 Open Access
Open Access
REVIEW
Cancer-associated fibroblasts of colorectal cancer: Translational prospects in liquid biopsy and targeted therapy
1 Human Genome Centre, School of Medical Sciences, Universiti Sains Malaysia, Kubang Kerian, 16150, Malaysia
2 Department of Internal Medicine, School of Medical Sciences, Universiti Sains Malaysia, Kubang Kerian, 16150, Malaysia
3 Department of Surgery, School of Medical Sciences, Universiti Sains Malaysia, Kubang Kerian, 16150, Malaysia
4 Department of Pathology, School of Medical Sciences, Universiti Sains Malaysia, Kubang Kerian, 16150, Malaysia
* Corresponding Author: MARAHAINI MUSA. Email:
(This article belongs to the Special Issue: Frontiers in cancer: tumor microenvironment)
BIOCELL 2023, 47(10), 2233-2244. https://doi.org/10.32604/biocell.2023.030541
Received 12 April 2023; Accepted 01 June 2023; Issue published 08 November 2023
Abstract
Colorectal cancer (CRC) is a major global health concern. Accumulation of cancer-associated fibroblasts (CAFs) in CRC is associated with poor prognosis and disease recurrence. CAFs are the main cellular component of the tumor microenvironment. CAF-tumor cell interplay, which is facilitated by various secretomes, drives colorectal carcinogenesis. The complexity of CAF populations contributes to the heterogeneity of CRC and influences patient survival and treatment response. Due to their significant roles in colorectal carcinogenesis, different clinical applications utilizing or targeting CAFs have been suggested. Circulating CAFs (cCAFs) which can be detected in blood samples, have been proposed to help in determining patient prognosis and enables the detection of cancer through liquid biopsy. Liquid biopsy is gaining traction as it is non-invasive, allows frequent and easy sampling, and shows concordance to tissue biopsy analysis. In addition, CAF-targeted therapy is currently being studied extensively to be used as one of the treatment avenues for CRC. Various mechanisms of CAF-targeted therapy have been reported, including blocking the signaling pathways involving CAFs and cancer cells, thus abolishing the CAF-tumor cell crosstalk and subsequently hindering tumorigenesis. These translational applications of cCAFs and utilization of CAFs as key targets for CRC therapy, although still in the early phases of development, will potentially improve CRC patient management in the future.Graphic Abstract
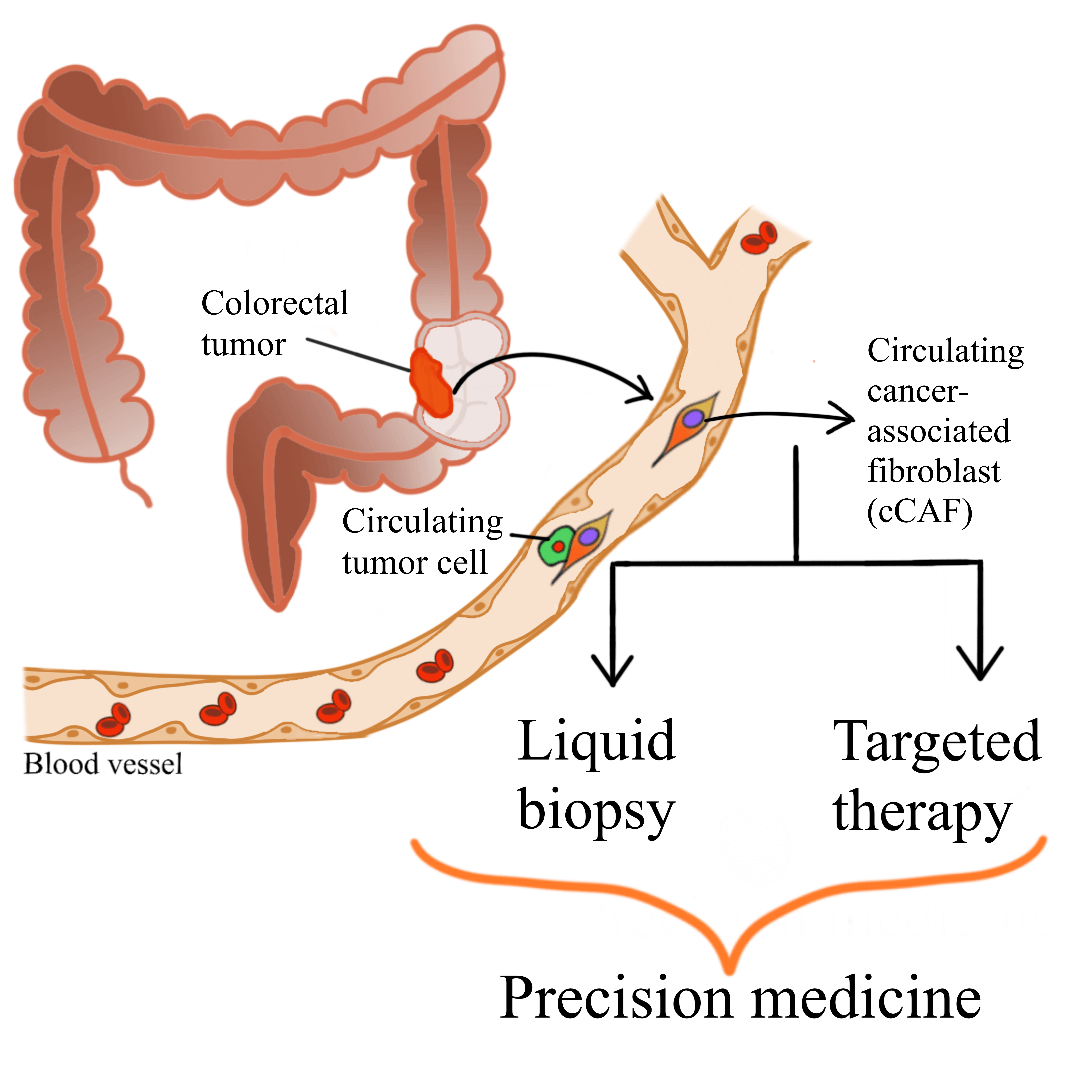
Keywords
Colorectal cancer (CRC) is one of the deadliest and commonly diagnosed malignancies, affecting men and women worldwide. It is predicted that the global new CRC cases will increase to 3.2 million in 2040. CRC incidence is higher in highly developed countries, although the cases are increasing steadily in middle- and low-income countries (Xi and Xu, 2021). In addition, there is an alarming global trend of early-onset CRC reported in individuals below 50 years of age (Akimoto et al., 2021). It is estimated that roughly 60% of CRC patients perished from their cancer (Riihimäki et al., 2012). Despite this, CRC mortality is steadily decreasing in developed countries, mainly due to efficient national screening initiatives, adaptation to a healthier lifestyle and diet, and significant uptake of colonoscopy, which is a gold standard procedure to screen for CRC (Ait Ouakrim et al., 2015).
CRC develops as the result of the accumulation of genetic and epigenetic changes involving oncogenes and tumor suppressor genes. The multistep process of classical adenoma-carcinoma transition in colorectal carcinogenesis occurs over a period of approximately 10–15 years (Fearon and Vogelstein, 1990; Binefa et al., 2014). CRC can spread systemically and metastasize to distant sites, most commonly to the liver and peritoneum (Riihimäki et al., 2016). Various risk factors ranging from genetics to environment, contributes to the onset and progression of CRC. The majority of CRC cases arise sporadically, without a family history of the disease or inherited genomic changes (Keum and Giovannucci, 2019; Tian et al., 2019). Despite extensive studies on CRC pathogenesis and the discovery of advanced cancer treatment, the survival rates of CRC patients are still poor. Multiple factors, such as late detection and low compliance in CRC screening, lead to high morbidity and mortality due to CRC.
CRC is a heterogeneous disease, and this influences patient prognosis and treatment efficacy (Xi and Xu, 2021). Genomic profiling on CRC has revealed the complex nature of this disease (Hinoi, 2021), which likely contributes to the patient’s lack of response to therapy.
A major factor that promotes CRC evolution and heterogeneity is the tumor microenvironment (TME). The accumulation of cancer-associated fibroblasts (CAFs), which are the main stromal cells of the TME, is associated with a worse prognosis and recurrence of CRC (Anderson and Simon, 2020; Musa and Ali, 2020).
Rather than focusing on cancer cells alone, scientists are shifting their focus to TME components, including CAFs. The CAF-cancer cell crosstalk has been reported widely as a key driver of CRC progression and metastasis (Bussard et al., 2016). Considering their vital role in colorectal carcinogenesis, CAFs can be potentially utilized in clinical setting, as a basis for liquid biopsy and targeted therapy. In this review, we discuss the biology of CAFs in CRC and unravel the translational prospects of CAFs, which could overcome present limitations in CRC management and thus may greatly improve the prognosis of CRC patients in the future.
The biology of colorectal cancer and tumor microenvironment
CRC is a heterogeneous entity and constantly evolving, which promotes its complex nature. Phenotypic plasticity exists in cancers, including CRC. Intra-tumor heterogeneity is a result of genetic and epigenetic variations as well as transcriptional plasticity (Black and McGranahan, 2021). The roles of these biological processes in tumor evolution remain unknown and is also yet to be fully explored in colorectal carcinogenesis. Dissecting this process will provide insight into CRC progression and the roles of the TME in promoting tumorigenesis (Ahmad Zawawi and Musa, 2022).
Over the past two decades, scientists have focused on the different components of the tumor, namely the TME, rather than studying the epithelial cancer cells alone. Previous reports have shown that the neoplastic cells can be reverted to a normal proliferation state despite having a highly mutated genome by blocking signaling pathways activated by the TME (Bizzarri et al., 2011; Northey et al., 2017). This highlights the essential role of the TME in dictating the fate of a tumor.
Rather than a silent bystander, the TME is considered an active driver of cancer progression. The TME is highly complex, heterogeneous, and continuously evolving. Its components vary between different tumors. Hallmark features of the TME include stromal, immune, and endothelial cells and extracellular matrix (ECM). The stromal cells have pro- and antitumorigenic properties, where cancer progression can be delayed or blocked by normal stroma, whereas aberrant stroma promotes carcinogenesis (Bissell and Radisky, 2001). However, evidence has suggested that stromal cells produce various factors that promote angiogenesis, tumor growth, invasion, and metastasis through reciprocal communication with cancer cells (Bussard et al., 2016; Anderson and Simon, 2020). Particularly, genetic mutation of cells can promote tumorigenesis, but nonmutant cells in the TME, such as CAFs, significantly influence the progression of cancer and treatment sensitivity (Bochet et al., 2013; Sahai et al., 2020).
Cancer-associated fibroblasts in the tumor microenvironment
CAFs are the most abundant cellular components of the TME. These cells are also known as activated fibroblasts or myofibroblasts in the tumor stroma and are reported to promote tumor development. The origin of CAFs is heavily debatable. Various cell types, including fibroblasts, endothelial cells, vascular mural cells, adipocytes pericytes, and mesenchymal stem cells, may give rise to CAFs (Kalluri, 2016; Nair et al., 2017; Öhlund et al., 2017; LeBleu and Kalluri, 2018; Miyazaki et al., 2020).
A build-up of stromal cells and activated fibroblasts in CRC is associated with poor prognosis and disease recurrence (Anderson and Simon, 2020; Musa and Ali, 2020). This corroborates previous report stating that the mesenchymal or stromal CRC subtype (consensus molecular subtype 4-CMS4) are associated with the worst survival outcomes compared to other CMSs (Guinney et al., 2015; Varga et al., 2020; Kasashima et al., 2021).
Interestingly, CAF-derived gene signatures also demonstrated an association with poor survival in CRC patients with advanced stage of the disease than in earlier stages (Herrera et al., 2021). Additionally, colorectal CAF activation is linked to metastasis, treatment outcome, and patient prognosis, solidifying their vital roles in CRC carcinogenesis (Calon et al., 2012; Tauriello et al., 2018; Khare et al., 2021).
Dienstmann et al. (2019) reported that the patterns of TME infiltration indicate the risk for distant dissemination in the early stages of CRC. Subsequent studies need to be conducted to examine the role of CMS groups and CAF and immune cell infiltration patterns in predicting survival over TNM staging and microsatellite status (MSI) status.
There is increasing evidence on the effects of CAFs on the immune response towards CRC. CAFs in the TME influence the prognosis of CRC patients by inhibiting immune response, subsequently driving tumor progression, and contributing to treatment failure (Chen et al., 2021; Maia et al., 2023). CAFs secrete a variety of cytokines and chemokines, including interleukin-6 (IL-6) and CC chemokine ligand 2 (CCL2), which promote tumor invasion and metastasis and lead to immunosuppressive TME (Kalluri, 2016). Additionally, CAFs also express checkpoint ligands such as PD-L1 and PD-L2, which inhibit T-cell activation and hinder autoimmunity (Latchman et al., 2001; Gorchs et al., 2019). In addition, in the CRC mice model, CXCL5 produced by CAFs was found to regulate the PD-L1 in tumor via PI3K/AKT signaling (Li et al., 2019). There are a number of clinical trials in different tumors involving the combination of immunotherapies with CAF-targeting for cancer treatment. Despite encouraging data from pre-clinical models, the effect of this mode of therapy is yet to be confirmed in human subjects (Maia et al., 2023).
Tumor cell-CAF crosstalk occurs through paracrine and autocrine signals (Fiori et al., 2019). This involves different molecular pathways such as transforming growth factor-beta (TGF-β) signaling and C-X-C motif chemokine 12- chemokine receptors C-X-C chemokine receptor type 4 (CXCL12-CXCR4) axis (Li et al., 2013; Khare et al., 2021).
CAFs are identified using various biomarkers. Depending on cancer type and staging, these CAF markers exhibit differential expression patterns, which corroborates the idea that different CAF subpopulations exist in the cancer tissue. The presence of heterogeneous populations of fibroblasts in a tumor can be partly attributed to the variations in the activation process of resident fibroblasts with organ-specific features (Miyashita and Saito, 2021).
Classical biomarkers of CAFs include the alpha-smooth muscle actin (α-SMA), fibroblast specific protein 1 (FSP-1), vimentin and fibroblast activation protein (FAP) (Herrera et al., 2013; Togo et al., 2013). Additionally, there are several CAF biomarkers with prognostic value and functional relevance, such as caveolin-1 (CAV1) and CD90 (Thy-1) (Huynh et al., 2016). CAV1 expression in the stroma is associated with poor survival (Goetz et al., 2011). A recent report has shown the association of multiple CAF markers, namely α-SMA, collagen I, and platelet-derived growth factor receptor-beta (PDGFR-β), which are widely distributed in the stroma, with high venous invasion in advanced CRC (Nishishita et al., 2018). This supports a previous observation of the correlation of α-SMA positive and desmin negative myofibroblasts in advanced CRC with malignancy potential (Takatsuna et al., 2016). On the contrary, the expression of podoplanin in CAFs indicates a favorable prognosis of CRC (Choi et al., 2013), although other reports have shown podoplanin expression as a marker for an unfavorable prognosis (Kitano et al., 2010; Schoppmann et al., 2013).
The gene set expressions associated with worse prognosis in CRC were found to increase in CAFs. Additionally, elevated levels of expression in genes upregulated in CAFs such as α-SMA and FAP indicate poor disease-free or lower survival for individuals with CRC (Herrera et al., 2013; Calon et al., 2015). Another potential marker is secreted glycoprotein stanniocalcin-1 (STC1), which is expressed in CAFs and acts as a mediator through platelet-derived growth factor (PDGF)-mediated signaling and drives the metastasis in CRC (Peña et al., 2013). Other recently discovered CAF-related prognostic markers for CRC are listed in Table 1.
Despite being used widely for CAF identification, conventional CAF biomarkers such as α-SMA are found to be heterogeneous in expression and have a lower specificity, whereby they are also expressed by other cell types besides CAFs, namely mesenchymal and hematopoietic cells (Nurmik et al., 2020). Therefore, it is impertinent to discover robust and specific markers, which can be applied in concordance with morphological evaluation for better identification and classification of heterogeneous CAF subpopulations.
CAFs have a complex nature and are thus poorly defined. Depending on the cancer type, phenotypic variation of CAFs may be found. Simon and Salhia (2022) broadly classified CAFs in the TME into four groups: immune, desmoplastic, contractile, and aggressive, where the immune and desmoplastic CAF subtypes have tumor-suppressive properties, while the contractile and aggressive are protumorigenic. Another study by Lavie et al. (2022) reported three major CAF subsets, identified using single-cell RNA sequencing analysis: (a) inflammatory CAFs, (b) antigen-presenting CAFs, and (c) myofibroblastic CAFs (myCAFs). Central characteristics of different CAF subgroups are found to be conserved across different organs and cancer types. Several hallmark genes which signify specific CAF features were identified. Nevertheless, the CAF signatures differ according to the organ. For example, specialized stromal cell subgroups (e.g., pericryptal Ptgs2-expressing fibroblasts and crypt-bottom fibroblasts) were reported to support epithelial stemness in the colon (Roulis et al., 2020).
As stated by various reports, different CAF populations in CRC have been identified using various markers. Table 2 summarizes these CAF subtypes, identified using different biomarkers, in the TME of animal and human CRC models.
Translational outlook of cancer-associated fibroblasts in colorectal cancer
CAFs are proven to be an active player in colorectal carcinogenesis, and studies surrounding this cell population have been extensively explored over the past two decades. This emphasizes several promising clinical applications of CAFs in CRC management.
Future CRC treatment will evolve to be more tailored to patients, which fulfills the need for personalized medicine. Recent discoveries in oncology have presented different strategies for CRC therapy, including targeting CAFs (Sahai et al., 2020), which potentially serve as a complement to conventional therapy. Conventional CRC therapy involves chemotherapeutic drugs targeting tumors more broadly, which renders it less effective and causes multiple adverse effects. Combining CAF-targeted therapy with a conventional mode of treatment hypothetically will increase the effectiveness of treatment and abolish cancer progression, growth, and metastasis.
Although the notion of CAF- or TME-targeted therapy, in general, is intriguing, existing treatments targeting the TME approved by the United States Food and Drug Administration (FDA) have limited efficacy. However, pre-clinical studies have shown promises results in employing the TME components, including chimeric antigen receptor-transduced natural killer (CAR-NK) cells, stellate cells, and fibroblasts as potential new targets for cancer treatment (Anderson and Simon, 2020; Zhang et al., 2022).
Designing an effective treatment against CAF populations would require robust biomarkers for identifying cells of the target. To fully characterize CAFs in CRC, multiple markers are suggested for their identification. Although various CAF markers have been established, their relationship and correlation with expression in CRC are still under investigation due to their complexity.
To improve patient management, further prognostic and/or predictive CRC markers are needed. Current diagnosis involves tumor, node, and metastasis (TNM) classification is limited as it relies on anatomical evaluation. Other lower-cost, simpler, and quick assessments, such as tumor-stroma ratio (TSR), have been suggested for patient stratification (Gao et al., 2021). However, this pathological marker may provide limited information regarding the tumor phenotype. Although other biomarkers or biomolecules based on molecular tumor profiling, such as genomic and transcriptomics analyses, acquire comprehensive and massive data, these are costly compared to conventional histopathological examination.
Circulating biomolecules of CRC have been reported to possess clinical utility. These biomolecules include carcinoembryonic antigen (CEA), calretinin, CDH17, MOC-31 (Ep-CAM), CK20, CK7, CA19-9, and MUC2 (enzyme) (Petrik et al., 2022). Mutational profiling of KRAS, BRAF, and PIK3CA also are being investigated due to their high prevalence in CRC. These biomarkers represent the epithelial origin of CRC instead of the fibroblast or CAF population of this cancer. Therefore, it is paramount to identify robust and specific biomarkers for CAFs for application in liquid biopsy for CRC.
Proposed translational prospects of CAF in CRC are elaborated further in the following section and depicted in Fig. 1.
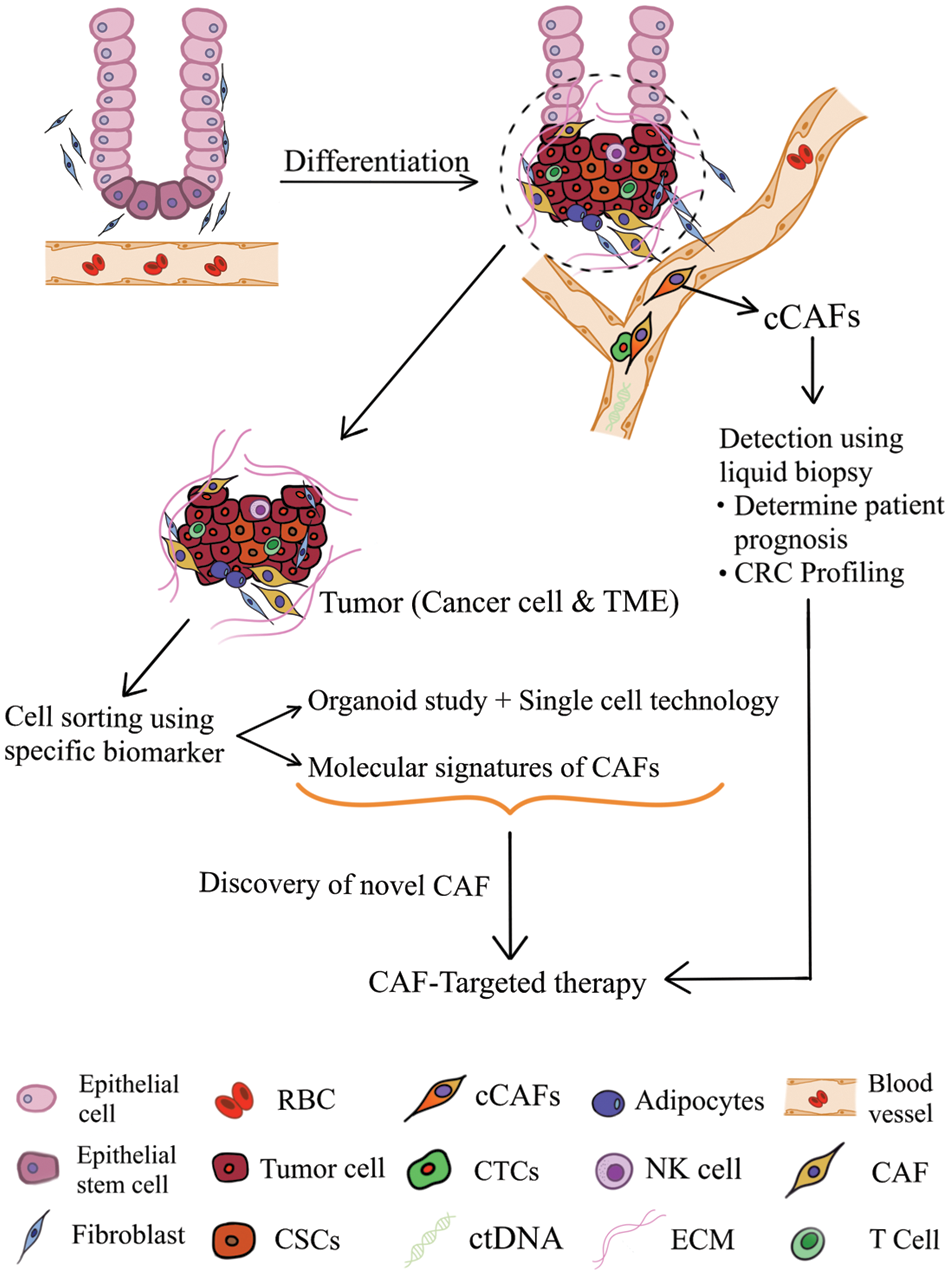
Figure 1: Translational prospects of cancer-associated fibroblasts in colorectal cancer (TME: tumor microenvironment; CAF: cancer-associated fibroblast; cCAF, circulating CAF; CTCs, circulating tumor cells; RBC, red blood cell; ctDNA, circulating tumor DNA; NK cell, natural killer cell; ECM, extracellular matrix; CSC, cancer stem cell).
Cancer-associated fibroblasts in liquid biopsy application for colorectal cancer
A significant number of CRC cases are diagnosed at the later stages (Meester et al., 2019; Siegel et al., 2020), largely due to a lack of screening and awareness of cancer prevention among the public. A high recurrence rate was found in CRC, and this may differ according to disease staging. Approximately 80% of stage IV CRC patients relapse within the first 2 years following surgery with curative intent (de Jong et al., 2009), whereas recurrence is lower (30%) for stages I–III patients (van der Stok et al., 2017). Existing clinical procedures to screen and diagnose CRC, such as colonoscopy, can be considered invasive, present a risk of complications, and cost ineffective and inconvenient, thus rendering CRC screening underused. The non-invasive option to screen for CRC is predicted be increasingly important in the future (Shaukat and Levin, 2022).
Liquid biopsy, which is the most current interest in oncology, allows a less invasive, more convenient, and frequent sampling compared to tissue biopsy and complements molecular imaging assessments (Cherry et al., 2018). The collected specimens include peripheral blood, urine, saliva, and other bodily fluid. The advancement in liquid biopsy aims to dissect spatial and temporal heterogeneity in tumor biology and can be achieved through serial blood test, rather than biopsy of the primary tumor, which is performed years prior on a single metastatic site when multiple metastases are potentially present before any drug treatment. This may improve patient management and treatment outcomes as well as enable early cancer detection (Pantel and Alix-Panabières, 2010; Wan et al., 2017). Despite great advances in liquid biopsy, the application of this approach in clinics has been slow (Ignatiadis et al., 2015).
Liquid biopsy involves the isolation and detection of tumor-derived analytes, including (a) circulating tumor cells (CTCs), (b) circulating nucleic acids such as circulating tumor DNA (ctDNA), cell-free DNA (cfDNA), cell-free RNAs (mRNAs, long non-coding RNAs, and microRNAs), (c) tumor extracellular vesicles, and (d) proteins and metabolites in the body fluids of individuals with cancer, followed by genomic and proteomic analyses (Ramalingam and Jeffrey, 2018; Lone et al., 2022). To date, a very little study has been conducted to explore circulating TME cellular components, such as CAFs in liquid biopsy, despite its promising clinical prospects.
CTCs, one of liquid biopsy analytes, refer to subsets of tumor- or metastasis-derived cells that circulate in the bloodstream. CTC isolation is less invasive and safer compared to conventional tissue biopsy. CTCs can be applied to monitor cancer progression and assess response to therapy in real-time. Detection of CTCs in peripheral blood serves as valuable tumor markers, including for CRC (Yap et al., 2014). CTCs in CRC include a vast range of cells, including epithelial tumor cells, tumor cells undergoing epithelial-mesenchymal transition (EMT), and cancerous stem cells (Hardingham et al., 2015). It is proposed that epithelial cancer cells migrate and invade the blood vessel after EMT, which is the main cause of metastasis in vivo (Masuda et al., 2016; Jia et al., 2017). Upon reaching a secondary organ, CTCs undergo a mesenchymal-epithelial transition, reacquire stemness properties and form a new metastatic site (Suarez-Carmona et al., 2017). Contrary to other cancer biomarkers, CTCs are viable tumor cells, carrying overall molecular and biological profiling of the tumor, promoting single-cell analysis, and directly indicates the ongoing alteration in tumor mass at various stages (Li et al., 2020).
Over the past 20 years, great advancements have been made in the detection methods of heterogeneous cells in colorectal tumors. Currently, tumor staging is performed by histology and radiological imaging. These techniques lack sensitivity for the detection of micrometastases or early tumor dissemination, which are vital for metastatic progression of the disease. The presence of CTCs indicates micrometastasis events and potentially have prognostic values independently of established staging factors (e.g., lymph node involvement) (Hardingham et al., 2015).
Vast arrays of advances in science, such as high-throughput sequencing (Jones et al., 2008), CRISPR/Cas9 editing tools (Chen et al., 2015), and single-cell technologies (Navin et al., 2011), have helped to gradually uncover biological phenomenon including mechanisms underlying metastasis as well as involvement of key driver genes, EMT of tumor cells, the role of exosomes in tumorigenesis, CTCs and complex interplay between tumor cells and the TME (Peinado et al., 2012; Quail and Joyce, 2013; Yu et al., 2013; Aceto et al., 2014). CTC detection is usually highly dependent on molecular markers, the most commonly used being the epithelial cell adhesion molecule markers. These molecular markers vary according to cancer types (Lin et al., 2021).
To date, there is very little knowledge of CAF-CTC interplay. The earliest report by Duda et al. (2010) on animal models of lung cancer showed that CTCs could interact and carry CAFs from the primary tumor to the metastatic site (Ortiz-Otero et al., 2020a). Molecular changes such as decreased expression of epithelial markers, including E-cadherin and claudins, as well as elevated levels of mesenchymal markers (vimentin, N-cadherin, FSP-1, and fibronectin) are observed during the EMT process (Mittal, 2018). A small fraction of CTCs interacts with different TME components, such as neutrophils, macrophages, and CAFs, to escape the immune system, thus driving their survival (Rejniak, 2016; Garrido-Navas et al., 2019).
In a study on human CRC samples, CAFs have been proposed to promote metastasis by creating gaps in the basement membrane that facilitates cancer cell invasion independently of matrix metalloproteinase (MMP) (Glentis et al., 2017). Additionally, during the dissemination process, CAFs are able to protect CTCs from the fluid shear force via stable intracellular contact and secretomes such as CXCL5, CCL2, and CCL7, which are linked to cell survival, invasion, and EMT (Ortiz-Otero et al., 2020a). A high proportion of mature CAFs was reported in intratumoral stroma (57.6%) and invasive front of CRC (60.3%). Interestingly, overexpression of epidermal growth factor receptor (EGFR) was found in the mature CAFs in the invasive front in comparison to immature CAFs. Increased lymphatic invasion directly correlates with the number of mature fibroblasts in the intratumoral stroma (Shin et al., 2019).
Currently, there is a lack of studies on circulating CAFs (cCAFs) of CRC, although reports have been previously published on cCAF detection in breast cancer. The isolation of cCAFs has been reported in the peripheral blood of metastatic breast cancer patients but not in individuals in their early stages (Ao et al., 2015). A recent report by Jiang et al. (2021) has demonstrated a novel, effective, label-free, and high throughput microstreaming platform for the isolation of cCAFs, CTCs, and immune cells from breast cancer patients. As stated in the previous section, CTCs represent the precursor of metastasis, where they can migrate as individual CTCs or in clusters (CTCs with stromal cells such as CAFs). However, until now, there is little direct evidence indicating CAF presence in the circulation of cancer patients in the clinical setting.
Ishii et al. (2007) have reported the presence of fibroblast progenitor cells in individuals with lung cancer, while Jones et al. (2013) demonstrated the detection of fibroblast-like cells in blood samples of metastatic prostate cancer but not in patients with localized prostate cancer or no known malignancy. Both studies employed vimentin as the marker for fibroblast population profiling despite this is not specific to fibroblast cells considering that it is also expressed by lymphocytes (Nieminen et al., 2006), as well as breast cancer cells and CTCs that have undergone EMT (Aktas et al., 2009). A more recent finding by Ao et al. (2015) used widely accepted CAF biomarkers, namely FAP+/α-SMA+/Cytokeratin−/CD45− for the detection of cCAFs in the peripheral blood of patients with metastatic breast cancer. In addition, their technique of evaluating the co-expression of FAP/α-SMA on the top of novel cell-size-based microfilter technology, which was initially used to capture CTCs (Zheng et al., 2007), has enabled the separation of CAFs from CTCs from blood samples of patients with breast, prostate, and colon cancer.
Using blood samples from various metastatic cancer patients, including CRC, followed by an assessment of CTC and CAF numbers, Ortiz-Otero et al. (2020b) established the correlation between cCAF levels with poor prognosis and lower survival rate in the selected samples. This report highly suggests CAF level as a predictive biomarker for CRC prognosis.
One of the challenges in studying CRC metastasis in vivo is the transient nature of carcinoma in situ, which rendered the onset of cancer invasion and basement membrane breakage laborious to study. Nevertheless, previous in vitro studies indicated that the metastasis process could be inhibited by hindering CAF formation and functions (Cazet et al., 2018; Li et al., 2019). Yang et al. (2021) reported different possible mechanisms of CAF metastasis in CRC. They further confirmed that the inhibition of CAF migration and invasion abolished the metastasis of both tumor cells and CAFs through an in vivo study. Worth noting that this study presents several limitations, including a lack of confirmation on the correlation between cCAF levels and the prognosis of CRC patients and the mechanism of CAF colonization and survival in the distant niche. This may be achieved by collecting peripheral blood from patients with CRC and isolating cCAFs for further analysis.
Based on the potential of cCAFs in liquid biopsy, we proposed that cCAF utilization in the clinical management of CRC can be potentially expanded to:
i) stratify patients more effectively
ii) indicate the staging and/or subtype of the disease
iii) be used as complementary to established genetic/genomic testing and tissue biopsy analyses
Cancer-associated fibroblasts-targeted therapy
Over the past decade, there is a great enthusiasm surrounding the idea of designing a targeted treatment against TME components in various cancers. Targeted therapies, which embody the concept of “precision medicine,” are preferred compared to traditional cancer treatment due to their higher efficacy and less adverse effect. Considering the major role of CAFs in colorectal carcinogenesis, it is not surprising that many are focusing on CAF-related treatment to eradicate CRC. Although studying cCAFs seems to be a good direction to pursue, there are still valid concerns on the lack of biomarkers for the detection of specific populations of CAFs. It is an established notion that CAFs are heterogeneous; thus, having robust and specific CAF biomarkers will ensure optimum specificity and sensitivity in the detection and isolation of cCAFs. This subsequently will drive the discovery of CAF-targeted therapy for CRC.
Taking into account that CRC has a molecular subset characterized by accumulation of stromal cells (CMS4), this serves as a great target for targeted treatment against CAFs. Moreover, as stated previously, this CRC subset is characterized by the worst prognosis among the four CMSs (Varga et al., 2020; Kasashima et al., 2021). In a recent report by Zhong et al. (2022) on transcription factor (TF) signatures of CMS4 has revealed nine TF-related genes as main regulators for the CMS4 subtype. These genes are MEIS2, MEIS3, SNAI1, KLF17, BARX1, ZNF532, HEYL, FOXL2 and LHX6. Interestingly, these genes are implicated as prognostic factors in CRC in both analyses using The Cancer Genome Atlas (TCGA) and International Cancer Genome Consortium (ICGC)-ARGO data. Additionally, these TF-related genes are able to discriminate patient survival after adjustment to clinical factors linked to prognosis, such as gender, TNM classification, MSI status, and the location of primary tumors (left versus right-sided). Microenvironment components in high-risk CRC patients with highly immunosuppressed profiles (overexpression of TIM3, CD39, and CD40) indicated that these individuals might not benefit from conventional immune checkpoint inhibitors (ICIs) such as anti-PD1 therapies. Stratification of CRC patients through gene expression profiling of CAFs, as reported, will drive the exploration of targeted drugs. Additionally, this information also can explain the behavior of high-risk CRC patients and predict drug response.
The interplay between CAF and cancer cells is facilitated by intricate autocrine and paracrine signaling pathways. These include TGF-β, phosphoinositide 3-kinase/AKT/mammalian target of rapamycin (PI3K/AKT/mTOR), mitogen-activated protein kinases (MAPK), Wnt, EGFR, Hippo, and nuclear factor kappa-light-chain-enhancer of activated B cells (NF-κB) pathways (Wu et al., 2021). These elaborate signaling pathways induce the transition of CAFs from stromal fibroblasts, which subsequently lead to CAF heterogeneity (Erez et al., 2010; Ringuette Goulet et al., 2018). Blocking these signaling pathways may impair the activation of CAFs and affect the CAF-tumor cell crosstalk. Thus, targeting the essential signal cascades involved in the bidirectional communication between cancer cells and CAFs may serve as an excellent strategy to explore treatment avenues for CRC.
The cancer-stromal interactions also promote CAF activation, which leads to CRC progression. It is an established notion that a tumor is a wound that never heal; thus, activated fibroblasts behave in a similar manner as they would to injury. This response involves CAF recruitment to tumor sites that mimic the fibrosis secondary to chronic inflammation (Dimmeler and Zeiher, 2017). Recruitment and activation of CAF are mainly facilitated by various growth factors, including TGF-β, chemokines, and cytokines. Despite many studies conducted on CAF-tumor cell crosstalk, the exact nature of CAF trans-differentiation from non-activated fibroblasts is yet to be fully described.
However, due to the complexity of molecular signaling in CRC, extensive studies must be conducted to further dissect CAFs’ role in tumor progression, their activation process, and their potential in antitumor therapy. Wu et al. (2021) have proposed three alternative mechanisms of CAF-targeted therapies, namely:
Epithelial-mesenchymal common target (EMCT)–targeting both tumor cells and adjacent stromal cells.
Sequential target perturbation (STP)–blocking protumorigenic process prior to treating cancers cells to acquire anticancer effects.
Crosstalk-directed signaling target (CDST)–targeting two different components of a signaling pathway in CAFs and tumor cells simultaneously.
This report by Wu et al. (2021) corroborated a recent publication by Wang et al. (2022). They suggested four main strategies for anticancer therapy against CAFs through:
i) CAF deletion (by targeting specific markers of CAFs such as α-SMA and FAP)
ii) CAF normalization (shift from pro-carcinogenic to tumor-suppressive state using all-trans retinoic acid (ATRA) or VDR ligands)
iii) blocking CAF activation (targeting cytokines and growth factor signaling)
iv) ECM remodeling
Although the prospect of targeting CAFs for CRC treatment is intriguing, the clinical data have not yet fully supported the benefit of these therapies. A pre-clinical study employing the co-culture method on CRC patient-derived organoids would be extremely helpful in discovering the activity of drugs against CAFs (Barnett and Vilar, 2017). An example of this approach, as reported by Biffi et al. (2019) using organoid and mouse model of pancreatic ductal adenocarcinoma (PDAC), demonstrated that IL-1-induced leukemia inhibitory factor (LIF) expression and downstream JAK/STAT activation resulted in inflammatory CAF (iCAFs) production. TGF-β is found to inhibit this signaling by suppressing IL-1 receptor type 1 (IL1R1) expression and suggesting a transition from iCAFs into myofibroblasts.
Organoid study on human or animal models must be accompanied by the advanced technique of flow cytometry that allows cell separation and identification of CAFs based on established molecular markers. Subsequent analysis at the single-cell level will provide insight into more accurate profiling of CAF populations in CRC. This step is essential, considering that various CAF subgroups with different molecular signatures were detected in CRC (Li et al., 2017; Joanito et al., 2022; Zhao et al., 2022), which highlights their functional diversity that may influence carcinogenesis differently.
CAFs in the TME of CRC represent highly complex and heterogeneous populations. The diversity in the CAF subtypes in the colorectal tumor niche and their interplay with malignant cells drive the evolution and progression of cancer, thus contributing to the heterogeneity of CRC. A repertoire of biomarkers is applied to identify various subgroups of CAF and to aid further understanding of molecular pathways involving CAF-tumor cell crosstalk in colorectal tumor progression. Deciphering the complexity of CAFs will provide insight into more accurate CRC patient profiling through a liquid biopsy approach and analysis of cCAFs. In addition, CAFs also potentially serve as targets for CRC therapy. Despite encouraging pre-clinical data and great promises of the translational prospects of CAFs, there are scientific challenges that need to be addressed in order to implement effective clinical applications utilizing CAFs in the future.
Acknowledgement: The authors would like to acknowledge the support of the GRA-Assist Scheme (Graduate Student Financial Assistance), Universiti Sains Malaysia (USM).
Funding Statement: This work was supported by the Ministry of Higher Education Malaysia for Fundamental Research Grant Scheme with Project Code: FRGS/1/2020/SKK0/USM/03/10 and USM.
Author Contributions: Manuscript conceptualization: MM; literature review: EAS, MM; draft manuscript preparation and review: EAS, MM, LYY, AZD, NACJ. All authors approved the final version of the manuscript.
Ethics Approval: Not applicable.
Conflicts of Interest: The authors declare that they have no conflicts of interest to report regarding the present study.
References
Aceto N, Bardia A, Miyamoto DT, Donaldson MC, Wittner BS et al. (2014). Circulating tumor cell clusters are oligoclonal precursors of breast cancer metastasis. Cell 158: 1110–1122. https://doi.org/10.1016/j.cell.2014.07.013 [Google Scholar] [PubMed] [CrossRef]
Ahmad Zawawi SS, Musa M (2022). Dynamic co-evolution of cancer cells and cancer-associated fibroblasts: Role in right- and left-sided colon cancer progression and its clinical relevance. Biology 11: 1014. https://doi.org/10.3390/biology11071014 [Google Scholar] [PubMed] [CrossRef]
Ait Ouakrim D, Pizot C, Boniol M, Malvezzi M, Boniol M, Negri E, Bota M, Jenkins MA, Bleiberg H, Autier P (2015). Trends in colorectal cancer mortality in Europe: Retrospective analysis of the WHO mortality database. British Medical Journal 351: 4970. https://doi.org/10.1136/bmj.h4970 [Google Scholar] [PubMed] [CrossRef]
Akimoto N, Ugai T, Zhong R, Hamada T, Fujiyoshi K, Giannakis M, Wu K, Cao Y, Ng K, Ogino S (2021). Rising incidence of early-onset colorectal cancer—A call to action. Nature Review Clinical Oncology 18: 230–243. https://doi.org/10.1038/s41571-020-00445-1 [Google Scholar] [PubMed] [CrossRef]
Aktas B, Tewes M, Fehm T, Hauch S, Kimmig R, Kasimir-Bauer S (2009). Stem cell and epithelial-mesenchymal transition markers are frequently overexpressed in circulating tumor cells of metastatic breast cancer patients. Breast Cancer Research 11: R46. https://doi.org/10.1186/bcr2333 [Google Scholar] [PubMed] [CrossRef]
Anderson NM, Simon MC (2020). The tumor microenvironment. Current Biology 30: 921–925. https://doi.org/10.1016/j.cub.2020.06.081 [Google Scholar] [PubMed] [CrossRef]
Ao Z, Shah SH, Machlin LM, Parajuli R, Miller PC et al. (2015). Identification of cancer-associated fibroblasts in circulating blood from patients with metastatic breast cancer. Cancer Research 78: 681–4687. https://doi.org/10.1158/0008-5472.CAN-15-1633 [Google Scholar] [PubMed] [CrossRef]
Barnett RM, Vilar E (2017). Targeted therapy for cancer-associated fibroblasts: Are we there yet? Journal of the National Cancer Institute 110: 11–13. https://doi.org/10.1093/jnci/djx131 [Google Scholar] [PubMed] [CrossRef]
Biffi G, Oni TE, Spielman B, Hao Y, Elyada E, Park Y, Tuveson DA (2019). IL1-Induced JAK/STAT signaling is antagonized by TGFβ to shape CAF heterogeneity in pancreatic ductal adenocarcinoma. Cancer Discovery 9: 282–301. https://doi.org/10.1158/2159-8290.CD-18-0710 [Google Scholar] [PubMed] [CrossRef]
Binefa G, Rodríguez-Moranta F, Teule A, Medina-Hayas M (2014). Colorectal cancer: From prevention to personalized medicine. World Journal of Gastroenterology 20: 6786–6808. https://doi.org/10.3748/wjg.v20.i22.6786 [Google Scholar] [PubMed] [CrossRef]
Bissell MJ, Radisky D (2001). Putting tumors in context. Nature Reviews Cancer 1: 46–54. https://doi.org/10.1038/35094059 [Google Scholar] [PubMed] [CrossRef]
Bizzarri M, Cucina A, Biava PM, Proietti S, D’Anselmi F, Dinicola S, Lisi E (2011). Embryonic morphogenetic field induces phenotypic reversion in cancer cells. Current Pharmaceutical Biotechnology 12: 243–253. https://doi.org/10.2174/138920111794295701 [Google Scholar] [PubMed] [CrossRef]
Black JRM, McGranahan N (2021). Genetic and non-genetic clonal diversity in cancer evolution. Nature Reviews Cancer 21: 379–392. https://doi.org/10.1038/s41568-021-00336-2 [Google Scholar] [PubMed] [CrossRef]
Bochet L, Lehuédé C, Dauvillier S, Wang YY, Dirat B et al. (2013). Adipocyte-derived fibroblasts promote tumor progression and contribute to the desmoplastic reaction in breast cancer. Cancer Research 73: 5657–5668. https://doi.org/10.1158/0008-5472.CAN-13-0530 [Google Scholar] [PubMed] [CrossRef]
Bussard KM, Mutkus L, Stumpf K, Gomez-Manzano C, Marini FC (2016). Tumor-associated stromal cells as key contributors to the tumor microenvironment. Breast Cancer Reserach 18: 84. https://doi.org/10.1186/s13058-016-0740-2 [Google Scholar] [PubMed] [CrossRef]
Calon A, Espinet E, Palomo-Ponce S, Tauriello DV, Iglesias M et al. (2012). Dependency of colorectal cancer on a TGF-β-driven program in stromal cells for metastasis initiation. Cancer Cell 22: 571–584. https://doi.org/10.1016/j.ccr.2012.08.013 [Google Scholar] [PubMed] [CrossRef]
Calon A, Lonardo E, Berenguer-Llergo A, Espinet E, Hernando-Momblona X et al. (2015). Stromal gene expression defines poor-prognosis subtypes in colorectal cancer. Nature Genetics 47: 320–329. https://doi.org/10.1038/ng.3225 [Google Scholar] [PubMed] [CrossRef]
Cazet AS, Hui MN, Elsworth BL, Wu SZ, Roden D et al. (2018). Targeting stromal remodeling and cancer stem cell plasticity overcomes chemoresistance in triple negative breast cancer. Nature Communications 9: 2897. https://doi.org/10.1038/s41467-018-05220-6 [Google Scholar] [PubMed] [CrossRef]
Chen S, Sanjana NE, Zheng K, Shalem O, Lee K et al. (2015). Genome-wide CRISPR screen in a mouse model of tumor growth and metastasis. Cell 160: 1246–1260. https://doi.org/10.1016/j.cell.2015.02.038 [Google Scholar] [PubMed] [CrossRef]
Chen YF, Yu ZL, Lv MY, Cai ZR, Zou YF, Lan P, Wu XJ, Gao F (2021). Cancer-associated fibroblasts impact the clinical outcome and treatment response in colorectal cancer via immune system modulation: A comprehensive genome-wide analysis. Molecular Medicine 27: 139. https://doi.org/10.1186/s10020-021-00402-3 [Google Scholar] [PubMed] [CrossRef]
Cherry SR, Jones T, Karp JS, Qi J, Moses WW, Badawi RD (2018). Total-body PET: Maximizing sensitivity to create new opportunities for clinical research and patient care. Journal of Nuclear Medicine 59: 3–12. https://doi.org/10.2967/jnumed.116.184028 [Google Scholar] [PubMed] [CrossRef]
Choi SY, Sung R, Lee SJ, Lee TG, Kim N, Yoon SM, Lee EJ, Chae HB, Youn SJ, Park SM (2013). Podoplanin, α-smooth muscle actin or S100A4 expressing cancer-associated fibroblasts are associated with different prognosis in colorectal cancers. Journal Korean Medical Science 28: 1293–1301. https://doi.org/10.3346/jkms.2013.28.9.1293 [Google Scholar] [PubMed] [CrossRef]
de Jong M C, Pulitano C, Ribero D, Strub J, Mentha G, Schulick R D, Choti M A, Aldrighetti L, Capussotti L, Pawlik T M (2009). Rates and patterns of recurrence following curative intent surgery for colorectal liver metastasis: An international multi-institutional analysis of 1669 patients. Annals of Surgery 250: 440–448. https://doi.org/10.1097/SLA.0b013e3181b4539b [Google Scholar] [PubMed] [CrossRef]
Dienstmann R, Villacampa G, Sveen A, Mason M J, Niedzwiecki D, Nesbakken A, Moreno V, Warren R S, Lothe R A, Guinney J (2019). Relative contribution of clinicopathological variables, genomic markers, transcriptomic subtyping and microenvironment features for outcome prediction in stage II/III colorectal cancer. Annals of Oncology 30: 1622–1629. https://doi.org/10.1093/annonc/mdz287 [Google Scholar] [PubMed] [CrossRef]
Dimmeler S, Zeiher AM (2017). Netting insights into fibrosis. New England Journal of Medicine 376: 1475–1477. https://doi.org/10.1056/NEJMcibr1616598 [Google Scholar] [PubMed] [CrossRef]
Duda DG, Duyverman AM, Kohno M, Snuderl M, Steller EJ, Fukumura D, Jain RK (2010). Malignant cells facilitate lung metastasis by bringing their own soil. Proceedings of the National Academy of Sciences of the United States of America 107: 21677–21682. https://doi.org/10.1073/pnas.1016234107 [Google Scholar] [PubMed] [CrossRef]
Erez N, Truitt M, Olson P, Arron ST, Hanahan D (2010). Cancer-associated fibroblasts are activated in incipient neoplasia to orchestrate tumor-promoting inflammation in an NF-κB-Dependent Manner. Cancer Cell 17: 135–147. https://doi.org/10.1016/j.ccr.2009.12.041 [Google Scholar] [PubMed] [CrossRef]
Fearon ER, Vogelstein B (1990). A genetic model for colorectal tumorigenesis. Cell 61: 759–767. https://doi.org/10.1016/0092-8674(90)90186-I [Google Scholar] [PubMed] [CrossRef]
Ferrer-Mayorga G, Gómez-López G, Barbáchano A, Fernández-Barral A, Peña C, Pisano DG, Cantero R, Rojo F, Muñoz A, Larriba MJ (2017). Vitamin D receptor expression and associated gene signature in tumour stromal fibroblasts predict clinical outcome in colorectal cancer. Gut 66: 1449–1462. https://doi.org/10.1136/gutjnl-2015-310977 [Google Scholar] [PubMed] [CrossRef]
Fiori ME, di Franco S, Villanova L, Bianca P, Stassi G, De Maria R (2019). Cancer-associated fibroblasts as abettors of tumor progression at the crossroads of EMT and therapy resistance. Molecular Cancer 18: 70. https://doi.org/10.1186/s12943-019-0994-2 [Google Scholar] [PubMed] [CrossRef]
Gao J, Shen Z, Deng Z, Mei L (2021). Impact of tumor-stroma ratio on the prognosis of colorectal cancer: A systematic review. Frontiers in Oncology 11: 738080. https://doi.org/10.3389/fonc.2021.738080 [Google Scholar] [PubMed] [CrossRef]
Garrido-Navas C, de Miguel-Perez D, Exposito-Hernandez J, Bayarri C, Amezcua V, Ortigosa A, Serrano MJ (2019). Cooperative and escaping mechanisms between circulating tumor cells and blood constituents. Cells 8: 1382. https://doi.org/10.3390/cells8111382 [Google Scholar] [PubMed] [CrossRef]
Giguelay A, Turtoi E, Khelaf L, Tosato G, Dadi I et al. (2022). The landscape of cancer-associated fibroblasts in colorectal cancer liver metastases. Theranostics 12: 7624–7639. https://doi.org/10.7150/thno.72853 [Google Scholar] [PubMed] [CrossRef]
Glentis A, Oertle P, Mariani P, Chikina A, El Marjou F, Attieh Y, Vignjevic DM (2017). Cancer-associated fibroblasts induce metalloprotease-independent cancer cell invasion of the basement membrane. Nature Communications 8: 924. https://doi.org/10.1038/s41467-017-00985-8 [Google Scholar] [PubMed] [CrossRef]
Goetz JG, Minguet S, Navarro-Lérida I, Lazcano JJ, Samaniego R et al. (2011). Biomechanical remodeling of the microenvironment by stromal caveolin-1 favors tumor invasion and metastasis. Cell 146: 148–163. https://doi.org/10.1016/j.cell.2011.05.040 [Google Scholar] [PubMed] [CrossRef]
Gorchs L, Fernández Moro C, Bankhead P, Kern KP, Sadeak I, Meng Q, Rangelova E, Kaipe H (2019). Human pancreatic carcinoma-associated fibroblasts promote expression of co-inhibitory markers on CD4+ and CD8+ T-cells. Frontiers in Immunology 24: 847. https://doi.org/10.3389/fimmu.2019.00847 [Google Scholar] [PubMed] [CrossRef]
Guinney J, Dienstman R, Wang X, de Reyniès A, Schlicker A et al. (2015). The consensus molecular subtypes of colorectal cancer. Nature Medicine 21: 1350–1356. https://doi.org/10.1038/nm.3967 [Google Scholar] [PubMed] [CrossRef]
Hardingham JE, Grover P, Winter M, Hewett PJ, Price TJ, Thierry B (2015). Detection and clinical significance of circulating tumor cells in colorectal cancer—20 years of progress. Molecular Medicine 21: S25–S31. https://doi.org/10.2119/molmed.2015.00149 [Google Scholar] [PubMed] [CrossRef]
Heichler C, Schmied A, Enderle K, Scheibe K, Murawska M, Schmid B, Neufert C (2022). Targeting STAT3 signaling in COL1+ fibroblasts controls colitis-associated cancer in mice. Cancers 14: 1472. https://doi.org/10.3390/cancers14061472 [Google Scholar] [PubMed] [CrossRef]
Herrera M, Berral-González A, López-Cade I, Galindo-Pumariño C, Bueno-Fortes S et al. (2021). Cancer-associated fibroblast-derived gene signatures determine prognosis in colon cancer patients. Molecular Cancer 20: 73. https://doi.org/10.1186/s12943-021-01367-x [Google Scholar] [PubMed] [CrossRef]
Herrera M, Herrera A, Domínguez G, Silva J, García V et al. (2013). Cancer-associated fibroblast and M2 macrophage markers together predict outcome in colorectal cancer patients. Cancer Science 104: 437–444. https://doi.org/10.1111/cas.12096 [Google Scholar] [PubMed] [CrossRef]
Hinoi T (2021). Cancer genomic profiling in colorectal cancer: Current challenges in subtyping colorectal cancers based on somatic and germline variants. Journal of the Anus, Rectum and Colon 5: 213–228. https://doi.org/10.23922/jarc.2021-009 [Google Scholar] [PubMed] [CrossRef]
Huynh PT, Beswick EJ, Coronado YA, Johnson P, O’Connell MR et al. (2016). CD90+ stromal cells are the major source of IL-6, which supports cancer stem-like cells and inflammation in colorectal cancer. International Journal of Cancer 138: 1971–1981. https://doi.org/10.1002/ijc.29939 [Google Scholar] [PubMed] [CrossRef]
Ignatiadis M, Lee M, Jeffrey SS (2015). Circulating tumor cells and circulating tumor DNA: Challenges and opportunities on the path to clinical utility. Clinical Cancer Research 21: 4786–4800. https://doi.org/10.1158/1078-0432.CCR-14-1190 [Google Scholar] [PubMed] [CrossRef]
Ishii G, Ito TK, Aoyagi K, Fujimoto H, Chiba H, Hasebe T, Fujii S, Nagai K, Sasaki H, Ochiai A (2007). Presence of human circulating progenitor cells for cancer stromal fibroblasts in the blood of lung cancer patients. Stem Cells 25: 1469–1477. https://doi.org/10.1634/stemcells.2006-0449 [Google Scholar] [PubMed] [CrossRef]
Jackstadt R, Norman JC (2021). Stromal WNTer keeps the tumor cold and drives metastasis. Developmental Cell 56: 3–4. https://doi.org/10.1016/j.devcel.2020.12.011 [Google Scholar] [PubMed] [CrossRef]
Jia S, Zhang R, Li Z, Li J (2017). Clinical and biological significance of circulating tumor cells, circulating tumor DNA, and exosomes as biomarkers in colorectal cancer. Oncotarget 8: 55632–55645. https://doi.org/10.18632/oncotarget.17184 [Google Scholar] [PubMed] [CrossRef]
Jiang R, Agrawal S, Aghaamoo M, Parajuli R, Agrawal A, Lee AP (2021). Rapid isolation of circulating cancer associated fibroblasts by acoustic microstreaming for assessing metastatic propensity of breast cancer patients. Lab on a Chip 21: 875–887. https://doi.org/10.1039/D0LC00969E [Google Scholar] [PubMed] [CrossRef]
Joanito I, Wirapati P, Zhao N, Nawaz Z, Yeo G et al. (2022). Single-cell and bulk transcriptome sequencing identifies two epithelial tumor cell states and refines the consensus molecular classification of colorectal cancer. Nature Genetics 54: 963–975. https://doi.org/10.1038/s41588-022-01100-4 [Google Scholar] [CrossRef]
Jones S, Chen WD, Parmigiani G, Diehl F, Beerenwinkel N et al. (2008). Comparative lesion sequencing provides insights into tumor evolution. Proceedings of National Academy of Sciences of the United States of America 105: 4283–4288. https://doi.org/10.1073/pnas.0712345105 [Google Scholar] [PubMed] [CrossRef]
Jones ML, Siddiqui J, Pienta KJ, Getzenberg RH (2013). Circulating fibroblast-like cells in men with metastatic prostate cancer. Prostate 73: 176–181. https://doi.org/10.1002/pros.22553 [Google Scholar] [PubMed] [CrossRef]
Kalluri R (2016). The biology and function of fibroblasts in cancer. Nature Reviews Cancer 16: 582–598. https://doi.org/10.1038/nrc.2016.73 [Google Scholar] [PubMed] [CrossRef]
Kasashima H, Duran A, Martinez-Ordoñez A, Nakanishi Y, Kinoshita H et al. (2021). Stromal SOX2 upregulation promotes tumorigenesis through the generation of a SFRP1/2-expressing cancer-associated fibroblast population. Developmental Cell 56: 95–110. https://doi.org/10.1016/j.devcel.2020.10.014 [Google Scholar] [PubMed] [CrossRef]
Keum N, Giovannucci E (2019). Global burden of colorectal cancer: Emerging trends, risk factors and prevention strategies. Nature Reviews Gastroenterology & Hepatology 16: 713–732. https://doi.org/10.1038/s41575-019-0189-8 [Google Scholar] [PubMed] [CrossRef]
Khaliq AM, Erdogan C, Kurt Z, Turgut SS, Grunvald MW et al. (2022). Refining colorectal cancer classification and clinical stratification through a single-cell atlas. Genome Biology 23: 113. https://doi.org/10.1186/s13059-022-02677-z [Google Scholar] [PubMed] [CrossRef]
Khare T, Bissonnette M, Khare S (2021). CXCL12-CXCR4/CXCR7 axis in colorectal cancer: Therapeutic target in preclinical and clinical studies. International Journal of Molecular Sciences 22: 7371. https://doi.org/10.3390/ijms22147371 [Google Scholar] [PubMed] [CrossRef]
Kitano H, Kageyama S-I, Hewitt SM, Hayashi R, Doki Y, Ozaki Y, Fujino S, Takikita M, Kubo H, Fukuoka J (2010). Podoplanin expression in cancerous stroma induces lymphangiogenesis and predicts lymphatic spread and patient survival. Archieves of Pathology & Laboratory Medicine 134: 1520–1527. https://doi.org/10.5858/2009-0114-OA.1 [Google Scholar] [PubMed] [CrossRef]
Latchman Y, Wood CR, Chernova T, Chaudhary D, Borde M et al. (2001). PD-L2 is a second ligand for PD-1 and inhibits T cell activation. Nature Immunology 2: 261–268. https://doi.org/10.1038/85330 [Google Scholar] [PubMed] [CrossRef]
Lavie D, Ben-Shmuel A, Erez N, Scherz-Shouval R (2022). Cancer-associated fibroblasts in the single-cell era. Nature Cancer 3: 793–807. https://doi.org/10.1038/s43018-022-00411-z [Google Scholar] [PubMed] [CrossRef]
LeBleu VS, Kalluri R (2018). A peek into cancer-associated fibroblasts: Origins, functions and translational impact. Disease Models & Mechanism 11: dmm029447. https://doi.org/10.1242/dmm.029447 [Google Scholar] [PubMed] [CrossRef]
Li H, Courtois ET, Sengupta D, Tan Y, Chen KH et al. (2017). Reference component analysis of single-cell transcriptomes elucidates cellular heterogeneity in human colorectal tumors. Nature Genetics 49: 708–718. https://doi.org/10.1038/ng.3818 [Google Scholar] [PubMed] [CrossRef]
Li XY, Dong M, Zang XY, Li MY, Zhou JY, Ma JJ, Wang GY (2020). The emerging role of circulating tumor cells in cancer management. American Journal of Translational Research 12: 332–342. [Google Scholar] [PubMed]
Li S, Ou Y, Liu S, Yin J, Zhuo W, Huang M, Zhu T, Zhang W, Zhou H, Liu Z (2019). The fibroblast TIAM2 promotes lung cancer cell invasion and metastasis. Journal of Cancer 10: 1879–1889. https://doi.org/10.7150/jca.30477 [Google Scholar] [PubMed] [CrossRef]
Li Q, Zhang D, Wang Y, Sun P, Hou X, Larner J, Mi J (2013). MiR-21/Smad 7 signaling determines TGF-β1-induced CAF formation. Scientific Reports 3: 2038. https://doi.org/10.1038/srep02038 [Google Scholar] [PubMed] [CrossRef]
Liang C, Ji D, Qin F, Chen G (2023). CAF signature predicts the prognosis of colorectal cancer patients: A retrospective study based on bulk RNA sequencing and single-cell RNA sequencing data. Medicine 102: e33149. https://doi.org/10.1097/MD.0000000000033149 [Google Scholar] [PubMed] [CrossRef]
Lin D, Shen L, Luo M, Zhang K, Li J et al. (2021). Circulating tumor cells: Biology and clinical significance. Signal Transduction and Target Therapy 6: 404. https://doi.org/10.1038/s41392-021-00817-8 [Google Scholar] [PubMed] [CrossRef]
Lone SN, Nisar S, Masoodi T, Singh M, Rizwan A et al. (2022). Liquid biopsy: A step closer to transform diagnosis, prognosis and future of cancer treatments. Molecular Cancer 21: 79. https://doi.org/10.1186/s12943-022-01543-7 [Google Scholar] [PubMed] [CrossRef]
Maia A, Schöllhorn A, Schuhmacher J, Gouttefangeas C (2023). CAF-immune cell crosstalk and its impact in immunotherapy. Seminars in Immunopathology 45: 203–214. https://doi.org/10.1007/s00281-022-00977-x [Google Scholar] [PubMed] [CrossRef]
Masuda T, Hayashi N, Iguchi T, Ito S, Eguchi H, Mimori K (2016). Clinical and biological significance of circulating tumor cells in cancer. Molecular Oncology 10: 408–417. https://doi.org/10.1016/j.molonc.2016.01.010 [Google Scholar] [PubMed] [CrossRef]
Meester RGS, Mannalithara A, Lansdorp-Vogelaar I, Ladabaum U (2019). Trends in incidence and stage at diagnosis of colorectal cancer in adults aged 40 through 49 years, 1975-2015. The Journal of the American Medical Association 321: 1933–1934. https://doi.org/10.1001/jama.2019.3076 [Google Scholar] [PubMed] [CrossRef]
Mittal V (2018). Epithelial mesenchymal transition in tumor metastasis. Annual Review of Pathology 13: 395–412. https://doi.org/10.1146/annurev-pathol-020117-043854 [Google Scholar] [PubMed] [CrossRef]
Miyashita N, Saito A (2021). Organ specificity and heterogeneity of cancer-associated fibroblasts in colorectal cancer. International Journal of Molecular Sciences 22: 10973. https://doi.org/10.3390/ijms222010973 [Google Scholar] [PubMed] [CrossRef]
Miyazaki K, Togo S, Okamoto R, Idiris A, Kumagai H, Miyagi Y (2020). Collective cancer cell invasion in contact with fibroblasts through integrin-α5β1/fibronectin interaction in collagen matrix. Cancer Science 111: 4381–4392. https://doi.org/10.1111/cas.14664 [Google Scholar] [PubMed] [CrossRef]
Musa M, Ali A (2020). Cancer-associated fibroblasts of colorectal cancer and their markers: Updates, challenges and translational outlook. Future Oncology 16: 2329–2344. https://doi.org/10.2217/fon-2020-0384 [Google Scholar] [PubMed] [CrossRef]
Nair N, Calle AS, Zahra MH, Prieto-Vila M, Oo AKK et al. (2017). Cancer stem cell model as the point of origin of cancer-associated fibroblasts in tumor microenvironment. Scientific Reports 7: 6838. https://doi.org/10.1038/s41598-017-07144-5 [Google Scholar] [PubMed] [CrossRef]
Navin N, Kendall J, Troge J, Andrews P, Rodgers L et al. (2011). Tumor evolution inferred by single-cell sequencing. Nature 472: 90–94. https://doi.org/10.1038/nature09807 [Google Scholar] [PubMed] [CrossRef]
Nieminen M, Henttinen T, Merinen M, Marttila-Ichihara F, Eriksson JE, Jalkanen S (2006). Vimentin function in lymphocyte adhesion and transcellular migration. Nature Cell Biology 8: 156–162. https://doi.org/10.1038/ncb1355 [Google Scholar] [PubMed] [CrossRef]
Nishishita R, Morohashi S, Seino H, Wu Y, Yoshizawa T, Haga T, Saito K, Hakamada K, Fukuda S, Kijima H (2018). Expression of cancer-associated fibroblast markers in advanced colorectal cancer. Oncology Letters 15: 6195–6202. https://doi.org/10.3892/ol.2018.8097 [Google Scholar] [PubMed] [CrossRef]
Northey JJ, Przybyla L, Weaver VM (2017). Tissue force programs cell fate and tumor aggression. Cancer Discovery 7: 1224–1237. https://doi.org/10.1158/2159-8290.CD-16-0733 [Google Scholar] [PubMed] [CrossRef]
Nurmik M, Ullmann P, Rodriguez F, Haan S, Letellier E (2020). Cancer-associated fibroblasts and their markers. International Journal of Cancer 146: 895–905. https://doi.org/10.1002/ijc.32193 [Google Scholar] [PubMed] [CrossRef]
Öhlund D, Handly-Santana A, Biffi G, Elyada E, Almeida AS et al. (2017). Distinct populations of inflammatory fibroblasts and myofibroblasts in pancreatic cancer. Journal of Experimental Medicine 214: 579–596. https://doi.org/10.1084/jem.20162024 [Google Scholar] [PubMed] [CrossRef]
Ortiz-Otero N, Clinch AB, Hope J, Wang W, Reinhart-King CA, King MR (2020a). Cancer associated fibroblasts confer shear resistance to circulating tumor cells during prostate cancer metastatic progression. Oncotarget 11: 1037–1050. https://doi.org/10.18632/oncotarget.27510 [Google Scholar] [PubMed] [CrossRef]
Ortiz-Otero N, Marshall JR, Lash B, King MR (2020b). Chemotherapy-induced release of circulating-tumor cells into the bloodstream in collective migration units with cancer-associated fibroblasts in metastatic cancer patients. BMC Cancer 20: 873. https://doi.org/10.1186/s12885-020-07376-1 [Google Scholar] [PubMed] [CrossRef]
Pantel K, Alix-Panabières C (2010). Circulating tumor cells in cancer patients: Challenges and perspectives. Trends in Molecular Medicine 16: 398–406. https://doi.org/10.1016/j.molmed.2010.07.001 [Google Scholar] [PubMed] [CrossRef]
Peinado H, Alečković M, Lavotshkin S, Matei I, Costa-Silva B et al. (2012). Melanoma exosomes educate bone marrow progenitor cells toward a pro-metastatic phenotype through MET. Nature Medicine 18: 883–891. https://doi.org/10.1038/nm.2753 [Google Scholar] [PubMed] [CrossRef]
Petrik J, Verbanac D, Fabijanec M, Hulina-Tomašković A, Čeri A, Somborac-Bačura A, Barišić K (2022). Circulating tumor cells in colorectal cancer: Detection systems and clinical utility. International Journal of Molecular Sciences 23: 13582. https://doi.org/10.3390/ijms232113582 [Google Scholar] [PubMed] [CrossRef]
Peña C, Céspedes MV, Lindh MB, Kiflemariam S, Mezheyeuski A, Edqvist PH, Ostman A (2013). STC1 expression by cancer-associated fibroblasts drives metastasis of colorectal cancer. Cancer Research 73: 1287–1297. https://doi.org/10.1158/0008-5472.CAN-12-1875 [Google Scholar] [PubMed] [CrossRef]
Qi J, Sun H, Zhang Y, Wang Z, Xun Z, Li Z, Su B (2022). Single-cell and spatial analysis reveal interaction of FAP+ fibroblasts and SPP1+ macrophages in colorectal cancer. Nature Communications 13: 1742. https://doi.org/10.1038/s41467-022-29366-6 [Google Scholar] [PubMed] [CrossRef]
Quail DF, Joyce JA (2013). Microenvironmental regulation of tumor progression and metastasis. Nature Medicine 19: 1423–1437. https://doi.org/10.1038/nm.3394 [Google Scholar] [PubMed] [CrossRef]
Ramalingam N, Jeffrey SS (2018). Future of liquid biopsies with growing technological and bioinformatics studies: Opportunities and challenges in discovering tumor heterogeneity with single-cell level analysis. The Cancer Journal 24: 104–108. https://doi.org/10.1097/PPO.0000000000000308 [Google Scholar] [PubMed] [CrossRef]
Rejniak KA (2016). Circulating tumor cells: When a solid tumor meets a fluid microenvironment. Advances in Experimental Medicine and Biology 936: 93–106. https://doi.org/10.1007/978-3-319-42023-3 [Google Scholar] [CrossRef]
Riihimäki M, Hemminki A, Sundquist J, Hemminki K (2016). Patterns of metastasis in colon and rectal cancer. Scientific Reports 6: 29765. https://doi.org/10.1038/srep29765 [Google Scholar] [PubMed] [CrossRef]
Riihimäki M, Thomsen H, Sundquist K, Hemminki K (2012). Colorectal cancer patients: What do they die of? Frontline Gastroenterology 3: 143–149. https://doi.org/10.1136/flgastro-2012-100141 [Google Scholar] [PubMed] [CrossRef]
Ringuette Goulet C, Bernard G, Tremblay S, Chabaud S, Bolduc S, Pouliot F (2018). Exosomes induce fibroblast differentiation into cancer-associated fibroblasts through TGFβ signaling. Molecular Cancer Research 16: 1196–1204. https://doi.org/10.1158/1541-7786.MCR-17-0784 [Google Scholar] [PubMed] [CrossRef]
Roulis M, Kaklamanos A, Schernthanner M, Bielecki P, Zhao J et al. (2020). Paracrine orchestration of intestinal tumorigenesis by a mesenchymal niche. Nature 580: 524–529. https://doi.org/10.1038/s41586-020-2166-3 [Google Scholar] [PubMed] [CrossRef]
Sahai E, Astsaturov I, Cukierman E, DeNardo DG, Egeblad M et al. (2020). A framework for advancing our understanding of cancer-associated fibroblasts. Nature Reviews Cancer 20: 174–186. https://doi.org/10.1038/s41568-019-0238-1 [Google Scholar] [PubMed] [CrossRef]
Schoppmann SF, Jesch B, Riegler MF, Maroske F, Schwameis K, Jomrich G, Birner P (2013). Podoplanin expressing cancer associated fibroblasts are associated with unfavourable prognosis in adenocarcinoma of the esophagus. Clinical Experimental Metastasis 30: 441–446. https://doi.org/10.1007/s10585-012-9549-2 [Google Scholar] [PubMed] [CrossRef]
Shaukat A, Levin TR (2022). Current and future colorectal cancer screening strategies. Nature Reviews Gastroenterology & Hepatology 19: 521–531. https://doi.org/10.1038/s41575-022-00612-y [Google Scholar] [PubMed] [CrossRef]
Shin N, Son GM, Shin DH, Kwon MS, Park BS, Kim HS, Kang CD (2019). Cancer-associated fibroblasts and desmoplastic reactions related to cancer invasiveness in patients with colorectal cancer. Annals of Coloproctology 35: 36–46. https://doi.org/10.3393/ac.2018.09.10 [Google Scholar] [PubMed] [CrossRef]
Siegel RL, Miller KD, Goding Sauer A, Fedewa SA, Butterly LF, Anderson JC, Jemal A (2020). Colorectal cancer statistics, 2020. CA: A Cancer Journal for Clinicians 70: 145–164. https://doi.org/10.3322/caac.21601 [Google Scholar] [PubMed] [CrossRef]
Simon T, Salhia B (2022). Cancer-associated fibroblast subpopulations with diverse and dynamic roles in the tumor microenvironment. Molecular Cancer Research 20: 183–192. https://doi.org/10.1158/1541-7786.MCR-21-0282 [Google Scholar] [PubMed] [CrossRef]
Suarez-Carmona M, Lesage J, Cataldo D, Gilles C (2017). EMT and inflammation: Inseparable actors of cancer progression. Molecular Oncology 11: 805–823. https://doi.org/10.1002/1878-0261.12095 [Google Scholar] [PubMed] [CrossRef]
Takatsuna M, Morohashi S, Yoshizawa T, Hirai H, Haga T et al. (2016). Myofibroblast distribution is associated with invasive growth types of colorectal cancer. Oncology Reports 36: 3154–3160. https://doi.org/10.3892/or.2016.5202 [Google Scholar] [PubMed] [CrossRef]
Tauriello DVF, Palomo-Ponce S, Stork D, Berenguer-Llergo A, Badia-Ramentol J et al. (2018). TGFβ drives immune evasion in genetically reconstituted colon cancer metastasis. Nature 554: 538–543. https://doi.org/10.1038/nature25492 [Google Scholar] [PubMed] [CrossRef]
Tian Y, Kharazmi E, Sundquist K, Sundquist J, Brenner H, Fallah M (2019). Familial colorectal cancer risk in half siblings and siblings: Nationwide cohort study. British Medical Journal (Clinical Research Edition) 364: l803. https://doi.org/10.1136/bmj.l803 [Google Scholar] [PubMed] [CrossRef]
Togo S, Polanska UM, Horimoto Y, Orimo A (2013). Carcinoma-associated fibroblasts are a promising therapeutic target. Cancers 5: 149–169. https://doi.org/10.3390/cancers5010149 [Google Scholar] [PubMed] [CrossRef]
van der Stok EP, Spaander MCW, Grünhagen DJ, Verhoef C, Kuipers EJ (2017). Surveillance after curative treatment for colorectal cancer. Nature Reviews Clinical Oncology 14: 297–315. https://doi.org/10.1038/nrclinonc.2016.199 [Google Scholar] [PubMed] [CrossRef]
Varga J, Nicolas A, Petrocelli V, Pesic M, Mahmoud A et al. (2020). AKT-dependent NOTCH3 activation drives tumor progression in a model of mesenchymal colorectal cancer. Journal of Experimental Medicine 217: e20191515. https://doi.org/10.1084/jem.20191515 [Google Scholar] [PubMed] [CrossRef]
Wan JCM, Massie C, Garcia-Corbacho J, Mouliere F, Brenton JD, Caldas C, Pacey S, Baird R, Rosenfeld N (2017). Liquid biopsies come of age: Towards implementation of circulating tumor DNA. Nature Reviews Cancer 17: 223–238. https://doi.org/10.1038/nrc.2017.7 [Google Scholar] [PubMed] [CrossRef]
Wang Y, Chen Z, Zhao G, Li Q (2023). Cancer-associated fibroblast risk model for prediction of colorectal carcinoma prognosis and therapeutic responses. Mediators of Inflammation 2023: 3781091. https://doi.org/10.1155/2023/3781091 [Google Scholar] [PubMed] [CrossRef]
Wang W, Cheng B, Yu Q (2022). Cancer-associated fibroblasts as accomplices to confer therapeutic resistance in cancer. Cancer Drug Resistance 5: 889–901. https://doi.org/10.20517/cdr.2022.67 [Google Scholar] [PubMed] [CrossRef]
Wu F, Yang J, Liu J, Wang Y, Mu J, Zeng Q, Zhou H (2021). Signaling pathways in cancer-associated fibroblasts and targeted therapy for cancer. Signal Transduction Target Therapy 6: 218. https://doi.org/10.1038/s41392-021-00641-0 [Google Scholar] [PubMed] [CrossRef]
Xi Y, Xu P (2021). Global colorectal cancer burden in 2020 and projections to 2040. Translational Oncology 14: 101174. https://doi.org/10.1016/j.tranon.2021.101174 [Google Scholar] [PubMed] [CrossRef]
Yang Y, Gu J, Li X, Xue C, Ba L, Gao Y, Zhou J, Bai C, Sun Z, Zhao R C (2021). HIF-1α promotes the migration and invasion of cancer-associated fibroblasts by miR-210. Aging and Disease 12: 1794–1807. https://doi.org/10.14336/AD.2021.0315 [Google Scholar] [PubMed] [CrossRef]
Yap TA, Lorente D, Omlin A, Olmos D, de Bono JS (2014). Circulating tumor cells: A multifunctional biomarker. Clinical Cancer Research 20: 2553–2568. https://doi.org/10.1158/1078-0432.CCR-13-2664 [Google Scholar] [PubMed] [CrossRef]
Yu M, Bardia A, Wittner BS, Stott SL, Smas ME et al. (2013). Circulating breast tumor cells exhibit dynamic changes in epithelial and mesenchymal composition. Science 339: 580–584. https://doi.org/10.1126/science.1228522 [Google Scholar] [PubMed] [CrossRef]
Yu M, Guo G, Huang L, Deng L, Chang CS, Achyut BR, Cui Y (2020). CD73 on cancer-associated fibroblasts enhanced by the A2B-mediated feedforward circuit enforces an immune checkpoint. Nature Communications 11: 515. https://doi.org/10.1038/s41467-019-14060-x [Google Scholar] [PubMed] [CrossRef]
Zhang L, Meng Y, Feng X, Han Z (2022). CAR-NK cells for cancer immunotherapy: From bench to bedside. Biomarker Research 10: 12. https://doi.org/10.1186/s40364-022-00364-6 [Google Scholar] [PubMed] [CrossRef]
Zhao J, Chen Y (2022). Systematic identification of cancer-associated-fibroblast-derived genes in patients with colorectal cancer based on single-cell sequencing and transcriptomics. Frontiers in Immunology 13: 988246. https://doi.org/10.3389/fimmu.2022.988246 [Google Scholar] [PubMed] [CrossRef]
Zhao Z, Li W, Zhu L, Xu B, Jiang Y, Ma N, Zhang M (2022). Construction and verification of a fibroblast-related prognostic signature model for colon cancer. Frontiers in Genetics 13: 908957. https://doi.org/10.3389/fgene.2022.908957 [Google Scholar] [PubMed] [CrossRef]
Zhao M, Liang F, Zhang B, Yan W, Zhang J (2015). The impact of osteopontin on prognosis and clinicopathology of colorectal cancer patients: A systematic meta-analysis. Scientific Report 5: 12713. https://doi.org/10.1038/srep12713 [Google Scholar] [PubMed] [CrossRef]
Zheng S, Lin H, Liu JQ, Balic M, Datar R, Cote RJ, Tai YC (2007). Membrane microfilter device for selective capture, electrolysis and genomic analysis of human circulating tumor cells. Journal of Chromatography A 1162: 154–161. https://doi.org/10.1016/j.chroma.2007.05.064 [Google Scholar] [PubMed] [CrossRef]
Zhong ME, Huang ZP, Wang X, Cai D, Li CH, Gao F, Wang W (2022). A transcription factor signature can identify the CMS4 subtype and stratify the prognostic risk of colorectal cancer. Frontiers in Oncology 12: 902974. https://doi.org/10.3389/fonc.2022.902974 [Google Scholar] [PubMed] [CrossRef]
Zhou Y, Bian S, Zhou X, Cui Y, Wang W, Wen L, Guo L, Fu W, Tang F (2020). Single-cell multiomics sequencing reveals prevalent genomic alterations in tumor stromal cells of human colorectal cancer. Cancer Cell 38: 818–828.e5. https://doi.org/10.1016/j.ccell.2020.09.015 [Google Scholar] [PubMed] [CrossRef]
Cite This Article
 Copyright © 2023 The Author(s). Published by Tech Science Press.
Copyright © 2023 The Author(s). Published by Tech Science Press.This work is licensed under a Creative Commons Attribution 4.0 International License , which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.


 Submit a Paper
Submit a Paper Propose a Special lssue
Propose a Special lssue View Full Text
View Full Text Download PDF
Download PDF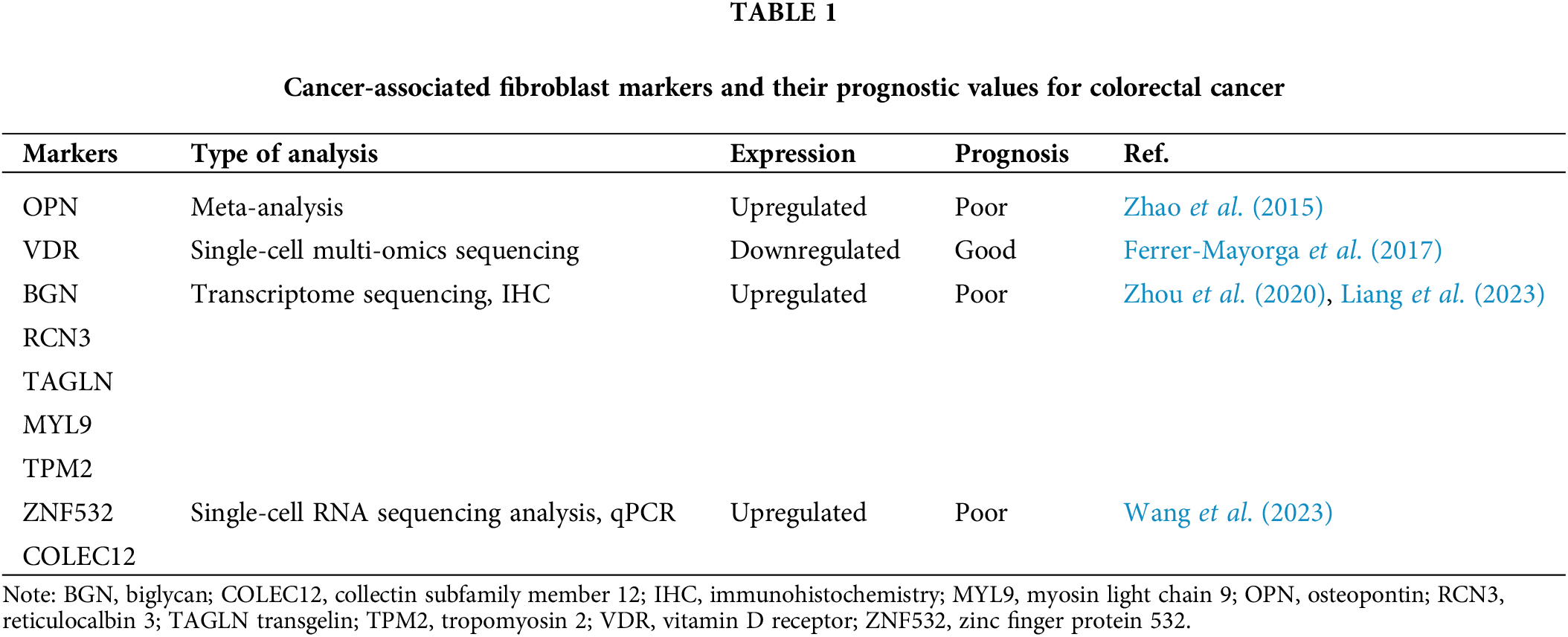
 Downloads
Downloads
 Citation Tools
Citation Tools