 Open Access
Open Access
REVIEW
Tumor Suppressor p53 and MicroRNAs Interaction in Breast Cancer
1 Herminio Ometto Foundation, Araras, São Paulo, 13607-339, Brazil
2 Laboratory of Cancer Genetics, School of Medical Sciences, University of Campinas, Campinas, São Paulo, 13083-888, Brazil
* Corresponding Author: Juliana Carron. Email:
Oncology Research 2026, 34(3), 6 https://doi.org/10.32604/or.2025.072133
Received 20 August 2025; Accepted 17 November 2025; Issue published 24 February 2026
Abstract
This literature review explores the complex interaction between p53 and microRNAs (miRNAs) in the occurrence and progression of breast cancer (BC), the most common and lethal tumor type among women. BC is a multifactorial disease resulting from a combination of genetic and epigenetic alterations in cell DNA, influencing proliferation, differentiation, and migration. TP53 gene, which codifies p53 protein, is a known tumor suppressor, and it plays an important role in cell maintenance as DNA repair, cell proliferation control, and apoptosis activation. TP53 expression can be modulated by several miRNAs, as miR-30c, miR-34a, and the miR-200 family, inhibiting p53 production and silencing its tumor suppressor effects. On the other hand, p53 protein can modulate several miRNAs expression, as miR-146a, miR-192, and the miR-200 family, by acting as a transcription factor or by modulating miRNA processing, interfering with BC aggressiveness and progression. Understanding the role of p53 and miRNAs in BC may aid in identifying new biomarkers and developing new targeted therapies for patient treatment.Keywords
Worldwide, breast cancer (BC) is the most common and most lethal tumor type among women, accounting for 23.8% of all cancer cases and 15.4% of deaths due to the disease [1]. The male population can also develop BC; however, it is less common, accounting for less than 1% of all BC cases [2].
BC can be classified according to the TNM system from the American Joint Committee on Cancer (AJCC) [3] and by its histopathological aspects, especially as invasive ductal, lobular, and tubular carcinoma [4]. In addition, BC classification is defined by the expression of hormone receptors and growth factors, as estrogen receptor (ER), progesterone receptor (PR), and epidermal growth factor receptor 2 (HER-2/ERBB2) [5]. The luminal A subtype (ER and PR positive, HER2 negative) is the most common molecular form, accounting for up to 50% of cases, and is generally associated with a less aggressive profile and better disease prognosis [6]. Luminal B tumors (ER, PR, and HER2 positive or negative) tend to be more aggressive than luminal A due to higher cellular proliferation rates and represent approximately 20% of cases [6]. HER2 positive tumors account for about 15% of cases and are associated with higher recurrence rates; however, they are highly responsive to anti-HER2 targeted therapy [6]. On the other hand, triple-negative or basal-like tumors are considered to have the worst prognosis, as they lack specific therapeutic targets and do not respond to current treatment options [6]. Triple-negative tumors comprise around 15% of cases and are commonly associated with genetic mutations [6].
BC-related cells comprise not only malignant epithelial cells but also a heterogeneous population of non-tumoral cells that constitute the tumor microenvironment (TME). The TME plays a critical role in BC development and progression, being composed of immune, stromal, and endothelial cells, and the extracellular matrix (ECM) [7]. Immune infiltrates include both lymphoid and myeloid-derived populations that mediate innate and adaptive immune responses. Cancer-associated fibroblasts (CAFs) establish reciprocal communication with tumor cells, promoting morphological and mechanical changes that enhance tumor invasiveness. Tumor-associated endothelial cells (TECs) contribute to angiogenesis and interact with other TME components to support vascular stability. Additionally, dynamic remodeling of the ECM by proteolytic enzymes facilitates tumor cell migration, invasion, and metastasis, underscoring the importance of TME in shaping BC behavior [7].
Patients’ therapeutic depends on the clinical, histopathological, and molecular characteristics of the tumor and may include surgery, chemotherapy, radiotherapy, endocrine and/or immunotherapy [8]. The survival rate ranges from 80% for early tumor diagnosis to 10% for late-stage detection [9,10].
The development of BC is multifactorial. Risk factors for BC include increasing age, reproductive characteristics, contraceptive and hormone replacement therapy, BC family history, alcoholism, smoking, and physical inactivity [11]. In addition to the environmental risk factors, genetic and epigenetic predisposition may also influence BC risk and progression [11]. Single-nucleotide variant (SNV), copy number variation (CNV); histone modification, DNA methylation, and post-transcriptional regulation by microRNAs (miRNAs) are important genetic and epigenetic, respectively, alterations in BC [11–13].
Breast cancer 1 (BRCA1) and breast cancer 2 (BRCA2) are well-known mutated genes in BC [11,14]. In addition, other genes are also associated with BC, as ataxia telangiectasia mutated (ATM), cadherin 1 (CDH1), CHEK2 (checkpoint kinase 2), phosphatase and tensin homolog (PTEN), and tumor protein p53 (TP53) [11,14]. Between them, TP53 gene is frequently altered in BC and may interfere directly with patients’ treatment success [14].
Current studies have presented substantial evidence that TP53 participates in a complex gene regulatory network involving miRNAs, which plays critical roles in BC development, progression, and response to therapy [15]. TP53 gene expression can be modulated through the binding of specific miRNAs, leading to changes in the levels and activity of the p53 protein, encoded by TP53. Conversely, the p53 protein can transcriptionally regulate a variety of miRNAs, thereby influencing multiple cellular processes, including cell proliferation, DNA repair, apoptosis, and metastasis [15]. This bidirectional regulation highlights the feedback loops that exist between TP53 and miRNAs, demonstrating their importance in maintaining cellular homeostasis and in tumorigenesis. In this review, we focused on this complex regulatory network of TP53 and miRNAs in BC, aiming to summarize current knowledge on their interactions and to discuss their potential implications for prognostic and therapeutic strategies.
TP53 codifies the tumor suppressor protein p53 that is activated in response to various cellular stresses, as lack of nutrients, hypoxia, and DNA damage [16]. The tumor-suppressor activity of p53 is mainly attributed to its ability to modulate the expression of several genes involved in cell cycle, DNA repair, apoptosis, senescence, and cell metabolism, in order to protect the DNA and maintain the tissue homeostasis [16].
The p53 belongs to a multiprotein family of transcription factors that also includes p63 and p73. p53 is a phosphorylation target of the kinases ATM, ataxia telangiectasia and Rad3-related (ATR), checkpoint kinase 1 (CHEK1), and CHEK2, which act in coordination to promote its stabilization [17]. These post-translational modifications play a crucial role in stabilizing p53 by disrupting its interaction with the negative regulators double minute 2 protein (MDM2) and double minute 4 protein (MDM4) [17]. MDM2 and MDM4 bind to the transactivation domains of p53, thereby suppressing its transcriptional activity. In addition, MDM2 functions as an E3 ubiquitin ligase that targets p53 for proteasome-dependent degradation [17]. In normal cells, p53 is kept at low levels by MDM2, which is transcriptionally regulated by p53, forming a negative-feedback loop [18]. Additionally, p53 can function as a transcriptional repressor, notably downregulating genes such as transcription factor AP-1 subunit C-Fos (FOS), MYC proto-oncogene, bHLH transcription factor (MYC), vascular endothelial growth factor A (VEGFA), and BIRC5 (survivin), inhibiting proliferation, survival, and angiogenesis [18].
TP53 importance in tumor progression is demonstrated by the high frequency of mutations found in several types of cancer, including BC [19]. SNVs are the most common mutation in TP53 gene, and it can influence protein-DNA binding domains (as p53R248Q and p53R273H mutations) or cause a partial or total distortion of the correct folding of p53 protein (as p53R175H and p53H179R mutations) [19].
SNVs in TP53 tend to accumulate in tumor cells, leading to the loss of tumor suppressor functions [19]. Besides, SNVs in TP53 may also favor BC progression, increasing proliferation, inflammation, angiogenesis, invasion, and chemotherapy resistance, and inhibiting apoptosis, through the called gain-of-function [20].
In addition, TP53 expression can be mediated by miRNAs binding [15]. The interaction of miRNAs and TP53 messenger RNA (mRNA) may decrease or even silence the production of p53 protein [15]. On the other hand, p53 protein can also regulate the expression of several miRNAs [14,20]. p53 can interact directly to the promoter region of miRNAs genes or by binding to Drosha complex, responsible for miRNAs maturation, influencing their transcription [15,21]. This complex interaction of TP53 and miRNAs impacts the occurrence and progression of BC [22–24].
miRNAs are endogenous small non-coding RNAs with approximately 22 nucleotides that silence gene expression post-transcriptionally through degradation or inhibition of genes’ mRNA [25], including those related to BC carcinogenesis [26,27].
In miRNA gene transcription, RNA polymerase II or III will process the primary miRNA (pri-miRNA) in a structure called hairpin [28]. In the nucleus, the pri-miRNA is processed by Drosha complex, forming the precursor miRNA (pre-miRNA), a large double-stranded RNA, which will be transported to the cytoplasm by Exportin-5 [28]. In the cytoplasm, pre-miRNA is cleaved by the ribonuclease Dicer, producing the mature miRNA [28]. The RNA-induced silencing complex (RISC) is responsible for separating the double strand from the mature miRNA, in which one strand is frequently cleaved and discarded and the other, called guide strand, is guided to the target mRNA [28].
Thus, miRNAs can either degrade target mRNA or repress protein translation through complete or partial base complementarity, respectively. In humans, complementarity between miRNAs and their target mRNAs is typically partial, leading to translational repression without degradation of the mRNA strand [29]. miRNAs contain complementary regions that bind primarily to the 3′ untranslated region (3′-UTR) of target genes, but can also interact with sequences within the 5′-UTR and coding regions, exerting silencing effects on gene expression [28]. Moreover, miRNAs’ interaction with promoter regions has been reported to induce transcription, highlighting their capacity to regulate gene expression through diverse and context-dependent mechanisms [30].
In the context of cancer, miRNAs could be divided into two groups: oncogenic miRNAs or oncomiRNAs, and tumor suppressive miRNAs, depending on their target genes and the resulting effects on tumor development. OncomiRNAs promote tumor progression by negatively regulating tumor suppressor genes that control cell survival, differentiation, apoptosis, invasion, and migration. In contrast, tumor suppressive miRNAs inhibit tumor progression by negatively regulating oncogenes that promote cell survival, differentiation, apoptosis, invasion, and migration [31].
4 TP53 Expression Modulated by miRNAs
Some miRNAs can bind directly to the TP53 mRNA, inhibiting p53 production and favoring BC development and progression, acting as oncomiRNAs [31].
Reports from the literature have demonstrated that miR-214 [32], miR-504 [33], miR-663a [34], and miR-1204 [35] can bind to the TP53 mRNA, inhibiting gene expression and increasing cell proliferation and invasion, and decreasing apoptosis of BC cells. In addition, miR-105, miR-200c, miR-659, miR-662, and miR-921 were associated with decreased TP53 expression, which may contribute to increased invasion and migration of BC cells [36].
miR-19a, miR-19b [37], miR-106a [38], and miR-8084 [39] were also associated with decreased TP53 expression, leading to increased cell proliferation and migration, along with decreased apoptosis and sensitivity to cisplatin chemotherapy in BC cell lines. Other oncomiRNAs, as miR-10b, miR-30c, miR-34a, miR-373 [40], miR-26a [41], and miR-155 [42], have been associated with decreased TP53 expression in BC; however, the exact mechanisms involved in BC carcinogenesis remain unclear. An overview of miRNAs involved in modulating TP53 is schematically illustrated in Fig. 1A.
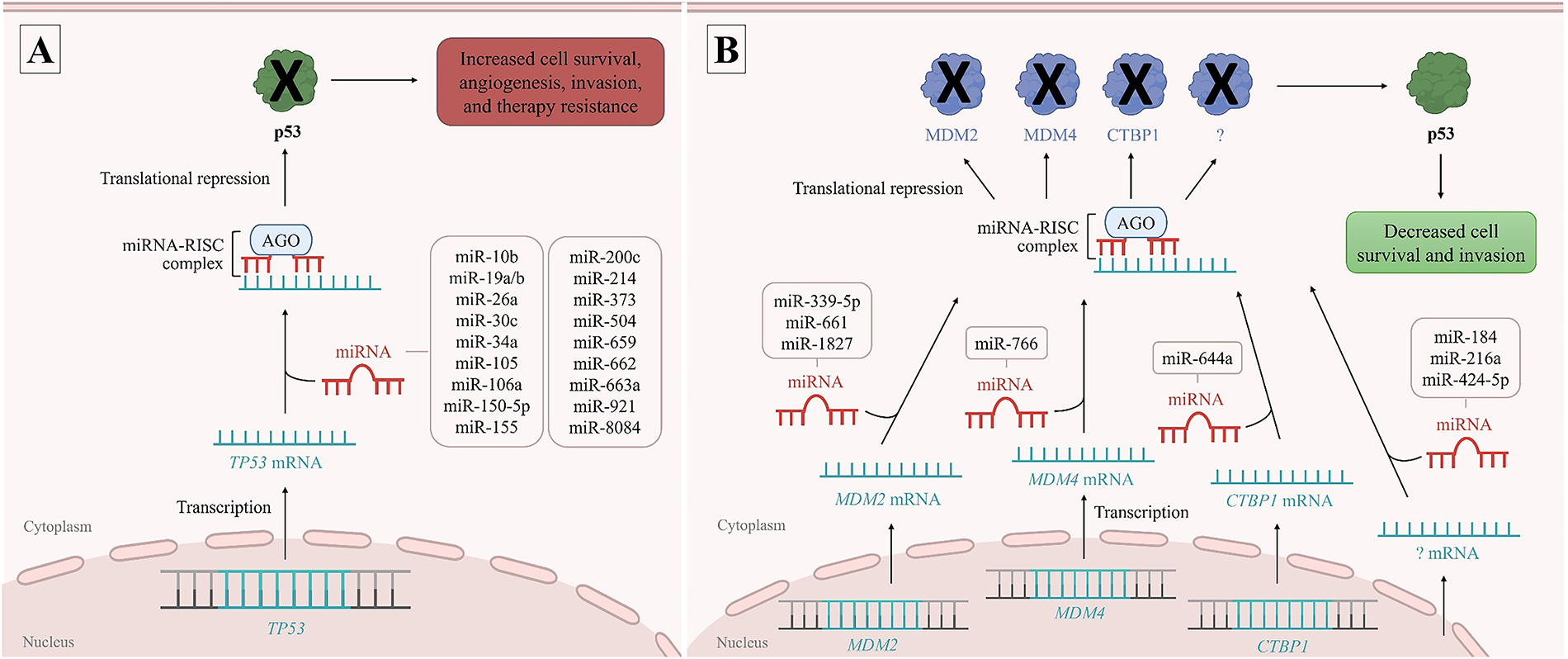
Figure 1: TP53 expression modulated by microRNAs (miRNAs). Several miRNAs can bind directly to the TP53 messenger RNA (mRNA), inhibiting p53 production and favoring breast cancer (BC) development and progression, acting as oncomiRNAs (A). Other miRNAs can regulate TP53 expression indirectly by binding and suppressing TP53 negative regulators, as MDM2, MDM4, and CTBP1 genes, thereby allowing p53 production and acting as tumor suppressor miRNAs (B). RISC complex: RNA-induced silencing complex, AGO: argonaute proteins. Figure made in PowerPoint (Microsoft 365) using icons sourced from the public domain
In contrast, other miRNAs can regulate TP53 expression indirectly. These miRNAs can bind to TP53 negative regulators, thereby allowing p53 production and acting as tumor suppressor miRNAs [31]. Among them, miRNAs miR-339-5p [43], miR-661 [44], miR-1827 [45], miR-766 [46], and miR-644a [47] were associated with TP53 upregulation by targeting the TP53 regulators MDM2, MDM4, and CTBP1 (C-terminal binding protein 1), respectively. Other miRNAs, such as miR-184 [48], miR-216a [49], and miR-424-5p [50] may exert similar effects, although the exact TP53 regulator involved remains unclear. Collectively, these tumor-suppressive miRNAs enhance p53-mediated apoptosis, cell cycle arrest, and inhibition of invasion in BC cells. These associations are schematically illustrated in Fig. 1B.
Interestingly, miR-193a-5p [51] and miR-3646 [52] were associated with TP53 downregulation and decreased cell proliferation and migration, and increased apoptosis and paclitaxel chemotherapy sensitivity. Similarly, miR-150-5p was associated with TP53 downregulation, but also with decreased tumor progression and improved patient survival [53]. These results counteract the expected effects of low TP53 expression in BC, and further studies are needed to explain this association. One possible explanation is that these miRNAs may downregulate TP53 while simultaneously targeting other cancer-related genes, thereby compensating for TP53 loss.
A comprehensive overview of miRNAs involved in modulating TP53 expression in BC can be found in Table 1.
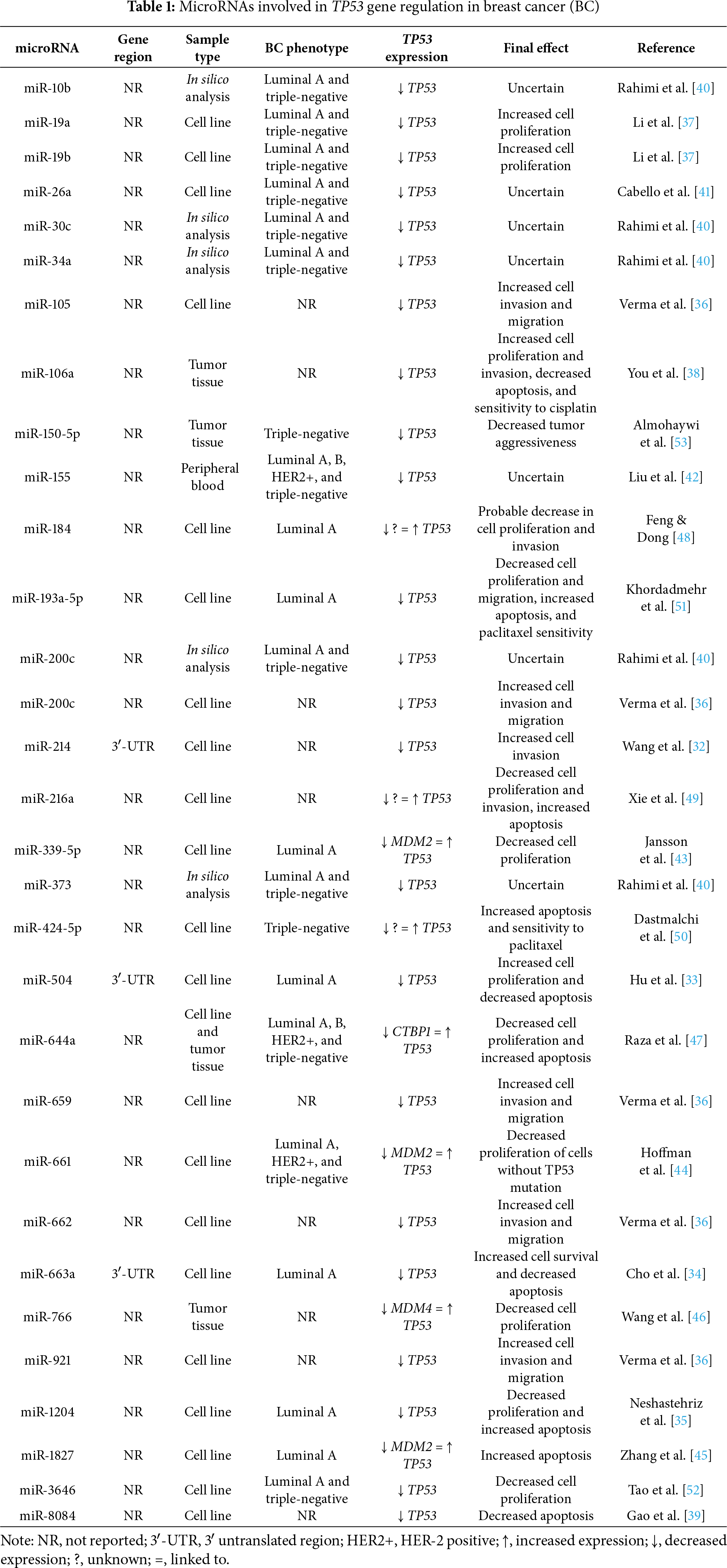
When considering BC subtypes, different miRNAs associated with TP53 regulation appear to display subtype-specific patterns. For instance, miR-10b [40], miR-19a, miR-19b [37], miR-26a [41], miR-30c, miR-34a, miR-200c, miR-373 [40], and miR-3646 [52] were reported in luminal A and triple-negative BC, whereas miR-184 [48], miR-193a-5p [51], miR-339-5p [43], miR-504 [33], miR-663a [34], and miR-1827 [45] were reported only in luminal A, and miR-150-5p [53] and miR-424-5p [50] only in triple-negative BC. On the other hand, miR-661 [44] was reported in luminal A, HER2-positive, and triple-negative BC, whereas miR-155 [42] and miR-644a [47] were reported in all four subtypes (luminal A, luminal B, HER2-positive, and triple-negative) (Table 1).
These observations indicate that the functional impact of each miRNA on TP53 regulation may depend not only on its molecular target but also on the intrinsic biology of each BC subtype. Understanding these relationships is particularly critical for triple-negative BC, where the absence of targeted treatment options makes the discovery of novel miRNA–TP53–based therapeutic strategies especially valuable.
5 miRNAs Expression Modulated by p53
The p53 protein can control the expression level of several miRNAs through two main mechanisms, particularly by acting as a transcription factor and by modulating miRNA processing [15,21]. In addition to regulating protein-coding genes, p53 also modulates the expression of several miRNA genes by directly binding to their DNA sequences and primarily promoting their transcription [54]. Although p53-mediated transcriptional repression has been described in the literature, the mechanisms underlying this process remain less well understood [54].
Beyond transcriptional regulation, p53 also plays a role in the post-transcriptional maturation of miRNAs. It has been demonstrated that p53 directly binds to the DEAD-box RNA helicase p68 (DDX5), facilitating its association with the Drosha complex, which is responsible for miRNAs maturation [54]. Through this interaction, p53 contributes to the conversion of pri-miRNAs into pre-miRNAs, thereby enhancing the production of mature miRNAs [54].
In BC, p53 induces the expression of miR-30a [55], miR-34a [56–58], miR-101, miR-124, miR-141 [59], miR-183 [60], miR-192 [59], miR-200b [61], miR-200c [60,62], miR-205 [63], miR-429 [61], and miR-506 [59] by binding to their promoters and inducing transcription. The upregulation of these miRNAs have been associated with the suppression of epithelial–mesenchymal transition (EMT), through the inhibition of transcription factors as Zeb1, Zeb2, and Snail, thereby interfering with the aggressiveness and progression of the disease, acting as tumor suppressive miRNAs [59]. p53 also induces the expression of miR-10, miR-22, miR-26a [22], miR-30c [64], miR-34b [65], and miR-148a [22] by binding to their promoters. These tumor suppressive miRNAs have been associated with a better prognosis in BC by decreasing cell proliferation, invasion, and therapy resistance. In addition, p53 promotes the processing of tumor suppressive miR-15a [23]; miR-16, miR-145, and miR-203 [22] from primary to precursor miRNA, leading to cell apoptosis in response to DNA damage; and decreasing cell proliferation, respectively. An overview of miRNAs expression modulated by p53 is schematically illustrated in Fig. 2A.
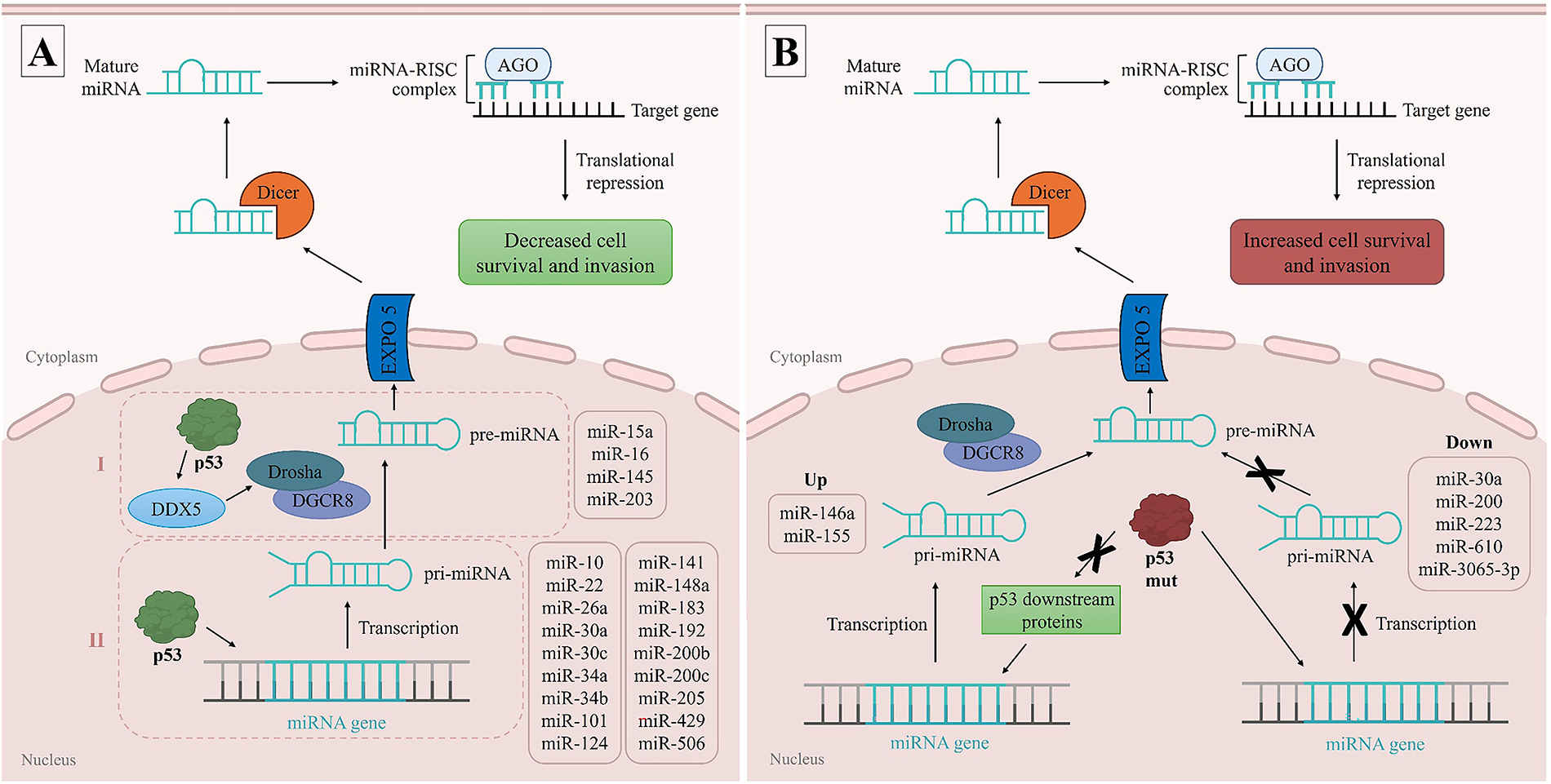
Figure 2: MicroRNAs (miRNAs) expression modulated by p53. The p53 protein can control the expression level of several miRNAs by directly binding to their DNA sequences and primarily promoting their transcription (I), or by interacting with the DEAD-box RNA helicase p68 (DDX5), facilitating its association with the Drosha complex and modulating miRNA processing (II) (A). The expression levels of certain miRNAs are influenced by genetic mutations in p53. Mutant p53 (p53 mut), as R273H variant, can bind to promoter regions and repress miRNA expression. On the other hand, p53 mut can also bind to downstream targets of p53, as tumor protein p63, allowing miRNA expression. These dysregulations contribute to the development and progression of BC (B). pri-miRNA: primary miRNA, pre-miRNA: precursor miRNA, EXPO5: exportin-5, RISC complex: RNA-induced silencing complex, AGO: argonaute proteins, up: upregulated miRNAs, down: repressed miRNAs. Figure made in PowerPoint (Microsoft 365) using icons sourced from the public domain
Moreover, the expression levels of some miRNAs, such as miR-30a [66], miR-146a [67], miR-155 [68], miR-200 [69], miR-223 [70], miR-610, and miR-3065-3p [71] are influenced by genetic mutations in p53. In BC cells, mutant p53, as the R273H variant, have been observed to bind to the promoter regions and repress the expression of miR-30a [66], miR-200 [69], miR-223 [70], miR-610, and miR-3065-3p [71] thereby increasing cell invasion and migration. On the other hand, the expression of miR-146a [67] and miR-155 [68] was found to be higher in BC cells harboring p53 mutations, likely due to the inactivation of downstream targets of p53, as tumor protein p63 [68]. This dysregulation contributes to increased cell proliferation and chemotherapy resistance, decreased apoptosis, and worse patient prognosis. These associations are schematically illustrated in Fig. 2B.
Finally, miR-10 [22,40], miR-26a [22,41], miR-30c [40,64], miR-34a [40,56], miR-155 [42,68], and miR-200c [40,72] were both regulated by p53 and able to regulate TP53 gene. This observation evidences the complex regulatory feedback loop between p53 and miRNAs.
A complete approach of miRNAs modulated by p53 in BC can be found in Table 2.
When considering BC subtypes, miR-15a [23], miR-124 [59], miR-146a [67], miR-192 [59], miR-200c [60,62,72], and miR-506 [59] modulation were reported in luminal A and triple-negative BC, whereas miR-200 [69] was reported only in luminal A, and miR-30a [55] and miR-205 [22,63] only in triple-negative BC. The modulation of miR-30c [64] was reported in luminal A, luminal B, and triple-negative BC; and miR-155 [68] in luminal B and triple-negative BC. miR-34b [65] modulation was reported in HER2-positive BC, whereas miR-223 [70] was reported in HER2 positive and triple-negative, and miR-34a [57], miR-610, and miR-3065–3p [71] were reported in all four subtypes (luminal A, luminal B, HER2 positive, and triple-negative) (Table 2).
It is important to highlight that this distribution may not be definitive, as future studies using other BC cell lines or patient cohorts may reveal that these miRNAs are also present in additional BC subtypes, thereby refining our current understanding of their subtype specificity.
6 miRNA-Based Therapies Targeting the p53-miRNA Axis
miRNA-based therapy is an emerging approach that aims to regulate gene expression by targeting miRNAs and is being studied in a variety of conditions. By correcting miRNA imbalances, this strategy holds promise for the treatment of several diseases, including BC [73]. Drug resistance and tumor relapse are major factors that compromise the survival of BC patients, underscoring the need for alternative therapeutic approaches to treat BC [73].
miRNA therapeutics are oligonucleotides capable of inhibiting oncogenic miRNAs or replacing tumor suppressive miRNAs. In the last decades, some miRNAs have been assessed in clinical studies, especially as biomarkers for diagnosis and prognosis. However, miRNA-based therapies have not yet led to any approvals by regulatory agencies, and their use in BC treatment remains an emerging field [73]. For example, miR-16 mimics (mesomiR-1 by TargomiRs/EnGeneIC), miR-34 mimics (MRX34 by Mirna Therapeutics), and miR-155 antagomir (MRG-106 by miRagen Therapeutics) have entered clinical trials for oncological diseases, but not for BC [73].
In p53-miRNA axis, miR-26a, miR-34a, miR-203, miR-205; miR-214, and miR-223 were described as potential antitumor or antimetastatic miRNA-based therapeutics, respectively, in animal models [73]. Along with presenting more consistent preclinical data, the development of miRNA-based therapies depends on overcoming major hurdles such as molecule stability and targeted delivery. In vivo application of miRNA molecules faces additional challenges, including poor bioavailability, enzymatic degradation, rapid clearance, and inefficient cellular uptake, which also need to be further explored [74].
In addition, although p53 does not possess typical drug target features, several strategies to manipulate p53 in cancer have been developed in recent years, including reactivation of mutant p53, induction of mutant p53 degradation, and exploitation of synthetic lethality in p53-deficient cells. Some of these drugs, such as eprenetapopt (APR-246), COTI-2, arsenic trioxide, nutlin-3a, and PC14586, are currently under evaluation in clinical trials, with the most advanced being eprenetapopt, primarily in myelodysplastic syndrome [75,76]. In BC, these p53-targeting drugs have not yet produced conclusive clinical results. Some strategies have been explored preclinically, particularly in triple-negative BC, such as zoledronic acid and atorvastatin, which may promote degradation of mutant p53 or inhibit pathways enhanced by mutant p53 [75,76]. No p53-based therapy has yet reached a clinically validated stage for BC, and most approaches remain experimental. Once available for BC, these therapies could potentially influence the p53–miRNA axis. Indeed, in a BC study, eprenetapopt was shown to restore the transcriptional activity of mutant p53, which in turn leads to the upregulation of miR-30c [77], a miRNA involved in mediating p53-dependent tumor suppressive effects [40,64].
The regulatory interplay between p53 and miRNAs represents a highly intricate network with critical implications for BC biology. Despite significant advances in our understanding of this axis, several knowledge gaps and methodological limitations remain that hinder its full translational potential.
One of the key challenges lies in the context-dependent behavior of specific miRNAs, which often produce outcomes that diverge from expected canonical effects. For instance, miR-193a-5p [51], miR-3646 [52], and miR-150-5p [53] have been reported to downregulate TP53 expression, yet paradoxically exhibit anti-proliferative and pro-apoptotic effects, along with increased chemosensitivity and improved patient survival. These findings challenge the conventional expectation that downregulation of TP53 necessarily promotes tumor progression. Moreover, miRNAs such as miR-10 [22,40], miR-26a [19,41], miR-30c [40,64], miR-34a [36,56], miR-155 [42,68], and miR-200c [36,72] are not only regulated by p53 but also regulate TP53 expression themselves, forming feedback loops that may shift functional outcomes depending on p53 status (wild-type vs. mutant) [78], epigenetic modulation [79], or external stress signals [80]. These regulatory paradoxes underscore the need for functional studies that integrate transcriptomic, proteomic, and phenotypic data.
In addition to biological complexity, there are technical limitations in current methodologies. The detection and quantification of circulating miRNAs remain inconsistent across studies due to variability in sample types (e.g., serum vs. plasma), normalization protocols, and analytical platforms [81,82]. This lack of standardization hampers reproducibility and delays the clinical validation of miRNAs as biomarkers or therapeutic agents. Moreover, miRNA-based therapies still face substantial delivery challenges, including low stability in circulation, enzymatic degradation, inefficient cellular uptake, and off-target effects [74].
Emerging technologies offer promising avenues to overcome these barriers. Single-cell RNA sequencing and spatial transcriptomics allow for the dissection of miRNA-p53 interactions at the resolution of individual cells and microenvironments, unveiling heterogeneity that is masked in bulk analyses [83]. CRISPR-based functional screens provide powerful tools to interrogate causal relationships between miRNAs, TP53, and downstream pathways in specific BC subtypes [84]. Integrating these approaches with machine learning models may further enable the identification of miRNA signatures predictive of prognosis, therapeutic response, or resistance mechanisms.
TP53 and miRNAs play an important and complex role in the gene regulation network in BC, which still requires further study. This knowledge will be extremely important for identifying new biomarkers for the diagnosis and follow-up of the disease and, above all, will assist in the development of potential targeted therapies for patient treatment, especially for those who do not respond to conventional therapy.
Acknowledgement: Not applicable.
Funding Statement: The authors received no specific funding for this study.
Author Contributions: Conceptualization: Marcia Eduarda Viana Luna, Juliana Carron; Literature search and data analysis: Marcia Eduarda Viana Luna, Juliana Carron; Writing—original draft preparation: Marcia Eduarda Viana Luna, Gustavo Jacob Lourenço, Juliana Carron; Writing—review and editing: Gustavo Jacob Lourenço, Juliana Carron. All authors reviewed the results and approved the final version of the manuscript.
Availability of Data and Materials: Not applicable.
Ethics Approval: Not applicable.
Conflicts of Interest: The authors declare no conflicts of interest to report regarding the present study.
References
1. Bray F, Laversanne M, Sung H, Ferlay J, Siegel RL, Soerjomataram I, et al. Global cancer statistics 2022: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2024;74(3):229–63. doi:10.3322/caac.21834. [Google Scholar] [PubMed] [CrossRef]
2. Hassett MJ, Somerfield MR, Baker ER, Cardoso F, Kansal KJ, Kwait DC, et al. Management of male breast cancer: ASCO guideline. J Clin Oncol. 2020;38(16):1849–63. doi:10.1200/jco.19.03120. [Google Scholar] [PubMed] [CrossRef]
3. Amin MB, Greene FL, Edge SB, Compton CC, Gershenwald JE, Brookland RK, et al. The Eighth Edition AJCC Cancer Staging Manual: continuing to build a bridge from a population-based to a more “personalized” approach to cancer staging. CA Cancer J Clin. 2017;67(2):93–9. doi:10.3322/caac.21388. [Google Scholar] [PubMed] [CrossRef]
4. Weigelt B, Reis-Filho JS. Histological and molecular types of breast cancer: is there a unifying taxonomy? Nat Rev Clin Oncol. 2009;6(12):718–30. doi:10.1038/nrclinonc.2009.166. [Google Scholar] [PubMed] [CrossRef]
5. Barzaman K, Karami J, Zarei Z, Hosseinzadeh A, Kazemi MH, Moradi-Kalbolandi S, et al. Breast cancer: biology, biomarkers, and treatments. Int Immunopharmacol. 2020;84(1):106535. doi:10.1016/j.intimp.2020.106535. [Google Scholar] [PubMed] [CrossRef]
6. Tsang JYS, Tse GM. Molecular classification of breast cancer. Adv Anat Pathol. 2020;27(1):27–35. [Google Scholar] [PubMed]
7. Rodríguez-Bejarano OH, Parra-López C, Patarroyo MA. A review concerning the breast cancer-related tumour microenvironment. Crit Rev Oncol Hematol. 2024;199:104389. doi:10.1016/j.critrevonc.2024.104389. [Google Scholar] [PubMed] [CrossRef]
8. Trayes KP, Cokenakes SEH. Breast Cancer Treatment. Am Fam Physician. 2021;104(2):171–8. [Google Scholar] [PubMed]
9. Zhai J, Wu Y, Ma F, Kaklamani V, Xu B. Advances in medical treatment of breast cancer in 2022. Cancer Innov. 2023;2(1):1–17. doi:10.1002/cai2.46. [Google Scholar] [PubMed] [CrossRef]
10. Soares LR, Freitas-Junior R, Curado MP, Paulinelli RR, Martins E, Oliveira JC. Low overall survival in women with de novo metastatic breast cancer: does this reflect tumor biology or a lack of access to health care? JCO Glob Oncol. 2020;6:679–87. doi:10.1200/jgo.19.00408. [Google Scholar] [PubMed] [CrossRef]
11. Britt KL, Cuzick J, Phillips KA. Key steps for effective breast cancer prevention. Nat Rev Cancer. 2020;20(8):417–36. doi:10.1038/s41568-020-0266-x. [Google Scholar] [PubMed] [CrossRef]
12. Kim I, Choi S, Kim S. BRCA-Pathway: a structural integration and visualization system of TCGA breast cancer data on KEGG pathways. BMC Bioinformatics. 2018;19(Suppl 1):42. doi:10.1186/s12859-018-2016-6. [Google Scholar] [PubMed] [CrossRef]
13. Terry MB, McDonald JA, Wu HC, Eng S, Santella RM. Epigenetic biomarkers of breast cancer risk: across the breast cancer prevention continuum. Adv Exp Med Biol. 2016;882(1):33–68. doi:10.1007/978-3-319-22909-6_2. [Google Scholar] [PubMed] [CrossRef]
14. Cai Y, Li J, Gao Y, Yang K, He J, Li N, et al. A systematic review of recommendations on screening strategies for breast cancer due to hereditary predisposition: who, when, and how? Cancer Med. 2021;10(10):3437–48. doi:10.1002/cam4.3898. [Google Scholar] [PubMed] [CrossRef]
15. Sargolzaei J, Etemadi T, Alyasin A. The P53/microRNA network: a potential tumor suppressor with a role in anticancer therapy. Pharmacol Res. 2020;160(3):105179. doi:10.1016/j.phrs.2020.105179. [Google Scholar] [PubMed] [CrossRef]
16. Blagih J, Buck MD, Vousden KH. p53, cancer and the immune response. J Cell Sci. 2020;133(5):jcs237453. doi:10.1242/jcs.237453. [Google Scholar] [PubMed] [CrossRef]
17. Bieging KT, Mello SS, Attardi LD. Unravelling mechanisms of p53-mediated tumour suppression. Nat Rev Cancer. 2014;14(5):359–70. doi:10.1038/nrc3711. [Google Scholar] [PubMed] [CrossRef]
18. Nag S, Qin J, Srivenugopal KS, Wang M, Zhang R. The MDM2-p53 pathway revisited. J Biomed Res. 2013;27(4):254–71. doi:10.7555/jbr.27.20130030. [Google Scholar] [PubMed] [CrossRef]
19. Stein Y, Rotter V, Aloni-Grinstein R. Gain-of-function mutant p53: all the roads lead to tumorigenesis. Int J Mol Sci. 2019;20(24). doi:10.3390/ijms20246197. [Google Scholar] [PubMed] [CrossRef]
20. Brosh R, Rotter V. When mutants gain new powers: news from the mutant p53 field. Nat Rev Cancer. 2009;9(10):701–13. doi:10.1038/nrc2693. [Google Scholar] [PubMed] [CrossRef]
21. Madrigal T, Hernández-Monge J, Herrera LA, González-De la Rosa CH, Domínguez-Gómez G, Candelaria M, et al. Regulation of miRNAs expression by mutant p53 gain of function in cancer. Front Cell Dev Biol. 2021;9:695723. doi:10.3389/fcell.2021.695723. [Google Scholar] [PubMed] [CrossRef]
22. Zhou E, Hui NA, Shu M, Wu B, Zhou J. Systematic analysis of the p53-related microRNAs in breast cancer revealing their essential roles in the cell cycle. Oncol Lett. 2015;10(6):3488–94. doi:10.3892/ol.2015.3751. [Google Scholar] [PubMed] [CrossRef]
23. Yang L, Zhao W, Wei P, Zuo W, Zhu S. Tumor suppressor p53 induces miR-15a processing to inhibit neuronal apoptosis inhibitory protein (NAIP) in the apoptotic response DNA damage in breast cancer cell. Am J Transl Res. 2017;9(2):683–91. [Google Scholar] [PubMed]
24. Zhang J, Su B, Gong C, Xi Q, Chao T. miR-214 promotes apoptosis and sensitizes breast cancer cells to doxorubicin by targeting the RFWD2-p53 cascade. Biochem Biophys Res Commun. 2016;478(1):337–42. doi:10.1016/j.bbrc.2016.07.054. [Google Scholar] [PubMed] [CrossRef]
25. Bartel DP. MicroRNAs: target recognition and regulatory functions. Cell. 2009;136(2):215–33. doi:10.1016/j.cell.2009.01.002. [Google Scholar] [PubMed] [CrossRef]
26. Chen H, Xie G, Luo Q, Yang Y, Hu S. Regulatory miRNAs, circRNAs and lncRNAs in cell cycle progression of breast cancer. Funct Integr Genomics. 2023;23(3):233. doi:10.1007/s10142-023-01130-z. [Google Scholar] [PubMed] [CrossRef]
27. Doloi R, Gupta SM. MicroRNAs: the key players regulating the crosstalk between Hippo and Wnt/β-catenin pathways in breast cancer. Life Sci. 2023;329(3):121980. doi:10.1016/j.lfs.2023.121980. [Google Scholar] [PubMed] [CrossRef]
28. Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004;116(2):281–97. [Google Scholar] [PubMed]
29. Moran Y, Agron M, Praher D, Technau U. The evolutionary origin of plant and animal microRNAs. Nat Ecol Evol. 2017;1(3):27. doi:10.1038/s41559-016-0027. [Google Scholar] [PubMed] [CrossRef]
30. O’Brien J, Hayder H, Zayed Y, Peng C. Overview of MicroRNA biogenesis, mechanisms of actions, and circulation. Front Endocrinol. 2018;9:402. doi:10.3389/fendo.2018.00402. [Google Scholar] [PubMed] [CrossRef]
31. Sharma S, Patnaik PK, Aronov S, Kulshreshtha R. ApoptomiRs of breast cancer: basics to clinics. Front Genet. 2016;7:175. doi:10.3389/fgene.2016.00175. [Google Scholar] [PubMed] [CrossRef]
32. Wang F, Lv P, Liu X, Zhu M, Qiu X. microRNA-214 enhances the invasion ability of breast cancer cells by targeting p53. Int J Mol Med. 2015;35(5):1395–402. doi:10.3892/ijmm.2015.2123. [Google Scholar] [PubMed] [CrossRef]
33. Hu W, Chan CS, Wu R, Zhang C, Sun Y, Song JS, et al. Negative regulation of tumor suppressor p53 by microRNA miR-504. Mol Cell. 2010;38(5):689–99. doi:10.1016/j.molcel.2010.05.027. [Google Scholar] [PubMed] [CrossRef]
34. Cho JG, Park S, Lim CH, Kim HS, Song SY, Roh TY, et al. ZNF224, Krüppel like zinc finger protein, induces cell growth and apoptosis-resistance by down-regulation of p21 and p53 via miR-663a. Oncotarget. 2016;7(21):31177–90. doi:10.18632/oncotarget.8870. [Google Scholar] [PubMed] [CrossRef]
35. Neshastehriz A, Hormozi-Moghaddam Z, Kichi ZA, Taheri SM, Amini SM, Aghaei A. Overcoming breast cancer cell treatment resistance by optimizing sonodynamic therapy and radiation sensitizers on lncRNA PVT1 and miR-1204 expression. Photodiagnosis Photodyn Ther. 2025;51(1):104433. doi:10.1016/j.pdpdt.2024.104433. [Google Scholar] [PubMed] [CrossRef]
36. Verma VK, Beevi SS, Nair RA, Kumar A, Kiran R, Alexander LE, et al. MicroRNA signatures differentiate types, grades, and stages of breast invasive ductal carcinoma (IDCmiRNA-target interacting signaling pathways. Cell Commun Signal. 2024;22(1):100. doi:10.1186/s12964-023-01452-2. [Google Scholar] [PubMed] [CrossRef]
37. Li X, Xie W, Xie C, Huang C, Zhu J, Liang Z, et al. Curcumin modulates miR-19/PTEN/AKT/p53 axis to suppress bisphenol A-induced MCF-7 breast cancer cell proliferation. Phytother Res. 2014;28(10):1553–60. doi:10.1002/ptr.5167. [Google Scholar] [PubMed] [CrossRef]
38. You F, Luan H, Sun D, Cui T, Ding P, Tang H. miRNA-106a promotes breast cancer cell proliferation, clonogenicity, migration, and invasion through inhibiting apoptosis and chemosensitivity. DNA Cell Biol. 2019;38(2):198–207. doi:10.1089/dna.2018.4282. [Google Scholar] [PubMed] [CrossRef]
39. Gao Y, Ma H, Gao C, Lv Y, Chen X, Xu R, et al. Tumor-promoting properties of miR-8084 in breast cancer through enhancing proliferation, suppressing apoptosis and inducing epithelial-mesenchymal transition. J Transl Med. 2018;16(1):38. doi:10.1186/s12967-018-1419-5. [Google Scholar] [PubMed] [CrossRef]
40. Rahimi M, Sharifi-Zarchi A, Firouzi J, Azimi M, Zarghami N, Alizadeh E, et al. An integrated analysis to predict micro-RNAs targeting both stemness and metastasis in breast cancer stem cells. J Cell Mol Med. 2019;23(4):2442–56. doi:10.1111/jcmm.14090. [Google Scholar] [PubMed] [CrossRef]
41. Cabello P, Pineda B, Tormo E, Lluch A, Eroles P. The antitumor effect of metformin is mediated by miR-26a in breast cancer. Int J Mol Sci. 2016;17(8). doi:10.3390/ijms17081298. [Google Scholar] [PubMed] [CrossRef]
42. Liu J, Mao Q, Liu Y, Hao X, Zhang S, Zhang J. Analysis of miR-205 and miR-155 expression in the blood of breast cancer patients. Chin J Cancer Res. 2013;25(1):46–54. [Google Scholar] [PubMed]
43. Jansson MD, Damas ND, Lees M, Jacobsen A, Lund AH. miR-339-5p regulates the p53 tumor-suppressor pathway by targeting MDM2. Oncogene. 2015;34(15):1908–18. doi:10.1038/onc.2014.130. [Google Scholar] [PubMed] [CrossRef]
44. Hoffman Y, Bublik DR, Pilpel Y, Oren M. miR-661 downregulates both Mdm2 and Mdm4 to activate p53. Cell Death Differ. 2014;21(2):302–9. doi:10.1038/cdd.2013.146. [Google Scholar] [PubMed] [CrossRef]
45. Zhang C, Liu J, Tan C, Yue X, Zhao Y, Peng J, et al. microRNA-1827 represses MDM2 to positively regulate tumor suppressor p53 and suppress tumorigenesis. Oncotarget. 2016;7(8):8783–96. doi:10.18632/oncotarget.7088. [Google Scholar] [PubMed] [CrossRef]
46. Wang Q, Selth LA, Callen DF. MiR-766 induces p53 accumulation and G2/M arrest by directly targeting MDM4. Oncotarget. 2017;8(18):29914–24. doi:10.18632/oncotarget.15530. [Google Scholar] [PubMed] [CrossRef]
47. Raza U, Saatci Ö, Uhlmann S, Ansari SA, Eyüpoğlu E, Yurdusev E, et al. The miR-644a/CTBP1/p53 axis suppresses drug resistance by simultaneous inhibition of cell survival and epithelial-mesenchymal transition in breast cancer. Oncotarget. 2016;7(31):49859–77. doi:10.18632/oncotarget.10489. [Google Scholar] [PubMed] [CrossRef]
48. Feng R, Dong L. Inhibitory effect of miR-184 on the potential of proliferation and invasion in human glioma and breast cancer cells in vitro. Int J Clin Exp Pathol. 2015;8(8):9376–82. [Google Scholar] [PubMed]
49. Xie Q, Wang S, Zhao Y, Zhang Z, Qin C, Yang X. MicroRNA-216a suppresses the proliferation and migration of human breast cancer cells via the Wnt/β-catenin signaling pathway. Oncol Rep. 2019;41(5):2647–56. doi:10.21203/rs.3.rs-823596/v1. [Google Scholar] [CrossRef]
50. Dastmalchi N, Safaralizadeh R, Hosseinpourfeizi MA, Baradaran B, Khojasteh SMB. MicroRNA-424-5p enhances chemosensitivity of breast cancer cells to Taxol and regulates cell cycle, apoptosis, and proliferation. Mol Biol Rep. 2021;48(2):1345–57. doi:10.1007/s11033-021-06193-4. [Google Scholar] [PubMed] [CrossRef]
51. Khordadmehr M, Shahbazi R, Baradaran B, Sadreddini S, Shanebandi D, Hajiasgharzadeh K. Restoring of miR-193a-5p sensitizes breast cancer cells to paclitaxel through P53 pathway. Adv Pharm Bull. 2020;10(4):595–601. doi:10.34172/apb.2020.071. [Google Scholar] [PubMed] [CrossRef]
52. Tao S, Liu YB, Zhou ZW, Lian B, Li H, Li JP, et al. miR-3646 promotes cell proliferation, migration, and invasion via regulating G2/M transition in human breast cancer cells. Am J Transl Res. 2016;8(4):1659–77. doi:10.1186/s12935-022-02657-0. [Google Scholar] [PubMed] [CrossRef]
53. Almohaywi M, Sugita BM, Centa A, Fonseca AS, Antunes VC, Fadda P, et al. Deregulated miRNA expression in triple-negative breast cancer of ancestral genomic-characterized latina patients. Int J Mol Sci. 2023;24(17). doi:10.3390/ijms241713046. [Google Scholar] [PubMed] [CrossRef]
54. Hermeking H. MicroRNAs in the p53 network: micromanagement of tumour suppression. Nat Rev Cancer. 2012;12(9):613–26. doi:10.1038/nrc3318. [Google Scholar] [PubMed] [CrossRef]
55. di Gennaro A, Damiano V, Brisotto G, Armellin M, Perin T, Zucchetto A, et al. A p53/miR-30a/ZEB2 axis controls triple negative breast cancer aggressiveness. Cell Death Differ. 2018;25(12):2165–80. doi:10.1038/s41418-018-0103-x. [Google Scholar] [PubMed] [CrossRef]
56. Avtanski DB, Nagalingam A, Tomaszewski JE, Risbood P, Difillippantonio MJ, Saxena NK, et al. Indolo-pyrido-isoquinolin based alkaloid inhibits growth, invasion and migration of breast cancer cells via activation of p53-miR34a axis. Mol Oncol. 2016;10(7):1118–32. doi:10.1158/1538-7445.am2016-1228. [Google Scholar] [CrossRef]
57. Peurala H, Greco D, Heikkinen T, Kaur S, Bartkova J, Jamshidi M, et al. MiR-34a expression has an effect for lower risk of metastasis and associates with expression patterns predicting clinical outcome in breast cancer. PLoS One. 2011;6(11):e26122. doi:10.1371/journal.pone.0026122. [Google Scholar] [PubMed] [CrossRef]
58. Hargraves KG, He L, Firestone GL. Phytochemical regulation of the tumor suppressive microRNA, miR-34a, by p53-dependent and independent responses in human breast cancer cells. Mol Carcinog. 2016;55(5):486–98. doi:10.1002/mc.22296. [Google Scholar] [CrossRef]
59. Parfenyev S, Singh A, Fedorova O, Daks A, Kulshreshtha R, Barlev NA. Interplay between p53 and non-coding RNAs in the regulation of EMT in breast cancer. Cell Death Dis. 2021;12(1):17. doi:10.1038/s41419-020-03327-7. [Google Scholar] [CrossRef]
60. Chang CJ, Chao CH, Xia W, Yang JY, Xiong Y, Li CW, et al. p53 regulates epithelial-mesenchymal transition and stem cell properties through modulating miRNAs. Nat Cell Biol. 2011;13(3):317–23. doi:10.1038/ncb2173. [Google Scholar] [PubMed] [CrossRef]
61. Tamura M, Sasaki Y, Kobashi K, Takeda K, Nakagaki T, Idogawa M, et al. CRKL oncogene is downregulated by p53 through miR-200s. Cancer Sci. 2015;106(8):1033–40. doi:10.1111/cas.12713. [Google Scholar] [PubMed] [CrossRef]
62. Kumar KJS, Vani MG, Hsieh HW, Lin CC, Wang SY. Antcin-A modulates epithelial-to-mesenchymal transition and inhibits migratory and invasive potentials of human breast cancer cells via p53-mediated miR-200c activation. Planta Med. 2019;85(9–10):755–65. doi:10.1055/a-0942-2087. [Google Scholar] [PubMed] [CrossRef]
63. Piovan C, Palmieri D, Di Leva G, Braccioli L, Casalini P, Nuovo G, et al. Oncosuppressive role of p53-induced miR-205 in triple negative breast cancer. Mol Oncol. 2012;6(4):458–72. doi:10.1016/j.molonc.2012.03.003. [Google Scholar] [PubMed] [CrossRef]
64. Lin S, Yu L, Song X, Bi J, Jiang L, Wang Y, et al. Intrinsic adriamycin resistance in p53-mutated breast cancer is related to the miR-30c/FANCF/REV1-mediated DNA damage response. Cell Death Dis. 2019;10(9):666. doi:10.1038/s41419-022-04543-z. [Google Scholar] [PubMed] [CrossRef]
65. Lee YM, Lee JY, Ho CC, Hong QS, Yu SL, Tzeng CR, et al. miRNA-34b as a tumor suppressor in estrogen-dependent growth of breast cancer cells. Breast Cancer Res. 2011;13(6):R116. doi:10.1186/bcr3059. [Google Scholar] [PubMed] [CrossRef]
66. Guo F, Chen H, Chang J, Zhang L. Mutation R273H confers p53 a stimulating effect on the IGF-1R-AKT pathway via miR-30a suppression in breast cancer. Biomed Pharmacother. 2016;78:335–41. doi:10.1016/j.biopha.2016.01.031. [Google Scholar] [PubMed] [CrossRef]
67. Sandhu R, Rein J, D’Arcy M, Herschkowitz JI, Hoadley KA, Troester MA. Overexpression of miR-146a in basal-like breast cancer cells confers enhanced tumorigenic potential in association with altered p53 status. Carcinogenesis. 2014;35(11):2567–75. doi:10.1093/carcin/bgu175. [Google Scholar] [PubMed] [CrossRef]
68. Neilsen PM, Noll JE, Mattiske S, Bracken CP, Gregory PA, Schulz RB, et al. Mutant p53 drives invasion in breast tumors through up-regulation of miR-155. Oncogene. 2013;32(24):2992–3000. doi:10.1038/onc.2012.305. [Google Scholar] [PubMed] [CrossRef]
69. Alam F, Mezhal F, El Hasasna H, Nair VA, Aravind SR, Saber Ayad M, et al. The role of p53-microRNA 200-Moesin axis in invasion and drug resistance of breast cancer cells. Tumour Biol. 2017;39(9):1010428317714634. doi:10.1177/1010428317714634. [Google Scholar] [PubMed] [CrossRef]
70. Masciarelli S, Fontemaggi G, Di Agostino S, Donzelli S, Carcarino E, Strano S, et al. Gain-of-function mutant p53 downregulates miR-223 contributing to chemoresistance of cultured tumor cells. Oncogene. 2014;33(12):1601–8. doi:10.1038/onc.2013.106. [Google Scholar] [PubMed] [CrossRef]
71. Zhang Y, Hu Y, Fang JY, Xu J. Gain-of-function miRNA signature by mutant p53 associates with poor cancer outcome. Oncotarget. 2016;7(10):11056–66. doi:10.18632/oncotarget.7090. [Google Scholar] [PubMed] [CrossRef]
72. Chao CH, Wang CY, Wang CH, Chen TW, Hsu HY, Huang HW, et al. Mutant p53 attenuates oxidative phosphorylation and facilitates cancer stemness through downregulating miR-200c-PCK2 axis in basal-like breast cancer. Mol Cancer Res. 2021;19(11):1900–16. doi:10.1158/1541-7786.mcr-21-0098. [Google Scholar] [PubMed] [CrossRef]
73. Grimaldi AM, Salvatore M, Incoronato M. miRNA-based therapeutics in breast cancer: a systematic review. Front Oncol. 2021;11:668464. doi:10.3389/fonc.2021.668464. [Google Scholar] [PubMed] [CrossRef]
74. Takahashi RU, Prieto-Vila M, Kohama I, Ochiya T. Development of miRNA-based therapeutic approaches for cancer patients. Cancer Sci. 2019;110(4):1140–7. doi:10.1111/cas.13965. [Google Scholar] [PubMed] [CrossRef]
75. Nishikawa S, Iwakuma T. Drugs targeting p53 Mutations with FDA approval and in clinical trials. Cancers. 2023;15(2):429. doi:10.3390/cancers15020429. [Google Scholar] [PubMed] [CrossRef]
76. Duffy MJ, Tang M, Rajaram S, O’Grady S, Crown J. Targeting mutant p53 for cancer treatment: moving closer to clinical use? Cancers. 2022;14(18):4499. doi:10.3390/cancers14184499. [Google Scholar] [PubMed] [CrossRef]
77. Walerych D, Lisek K, Sommaggio R, Piazza S, Ciani Y, Dalla E, et al. Proteasome machinery is instrumental in a common gain-of-function program of the p53 missense mutants in cancer. Nat Cell Biol. 2016;18(8):897–909. doi:10.1038/ncb3380. [Google Scholar] [PubMed] [CrossRef]
78. Jones M, Lal A. MicroRNAs, wild-type and mutant p53: more questions than answers. RNA Biol. 2012;9(6):781–91. doi:10.4161/rna.20146. [Google Scholar] [PubMed] [CrossRef]
79. Baer C, Claus R, Plass C. Genome-wide epigenetic regulation of miRNAs in cancer. Cancer Res. 2013;73(2):473–7. doi:10.1158/0008-5472.can-12-3731. [Google Scholar] [PubMed] [CrossRef]
80. Capaccia C, Diverio S, Zampini D, Guelfi G. The complex interaction between P53 and miRNAs joins new awareness in physiological stress responses. Cells. 2022;11(10). doi:10.3390/cells11101631. [Google Scholar] [PubMed] [CrossRef]
81. Kim DJ, Linnstaedt S, Palma J, Park JC, Ntrivalas E, Kwak-Kim JY, et al. Plasma components affect accuracy of circulating cancer-related microRNA quantitation. J Mol Diagn. 2012;14(1):71–80. doi:10.1016/j.jmoldx.2011.09.002. [Google Scholar] [PubMed] [CrossRef]
82. McDermott AM, Kerin MJ, Miller N. Identification and validation of miRNAs as endogenous controls for RQ-PCR in blood specimens for breast cancer studies. PLoS One. 2013;8(12):e83718. doi:10.1371/journal.pone.0083718. [Google Scholar] [PubMed] [CrossRef]
83. Moore R, Ooi HK, Kang T, Bleris L, Ma L. MiR-192-mediated positive feedback loop controls the robustness of stress-induced p53 oscillations in breast cancer cells. PLoS Comput Biol. 2015;11(12):e1004653. doi:10.1371/journal.pcbi.1004653. [Google Scholar] [PubMed] [CrossRef]
84. Samuels M, Besta S, Betrán AL, Nia RS, Xie X, Gu X, et al. CRISPR screening approaches in breast cancer research. Cancer Metastasis Rev. 2025;44(3):59. doi:10.1007/s10555-025-10275-1. [Google Scholar] [PubMed] [CrossRef]
Cite This Article
 Copyright © 2026 The Author(s). Published by Tech Science Press.
Copyright © 2026 The Author(s). Published by Tech Science Press.This work is licensed under a Creative Commons Attribution 4.0 International License , which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.


 Submit a Paper
Submit a Paper Propose a Special lssue
Propose a Special lssue View Full Text
View Full Text Download PDF
Download PDF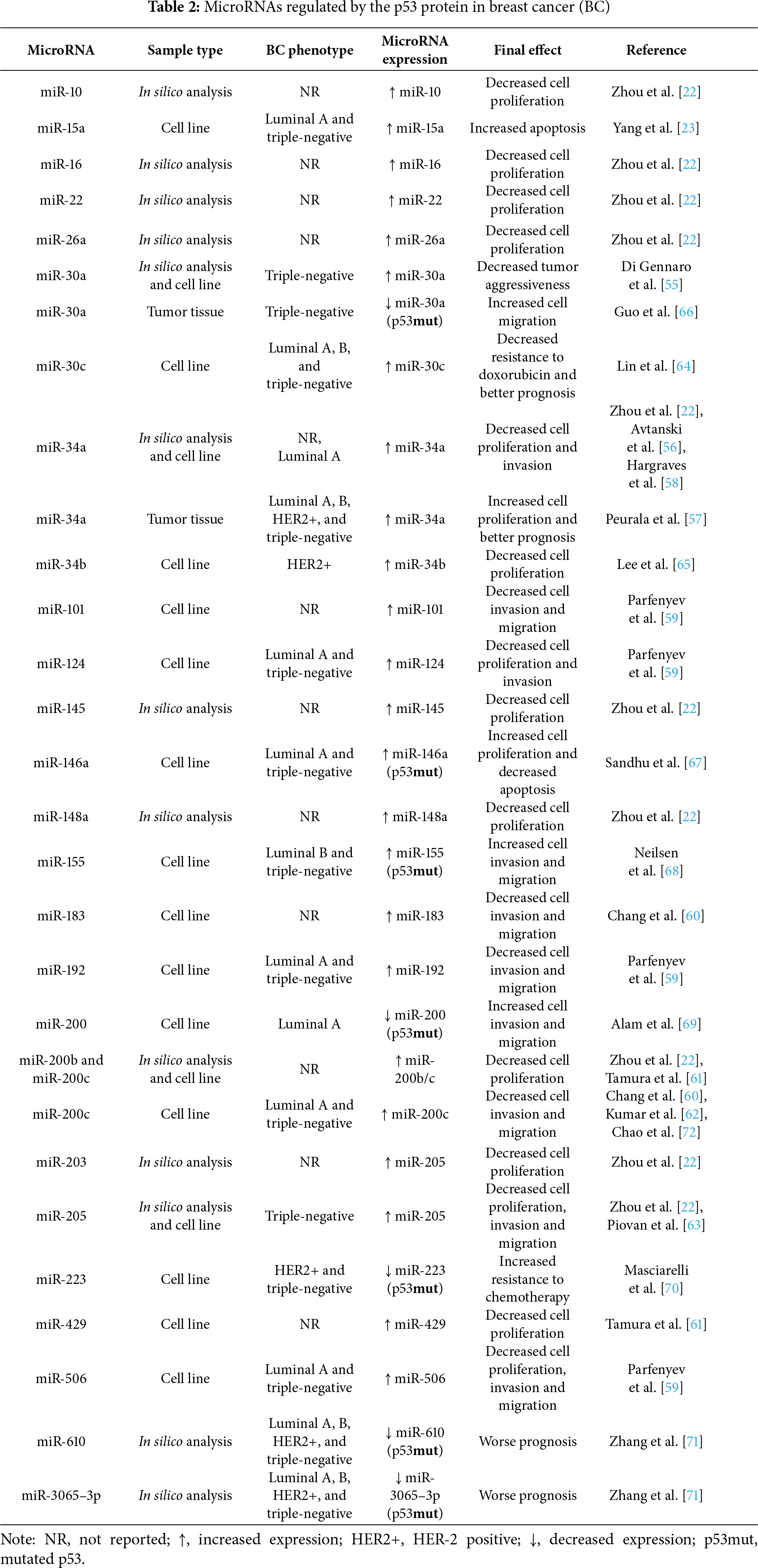
 Downloads
Downloads
 Citation Tools
Citation Tools