition, and accelerates hepatic fibrosis, thereby linking intestinal barrier dysfunction to the pathogenesis of obesity, T2DM, and NAFLD [51].
3.2 Disease-Specific Microbial and Metabolic Signatures
Despite these shared mechanisms, distinct metabolic disorders exhibit unique gut microbiota profiles and metabolite signatures. These microbial configurations and metabolic signatures modulate the relative dominance of SCFA, BA, and LPS-driven pathways.
3.2.1 Obesity-Specific Patterns
Obesity is a complex disease characterized by abnormal and excessive accumulation of body fat. The underlying cause of obesity is the long-term imbalance between energy intake and energy expenditure [52]. The disease is driven by both genetic and non-genetic (such as environmental) factors. According to the World Health Organization (WHO) criteria, obesity is defined as a body mass index (BMI) ≥30 kg/m2, but this threshold varies by region. For example, in China, obesity is defined as BMI ≥28 kg/m2. Globally, obesity has become a major public health challenge. A comprehensive survey shows that approximately 10% of the global population suffers from obesity, and nearly one-third of the population is overweight [53].
Mechanistically, obesity-associated microbiota can increase calories extracted from indigestible polysaccharides, alter the SCFA profile, thereby affecting appetite-regulating hormones (GLP-1, PYY, ghrelin) and adipose tissue thermogenesis, while remodeling the bile acid pool to promote FXR-driven lipid synthesis [21,31]. SCFAs are essential for maintaining energy balance. Acetate esters in particular promote lipid oxidation, reduce pro-inflammatory cytokine levels, and enhance lipolysis in vivo, thereby boosting energy expenditure [54]. Propionate promotes intestinal lipid solubility and energy balance in mice via the AMPK/LSD1 pathway [55]. The colon’s primary energy source is butyrate, which provides most of the energy required by intestinal epithelial cells through oxidation. By increasing butyrate production, butyrate-producing gut bacteria enhance lipid metabolism via the butyrate-SESN2/CRTC2 pathway. However, high butyrate concentrations may paradoxically impair this metabolic benefit and reduce the proportion of beneficial bacteria [56]. Rasmussen et al. [57] have shown that germ-free (GF) mice exhibit resistance to diet-induced obesity (DIO). Notably, FMT experiments revealed a donor-dependent effect: GF mice receiving microbiota from obese human donors gained significantly more body weight than those receiving microbiota from lean donors. Kulecka et al. [58] demonstrated that transferring gut microbiota from healthy mice to obese mice resulted in significant weight loss and markedly increased gut microbial diversity compared to baseline levels. This finding supports the causal role of the microbiota in obesity-related traits.
Large-scale metagenomic and 16S rRNA sequencing studies have consistently demonstrated that obese individuals exhibit reduced gut microbiota diversity, with enrichment of functional microbial groups that enhance energy extraction, bile salt hydrolase activity, and pro-inflammatory potential [18,31]. However, not all obese individuals exhibit the same microbiota dysbiosis, and microbial community profiles often overlap with those of metabolically ‘healthy’ obesity or metabolic syndrome [59,60]. The findings of different cohorts are inconsistent on the ratio of Firmicutes to Bacteroidetes (F/B ratio) [61–63]. Methodological heterogeneity (including sample processing, sequencing platforms, and bioinformatics workflows), along with strong confounding effects from dietary, pharmacological, and geographic factors, may explain the conflicting results. To ensure reproducibility, future studies should standardize experimental protocols and conduct large-scale prospective cohorts. Furthermore, microbial and metabolite profiles (rather than single microbial communities or F/B ratios) should be used to characterize obesity-associated microbiota dysbiosis [18,64].
T2DM is a metabolic disorder characterized by impaired pancreatic β-cell function and/or adipose tissue dysfunction, manifesting as insulin secretion defects and resistance, leading to chronic hyperglycemia [65]. Over the past three decades, the global prevalence of T2DM and prediabetes has risen significantly, with adult diabetes rates more than doubling since the 1980s, now posing a growing public health crisis [66,67]. From the perspective of pathogenesis, T2DM and obesity share common underlying pathological mechanisms, both involving metabolic dysregulation and alterations in gut microbiota [68,69]. Notably, fat accumulation exacerbates insulin resistance—a key driver of T2DM.
Comparative microbiome analysis revealed significant microbial dysbiosis between T2DM patients and healthy controls [70,71]. Compared with healthy controls, individuals with T2DM are frequently observed to exhibit a reduced abundance of butyrate-producing bacteria and a diminished functional potential for butyrate production; this alteration may attenuate the ability of butyrate to stimulate L-cell secretion of GLP-1 and PYY via G protein–coupled receptor (GPCR) pathways, thereby being unfavorable for the maintenance of glucose tolerance and insulin sensitivity [72,73]. In T2DM patients, the levels of SCFA-producing bacteria such as Myxobacterium mucosae and Enterococcus faecalis are reduced, while opportunistic or LPS-producing bacterial populations are elevated [49,74]. These changes correlate with alterations in the SCFA profile, metabolism of branched-chain amino acids (BCAA) and aromatic amino acids, as well as insulin resistance markers such as the insulin resistance index (HOMA-IR) [75,76]. SCFAs can activate FFAR2/GPR43 expressed in pancreatic islet cells (including β cells) and modulate glucose-stimulated insulin secretion through receptor-mediated downstream signaling. In addition, some experimental studies suggest that FFAR2-related pathways may participate in the regulation of β-cell mass and homeostasis; however, the available evidence is context dependent and remains somewhat controversial, and further population-based and mechanistic studies are required to determine whether FFAR2 activation confers long-term benefits for insulin sensitivity [77,78].
Multidisciplinary research has begun to identify specific microbial metabolites that link microbiota dysbiosis with insulin signaling dysfunction. For example, imidazolpropionate derived from histidine fermentation has been shown to antagonize insulin signaling in both mice and humans by activating stress kinases, with its circulating levels correlating with T2DM and metformin treatment response [79]. Microbial tryptophan metabolites (e.g., indole derivatives) can modulate mucosal inflammatory responses and epithelial barrier integrity and, via receptor-related pathways such as AHR and PXR signaling, influence islet/β-cell–associated phenotypes and insulin sensitivity [80]. Recent research titled “Microbiota alters mouse metabolome in age- and sex-dependent ways” further highlights that microbial effects on systemic metabolites, including branched-chain amino acids (BCAAs) and bile acid derivatives, vary significantly with host age and sex [81], adding another layer of complexity to the pathophysiology of T2DM.
In clinical practice, compared with obesity alone, patients with T2DM typically exhibit an impaired incretin effect and more pronounced hepatic insulin resistance (e.g., elevated HIRI), leading to an insufficient postprandial insulin response and dysregulated control of hepatic glucose output [82,83]. Microbial disturbances may either reduce GLP-1 secretion (e.g., through BA-FXR/TGR5 dysregulation) or enhance hepatic gluconeogenesis and low-grade inflammation, resulting in disproportionate effects [84,85]. This aligns with the earlier emphasis on the SCFA-FFAR2/3 and BA-FXR/TGR5 signaling pathways as key nodes connecting gut metabolism and glucose homeostasis [24,25,37].
NAFLD, also known as metabolic dysfunction-related fatty liver disease, is a chronic liver disorder characterized by excessive lipid accumulation in hepatocytes, diagnosed after ruling out heavy alcohol consumption and other identifiable causes of liver damage [86]. NAFLD is recognized as a new global health problem and a leading cause of chronic liver disease [87]. The prevalence in Western countries and Asia is 25% and 34%, respectively [88,89]. Chronic NAFLD can progress to non-alcoholic steatohepatitis (NASH) and other persistent liver conditions, eventually leading to cirrhosis and hepatocellular carcinoma (HCC) [90,91]. The “multiple hits” hypothesis suggests that in genetically susceptible individuals, a combination of factors—such as insulin resistance, dyslipidemia, nutritional imbalances, gut microbiota dysbiosis, and genetic or epigenetic alterations—collectively drive the development of NAFLD [86]. Dysbiosis increases the permeability of gut bacteria and increases exposure to harmful substances that exacerbate liver inflammation and fibrosis [92]. Insulin resistance and obesity are key pathophysiological factors in the development and progression of NAFLD, which is commonly regarded as the hepatic manifestation of metabolic syndrome [93].
Mechanistically, gut-derived bacteria and their metabolites can be transferred to the liver via the portal vein system, thereby modulating hepatic pathophysiological processes [94]. The Gut Microbiota and Their Metabolites Act as Molecular Messengers in the Gut-Hepatic Axis [95,96]. Multiple comparative studies have demonstrated that disease-specific alterations in gut microbiota communities and their metabolites—including SCFAs, BAs, LPS, choline, and trimethylamine (TMA)—are associated with disease severity and progression to liver fibrosis [97–99]. The reduction of gut microbial diversity, enrichment of ethanol-and LPS-producing bacteria, and depletion of choline-and SCFA-metabolizing bacteria are particularly significant in NAFLD, leading to barrier dysfunction and metabolic endotoxemia [19,100]. Increased intestinal permeability allows LPS and other pathogen-associated molecular patterns (PAMPs) to reach the liver via the portal vein. LPS triggers the TLR4/MyD88/NF-κB cascade, which promotes the release of pro-inflammatory factors (IL-6, TNF-α) and subsequently drives inflammation and hepatic fibrosis [101]. Meanwhile, the disruption of BA homeostasis and the FXR/TGR5-FGF15/19 signaling pathway alters hepatic lipid oxidation, very low-density lipoprotein (VLDL) secretion, and neofatogenesis, thereby creating a fatty degenerative environment [37,102].
Several gut-derived metabolites exhibit stage-specific variations in NAFLD. 5-Aminolevulinic acid trimethyl ester (TMAVA), produced by Enterococcus faecalis and Pseudomonas aeruginosa, elevates plasma levels in patients with hepatic steatosis and impairs carnitine-dependent fatty acid oxidation, thereby exacerbating hepatic steatosis in mice. Supplementation with carnitine or gene knockout of γ-butyryl betaine hydroxylase alleviates TMAVA-induced hepatic fat accumulation [103]. In a prospective clinical study, Meijnikman et al. [104] measured ethanol levels in 146 individuals. Using selective ADH inhibition, they demonstrated that an increase in portal vein ethanol concentration driven by gut microbiota (e.g., Lactobacillaceae) rises with NAFLD progression and plays a significant role in its pathogenesis. NAFLD can be induced in mice with Klebsiella pneumoniae infection through oral gastric pouch transplantation or FMT.
Other candidate substances include phenylacetic acid and specific sphingolipids, which have been associated with the severity and fibrosis of NAFLD in recent multi-omics studies [100,105]. Compared with obesity and T2DM, NAFLD is more closely associated with alterations in bile acid pools, the BA-FXR/FGF15 signaling pathway, and specific microbial metabolites (e.g., TMAVA), which directly regulate hepatic lipid processing and inflammation [18].
In conclusion, obesity, T2DM, and NAFLD share a core microbiota-driven mechanism involving SCFA production disruption, BA-FXR/TGR5-FGF15/19 signaling, and LPS-induced hyperinflammation. These mechanisms ultimately lead to compromised gut barrier integrity, chronic low-grade inflammation, and energy metabolism disorders [18]. However, the relative contribution and manifestation of these mechanisms vary from disease to disease.
3.3 Comparative Overview: Commonalities and Disease-Specific Differences
In obesity, changes in microbiota-associated energy acquisition, SCFA profiles, and appetite-regulating hormones are predominant, while BA remodeling and metabolic inflammation exacerbate adipose tissue expansion [19]. In T2DM, alterations in SCFA and BCA signaling interact with age-and sex-dependent metabolic changes in BCAA and aromatic amino acids, as well as β-cell dysfunction, ultimately leading to insulin resistance and impaired glucose regulation [19]. In NAFLD, these axes are mediated by the highly exposed gut-liver axis, where portal vein endotoxemia, BA imbalance, and specific microbial metabolites (e.g., TMAVA) disproportionately contribute to hepatic steatosis, inflammation, and fibrosis [38]. In conclusion, obesity, T2DM, and NAFLD share a core gut microbiota-host metabolic interaction axis centered on SCFAs, bile acids, and LPS [62]. However, each disease exhibits distinct microbial ecology and metabolic profiles (Table 1): obesity is characterized by dysregulation of energy acquisition and appetite hormone control; T2DM is strongly associated with insulin resistance and amino acid metabolic disorders; whereas NAFLD is predominantly marked by abnormal gut-liver axis communication and accumulation of specific hepatotoxic metabolites (e.g., TMAVA). Understanding these shared and unique features is essential for developing precision-targeted microbiome therapies.
4 Microbiota-Targeted Therapies in Obesity, T2DM, and NAFLD
Given the pivotal role of SCFAs, BAs, and endotoxins in linking gut microbiota dysbiosis to metabolic inflammation, various microbiota-targeting interventions have been explored, including dietary modifications, prebiotics, probiotics, synbiotics, FMT, and engineered live biotherapeutics. This section outlines the major therapeutic categories, with particular emphasis on (i) the magnitude and heterogeneity of clinical benefits, and (ii) safety considerations that are sometimes underestimated.
4.1 Lifestyle and Dietary Strategies: The Therapeutic Backbone
Lifestyle changes remain the cornerstone of treatment for obesity, T2DM, and NAFLD, with many of their metabolic benefits achieved through changes in the composition and function of the gut microbiota [111,112]. Recent evidence reveals that distinct dietary patterns exert varying effects on gut microbiota composition and SCFA production. For instance, animal-based diets have been shown to significantly reduce fecal concentrations of acetate and butyrate [68,113], whereas plant-based diets demonstrate greater efficacy in microbial modulation. Studies indicate that the Mediterranean diet (35% fat) and traditional low-fat diets (<30% fat) restore SCFA production capacity in metabolic syndrome patients by increasing beneficial bacterial populations such as Bacteroides, Prevotella, and Enterobacter cloacae, thereby improving gut microbiota health [114,115]. Conversely, there is a potential link between a high-fat diet and the development of endotoxemia and barrier dysfunction [116]. Food components interact closely with epithelial cells, serving as key triggers for intestinal barrier regulation. Moreover, a high-fat diet reduces beneficial bacteria and SCFAs, disrupts bile acid metabolism, induces inflammation, and accelerates the progression from NAFLD to NASH [117,118]. Dietary fiber plays a pivotal role in promoting the proliferation of SCFA-producing bacteria, enhancing microbial diversity and richness, and improving glycated hemoglobin levels in patients with T2DM [119]. Furthermore, it can reduce the production of harmful metabolites such as indole and hydrogen sulfide [120]. A meta-analysis showed that fiber intake led to increased abundance of Bifidobacterium and Lactobacillus genera, which may lead to increased butyrate concentrations in feces [121]. Conversely, a low-fiber diet not only reduces the production of SCFAs but also increases the formation of mucosal-damaging metabolites, potentially compromising the mucosal barrier function of the colon [122].
In the realm of dietary components, digested dietary proteins release amino acids that serve a dual purpose: they act as substrates for microbial protein synthesis and nitrogen cycling, while also participating in colonic fermentation to produce metabolites such as SCFAs, hydrogen sulfide, and ammonia. These metabolites influence microbial composition and function, thereby regulating intestinal barrier integrity and immune responses, ultimately affecting host health [123]. Extensive research consistently emphasizes the indispensable role of dietary fiber in blood glucose regulation [124,125]. Increasing intake of whole grains and nuts (foods rich in dietary fiber) promotes gut microbiota fermentation, which generates short-chain fatty acids such as butyrate and propionate. This process enhances glucose metabolism, improves insulin sensitivity, and reduces the risk of T2DM [126]. Specifically, adopting healthier dietary choices—including replacing saturated and trans fats with unsaturated fats, selecting low glycemic index (GI) carbohydrates, and consuming adequate high-quality protein—can effectively improve insulin sensitivity. Additionally, dietary supplementation with inulin has been shown to lower lipid levels (triglycerides and cholesterol), a mechanism related to SCFAs produced by gut microbiota fermentation. Specifically, propionate and butyrate improve glucose metabolism, while acetate reduces hepatic lipid accumulation by inhibiting SREBP-1 [127]. Randomized controlled trials demonstrate that daily intake of 220 g of yogurt is more effective than milk in improving insulin resistance and hepatic fat accumulation in obese women with NAFLD and metabolic syndrome (MetS). Potential mechanisms may involve regulation of gut microbiota, reduction of LPS levels, improvement of lipid metabolism, and alleviation of inflammation and oxidative stress [128].
In the field of dietary intervention strategies, emerging evidence highlights the profound impact of dietary modifications on metabolic regulation in T2DM. Studies demonstrate that calorie restriction (CR) yields significant results, with approximately 80% of obese T2DM patients achieving diabetes remission and an average weight loss of 15 kg [128,129]. The underlying mechanisms of CR involve multiple pathways: altering the gut microbiota composition, specifically increasing Bifidobacterium levels by 50%, Rosella bacteria levels by 30%, and elevating the proportion of non-12α-hydroxy bile acids by 60%–75%, thereby enhancing the production of SCFAs [129]. Similarly, the very low carbohydrate ketogenic diet (VLCK) improves the metabolic profile by elevating β-hydroxybutyrate (BHB) levels to 1.5–3.0 mM and reducing pro-inflammatory Th17 cells by 25%–40%, thereby exerting beneficial effects [130]. Conversely, HFDs induce adverse microbiota changes, with pathogenic bacteria like Escherichia coli increasing 2–3-fold while beneficial lactic acid bacteria decrease by 50%–60%. However, recent studies suggest that immune checkpoint modulation may confer immunomodulatory benefits, resulting in a 20%–25% increase in β-cell numbers [131,132]. Intermittent fasting (IF) and time-restricted feeding (TRF) have also shown potential, with studies demonstrating their ability to modulate the composition and function of gut microbiota, thereby leading to weight loss, fat reduction, and improved insulin sensitivity [132]. While these findings have been validated in animal models, further clinical studies are required to determine their applicability in humans. Compared with conventional calorie restriction, intermittent fasting (IF) demonstrates distinct advantages in blood glucose control, weight management, and reducing insulin resistance in T2DM patients [133,134].
A recent meta-analysis of nine studies [135] revealed the dual effects of VLCKD on gut microbiota. While VLCKD significantly reduced the α-diversity and abundance of beneficial bacteria like Bifidobacterium in the gut microbiota, it also promoted the growth of pro-inflammatory Enterobacteriaceae. Additionally, VLCKD was associated with elevated levels of potentially harmful metabolites, including branched-chain fatty acids and indole-3-acetic acid. These findings suggest that VLCKD may be linked to the risk of gut microbiota dysbiosis; however, existing evidence is limited by small sample sizes and the inclusion of both human and animal studies. Therefore, further large-scale clinical studies are needed to validate these observations and clarify the long-term effects of VLCKD in the management of T2DM.
In summary, these studies demonstrate that a balanced diet helps maintain the healthy structure and functional metabolites of gut microbiota, thereby effectively reducing the risks of obesity, NAFLD, and T2DM while optimizing glycemic control [136,137]. Furthermore, the research highlights that dietary modifications are mechanistically sound and clinically meaningful, though they require sustained behavioral changes and a baseline diet [117].
4.2 Probiotics, Prebiotics, and Synbiotics: Modest Effects, Marked Heterogeneity
Probiotics are non-pathogenic microorganisms with specific activities that have attracted much attention for their potential therapeutic value [138]. These microbes, including bacteria, fungi, and yeasts, when administered in sufficient doses (minimum 106 CFU/g), can actively regulate the host’s physical and chemical internal environment, interact with the host’s metabolism, and bring health benefits, sometimes as a complementary therapy to traditional treatments [139]. These health benefits include inhibiting pathogenic bacteria, regulating the immune system, reducing blood cholesterol, and enhancing the intestinal mucosal barrier function [93]. The most commonly used strains in practice are lactic acid bacteria and bifidobacteria, which are particularly advantageous because of their excellent adaptability in the human gut and the strong evidence supporting them [140].
Lactobacillus plantarum KC28 and Lactobacillus rhamnosus GG have been shown to prevent obesity in mice through their beneficial metabolic effects [141,142]. Recent studies have found that Bifidobacterium adolescentis FJSSZ23M10 [143] can also alleviate obesity through gut microbiota modulation and metabolic regulation. Numerous studies have highlighted the profound effects of probiotics on blood glucose regulation. For instance, animal research demonstrates that Lactobacillus bulgaricus strains G15 and Q14 enhance SCFA production by activating the GPR43 pathway, thereby improving intestinal barrier function and metabolic profiles [144]. Similarly, in diet-induced obesity models, combined intervention with Bifidobacterium animalis and Lactobacillus gannari with prebiotics improved metabolic and immunomodulatory benefits [145]. Clinical evidence corroborates these findings: Lactobacillus acidophilus strains LA-5 and BB-12 have been shown to improve glucose tolerance in T2DM patients [146], while Bifidobacterium lactis subsp lactis strain GCL2505 in animal studies reduced visceral fat and enhanced glucose tolerance [147]. The glucose regulation mechanisms mediated by probiotics involve multiple pathways. Studies have shown that probiotics can enhance GLP-1 secretion and upregulate the GPR41/43 signaling pathway, thereby improving insulin secretion and glucose metabolism [148]. Furthermore, probiotics may modulate immune responses in diabetic patients by regulating gut microbiota composition [149,150]. Their capacity to promote beneficial metabolites such as SCFAs further highlights their therapeutic potential. Clinical trials have demonstrated that probiotic interventions effectively reduce glycemic parameters—including blood glucose, HbA1c, and related symptoms—in both T2DM and gestational diabetes mellitus (GDM) patients [151,152]. A meta-analysis by Zhang et al. further validated these benefits, demonstrating significant improvements in fasting glucose, insulin levels, and HOMA-IR among T2DM patients after probiotic treatment [153].
Studies have also investigated the synergistic effects of combining probiotics with metformin in the management of T2DM. Mechanistically, metformin’s blood glucose regulation is partially mediated through modulation of gut microbiota composition [154,155]. Research indicates that targeted correction of gut microbiota dysbiosis can enhance the efficacy of lifestyle interventions and improve the therapeutic outcomes of metformin therapy [85,156]. Clinical evidence shows that this combination therapy has multiple advantages: it can not only improve glycemic control, insulin sensitivity, and lipid profile, but also reduce inflammatory responses and metabolic endotoxemia, while minimizing gastrointestinal adverse effects [157]. Other microbial strains, such as Enterococcus, Streptococcus, and Bacillus, also show therapeutic potential under specific conditions [138,158].
Probiotics are live bacteria that provide health benefits when consumed in sufficient amounts, while prebiotics are indigestible carbohydrates that support beneficial gut flora [159]. The use of prebiotics increases the number of beneficial microbes such as Acremonia, Fakalibacterium, Bifidobacterium, Lactobacillus, and Rosea, while reducing the number of pathogenic microbes [93,160]. Among prebiotics, inulin has garnered significant attention for its unique mechanisms, including promoting GLP-1 secretion to regulate blood glucose levels. Additionally, inulin exerts protective effects on T2DM through gut microbiota remodeling and anti-inflammatory actions [161,162]. Fructooligosaccharide (FOS) is known to improve liver pathology, inhibit adipocyte enlargement, and increase butyrate and acetate concentrations in feces as well as propionate levels in serum [163]. Studies indicate that dietary FOS can reduce steatohepatitis by restoring the intestinal epithelial barrier function and normalizing the gastrointestinal microbiota [164]. Administration of oligofructose in NAFLD patients has been shown to improve hepatic steatosis, increase Bifidobacterium abundance, and reduce Clostridium spp. cluster XI and I abundance [165]. Aoki et al. [166] found that inulin intake induces changes in the gut microbiota, including enrichment of Acidobacter and Blautia producta, which synergistically enhances acetyl ester production—a SCFA that protects NAFLD via the FFAR2 pathway. Hao et al. [167] reported that Crataegus pinnatifida polysaccharides effectively reduced hepatic steatosis by lowering serum triglycerides, total cholesterol, AST, ALT, and LDL-C levels. The intervention inhibited fatty acid synthesis gene expression, activated fatty acid oxidation gene expression, and increased total SCFA levels in NAFLD mice. Furthermore, the intervention restored the gut microbiota by reducing the ratio of Helicobacter pylori to Bacteroides and increasing the abundance of Akkermansia. Zuo et al. [168] discovered that epigallocatechin gallate in green tea inhibits intestinal barrier dysfunction and inflammation, improves the phenotype of non-alcoholic fatty liver disease, and alleviates metabolic disorders in high-fat diet-fed rats. Additionally, it promotes beneficial microorganisms, particularly SCFA-producing bacteria such as Lactobacillus, and inhibits suppressed Gram-negative bacteria like Desulfovibrio.
In addition, the synergistic effects of probiotics and prebiotics in promoting gut health have been well documented, and they can enhance the therapeutic effects of traditional medicines and inhibit pathogen colonization [169]. Compared with using either intervention alone, the combined use of prebiotics and probiotics demonstrated greater efficacy in reducing fasting blood glucose and HbA1c levels in patients with T2DM [126]. The synergistic effect of prebiotics and probiotics (known as synbiotics) has shown promising clinical applications in managing obesity and metabolic disorders. A randomized controlled trial demonstrated that a novel synbiotic formulation—combining inulin with Akkermansia mucilaginosa, Clostridium beilii, Clostridium butyricum, Bifidobacterium infantis, and Clostridium perfringens—significantly improves glycemic control in patients with T2DM [170]. A systematic review by Ferro et al. [171] demonstrated that adding specific prebiotics—particularly the galactooligosaccharide (GOS)/GOS-FOS combination—to infant formula promotes bifidobacterial colonization. The synbiotic intervention showed superior synergistic effects, with 40%–60% of infants exhibiting softer stool consistency.
Engineered probiotics (including Lactobacillus rhamnosus producing IL-22, Lactobacillus casei producing GLP-1, Escherichia coli 1917 producing NPEs, and Lactococcus lactis producing proinsulin/IL-10) and phage transplantation technology represent the fashionable engineering methods for precise regulation of microbial functions [172,173]. These strategies have demonstrated clinically significant efficacy in treating metabolic diseases (e.g., obesity, T2DM, and NAFLD) and intestinal inflammation [174]. Notably, probiotics and their derivatives exert therapeutic effects through multi-target mechanisms, showing great potential for the prevention and treatment of metabolic diseases. Moving forward, efforts must focus on developing personalized treatment strategies and accelerating their clinical translation to maximize their potential benefits.
RCTs on probiotics and prebiotics in obesity, T2DM, and metabolic syndrome show moderate and heterogeneous effects on body weight, insulin sensitivity, and liver histology, which are associated with significant differences in strains, doses, duration of treatment, and background therapy [175] (Table 2). These differences underscore the critical need for standardized formulations, strain-specific protocols, and patient stratification based on baseline microbiome profiles and metabolic signatures. Furthermore, deciphering the molecular mechanisms of host-microbe interactions will help identify key functional strains and their bioactive metabolites. Integrating multi-omics technologies with AI analytics can enhance predictions of individualized response variations. Future research should prioritize high-quality, large-scale, long-term clinical trials with standardized efficacy evaluation criteria. Concurrently, strengthening regulatory science frameworks to ensure verifiable safety and efficacy of probiotic products will accelerate their clinical translation in precision nutrition and chronic disease management.
4.3 FMT: Signal, Noise, and Safety
FMT is the most invasive method for microbiota modulation. Numerous studies have highlighted FMT’s therapeutic potential in metabolic diseases. For instance, preclinical models demonstrate that FMT treatment can lower blood glucose levels, enhance insulin sensitivity, and reduce pancreatic β-cell apoptosis in mice [195,196]. Mechanistically, transplantation of the normal human gut microbiota into diabetic mice improved glucose control [197–199] homeostasis by increasing SCFA production and stimulating the release of GLP-1 through the GPR43 receptor [148]. FMT from Healthy Donors Is Associated with Transcriptional Regulation of Genes Related to Liver Inflammation and Lipid Metabolism [200]. FMT restored microbial community balance, re-established bacterial colonization, and prevented excessive bacterial metabolites from migrating to the liver. By enhancing intestinal barrier integrity, FMT improved lipid metabolism, reduced insulin resistance, and suppressed inflammatory responses, thereby improving NAFLD [201].
Although it is now the standard of care for recurrent Clostridium difficile infection [202,203], its role in metabolic diseases is still experimental. A proof-of-concept randomized controlled trial in patients with metabolic syndrome [196] demonstrated that a single dose of FMT from a lean donor administered via nasoduodenal tube temporarily improved peripheral insulin sensitivity and increased microbial diversity in participants with low baseline microbial richness [204]. These early signals support the hypothesis that transferring an “insulin-sensitive” microbiome may improve host metabolism.
However, subsequent trial results were inconsistent [205]. A recent meta-analysis of randomized controlled trials (RCTs) on FMT for obesity [206], T2DM, and metabolic syndrome demonstrated short-term modest improvements in fasting blood glucose, HbA1c, and high-density lipoprotein cholesterol (HDL-cholesterol). However, no significant associations were observed with body weight, BMI, or HOMA-IR, and no evidence supported sustained benefits beyond 3–6 months. Several NAFLD/MAFLD studies [207] reported no improvement in insulin resistance or liver fat markers after allogeneic FMT compared to the autologous control group, with some even showing transient deterioration [208]. This highlights the potential for neutral or adverse metabolic trajectories [209].
The heterogeneity in FMT efficacy stems from multiple factors: donor selection (single versus mixed donors, lean versus metabolically healthy obese individuals), administration routes and frequency (upper gastrointestinal tract, colonoscopy, capsules), use of antibiotic pretreatment, disease stage, concomitant medications, and the critical baseline microbiome composition of the recipient [209]. Hierarchical analysis revealed that only a subset of recipients (‘responders’) achieved sustained colonization and metabolic benefits, typically those with pre-FMT low microbial diversity or specific composition defects [204]. Furthermore, the dynamic experimental results of gut bacteriophage communities in metabolic syndrome subjects after FMT [210] showed that although most individuals with high levels of these bacteriophages exhibited clinical improvement, their overall discriminative power was insufficient to serve as a reliable biomarker for predicting treatment outcomes.
Safety is a major concern. Since 2019, regulators have issued safety alerts on multiple occasions due to the spread of multidrug-resistant organisms (MDRO) via FMT, leading to severe infections and deaths in immunocompromised recipients [211]. These events have prompted strengthened donor screening requirements for multidrug-resistant organisms (MDROs), intestinal pathogens, and (during the COVID-19 pandemic) SARS-CoV-2, with most current guidelines recommending that non-Clostridium difficile FMT should only be performed in strictly monitored clinical trials and with detailed adverse event reporting [212,213].
Available evidence suggests that FMT can induce significant but unpredictable host metabolic changes in rigorously selected individuals, but entails considerable infection risks and complex implementation procedures [213,214]. For metabolic diseases, FMT should be considered a research tool rather than a routine therapy, with future research prioritizing: (i) rational donor-recipient matching based on microbiome/viroome characteristics; (ii) the use of clearly defined microbial consortia as safer, standardized alternatives; and (iii) assessing its durability and safety through long-term follow-up.
4.4 Emerging and Precision Microbiome-Based Strategies
To overcome the limitations of empirical probiotics and crude FMT, a variety of next-generation strategies are being developed to achieve more precise and safer microbiota modulation.
Defined microbial consortia and next-generation probiotics—such as Akkermansia muciniphila, Blautia wexlerae, and SCFA-producing Clostridia—are being developed as fixed-composition, GMP-manufactured live biotherapeutic products [215]. Earlier studies on obesity and NAFLD [216,217] demonstrated that these gut microbiota could improve insulin sensitivity, hepatic steatosis, and inflammatory markers, while avoiding the unpredictability of whole fecal microbiota transplantation, though the data remain limited and inconsistent.
Phage- and virome-based targeted therapies can selectively eliminate pathogenic or metabolically deleterious bacteria without widespread disruption. In a double-blind pilot trial [218], researchers safely transplanted a sterile fecal filtrate rich in donor bacteriophages into patients with metabolic syndrome. The therapy not only induced measurable changes in the bacteriophage-bacterial network but also showed no significant difference from placebo in the primary glucose tolerance metric. This highlights the therapy’s potential while indicating its current efficacy remains limited [219]. Phage mixtures targeting specific ethanol-producing or endotoxin-producing groups, associated with NAFLD and metabolic endotoxemia [220], are now in early clinical development.
Engineering Probiotics and GLP-1-Centric Design. Synthetic biology enables the development of engineered probiotics capable of secreting incretin hormones, antimicrobial peptides, or other therapeutic molecules in situ. Proof-of-concept studies demonstrate that genetically modified strains of Lactococcus lactis, Lactobacillus casei, or Bacillus subtilis can secrete GLP-1 or GLP-1 analogs, which effectively reduce blood glucose levels, improve dyslipidemia, and alleviate hepatic steatosis in rodent models of obesity and T2DM [221]. In murine colitis models, engineered Escherichia coli strains expressing GLP-1 signaling have demonstrated efficacy in restoring intestinal barrier integrity and metabolic parameters. These findings underscore the broader applicability of such microbial designs for addressing both barrier dysfunction and systemic metabolic disorders. While these approaches remain at a preliminary stage before clinical translation, they represent a paradigm shift from community-wide microbial regulation to pathway-specific, programmable microbiome therapies [222].
Towards precision microbiome intervention. Multi-omics and machine learning frameworks are increasingly being applied to identify microbial and metabolomic signatures that predict disease progression and treatment response in NAFLD/MASLD and T2DM [223–225]. In the future, microbiome-targeted therapies may be deployed as precision medicine: baseline microbiome and metabolomics analyses will be used to (i) stratify patients into subgroups (e.g., short-chain fatty acid deficiency, bile acid imbalance, barrier leakage), (ii) select customized combinations of diets, prebiotics, probiotics/prebiotics, specific microbiota, or phages/engineered bacterial strains, and (iii) monitor dynamic changes in intermediate endpoints such as short-chain fatty acids, bile acids, lipopolysaccharides, and host inflammatory and metabolic biomarkers [226].
In conclusion, current microbiota-targeted therapies for obesity, T2DM, and NAFLD demonstrate limited average efficacy with significant interindividual variations [227]. Probiotics and prebiotics are generally safe but have mixed effects; FMT can be very effective, but its effects are difficult to predict, and there are significant safety risks [227]. Emerging precision strategies offer hope for more targeted and sustained metabolic reprogramming, though most remain in early development. Future trials must incorporate mechanistic biomarkers, rigorous safety monitoring, and patient stratification to transition from empirical ‘one-size-fits-all’ interventions to microbiome-based, rational personalized therapies.
The dysbiosis of the gut microbiota has been identified as a significant driver of metabolic disorders such as obesity, T2DM, and NAFLD operating through mechanisms involving disrupted energy metabolism, compromised gut barrier integrity, and systemic inflammation. Various therapeutic strategies have demonstrated potential in modulating the gut microbiota, including probiotics, prebiotics, synbiotics, dietary interventions (e.g., Mediterranean diet), and FMT.
Notably, FMT has shown promise in improving insulin sensitivity by promoting colonization of beneficial bacteria such as Akkermansia muciniphila [228], while specific dietary patterns have been found to significantly reduce liver fat production. However, current research is hampered by several critical limitations.
Despite impressive progress, several controversies remain unresolved. First, the Firmicutes/Bacteroidetes ratio is still widely cited as a hallmark of obesity, yet meta-analyses fail to support its utility as a robust, disease-specific biomarker [60].
Second, clinical trials of probiotics, prebiotics, and synbiotics are heterogeneous in strains, endpoints, and study populations, making it difficult to derive evidence-based recommendations for specific products or indications.
Third, FMT and related microbiota-based therapies show variable efficacy and raise safety concerns, underscoring the need for well-characterized microbial consortia and rigorous long-term follow-up [229].
Finally, host genetics, diet, medication use, and circadian rhythms profoundly modulate microbiota-metabolism interactions, but are insufficiently controlled in most current studies.
Firstly, substantial differences exist between animal models and human microbiota, which complicate the translation of findings from preclinical studies to clinical applications. Additionally, there is a paucity of long-term safety and resistance data, which are essential for ensuring the efficacy and sustainability of microbiome-based therapies. Another major challenge lies in the limitations of detection technologies, such as 16S rRNA sequencing, which lack the resolution needed for precise functional characterization of microbial strains. Furthermore, the role of the virome and other microbial components remains understudied, and approximately 80% of the mechanisms underlying microbiome-host interactions remain to be elucidated [51,230]. These gaps underscore the need for further research to fully harness the therapeutic potential of the gut microbiota in addressing metabolic disorders.
6 Future Research Direction and Clinical Transformation Challenges
Future advancements in microbiome therapy will require a concerted, multidisciplinary effort to drive innovation and overcome current limitations. In the realm of basic research, there is a critical need to enhance our understanding of the causal mechanisms underlying specific bacterial and microbial metabolite functions. Functional assessments utilizing germ-free animal models coupled with organoid systems, particularly at the strain level, will be essential for elucidating mechanisms involving key metabolites such as short-chain fatty acids.
More specifically, three critical challenges must be addressed to facilitate successful clinical translation: First, moving from correlation to causation requires well-controlled longitudinal cohorts, gnotobiotic and humanized animal models, and interventional trials with defined microbial strains or metabolites to disentangle causal from passenger microbes. Second, integrative multi-omics—combining metagenomics, metatranscriptomics, metabolomics, and host genomics—will be essential to derive mechanistically grounded, metabolite-centered endotypes of metabolic disease [231]. Untargeted and targeted metabolomics can identify microbial metabolites, such as SCFAs, bile acid derivatives, and amino acid catabolites, that serve as proximal biomarkers and innovative therapeutic targets [81].
Third, emerging microbiota-targeted therapies, including bacteriophage cocktails against pathobionts, engineered probiotics that deliver GLP-1 or bile acid-modifying enzymes, and microbiome-informed precision nutrition, hold promise but are still in early development stages [232].
While existing interventions like bariatric surgery have demonstrated efficacy in reshaping the gut microbiota and improving metabolic profiles, their broad applicability remains constrained by their invasive nature. Consequently, the development of non-invasive targeted therapies, such as microbial metabolite encapsulation, and the identification of precision biomarkers will be key areas of focus.
Ultimately, the integration of metagenomic, metabolomic, and clinical trial data will be instrumental in bridging the gap between observational and interventional studies, enabling the critical transition from association to causation in microbiome-based therapies [51].
Acknowledgement: Not applicable.
Funding Statement: This study was supported by the “Medical and Health Talent Cultivation Program Project”, Department of Science and Technology Aid to Tibet, Yangtze University (2023YZ02), Project of Graduate Student Innovation Fund of Yangtze University (202406), and the Project of Innovation and Entrepreneurship Training Program for College Students of Yangtze University (Yz2023315, ydc202410).
Author Contributions: Research conception and design: Wangui Yu, Shuyu Yuan; Manuscript drafting: Shuyu Yuan, Guoxiao Han; Review and editing: Huimin Qiu, Xiaochun Peng, Henan Zheng; Visualization: Rongzhi Fang, Wangmiao Xie; Supervision: Xiaochun Peng, Wangui Yu. All authors reviewed and approved the final version of the manuscript.
Availability of Data and Materials: Not applicable.
Ethics Approval: Not applicable.
Conflicts of Interest: The authors declare no conflicts of interest to report regarding the present study.
Abbreviations
| Abbreviation | Full Term |
| ADH | Alcohol dehydrogenase |
| ALT | Alanine aminotransferase |
| AMPK | AMP-activated protein kinase |
| AST | Aspartate aminotransferase |
| BA(s) | Bile acid(s) |
| BCAA(s) | Branched-chain amino acid(s) |
| BHB | β-Hydroxybutyrate |
| BMI | Body mass index |
| CCK | Cholecystokinin |
| CD14 | Cluster of differentiation 14 |
| CFU | Colony-forming unit |
| CR | Calorie restriction |
| CRP | C-reactive protein |
| CRTC2 | CREB-regulated transcription coactivator 2 |
| DIO | Diet-induced obesity |
| F/B ratio | Firmicutes/Bacteroidetes ratio |
| FGF15/19 | Fibroblast growth factor 15/19 |
| FGF19 | Fibroblast growth factor 19 |
| FINS | Fasting insulin |
| FFAR2 | Free fatty acid receptor 2 (also known as GPR43) |
| FFAR3 | Free fatty acid receptor 3 (also known as GPR41) |
| FMT | Fecal microbiota transplantation |
| FOS | Fructooligosaccharides |
| FXR | Farnesoid X receptor |
| GDM | Gestational diabetes mellitus |
| GI | Glycemic index |
| GLP-1 | Glucagon-like peptide-1 |
| GGT | Gamma-glutamyl transferase |
| GMP | Good Manufacturing Practice |
| GPBAR1 | G protein-coupled bile acid receptor 1 (Takeda G-protein receptor 5) |
| GPCR | G protein-coupled receptor |
| GPR41 | G protein-coupled receptor 41 (FFAR3) |
| GPR43 | G protein-coupled receptor 43 (FFAR2) |
| HbA1c | Hemoglobin A1c (glycated hemoglobin) |
| HCC | Hepatocellular carcinoma |
| HDAC | Histone deacetylase |
| HDL-C | High-density lipoprotein cholesterol |
| HFD | High-fat diet |
| HOMA-IR | Homeostasis model assessment of insulin resistance |
| IF | Intermittent fasting |
| IGF | Insulin-like growth factor |
| IL-1β | Interleukin-1 beta |
| IL-6 | Interleukin-6 |
| IL-10 | Interleukin-10 |
| IL-22 | Interleukin-22 |
| ImP | Imidazole propionate |
| IR | Insulin resistance |
| IS | Insulin sensitivity |
| ITFs | Inulin-type fructans |
| iHMP | Integrative Human Microbiome Project |
| JNK | c-Jun N-terminal kinase |
| LDL-C | Low-density lipoprotein cholesterol |
| LPS | Lipopolysaccharide |
| MAFLD | Metabolic dysfunction-associated fatty liver disease |
| MASLD | Metabolic dysfunction-associated steatotic liver disease |
| MDRO(s) | Multidrug-resistant organism(s) |
| MetS | Metabolic syndrome |
| MRI-PDFF | Magnetic resonance imaging-proton density fat fraction |
| mTORC1 | Mechanistic target of rapamycin complex 1 |
| MyD88 | Myeloid differentiation primary response 88 |
| NAFLD | Non-alcoholic fatty liver disease |
| NASH | Non-alcoholic steatohepatitis |
| NLRP3 | NOD-, LRR- and pyrin domain-containing protein 3 (inflammasome) |
| PAMP(s) | Pathogen-associated molecular pattern(s) |
| PPARγ | Peroxisome proliferator-activated receptor gamma |
| PYY | Peptide YY |
| QUICKI | Quantitative insulin sensitivity check index |
| RCT(s) | Randomized controlled trial(s) |
| rRNA | Ribosomal RNA |
| SARS-CoV-2 | Severe acute respiratory syndrome coronavirus 2 |
| SCFA(s) | Short-chain fatty acid(s) |
| SESN2 | Sestrin 2 |
| SREBP-1 | Sterol regulatory element-binding protein 1 |
| T2DM | Type 2 diabetes mellitus |
| Th17 | T helper 17 (cell) |
| TGR5 | Takeda G protein-coupled receptor 5 (also known as GPBAR1) |
| TLR4 | Toll-like receptor 4 |
| TMAVA | N,N,N-Trimethyl-5-aminovaleric acid |
| TMA | Trimethylamine |
| TMAO | Trimethylamine N-oxide |
| TNF-α | Tumor necrosis factor alpha |
| Treg | Regulatory T cell |
| TRF | Time-restricted feeding |
| VLDL | Very low-density lipoprotein |
| VLCK | Very low-carbohydrate ketogenic diet |
| VLCKD | Very low-carbohydrate ketogenic diet (diet regimen) |
| WC | Waist circumference |
| WHO | World Health Organization |
| XOS | Xylo-oligosaccharides |
References
1. Chew NWS, Ng CH, Tan DJH, Kong G, Lin C, Chin YH, et al. The global burden of metabolic disease: data from 2000 to 2019. Cell Metab. 2023;35(3):414–28.e3. doi:10.1016/j.cmet.2023.02.003. [Google Scholar] [PubMed] [CrossRef]
2. Chen Y, Zhou J, Wang L. Role and mechanism of gut microbiota in human disease. Front Cell Infect Microbiol. 2021;11:625913. doi:10.3389/fcimb.2021.625913. [Google Scholar] [PubMed] [CrossRef]
3. Gomaa EZ. Human gut microbiota/microbiome in health and diseases: a review. Antonie Van Leeuwenhoek. 2020;113(12):2019–40. doi:10.1007/s10482-020-01474-7. [Google Scholar] [PubMed] [CrossRef]
4. Knight R, Vrbanac A, Taylor BC, Aksenov A, Callewaert C, Debelius J, et al. Best practices for analysing microbiomes. Nat Rev Microbiol. 2018;16(7):410–22. doi:10.1038/s41579-018-0029-9. [Google Scholar] [PubMed] [CrossRef]
5. Bharti R, Grimm DG. Current challenges and best-practice protocols for microbiome analysis. Brief Bioinform. 2021;22(1):178–93. doi:10.1093/bib/bbz155. [Google Scholar] [PubMed] [CrossRef]
6. Sharpton TJ. An introduction to the analysis of shotgun metagenomic data. Front Plant Sci. 2014;5:209. doi:10.3389/fpls.2014.00209. [Google Scholar] [PubMed] [CrossRef]
7. Shakya M, Lo CC, Chain PSG. Advances and challenges in metatranscriptomic analysis. Front Genet. 2019;10:904. doi:10.3389/fgene.2019.00904. [Google Scholar] [PubMed] [CrossRef]
8. Chen MX, Wang SY, Kuo CH, Tsai IL. Metabolome analysis for investigating host-gut microbiota interactions. J Formos Med Assoc. 2019;118(Suppl 1):S10–22. doi:10.1016/j.jfma.2018.09.007. [Google Scholar] [PubMed] [CrossRef]
9. Chetty A, Blekhman R. Multi-omic approaches for host-microbiome data integration. Gut Microbes. 2024;16(1):2297860. doi:10.1080/19490976.2023.2297860. [Google Scholar] [PubMed] [CrossRef]
10. Lee KA, Luong MK, Shaw H, Nathan P, Bataille V, Spector TD. The gut microbiome: what the oncologist ought to know. Br J Cancer. 2021;125(9):1197–209. doi:10.1038/s41416-021-01467-x. [Google Scholar] [PubMed] [CrossRef]
11. Luckey TD. Introduction to intestinal microecology. Am J Clin Nutr. 1972;25(12):1292–4. doi:10.1093/ajcn/25.12.1292. [Google Scholar] [PubMed] [CrossRef]
12. Sender R, Fuchs S, Milo R. Revised estimates for the number of human and bacteria cells in the body. PLoS Biol. 2016;14(8):e1002533. doi:10.1371/journal.pbio.1002533. [Google Scholar] [PubMed] [CrossRef]
13. The Human Microbiome Project Consortium. Structure, function and diversity of the healthy human microbiome. Nature. 2012;486(7402):207–14. doi:10.1038/nature11234. [Google Scholar] [PubMed] [CrossRef]
14. Lloyd-Price J, Abu-Ali G, Huttenhower C. The healthy human microbiome. Genome Med. 2016;8(1):51. doi:10.1186/s13073-016-0307-y. [Google Scholar] [PubMed] [CrossRef]
15. Fujisaka S, Watanabe Y, Tobe K. The gut microbiome: a core regulator of metabolism. J Endocrinol. 2023;256(3):e220111. doi:10.1530/JOE-22-0111. [Google Scholar] [PubMed] [CrossRef]
16. Li Y, Wang L, Yi Q, Luo L, Xiong Y. Regulation of bile acids and their receptor FXR in metabolic diseases. Front Nutr. 2024;11:1447878. doi:10.3389/fnut.2024.1447878. [Google Scholar] [PubMed] [CrossRef]
17. Lynch SV, Pedersen O. The human intestinal microbiome in health and disease. N Engl J Med. 2016;375(24):2369–79. doi:10.1056/NEJMra1600266. [Google Scholar] [PubMed] [CrossRef]
18. Fan Y, Pedersen O. Gut microbiota in human metabolic health and disease. Nat Rev Microbiol. 2021;19(1):55–71. doi:10.1038/s41579-020-0433-9. [Google Scholar] [PubMed] [CrossRef]
19. Wu J, Wang K, Wang X, Pang Y, Jiang C. The role of the gut microbiome and its metabolites in metabolic diseases. Protein Cell. 2021;12(5):360–73. doi:10.1007/s13238-020-00814-7. [Google Scholar] [PubMed] [CrossRef]
20. de Vos WM, Tilg H, Van Hul M, Cani PD. Gut microbiome and health: mechanistic insights. Gut. 2022;71(5):1020–32. doi:10.1136/gutjnl-2021-326789. [Google Scholar] [PubMed] [CrossRef]
21. Birkeland E, Gharagozlian S, Valeur J, Aas AM. Short-chain fatty acids as a link between diet and cardiometabolic risk: a narrative review. Lipids Health Dis. 2023;22(1):40. doi:10.1186/s12944-023-01803-5. [Google Scholar] [PubMed] [CrossRef]
22. Mukhopadhya I, Louis P. Gut microbiota-derived short-chain fatty acids and their role in human health and disease. Nat Rev Microbiol. 2025;23(10):635–51. doi:10.1038/s41579-025-01183-w. [Google Scholar] [PubMed] [CrossRef]
23. Vallianou N, Stratigou T, Christodoulatos GS, Dalamaga M. Understanding the role of the gut microbiome and microbial metabolites in obesity and obesity-associated metabolic disorders: current evidence and perspectives. Curr Obes Rep. 2019;8(3):317–32. doi:10.1007/s13679-019-00352-2. [Google Scholar] [PubMed] [CrossRef]
24. Mann ER, Lam YK, Uhlig HH. Short-chain fatty acids: linking diet, the microbiome and immunity. Nat Rev Immunol. 2024;24(8):577–95. doi:10.1038/s41577-024-01014-8. [Google Scholar] [PubMed] [CrossRef]
25. Kimura I, Ichimura A, Ohue-Kitano R, Igarashi M. Free fatty acid receptors in health and disease. Physiol Rev. 2020;100(1):171–210. doi:10.1152/physrev.00041.2018. [Google Scholar] [PubMed] [CrossRef]
26. Kalkan AE, BinMowyna MN, Raposo A, Ahmad MF, Ahmed F, Otayf AY, et al. Beyond the gut: unveiling butyrate’s global health impact through gut health and dysbiosis-related conditions: a narrative review. Nutrients. 2025;17(8):1305. doi:10.3390/nu17081305. [Google Scholar] [PubMed] [CrossRef]
27. SIMãO VA, De Almeida Chuffa LG, Ferder L, Inserra F, Manucha W. Mitochondrial-epigenetic crosstalk as an integrative standpoint into gut microbiome dysbiosis and related diseases. Biocell. 2024;48(10):1429–42. doi:10.32604/biocell.2024.053478. [Google Scholar] [CrossRef]
28. Münte E, Hartmann P. The role of short-chain fatty acids in metabolic dysfunction-associated steatotic liver disease and other metabolic diseases. Biomolecules. 2025;15(4):469. doi:10.3390/biom15040469. [Google Scholar] [PubMed] [CrossRef]
29. Tamanai-Shacoori Z, Smida I, Bousarghin L, Loreal O, Meuric V, Fong SB, et al. Roseburia spp.: a marker of health? Future Microbiol. 2017;12(2):157–70. doi:10.2217/fmb-2016-0130. [Google Scholar] [CrossRef]
30. Hu W, Gao W, Liu Z, Fang Z, Wang H, Zhao J, et al. Specific strains of Faecalibacterium prausnitzii ameliorate nonalcoholic fatty liver disease in mice in association with gut microbiota regulation. Nutrients. 2022;14(14):2945. doi:10.3390/nu14142945. [Google Scholar] [PubMed] [CrossRef]
31. Ilyés T, Silaghi CN, Crăciun AM. Diet-related changes of short-chain fatty acids in blood and feces in obesity and metabolic syndrome. Biology. 2022;11(11):1556. doi:10.3390/biology11111556. [Google Scholar] [PubMed] [CrossRef]
32. Simbrunner B, Trauner M, Reiberger T. Review article: therapeutic aspects of bile acid signalling in the gut-liver axis. Aliment Pharmacol Ther. 2021;54(10):1243–62. doi:10.1111/apt.16602. [Google Scholar] [PubMed] [CrossRef]
33. Li T, Chiang JYL. Bile acid signaling in metabolic and inflammatory diseases and drug development. Pharmacol Rev. 2024;76(6):1221–53. doi:10.1124/pharmrev.124.000978. [Google Scholar] [PubMed] [CrossRef]
34. Fleishman JS, Kumar S. Bile acid metabolism and signaling in health and disease: molecular mechanisms and therapeutic targets. Signal Transduct Target Ther. 2024;9(1):97. doi:10.1038/s41392-024-01811-6. [Google Scholar] [PubMed] [CrossRef]
35. Simbrunner B, Hofer BS, Schwabl P, Zinober K, Petrenko O, Fuchs C, et al. FXR-FGF19 signaling in the gut-liver axis is dysregulated in patients with cirrhosis and correlates with impaired intestinal defence. Hepatol Int. 2024;18(3):929–42. doi:10.1007/s12072-023-10636-4. [Google Scholar] [PubMed] [CrossRef]
36. Han B, Lv X, Liu G, Li S, Fan J, Chen L, et al. Gut microbiota-related bile acid metabolism-FXR/TGR5 axis impacts the response to anti-α4β7-integrin therapy in humanized mice with colitis. Gut Microbes. 2023;15(1):2232143. doi:10.1080/19490976.2023.2232143. [Google Scholar] [PubMed] [CrossRef]
37. Chiang JYL, Ferrell JM. Bile acid receptors FXR and TGR5 signaling in fatty liver diseases and therapy. Am J Physiol Gastrointest Liver Physiol. 2020;318(3):G554–73. doi:10.1152/ajpgi.00223.2019. [Google Scholar] [PubMed] [CrossRef]
38. Ni Y, Lu M, Xu Y, Wang Q, Gu X, Li Y, et al. The role of gut microbiota-bile acids axis in the progression of non-alcoholic fatty liver disease. Front Microbiol. 2022;13:908011. doi:10.3389/fmicb.2022.908011. [Google Scholar] [PubMed] [CrossRef]
39. Chen B, Bai Y, Tong F, Yan J, Zhang R, Zhong Y, et al. Glycoursodeoxycholic acid regulates bile acids level and alters gut microbiota and glycolipid metabolism to attenuate diabetes. Gut Microbes. 2023;15(1):2192155. doi:10.1080/19490976.2023.2192155. [Google Scholar] [PubMed] [CrossRef]
40. Zhao M, Zhao J, Yang H, Ouyang Z, Lv C, Geng Z, et al. The bile acid-gut microbiota axis: a central hub for physiological regulation and a novel therapeutic target for metabolic diseases. Biomed Pharmacother. 2025;188:118182. doi:10.1016/j.biopha.2025.118182. [Google Scholar] [PubMed] [CrossRef]
41. Molinaro A, Wahlström A, Marschall HU. Role of bile acids in metabolic control. Trends Endocrinol Metab. 2018;29(1):31–41. doi:10.1016/j.tem.2017.11.002. [Google Scholar] [PubMed] [CrossRef]
42. Parséus A, Sommer N, Sommer F, Caesar R, Molinaro A, Ståhlman M, et al. Microbiota-induced obesity requires farnesoid X receptor. Gut. 2017;66(3):429–37. doi:10.1136/gutjnl-2015-310283. [Google Scholar] [PubMed] [CrossRef]
43. Zanoni I, Ostuni R, Marek LR, Barresi S, Barbalat R, Barton GM, et al. CD14 controls the LPS-induced endocytosis of Toll-like receptor 4. Cell. 2011;147(4):868–80. doi:10.1016/j.cell.2011.09.051. [Google Scholar] [PubMed] [CrossRef]
44. Cani PD, Bibiloni R, Knauf C, Waget A, Neyrinck AM, Delzenne NM, et al. Changes in gut microbiota control metabolic endotoxemia-induced inflammation in high-fat diet-induced obesity and diabetes in mice. Diabetes. 2008;57(6):1470–81. doi:10.2337/db07-1403. [Google Scholar] [PubMed] [CrossRef]
45. Scheithauer TPM, Rampanelli E, Nieuwdorp M, Vallance BA, Verchere CB, van Raalte DH, et al. Gut microbiota as a trigger for metabolic inflammation in obesity and type 2 diabetes. Front Immunol. 2020;11:571731. doi:10.3389/fimmu.2020.571731. [Google Scholar] [PubMed] [CrossRef]
46. Mohammad S, Thiemermann C. Role of metabolic endotoxemia in systemic inflammation and potential interventions. Front Immunol. 2021;11:594150. doi:10.3389/fimmu.2020.594150. [Google Scholar] [PubMed] [CrossRef]
47. Rohr MW, Narasimhulu CA, Rudeski-Rohr TA, Parthasarathy S. Negative effects of a high-fat diet on intestinal permeability: a review. Adv Nutr. 2020;11(1):77–91. doi:10.1093/advances/nmz061. [Google Scholar] [PubMed] [CrossRef]
48. Guo S, Al-Sadi R, Said HM, Ma TY. Lipopolysaccharide causes an increase in intestinal tight junction permeability in vitro and in vivo by inducing enterocyte membrane expression and localization of TLR-4 and CD14. Am J Pathol. 2013;182(2):375–87. doi:10.1016/j.ajpath.2012.10.014. [Google Scholar] [PubMed] [CrossRef]
49. Gomes JMG, de Assis Costa J, de Cássia Gonçalves Alfenas R. Metabolic endotoxemia and diabetes mellitus: a systematic review. Metabolism. 2017;68:133–44. doi:10.1016/j.metabol.2016.12.009. [Google Scholar] [PubMed] [CrossRef]
50. Di Vincenzo F, Del Gaudio A, Petito V, Lopetuso LR, Scaldaferri F. Gut microbiota, intestinal permeability, and systemic inflammation: a narrative review. Intern Emerg Med. 2024;19(2):275–93. doi:10.1007/s11739-023-03374-w. [Google Scholar] [PubMed] [CrossRef]
51. Zhang L, Wang P, Huang J, Xing Y, Wong FS, Suo J, et al. Gut microbiota and therapy for obesity and type 2 diabetes. Front Endocrinol. 2024;15:1333778. doi:10.3389/fendo.2024.1333778. [Google Scholar] [PubMed] [CrossRef]
52. Lin X, Li H. Obesity: epidemiology, pathophysiology, and therapeutics. Front Endocrinol. 2021;12:706978. doi:10.3389/fendo.2021.706978. [Google Scholar] [PubMed] [CrossRef]
53. Afshin A, Forouzanfar MH, Reitsma MB, Sur P, Estep K, Lee A, et al. Health effects of overweight and obesity in 195 countries over 25 years. N Engl J Med. 2017;377(1):13–27. doi:10.1056/NEJMoa1614362. [Google Scholar] [PubMed] [CrossRef]
54. Hernández MAG, Canfora EE, Jocken JWE, Blaak EE. The short-chain fatty acid acetate in body weight control and insulin sensitivity. Nutrients. 2019;11(8):1943. doi:10.3390/nu11081943. [Google Scholar] [PubMed] [CrossRef]
55. Wang L, Yang H, Huang H, Zhang C, Zuo HX, Xu P, et al. Inulin-type fructans supplementation improves glycemic control for the prediabetes and type 2 diabetes populations: results from a GRADE-assessed systematic review and dose-response meta-analysis of 33 randomized controlled trials. J Transl Med. 2019;17(1):410. doi:10.1186/s12967-019-02159-0. [Google Scholar] [PubMed] [CrossRef]
56. Yu C, Liu S, Chen L, Shen J, Niu Y, Wang T, et al. Effect of exercise and butyrate supplementation on microbiota composition and lipid metabolism. J Endocrinol. 2019;243(2):125–35. doi:10.1530/JOE-19-0122. [Google Scholar] [PubMed] [CrossRef]
57. Rasmussen TS, Mentzel CMJ, Kot W, Castro-Mejía JL, Zuffa S, Swann JR, et al. Faecal virome transplantation decreases symptoms of type 2 diabetes and obesity in a murine model. Gut. 2020;69(12):2122–30. doi:10.1136/gutjnl-2019-320005. [Google Scholar] [PubMed] [CrossRef]
58. Kulecka M, Paziewska A, Zeber-Lubecka N, Ambrozkiewicz F, Kopczynski M, Kuklinska U, et al. Prolonged transfer of feces from the lean mice modulates gut microbiota in obese mice. Nutr Metab. 2016;13(1):57. doi:10.1186/s12986-016-0116-8. [Google Scholar] [PubMed] [CrossRef]
59. Sze MA, Schloss PD. Erratum for sze and schloss, “looking for a signal in the noise: revisiting obesity and the microbiome”. mBio. 2017;8(6):e01995–17. doi:10.1128/mBio.01995-17. [Google Scholar] [PubMed] [CrossRef]
60. Magne F, Gotteland M, Gauthier L, Zazueta A, Pesoa S, Navarrete P, et al. The firmicutes/bacteroidetes ratio: a relevant marker of gut dysbiosis in obese patients? Nutrients. 2020;12(5):1474. doi:10.3390/nu12051474. [Google Scholar] [PubMed] [CrossRef]
61. Wang PX, Deng XR, Zhang CH, Yuan HJ. Gut microbiota and metabolic syndrome. Chin Med J. 2020;133(7):808–16. doi:10.1097/CM9.0000000000000696. [Google Scholar] [PubMed] [CrossRef]
62. Ezenabor EH, Adeyemi AA, Adeyemi OS. Gut microbiota and metabolic syndrome: relationships and opportunities for new therapeutic strategies. Scientifica. 2024;2024(1):4222083. doi:10.1155/2024/4222083. [Google Scholar] [PubMed] [CrossRef]
63. Sasidharan Pillai S, Gagnon CA, Foster C, Ashraf AP. Exploring the gut microbiota: key insights into its role in obesity, metabolic syndrome, and type 2 diabetes. J Clin Endocrinol Metab. 2024;109(11):2709–19. doi:10.1210/clinem/dgae499. [Google Scholar] [PubMed] [CrossRef]
64. Castaner O, Goday A, Park YM, Lee SH, Magkos F, Shiow STE, et al. The gut microbiome profile in obesity: a systematic review. Int J Endocrinol. 2018;2018:4095789. doi:10.1155/2018/4095789. [Google Scholar] [PubMed] [CrossRef]
65. Donath MY, Dalmas É, Sauter NS, Böni-Schnetzler M. Inflammation in obesity and diabetes: islet dysfunction and therapeutic opportunity. Cell Metab. 2013;17(6):860–72. doi:10.1016/j.cmet.2013.05.001. [Google Scholar] [PubMed] [CrossRef]
66. Zhou B, Lu Y, Hajifathalian K, Bentham J, Di Cesare M, Danaei G, et al. Worldwide trends in diabetes since 1980: a pooled analysis of 751 population-based studies with 4·4 million participants. Lancet. 2016;387(10027):1513–30. doi:10.1016/S0140-6736(16)00618-8. [Google Scholar] [PubMed] [CrossRef]
67. Saeedi P, Petersohn I, Salpea P, Malanda B, Karuranga S, Unwin N, et al. Global and regional diabetes prevalence estimates for 2019 and projections for 2030 and 2045: results from the international diabetes federation diabetes atlas. Diabetes Res Clin Pract. 2019;157:107843. doi:10.1016/j.diabres.2019.107843. [Google Scholar] [PubMed] [CrossRef]
68. Qi C, Wang P, Fu T, Lu M, Cai Y, Chen X, et al. A comprehensive review for gut microbes: technologies, interventions, metabolites and diseases. Brief Funct Genomics. 2021;20(1):42–60. doi:10.1093/bfgp/elaa029. [Google Scholar] [PubMed] [CrossRef]
69. Massey W, Brown JM. The gut microbial endocrine organ in type 2 diabetes. Endocrinology. 2021;162(2):bqaa235. doi:10.1210/endocr/bqaa235. [Google Scholar] [PubMed] [CrossRef]
70. Wu X, Ma C, Han L, Nawaz M, Gao F, Zhang X, et al. Molecular characterisation of the faecal microbiota in patients with type II diabetes. Curr Microbiol. 2010;61(1):69–78. doi:10.1007/s00284-010-9582-9. [Google Scholar] [PubMed] [CrossRef]
71. Ahmad A, Yang W, Chen G, Shafiq M, Javed S, Ali Zaidi SS, et al. Analysis of gut microbiota of obese individuals with type 2 diabetes and healthy individuals. PLoS One. 2019;14(12):e0226372. doi:10.1371/journal.pone.0226372. [Google Scholar] [PubMed] [CrossRef]
72. Arora T, Tremaroli V. Therapeutic potential of butyrate for treatment of type 2 diabetes. Front Endocrinol. 2021;12:761834. doi:10.3389/fendo.2021.761834. [Google Scholar] [PubMed] [CrossRef]
73. Zeng Y, Wu Y, Zhang Q, Xiao X. Crosstalk between glucagon-like peptide 1 and gut microbiota in metabolic diseases. mBio. 2024;15(1):e02032–23. doi:10.1128/mbio.02032-23. [Google Scholar] [PubMed] [CrossRef]
74. Horvath A, Zukauskaite K, Hazia O, Balazs I, Stadlbauer V. Human gut microbiome: therapeutic opportunities for metabolic syndrome-Hype or hope? Endocrinol Diabetes Metab. 2024;7(1):e436. doi:10.1002/edm2.436. [Google Scholar] [PubMed] [CrossRef]
75. Newgard CB, An J, Bain JR, Muehlbauer MJ, Stevens RD, Lien LF, et al. A branched-chain amino acid-related metabolic signature that differentiates obese and lean humans and contributes to insulin resistance. Cell Metab. 2009;9(4):311–26. doi:10.1016/j.cmet.2009.02.002. [Google Scholar] [PubMed] [CrossRef]
76. Wu Z, Zhang B, Chen F, Xia R, Zhu D, Chen B, et al. Fecal microbiota transplantation reverses insulin resistance in type 2 diabetes: a randomized, controlled, prospective study. Front Cell Infect Microbiol. 2023;12:1089991. doi:10.3389/fcimb.2022.1089991. [Google Scholar] [PubMed] [CrossRef]
77. Teyani R, Moniri NH. Gut feelings in the islets: the role of the gut microbiome and the FFA2 and FFA3 receptors for short chain fatty acids on β-cell function and metabolic regulation. Br J Pharmacol. 2023;180(24):3113–29. doi:10.1111/bph.16225. [Google Scholar] [PubMed] [CrossRef]
78. Li Y, Liu Q, Pan CY, Lan XY. The free fatty acid receptor 2 (FFA2mechanisms of action, biased signaling, and clinical prospects. Pharmacol Ther. 2025;272(4):108878. doi:10.1016/j.pharmthera.2025.108878. [Google Scholar] [PubMed] [CrossRef]
79. Koh A, Molinaro A, Ståhlman M, Khan MT, Schmidt C, Mannerås-Holm L, et al. Microbially produced imidazole propionate impairs insulin signaling through mTORC1. Cell. 2018;175(4):947–61.e17. doi:10.1016/j.cell.2018.09.055. [Google Scholar] [PubMed] [CrossRef]
80. Wu Y, Li T, Chen B, Sun Y, Song L, Wang Y, et al. Tryptophan indole derivatives: key players in type 2 diabetes mellitus. Diabetes Metab Syndr Obes. 2025;18:1563–74. doi:10.2147/DMSO.S511068. [Google Scholar] [PubMed] [CrossRef]
81. Brown K, Thomson CA, Wacker S, Drikic M, Groves R, Fan V, et al. Microbiota alters the metabolome in an age- and sex-dependent manner in mice. Nat Commun. 2023;14(1):1348. doi:10.1038/s41467-023-37055-1. [Google Scholar] [PubMed] [CrossRef]
82. Nauck MA, Müller TD. Incretin hormones and type 2 diabetes. Diabetologia. 2023;66(10):1780–95. doi:10.1007/s00125-023-05956-x. [Google Scholar] [PubMed] [CrossRef]
83. Mittendorfer B, van Vliet S, Smith GI, Petersen MC, Patterson BW, Klein S. Impaired plasma glucose clearance is a key determinant of fasting hyperglycemia in people with obesity. Obesity. 2024;32(3):540–6. doi:10.1002/oby.23963. [Google Scholar] [PubMed] [CrossRef]
84. Holst JJ, Vilsbøll T, Deacon CF. The incretin system and its role in type 2 diabetes mellitus. Mol Cell Endocrinol. 2009;297(1–2):127–36. doi:10.1016/j.mce.2008.08.012. [Google Scholar] [PubMed] [CrossRef]
85. Sun L, Xie C, Wang G, Wu Y, Wu Q, Wang X, et al. Gut microbiota and intestinal FXR mediate the clinical benefits of metformin. Nat Med. 2018;24(12):1919–29. doi:10.1038/s41591-018-0222-4. [Google Scholar] [PubMed] [CrossRef]
86. Buzzetti E, Pinzani M, Tsochatzis EA. The multiple-hit pathogenesis of non-alcoholic fatty liver disease (NAFLD). Metabolism. 2016;65(8):1038–48. doi:10.1016/j.metabol.2015.12.012. [Google Scholar] [PubMed] [CrossRef]
87. Powell EE, Wong VW, Rinella M. Non-alcoholic fatty liver disease. Lancet. 2021;397(10290):2212–24. doi:10.1016/S0140-6736(20)32511-3. [Google Scholar] [PubMed] [CrossRef]
88. Kasper P, Martin A, Lang S, Kütting F, Goeser T, Demir M, et al. NAFLD and cardiovascular diseases: a clinical review. Clin Res Cardiol. 2021;110(7):921–37. doi:10.1007/s00392-020-01709-7. [Google Scholar] [PubMed] [CrossRef]
89. Yip TC, Lee HW, Chan WK, Wong GL, Wong VW. Asian perspective on NAFLD-associated HCC. J Hepatol. 2022;76(3):726–34. doi:10.1016/j.jhep.2021.09.024. [Google Scholar] [PubMed] [CrossRef]
90. Friedman SL, Neuschwander-Tetri BA, Rinella M, Sanyal AJ. Mechanisms of NAFLD development and therapeutic strategies. Nat Med. 2018;24(7):908–22. doi:10.1038/s41591-018-0104-9. [Google Scholar] [PubMed] [CrossRef]
91. Tacke F, Horn P, Wai-Sun Wong V, Ratziu V, Bugianesi E, Francque S, et al. EASL-EASD–EASO clinical practice guidelines on the management of metabolic dysfunction-associated steatotic liver disease (MASLD). J Hepatol. 2024;81(3):492–542. doi:10.1016/j.jhep.2024.04.031. [Google Scholar] [PubMed] [CrossRef]
92. Kolodziejczyk AA, Zheng D, Shibolet O, Elinav E. The role of the microbiome in NAFLD and NASH. EMBO Mol Med. 2019;11(2):e9302. doi:10.15252/emmm.201809302. [Google Scholar] [PubMed] [CrossRef]
93. Pandey H, Goel P, Srinivasan VM, Tang DWT, Wong SH, Lal D. Gut microbiota in non-alcoholic fatty liver disease: pathophysiology, diagnosis, and therapeutics. World J Hepatol. 2025;17(6):106849. doi:10.4254/wjh.v17.i6.106849. [Google Scholar] [PubMed] [CrossRef]
94. Han H, Jiang Y, Wang M, Melaku M, Liu L, Zhao Y, et al. Intestinal dysbiosis in nonalcoholic fatty liver disease (NAFLDfocusing on the gut-liver axis. Crit Rev Food Sci Nutr. 2023;63(12):1689–706. doi:10.1080/10408398.2021.1966738. [Google Scholar] [PubMed] [CrossRef]
95. Fang J, Yu CH, Li XJ, Yao JM, Fang ZY, Yoon SH, et al. Gut dysbiosis in nonalcoholic fatty liver disease: pathogenesis, diagnosis, and therapeutic implications. Front Cell Infect Microbiol. 2022;12:997018. doi:10.3389/fcimb.2022.997018. [Google Scholar] [PubMed] [CrossRef]
96. Nawrot M, Peschard S, Lestavel S, Staels B. Intestine-liver crosstalk in Type 2 Diabetes and non-alcoholic fatty liver disease. Metabolism. 2021;123(1):154844. doi:10.1016/j.metabol.2021.154844. [Google Scholar] [PubMed] [CrossRef]
97. Del Chierico F, Nobili V, Vernocchi P, Russo A, De Stefanis C, Gnani D, et al. Gut microbiota profiling of pediatric nonalcoholic fatty liver disease and obese patients unveiled by an integrated meta-omics-based approach. Hepatology. 2017;65(2):451–64. doi:10.1002/hep.28572. [Google Scholar] [PubMed] [CrossRef]
98. Loomba R, Seguritan V, Li W, Long T, Klitgord N, Bhatt A, et al. Gut microbiome-based metagenomic signature for non-invasive detection of advanced fibrosis in human nonalcoholic fatty liver disease. Cell Metab. 2019;30(3):607. doi:10.1016/j.cmet.2019.08.002. [Google Scholar] [PubMed] [CrossRef]
99. Lee G, You HJ, Bajaj JS, Joo SK, Yu J, Park S, et al. Distinct signatures of gut microbiome and metabolites associated with significant fibrosis in non-obese NAFLD. Nat Commun. 2020;11(1):4982. doi:10.1038/s41467-020-18754-5. [Google Scholar] [PubMed] [CrossRef]
100. Jeong MK, Min BH, Choi YR, Hyun JY, Park HJ, Eom JA, et al. Food and gut microbiota-derived metabolites in nonalcoholic fatty liver disease. Foods. 2022;11(17):2703. doi:10.3390/foods11172703. [Google Scholar] [PubMed] [CrossRef]
101. Arab JP, Martin-Mateos RM, Shah VH. Gut-liver axis, cirrhosis and portal hypertension: the chicken and the egg. Hepatol Int. 2018;12(Suppl 1):24–33. doi:10.1007/s12072-017-9798-x. [Google Scholar] [PubMed] [CrossRef]
102. Shao JW, Ge TT, Chen SZ, Wang G, Yang Q, Huang CH, et al. Role of bile acids in liver diseases mediated by the gut microbiome. World J Gastroenterol. 2021;27(22):3010–21. doi:10.3748/wjg.v27.i22.3010. [Google Scholar] [PubMed] [CrossRef]
103. Zhang X, Coker OO, Chu ES, Fu K, Lau HCH, Wang YX, et al. Dietary cholesterol drives fatty liver-associated liver cancer by modulating gut microbiota and metabolites. Gut. 2021;70(4):761–74. doi:10.1136/gutjnl-2019-319664. [Google Scholar] [PubMed] [CrossRef]
104. Meijnikman AS, Davids M, Herrema H, Aydin O, Tremaroli V, Rios-Morales M, et al. Microbiome-derived ethanol in nonalcoholic fatty liver disease. Nat Med. 2022;28(10):2100–6. doi:10.1038/s41591-022-02016-6. [Google Scholar] [PubMed] [CrossRef]
105. Ezzaidi N, Zhang X, Coker OO, Yu J. New insights and therapeutic implication of gut microbiota in non-alcoholic fatty liver disease and its associated liver cancer. Cancer Lett. 2019;459:186–91. doi:10.1016/j.canlet.2019.114425. [Google Scholar] [PubMed] [CrossRef]
106. Van Hul M, Cani PD. The gut microbiota in obesity and weight management: microbes as friends or foe? Nat Rev Endocrinol. 2023;19(5):258–71. doi:10.1038/s41574-022-00794-0. [Google Scholar] [PubMed] [CrossRef]
107. Qin J, Li Y, Cai Z, Li S, Zhu J, Zhang F, et al. A metagenome-wide association study of gut microbiota in type 2 diabetes. Nature. 2012;490(7418):55–60. doi:10.1038/nature11450. [Google Scholar] [PubMed] [CrossRef]
108. Gurung M, Li Z, You H, Rodrigues R, Jump DB, Morgun A, et al. Role of gut microbiota in type 2 diabetes pathophysiology. EBioMedicine. 2020;51(10):102590. doi:10.1016/j.ebiom.2019.11.051. [Google Scholar] [PubMed] [CrossRef]
109. Thomas C, Gioiello A, Noriega L, Strehle A, Oury J, Rizzo G, et al. TGR5-mediated bile acid sensing controls glucose homeostasis. Cell Metab. 2009;10(3):167–77. doi:10.1016/j.cmet.2009.08.001. [Google Scholar] [PubMed] [CrossRef]
110. Zhao M, Zhao L, Xiong X, He Y, Huang W, Liu Z, et al. TMAVA, a metabolite of intestinal microbes, is increased in plasma from patients with liver steatosis, inhibits γ-butyrobetaine hydroxylase, and exacerbates fatty liver in mice. Gastroenterology. 2020;158(8):2266–81.e27. doi:10.1053/j.gastro.2020.02.033. [Google Scholar] [PubMed] [CrossRef]
111. European Association for the Study of the Liver (EASLEuropean Association for the Study of Diabetes (EASDEuropean Association for the Study of Obesity (EASO). EASL–EASD–EASO clinical practice guidelines for the management of non-alcoholic fatty liver disease. J Hepatol. 2016;64(6):1388–402. doi:10.1016/j.jhep.2015.11.004. [Google Scholar] [PubMed] [CrossRef]
112. ElSayed NA, Aleppo G, Bannuru RR, Bruemmer D, Collins BS, Ekhlaspour L, et al. 2. diagnosis and classification of diabetes: standards of care in diabetes—2024. Diabetes Care. 2024;47(Suppl_1):S20–42. doi:10.2337/dc24-s002. [Google Scholar] [PubMed] [CrossRef]
113. Yuan L, Li Y, Chen M, Xue L, Wang J, Ding Y, et al. Therapeutic applications of gut microbes in cardiometabolic diseases: current state and perspectives. Appl Microbiol Biotechnol. 2024;108(1):156. doi:10.1007/s00253-024-13007-7. [Google Scholar] [PubMed] [CrossRef]
114. De Filippis F, Pellegrini N, Vannini L, Jeffery IB, La Storia A, Laghi L, et al. High-level adherence to a Mediterranean diet beneficially impacts the gut microbiota and associated metabolome. Gut. 2016;65(11):1812–21. doi:10.1136/gutjnl-2015-309957. [Google Scholar] [PubMed] [CrossRef]
115. Haro C, García-Carpintero S, Rangel-Zúñiga OA, Alcalá-Díaz JF, Landa BB, Clemente JC, et al. Consumption of two healthy dietary patterns restored microbiota dysbiosis in obese patients with metabolic dysfunction. Mol Nutr Food Res. 2017;61(12):1700300. doi:10.1002/mnfr.201700300. [Google Scholar] [PubMed] [CrossRef]
116. Barrea L, Annunziata G, Muscogiuri G, Di Somma C, Laudisio D, Maisto M, et al. Trimethylamine-N-oxide (TMAO) as novel potential biomarker of early predictors of metabolic syndrome. Nutrients. 2018;10(12):1971. doi:10.3390/nu10121971. [Google Scholar] [PubMed] [CrossRef]
117. Long C, Zhou X, Xia F, Zhou B. Intestinal barrier dysfunction and gut microbiota in non-alcoholic fatty liver disease: assessment, mechanisms, and therapeutic considerations. Biology. 2024;13(4):243. doi:10.3390/biology13040243. [Google Scholar] [PubMed] [CrossRef]
118. Garcia-Mantrana I, Selma-Royo M, Alcantara C, Collado MC. Shifts on gut microbiota associated to Mediterranean diet adherence and specific dietary intakes on general adult population. Front Microbiol. 2018;9:890. doi:10.3389/fmicb.2018.00890. [Google Scholar] [PubMed] [CrossRef]
119. Makki K, Deehan EC, Walter J, Bäckhed F. The impact of dietary fiber on gut microbiota in host health and disease. Cell Host Microbe. 2018;23(6):705–15. doi:10.1016/j.chom.2018.05.012. [Google Scholar] [PubMed] [CrossRef]
120. Zhao L, Zhang F, Ding X, Wu G, Lam YY, Wang X, et al. Gut bacteria selectively promoted by dietary fibers alleviate type 2 diabetes. Science. 2018;359(6380):1151–6. doi:10.1126/science.aao5774. [Google Scholar] [PubMed] [CrossRef]
121. So D, Whelan K, Rossi M, Morrison M, Holtmann G, Kelly JT, et al. Dietary fiber intervention on gut microbiota composition in healthy adults: a systematic review and meta-analysis. Am J Clin Nutr. 2018;107(6):965–83. doi:10.1093/ajcn/nqy041. [Google Scholar] [PubMed] [CrossRef]
122. Desai MS, Seekatz AM, Koropatkin NM, Kamada N, Hickey CA, Wolter M, et al. A dietary fiber-deprived gut microbiota degrades the colonic mucus barrier and enhances pathogen susceptibility. Cell. 2016;167(5):1339–53.e21. doi:10.1016/j.cell.2016.10.043. [Google Scholar] [PubMed] [CrossRef]
123. Blakeley-Ruiz JA, Bartlett A, McMillan AS, Awan A, Walsh MV, Meyerhoffer AK, et al. Dietary protein source alters gut microbiota composition and function. ISME J. 2025;19(1):wraf048. doi:10.1093/ismejo/wraf048. [Google Scholar] [PubMed] [CrossRef]
124. McRae MP. Dietary fiber intake and type 2 diabetes mellitus: an umbrella review of meta-analyses. J Chiropr Med. 2018;17(1):44–53. doi:10.1016/j.jcm.2017.11.002. [Google Scholar] [PubMed] [CrossRef]
125. Kyrø C, Tjønneland A, Overvad K, Olsen A, Landberg R. Higher whole-grain intake is associated with lower risk of type 2 diabetes among middle-aged men and women: the Danish diet, cancer, and health cohort. J Nutr. 2018;148(9):1434–44. doi:10.1093/jn/nxy112. [Google Scholar] [PubMed] [CrossRef]
126. Li SX, Guo Y. Gut microbiome: new perspectives for type 2 diabetes prevention and treatment. World J Clin Cases. 2023;11(31):7508–20. doi:10.12998/wjcc.v11.i31.7508. [Google Scholar] [PubMed] [CrossRef]
127. Bergentall M, Tremaroli V, Sun C, Henricsson M, Khan MT, Mannerås Holm L, et al. Gut microbiota mediates SREBP-1c-driven hepatic lipogenesis and steatosis in response to zero-fat high-sucrose diet. Mol Metab. 2025;97:102162. doi:10.1016/j.molmet.2025.102162. [Google Scholar] [PubMed] [CrossRef]
128. Chen Y, Feng R, Yang X, Dai J, Huang M, Ji X, et al. Yogurt improves insulin resistance and liver fat in obese women with nonalcoholic fatty liver disease and metabolic syndrome: a randomized controlled trial. Am J Clin Nutr. 2019;109(6):1611–9. doi:10.1093/ajcn/nqy358. [Google Scholar] [PubMed] [CrossRef]
129. Yao B, Fang H, Xu W, Yan Y, Xu H, Liu Y, et al. Dietary fiber intake and risk of type 2 diabetes: a dose-response analysis of prospective studies. Eur J Epidemiol. 2014;29(2):79–88. doi:10.1007/s10654-013-9876-x. [Google Scholar] [PubMed] [CrossRef]
130. Ang QY, Alexander M, Newman JC, Tian Y, Cai J, Upadhyay V, et al. Ketogenic diets alter the gut microbiome resulting in decreased intestinal Th17 cells. Cell. 2020;181(6):1263–75.e16. doi:10.1016/j.cell.2020.04.027. [Google Scholar] [PubMed] [CrossRef]
131. Clark AL, Yan Z, Chen SX, Shi V, Kulkarni DH, Diwan A, et al. High-fat diet prevents the development of autoimmune diabetes in NOD mice. Diabetes Obes Metab. 2021;23(11):2455–65. doi:10.1111/dom.14486. [Google Scholar] [PubMed] [CrossRef]
132. Ding JT, Yang KP, Lin KL, Cao YK, Zou F. Mechanisms and therapeutic strategies of immune checkpoint molecules and regulators in type 1 diabetes. Front Endocrinol. 2023;13:1090842. doi:10.3389/fendo.2022.1090842. [Google Scholar] [PubMed] [CrossRef]
133. Borgundvaag E, Mak J, Kramer CK. Metabolic impact of intermittent fasting in patients with type 2 diabetes mellitus: a systematic review and meta-analysis of interventional studies. J Clin Endocrinol Metab. 2021;106(3):902–11. doi:10.1210/clinem/dgaa926. [Google Scholar] [PubMed] [CrossRef]
134. Liu Z, Dai X, Zhang H, Shi R, Hui Y, Jin X, et al. Gut microbiota mediates intermittent-fasting alleviation of diabetes-induced cognitive impairment. Nat Commun. 2020;11(1):855. doi:10.1038/s41467-020-14676-4. [Google Scholar] [PubMed] [CrossRef]
135. Croci S, D’Apolito LI, Gasperi V, Catani MV, Savini I. Dietary strategies for management of metabolic syndrome: role of gut microbiota metabolites. Nutrients. 2021;13(5):1389. doi:10.3390/nu13051389. [Google Scholar] [PubMed] [CrossRef]
136. Magkos F, Hjorth MF, Astrup A. Diet and exercise in the prevention and treatment of type 2 diabetes mellitus. Nat Rev Endocrinol. 2020;16(10):545–55. doi:10.1038/s41574-020-0381-5. [Google Scholar] [PubMed] [CrossRef]
137. Glenn AJ, Li J, Lo K, Jenkins DJA, Boucher BA, Hanley AJ, et al. The portfolio diet and incident type 2 diabetes: findings from the women’s health initiative prospective cohort study. Diabetes Care. 2023;46(1):28–37. doi:10.2337/dc22-1029. [Google Scholar] [PubMed] [CrossRef]
138. Dai Y, Quan J, Xiong L, Luo Y, Yi B. Probiotics improve renal function, glucose, lipids, inflammation and oxidative stress in diabetic kidney disease: a systematic review and meta-analysis. Ren Fail. 2022;44(1):862–80. doi:10.1080/0886022X.2022.2079522. [Google Scholar] [PubMed] [CrossRef]
139. Peña-Durán E, García-Galindo JJ, López-Murillo LD, Huerta-Huerta A, Balleza-Alejandri LR, Beltrán-Ramírez A, et al. Microbiota and inflammatory markers: a review of their interplay, clinical implications, and metabolic disorders. Int J Mol Sci. 2025;26(4):1773. doi:10.3390/ijms26041773. [Google Scholar] [PubMed] [CrossRef]
140. Stavropoulou E, Bezirtzoglou E. Probiotics in medicine: a long debate. Front Immunol. 2020;11:2192. doi:10.3389/fimmu.2020.02192. [Google Scholar] [PubMed] [CrossRef]
141. Ji Y, Park S, Park H, Hwang E, Shin H, Pot B, et al. Modulation of active gut microbiota by Lactobacillus rhamnosus GG in a diet induced obesity murine model. Front Microbiol. 2018;9:710. doi:10.3389/fmicb.2018.00710. [Google Scholar] [PubMed] [CrossRef]
142. Huang E, Kim S, Park H, Park S, Ji Y, Todorov SD, et al. Modulation of the gut microbiome and obesity biomarkers by Lactobacillus Plantarum KC28 in a diet-induced obesity murine model. Probiotics Antimicrob Proteins. 2021;13(3):677–97. doi:10.1007/s12602-020-09720-0. [Google Scholar] [PubMed] [CrossRef]
143. Zhang B, Qiu J, Qu Z, Xiao R, Wang L, Tian P, et al. Bifidobacterium adolescentis FJSSZ23M10 modulates gut microbiota and metabolism to alleviate obesity through strain-specific genomic features. Food Funct. 2025;16(6):2415–31. doi:10.1039/d4fo06449f. [Google Scholar] [PubMed] [CrossRef]
144. Wu J, Yang K, Fan H, Wei M, Xiong Q. Targeting the gut microbiota and its metabolites for type 2 diabetes mellitus. Front Endocrinol. 2023;14:1114424. doi:10.3389/fendo.2023.1114424. [Google Scholar] [PubMed] [CrossRef]
145. Ke X, Walker A, Haange SB, Lagkouvardos I, Liu Y, Schmitt-Kopplin P, et al. Synbiotic-driven improvement of metabolic disturbances is associated with changes in the gut microbiome in diet-induced obese mice. Mol Metab. 2019;22:96–109. doi:10.1016/j.molmet.2019.01.012. [Google Scholar] [PubMed] [CrossRef]
146. Tonucci LB, Olbrich Dos Santos KM, Licursi de Oliveira L, Rocha Ribeiro SM, Duarte Martino HS. Clinical application of probiotics in type 2 diabetes mellitus: a randomized, double-blind, placebo-controlled study. Clin Nutr. 2017;36(1):85–92. doi:10.1016/j.clnu.2015.11.011. [Google Scholar] [PubMed] [CrossRef]
147. Aoki R, Kamikado K, Suda W, Takii H, Mikami Y, Suganuma N, et al. A proliferative probiotic Bifidobacterium strain in the gut ameliorates progression of metabolic disorders via microbiota modulation and acetate elevation. Sci Rep. 2017;7(1):43522. doi:10.1038/srep43522. [Google Scholar] [PubMed] [CrossRef]
148. Zhang PP, Li LL, Han X, Li QW, Zhang XH, Liu JJ, et al. Fecal microbiota transplantation improves metabolism and gut microbiome composition in db/db mice. Acta Pharmacol Sin. 2020;41(5):678–85. doi:10.1038/s41401-019-0330-9. [Google Scholar] [PubMed] [CrossRef]
149. Kerry RG, Patra JK, Gouda S, Park Y, Shin HS, Das G. Benefaction of probiotics for human health: a review. J Food Drug Anal. 2018;26(3):927–39. doi:10.1016/j.jfda.2018.01.002. [Google Scholar] [PubMed] [CrossRef]
150. Wang X, Zhang P, Zhang X. Probiotics regulate gut microbiota: an effective method to improve immunity. Molecules. 2021;26(19):6076. doi:10.3390/molecules26196076. [Google Scholar] [PubMed] [CrossRef]
151. Hsieh MC, Tsai WH, Jheng YP, Su SL, Wang SY, Lin CC, et al. The beneficial effects of Lactobacillus reuteri ADR-1 or ADR-3 consumption on type 2 diabetes mellitus: a randomized, double-blinded, placebo-controlled trial. Sci Rep. 2018;8(1):16791. doi:10.1038/s41598-018-35014-1. [Google Scholar] [PubMed] [CrossRef]
152. Hasain Z, Raja Ali RA, Ahmad HF, Abdul Rauf UF, Oon SF, Mokhtar NM. The roles of probiotics in the gut microbiota composition and metabolic outcomes in asymptomatic post-gestational diabetes women: a randomized controlled trial. Nutrients. 2022;14(18):3878. doi:10.3390/nu14183878. [Google Scholar] [PubMed] [CrossRef]
153. Zhang Q, Wu Y, Fei X. Effect of probiotics on glucose metabolism in patients with type 2 diabetes mellitus: a meta-analysis of randomized controlled trials. Medicina. 2016;52(1):28–34. doi:10.1016/j.medici.2015.11.008. [Google Scholar] [PubMed] [CrossRef]
154. Buse JB, DeFronzo RA, Rosenstock J, Kim T, Burns C, Skare S, et al. The primary glucose-lowering effect of metformin resides in the gut, not the circulation: results from short-term pharmacokinetic and 12-week dose-ranging studies. Diabetes Care. 2016;39(2):198–205. doi:10.2337/dc15-0488. [Google Scholar] [PubMed] [CrossRef]
155. He D, Han H, Fu X, Liu A, Zhan Y, Qiu H, et al. Metformin reduces blood glucose in treatment-naive type 2 diabetes by altering the gut microbiome. Can J Diabetes. 2022;46(2):150–6. doi:10.1016/j.jcjd.2021.08.001. [Google Scholar] [PubMed] [CrossRef]
156. Bin Lee C, Chae SU, Jo SJ, Jerng UM, Bae SK. The relationship between the gut microbiome and metformin as a key for treating type 2 diabetes mellitus. Int J Mol Sci. 2021;22(7):3566. doi:10.3390/ijms22073566. [Google Scholar] [PubMed] [CrossRef]
157. Memon H, Abdulla F, Reljic T, Alnuaimi S, Serdarevic F, Asimi ZV, et al. Effects of combined treatment of probiotics and metformin in management of type 2 diabetes: a systematic review and meta-analysis. Diabetes Res Clin Pract. 2023;202(3):110806. doi:10.1016/j.diabres.2023.110806. [Google Scholar] [PubMed] [CrossRef]
158. Wang Q, Wang Z, Pang B, Zheng H, Cao Z, Feng C, et al. Probiotics for the improvement of metabolic profiles in patients with metabolic-associated fatty liver disease: a systematic review and meta-analysis of randomized controlled trials. Front Endocrinol. 2022;13:1014670. doi:10.3389/fendo.2022.1014670. [Google Scholar] [PubMed] [CrossRef]
159. Al-Habsi N, Al-Khalili M, Haque SA, Elias M, Al Olqi N, Al Uraimi T. Health benefits of prebiotics, probiotics, synbiotics, and postbiotics. Nutrients. 2024;16(22):3955. doi:10.3390/nu16223955. [Google Scholar] [PubMed] [CrossRef]
160. Eslick S, Thompson C, Berthon B, Wood L. Short-chain fatty acids as anti-inflammatory agents in overweight and obesity: a systematic review and meta-analysis. Nutr Rev. 2022;80(4):838–56. doi:10.1093/nutrit/nuab059. [Google Scholar] [PubMed] [CrossRef]
161. Li K, Zhang L, Xue J, Yang X, Dong X, Sha L, et al. Dietary inulin alleviates diverse stages of type 2 diabetes mellitus via anti-inflammation and modulating gut microbiota in db/db mice. Food Funct. 2019;10(4):1915–27. doi:10.1039/c8fo02265h. [Google Scholar] [PubMed] [CrossRef]
162. Morshedi M, Saghafi-Asl M, Hosseinifard ES. The potential therapeutic effects of the gut microbiome manipulation by synbiotic containing-Lactobacillus plantarum on neuropsychological performance of diabetic rats. J Transl Med. 2020;18(1):18. doi:10.1186/s12967-019-02169-y. [Google Scholar] [PubMed] [CrossRef]
163. Takai A, Kikuchi K, Ichimura M, Tsuneyama K, Moritoki Y, Matsumoto K, et al. Fructo-oligosaccharides ameliorate steatohepatitis, visceral adiposity, and associated chronic inflammation via increased production of short-chain fatty acids in a mouse model of non-alcoholic steatohepatitis. BMC Gastroenterol. 2020;20(1):46. doi:10.1186/s12876-020-01194-2. [Google Scholar] [PubMed] [CrossRef]
164. Huang X, Chen Q, Fan Y, Yang R, Gong G, Yan C, et al. Fructooligosaccharides attenuate non-alcoholic fatty liver disease by remodeling gut microbiota and association with lipid metabolism. Biomed Pharmacother. 2023;159(9):114300. doi:10.1016/j.biopha.2023.114300. [Google Scholar] [PubMed] [CrossRef]
165. Bomhof MR, Parnell JA, Ramay HR, Crotty P, Rioux KP, Probert CS, et al. Histological improvement of non-alcoholic steatohepatitis with a prebiotic: a pilot clinical trial. Eur J Nutr. 2019;58(4):1735–45. doi:10.1007/s00394-018-1721-2. [Google Scholar] [PubMed] [CrossRef]
166. Aoki R, Onuki M, Hattori K, Ito M, Yamada T, Kamikado K, et al. Commensal microbe-derived acetate suppresses NAFLD/NASH development via hepatic FFAR2 signalling in mice. Microbiome. 2021;9(1):188. doi:10.1186/s40168-021-01125-7. [Google Scholar] [PubMed] [CrossRef]
167. Hao P, Yang X, Yin W, Wang X, Ling Y, Zhu M, et al. A study on the treatment effects of Crataegus pinnatifida polysaccharide on non-alcoholic fatty liver in mice by modulating gut microbiota. Front Vet Sci. 2024;11:1383801. doi:10.3389/fvets.2024.1383801. [Google Scholar] [PubMed] [CrossRef]
168. Zuo G, Chen M, Zuo Y, Liu F, Yang Y, Li J, et al. Tea polyphenol epigallocatechin gallate protects against nonalcoholic fatty liver disease and associated endotoxemia in rats via modulating gut microbiota dysbiosis and alleviating intestinal barrier dysfunction and related inflammation. J Agric Food Chem. 2024;acs.jafc.3c04832. doi:10.1021/acs.jafc.3c04832. [Google Scholar] [PubMed] [CrossRef]
169. Rahman MN, Barua N, Tin MCF, Dharmaratne P, Wong SH, Ip M. The use of probiotics and prebiotics in decolonizing pathogenic bacteria from the gut; a systematic review and meta-analysis of clinical outcomes. Gut Microbes. 2024;16(1):2356279. doi:10.1080/19490976.2024.2356279. [Google Scholar] [PubMed] [CrossRef]
170. Perraudeau F, McMurdie P, Bullard J, Cheng A, Cutcliffe C, Deo A, et al. Improvements to postprandial glucose control in subjects with type 2 diabetes: a multicenter, double blind, randomized placebo-controlled trial of a novel probiotic formulation. BMJ Open Diabetes Res Care. 2020;8(1):e001319. doi:10.1136/bmjdrc-2020-001319. [Google Scholar] [PubMed] [CrossRef]
171. Ferro LE, Crowley LN, Bittinger K, Friedman ES, Decker JE, Russel K, et al. Effects of prebiotics, probiotics, and synbiotics on the infant gut microbiota and other health outcomes: a systematic review. Crit Rev Food Sci Nutr. 2023;63(22):5620–42. doi:10.1080/10408398.2021.2022595. [Google Scholar] [PubMed] [CrossRef]
172. Ma J, Lyu Y, Liu X, Jia X, Cui F, Wu X, et al. Engineered probiotics. Microb Cell Fact. 2022;21(1):72. doi:10.1186/s12934-022-01799-0. [Google Scholar] [PubMed] [CrossRef]
173. Murali SK, Mansell TJ. Next generation probiotics: engineering live biotherapeutics. Biotechnol Adv. 2024;72:108336. doi:10.1016/j.biotechadv.2024.108336. [Google Scholar] [PubMed] [CrossRef]
174. Yang G, Wei J, Liu P, Zhang Q, Tian Y, Hou G, et al. Role of the gut microbiota in type 2 diabetes and related diseases. Metabolism. 2021;117:154712. doi:10.1016/j.metabol.2021.154712. [Google Scholar] [PubMed] [CrossRef]
175. Ahmed N, Gaur V, Kamle M, Chauhan A, Chauhan R, Kumar P, et al. Microbiome-based therapeutics for metabolic disorders: harnessing microbial intrusions for treatment. Front Med Technol. 2025;7:1695329. doi:10.3389/fmedt.2025.1695329. [Google Scholar] [PubMed] [CrossRef]
176. Depommier C, Everard A, Druart C, Plovier H, Van Hul M, Vieira-Silva S, et al. Supplementation with Akkermansia muciniphila in overweight and obese human volunteers: a proof-of-concept exploratory study. Nat Med. 2019;25(7):1096–103. doi:10.1038/s41591-019-0495-2. [Google Scholar] [PubMed] [CrossRef]
177. Hadi A, Arab A, Khalesi S, Rafie N, Kafeshani M, Kazemi M. Effects of probiotic supplementation on anthropometric and metabolic characteristics in adults with metabolic syndrome: a systematic review and meta-analysis of randomized clinical trials. Clin Nutr. 2021;40(7):4662–73. doi:10.1016/j.clnu.2021.05.027. [Google Scholar] [PubMed] [CrossRef]
178. Guo M, Li J, Zhang L, Chen C, Wei Y, Shen ZA. Effects of oral supplementation of probiotics on body weight and visceral fat in obese patients: a meta-analysis and systematic review. Sci Rep. 2025;15(1):6355. doi:10.1038/s41598-025-90820-8. [Google Scholar] [PubMed] [CrossRef]
179. Cao N, Zhao F, Kwok LY, Wang H, Sun Z. Impact of probiotics on weight loss, glucose and lipid metabolism in overweight or obese women: a meta-analysis of randomized controlled trials. Curr Res Food Sci. 2024;9:100810. doi:10.1016/j.crfs.2024.100810. [Google Scholar] [PubMed] [CrossRef]
180. Zhang Y, Gu Y, Ren H, Wang S, Zhong H, Zhao X, et al. Gut microbiome-related effects of berberine and probiotics on type 2 diabetes (the PREMOTE study). Nat Commun. 2020;11(1):5015. doi:10.1038/s41467-020-18414-8. [Google Scholar] [PubMed] [CrossRef]
181. Firouzi S, Majid HA, Ismail A, Kamaruddin NA, Barakatun-Nisak MY. Effect of multi-strain probiotics (multi-strain microbial cell preparation) on glycemic control and other diabetes-related outcomes in people with type 2 diabetes: a randomized controlled trial. Eur J Nutr. 2017;56(4):1535–50. doi:10.1007/s00394-016-1199-8. [Google Scholar] [PubMed] [CrossRef]
182. Liang T, Wu L, Xi Y, Li Y, Xie X, Fan C, et al. Probiotics supplementation improves hyperglycemia, hypercholesterolemia, and hypertension in type 2 diabetes mellitus: an update of meta-analysis. Crit Rev Food Sci Nutr. 2021;61(10):1670–88. doi:10.1080/10408398.2020.1764488. [Google Scholar] [PubMed] [CrossRef]
183. Carpi RZ, Barbalho SM, Sloan KP, Laurindo LF, Gonzaga HF, Grippa PC, et al. The effects of probiotics, prebiotics and synbiotics in non-alcoholic fat liver disease (NAFLD) and non-alcoholic steatohepatitis (NASHa systematic review. Int J Mol Sci. 2022;23(15):8805. doi:10.3390/ijms23158805. [Google Scholar] [PubMed] [CrossRef]
184. Pan Y, Yang Y, Wu J, Zhou H, Yang C. Efficacy of probiotics, prebiotics, and synbiotics on liver enzymes, lipid profiles, and inflammation in patients with non-alcoholic fatty liver disease: a systematic review and meta-analysis of randomized controlled trials. BMC Gastroenterol. 2024;24(1):283. doi:10.1186/s12876-024-03356-y. [Google Scholar] [PubMed] [CrossRef]
185. Li HY, Zhou DD, Gan RY, Huang SY, Zhao CN, Shang A, et al. Effects and mechanisms of probiotics, prebiotics, synbiotics, and postbiotics on metabolic diseases targeting gut microbiota: a narrative review. Nutrients. 2021;13(9):3211. doi:10.3390/nu13093211. [Google Scholar] [PubMed] [CrossRef]
186. Leyrolle Q, Cserjesi R, Mulders DGHM, Zamariola G, Hiel S, Gianfrancesco MA, et al. Prebiotic effect on mood in obese patients is determined by the initial gut microbiota composition: a randomized, controlled trial. Brain Behav Immun. 2021;94:289–98. doi:10.1016/j.bbi.2021.01.014. [Google Scholar] [PubMed] [CrossRef]
187. Hall CV, Twelves JL, Saxena M, Scapozza L, Gurry T. Effects of a diverse prebiotic fibre supplement on HbA1c, insulin sensitivity and inflammatory biomarkers in pre-diabetes: a pilot placebo-controlled randomised clinical trial. Br J Nutr. 2024;132(1):68–76. doi:10.1017/S0007114524000904. [Google Scholar] [PubMed] [CrossRef]
188. Teparak C, Uriyapongson J, Phoemsapthawee J, Tunkamnerdthai O, Aneknan P, Tong-Un T, et al. Diabetes therapeutics of prebiotic soluble dietary fibre and antioxidant anthocyanin supplement in patients with type 2 diabetes: randomised placebo-controlled clinical trial. Nutrients. 2025;17(7):1098. doi:10.3390/nu17071098. [Google Scholar] [PubMed] [CrossRef]
189. Stachowska E, Portincasa P, Jamioł-Milc D, Maciejewska-Markiewicz D, Skonieczna-Żydecka K. The relationship between prebiotic supplementation and anthropometric and biochemical parameters in patients with NAFLD-a systematic review and meta-analysis of randomized controlled trials. Nutrients. 2020;12(11):3460. doi:10.3390/nu12113460. [Google Scholar] [PubMed] [CrossRef]
190. Visuthranukul C, Sriswasdi S, Tepaamorndech S, Chamni S, Leelahavanichkul A, Joyjinda Y, et al. Enhancing gut microbiota and microbial function with inulin supplementation in children with obesity. Int J Obes. 2024;48(12):1696–704. doi:10.1038/s41366-024-01590-8. [Google Scholar] [PubMed] [CrossRef]
191. Rasaei N, Heidari M, Esmaeili F, Khosravi S, Baeeri M, Tabatabaei-Malazy O, et al. The effects of prebiotic, probiotic or synbiotic supplementation on overweight/obesity indicators: an umbrella review of the trials’ meta-analyses. Front Endocrinol. 2024;15:1277921. doi:10.3389/fendo.2024.1277921. [Google Scholar] [PubMed] [CrossRef]
192. Niu X, Zhang Q, Liu J, Zhao Y, Shang N, Li S, et al. Effect of synbiotic supplementation on obesity and gut microbiota in obese adults: a double-blind randomized controlled trial. Front Nutr. 2024;11:1510318. doi:10.3389/fnut.2024.1510318. [Google Scholar] [PubMed] [CrossRef]
193. Mirmiranpour H, Huseini HF, Derakhshanian H, Khodaii Z, Tavakoli-Far B. Effects of probiotic, cinnamon, and synbiotic supplementation on glycemic control and antioxidant status in people with type 2 diabetes; a randomized, double-blind, placebo-controlled study. J Diabetes Metab Disord. 2019;19(1):53–60. doi:10.1007/s40200-019-00474-3. [Google Scholar] [PubMed] [CrossRef]
194. Naseri K, Saadati S, Ashtary-Larky D, Asbaghi O, Ghaemi F, Pashayee-Khamene F, et al. Probiotics and synbiotics supplementation improve glycemic control parameters in subjects with prediabetes and type 2 diabetes mellitus: a GRADE-assessed systematic review, meta-analysis, and meta-regression of randomized clinical trials. Pharmacol Res. 2022;184:106399. doi:10.1016/j.phrs.2022.106399. [Google Scholar] [PubMed] [CrossRef]
195. Wang H, Lu Y, Yan Y, Tian S, Zheng D, Leng D, et al. Promising treatment for type 2 diabetes: fecal microbiota transplantation reverses insulin resistance and impaired islets. Front Cell Infect Microbiol. 2020;9:455. doi:10.3389/fcimb.2019.00455. [Google Scholar] [PubMed] [CrossRef]
196. Vrieze A, Van Nood E, Holleman F, Salojärvi J, Kootte RS, Bartelsman JFWM, et al. Transfer of intestinal microbiota from lean donors increases insulin sensitivity in individuals with metabolic syndrome. Gastroenterology. 2012;143(4):913–6.e7. doi:10.1053/j.gastro.2012.06.031. [Google Scholar] [PubMed] [CrossRef]
197. Bakhshimoghaddam F, Shateri K, Sina M, Hashemian M, Alizadeh M. Daily consumption of synbiotic yogurt decreases liver steatosis in patients with nonalcoholic fatty liver disease: a randomized controlled clinical trial. J Nutr. 2018;148(8):1276–84. doi:10.1093/jn/nxy088. [Google Scholar] [PubMed] [CrossRef]
198. Abhari K, Saadati S, Yari Z, Hosseini H, Hedayati M, Abhari S, et al. The effects of Bacillus coagulans supplementation in patients with non-alcoholic fatty liver disease: a randomized, placebo-controlled, clinical trial. Clin Nutr ESPEN. 2020;39(1 Suppl):53–60. doi:10.1016/j.clnesp.2020.06.020. [Google Scholar] [PubMed] [CrossRef]
199. Mitrović M, Dobrosavljević A, Odanović O, Knežević-Ivanovski T, Kralj Đ, Erceg S, et al. The effects of synbiotics on the liver steatosis, inflammation, and gut microbiome of metabolic dysfunction-associated liver disease patients-randomized trial. Rom J Intern Med. 2024;62(2):184–93. doi:10.2478/rjim-2024-0004. [Google Scholar] [PubMed] [CrossRef]
200. Witjes JJ, Smits LP, Pekmez CT, Prodan A, Meijnikman AS, Troelstra MA, et al. Donor fecal microbiota transplantation alters gut microbiota and metabolites in obese individuals with steatohepatitis. Hepatol Commun. 2020;4(11):1578–90. doi:10.1002/hep4.1601. [Google Scholar] [PubMed] [CrossRef]
201. Wang LJ, Sun JG, Chen SC, Sun YL, Zheng Y, Feng JC. The role of intestinal flora in metabolic dysfunction-associated steatotic liver disease and treatment strategies. Front Med. 2025;11:1490929. doi:10.3389/fmed.2024.1490929. [Google Scholar] [PubMed] [CrossRef]
202. van Nood E, Vrieze A, Nieuwdorp M, Fuentes S, Zoetendal EG, de Vos WM, et al. Duodenal infusion of donor feces for recurrent Clostridium difficile. N Engl J Med. 2013;368(5):407–15. doi:10.1056/NEJMoa1205037. [Google Scholar] [CrossRef]
203. Cammarota G, Ianiro G, Tilg H, Rajilić-Stojanović M, Kump P, Satokari R, et al. European consensus conference on faecal microbiota transplantation in clinical practice. Gut. 2017;66(4):569–80. doi:10.1136/gutjnl-2016-313017. [Google Scholar] [PubMed] [CrossRef]
204. Kootte RS, Levin E, Salojärvi J, Smits LP, Hartstra AV, Udayappan SD, et al. Improvement of insulin sensitivity after lean donor feces in metabolic syndrome is driven by baseline intestinal microbiota composition. Cell Metab. 2017;26(4):611–9.e6. doi:10.1016/j.cmet.2017.09.008. [Google Scholar] [PubMed] [CrossRef]
205. Napolitano M, Covasa M. Microbiota transplant in the treatment of obesity and diabetes: current and future perspectives. Front Microbiol. 2020;11:590370. doi:10.3389/fmicb.2020.590370. [Google Scholar] [PubMed] [CrossRef]
206. Mederle AL, Dima M, Stoicescu ER, Capastraru BF, Levai CM, Hategan OA, et al. Impact of gut microbiome interventions on glucose and lipid metabolism in metabolic diseases: a systematic review and meta-analysis. Life. 2024;14(11):1485. doi:10.3390/life14111485. [Google Scholar] [PubMed] [CrossRef]
207. Le Roy T, Llopis M, Lepage P, Bruneau A, Rabot S, Bevilacqua C, et al. Intestinal microbiota determines development of non-alcoholic fatty liver disease in mice. Gut. 2013;62(12):1787–94. doi:10.1136/gutjnl-2012-303816. [Google Scholar] [PubMed] [CrossRef]
208. Craven L, Rahman A, Nair Parvathy S, Beaton M, Silverman J, Qumosani K, et al. Allogenic fecal microbiota transplantation in patients with nonalcoholic fatty liver disease improves abnormal small intestinal permeability: a randomized control trial. Am J Gastroenterol. 2020;115(7):1055–65. doi:10.14309/ajg.0000000000000661. [Google Scholar] [PubMed] [CrossRef]
209. Lanthier N, Delzenne N. Targeting the gut microbiome to treat metabolic dysfunction-associated fatty liver disease: ready for prime time? Cells. 2022;11(17):2718. doi:10.3390/cells11172718. [Google Scholar] [PubMed] [CrossRef]
210. Manrique P, Zhu Y, van der Oost J, Herrema H, Nieuwdorp M, de Vos WM, et al. Gut bacteriophage dynamics during fecal microbial transplantation in subjects with metabolic syndrome. Gut Microbes. 2021;13(1):1897217. doi:10.1080/19490976.2021.1897217. [Google Scholar] [PubMed] [CrossRef]
211. DeFilipp Z, Bloom PP, Torres Soto M, Mansour MK, Sater MRA, Huntley MH, et al. Drug-ResistantE. coliBacteremia transmitted by fecal microbiota transplant. N Engl J Med. 2019;381(21):2043–50. doi:10.1056/nejmoa1910437. [Google Scholar] [PubMed] [CrossRef]
212. Park SY, Seo GS. Fecal microbiota transplantation: is it safe? Clin Endosc. 2021;54(2):157–60. doi:10.5946/ce.2021.072. [Google Scholar] [PubMed] [CrossRef]
213. Cammarota G, Ianiro G, Kelly CR, Mullish BH, Allegretti JR, Kassam Z, et al. International consensus conference on stool banking for faecal microbiota transplantation in clinical practice. Gut. 2019;68(12):2111–21. doi:10.1136/gutjnl-2019-319548. [Google Scholar] [PubMed] [CrossRef]
214. Aron-Wisnewsky J, Clément K, Nieuwdorp M. Fecal microbiota transplantation: a future therapeutic option for obesity/diabetes? Curr Diab Rep. 2019;19(8):51. doi:10.1007/s11892-019-1180-z. [Google Scholar] [PubMed] [CrossRef]
215. Min H, Choi KS, Yun S, Jang S. Live biotherapeutic products for metabolic diseases: development strategies, challenges, and future directions. J Microbiol Biotechnol. 2025;35:e2410054. doi:10.4014/jmb.2410.10054. [Google Scholar] [PubMed] [CrossRef]
216. Everard A, Belzer C, Geurts L, Ouwerkerk JP, Druart C, Bindels LB, et al. Cross-talk between Akkermansia muciniphila and intestinal epithelium controls diet-induced obesity. Proc Natl Acad Sci USA. 2013;110(22):9066–71. doi:10.1073/pnas.1219451110. [Google Scholar] [PubMed] [CrossRef]
217. Plovier H, Everard A, Druart C, Depommier C, Van Hul M, Geurts L, et al. A purified membrane protein from Akkermansia muciniphila or the pasteurized bacterium improves metabolism in obese and diabetic mice. Nat Med. 2017;23(1):107–13. doi:10.1038/nm.4236. [Google Scholar] [PubMed] [CrossRef]
218. Ott SJ, Waetzig GH, Rehman A, Moltzau-Anderson J, Bharti R, Grasis JA, et al. Efficacy of sterile fecal filtrate transfer for treating patients with Clostridium difficile infection. Gastroenterology. 2017;152(4):799–811.e7. doi:10.1053/j.gastro.2016.11.010. [Google Scholar] [PubMed] [CrossRef]
219. Shkoporov AN, Hill C. Bacteriophages of the human gut: the known unknown of the microbiome. Cell Host Microbe. 2019;25(2):195–209. doi:10.1016/j.chom.2019.01.017. [Google Scholar] [PubMed] [CrossRef]
220. Duan Y, Llorente C, Lang S, Brandl K, Chu H, Jiang L, et al. Bacteriophage targeting of gut bacterium attenuates alcoholic liver disease. Nature. 2019;575(7783):505–11. doi:10.1038/s41586-019-1742-x. [Google Scholar] [PubMed] [CrossRef]
221. Charbonneau MR, Isabella VM, Li N, Kurtz CB. Developing a new class of engineered live bacterial therapeutics to treat human diseases. Nat Commun. 2020;11(1):1738. doi:10.1038/s41467-020-15508-1. [Google Scholar] [PubMed] [CrossRef]
222. Isabella VM, Ha BN, Castillo MJ, Lubkowicz DJ, Rowe SE, Millet YA, et al. Development of a synthetic live bacterial therapeutic for the human metabolic disease phenylketonuria. Nat Biotechnol. 2018;36(9):857–64. doi:10.1038/nbt.4222. [Google Scholar] [PubMed] [CrossRef]
223. Leung H, Long X, Ni Y, Qian L, Nychas E, Siliceo SL, et al. Risk assessment with gut microbiome and metabolite markers in NAFLD development. Sci Transl Med. 2022;14(648):eabk0855. doi:10.1126/scitranslmed.abk0855. [Google Scholar] [PubMed] [CrossRef]
224. Long J, Fang Q, Shi Z, Miao Z, Yan D. Integrated biomarker profiling for predicting the response of type 2 diabetes to metformin. Diabetes Obes Metab. 2024;26(8):3439–47. doi:10.1111/dom.15689. [Google Scholar] [PubMed] [CrossRef]
225. Jeong K, Moon SJ, Rachim VP, Song Y, Cho YM, Park SM. Enhanced post-prandial glycemic response prediction in type 2 diabetes with microbiome data and deep learning. IEEE J Biomed Health Inform. 2025. doi:10.1109/JBHI.2025.3602827. [Google Scholar] [PubMed] [CrossRef]
226. Bar N, Korem T, Weissbrod O, Zeevi D, Rothschild D, Leviatan S, et al. A reference map of potential determinants for the human serum metabolome. Nature. 2020;588(7836):135–40. doi:10.1038/s41586-020-2896-2. [Google Scholar] [PubMed] [CrossRef]
227. Sanders ME, Merenstein DJ, Reid G, Gibson GR, Rastall RA. Probiotics and prebiotics in intestinal health and disease: from biology to the clinic. Nat Rev Gastroenterol Hepatol. 2019;16(10):605–16. doi:10.1038/s41575-019-0173-3. [Google Scholar] [PubMed] [CrossRef]
228. Zhang Z, Mocanu V, Cai C, Dang J, Slater L, Deehan EC, et al. Impact of fecal microbiota transplantation on obesity and metabolic syndrome-a systematic review. Nutrients. 2019;11(10):2291. doi:10.3390/nu11102291. [Google Scholar] [PubMed] [CrossRef]
229. Rahaman MM, Wangchuk P, Sarker S. Therapeutic potential targeting gut microbiota modulation with emphasis on Lactobacillus spp. in common metabolic disorders: a systematic review. Scientifica. 2025;2025(1):3367875. doi:10.1155/sci5/3367875. [Google Scholar] [PubMed] [CrossRef]
230. Park G, Jung S, Wellen KE, Jang C. The interaction between the gut microbiota and dietary carbohydrates in nonalcoholic fatty liver disease. Exp Mol Med. 2021;53(5):809–22. doi:10.1038/s12276-021-00614-x. [Google Scholar] [PubMed] [CrossRef]
231. Kulkarni AS, Carrara GMP, Jin J, Laro J, Peramuna T, McCall LI, et al. Mass spectrometry-based metabolomics approaches to interrogate host-microbiome interactions in mammalian systems. Nat Prod Rep. 2025. doi:10.1039/d5np00021a. [Google Scholar] [PubMed] [CrossRef]
232. Ciocan D, Elinav E. Engineering bacteria to modulate host metabolism. Acta Physiol. 2023;238(3):e14001. doi:10.1111/apha.14001. [Google Scholar] [PubMed] [CrossRef]
Cite This Article
 Copyright © 2026 The Author(s). Published by Tech Science Press.
Copyright © 2026 The Author(s). Published by Tech Science Press.This work is licensed under a Creative Commons Attribution 4.0 International License , which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.


 Submit a Paper
Submit a Paper Propose a Special lssue
Propose a Special lssue View Full Text
View Full Text Download PDF
Download PDF

 Downloads
Downloads
 Citation Tools
Citation Tools